Abstract
Cells use mitophagy to remove dysfunctional or excess mitochondria, frequently in response to imposed stresses, such as hypoxia and nutrient deprivation. Mitochondrial cargo receptors (MCR) induced by these stresses target mitochondria to autophagosomes through interaction with members of the LC3/GABARAP family. There are a growing number of these MCRs, including BNIP3, BNIP3L, FUNDC1, Bcl2-L-13, FKBP8, Prohibitin-2, and others, in addition to mitochondrial protein targets of PINK1/Parkin phospho-ubiquitination. There is also an emerging link between mitochondrial lipid signaling and mitophagy where ceramide, sphingosine-1-phosphate, and cardiolipin have all been shown to promote mitophagy. Here, we review the upstream signaling mechanisms that regulate mitophagy, including components of the mitochondrial fission machinery, AMPK, ATF4, FoxOs, Sirtuins, and mtDNA release, and address the significance of these pathways for stress responses in tumorigenesis and metastasis. In particular, we focus on how mitophagy modulators intersect with cell cycle control and survival pathways in cancer, including following ECM detachment and during cell migration and metastasis. Finally, we interrogate how mitophagy affects tissue atrophy during cancer cachexia and therapy responses in the clinic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
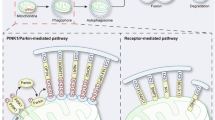
Mitophagy is a selective form of macro-autophagy in which mitochondria are preferentially targeted for degradation at the autophagolysosome [1, 2]. Mitophagy is required to eliminate mitochondria that have become dysfunctional, and thus serves an important housekeeping role that preserves mitochondrial function and limits production of damaging reactive oxygen species (ROS). Mitophagy can also eliminate healthy mitochondria to reduce overall mitochondrial mass as an adaptive response to stresses such as hypoxia and nutrient deprivation, to ensure efficient use of scarce metabolites and oxygen, and again limit production of excess ROS. The turnover of mitochondria by mitophagy relies on the activity of a growing list of mitochondrial cargo receptors (MCRs), that include BNIP3, BNIP3L (NIX), FUNDC1, Bcl2-L-13, FKBP8, FANC-C, Prohibitin-2 (PHB-2), Metaxin-1, Cardiolipin, and others, in addition to mitochondrial proteins that are substrates of PINK1/Parkin-mediated ubiquitination. These MCRs localize to the mitochondria and promote mitophagy by interacting directly with processed LC3/GABARAP proteins through conserved LC3 Interaction Regions (LIR) (Fig. 1). Importantly, the expression and/or processing of these MCRs is modulated by stresses that induce mitophagy, including mitochondrial depolarization, hypoxia, nutrient deprivation, etc. For example, BNIP3, BNIP3L, and FUNDC1-mediated mitophagy is activated by hypoxia, while cardiolipin and PHB-2 are exposed by mitochondrial damage. In addition to these MCRs that bind LC3/GABARAP proteins directly, there are other signaling molecules that contribute to activation of mitophagy including DRP1, FIS1, and MFF that are better known for their role in other aspects of mitochondrial dynamics. Ceramide and sphingolipids modulate mitophagy partly by recruiting LC3 to mitochondria, while other novel mitophagy modulators such as the adenine nucleotide transporter (ANT) affect mitophagy by controlling PINK1 import into mitochondria for degradation. The growing list of mitophagy modulators (Table 1) underlines the extent to which it is important for cells to control mitochondrial turnover and mitochondrial function in response to different stresses and signals. That there are so many different mitophagy modulators also causes one to question the extent to which these mitophagy pathways are redundant and interact with each other, or alternatively are uniquely required for mitophagy in response to distinct stresses. In this review, we will examine some of these questions in the context of tumorigenesis and metastasis to assess whether mitophagy may be a valid and more targeted therapy in cancer treatment than general autophagy.
Modulation of mitochondrial cargo receptor interactions with LC3 family members by post-translational modification. a Alignment of the LIR motifs (blue) in BNIP3, BNIP3L, Bcl1L13 and FUNDC1 mitochondrial cargo receptors that are required for them to interact with processed LC3/GABARAP family members, and highlighting the serine residue (red) at − 1 in each LIR motif that is subject to phosphorylation to further promote LC3/GABARAP interaction. b BNIP3 is phosphorylated on S17 (and S24) adjacent to the critical tryptophan reside at W18 which promotes its interaction with LC3B-II. The kinase responsible is currently unknown. c Similarly, BNIP3L is phosphorylated on S35 (and S34) adjacent to W36 of its LIR motif that promotes its interaction with LC3B and GABARAP. d ULK1 phosphorylates FUNDC1 on S17 to promote LC3 interaction, while PGAM5 (not shown) de-phosphorylates inhibitory phospho-S13, also to promote LC3 interaction. e Interaction of FUNDC1 with LC3B is repressed by phosphorylation on Y18 by SRC kinase within its LIR motif and also inhibited by CK2 phosphorylation on S13. f Bcl2-L-13 is phosphorylated on S272 adjacent to its LIR motif and also interacts with ULK1, although whether ULK1 phosphorylates Bcl2-L-13 on S272 has not been shown
Mitophagy pathways and mitochondrial cargo receptors
Before addressing the role of different mitophagy modulators in cancer, we first review the major molecular pathways that regulate mitophagy with a particular focus on mitochondrial cargo receptors that localize to mitochondria in response to mitochondrial stresses. Table 1 summarizes known mitophagy modulators more comprehensively.
PINK1/Parkin-dependent mitophagy
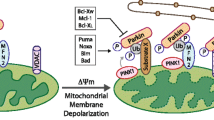
Mitochondrial membrane depolarization (ΔΨmt) induces mitophagy through activation of PTEN-induced putative kinase-1 (PINK1), a serine/threonine ubiquitin kinase encoded by the PARK6 locus, which accumulates at the outer mitochondrial membrane (OMM) where it recruits E3 ubiquitin ligases, most notably Parkin, encoded by the PARK2 locus [3, 4]. PINK1 phosphorylates Parkin on S65 within its ubiquitin-like (UBL) domain, derepressing Parkin auto-inhibition and amplifying recruitment of active Parkin to the OMM [4]. This leads to the ubiquitination and phosphorylation of proteins at the OMM, such as Mitofusin-2 (Mfn2) and the voltage-dependent anion channel-1 (VDAC-1), that then interact with the Optineurin (OPTN) and NDP52 cargo receptors, but also p62/Sqstm1 in some systems, which contain critical LIR motifs [3, 5,6,7]. PINK1/Parkin-dependent mitophagy is the major mechanism in the cell for elimination of depolarized mitochondria (Fig. 2).
PINK1 degradation controls turnover of depolarized mitochondria. a PINK1 is a ubiquitin kinase that is degraded at functional mitochondria. PINK1 import via the TOM/TIM protein complexes is highly regulated and dependent on mitochondrial membrane polarization. PINK1 uptake into the mitochondria is positively regulated by the Atad3a AAA + ATPase, and negatively regulated by the adenine nucleotide translocase (ANT) that interferes with TOM23 and TIM40 to block PINK1 uptake. This function of ANT is separable from its function in ADP/ATP exchange at the IMM. The MTS of PINK1 is proteolytically cleaved off in the matrix by MPP, while the transmembrane part of PINK1 is cleaved by PARL protease and then retrotranslocated back to the cytosol for proteasomal degradation. b In response to loss of mitochondrial membrane potential (∆Ψmt), PINK1 uptake at the mitochondria is blocked and it accumulates at the OMM, where it recruits the Parkin E3 ubiquitin ligase. PINK1 phosphorylates Parkin within it UBL domain derepressing its activity and also phosphorylates ubiquitin groups on Parkin substrates (S). Parkin ubiquitinates substrates (S) at the OMM such as MFN-2 and Vdac1 and other proteins and these phospho-ubiquitinated substrates are bound by cargo receptors, such as p62/Sqstm1 and OPTN1, which also interact with LC3 to promote mitophagy. Recruitment of OPTN1 and other cargo receptors to the OMM is dependent on TBK1 that is recruited away from centrosomes and its role in mitosis to the OMM in PINK1/Parkin-dependent manner
At healthy mitochondria, the amino-terminal mitochondrial targeting sequence (MTS) of PINK1 is translocated to the mitochondrial matrix where it is cleaved off by mitochondrial processing peptidase (MPP), while the transmembrane portion of PINK1 is cleaved by PINK/PGAM5-associated rhomboid-like (PARL) proteases, and then, the remaining 52 kD fragment of PINK1 is retrotranslocated back out to the cytosol and degraded at the proteasome according to the N-end rule [4, 8, 9]. Mitochondrial import of PINK1 at healthy mitochondria is facilitated by Atad3a, an AAA + ATPase that spans the IMM and OMM [10]. Inactivation of Atad3a increases mitophagy of both depolarized and healthy mitochondria, which is partially rescued by PINK1 deletion, indicating that Atad3a plays an important role in preventing aberrant accumulation of PINK1 [10]. Conversely, the adenine nucleotide transporter-1 and -2 (ANT) plays a role in promoting PINK1 accumulation by suppressing translocase of inner membrane (TIM) proteins, TIM23 and TIM44, that are involved in PINK1 mitochondrial import [11]. This function of the ANT was independent of its role in ADP/ATP exchange at the IMM and loss of ANT decreased levels of both PINK1 and Parkin in mouse brain and heart, causing the accumulation of dysfunctional mitochondria and cardiomyopathy [11]. Interestingly, mutant forms of ANT found in patients with cardiomyopathy and opthalmoplegia were unable to rescue mitophagy defects in ANT null cells, whereas mutant ANT that cannot bind ATP or promote ADP/ATP exchange was able to rescue mitophagy defects. Together with evidence showing mitochondrial abnormalities in these patients, this work indicates that defective mitophagy arising from mutant ANT causes the pathologies in these patients [11].
In addition to their activation by mitochondrial depolarization, PINK1 and Parkin-dependent mitophagy are also activated by accumulation of unfolded proteins in the mitochondrial matrix [12, 13] suggesting a link between the mitochondrial unfolded protein response (UPRmt) and mitophagy, two key mitochondrial quality control processes in the cell.
While most research has focused on the interaction of PINK1 with Parkin, PINK1 interacts functionally with other mitochondrial E3 ubiquitin ligases involved in mitophagy, including ARIH1, MARCH5, and MUL1, which can substitute for Parkin in mitophagy in various contexts [14,15,16,17].
BNIP3/BNIP3L-dependent mitophagy
Originally categorized as pro-apoptotic BH3-only proteins, BNIP3 and BNIP3L (NIX) function primarily as stress-induced MCRs. BNIP3 and BNIP3L are tail-anchored proteins that integrate into the OMM via their carboxy terminal transmembrane domains (TMD) with the remaining bulk of both proteins facing into the cytosol where a tightly conserved LIR motif (WVEL) near the amino-terminal end of each protein (W18 to L21 in BNIP3, and W36 to L39 in BNIP3L; Fig. 1a) interacts with processed LC3/GABARAP [18,19,20]. The putative BH3 domains in BNIP3 and BNIP3L are poorly conserved (2 amino acids out of 11) and are redundant for BNIP3/BNIP3L function in mitophagy [21, 22]. Furthermore, the interaction of BNIP3 and BNIP3L with Bcl-XL (and Bcl-2) is mediated via the amino-terminal region overlapping with the LIR motif, and not dependent on the putative BH3 domains in BNIP3/BNIP3L [23]. This work also suggested that the interaction of BNIP3/BNIP3L with Bcl-XL enhances their interaction with LC3/GABARAP family members to promote mitophagy [23]. The BNIP3 TMD consists of a glycine zipper that mediates homo-dimerization via hydrogen bonding between polar residues on the inside of the glycine zipper, while hydrophobic residues on the outside of the zipper stabilize the dimer in the OMM lipid bilayer [24]. Remarkably, BNIP3 and BNIP3L dimers are resistant to SDS, DDT, and urea, and can be visualized on denaturing gels [25, 26] but to what extent BNIP3 or BNIP3L dimers are stable in vivo has not been examined. While dimerization is not required for integration into the OMM, recent evidence suggests that dimerization is required for efficient LC3/GABARAP interaction [26]. Also, intriguingly, BNIP3 and BNIP3L have been shown to heterodimerize [27], but when and where such heterodimers form in a physiological context has not been determined.
BNIP3 and BNIP3L are rapidly induced at a transcriptional level by stresses, such as hypoxia and nutrient deprivation, but like many stress response proteins, both proteins are also rapidly turned over by mitophagy and at the proteasome, requiring the stressed cell to maintain transcription of BNIP3 and BNIP3L until the stress is removed [2]. Both BNIP3 and BNIP3L are transcriptionally activated by HIF-1 in response to hypoxia and elevated ROS [28, 29], and BNIP3 in particular is a strong hypoxia marker that gets induced rapidly and to high levels as oxygen drops below 2% [30,31,32]. BNIP3 is also a validated transcriptional target of PPARα explaining its induction in the liver in response to nutrient deprivation and fasting [22, 33, 34], and is induced by FoxO3A in atrophying and cachectic muscle [35,36,37]. BNIP3 and BNIP3L are both transcriptionally regulated by p53 [38, 39], but while BNIP3L is upregulated transcriptionally by p53 [38], BNIP3 is repressed [39], suggesting that they play differential roles in the response to DNA damage and other signals that activate p53.
