Abstract
TIMAP (TGFβ-inhibited membrane-associated protein) is an endothelium-enriched TGFβ downstream protein and structurally belongs to the targeting subunit of myosin phosphatase; however, the mechanism of TGFβ repressing TIMAP and its functional relevance to TGFβ bioactivity remain largely unknown. Here, we report that TIMAP is reduced in TGFβ-elevated mouse fibrotic kidney and highly expressed in macrophages. TGFβ repression of TIMAP is associated with HDAC3 upregulation and its recruitment by Smad2/3 at the Smad binding element on TIMAP promoter, whereas specific HDAC3 inhibition reversed the TIMAP repression, suggesting that TGFβ transcriptionally downregulates TIMAP through HDAC3-associated Smad signaling. Further investigation showed that TIMAP over-expression interrupted TGFβ-associated Smad signaling and TIMAP repression by TGFβ correlated with TGFβ-induced macrophage M2 polarization markers, migration, and phagocytosis—the processes promoted by phosphorylation of the putative TIMAP substrate myosin light chain (MLC). Consistently, TIMAP dephosphorylated MLC in macrophages and TGFβ induced macrophage migration and phagocytosis in TIMAP- and MLC phosphorylation-dependent manners, suggesting that TIMAP dephosphorylation of MLC constitutes an essential regulatory loop mitigating TGFβ-associated macrophage M2 phenotypic activities. Given that hyperactive TGFβ often causes excessive macrophage phagocytic activities potentially leading to various chronic disorders, the strategies targeting HDAC3/TIMAP axis might improve TGFβ-associated pathological processes.
Key message
-
TIMAP is enriched in the endothelium and highly expressed in macrophages.
-
TIMAP is suppressed by TGFβ via HDAC3-associated Smad signaling.
-
TIMAP inhibits TGFβ signaling and TGFβ-associated macrophage M2 polarization.
-
TIMAP dephosphorylation of MLC counteracts TGFβ-induced macrophage phagocytosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
TIMAP (TGFβ-inhibited membrane-associated protein) is originally identified from glomerular endothelial cells as a CAAX box protein, and its expression is downregulated by TGFβ [1], an important cytokine whose deregulation is involved in many chronic clinic diseases such as chronic inflammation, tissue fibrosis, and cancer [2]. TIMAP structurally belongs to the myosin phosphatase targeting (MYPT) protein family [3] and is thought to play a regulatory role in TGFβ-mediated cellular processes. Past studies have identified several plasma membrane proteins—the non-integrin laminin receptor 1 (LAMR1) [4, 5], Moesin [6], and receptor for activated C kinase 1 (RACK1) [7] as the TIMAP interacting partners or substrates. These studies indicate that TIMAP is enriched in endothelial filopodia [5], promotes angiogenesis of glomerular endothelial cells [8], and functions as a protector in LPS-induced lung endothelial barrier leakage [6]; however, the information about TGFβ-regulated cellular process involving TIMAP is still lacking.
TGFβ induces various cellular responses mainly through Smad signaling pathway. Upon activation, regulatory Smad2 and 3 (R-Smads) in complex with common Smad4 translocate to the nucleus and positively or negatively transcribe TGFβ downstream genes through Smad binding element (SBE) located on the promoter [9]. R-Smads can recruit transcription co-activator CBP (CREB-binding protein), p300, or P/CAF (p300/CBP-associated factor), which contain intrinsic histone acetyl-transferase, to promote transactivation [10]. Alternatively R-Smads can recruit transcription co-repressors such as Ski, NCoR/SMRT, or mSin3A, in association with histone deacetylase (HDAC), to repress the transcription [11, 12]. TGFβ repression of TIMAP is sensitive to HDAC inhibition [1], suggesting the potential involvement of Smad/HDAC signaling.
From a developmental point of view, endothelia and hematopoietic cells share a common precursor—hemangioblast [13]. Our initial study showed that TIMAP is also expressed in hematopoietic cell line KG1a (human myelogenous leukemia cell), Molt4 (human lymphoblastic T cell), and Dami (human megakaryocytic cell) [1]. Gene expression analysis from Gene Atlas database (www.biogps.org) reveals that the highest TIMAP level is found in mouse bone marrow-derived macrophages, which are the professional phagocytic cells regulated by TGFβ. Macrophage can polarize to pro-inflammatory M1 phenotype, or alternatively to M2 phenotype influenced by TGFβ or other cytokines, in which macrophage migrates to the injured site to clean harmful cell debris through phagocytosis and repair injury by depositing extracellular matrix proteins [14]. Macrophage migration and phagocytosis involve active cytoskeleton reorganization and cell membrane movement mediated by myosin-involved actin polymerization [15, 16], which is regulated by the phosphorylation of myosin light chain (MLC) [17, 18]. Recent in vitro study showed that TIMAP regulated PP1C activity towards MLC phosphorylation [19], implying that TIMAP might affect TGFβ-regulated cell migration and phagocytosis through regulating MLC phosphorylation-associated cell cytoskeleton rearrangements.
In this study, we sought to investigate the mechanism of TGFβ repressing TIMAP and its functional significance in TGFβ-associated cellular processes. We showed for the first time that TIMAP is reduced in TGFβ-elevated fibrotic mouse kidney and highly expressed in macrophages. We investigated how TGFβ HDAC-sensitively repressed TIMAP transcription and further identified HDAC3 as the subtype involved. We then tested if TIMAP affected TGFβ-induced macrophage M2 polarization and determined whether TIMAP regulated MLC phosphorylation and its role in TGFβ-associated macrophage migration and phagocytosis. These results reveal a novel regulatory pathway and shed new lights on TGFβ-associated macrophage functions potentially related to chronic disorders.
Materials and methods
Animal work
The use of animals is in accordance with the university guidelines and regulations and the animal protocols were approved by the Nanjing University Animal Welfare and Ethics committee. C57BL6/J male mice aged at 6–8 weeks were from the University Model Animal Research Center. Mouse UUO (unilateral ureteral occlusion) model was established as previously [20]. Sham- and UUO-operated mice were composed of six animals in each group. After 7 days of experiments, all mice were euthanized by CO2 inhalation and mouse kidneys were removed surgically and stored at −80 °C for further examination.
Histological and immunofluorescent staining
Kidney tissues were prepared as before [20]. Sections (2 μm) were cut and subjected to Masson’s trichrome staining by standard procedure. Renal fibrosis areas (the ratio of blue-colored area of collagen deposition over total renal area) were measured quantitatively from 10 randomly selected fields from each kidney section and analyzed by the ImageJ software.