In addition to regulation at a transcriptional level, BNIP3 and BNIP3L are modulated post-translationally by phosphorylation [23, 40]. Both proteins are phosphorylated on the serine residue (S17 in BNIP3 and S35 in BNIP3L) adjacent to the critical tryptophan residue in their LIR motifs (W18 in BNIP3 and W36 in BNIP3L) [23, 40] (Fig. 1b, c). These phosphorylation events promote their interaction with LC3/GABARAP [23, 40], and also their interaction with Bcl-XL [23]. BNIP3 and BNIP3L are multiply phosphorylated, but it remains to be determined what kinases are involved in modulating BNIP3/BNIP3L levels and activity.
Beyond phosphorylation, BNIP3L is ubiquitinated by Parkin and promotes PINK1/Parkin-induced mitophagy [41, 42]. However, neither BNIP3 nor BNIP3L requires PINK1 or Parkin to elicit mitophagy [43], and conversely, BNIP3 is not required for PINK1 accumulation, or for Parkin recruitment to the OMM [43]. Nevertheless, Parkin/PINK1 loss causes HIF-1α stabilization [44, 45] but whether BNIP3 or BNIP3L, that are both HIF targets, are upregulated by PINK1/Parkin loss has not been examined, although BNIP3 can compensate in mitophagy for Parkin loss [43].
BNIP3 and BNIP3L play important functions in tissue differentiation and stress responses with each protein exhibiting distinct tissue expression patterns [2]. BNIP3 is induced in a PPAR-α-dependent manner in response to fasting [22, 33, 34] and is required for glucagon-induced mitophagy in the liver [22]. BNIP3 exhibits zonal expression patterning in the liver with high expression around the central vein of the liver lobule where cells are most hypoxic [46], and low expression around the more oxygenated periportal vein [22]. As a result, mitochondrial mass is zonal in the liver with higher mitochondrial mass around the central vein and lower mitochondrial mass around the periportal vein [22]. Metabolic processes are spatially organized across the liver lobule, such that opposing activities, such as urea cycle and glutamine synthesis, fatty acid oxidation and lipogenesis, and gluconeogenesis and glycolysis, are physically separated [47]. This likely improves the efficiency of metabolite re-cycling and avoids competition between metabolic pathways for common substrates [47]. Loss of BNIP3-dependent mitophagy caused mitochondria to accumulate aberrantly and disrupted metabolic zonation in the liver, such that periportal processes such as urea cycle were expanded at the expense of central vein concentrated processes, such as glutamate-glutamine cycling and glycolysis [22].
BNIP3L is markedly upregulated and essential for mitophagy during normal red blood cell differentiation such that loss of BNIP3L resulted in anemia and splenomegaly due to production of defective reticulocytes [48,49,50]. Sphingosine kinase (SPHK1) contributes to BNIP3L induction during erythroid differentiation [51]. SPHK1 is required for the UPRmt in worms which is a transcriptional response to mitochondrial stress mediated by transcription factors such as ATF4, NRF2, and FoxOs that induce expression of genes that mitigate against mitochondrial stress, including mitochondrial proteases, mitochondrial protein chaperones, autophagy genes, anti-oxidants, etc. [52, 53]. Taken together with the mitochondrial localization of SPHK1 in response to stress, this suggests a mechanism linking activation of the UPRmt to mitophagy through induction of BNIP3L. BNIP3L is also required for differentiation of retinal ganglion cells during embryogenesis, where it is induced in mid-gestation by hypoxia [54]. This results in a BNIP3L-dependent glycolytic switch that is also seen during the polarization of M1 macrophages [54]. BNIP3/BNIP3L-dependent mitophagy was also required for survival of long-lived NK memory cells in response to viral infection [55].
Both BNIP3 and BNIP3L interact with Rheb [56, 57], the small GTPase required to promote lysosomal localization and activity of mTOR, a major growth-promoting kinase in the cell [58]. BNIP3 binding repressed Rheb activity and disrupting the BNIP3–Rheb interaction promoted mTOR activity and cell growth, consistent with a growth suppressive role for BNIP3 via its inhibitory interaction with Rheb [56]. By contrast, the interaction of BNIP3L with Rheb elicited mTOR-independent effects on cell growth [57]. BNIP3L-dependent mitophagy induced by switching cells from growth in glucose-only media to growth in glutamine plus galactose required interaction of BNIP3L with Rheb at the OMM in a tri-molecular complex with processed LC3 [57]. The relative importance of BNIP3/BNIP3L interactions with Rheb has not yet been addressed in a physiological setting.
FUNDC1
Like BNIP3 and BNIP3L, FUNDC1 promotes hypoxia-induced mitophagy, but in contrast to BNIP3 and BNIP3L, the activation of FUNDC1 by hypoxia is mediated primarily at a post-translational level, via regulation of its phosphorylation state [59]. Like BNIP3 and BNIP3L, FUNDC1 integrates into the OMM, although FUNDC1 is a multi-pass protein with three TMDs, and has an amino-terminal LIR motif (YVEL from Y18 to L21), which projects into the cytosol to interact with LC3 family members [59]. Its interaction with LC3B is tightly modulated by phosphorylation on S17 immediately adjacent to its LIR motif that is phosphorylated by ULK1 (Fig. 1d), the catalytic component of the autophagy pre-initiation complex [60, 61], in response to hypoxia to enhance LC3B interaction and mitophagy [62]. FUNDC1 is also phosphorylated by oncogenic SRC on Y18 within the LIR motif (Fig. 1e) to block LC3 interaction and mitophagy [62]. The FUNDC1–LC3B interaction is also inhibited by phosphorylation on S13 by Casein Kinase 2 (CK2). Conversely, PGAM5 phosphatase de-phosphorylates S13 to promote LC3 interaction and mitophagy [63]. Interestingly, PGAM5 also plays a role in mitophagy by promoting PINK1 accumulation [64]. FUNDC1 is also modulated by the MARCH5 E3 Ub ligase that ubiquitinates FUNDC1 on K119 to promote its turnover and limit hypoxia-induced mitophagy [17]. In addition to interacting with LC3B, FUNDC1 interacts with DRP1 and Calnexin at mitochondrial–ER junctions where it appears to be required for recruiting DRP1, since mutant forms of FUNDC1 that cannot bind DRP1 cannot promote mitophagy [65]. FUNDC1 may also play a role in the cellular response to proteostatic stress through its interaction with HSC70 [66]. While the hypoxia-induced microRNA, miR-137 represses both FUNDC1 and BNIP3L expression during hypoxia possibly limiting mitophagy [67], the extent to which FUNDC1 function in hypoxia-induced mitophagy overlaps with that of BNIP3 and BNIP3L has not been examined.
Bcl2-L-13
Bcl2-L-13 is a Bcl-2-related homolog of ATG32 in yeast that integrates into the OMM via a C-terminal TMD and interacts with LC3B via a conserved LIR motif (WQQI) at amino acids 273–276 (Fig. 1f) [68]. Bcl2-L-13 over-expression induces mitochondrial fragmentation and mitophagy, independent of DRP-1 and Parkin, and rescues mitophagy in Atg32-deleted yeast [68]. Bcl2-L-13 expression is induced by mitochondrial depolarization and knockdown of Bcl2-L-13 impeded CCCP-induced mitophagy [68]. Given that Bcl1-L-13 acts independently of Parkin, these results suggest that Parkin is not essential for mitophagy induced by mitochondrial membrane depolarization, but further work is needed to determine whether Bcl2-L-13 depends on PINK1 and other mitochondrial E3 Ub ligases to function. The induction of Bcl2-L-13 by mitochondrial depolarization was associated with its phosphorylation on S272, adjacent to its LIR motif at 273–276 [68]. Recent work has shown that ULK1 binds to Bcl2-L-13 in a manner dependent on the kinase activity of ULK1, in a tri-molecular complex with LC3B, and that ULK1 is required for Bcl2-L-13-dependent mitophagy [69], but whether ULK1 phosphorylates Bcl2-L-13 on S272 was not determined. Physiological studies addressing the importance of Bcl2-L-13 for mitophagy in vivo have not been reported.
Prohibitin-2
Prohibitin-2 (PHB-2) promotes cristae structure and supports electron transport chain (ETC) complex formation at the IMM [70]. PHB-2 interacts directly with LC3B via an LIR motif (YQRL) at amino acids 121–124 that is required for mitophagy induced by mitochondrial membrane depolarization and to eliminate paternal mitochondria in worms [71]. It has been suggested that proteasomal degradation of OMM proteins during mitophagy is required to expose IMM proteins, such as PHB-2 that are turned over by mitophagy [72], and indeed, PHB-2 relies on Parkin-mediated rupture of the OMM to promote mitophagy [71]. Various stresses known to induce mitophagy, including mitochondrial depolarization, happen at the IMM, and thus, it is significant that IMM molecules like PHB-2 and cardiolipin induce mitophagy.
Cardiolipin, ceramide, and lipid-dependent mitophagy
Cardiolipin is a phospholipid required to properly anchor proteins to the IMM, including components of the ETC, and to stabilize mitochondrial cristae [73]. Cardiolipin relocalizes to the OMM in response to stresses, such as mitochondrial membrane depolarization and respiratory chain inhibition, where it binds directly to processed LC3B to promote mitophagy [74]. LC3-dependent mitophagy was attenuated by blocking the LC3B–cardiolipin interaction, knocking down cardiolipin synthase, or preventing transport of cardiolipin to the OMM [74]. LC3 interacts with mature tetralinoleoyl–cardiolipin that is dependent on processing from immature monolyso-cardiolipin by the Tafazzin (TAZ) phospholipid trans-acylase, an enzyme found mutated in patients with Barth syndrome. TAZ deficiency prevents cardiolipin processing and blocks mitophagy leading to mitochondrial dysfunction (reduced respiration and ROS production), cardiomyopathy, and other pathologies found in Barth syndrome patients [75].
Ceramide is a bioactive sphingolipid known to induce caspase-independent cell death that may be attributed to a lethal form of mitophagy [76]. Ceramide C18 induced expression of LC3B-II, co-localized with LC3B-II at mitochondria, and caused recruitment of autophagosomes to mitochondria. The interaction of ceramide with LC3B-II relied on LC3 lipidation, since ceramide did not interact with LC3B-I. Over-expression of ceramide synthase-1 (CS1), but not catalytically dead CS1 (H183A), induced LC3B-II, LC3 puncta and mitophagy [76]. While ceramide-induced mitophagy required DRP1, it did not require p62/Sqstm1 or BNIP3L [76]. Interestingly, serine deprivation of tumor cells caused a marked decrease in ceramide, as well as lower levels of sphingolipids, cardiolipin, and phenylethanolamine, and was associated with mitochondrial fragmentation, mitochondrial dysfunction, and reduced cell proliferation [77]. These defects were rescued by providing ceramide (C16) to cells, although the effects of ceramide on mitophagy were not examined in this study.
Another lipid-dependent modulator of mitophagy is Lipin-1, a phosphatidic acid phosphatase that catalyzes the conversion of phosphatidic acid (PA) to diacylglycerol (DAG), which is required for de novo synthesis of phospholipids and triglycerides [78, 79]. Patients with heritable Lipin-1 mutations develop muscle atrophy that is associated with mitochondrial dysfunction and lipid droplet accumulation due to a failure in autophagy [80]. Muscle-specific over-expression of a Lipin-1 transgene rescued these defects in mice and also ameliorated the myopathy that develops following statin treatment [80]. Ceramide and other phospholipids accumulated in Lipin-1-deficient muscle following statin treatment but the extent to which this contributed to defects in mitophagy was not examined. Lipin-1 is required for activation of Protein Kinase D (PKD) and VPS34 to promote autophagosome fusion with lysosomes [80], but recent work also shows that Lipin-1 is required for the interaction of BNIP3 with LC3 in muscle [81]. How the role of Lipin-1 in mitophagy intersects with its role in control of nuclear transcription factor activity or in lipid signaling pathways is not clear.
Sphingosine kinase 1 (SPHK-1) catalyzes the formation of sphingosine-1-phosphate (S1P) from sphingosine, and while sphingosine (and ceramide) promote cellular senescence or apoptosis, S1P stimulates tumor cell proliferation and migration [82, 83]. SPHK1 is hypoxia-inducible [82, 84] and, as mentioned above, promotes mitophagy during erythropoiesis in part by inducing BNIP3L expression [51]. SPHK1 also activates the UPRmt in response to respiratory chain inhibition, inhibition of mitochondrial protein import or inhibition of mitochondrial fission or fusion [85]. SPHK1 localizes to mitochondria in response to mitochondrial stress and production of S1P regulates a number of mitochondrial proteins involved in the UPRmt to promote mitochondrial functionality [85]. In particular, S1P interacts directly with PHB-2 to promote ETC complex formation, suggesting that PHB-2 (and BNIP3L) may act downstream of lipid signaling to promote mitochondrial function and mitophagy, but much remains to be investigated in this area to understand how signaling lipids regulate mitophagy.