Mouse kidney sections or RAW cells seeded on cover-slips were incubated with anti-TIMAP or anti-phosphorylated MLC (P-MLC) antibody (Santa Cruz, USA) overnight at 4 °C, followed by incubating with the Alexa Fluor 594-conjugated AffiniPure donkey anti-goat or anti-rabbit secondary antibody (Jackson Immuno Research Lab., USA) for additional 1 h at room temperature. Cultured cells were counter-stained with DAPI (4′,6-diamidino-2-phenylindole) for nucleus verification. Images were taken by a light or a fluorescence confocal microscope. The levels of fluorescent staining of kidney sections were measured quantitatively from 10 randomly selected fields from each kidney section. The expression levels of TIMAP and P-MLC from immunofluorescence-stained RAW cells were measured quantitatively from 50 cells each time from at least four randomly selected fields from three repeated experiments and analyzed by the ImageJ software.
Cell and cell culture
Human embryonic kidney HEK293 and mouse macrophage RAW cells (ATCC, USA) were cultured in MEM medium with 10 % fetal bovine serum or DMEM containing 10 % newborn calf serum, respectively. Mouse peritoneal macrophages were collected and cultured basically as before [21]. All cells were cultured at 37 °C in a 5 % CO2 incubator. TGFβ or lipopolysaccharide (LPS) were added 1 h after TSA (Sigma Aldrich, USA), SAHA (suberoylanilide hydroxamic acid), or RGFP966 (Selleck, USA).
Western blotting
Western blotting was performed essentially as before [20] with antibodies to HDACs, phosphorylated Smad3 and Smad3 (Cell Signaling Technology, USA), TIMAP, MLC, P-MLC, Smad2/3, PAI-1 (Santa Cruz, USA), fibronectin (Protein Technology, China), and goat anti-mouse IgG-HRP, goat anti-rabbit IgG-HRP, or rabbit anti-goat IgG-HRP secondary antibodies (Yifeixue Biotech, China).
RT-PCR and quantitative real-time PCR (qRT-PCR)
Total RNA was extracted using TRIzol reagent (Invitrogen, USA) according to the manufacturer’s instructions. RT-PCR and qRT-PCR were carried out with a Superscript Kit and RT-PCR/qRT-PCR mix reagents (Vazyme, China). The primer sequences were listed in Table 1. RT-PCR products were analyzed by agarose gel electrophoresis.
Real-time PCR was performed in 20 μl SYBR Green PCR Master Mix (Vazyme, China). The primers for TGFβ (TGFβ-F/TGFβ-R), Arg-1 (Arg-1-F/Arg-1-R), IL-10 (IL-10-F/IL-10-R), or internal control GAPDH (GADPH-F/GADPH-R) were listed in Table 1. The original Ct (cycle of the threshold) values were adjusted to GAPDH. Data were converted and presented as the fold changes related to control.
Plasmid construction and luciferase reporter assay
Mouse TIMAP (mPPP1R16B) encodes a 568 amino acid protein, same as bovine TIMAP [1]. HA-tagged mouse wild-type and mutant TIMAP expression plasmids (HA-TIMAP/HA-TIMAPC565S) were constructed in pUSE-HA vector [22] by RT-PCR from RAW cell mRNA using primer pTIMAP-F, pTIMAP-R, or pmTIMAP-R (see Table 1). The mutated TIMAP has a serine substitution for cysteine in CAAX box [1]. Mouse TIMAP promoter (pTIMAP-luc, -5000 to +96, related to the transcription starting site), and the 5′- or 3′- deletion (pTimap5d-luc, -2474 to +96 and pTimap3d-luc, -5000 to -2453) reporter plasmids were constructed in pGL3 basic vector at the XhoI/HindIII sites. The primer sequences for full-length (pFull-F/R) or 5′/3′ deletions (p5d-F/R) were listed in Table 1. Two TIMAP-shRNA plasmids were constructed in GV102 vector at the BamHI and HindIII sites with the following oligos: GATCCCGTTCGCTACTTCCTGAAGATTCAAGAGATCTTCAGGAAG TAGCGAACTTTTTTGGAAA and AGCTTTTCCAAAAAAGTTCGCTACTTCCTGAAGAT CTCTTGAATCTTCAGGAAGTAGCGAACGG (shRNA-TIMAP1, the target sequences corresponding to TIMAP open reading frame 250-268 were underlined) and GATC CCGCCTATACCGGAAAGAATATTTCAAGAGAATATTCTTTCCGGTATAGGTTTTTT GGAAA and AGCTTTTCCAA AAAACCTATACCGGAAAGAATATTCTCTTGAAATAT TCTTTCCGGTATAGGCGG (shRNA-TIMAP2, the underlined sequences targeted 1068-1086 of TIMAP). A plasmid containing a scrambled sequence was used as a control (shRNA control).
For luciferase assay, HEK293 cells were transfected with reporter plasmids plus a renilla luciferase plasmid by Lipofectamine 2000 reagent (Invitrogen, USA). The cells were then treated with TGFβ and/or TSA for 24 h before the luciferase activity was assessed using a dual luciferase assay system (Promega, USA).
Chromatin immunoprecipitation (ChIP) assay
ChIP assay was carried out with a ChIP Kit from Millipore (USA) following manufacturer’s protocol. The immunoprecipitation was performed with anti-Smad2/3, anti-HDAC3, anti-acetylated histone 3, or an isoform-matched immunoglobulin, respectively. After precipitation, the antibody-associated DNA fragments were PCR-amplified using primer sets for mouse SBEI (ChIP-mSBEI F/R for -4470/-4160 region), SBEII (ChIP-mSBEII F/R for -3982/-3511 region), the negative control (mChIP-negF/R, -3532/-3072), or for SBE on human TIMAP promoter (hChIP-SBE F/R, -384/-919). The primer sequences were listed in Table 1. PCR products were analyzed by agarose gel electrophoresis.
Co-immunoprecipitation (Co-IP) assay
Co-immunoprecipitation assay was performed reciprocally with antibodies specific for TIMAP or MLC as before [23]. The immunoprecipitants were analyzed by Western blotting using antibodies to MLC or TIMAP respectively.
Macrophage migration assay
RAW cells stably expressing either wild-type or mutant TIMAP (C565S) were established as before [22]. The confluent cell monolayer was stroked with a pipette tip. Cells were washed to remove detached and damaged cells and then treated with TGFβ and/or ML-7 (Santa Cruz, USA) for 24 h. The cell migrations were monitored microscopically and the migration distance was measured from five preset positions for each treatment condition by the ImageJ software.
Endocytosis assay
Neutral red endocytosis: RAW cells were treated with either TGFβ or RGFP966 for 24 h before neutral red reagent (100 μl of 0.1 % solution) was added. The endocytosis underwent at 37 °C for 2 h. Cells were then lysed and the sample absorbance was measured at 540 nm on a micro-plate reader [24].
Phagocytosis assay: RAW cells, RAW cells stably expressing either wild-type or mutant TIMAP (C565S) or transfected with shRNA plasmids were treated with M-CSF (Sino Biological, China) to potentiate M2 polarization [25] before TGFβ, RGFP966, and/or ML-7 treatment. Zymosan particles (Sigma Aldrich, USA) were added to the cells at a ratio of 10:1 (zymosan to macrophage) [18, 26]. Phagocytosis was allowed to proceed for 2 h at 37 °C and then terminated by washing off non-ingested particles. Cells were stained by Wright staining and examined and photographed under a light microscopy at ×600 magnifications. A total of 100 cells from randomly selected fields for each condition were examined and phagocytosis index was calculated as the product of percentage of phagocytic cells multiplied by the mean of particles per cell [27].
Statistics
Data were expressed as the mean ± SD (standard deviation). The statistical significance of difference between two groups was analyzed by unpaired Student’s t test. The difference is considered statistically significant or very significant at p < 0.05 or p < 0.01, respectively.
Results
TGFβ downregulates TIMAP in mouse macrophages
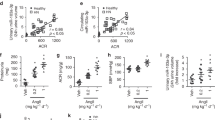
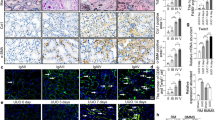
Our previous study indicated that TGFβ strongly inhibited TIMAP expression in cultured endothelial cells [1]. To gain insights into the possibility that TGFβ decreases TIMAP in vivo, we examined TIMAP expression from healthy or UUO mouse kidney in which TGFβ was a crucial mediator of disease progression [28]. We found that UUO mouse kidney displayed severe tubule dystrophy and increased interstitial fibrosis represented by blue-stained collagen depositions (Fig. 1a, upper panel and Fig. 1b, left panel). TIMAP was preferentially expressed in glomerulus in control kidney, but the expression was significantly reduced in UUO kidney (Fig. 1a, right-pointed arrows in lower panel and 1b, right). In addition, UUO kidney displayed sporadic, although dimmed, positive staining outside glomerulus where macrophages often infiltrated to (Fig. 1a, lower panel right, up-headed arrows), accompanied by increased TGFβ mRNA levels (Fig. 1c). To directly test whether macrophage expresses TIMAP and whether TGFβ represses its level, we treated mouse macrophage RAW cells with TGFβ and found that TGFβ caused remarkable TIMAP suppression (Fig. 1d, e). TGFβ also time-dependently reduced TIMAP mRNA in RAW cells and mouse peritoneal macrophages (Fig. 1f). Interestingly, the general HDAC inhibitor TSA (trichostatin A) and SAHA (suberoylanilide hydroxamic acid) prevented TGFβ inhibition of TIMAP at both mRNA and protein levels in RAW and peritoneal macrophages, respectively (Fig. 1g, h). Taken together, these results suggest that TIMAP levels inversely correlate with renal fibrosis severity in UUO kidney and TGFβ HDAC-dependently inhibits TIMAP transcription.
TGFβ downregulates TIMAP in mouse macrophages. a Kidney expression of TIMAP. Representative kidney sections from sham and UUO mice were stained by Masson trichrome staining (upper panel) or by immunofluorescent staining with anti-TIMAP antibody. The arrow heads indicated the positive TIMAP staining. b Quantitations of kidney fibrosis lesions (ratio of blue color area over total area) and TIMAP staining intensities from sham and UUO kidneys. c Kidney expression of TGFβ mRNA. Total RNAs were isolated from sham and UUO mouse kidneys (n = 6 in each group) and analyzed by qRT-PCR. The original Ct (cycle of the threshold) values were adjusted to GAPDH. Data were converted and presented as the fold changes related to sham control. d RAW cells were treated with TGFβ (5 ng/ml) for 24 h. TIMAP proteins were examined by immunofluorescent staining using a specific anti-TIMAP antibody (left panel). Cells were counter-stained with DAPI (middle panel) and merged (right panel). e Quantification of TIMAP staining intensities in d. f TGFβ inhibits TIMAP mRNA. RAW cells and mouse peritoneal macrophages (PM) were treated with TGFβ (5 ng/ml) for 6 and 12 h. TIMAP mRNA levels were analyzed by RT-PCR. GAPDH was used as internal control. Quantifications were underneath the figures. g RAW cells and h peritoneal macrophages (PM) were treated with TSA (30 ng/ml) or SAHA (1 μM) in the presence or absence of TGFβ (5 ng/ml) for 12 h. TIMAP mRNA or protein levels were analyzed by RT-PCR or Western blotting (WB), respectively. TIMAP mRNA or protein levels were normalized to GAPDH or β-actin and quantifications were listed under the figures. All the cell-based experiments were repeated at least three times and the representative results are shown. *p < 0.05 versus control or sham, # p < 0.05 versus TGFβ treatment
TGFβ represses TIMAP transcription through Smad signaling
To gain further insights into the TGFβ transcriptional regulation of TIMAP, we constructed a mouse TIMAP promoter (5 kbp) luciferase reporter, which contained two putative Smad binding sites located at -4347/-4340 and -3817/-3812 related to the transcription starting site, and two additional deletion mutants that covered 5′ or 3′ half of the 5 K promoter (Fig. 2a). We transfected the plasmids into HEK293 cells and found that TGFβ significantly inhibited the transcriptions of both full length (pTIMAP-luc) and 3′ deletion (pTIMAP3d-luc) plasmids, while only marginally affecting the 5′ deletion plasmid (pTIMAP5d-luc) (Fig. 2b). Since both full length and 3′ deletion plasmids share the same SBEs, we further tested the Smad2/3 binding of SBE on mouse and human TIMAP promoter by ChIP assay using a ChIP grade antibody that recognized both Smad2 and Smad3. The result showed that TGFβ treatment caused significant increase of Smad2/3 accumulation at both SBEs in mouse RAW cells (Fig. 2c, d) and at the SBE site in human HEK293 cells (Fig. 2e, f), suggesting that TGFβ transcriptionally downregulated TIMAP through Smad signaling.