Upstream sensing of mitochondrial stress to induce mitophagy
Mitochondria are subject to many different kinds of stresses, including oxidative damage caused by respiratory chain inhibition, mutation of the mitochondrial genome that can repress respiration, failure to properly uptake mitochondrial proteins that then accumulate in the cytosol and cause an imbalance in mitochondrial to nuclear encoded proteins, defects in mitochondrial translation such as occur in response to certain antibiotics, as well as stresses shown to induce mitophagy, such as low oxygen and membrane depolarization [53]. As discussed above, much is known about how the cell promotes mitophagy in response to membrane depolarization (PINK1 accumulation) or hypoxia (induction of BNIP3/BNIP3L). Here, we examine several signaling pathways that have been linked to induction of mitophagy downstream of mitochondrial stresses (Fig. 3).
Mitochondrial stress-sensing pathways that modulate mitophagy. Mitochondrial stress and dysfunction is sensed in various ways and feeds forward to activate mitophagy, and other stress response pathways, such as the UPRmt. a Excess ROS produced following inhibition of the electron transport chain (ETC) results in escape of electrons (e−) primarily from complex I and complex III, that react with oxygen and water to form superoxide and peroxides that may be quenched by mitochondrial anti-oxidants or escape to the matrix and cytosol. ROS is also induced in tumor cells by matrix detachment and by mitophagy defects that cause dysfunctional mitochondria to accumulate. ROS activates transcription factors, including HIF-1, FoxOs, NRF2, that induce stress responses, including BNIP3-dependent mitophagy and NRF2 anti-oxidant responses. b While ATF4 is a key component of the cellular integrated stress response, and is activated by amino acid deprivation, it also plays a unique role in sensing mitochondrial stress that is mediated via HRI (heme-regulated eIF2α kinase) that is activated by a cleaved cytosolic form of DELE1, released from damaged mitochondria following OMA1 cleavage. ATF4 induces genes involved in amino acid biosynthesis and transport, but also induces genes that promote autophagy, OXPHOS and the UPRmt. c NAD+ is depleted by mitochondrial dysfunction, particularly in response to defects in respiration and lower complex I activity. NAD+ levels in cells are further decreased by DNA damage although the extent to which the nuclear/cytosolic pools of NAD + affect the mitochondrial pools and vice versa is unclear. Decreased NAD+ limits Sirtuin activity in the nucleus and at the mitochondria, resulting in lower FoxO activity and reduced mitophagy, which can be rescued by replenishing NAD+ with NR or by PARP inhibition. d Decreased ATP levels arising from lower respiratory chain activity and increased AMP activates AMPK, that can promote mitophagy by activitating FIS1 and MFF, or by activating the TFEB/TFE3 transcription factors, amongst other functions. e Mitophagy plays a key role in preventing activation of the cGAS-STING and NRLP3 inflammasome in response to release of mtDNA from damaged mitochondria during innate immune responses and inflammation
Activation of mitophagy by AMPK
AMPK is activated by defects in respiration caused by agents, such as Metformin [86, 87], that can lead to induction of general autophagy via ULK1 activation [60]. AMPK promotes mitophagy specifically via phosphorylation of the FIS1 and MFF receptors for DRP1 [88, 89] and AMPK phosphorylation of glycolysis regulator PFKFB3 promoted mitophagy and survival following extended mitotic arrest [90]. AMPK also activates TFEB and TFE3 in response to glucose deprivation to switch on the CLEAR network of genes that includes many autophagy genes [91]. Inactivation of AMPK caused unprocessed autophagosomes to accumulate in lung tumors driven by Ras and mutant p53 [91]. Loss of LKB1, the regulatory kinase upstream of AMPK [87], also caused mitochondrial dysfunction, including decreased oxygen consumption, increased ROS, and altered mitochondrial metabolism [92,93,94,95], but it is unclear if this was due to a failure to activate mitophagy [96, 97]. Nevertheless, the LKB1–AMPK signaling pathway has emerged as a key sensing mechanism of mitochondrial dysfunction that activates mitophagy.
ATF4 and the mitochondrial stress response
To obtain unbiased insight to how mitochondria respond to different kinds of stress, including ΔΨmt, mitochondrial protein translation inhibition, or respiratory chain inhibition, Quiros and colleagues performed transcriptomic, proteomic, and metabolomics profiling of HeLa cells exposed to agents that induce these stresses [98]. Genes involved in amino acid biosynthesis and handling, including ASNS and ASS1, correlated most strongly with mitochondrial stress and many of the induced genes were known ATF4 targets [98]. Consistently, levels of many amino acids were also elevated in response to mitochondrial stress. This ATF4 transcriptional signature was also upregulated in HEK-293T cells depleted for mtDNA or treated with oligomycin to repress respiration, where serine biosynthetic enzymes (PSAT1, PHGDH etc.) in particular were upregulated [99].
The ATF4 signature was also induced in muscle following OPA1 deletion where loss of OPA1 caused small, disrupted mitochondria to accumulate [100]. Interestingly, BNIP3 and LC3 were induced in Opa1 null muscle and deletion of FoxO (known to induce BNIP3) rescued atrophy of Opa1 null muscle fibers. The ATF4 signature has also been seen in other myopathy models [101, 102] and deletion of ATF4 protects muscle from fasting induced atrophy, where again BNIP3 and LC3 were induced [103]. These findings suggest that mitochondrial stress is a major factor in muscle atrophy and myopathies.
Recent work has shed light on how ATF4 is activated by mitochondrial stress (such as treatment with oligomycin and inhibition of respiration) [104, 105]. Mitochondrial stress activates the OMA1 protease to cleave DELE1 at the mitochondria. DELE1 was previously identified in a yeast two-hybrid system as a DAP3-interacting protein involved in modulating rates of apoptosis [106], but until now, little else has been reported about DELE1. Cleavage of DELE1 by OMA1 releases the short form of DELE1 to the cytosol where it interacts directly with HRI (Heme Regulated eIF2a kinase), one of four kinases known to regulate ATF4 activity [104, 105]. Knockdown of DELE1 blocked ATF4 induction by mitochondrial stress, but was not required for ATF4 activation by ER stress [104].
ATF4 has been interrogated as a functional homolog in mammalian systems of Atfs-1, the master regulator of the UPRmt in worms [107, 108]. While ATF4 has not been shown to localize to mitochondria like Atfs-1, both ATF4 and Atfs-1 are induced in response to inhibition of mitochondrial protein translation [109]. Atfs-1 accumulates in the nucleus of cells in response to mitochondrial stresses that block its import to the mitochondrial matrix, where it would be degraded under non-stressed conditions [110]. Atfs-1 target genes include mitochondrial chaperones that are activated by Atfs-1 and components of the ETC that are repressed by Atfs-1 [111]. Atfs-1 was also required to activate the UPRmt in a worm model of Alzheimers Disease (AD) in which amyloid-β was over-expressed [108]. In addition to activating the UPRmt in this model, Atfs-1 was also required for mitophagy via induction of dct-1, the worm homolog of BNIP3/BNIP3L [112]. In a mouse model of AD, mitophagy, UPRmt and OXPHOS gene sets (Mitophagy: Map1lc3b/Lc3b, Bnip3, Sqstm1/p62, Pink1/Park6; UPRmt: Hspa9, Hsp60, Yme1L1, Lonp1; OXPHOS: Cox5a, Cox2, Nd1, Sdhc) were induced [108], although the extent to which their expression was dependent on ATF4 was not examined. The simultaneous induction of the UPRmt and mitophagy in multiple systems in response to mitochondrial stress suggests joint regulation of these survival mechanisms, but there is much more to uncover about how these responses are coordinated.
NAD+ promotes mitophagy
Unresolved DNA damage occurring in cells mutated for enzymes involved in DNA repair, such as the ATM kinase in patients with Ataxia telangiectasia (AT), depletes NAD+ due to constitutive activity of the DNA repair protein Poly-ADP Ribose Polymerase (PARP) that consumes NAD+ [113]. Deficiency for atm1 in worms caused neuronal defects that were rescued by replenishing NAD+ levels with nicotinamide riboside (NR) or via PARP inhibition (Olaprarib/AZD2281) that promoted expression of dct-1, mitophagy, and mitochondrial function [114]. Boosting NAD+ levels also induced mitophagy and the UPRmt in a worm model of AD where amyloid-β was over-expressed, leading to mitochondrial proteostasis that both resolved degenerative disease and extended worm lifespan [108]. Dct-1, sqstm1 and pink1 were among the mitophagy genes induced in response to NR where their induction was dependent on Atfs-1, as described above [108].
Depletion of nuclear NAD+ through elevated PARP activity inhibits NAD+-dependent Sirtuins [113, 115]. Sirtuins modulate many metabolic processes in the cell, including via NAD+-dependent de-acetylation of PGC-1α required for expression of genes involved in mitochondrial biogenesis, fatty acid oxidation, and oxidative phosphorylation [116]. Sirtuins also de-acetylate FoxO3a that is a known inducer of autophagy genes, including BNIP3, LC3B, and Beclin1 [35]. The worm homolog of FoxOs, daf-16, is also activated by sirtuins resulting in dct-1 upregulation that limited aging and extended worm lifespan [112, 114]. Thus, both Atfs-1 and daf-16 induce mitophagy genes in worms in response to nutrient and mitochondrial stress, in a manner modulated by NAD+. Further characterization of these pathways linking NAD+ availability and mitophagy is necessary in mammalian systems [35, 117,118,119].
Activation of mitophagy in response to cytosolic mtDNA during inflammatory responses
Release of mtDNA to the cytosol following mitochondrial damage or dysfunction activates the cGAS–STING pathway and the NLRP3 inflammasome, and thus by eliminating dysfunctional mitochondria, mitophagy limits activation of cGAS–STING and inflammasomes [120, 121]. During apoptosis, efflux of mtDNA from the mitochondria occurs via IMM balloons that herniate out through BAX/BAK macropores in the OMM [122], but how mtDNA escapes the mitochondria during innate immune responses is not clear, although unlike apoptosis, cytochrome c is not released [123]. Newly synthesized mtDNA not yet packaged into Tfam-positive nucleoids may be most susceptible to oxidation, cleavage and release to activate either cGAS–STING or the NLRP3 inflammasome [124].
Oxidized mtDNA binds directly to the NLRP3 inflammasome, inducing pro-caspase-1 activation at the mitochondria and stimulating IL-1β release [125,126,127]. Mitochondrial caspase-1 activity causes further mitochondrial damage, including mitochondrial membrane depolarization that induces Parkin-dependent mitophagy [128], although since Parkin is cleaved by caspase-1, mitophagy may be limited depending on the scale of mitochondrial damage and onset of pyroptosis [129].
Activation of cGAS-STING by cytosolic mtDNA induces a Type I interferon response mediated by TANK-binding kinase-1 (TBK1) and IRF3 [121]. Amongst other activities, TBK1 promotes mitophagy by recruiting cargo receptors, such as OPTN to the OMM of damaged mitochondria [130, 131]. STING trafficking also promotes autophagy independent of TBK1 by stimulating autophagosome biogenesis at the ER-Golgi [132]. cGAS–STING and NLRP3 pathways communicate with each other during inflammatory responses, via effects of STING on release of potassium from the lysosome that activates NLRP3 [133]. Thus, by eliminating damaged mitochondria, mitophagy prevents further mtDNA release and attenuates activation of both cGAS–STING and the NLRP3 inflammasome [120, 121, 129, 134].
Mitophagy in cancer
PINK1/Parkin-dependent mitophagy in cancer
PINK1 and Parkin are encoded by genetic loci (PARK6 and PARK2, respectively) that are mutated in human Parkinson’s disease (PD), and also disrupted in various human cancers [135]. The PARK2 locus at 6q25-q26 is frequently deleted in bladder, breast, lung, ovarian, and other cancers [136], while PARK2 mutations have been linked to glioblastoma, colon cancer, and lung cancer [137]. Similarly, PARK6 expression (PINK1) is down-regulated in glioblastoma and ovarian cancer, with rare mutations found in neuroblastoma [138]. Analysis of tumor phenotypes in mouse models of cancer showed Parkin null mice to be susceptible to spontaneous hepatocellular carcinoma (HCC) and sensitized to irradiation-induced lymphomagenesis [139, 140]. Loss of either Parkin or PINK1 increased tumor burden and metastasis in KRas-driven PDAC [45, 141]. Parkin also suppressed the growth and metastatic progression of breast cancer cells [44]. Taken together, these findings indicate that Parkin and PINK1 play tumor suppressor functions in both mouse and human.
Given their canonical function in mitophagy, the tumor suppressor functions of Parkin and PINK1 have largely been attributed to their role in maintaining mitochondrial function and metabolic homeostasis by clearing depolarized mitochondria to prevent Warburg metabolism and excess ROS [45, 140, 142]. However, Parkin and PINK1 also suppress HIF-1α stabilization [44, 45, 138]; Parkin ubiquitinates HIF-1α on K477 to promote its degradation and low Parkin levels correlated with high HIF-1α levels and reduced metastasis-free survival in human breast cancer [44]. Deletion of HIF-1α (or iron chelation) rescued elevated glycolysis and ROS detected in PDAC following deletion of Parkin or PINK1 [45]. These observations indicate that Parkin/PINK1 may suppress tumorigenesis in part by promoting HIF-1α turnover (Fig. 4).