TGFβ represses TIMAP transcription through Smad signaling. a Schematic diagram of mouse TIMAP promoter reporter constructs. The locations of Smad binding element (SBE) were indicated (-4347/-4340, -3817/-3812). Three reporter plasmids pTIMAP-luc (-5000/+96), pTIMAP5d-luc (-2474/+96), and pTIMAP3d-luc (-5000/-2453) were depicted. b HEK293 cells were transfected with control PGL3-basic or TIMAP promoter reporter plasmids, respectively. After 12 h, the cells were treated with 5 ng/ml of TGFβ for 24 h and then analyzed for luciferase activity. The luciferase activities are normalized by dividing the reporter firefly luciferase activities by the renilla luciferase activities and represented as fold change. c, e ChIP analysis of Smad2/3 interaction with SBEs on mouse (c) or human (e) TIMAP promoter. RAW (c) or HEK293 cells (e) were treated with TGFβ (5 ng/ml) for 3 h and then subjected to ChIP assay using an anti-Smad2/3 antibody or an isoform-matched IgG antibody. Immunoprecipitated DNA fragments were analyzed by PCR using specific primer sets for mouse SBEI (-4160 to -4470), SBEII (-3982 to -3511), or negative control SBEn (-3532 to -3072) that contained no SBE, or SBE (-384 to -919) on human TIMAP promoter. d, f Quantifications of Smad2/3 DNA binding from ChIP assay of RAW (d) or HEK293 cells (f). The bindings were normalized by dividing Smad2/3 bindings by the input PCR signaling. All the experiments were repeated at least three times and the representative results are shown. *p < 0.05 versus control. # p < 0.05 versus TGFβ treatment
TGFβ downregulation of TIMAP involves HDAC3-associated transcription repression
TGFβ/Smad transcriptionally represses downstream gene in association with transcription repressors and HDACs [29]. In order to identify the HDAC subtype involved in TIMAP repression, we tested the expression of a panel of HDACs that were reported to associate with Smad signaling. We found that the expression of HDAC1, 2, 4, 6, and 8 remained constant, but HDAC3 was notably upregulated by TGFβ in RAW cells (Fig. 3a) and in UUO kidney (Fig. 3c). Further, ChIP assay detected that TGFβ treatment caused a significant HDAC3 association with the SBE sites on both mouse and human TIMAP promoters (Fig. 3d, f). Accordingly, the local histone3 acetylation was decreased (Fig. 3d, lower panel), presumably due to the increased local HDAC3 activity. To prove HDAC3 is responsible for TGFβ repression of TIMAP, we tested the effect of a specific HDAC3 inhibitor RGFP966 on the repression. The results showed that TGFβ inhibition of TIMAP mRNA and protein was almost completely restored by RGFP966 (Fig. 3h, i), suggesting that HDAC3 plays an essential role in TGFβ transcriptional repression of TIMAP.
TGFβ downregulation of TIMAP involves HDAC3-associated transcriptional repression. a HDAC subtype expression in RAW cells or c in UUO mouse kidney. RAW cells were treated with 1 or 5 ng/ml of TGFβ for 24 h. The kidney tissue lysates were prepared from control or UUO mice (7 days). The lysates were then analyzed for HDAC subtype expression by Western blotting. b Quantification of HDAC3 expression in a. The expression levels were normalized to β-actin. d, f ChIP analysis of HDAC3 or acetylated histone3 binding to SBE on mouse or human TIMAP promoter. RAW (d) or HEK293 cells (f) were treated with TGFβ (5 ng/ml) for 3 h and then subjected to ChIP assay using either an anti-HDAC3 or an anti-acetylated histone3 antibody. Immunoprecipitated DNA fragments were analyzed by PCR using specific primer sets for mouse SBEI (-4160 to -4470), SBEII (-3982 to -3511), negative control SBEn (-3532 to -3072) (d), or SBE (-384 to -919) on human TIMAP promoter (f). e, g Quantification analyses for (d) or (f), respectively. h HDAC3 inhibition relieves TGFβ repression of TIMAP. RAW cells were treated with RGFP966 (10 μM) and/or TGFβ (5 ng/ml) for 24 h. TIMAP mRNA or protein levels were assayed by RT-PCR or Western blotting respectively. i Quantification of TIMAP mRNA and protein levels in h. The TIMAP levels in RT-PCR or Western blotting were normalized to GAPDH or β-actin, respectively. All the experiments were repeated at least three times and representative results are shown. *p < 0.05 versus control, # p < 0.05 versus TGFβ treatment
TIMAP affects TGFβ regulation of M2 markers of macrophage polarization
TGFβ is a potent inducer of macrophage M2c polarization [30]. RAW cells are able to express M1 biomarker NOS2 (nitric oxide synthase 2) and IL-6 stimulated by LPS or M2 marker Arginase-1 (Arg-1) and IL-10 in response to TGFβ, respectively (Fig. 4a, b, left panel). TSA or RGFP966 treatment inhibited TGFβ-induced expression of Arg-1 and IL-10 (Fig. 4b, middle and right panel and Fig. 4c). To further determine whether TIMAP causatively affects TGFβ-induced macrophage M2 polarization, we employed TIMAP gain or loss strategies by TIMAP over-expression (HA-TIMAP) or small hairpin RNA (shRNA-TIMAP) knockdown technique, respectively. To ensure the result accuracy, we performed the assay by qRT-PCR. The results showed that TGFβ induction of both Arg-1 and IL-10 were enhanced when TIMAP was knocked down by two specific shRNAs that target different areas on TIMAP mRNA (Fig. 4d). On the contrary, TIMAP over-expression markedly reduced the TGFβ induction of Arg-1 and IL-10 (Fig. 4e). TIMAP over-expression also repressed TGFβ-mediated Smad3 phosphorylation, PAI-1 (plasminogen activator inhibitor -1) and fibronectin (Fig. 4f, g) expression, the major TGFβ signaling molecule and downstream proteins highly expressed in M2 phase. Taken together, these results suggest that TIMAP negatively affects TGFβ signaling and its regulation of macrophage M2 polarization.
TIMAP affects TGFβ regulation of M2 markers of macrophage polarization. a RAW cells were treated with 100 or 200 ng/ml of LPS for 24 h. NOS2 and IL-6 mRNAs were assayed by RT-PCR. b RAW cells were treated with 2 or 5 ng/ml of TGFβ for 24 h. (left panel), or with TGFβ (5 ng/ml) and/or TSA (30 ng/ml) for 24 h (middle panel), or with RGFP966 (10 μM) and/or TGFβ (5 ng/ml) for 24 h (right panel). The mRNA levels of Arg-1, IL-10, and TIMAP were assayed by RT-PCR. c Quantifications of b. d Quantitative real-time PCR (qRT-PCR) analysis of M2 markers. RAW cells were transfected with a shRNA control (containing a scrambled sequence) or two shRNA-TIMAP plasmids (shRNA-TIMAP1 and shRNA-TIMAP2) and then treated with or without TGFβ (5 ng/ml) for 24 h. Arg-1 and IL-10 mRNA levels were determined by qRT-PCR. TIMAP protein expressions were verified by Western blotting (bottom). e RAW cells were transfected with HA vector (HA-control) or HA-tagged TIMAP (HA-TIMAP) expression plasmid for 24 h and then treated with or without TGFβ (5 ng/ml) for 24 h. Arg-1 and IL-10 mRNA levels were determined by qRT-PCR. HA-TIMAP protein expressions were verified by Western blotting (bottom). f The cell lysates treated as in e for different times were analyzed for phosphorylated Smad3, total Smad3 (30 min), TGFβ downstream protein plasminogen activator inhibitor-1 (PAI-1), and fibronectin (FN) expression (24 h) by Western blotting. g Quantifications of f. The expression levels of RT-PCR or Western blotting were normalized to GAPDH or β-actin, respectively, and presented as the fold changes related to controls. All the experiments were repeated at least three times and representative results are shown. *p < 0.05 and **p < 0.01 versus control, # p < 0.05 and ## p < 0.01 versus TGFβ treatment
TIMAP regulates TGFβ-induced macrophage M2 phenotypic migration and phagocytosis
Promotion of macrophage migration to the injured site and removal of dead cell debris are characteristic properties of macrophage M2 phenotypes incurred by TGFβ during tissue remodeling [31]. We next investigated whether TGFβ repression of TIMAP affected TGFβ-induced macrophage migration and phagocytosis. The results showed that TGFβ treatment led to increased RAW migration, which was inhibited by RGFP966 (Fig. 5a, b). Macrophage phagocytosis refers to receptor-mediated internalization of particles ≥0.5 μm in diameter [32], while fluid-phase and soluble materials are taken up by a process called macropinocytosis [33]. We found that TGFβ promoted the macropinocytosis of neutral red (Fig. 5c) [24] and phagocytosis of zymosan particles (average 3 μm in diameter, Fig. 5d), which were significantly inhibited by TSA or RGFP966. To further verify these observations, we assayed the effects of TIMAP over-expression on TGFβ-induced macrophage phagocytosis. To avoid the interference of excess protein on cell motility, we used a mutant TIMAP expression plasmid as control, in which the cysteine (C565) in CAAX box was substituted with serine (S), hence prohibiting its functional membrane localization [1]. The results showed that over-expression of wild-type TIMAP prevented basal and TGFβ-induced macrophage phagocytosis (Fig. 5e). On the marked contrary, TIMAP knockdown by shRNA enhanced the phagocytosis (Fig. 5f). Taken together, these results indicate that TIMAP plays an important role in TGFβ-induced macrophage migration and phagocytosis.
TIMAP regulates TGFβ-induced macrophage M2 phenotypic migration and phagocytosis. a Cell migration assay. RAW cells were treated with RGFP966 (10 μM) in the presence or absence of TGFβ (5 ng/ml). The cell monolayer was wounded by scraping with a pipette tip and the cell migrations were monitored at time zero (0 h) and 24 h. b Cell migration distances were measured and analyzed by the ImageJ software. The number represented the average of 5 measurements from 5 preset locations for each field. c Neutral red endocytosis assay. RAW cells were first stimulated with M-CSF (50 ng/ml) for 12 h and then treated with TGFβ (5 ng/ml) in the presence or absence of TSA (30 ng/ml) or RGFP966 (10 μM) for additional 24 h. Cells were then subjected to neutral red endocytosis assay. d RAW cells were stimulated with M-CSF (50 ng/ml) for 12 h and then treated with TGFβ (5 ng/ml) in the presence or absence of RGFP966 (10 μM) for 24 h before addition of zymosan particles. Phagocytosis was allowed to proceed for 2 h, then the photographs were taken at ×600 magnification. The quantification was underneath the figures. e RAW cells stably expressing either mutant or wild-type TIMAP or f transfected with shRNA control or shRNA-TIMAP plasmid were subjected to zymosan phagocytosis assay as in d. The expressions of exogenous HA-TIMAP or endogenous TIMAP were verified by Western blotting and the quantifications were underneath the figures, respectively. All the experiments were repeated at least three times and representative results are shown. *p < 0.05 versus control, # p < 0.05 versus TGFβ treatment
TIMAP regulates myosin light chain phosphorylation in macrophages
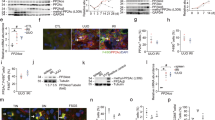
The phosphorylation of myosin light chain (MLC) is critical in controlling cell membrane movement during cell migration and phagocytosis [17]. Since MLC phosphorylation is regulated by TIMAP in vitro [19], we assume that TIMAP might mediate TGFβ-induced cell migration and phagocytosis through modulating MLC phosphorylation in macrophages. To test this idea, we first treated RAW cells with TGFβ and measured the phosphorylated MLC. TGFβ dose and time dependently increased MLC phosphorylation (Fig. 6a), while RGFP966 inhibited the increment (Fig. 6b). TIMAP over-expression blunted TGFβ-incurred MLC phosphorylation (Fig. 6c–e) and on the contrary, TIMAP knockdown by shRNA increased the basal MLC phosphorylation and sustained the level in presence of TGFβ (Fig. 6f). To further determine the relationship between TIMAP and MLC phosphorylation, we performed co-immunoprecipitation assay. We found that TIMAP was physically associated with MLC in resting RAW cells and TGFβ treatment reduced their associations (Fig. 6g, h) with a concomitant increase of MLC phosphorylation, suggesting that MLC is a TIMAP substrate in macrophages that potentially mediates TIMAP regulation of TGFβ-associated macrophage M2 phenotypes.