PINK1/Parkin functions relevant to tumor cell growth and tumorigenesis. In addition to their canonical role in promoting mitophagy of depolarized mitochondria, Pink1/Parkin also promote turnover of the HIF-1α transcriptional sub-unit that is known to promote tumorigenesis via induction of HIF target genes that enhance glycolysis, angiogenesis, and metastasis. Loss of Parkin or Pink1 could thus promote tumorigenesis as a result of increased HIF-1α levels. Furthermore, Parkin Pink1/Parkin-induced mitophagy modulates cell cycle checkpoints by sequestering TBK1 away from centrosomes where it promotes spindle assembly and mitosis. Thus increased mitophagy can inhibit cell cycle progression. Parkin also has Pink1-independent roles in regulating cell cycle by promoting Cdc20 and Cdh1 activity
Mitochondrial damage blocked G2-to-M-phase cell cycle transition in a PINK1/Parkin-dependent manner, while loss of Parkin or PINK1 increased rates of cell division [143]. Parkin’s E3 Ub ligase activity modulates cyclin levels and the mitotic checkpoint independently of the Anaphase-Promoting Complex (APC) and its E3 Ub ligase activity, via direct interaction of Parkin with Cdc20 and Cdh1 [144, 145]. This mitotic function for Parkin was independent of PINK1 and required activation of Parkin via phosphorylation on S378 by Polo-like kinase-1 [145]. Thus, Parkin has dual roles in mitophagy and in control of cell cycle. More recently, PINK1-dependent Parkin activity has been reported to modulate mitosis indirectly by sequestering TBK1 at damaged mitochondria [143]. In addition to its role in recruiting OPTN and other cargo receptors to damaged mitochondria [130, 131], TBK1 is a centrosome-associated protein required for microtubule dynamics and mitosis [146]. By sequestering TBK1 at damaged mitochondria and away from centrosomes, PINK1/Parkin activity caused a cell cycle arrest and failure to enter mitosis [143]. Artificially tethering TBK1 to mitochondria blocked cell cycle progression into mitosis, while deletion of Parkin rescued cell cycle progression defects in Drosophila lacking ik2, the fly homoplog of TBK1 [143]. Thus, negative control of cell cycle and de-stabilization of HIF-1α are added aspects to the tumor suppressor functions of PINK1/Parkin, although, arguably, these functions are secondary to their function in eliminating dysfunctional mitochondria (Fig. 4). How the PINK1-independent function of Parkin in promoting anaphase transition via interaction with Cdc20 [145] is coordinated with its PINK1-dependent function in blocking mitosis via TBK1 sequestration [143] may depend on whether mitochondria are functional or not.
There is a growing appreciation that E3 Ub ligases, other than Parkin, play a role in mitophagy and can substitute for Parkin [147]. For example, the MUL1 E3 ubiquitin ligase can compensate for Parkin loss in Drosophila and is required for elimination of paternal mitochondria during embryogenesis [14, 15]. While Parkin is poorly expressed in most cancers and cancer cell lines, ARIH1 is an E3 ubiquitin ligase recruited by PINK1 to mitochondria to promote mitophagy in response to chemotherapeutic agents and in doing so protects cancer cells and causes drug resistance [16]. ARIH1 thus represents a valid therapeutic target. Interestingly, while ARIH1 functions similarly to Parkin in requiring PINK1 and ubiquitinating mitochondrial proteins, its substrates at the mitochondria are different, but remain uncharacterized [147]. The extent to which other mitochondrial E3 Ub ligases perform Parkin-like functions in mitophagy that are relevant to cancer is an area of growing interest.
BNIP3/BNIP3L in cancer
BNIP3 protein levels are induced at early pre-malignant stages of various human solid cancers as tumors become hypoxic, including breast cancer and pancreatic cancer, but frequently down-regulated as these tumors progress to becoming invasive [148,149,150]. Hyper-methylation of the BNIP3 promoter is a common mechanism of BNIP3 inactivation in human cancers, including in gastric, pancreatic, liver, lung, and hematological malignancies [149, 151]. Epigenetic silencing of BNIP3 in human pancreatic cancer was linked to chemoresistance and poor prognosis [150, 152], consistent with tumor-suppressive functions for BNIP3. In contrast to BNIP3, elevated BNIP3L expression in a cohort of PDAC patients was linked to worse prognosis and loss of BNIP3L in the KPC model of PDAC (LSL-KRasG12D; p53R172H; Pdx1-Cre) delayed tumor progression from the PanIN stage to malignant PDAC [153]. These findings suggest that BNIP3 and BNIP3L may play different roles in PDAC, possibly due to their expression in different cell types or at different stages of disease progression, although this requires further analysis.
The BNIP3 locus at the end of chromosome 10 (chr.10q26.3) is also subject to deletion in a number of cancers including in breast and prostate cancer [154]. BNIP3 is most frequently deleted in triple-negative breast cancer (TNBC) [155, 156]. TNBC is the sub-type of breast cancer that exhibits the strongest HIF signature and Warburg metabolism [157] and loss of BNIP3 combined with high HIF-1 expression predicted poor metastasis-free survival specifically for TNBC patients [156]. TNBC is also the breast cancer sub-type with the highest degree of mitochondrial dysfunction, possibly due to loss of BNIP3 and defective mitophagy, and this could potentially be exploited therapeutically to target mitochondrial function in TNBC and bring a novel treatment option to this deadly form of breast cancer [158, 159]. Supporting data from human studies, BNip3 knockdown promoted tumor growth and metastasis in the 4T1 orthotopic model of mammary tumorigenesis [160]. Similarly, loss of BNip3 in the MMTV-PyMT mouse model of breast cancer increased tumor burden and accelerated progression to invasive carcinoma and lung metastasis [156]. Increased tumor burden and invasiveness of BNip3 null mammary tumors were associated with reduced mitophagy, increased ROS, and elevated HIF-1 levels, contributing to increased glycolysis and angiogenesis [156]. Quenching ROS attenuated HIF-1 levels and reduced metastasis in the BNip3 null mice, indicating that BNIP3 suppressed tumorigenesis in this model by reducing mitochondrial mass and limiting ROS production and explaining why there is selective pressure to inactivate BNIP3 downstream of HIF-1, as tumors progress to becoming invasive [156].
BNIP3 has also been shown in other tumor types to promote tumor cell survival following detachment from extracellular matrix (ECM) and high BNIP3 correlated in these primary human cancers (kidney, sarcoma) with worse patient outcomes [161]. These dichotomies may be explained by differential roles for BNIP3 depending on tumor type or in different cell types within a tumor (cancer stem cell versus non-stem cell, for example). As we have discussed previously, mitophagy plays an important role in maintaining the cancer stem cell phenotype [162]. For example, mitophagy limited p53 levels in HCC stem cells via PINK1-mediated phosphorylation of mitochondrial p53 that gets turned over by mitophagy resulting in de-repression of the stem cell factor, Nanog [163]. Mitophagy also enhances stemness by promoting glycolysis and limiting oxidative metabolism, thus engendering the low energy state required for stemness [162] and is required for segregation of young mitochondria to daughter stem cells that is required to maintain stem cell fitness [164]. Consistently, autophagy reduced replication stress and aging in hematopoietic stem cells [165,166,167]. Specifically, stimulating mitophagy with nicotinamide riboside (NR) that increases the cellular NAD+/NADH ratio and stimulates Sirtuin activity promoted hematopoietic stem cell self-renewal [168]. The extent to which BNIP3/BNIP3L or other mitophagy pathways promote cancer stemness remains an open area of research.
FUNDC1 in tumor cell proliferation and migration/invasion
The interaction of FUNDC1 with LC3 is inhibited by SRC-mediated phosphorylation on Y18 within the LIR motif of FUNDC1 [62], suggesting that FUNDC1-dependent mitophagy is antagonistic to SRC-driven effects on tumor cell migration and invasion. FUNDC1 also recently emerged from an shRNA screen for genes that modulate tumor cell migration and invasion in response to mitochondrial proteotoxic stress [169]. Silencing FUNDC1 increased rates of focal adhesion assembly and disassembly, and increased cellular motility and invasion in a DRP1-dependent manner through Matrigel of both prostate and glioblastoma cell lines [169]. Conversely, over-expressing FUNDC1 limited cell migration and invasion. The increased migration and invasion caused by decreased FUNDC1 expression was associated with increased liver metastasis in vivo in xenograft models, but conversely also resulted in decreased tumor cell proliferation. RNA-Seq analysis of 33 different tumor types in the TCGA database showed increased FUNDC1 levels to be associated with high levels of expression of genes involved in mitochondrial bioenergetics, while low levels of FUNDC1 were associated with increased ROS signaling and metastasis, particularly in lung cancer. These findings are at odds with previous work in which elevated FUNDC1 in human breast cancer was associated with increased tumor cell proliferation, migration, and invasion and with worse prognosis for breast cancer patients [170], and additional work will be needed to explain these divergent findings.
Nevertheless, it has been suggested that the role of FUNDC1 in tumor cell proliferation and motility is mitophagy-independent and reflects a more significant role for FUNDC1 in mitochondrial bioenergetics [169]. Knockdown of FUNDC1 decreased levels of TCA cycle intermediates, lowered respiration rates, and increased ROS production in tumor cells that was associated with increased degradation of electron transport chain complexes. This was rescued by exogenous expression of the LonP protease that is known to maintain the integrity of electron transport chain complexes and was misfolded as a result of FUNDC1 loss, possibly related to its role in resolving mitochondrial proteostatic stress via interaction with HSC70 [66]. Interestingly, exogenous LonP also rescued the effect of FUNDC1 loss on tumor cell proliferation and invasion, suggesting that FUNDC1 modulates tumor cell behavior independent of its function in mitophagy [169].
Mitophagy defects in XP, AT, and FA cancer predisposition syndromes
Mitophagy defects have emerged as a feature of the cancer predisposition syndromes Xeroderma pigmentosum (XP), Ataxia telangiectasia (AT), and Fanconi Anemia (FA) that arise due to defects in nucleotide excision repair (XP) and double-strand break repair (AT, FA) [114, 115, 171,172,173,174].
Work from the lab of Beth Levine identified FANC-C and other Fanconi complex components (FANC-A, FANC-D2, FANC-F, FANC-L, BRCA1, and BRCA2) as required for Parkin-mediated mitophagy as well as virophagy of Sindbis virus [174, 175]. FANC-C co-localized with Parkin at depolarized mitochondria and loss of FANC-C caused dysfunctional mitochondria to accumulate and elevated mitochondrial ROS production, as a result of defective mitophagy. This role of FANC-C in mitophagy was genetically separable from its function in DNA repair [174]. Possibly due to the defect in mitophagy, there was also increased inflammasome activation in FA cells that, along with elevated ROS levels, contributed to bone marrow failure and increased incidence of hematological pathologies in FA individuals, while defective virophagy could explain why FA patients are more prone to infection than others [174]. The FANC-C c.67delG mutation is common in Dutch FA patients and is associated with a milder form of the disease, including older age at diagnosis, lower rate of bone marrow failure, and reduced incidence of malignancy [176]. Intriguingly, this mutant form of FANC-C retained its mitophagy function, but had lost its ability to promote DNA repair, indicating that the loss of mitophagy does contribute significantly to disease pathology [174].
Loss of ATM kinase increased mitochondrial ROS and caused mtDNA depletion [177,178,179] and neurons lacking ATM exhibited reduced mitophagy [114]. Replenishing NAD+ levels with NR or via PARP inhibition promoted mitophagy, rescued cell death, and stimulated neuronal differentiation of ATM-deficient neurons [114]. Similarly, PARP inhibition rescued mitophagy and neurodegeneration in XPA patients [115]. Boosting NAD+ levels also induced mitophagy in long-term repopulating hematopoietic stem cells and promoted long-term hematopoiesis in mice fed NR, consistent with a role for mitophagy in promoting stemness and longevity [162, 168]. The extent to which NAD+ depletion contributes to the mitophagy defect in FA and whether it also can be rescued by NR or PARP inhibition has not been examined.
From a therapeutic angle, inducing mitophagy with NR or PARP inhibition could be beneficial if it prevents accumulation of DNA mutations arising from mitochondrial dysfunction, but could be detrimental if it promotes tumor cell survival. For example, DNA-damaging agents by depleting NAD+ would be predicted to cause mitochondrial defects and loss of survival if mitophagy gene induction is inhibited. Mitophagy induced via replenishment of NAD+ (via NR or PARP inhibition) may thus limit the efficacy of genotoxic agents, such as cisplatin, or 5-fluorouracil.
Intersection of mitophagy with mitochondrial dynamics
Mitochondrial fission is required for mitophagy in response to certain stresses [180,181,182] and knocking out DRP1 in the liver resulted in unopposed mitochondrial fusion and caused p62-positive mitophagy intermediates to accumulate [182]. This was rescued by liver-specific knockout of OPA1 [182], consistent with previous work, showing that healthy mitochondria are protected from turnover during nutrient deprivation by OPA1-dependent fusion [180, 181].
Mitophagy may require mitochondrial fission due to the limiting size constraints on autophagosomal cargo that can be engulfed successfully, and since mitochondrial fusion requires membrane polarization, this would also selectively target depolarized mitochondria for turnover [180, 181]. Indeed, DRP1 is activated by loss of MMP (ΔΨmt) and recruited to the OMM via the MFF receptor [183, 184].