TIMAP mediates TGFβ regulation of myosin light chain phosphorylation in macrophages. a RAW cells were treated with 2 or 5 ng/ml of TGFβ for 24 h (upper panel) or with 5 ng/ml of TGFβ for 6 or 12 h (lower panel). The phosphorylated MLC (P-MLC) levels were assayed by Western blotting using a specific anti-phosphorylated MLC antibody. The changes of TIMAP expression were also monitored. b RAW cells were treated with specific HDAC3 inhibitor RGFP966 (10 μM) and/or TGFβ (5 ng/ml) for 24 h, and then P-MLC levels were assayed by Western blotting. The P-MLC levels were normalized to β-actin and the quantification was underneath the figure. c RAW cells were transfected with a vector control or an HA-tagged TIMAP expression plasmid and then treated with or without TGFβ (5 ng/ml) for 24 h. P-MLC levels were assayed by Western blotting. The levels were normalized to β-actin and the quantification was underneath the figure. d Immunofluorescent staining of P-MLC. RAW cells stably transcfected with control or HA-TIMAP plasmid were treated with or without TGFβ (5 ng/ml) for 24 h and then stained with anti-P-MLC antibody (upper panel), counter-stained by DAPI for nucleus DNA (middle panel) and merged (lower panel). e Quantification of d. f RAW cells were transfected with a shRNA control or shRNA-TIMAP plasmid and then treated with or without TGFβ (5 ng/ml) for 24 h. TIMAP and P-MLC levels were examined by Western blotting. P-MLC levels were normalized to β-actin and the quantification was underneath the figure. g Co-immunoprecipitation of TIMAP and MLC. RAW cells were untreated or treated with TGFβ (5 ng/ml) for 8 h. The cell lysates were reciprocally subjected to co-immunoprecipitation with anti-MLC, anti-TIMAP, or an isoform (IgG)-matched control antibody followed by Western blotting with TIMAP or MLC antibodies respectively (upper panel). The total cell lysates were also examined for P-MLC, TIMAP, or MLC expression by Western blotting (lower panel). h Quantification of TIMAP and MLC of co-IP assay in g. All the experiments were repeated at least three times and representative results are shown. *p < 0.05 versus control, # p < 0.05 versus TGFβ treatment
TIMAP regulation of MLC phosphorylation contributes to TGFβ-induced macrophage migration and phagocytosis
To confirm that TIMAP dephosphorylation of MLC contributes to TGFβ-mediated macrophage M2 phenotypic activities, we tested the effect of a specific MLC kinase inhibitor ML-7 on TGFβ-induced macrophage migration and phagocytosis. The results showed that ML-7 significantly inhibited the basal and TGFβ-induced macrophage migration, even when cells over-expressed wilt-type TIMAP (Fig. 7a, b). In addition, ML-7 also similarly inhibited TGFβ-induced RAW phagocytosis of zymosan particles (Fig. 7c, d). Taken together, these results indicate that TIMAP regulation of MLC phosphorylation plays an essential role in TGFβ-induced macrophage migration and phagocytosis.
TIMAP regulation of MLC phosphorylation contributes to TGFβ-induced macrophage migration and phagocytosis. a Cell migration assay. RAW cells were stably transfected with an HA-tagged mutant (HA-TIMAPC565S) or wild-type TIMAP (HA-TIMAPWT) expression plasmid and the positive cells were selected. The cells were treated with TGFβ (5 ng/ml) and/or MLCK inhibitor ML-7 (10 μM), respectively. The cell monolayer was wounded by scraping with a pipette tip and the cell migrations were monitored at time zero (0 h) and 24 h. b Cells migration distances were measured and analyzed by the ImageJ software as before. TIMAP protein expressions were verified by Western blotting (bottom). c RAW cells were first stimulated with M-CSF (50 ng/ml) for 12 h and then treated with TGFβ (5 ng/ml) in the presence or absence of ML-7 (10 μM) for 24 h before addition of zymosan particles. Phagocytosis was allowed to proceed for 2 h, and photograph was taken at ×600 magnification. d Quantification of c. All the experiments were repeated at least three times and representative results are shown. *p < 0.05 versus control, # p < 0.05 versus TGFβ treatment
Discussion
In this study, we have made several novel findings: (a) TIMAP level is reduced in TGFβ-elevated fibrotic mouse kidney, providing the first in vivo evidence that TGFβ represses TIMAP in diseased kidneys. (b) TIMAP is highly expressed in macrophages and TGFβ represses its transcription through HDAC3-dependent Smad signaling. (c) TIMAP interrupts TGFβ/Smad signaling and negatively affects the TGFβ-induced macrophage M2 polarization, suggesting that TIMAP participates in TGFβ-associated wound repair/tissue remodeling processes. (d) TIMAP dephosphorylates MLC in macrophages and TGFβ repression of TIMAP increases MLC phosphorylation, which constitutes a critical part of TGFβ/Smad signaling pathways promoting macrophage M2 phenotypic phagocytosis (Fig. 8).
A schematic representation of TGFβ repression of TIMAP and its regulation of macrophage phagocytosis. TGFβ induces HDAC3-associated Smad signaling that transcriptionally represses TIMAP, a regulatory subunit of myosin phosphatase (MP) displaying inhibiting properties against TGFβ/Smad signaling and macrophage M2 polarization. In turn, TIMAP dephosphorylation of MLC is abrogated, which facilitates TGFβ and myosin light chain kinase (MLCK) promotion of macrophage migration and phagocytosis
TGFβ plays a key role in tissue homeostasis, adult immunity, and wound repair [2]. TGFβ regulates various cellular processes through either Smad-dependent or Smad-independent signaling pathways, such as MAPK, RhoA, or PKC [34]. The identification of the Smad binding elements on TIMAP promoter and further confirmation of Smad2/3-dependent repression of TIMAP transcription establish that TIMAP is a direct TGFβ/Smad target. Discovery of TIMAP preferential expression in macrophage and its participation in macrophage M2 polarization are important because TGFβ-regulated macrophage activity plays an essential role in various inflammatory disorders. It is generally accepted that macrophages polarized to M1 phenotype early during inflammation and shifted to M2 polarization later, which released high level TGFβ, to repair injured tissues [30, 35]. We clearly demonstrated that TGFβ HDAC3-dependently represses TIMAP, which blocked TIMAP dephosphorylation of MLC and promoted macrophage migration and phagocytosizing activities. Therefore, our results elaborate a novel regulatory loop that provides additional control over TGFβ-associated injury repair and tissue remodeling processes.
HDAC, as a protein acetylation modifier, is emerging as a key epigenetic regulator of various physiological or pathological processes. HDAC inhibitions are protective against various inflammatory disorders, tissue fibrosis, and cancer, and especially, TGFβ/Smad signaling is markedly affected by HDAC activities [36, 37]. Intriguingly HDAC either positively or negatively affects TGFβ signaling [38]. For example, the inductions of HDAC2 [39], HDAC4 [40] and HDAC6 [41] are associated with TGFβ-induced upregulation of pro-fibrogenic proteins and epithelial mesenchymal transition after kidney injury, which worsen renal fibrosis, while HDAC3 upregulation, presumably complexed with R-Smad/transcriptional co-repressors, correlates with the inhibitions of some TGFβ-regulated renal protective molecules such as miR-30 [42], PPARγ [43], and TIMAP as demonstrated in this study. Based on these observations, it seems that HDAC subtype involvements in TGFβ-associated cellular processes are cell type- and target gene-specific and the protective effects of general HDAC inhibition under a specific disease setting are achieved through its multiple actions on distinct signaling or effecter molecules. Likewise, the concomitant inhibition of HDAC3 and reversal of TIMAP repression contribute significantly to the protection.