While FIS1 is now considered redundant for recruitment of DRP1 to mitochondria [185], FIS1 is required for PINK1 recruitment to mitochondria, acts downstream of Parkin to promote mitophagy and FIS1-dependent mitophagy in leukemia stem cells (LSCs) promoted responses to chemotherapy [89]. FIS1 promotes mitophagy mainly via recruitment of Rab-GAPs TBC1D15 and TBC1D17 that dimerize to modulate Rab7 GTPase activity and also interact with LC3/GABARAP family members ensuring the complete encapsulation of mitochondria by autophagosomes [186]. In this way, FIS1 is less of a fission factor and more a direct mitophagy regulator. Indeed, mutation of FIS1 led to accumulation of large LC3-positive aggregates [187].
At the OMM, DRP1 interacts with ZIP1 to modulate MMP via uptake of Zn2+ through the mitochondrial calcium uniporter (MCU) [188]. DRP1 interacts with ZIP1 at sites of mitochondrial fission, although GTPase-deficient DRP1 retains ability to interact with ZIP1 and to induce ΔΨmt, indicating two separable functions for DRP1 in mitophagy [188]. The DRP1–ZIP1 interaction was required for Parkin recruitment to mitochondria in response to hyperglycemia, and chelating Zn2+ or blocking the DRP1–ZIP1 interaction inhibited Parkin recruitment and mitophagy [188]. As a result, fragmented mitochondria with reduced MMP accumulated as “damaged” mitochondria that could not be turned over by mitophagy. BNIP3 and FUNDC1 also interact with DRP1 and may function as alternative receptors for DRP1, since DRP1 is required for hypoxia-induced mitophagy [65, 189]. These findings highlight how control of mitochondrial dynamics is coordinated to promote mitophagy.
Mitochondrial fission and DRP1 phosphorylation on S616 by ERK2 are required for oncogenic RAS transformation [190, 191]. DRP1-dependent fission was also required for B-RAFV600E transformation in melanoma [192, 193]. Inhibition of DRP1 phosphorylation on S616 induced mitochondrial fusion and blocked tumor cell growth in xenografts, while pancreas-specific deletion of Drp1 inhibited progression from PanIN to pancreatic ductal adenocarcinoma (PDAC) in KPC mice [190, 194]. At this time, it is not understood why mitochondrial fission is required for Ras transformation, but this is clearly of major interest, since ERK inhibitors are used therapeutically against Ras-driven cancers, including human PDAC [195]. Intriguingly, inhibition of Ras or ERK1/2 inhibition induces autophagy that was associated with reduced mitochondrial mass [196], and thus, one could speculate that mitochondrial fission is required for Ras transformation to promote mitophagy and mitochondrial homeostasis. Ras transformation induces expression of BNIP3 and BNIP3L [153, 197] and loss of BNip3L attenuates PDAC formation in KPC mice [153], but studies to demonstrate a requirement for BNIP3/BNIP3L downstream of DRP1 in Ras transformation have not been performed. The combination of ERK1/2 inhibition with autophagy inhibition was shown to be strikingly effective at blocking tumor cell growth in vitro [195, 196], and in patients where the combination of Trametinib with hydroxychloroquine significantly reduced PDAC tumor burden [195]; these studies are now being expanded in clinical trials (NCT03825289).
Mitophagy, ROS, and metastasis
Mitochondrial dysfunction has been linked to metastasis most frequently as a result of increased ROS production, for example arising due to mtDNA mutation [198] or ETC inhibition [199]. Indeed, mitochondrial ROS generation following recovery of host mtDNA in mitochondria-depleted tumor cells was required for metastatic progression in both the 4T1 and B16-F10 orthotopic mouse models of metastasis [200, 201]. Increased ROS has been shown to drive the progression to metastasis in a number of other tumor models also, including in PDAC where loss of TIGAR promoted ROS-driven EMT, invasiveness, and metastasis to the lungs, which was mitigated by treatment with N-acetyl cysteine (NAC) [202]. ROS drives metastasis in part by activating HIF-1 and NF-κB to induce EMT and tumor cell invasion, respectively [203, 204]. ROS also inactivates protein phosphatases and activates key signaling kinases, such as ERK, JNK, and p38 that stimulate various pro-metastatic pathways [205, 206]. However, it has also been argued that ROS limits metastasis in melanoma where circulating tumor cells showed higher levels of oxidative stress than primary tumor cells, and treatment of mice with NAC increased metastasis without affecting primary tumor burden, suggesting that many circulating metastatic cells were not surviving in the circulation due to excess ROS [207, 208]. Successfully metastasizing tumor cells had upregulated pathways (such as folate metabolism to produce more NADPH) to manage ROS and survive [207]. Taken together, this suggests that while ROS is required for metastasis, too much ROS is deleterious to the disseminated tumor cell and limits metastasis (Fig. 5).
ROS, mitophagy, and metastasis. Extensive evidence shows that ROS is required for tumor cell migration and metastasis. However, other data shows that excess ROS can limit metastasis, which has lead to a model in which levels of ROS need to be maintained within a range, the so-called hormetic zone, to promote metastasis. ROS induces mitohormetic responses, including the UPRmt that promote mitochondrial fitness and limit ROS allowing tumor cells to survive and progress to metastasis. By limiting ROS, mitophagy may inhibit these adaptive responses and work suggests there is selection for BNIP3 loss during progression to invasiveness in breast cancer. However, the ensuing higher levels of ROS that occur once BNIP3 or other mitophagy genes are lost, would not be expected to restrict metastasis due to upregulation of other components of the UPRmt. Selection for tumor cells that survive, possibly by upregulating stress response pathways that manage ROS, such as the NRF2 anti-oxidant response, or ATF4 that promotes amino acid biosynthesis as part of the UPRmt, would promote cell survival and be permissive for metastasis again
This is consistent with data, showing that mitochondrial ROS needs to be within the hormetic zone to promote metastasis [209], as shown in studies reporting how high ROS levels promote breast cancer invasiveness in vitro and in vivo, in part by inducing the UPRmt [210]. ROS-induced UPRmt consisted of a transcriptional response involving CHOP, ATF5, NRF2, FOXO3A, and SIRT3, which activated genes that restored mitochondrial function [209]. Kenny et al. showed that the UPRmt is upregulated in metastatic tumors relative to primary tumors, consistent with the UPRmt conferring a metastatic advantage and high UPRmt in primary human cancer correlated with poor prognosis [210]. Thus, ROS at non-lethal levels may promote mitochondrial fitness, tumor cell viability, and metastasis in part by activating the UPRmt.
Mitophagy genes are induced as part of the UPRmt in response to ROS and nutrient deprivation, including induction of p62/Sqstm1 by NRF2 and BNIP3 by FOXO3A. As discussed above, loss of BNip3 promoted more rapid progression to metastasis in both the autochtonous MMTV-PyMT mammary tumor model and in the 4T1 orthotopic model of breast cancer [156, 160]. Significantly, mammary tumors forming in MMTV-PyMT;BNip3−/− mice had higher mitochondrial mass and higher ROS levels than wild-type control tumors, and expressed elevated nuclear HIF-1 and HIF target genes [156]. Treatment of MMTV-PyMT mice with anti-oxidant attenuated mammary tumor metastasis to the lungs in BNip3 null mice to that seen in wild-type mice, without affecting primary tumor growth [156]. This work indicated that BNIP3 was suppressing metastasis by promoting mitochondrial turnover and limiting ROS that would otherwise cause HIF-1α stabilization [211].
However, BNIP3 and BNIP3L are induced in tumor cells following detachment from the extracellular matrix (ECM) [212] and required for mitophagy and tumor cell survival following detachment [161] suggesting a pro-metastatic function for BNIP3 and BNIP3L. The induction of BNIP3 and BNIP3L in response to detachment was HIF-1 dependent due to hypoxia within tumor cell clusters and associated with a switch to glycolysis and reductive carboxylation of glutamine [161]. Preventing cell clustering, knocking down HIF-1α, or knocking down BNIP3 and BNIP3L inhibited mitophagy and increased mitochondrial mass and ROS levels leading to cell death [161]. BNIP3 has also been shown to promote the migration of B16-F10 melanoma cells through effects on the actin cytoskeleton [213]. Similarly, BNIP3 is induced in the skin by hypoxia where BNIP3-dependent mitophagy is required for keratinocyte migration during wound healing [214]. It is not yet clear how mitophagy modulates the actin cytoskeleton or is otherwise modulating cell migration and this is an avenue for future investigation.
Thus, similar to the impact of ROS on metastasis, these results indicate that BNIP3 (or BNIP3L) may be pro- or anti-metastatic depending on context and whether BNIP3 (or BNIP3L) is lost acutely, or is stably deleted in systems where tumor cells are then able to adapt by upregulating other pathways that may compensate for BNIP3/BNIP3L loss, such as increasing NRF2 target gene expression to mitigate against ROS (Fig. 5). Recent work has identified NRF2 activation as a mechanism by which tumor cells survive autophagy inhibition [215, 216]. NRF2 is upregulated in autophagy-deficient cells due to elevated p62/Sqstm2 levels [215, 217] and may contribute to ATF4 induction in autophagy-deficient cells both due to depletion of amino acids but also by interacting directly with ATF4 [215]. Whether mitophagy inhibition is as effective as inhibition of general autophagy at inducing NRF2, through increased p62/Sqstm1 levels, has not been examined but if not, may indicate that specific mitophagy inhibition would be more effective than autophagy inhibition in cancer therapy.
Mitophagy and cancer cachexia
Another debilitating aspect to terminal cancer is the development of cancer cachexia that is found most commonly in lung, pancreas, and colorectal cancer patients [218, 219]. Cancer cachexia entails rapid body weight loss due to peripheral tissue wasting that negatively affects the long-term survival of cancer patients, partly due to reduced tolerance of and response to therapy [218, 220, 221]. There are many factors playing into cachexia, including release of pro-inflammatory cytokines that affect brain, liver, and immune cell function that feed into breakdown of lipid in adipose tissue and protein mass in skeletal muscle [222]. Previous work identified BNIP3 as a transcriptional target of FoxO3A during skeletal muscle atrophy [35]. It was also shown that FoxO transcriptional activity was required for muscle atrophy in C57/B6 mice implanted with Lewis lung carcinoma (LLC) and that BNIP3 mRNA was elevated in an FoxO-dependent manner in the muscle of these tumor-bearing mice [36, 37]. BNIP3 upregulation has also been detected in cachectic muscle from PDAC patients [223]. Muscle atrophy in cancer cachexia has been linked to reduced oxidative capacity and mitochondrial dysfunction in skeletal muscle [37, 224, 225]. Taken together, this may suggest that BNIP3-dependent mitophagy and lower mitochondrial mass promotes muscle atrophy during cancer cachexia by facilitating the switch away from oxidative metabolism to glycolysis. This awaits further testing in the laboratory, but is clearly an avenue of high significance given its relevance to disease and patient suffering.
Targeting mitophagy as a therapeutic approach
Inhibiting general autophagy is advancing in clinical trials for many different human cancers, primarily in combination with other drug regimens [226]. As mentioned above, autophagy inhibition (with chloroquine) in combination with Trametinib to inhibit ERK signaling showed significant efficacy in reducing tumor burden in KRas-driven PDAC [195]. Autophagy is induced by many therapeutic agents, and by promoting survival elicits drug resistance [227]. This has provided the rationale for combining autophagy inhibitors, such as chloroquine with drugs like temozolomide in the treatment of glioblastoma, where the combined therapy more than doubled patient survival times [228,229,230]. To what extent these effects are mediated via inhibition of mitophagy versus inhibition of other autophagy functions is not clear.
Functional mitochondria are required for the biosynthesis of certain amino acids, such as aspartate that is required for nucleotide biosynthesis, upon which tumor cells rely for growth, and recent studies showed that inhibiting autophagy with chloroquine synergized with agents that induce replication stress that depletes reserves of nucleotides that rely on amino acids for their synthesis [231]. Furthermore, ferroptosis is a necrosis-like cell death that is activated by oxidized mitochondrial lipids and preceded by ΔΨmt, mitochondrial fragmentation, and disrupted cristae formation, that can be prevented by activating mitophagy through over-expression of Parkin [232]. These studies suggest that inhibiting mitophagy may promote ferroptosis, although it is important to note that mitochondria were only required for ferroptosis induced by cysteine deprivation and not in response to glutathione peroxidase-4 (GPX4) inhibition. Findings such as these provide rationale for developing drugs that specifically inhibit mitophagy to achieve a more targeted effect than with targeting autophagy in general [233] (Fig. 6).
Targeting mitophagy in cancer therapy. Mitophagy is induced physiologically in response to hypoxia and nutrient deprivation or can be activated with drugs that induce mitochondrial stress (see Fig. 3), such as Metformin or PARP inhibitors. Alternatively, mitophagy can be inhibited using generic autophagy inhibitors such as chloroquine since as yet specific mitophagy inhibitors remain to be developed. Activating mitophagy promotes mitochondrial homeostasis and cell survival while inhibiting mitophagy, by causing mitochondrial dysfunction, elevated ROS production, metabolite deficiencies, including deficiency for key amino acids, such as cysteine, frequently results in cell death. Based on this rationale, inhibiting mitophagy seems more likely to elicit a clinical benefit in cancer therapy, while activating mitophagy would likely promote cell survival and therefore not be advisable as a cancer therapy, although possibly useful for treatments of myopathies where mitophagy has been shown to promote tissue homeostasis. However, it is likely not a simple as this, since cancer cells may adapt to mitophagy inhibition, for example by upregulating NRF2 signaling to allow cancer cell survival. These studies require much more extensive analysis in the lab and clinic before conclusions can be made
The use of such drugs to inhibit mitophagy would be expected to cause excess dysfunctional mitochondria to accumulate and thus would be expected to generate high levels of ROS as seen when mitophagy is inhibited genetically [156, 161]. This could promote cell killing, but could also induce progression to metastasis depending on the level of ROS generated. As we discussed previously [234], the combination of a drug to inhibit mitophagy with compounds that induce mitochondrial dysfunction and elevated ROS production, such as Metformin or Mitoquinone, may push the tumor cell further in the direction of cell death rather than adaptation [234], although activation of NRF2 in response to some of these agents [235] remains a challenge.
There is also an argument that promoting mitophagy could be beneficial. For example, there was selection for silencing of BNIP3 in drug-resistant PDAC, while re-expression of BNIP3 promoted drug sensitivity [150, 152], although the mechanistic underpinnings of this were not further pursued. The cotton seed derived compound AT-101 induces BNIP3/BNIP3L-dependent mitophagy leading to autophagic cell death in apoptosis-deficient tumor cells [236]. Mitophagy is elicited in response to AT-101 following rapid loss of membrane potential and marked induction of heme oxygenase (HMOX1) [236]. HMOX1 has been shown to induce mitophagy in other systems and the concern here would be the effect of AT-101 on normal tissues with high redox potential, such as the brain and red blood cells. Most of the drugs identified as inducing mitophagy, like AT-101, do so indirectly by depolarizing mitochondria [236] or modulating levels of ROS or NAD+ that activate NRF2 and Sirtuins, respectively [233]. For example, PMI is a Keap1 inhibitor that induces mitophagy through activation of NRF2, and thus, mitophagy is unlikely to be the only effect of such a drug [237]. Compounds that directly activate or modulate MCRs for example are lacking. However, as discussed above by maintaining mitochondrial fitness, stresses that induce mitophagy promote survival [53], which may be a useful approach in the treatment of myopathies and other diseases, but would not be a good outcome in cancer therapy. Overall, acute inhibition of mitophagy in combination with other drugs/stresses that depend on mitophagy for survival remains the approach most likely to yield efficacy in cancer treatment (Fig. 6).
Summary and future directions
Mitophagy is a core component of the mitochondrial quality control system in the cell alongside the UPRmt and other mitochondrial surveillance mechanisms that limit mitochondrial ROS production, ensure metabolic homeostasis, and prevent apoptosis. There are numerous mitophagy pathways in the cell and how each of these different pathways interact with each other, are redundant versus overlapping or are activated by oncogenic signaling specifically remains to be determined in most instances. Furthermore, how mitophagy pathways interact with the UPRmt and other mitochondrial surveillance processes also requires continued investigation. In cancer, various mitophagy pathways appear to be tumor-promoting or tumor-suppressive depending on tumor type or indeed stage of progression, or importantly, in cancer stem cells versus non-stem cells. Teasing these nuanced roles apart will require further study through use of relevant pre-clinical cancer models that target these pathways in appropriate spatial and temporal ways. The role of mitophagy in cancer cachexia is emerging as an area of significant interest given the debilitating effect of tissue wasting on a cancer patient’s long-term survival. Finally, work is on-going to determine how mitophagy might be manipulated therapeutically to better treat cancer, but again, this may depend on tumor type and stage of progression, and this is where the development and use of improved pre-clinical cancer models targeting mitophagy pathways will be essential.
Abbreviations
- AD:
-
Alzheimers disease
- ADP:
-
Adenosine diphosphate
- ALS:
-
Amyotrophic lateral sclerosis
- AMPK:
-
5′ AMP-activated protein kinase
- ANT:
-
Adenine nucleotide transporter
- ARIH1:
-
Ariadne RBR E3 ubiquitin protein ligase 1
- ASNS:
-
Asparagine synthetase
- ASS1:
-
Argininosuccinate synthase-1
- Atad3a:
-
ATPase family AAA-domain-containing protein 3 A
- ATF4:
-
Activating transcription factor 4
- ATM:
-
Ataxia telangiectasia mutated
- ATP:
-
Adenosine triphosphate
- BCL-2:
-
Breakpoint cluster locus-2
- BCL-XL:
-
BCL2-like 1 long
- BNIP3:
-
BCL2 interacting protein 3
- BNIP3L:
-
BCL2 interacting protein 3 like
- BCL2-L-13:
-
BCL2-like-13
- BRCA1:
-
Breast cancer 1
- CCCP:
-
Carbonyl cyanide 3-chlorophenylhydrazone
- CK2:
-
Casein kinase-2
- CL:
-
Cardiolipin
- DAG:
-
Diacyl glycerol
- DELE1:
-
DAP3-binding cell death enhancer 1
- DTT:
-
Dithiothreitol
- DRP1:
-
Dynamin-related protein 1
- ECM:
-
Extracellular matrix
- ERK:
-
Extracellular signal-regulated kinase
- ETC:
-
Electron transport chain
- FA:
-
Fanconi anemia
- FANC-C:
-
FA protein C
- FIS1:
-
Mitochondrial fission 1 protein
- FoxO3:
-
Forkhead box protein O3
- FKBP8:
-
FK506-binding protein 8
- FUNDC1:
-
FUN 14 domain-containing 1
- GABARAP:
-
GABA Type A Receptor-Associated Protein
- GAP:
-
GTPase activating protein
- cGAS:
-
Cyclic GMP–AMP synthase
- GPX4:
-
Glutathione peroxidase-4, GTPase activating protein
- HIF:
-
Hypoxia inducible factor
- HCC:
-
Hepatocellular carcinoma
- HRI:
-
Heme-regulated inhibitor
- IMM:
-
Inner mitochondrial membrane
- IRF3:
-
Interferon response factor 3
- LC3:
-
Microtubule-associated protein 1 light chain 3
- LIR:
-
LC3 interacting region
- LKB1:
-
Liver kinase B1
- LLC:
-
Lewis Lung carcinoma
- MARCH5:
-
Membrane-associated ring finger (C3HC4) 5
- MCR:
-
Mitochondrial cargo receptor
- MITF:
-
Melanocyte inducing transcription factor
- MFF:
-
Mitochondrial fission factor
- MFN:
-
Mitofusin
- MMP:
-
Mitochondrial processing peptidase
- MPP:
-
Mitochondrial processing peptidase
- mtDNA:
-
Mitochondrial DNA
- MUL1:
-
Mitochondrial E3 ubiquitin ligase 1
- NAC:
-
N-acetyl cysteine
- NAD + :
-
Nicotinamde adenine dinucleotide
- NBR1:
-
Near BRCA1
- NDP52:
-
Nuclear dot protein 52
- NLRP3:
-
Nucleotide binding domain and leucine rich repeat pyrin domain-containing 3
- NR:
-
Nicotinamide riboside
- NRF2:
-
Nuclear factor, erythroid 2 like
- OMM:
-
Outer mitochondrial membrane
- OPTN:
-
Optineurin
- OXPHOS:
-
Oxidative phosphorylation, p62/SQSTM1
- PA:
-
Phosphatidic acid
- PARK2:
-
Parkin encoding locus
- PARK6:
-
PINK1 encoding locus
- PARK8:
-
LRRK2 encoding locus
- PARL:
-
PINK/PGAM5-associated rhomboid-like proteases
- PARP:
-
Poly-ADP-ribose polymerase
- PD:
-
Parkinson’s disease
- PDAC:
-
Pancreatic ductal adenocarcinoma
- PFKFB3:
-
6-Phosphofructo-2-kinase/fructose-2,6-biphosphatase 3
- PGAM5:
-
Phosphoglycerate mutase family member 5
- PHB-2:
-
Prohibitin-2
- PHGDH:
-
D-3-phosphoglycerate dehydrogenase
- PINK1:
-
PTEN-induced putative kinase 1
- PMI:
-
P62-mediated mitophagy inducer
- PSAT1:
-
Phosphoserine aminotransferase 1
- RAF:
-
Rapidly accelerated fibrosarcoma oncogene
- RHEB:
-
Ras homolog enriched in brain
- ROS:
-
Reactive oxygen species
- S1P:
-
Sphingosine-1-phosphate
- SDS:
-
Sodium dodecyl sulfide
- SPHK1:
-
Sphingosine kinase 1
- SRC:
-
Rous sarcoma oncogene
- STING:
-
Stimulator of Interferon Genes
- TAX1BP1:
-
Tax1-binding protein 1
- TAZ:
-
Tafazzin
- TBK:
-
TANK-binding kinase 1
- TCA:
-
Tricarboxylic acid cycle
- TIM:
-
Translocase of inner membrane
- TFE:
-
Transcription factor E
- TMD:
-
Transmembrane domain
- TNBC:
-
Triple-negative breast cancer
- TOM:
-
Translocase of outer membrane
- ULK1:
-
Unc-51-like autophagy activating kinase 1
- UPRmt :
-
Mitochondrial unfolded protein response
- VPS34:
-
Vacuolar protein sorting 34
- XP:
-
Xeroderma pigmentosum
- ZIP1:
-
Zinc transporter-1 precursor
References
Palikaras K, Lionaki E, Tavernarakis N (2018) Mechanisms of mitophagy in cellular homeostasis, physiology and pathology. Nat Cell Biol 20(9):1013–1022. https://doi.org/10.1038/s41556-018-0176-2
Macleod KF (2020) Mitophagy and mitochondrial dysfunction in cancer. Ann Rev Cancer Biol 4:41–60
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, Sideris DP, Fogel AI, Youle RJ (2015) The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524:309–314
Harper JW, Ordureau A, Heo JM (2018) Building and decoding ubiquitin chains for mitophagy. Nat Rev Mol Cell Biol 19(2):93–108. https://doi.org/10.1038/nrm.2017.129
Geisler S, Holmström KM, Skujat D, Fiesel FC, Rothfuss OC, Kahle PJ, Springer W (2010) PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat Cell Biol 12(2):119–131
Chen Y, Dorn GW (2013) PINK1-phosphorylated Mitofusin-2 is a Parkin receptor for culling damaged mitochondria. Science 340:471–475
Sarraf SA, Raman M, Guarani-Pereira V, Sowa ME, Huttlin EL, Gygi SP, Harper JW (2013) Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496:372–376
Sekine S, Kanamaru Y, Koike M, Nishihara A, Okada M, Kinoshita H, Kamiyama M, Maruyama J, Uchiyama Y, Ishihara N, Takeda K, Ichijo H (2012) Rhomboid protease PARL mediates the mitochondrial membrane potential loss-induced cleavage of PGAM5. J Biol Chem 287(41):34635–34645. https://doi.org/10.1074/jbc.M112.357509
Sekine S, Wang C, Sideris DP, Bunker E, Zhang Z, Youle RJ (2019) Reciprocal roles of Tom7 and OMA1 during mitochondrial import and activation of PINK1. Mol Cell 73(5):1028-1043.e1025. https://doi.org/10.1016/j.molcel.2019.01.002
Jin G, Xu C, Zhang X, Long J, Rezaeian AH, Liu C, Furth ME, Kridel S, Pasche B, Bian XW, Lin HK (2018) Atad3a suppresses Pink1-dependent mitophagy to maintain homeostasis of hematopoietic progenitor cells. Nat Immunol 19(1):29–40. https://doi.org/10.1038/s41590-017-0002-1
Hoshino A, Wang WJ, Wada S, McDermott-Roe C, Evans CS, Gosis B, Morley MP, Rathi KS, Li J, Li K, Yang S, McManus MJ, Bowman C, Potluri P, Levin M, Damrauer S, Wallace DC, Holzbaur ELF, Arany Z (2019) The ADP/ATP translocase drives mitophagy independent of nucleotide exchange. Nature 575(7782):375–379. https://doi.org/10.1038/s41586-019-1667-4
Jin SM, Youle RJ (2013) The accumulation of misfolded proteins in the mitochondrial matrix is sensed by PINK1 to induce PARK2/Parkin-mediated mitophagy of polarized mitochondria. Autophagy 9(11):1750–1757. https://doi.org/10.4161/auto.26122
Fiesel FC, James ED, Hudec R, Springer W (2017) Mitochondrial targeted HSP90 inhibitor Gamitrinib-TPP (G-TPP) induces PINK1/Parkin-dependent mitophagy. Oncotarget 8(63):106233–106248. https://doi.org/10.18632/oncotarget.22287
Yun J, Puri R, Yang H, Lizzio MA, Wu C, Sheng ZH, Guo M (2014) MUL1 acts in parallel to the PINK1/parkin pathway in regulating mitofusin and compensates for loss of PINK1/parkin. eLife 3:e01958. https://doi.org/10.7554/eLife.01958
Rojansky R, Cha MY, Chan DC (2016) Elimination of paternal mitochondria in mouse embryos occurs through autophagic degradation dependent on PARKIN and MUL1. eLife. https://doi.org/10.7554/eLife.17896