Macrophage migration and phagocytosis are the crucial features of TGFβ-associated M2 phenotypes, which involve cytoskeleton remodeling and cell membrane movement driven by myosin-associated actin polymerization and contractivity [44]. MLC phosphorylation promotes actin/myosin filament contraction and cell motility [45] and its level is controlled by MLC kinase and myosin phosphatase (MP) [46]. MP holoenzyme is composed of a catalytic subunit of type 1 phosphatase isoform (PP1c), a small subunit (M20) of unknown function, and a target subunit which determines MP substrate specificity [47]. Although TIMAP is a member of MP target subunit family, its substrate in macrophage is unknown and there is paucity of information regarding the regulatory cascade mediating TGFβ-induced macrophage phagocytosis. Our results indicate that MLC is a TIMAP substrate in macrophages since TIMAP physically associates with MLC (Fig. 6g, h) and TIMAP levels inversely correlate with MLC phosphorylation (Fig. 6d–f). More importantly, TGFβ induces macrophage migration and phagocytosis in TIMAP- and MLC phosphorylation-sensitive manners (Fig. 7a, c, d), indicating that TGFβ repression of TIMAP leads to increased MLC phosphorylation and subsequent macrophage migration and phagocytosis. Therefore, our study uncovered TIMAP dephosphorylation of MLC as a critical component mediating TGFβ-induced macrophage migration and phagocytic processes.
In conclusion, our studies have identified TIMAP as a macrophage-enriched protein whose transcription is suppressed by TGFβ through HDAC3-associated Smad signaling. We further demonstrated that TGFβ repression of TIMAP increased MLC phosphorylation and accelerated macrophage M2 phenotypic migration and phagocytosis. Macrophage M2 polarization-associated wound repair and tissue remodeling processes are generally considered beneficial for the host; however, macrophages often become hyper-responsive, resulting in uncontrolled or deregulated release of TGFβ that exacerbates acute tissue injury and promotes the development of chronic diseases [48, 49]. Intervention strategies inhibiting HDAC and TGFβ activity have been considered the potential medications in anti-cancer and anti-fibrosis therapies [2, 50]; hence, the strategies targeting HDAC3/TIMAP axis might beneficially affect TGFβ-associated pathological outcomes.
References
Cao W, Mattagajasingh SN, Xu H, Kim K, Fierlbeck W, Deng J, Lowenstein CJ, Ballermann BJ (2002) TIMAP, a novel CAAX box protein regulated by TGF-beta1 and expressed in endothelial cells. Am J Physiol Cell Physiol 283: C327–C337
Hawinkels LJ, Ten Dijke P (2011) Exploring anti-TGF-beta therapies in cancer and fibrosis. Growth Factors 29:140–152
Poirier C, Gorshkov BA, Zemskova MA, Bogatcheva NV, Verin AD (2011) TIMAP protects endothelial barrier from LPS-induced vascular leakage and is down-regulated by LPS. Respir Physiol Neurobiol 179:334–337
Kim K, Li L, Kozlowski K, Suh HS, Cao W, Ballermann BJ (2005) The protein phosphatase-1 targeting subunit TIMAP regulates LAMR1 phosphorylation. Biochem Biophys Res Commun 338: 1327–1334
Li L, Kozlowski K, Wegner B, Rashid T, Yeung T, Holmes C, Ballermann BJ (2007) Phosphorylation of TIMAP by glycogen synthase kinase-3beta activates its associated protein phosphatase 1. J Biol Chem 282: 25960–25969
Csortos C, Czikora I, Bogatcheva NV, Adyshev DM, Poirier C, Olah G, Verin AD (2008) TIMAP is a positive regulator of pulmonary endothelial barrier function. Am J Physiol Lung Cell Mol Physiol 295:L440–L450
Boratko A, Gergely P, Csortos C (2013) RACK1 is involved in endothelial barrier regulation via its two novel interacting partners. Cell Commun Signal 11:2
Obeidat M, Li L, Ballermann BJ (2014) TIMAP promotes angiogenesis by suppressing PTEN-mediated Akt inhibition in human glomerular endothelial cells. Am J Physiol Renal Physiol 307:F623–F633
Davis BN, Hilyard AC, Nguyen PH, Lagna G, Hata A (2010) Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Mol Cell 39:373–384
Simonsson M, Kanduri M, Gronroos E, Heldin CH, Ericsson J (2006) The DNA binding activities of Smad2 and Smad3 are regulated by coactivator-mediated acetylation. J Biol Chem 281:39870–39880
Nomura T, Khan MM, Kaul SC, Dong HD, Wadhwa R, Colmenares C, Kohno I, Ishii S (1999) Ski is a component of the histone deacetylase complex required for transcriptional repression by Mad and thyroid hormone receptor. Genes Dev 13:412–423
Tabata T, Kokura K, Ten Dijke P, Ishii S (2009) Ski co-repressor complexes maintain the basal repressed state of the TGF-beta target gene, SMAD7, via HDAC3 and PRMT5. Genes Cells 14:17–28
Tavian M, Cortes F, Robin C, Schiavon V, Hallais MF, Coulombel L, Charbord P, Labastie MC, Peault B (2000) The hemangioblast, common precursor of endothelial and hematopoietic cells. Transfus Clin Biol 7:238–241
Bohdanowicz M, Grinstein S (2013) Role of phospholipids in endocytosis, phagocytosis, and macropinocytosis. Physiol Rev 93:69–106
Niedergang F, Chavrier P (2004) Signaling and membrane dynamics during phagocytosis: many roads lead to the phagos(R)ome. Curr Opin Cell Biol 16:422–428
Stuart LM, Ezekowitz RA (2005) Phagocytosis: elegant complexity. Immunity 22:539–550
Sun C, MH W, Yuan SY (2011) Nonmuscle myosin light-chain kinase deficiency attenuates atherosclerosis in apolipoprotein E-deficient mice via reduced endothelial barrier dysfunction and monocyte migration. Circulation 124:48–57
Sharma L, Wu W, Dholakiya SL, Gorasiya S, Wu J, Sitapara R, Patel V, Wang M, Zur M, Reddy S et al (2014) Assessment of phagocytic activity of cultured macrophages using fluorescence microscopy and flow cytometry. Methods Mol Biol 1172:137–145
Shopik MJ, Li L, Luu HA, Obeidat M, Holmes CF, Ballermann BJ (2013) Multi-directional function of the protein phosphatase 1 regulatory subunit TIMAP. Biochem Biophys Res Commun 435: 567–573