Villa E, Proics E, Rubio-Patino C, Obba S, Zunino B, Bossowski JP, Rozier RM, Chiche J, Mondragon L, Riley JS, Marchetti S, Verhoeyen E, Tait SWG, Ricci JE (2017) Parkin-independent mitophagy controls chemotherapeutic response in cancer cells. Cell Rep 20(12):2846–2859. https://doi.org/10.1016/j.celrep.2017.08.087
Chen Z, Liu L, Cheng Q, Li Y, Wu H, Zhang W, Wang Y, Sehgal SA, Siraj S, Wang X, Wang J, Zhu Y, Chen Q (2017) Mitochondrial E3 ligase MARCH5 regulates FUNDC1 to fine-tune hypoxic mitophagy. EMBO Rep 18(3):495–509. https://doi.org/10.15252/embr.201643309
Schwarten M, Mohrluder J, Ma P, Stoldt M, Thielman Y, Stangler T, Hersch N, Hoffman B, Merkel R, Wilbold D (2009) Nix binds to GABARAP: a possible crosstalk between apoptosis and autophagy. Autophagy 5(5):690–698
Novak I, Kirkin V, McEwan DG, Zhang J, Wild P, Rozenknop A, Rogov V, Löhr F, Popovic D, Occhipinti A, Reichert AS, Terzic J, Dötsch V, Ney PA, Dikic I (2010) Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep 11(1):45–51
Hanna RA, Quinsay MN, Orogo AM, Giang K, Rikka S, Gustafsson AB (2012) Microtubule-associated protein 1 light chain 3 (LC3) interacts with BNip3 protein to selectively remove endoplasmic reticulum and mitochondria via autophagy. J Biol Chem 287:19094–19104
Ney PA (2015) Mitochondrial autophagy: origins, significance, and role of BNIP3 and NIX. Biochim Biophy Acta 1853:2775–2783
Springer MZ, Poole LP, Drake LE, Bock-Hughes A, Boland ML, Smith AG, Hart J, Chourasia AH, Liu I, Bozek G, Macleod KF (2021) BNIP3-dependent mitophagy promotes cytosolic localization of LC3B and metabolic homeostasis in the liver. Autophagy Jan 17. https://doi.org/10.1080/15548627.2021.1877469
Zhu Y, Massen S, Terenzio M, Lang V, Chen-Lindner S, Eils R, Novak I, Dikic I, Hamacher-Brady A, Brady NR (2012) Modulation of serines 17 and 24 in the LC3-interacting region of BNIP3 determines pro-survival mitophagy VS apoptosis. J Biol Chem 288(2):1099–1113
Sulistijo ES, MacKenzie KR (2009) Structural basis for dimerization of the BNIP3 transmembrane domain. Biochem 48:5106–5120
Vande Velde C, Cizeau J, Dubik D, Alimonti J, Brown T, Israels S, Hakem R, Greenberg AH (2000) BNIP3 and genetic control of necrosis-like cell death through the mitochondrial permeability transition pore. Mol Cell Biol 20(15):5454–5468
Marinković M, Šprung M, Novak I (2020) Dimerization of mitophagy receptor BNIP3L/NIX is essential for recruitment of autophagic machinery. Autophagy. https://doi.org/10.1080/15548627.2020.1755120
Ohi N, Tokunaga A, Tsunoda H, Nakano K, Haraguchi K, Oda K, Motoyama N, Nakajima T (1999) A novel adenovirus E1B19K-binding protein B5 inhibits apoptosis induced by Nip3 by forming a heterodimer through the C-terminal hydrophobic region. Cell Death Differ 6(4):314–325. https://doi.org/10.1038/sj.cdd.4400493
Guo K, Searfoss G, Krolikowski D, Pagnoni M, Franks C, Clark K, Yu KT, Jaye M, Ivashchenko Y (2001) Hypoxia induces the expression of the pro-apoptotic gene BNIP3. Cell Death Diff 8:367–376
Dayan F, Roux D, Brahimi-Horn C, Pouyssegur J, Mazure NM (2006) The oxygen sensing factor-inhibiting hypoxia-inducible factor-1 controls expression of distinct genes through the bi-functional transcriptional character of hypoxia-inducible factor-1a. Cancer Res 66:3688–3698
Bruick RK (2000) Expression of the gene encoding the proapoptotic Nip3 protein is induced by hypoxia. Proc Natl Acad Sci USA 97(6):9082–9087
Kasper LH, Boussouar F, Boyd K, Xu W, Biesen M, Rehg J, Baudino T, Cleveland JL, Brindle PK (2005) Two transcriptional mechanisms cooperate for the bulk of HIF-1-responsive gene expression. EMBO J 24:3846–3858
Pouyssegur J, Dayan F, Mazure NM (2006) Hypoxia signalling in cancer and approaches to enforce tumour regression. Nature 441:437–443
Glick D, Zhang W, Beaton M, Marsboom G, Gruber M, Simon MC, Hart J, Dorn GW II, Brady MJ, Macleod KF (2012) BNip3 regulates mitochondrial function and lipid metabolism in the liver. Mol Cell Biol 32(13):2570–2584
Lee JM, Wagner M, Xiao R, Kim KH, Lazar MA, Moore DD (2014) Nutrient-sensing nuclear receptors coordinate autophagy. Nature 516:112–115
Mammucari C, Milan G, Romanello V, Masiero E, Rudolf R, Del Piccolo P, Burden SJ, Di Lisi R, Sandri C, Zhao J, Goldberg AL, Schiaffino S, Sandri M (2007) FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab 6:458–471
Reed SA, Sandesara PB, Senf SM, Judge AR (2012) Inhibition of FoxO transcriptional activity prevents muscle fiber atrophy during cachexia and induces hypertrophy. FASEB J 26(3):987–1000. https://doi.org/10.1096/fj.11-189977
Brown JL, Rosa-Caldwell ME, Lee DE, Blackwell TA, Brown LA, Perry RA, Haynie WS, Hardee JP, Carson JA, Wiggs MP, Washington TA, Greene NP (2017) Mitochondrial degeneration precedes the development of muscle atrophy in progression of cancer cachexia in tumour-bearing mice. J Cachexia Sarcopenia Muscle 8(6):926–938. https://doi.org/10.1002/jcsm.12232
Fei P, Wang W, Kim SH, Wang S, Burns TF, Sax JK, Buzzai M, Dicker DT, McKenna WG, Bernhard EJ, El-Deiry WS (2005) Bnip3L is induced by p53 under hypoxia, and its knockdown promotes tumor growth. Cancer Cell 6:597–609
Feng X, Liu X, Zhang W, Xiao W (2011) p53 directly suppresses BNIP3 expression to protect against hypoxia-induced cell death. EMBO J 30:3397–3415
Rogov VV, Suzuki H, Marinkovic M, Lang V, Kato R, Kawasaki M, Buljubasic M, Sprung M, Rogova N, Wakatsuki S, Hamacher-Brady A, Dotsch V, Dikic I, Brady NR, Novak I (2017) Phosphorylation of the mitochondrial autophagy receptor Nix enhances its interaction with LC3 proteins. Sci Rep 7(1):1131. https://doi.org/10.1038/s41598-017-01258-6
Ding WX, Ni HM, Li M, Liao Y, Chen X, Stolz DB, Dorn GW 2nd, Yin XM (2010) Nix is critical to two distinct phases of mitophagy, reactive oxygen species-mediated autophagy induction and Parkin-ubiquitin-p62-mediated mitochondrial priming. J Biol Chem 285(36):27879–27890. https://doi.org/10.1074/jbc.M110.119537
Gao F, Chen D, Si J, Hu Q, Qin Z, Fang M, Wang G (2015) The mitochondrial protein BNIP3L is the substrate of PARK2 and mediates mitophagy in PINK1/PARK2 pathway. Hum Mol Genet 24(9):2528–2538. https://doi.org/10.1093/hmg/ddv017
Zhang T, Xue L, Li L, Tang C, Wan Z, Wang R, Tan J, Tan Y, Han H, Tian R, Billiar TR, Tao WA, Zhang Z (2016) BNIP3 protein suppresses PINK1 kinase proteolytic cleavage to promote mitophagy. J Biol Chem 291(41):21616–21629. https://doi.org/10.1074/jbc.M116.733410
Liu J, Zhang C, Zhao Y, Yue X, Wu H, Huang S, Chen J, Tomsky K, Xie H, Khella CA, Gatza ML, Xia D, Gao J, White E, Haffty BG, Hu W, Feng Z (2017) Parkin targets HIF-1alpha for ubiquitination and degradation to inhibit breast tumor progression. Nat Commun 8(1):1823. https://doi.org/10.1038/s41467-017-01947-w
Li C, Zhang Y, Cheng X, Yuan H, Zhu S, Liu J, Wen Q, Xie Y, Liu J, Kroemer G, Klionsky DJ, Lotze MT, Zeh HJ, Kang R, Tang D (2018) PINK1 and PARK2 suppress pancreatic tumorigenesis through control of mitochondrial iron-mediated immunometabolism. Dev Cell 46(4):441-455.e448. https://doi.org/10.1016/j.devcel.2018.07.012
Kietzmann T (2017) Metabolic zonation of the liver: the oxygen gradient revisited. Redox Biol 11:622–630. https://doi.org/10.1016/j.redox.2017.01.012
Ben-Moshe S, Itzkovitz S (2019) Spatial heterogeneity in the mammalian liver. Nat Rev Gastroenterol Hepatol 16(7):395–410. https://doi.org/10.1038/s41575-019-0134-x
Diwan A, Koesters AG, Odley AM, Pushkaran S, Baines CP, Spike BT, Daria D, Jegga AG, Geiger H, Aronow BJ, Molkentin JD, Macleod KF, Kalfa TA, Dorn GW (2007) Unrestrained erythroblast development in Nix-/- mice reveals a mechanism for apoptotic modulation of erythropoiesis. Proc Natl Acad Sci USA 104:6794–6799
Schweers RL, Zhang J, Randall MS, Loyd MR, Li W, Dorsey FC, Kundu M, Opferman JT, Cleveland JL, Miller JL, Ney PA (2007) NIX is required for programmed mitochondrial clearance during reticulocyte maturation. Proc Natl Acad Sci USA 104:19500–19505
Sandoval H, Thiagarajan P, Dasgupta SK, Scumacker A, Prchal JT, Chen M, Wang J (2008) Essential role for Nix in autophagic maturation of red cells. Nature 454:232–235
Yang C, Hashimoto M, Lin QXX, Tan DQ, Suda T (2019) Sphingosine-1-phosphate signaling modulates terminal erythroid differentiation through the regulation of mitophagy. Exp Hematol 72:47-59.e41. https://doi.org/10.1016/j.exphem.2019.01.004
Quiros PM, Langer T, Lopez-Otin C (2015) New roles for mitochondrial proteases in health, ageing and disease. Nat Rev Mol Cell Biol 16(6):345–359. https://doi.org/10.1038/nrm3984
Mottis A, Herzig S, Auwerx J (2019) Mitocellular communication: shaping health and disease. Science 366(6467):827–832. https://doi.org/10.1126/science.aax3768
Esteban-Martinez L, Sierra-Filardi E, McGreal RS, Salazar-Roa M, Marino G, Seco E, Durand S, Enot D, Grana O, Malumbres M, Cvekl A, Cuervo AM, Kroemer G, Boya P (2017) Programmed mitophagy is essential for the glycolytic switch during cell differentiation. Embo J 36(12):1688–1706. https://doi.org/10.15252/embj.201695916
O’Sullivan TE, Johnson LR, Kang HH, Sun JC (2015) BNIP3 and BNIP3L-mediated mitophagy promotes the generation of natural killer cell memory. Immunity 43:331–342
Li Y, Wang Y, Kim E, Beemiller P, Wang CY, Swanson J, You M, Guan KL (2007) Bnip3 mediates the hypoxia-induced inhibition on mTOR by interacting with Rheb. J Biol Chem 282:35803–35813
Melser S, Chatelain EH, Lavie J, Mahfouf W, Jose C, Obre E, Goorden S, Priault M, Elgersma Y, Rezvani HR, Rossignol R, Bénard G (2013) Rheb regulates mitophagy induced by mitochondrial energetic status. Cell Metab 17(5):719–730
Saxton RA, Sabatini DM (2017) mTOR signaling in growth, metabolism, and disease. Cell 168(6):960–976. https://doi.org/10.1016/j.cell.2017.02.004
Liu L, Feng D, Chen G, Chen M, Zheng Q, Song P, Ma Q, Zhu C, Wang R, Qi W, Huang L, Xue P, Li B, Wang X, Jin H, Wang J, Yang F, Liu P, Zhu Y, Sui S, Chen Q (2012) Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat Cell Biol 14(2):177–185
Egan DF, Shackelford DB, Mihaylova MM, Gelino S, Kohnz RA, Mair W, Vasquez DS, Joshi A, Gwinn DM, Taylor RC, Asara JM, Fitzpatrick J, Dillin A, Viollet B, Kundu M, Hansen M, Shaw RJ (2011) Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331:456–461
Russell RC, Tian Y, Yuan H, Park HW, Chang YY, Kim J, Kim H, Neufeld TP, Dillin A, Guan KL (2013) ULK1 induces autophagy by phosphorylating Beclin-1 and activating VPS34 lipid kinase. Nat Cell Biol 15(7):741–750. https://doi.org/10.1038/ncb2757