Qin T, Du R, Huang F, Yin S, Yang J, Qin S, Cao W (2016) Sinomenine activation of Nrf2 signaling prevents hyperactive inflammation and kidney injury in a mouse model of obstructive nephropathy. Free Radic Biol Med 92:90–99
Qin S, Du R, Yin S, Liu X, Xu G, Cao W (2015) Nrf2 is essential for the anti-inflammatory effect of carbon monoxide in LPS-induced inflammation. Inflamm Res 64:537–548
Cao W, Bao C, Padalko E, Lowenstein CJ (2008) Acetylation of mitogen-activated protein kinase phosphatase-1 inhibits Toll-like receptor signaling. J Exp Med 205:1491–1503
Yin S, Cao W (2015) TLR signaling induces Nrf2 pathway activation through p62-triggered Keap1 degradation. Mol Cell Biol. doi:10.1128/MCB.00105-15
Konopski Z, Fandrem J, Seljelid R, Eskeland T (1995) Interferon-gamma inhibits endocytosis of soluble animated beta-1,3-D-glucan and neutral red in mouse peritoneal macrophages. J Interf Cytokine Res 15:597–603
Fleetwood AJ, Lawrence T, Hamilton JA, Cook AD (2007) Granulocyte-macrophage colony-stimulating factor (CSF) and macrophage CSF-dependent macrophage phenotypes display differences in cytokine profiles and transcription factor activities: implications for CSF blockade in inflammation. J Immunol 178:5245–5252
Gitik M, Reichert F, Rotshenker S (2010) Cytoskeleton plays a dual role of activation and inhibition in myelin and zymosan phagocytosis by microglia. FASEB J 24:2211–2221
Fuentes AL, Millis L, Vapenik J, Sigola L (2014) Lipopolysaccharide-mediated enhancement of zymosan phagocytosis by RAW 264.7 macrophages is independent of opsonins, laminarin, mannan, and complement receptor 3. J Surg Res 189:304–312
Klahr S, Morrissey J (2003) Obstructive nephropathy and renal fibrosis: the role of bone morphogenic protein-7 and hepatocyte growth factor. Kidney Int Suppl 87:S105–S112
Shi Y, Massague J (2003) Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 113:685–700
Gordon S (2003) Alternative activation of macrophages. Nat Rev Immunol 3:23–35
Li C, Yang G, Ruan J (2012) Sphingosine kinase-1/sphingosine-1-phosphate receptor type 1 signalling axis is induced by transforming growth factor-beta1 and stimulates cell migration in RAW264.7 macrophages. Biochem Biophys Res Commun 426:415–420
Levin R, Grinstein S, Schlam D (2015) Phosphoinositides in phagocytosis and macropinocytosis. Biochim Biophys Acta 1851:805–823
Ashcroft GS (1999) Bidirectional regulation of macrophage function by TGF-beta. Microbes Infect 1:1275–1282
Derynck R, Zhang YE (2003) Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 425:577–584
Braga TT, Agudelo JS, Camara NO (2015) Macrophages during the fibrotic process: M2 as friend and foe. Front Immunol 6:602
Liu N, Zhuang S (2015) Treatment of chronic kidney diseases with histone deacetylase inhibitors. Front Physiol 6:121
Pang M, Kothapally J, Mao H, Tolbert E, Ponnusamy M, Chin YE, Zhuang S (2009) Inhibition of histone deacetylase activity attenuates renal fibroblast activation and interstitial fibrosis in obstructive nephropathy. Am J Physiol Renal Physiol 297:F996–F1005
Massague J, Wotton D (2000) Transcriptional control by the TGF-beta/Smad signaling system. EMBO J 19:1745–1754
Ryu JK, Kim WJ, Choi MJ, Park JM, Song KM, Kwon MH, Das ND, Kwon KD, Batbold D, Yin GN et al (2013) Inhibition of histone deacetylase 2 mitigates profibrotic TGF-beta1 responses in fibroblasts derived from Peyronie’s plaque. Asian J Androl 15:640–645
Glenisson W, Castronovo V, Waltregny D (2007) Histone deacetylase 4 is required for TGFbeta1-induced myofibroblastic differentiation. Biochim Biophys Acta 1773:1572–1582
Shan B, Yao TP, Nguyen HT, Zhuo Y, Levy DR, Klingsberg RC, Tao H, Palmer ML, Holder KN, Lasky JA (2008) Requirement of HDAC6 for transforming growth factor-beta1-induced epithelial-mesenchymal transition. J Biol Chem 283:21065–21073
Liu L, Lin W, Zhang Q, Cao W, Liu Z (2015) TGF-beta induces miR-30d down-regulation and podocyte injury through Smad2/3 and HDAC3-associated transcriptional repression. J Mol Med (Berl). doi:10.1007/s00109-015-1340-9
Jiang X, Ye X, Guo W, Lu H, Gao Z (2014) Inhibition of HDAC3 promotes ligand-independent PPARgamma activation by protein acetylation. J Mol Endocrinol 53:191–200
Bright MD, Frankel G (2011) PAK4 phosphorylates myosin regulatory light chain and contributes to Fcgamma receptor-mediated phagocytosis. Int J Biochem Cell Biol 43:1776–1781
Deery WJ, Heath JP (1993) Phagocytosis induced by thyrotropin in cultured thyroid cells is associated with myosin light chain dephosphorylation and stress fiber disruption. J Cell Biol 122:21–37
Mansfield PJ, Shayman JA, Boxer LA (2000) Regulation of polymorphonuclear leukocyte phagocytosis by myosin light chain kinase after activation of mitogen-activated protein kinase. Blood 95:2407–2412
Ito M, Nakano T, Erdodi F, Hartshorne DJ (2004) Myosin phosphatase: structure, regulation and function. Mol Cell Biochem 259:197–209
Ricevuti G (1997) Host tissue damage by phagocytes. Ann N Y Acad Sci 832:426–448
Laskin DL, Sunil VR, Gardner CR, Laskin JD (2011) Macrophages and tissue injury: agents of defense or destruction? Annu Rev Pharmacol Toxicol 51:267–288
Meng XM, Nikolic-Paterson DJ, Lan HY (2016) TGF-beta: the master regulator of fibrosis. Nat Rev Nephrol. doi:10.1038/nrneph.2016.48
Acknowledgments
This work was supported by research grants from the National Nature Science Foundation of China (81271301, 81470940, and 81670672) to W. Cao.
Authors’ contributions
JY performed most experiments and participated in manuscript writing; SY, FB, LL, and QT assisted in animal work, data collection, and some experiments. HW provided critical advice; WC designed and conceived the project, analyzed and arranged results, and wrote the manuscript. All the authors reviewed the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Yang, J., Yin, S., Bi, F. et al. TIMAP repression by TGFβ and HDAC3-associated Smad signaling regulates macrophage M2 phenotypic phagocytosis. J Mol Med 95, 273–285 (2017). https://doi.org/10.1007/s00109-016-1479-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-016-1479-z