Wu WK, Tian W, Hu Z, Chen G, Huang L, Li W, Zhang X, Xue P, Zhou C, Liu L, Zhu Y, Zhang X, Li L, Zhang L, Sui S, Zhao B, Feng D (2014) ULK1 translocates to mitochondria and phosphorylates FUNDC1 to regulate mitophagy. EMBO Rep 15(5):566–575
Chen G, Han Z, Feng D, Chen Y, Chen L, Wu H, Huang L, Zhou C, Cai X, Fu C, Duan L, Wang X, Liu L, Liu X, Shen Y, Zhu Y, Chen Q (2014) A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol Cell 54(3):362–377. https://doi.org/10.1016/j.molcel.2014.02.034
Lu W, Karuppagounder SS, Springer DA, Allen MD, Zheng L, Chao B, Zhang Y, Dawson VL, Dawson TM, Lenardo M (2014) Genetic deficiency of the mitochondrial protein PGAM5 causes a Parkinson’s-like movement disorder. Nat Commun 5:4930. https://doi.org/10.1038/ncomms5930
Wu W, Lin C, Wu K, Jiang L, Wang X, Li W, Zhuang H, Zhang X, Chen H, Li S, Yang Y, Lu Y, Wang J, Zhu R, Zhang L, Sui S, Tan N, Zhao B, Zhang J, Li L, Feng D (2016) FUNDC1 regulates mitochondrial dynamics at the ER-mitochondrial contact site under hypoxic conditions. Embo J 35(13):1368–1384. https://doi.org/10.15252/embj.201593102
Li Y, Xue Y, Xu X, Wang G, Liu Y, Wu H, Li W, Wang Y, Chen Z, Zhang W, Zhu Y, Ji W, Xu T, Liu L, Chen Q (2019) A mitochondrial FUNDC1/HSC70 interaction organizes the proteostatic stress response at the risk of cell morbidity. Embo J. https://doi.org/10.15252/embj.201798786
Li W, Zhang X, Zhuang H, Chen HG, Chen Y, Tian W, Wu WK, Li Y, Wang S, Zhang L, Chen Y, Li L, Zhao B, Sui S, Feng D (2014) MicroRNA-137 is a novel hypoxia-responsive microRNA that inhibits mitophagy via regulation of two mitophagy receptors FUNDC1 and NIX. J Biol Chem 289:10691–10701
Murakawa T, Yamaguchi O, Hashimoto A, Hikoso S, Takeda T, Oka T, Yasui H, Ueda H, Akazawa Y, Nakayama H, Taneike M, Misaka T, Omiya S, Shah AM, Yamamoto A, Nishida K, Ohsumi Y, Okamoto K, Sakata Y, Otsu K (2015) Bcl-2-like protein 13 is a mammalian Atg32 homologue that mediates mitophagy and mitochondrial fragmentation. Nat Commun 6:7527. https://doi.org/10.1038/ncomms8527
Murakawa T, Okamoto K, Omiya S, Taneike M, Yamaguchi O, Otsu K (2019) A mammalian mitophagy receptor, Bcl2-L-13, recruits the ULK1 complex to induce mitophagy. Cell Rep 26(2):338-345.e336. https://doi.org/10.1016/j.celrep.2018.12.050
Merkwirth C, Dargazanli S, Tatsuta T, Geimer S, Löwer B, Wunderlich FT, von Kleist-Retzow JC, Waisman A, Westermann B, Langer T (2008) Prohibitins control cell proliferation and apoptosis by regulating OPA1-dependent cristae morphogenesis in mitochondria. Genes Dev 22(4):476–488
Wei Y, Chiang WC, Sumpter R Jr, Mishra P, Levine B (2016) Prohibitin 2 is an inner mitochondrial membrane mitophagy receptor. Cell. https://doi.org/10.1016/j.cell.2016.11.042
Yoshii SR, Kishi C, Ishihara N, Mizushima N (2011) Parkin mediates proteasome-dependent protein degradation and rupture of the outer mitochondrial membrane. J Biol Chem 286(22):19630–19640. https://doi.org/10.1074/jbc.M110.209338
Dumas JF, Peyta L, Couet C, Servais S (2013) Implication of liver cardiolipins in mitochondrial energy metabolism disorder in cancer cachexia. Biochimie 95(1):27–32. https://doi.org/10.1016/j.biochi.2012.07.009
Chu CT, Ji J, Dagda RK, Jiang JF, Tyurina YY, Kapralov AA, Tyurin VA, Yanamala N, Shrivastava IH, Mohammadyani D, Qiang Wang KZ, Zhu J, Klein-Seetharaman J, Balasubramanian K, Amoscato AA, Borisenko G, Huang Z, Gusdon AM, Cheikhi A, Steer EK, Wang R, Baty C, Watkins S, Bahar I, Bayir H, Kagan VE (2013) Cardiolipin externalization to the outer mitochondrial membrane acts as an elimination signal for mitophagy in neuronal cells. Nat Cell Biol 15(10):1197–1205. https://doi.org/10.1038/ncb2837
Hsu P, Liu X, Zhang J, Wang HG, Ye JM, Shi Y (2015) Cardiolipin remodeling by TAZ/tafazzin is selectively required for the initiation of mitophagy. Autophagy 11(4):643–652. https://doi.org/10.1080/15548627.2015.1023984
Sentelle RD, Senkal CE, Jiang W, Ponnusamy S, Gencer S, Selvam SP, Ramshesh VK, Peterson YK, Lemasters JJ, Szulc ZM, Bielawski J, Ogretmen B (2012) Ceramide targets autophagosomes to mitochondria and induces lethal mitophagy. Nat Chem Biol 8(10):831–838. https://doi.org/10.1038/nchembio.1059
Gao X, Lee K, Reid MA, Sanderson SM, Qiu C, Li S, Liu J, Locasale JW (2018) Serine availability influences mitochondrial dynamics and function through lipid metabolism. Cell Rep 22(13):3507–3520. https://doi.org/10.1016/j.celrep.2018.03.017
Csaki LS, Reue K (2010) Lipins: multifunctional lipid metabolism proteins. Annu Rev Nutr 30:257–272. https://doi.org/10.1146/annurev.nutr.012809.104729
Harris TE, Finck BN (2011) Dual function lipin proteins and glycerolipid metabolism. Trends Endocrinol Metab TEM 22(6):226–233. https://doi.org/10.1016/j.tem.2011.02.006
Zhang P, Verity MA, Reue K (2014) Lipin-1 regulates autophagy clearance and intersects with statin drug effects in skeletal muscle. Cell Metab 20(2):267–279. https://doi.org/10.1016/j.cmet.2014.05.003
Alshudukhi AA, Zhu J, Huang D, Jama A, Smith JD, Wang QJ, Esser KA, Ren H (2018) Lipin-1 regulates Bnip3-mediated mitophagy in glycolytic muscle. FASEB J. https://doi.org/10.1096/fj.201800374
Leong WI, Saba JD (2010) S1P metabolism in cancer and other pathological conditions. Biochimie 92(6):716–723. https://doi.org/10.1016/j.biochi.2010.02.014
Hart PC, Chiyoda T, Liu X, Weigert M, Curtis M, Chiang CY, Loth R, Lastra R, McGregor SM, Locasale JW, Lengyel E, Romero IL (2019) SPHK1 is a novel target of metformin in ovarian cancer. Mol Cancer Res MCR 17(4):870–881. https://doi.org/10.1158/1541-7786.mcr-18-0409
Ader I, Malavaud B, Cuvillier O (2009) When the sphingosine kinase 1/sphingosine 1-phosphate pathway meets hypoxia signaling: new targets for cancer therapy. Can Res 69(9):3723–3726. https://doi.org/10.1158/0008-5472.can-09-0389
Kim S, Sieburth D (2018) Sphingosine kinase activates the mitochondrial unfolded protein response and is targeted to mitochondria by stress. Cell Rep 24(11):2932-2945.e2934. https://doi.org/10.1016/j.celrep.2018.08.037
Martinez-Reyes I, Diebold LP, Kong H, Schieber M, Huang H, Hensley CT, Mehta MM, Wang T, Santos JH, Woychik R, Dufour E, Spelbrink JN, Weinberg SE, Zhao Y, DeBerardinis RJ, Chandel NS (2016) TCA cycle and mitochondrial membrane potential are necessary for diverse biological functions. Mol Cell 61(2):199–209. https://doi.org/10.1016/j.molcel.2015.12.002
Herzig S, Shaw RJ (2018) AMPK: guardian of metabolism and mitochondrial homeostasis. Nat Rev Mol Cell Biol 19(2):121–135. https://doi.org/10.1038/nrm.2017.95
Toyama EQ, Herzig S, Courchet J, Lewis TL Jr, Loson OC, Hellberg K, Young NP, Chen H, Polleux F, Chan DC, Shaw RJ (2016) Metabolism. AMP-activated protein kinase mediates mitochondrial fission in response to energy stress. Science 351(6270):275–281. https://doi.org/10.1126/science.aab4138
Pei S, Minhajuddin M, Adane B, Khan N, Stevens BM, Mack SC, Lai S, Rich JN, Inguva A, Shannon KM, Kim H, Tan AC, Myers JR, Ashton JM, Neff T, Pollyea DA, Smith CA, Jordan CT (2018) AMPK/FIS1-mediated mitophagy is required for self-renewal of human AML stem cells. Cell Stem Cell 23(1):86-100.e106. https://doi.org/10.1016/j.stem.2018.05.021
Domenech E, Maestre C, Esteban-Martinez L, Partida D, Pascual R, Fernandez-Miranda G, Seco E, Campos-Olivas R, Perez M, Megias D, Allen K, Lopez M, Saha AK, Velasco G, Rial E, Mendez R, Boya P, Salazar-Roa M, Malumbres M (2015) AMPK and PFKFB3 mediate glycolysis and survival in response to mitophagy during mitotic arrest. Nat Cell Biol 17(10):1304–1316. https://doi.org/10.1038/ncb3231
Eichner LJ, Brun SN, Herzig S, Young NP, Curtis SD, Shackelford DB, Shokhirev MN, Leblanc M, Vera LI, Hutchins A, Ross DS, Shaw RJ, Svensson RU (2019) Genetic analysis reveals AMPK is required to support tumor growth in murine kras-dependent lung cancer models. Cell Metab 29(2):285-302.e287. https://doi.org/10.1016/j.cmet.2018.10.005
Nakada D, Saunders TL, Morrison SJ (2010) Lkb1 regulates cell cycle and energy metabolism in haematopoietic stem cells. Nature 468(7324):653–658. https://doi.org/10.1038/nature09571
Gurumurthy S, Xie SZ, Alagesan B, Kim J, Yusuf RZ, Saez B, Tzatsos A, Ozsolak F, Milos P, Ferrari F, Park PJ, Shirihai OS, Scadden DT, Bardeesy N (2010) The Lkb1 metabolic sensor maintains haematopoietic stem cell survival. Nature 468(7324):659–663. https://doi.org/10.1038/nature09572
Masand R, Paulo E, Wu D, Wang Y, Swaney DL, Jimenez-Morales D, Krogan NJ, Wang B (2018) Proteome imbalance of mitochondrial electron transport chain in brown adipocytes leads to metabolic benefits. Cell Metab 27(3):616-629.e614. https://doi.org/10.1016/j.cmet.2018.01.018
Kottakis F, Nicolay BN, Roumane A, Karnik R, Gu H, Nagle JM, Boukhali M, Hayward MC, Li YY, Chen T, Liesa M, Hammerman PS, Wong KK, Hayes DN, Shirihai OS, Dyson NJ, Haas W, Meissner A, Bardeesy N (2016) LKB1 loss links serine metabolism to DNA methylation and tumorigenesis. Nature 539(7629):390–395. https://doi.org/10.1038/nature20132
Yang K, Blanco DB, Neale G, Vogel P, Avila J, Clish CB, Wu C, Shrestha S, Rankin S, Long L, Kc A, Chi H (2017) Homeostatic control of metabolic and functional fitness of Treg cells by LKB1 signalling. Nature 548(7669):602–606. https://doi.org/10.1038/nature23665
He N, Fan W, Henriquez B, Yu RT, Atkins AR, Liddle C, Zheng Y, Downes M, Evans RM (2017) Metabolic control of regulatory T cell (Treg) survival and function by Lkb1. Proc Natl Acad Sci USA 114(47):12542–12547. https://doi.org/10.1073/pnas.1715363114
Quiros PM, Prado MA, Zamboni N, D’Amico D, Williams RW, Finley D, Gygi SP, Auwerx J (2017) Multi-omics analysis identifies ATF4 as a key regulator of the mitochondrial stress response in mammals. J Cell Biol 216(7):2027–2045. https://doi.org/10.1083/jcb.201702058
Bao XR, Ong SE, Goldberger O, Peng J, Sharma R, Thompson DA, Vafai SB, Cox AG, Marutani E, Ichinose F, Goessling W, Regev A, Carr SA, Clish CB, Mootha VK (2016) Mitochondrial dysfunction remodels one-carbon metabolism in human cells. eLife. https://doi.org/10.7554/eLife.10575
Tezze C, Romanello V, Desbats MA, Fadini GP, Albiero M, Favaro G, Ciciliot S, Soriano ME, Morbidoni V, Cerqua C, Loefler S, Kern H, Franceschi C, Salvioli S, Conte M, Blaauw B, Zampieri S, Salviati L, Scorrano L, Sandri M (2017) Age-associated loss of OPA1 in muscle impacts muscle mass, metabolic homeostasis, systemic inflammation, and epithelial senescence. Cell Metab 25(6):1374-1389.e1376. https://doi.org/10.1016/j.cmet.2017.04.021
Khan NA, Nikkanen J, Yatsuga S, Jackson C, Wang L, Pradhan S, Kivela R, Pessia A, Velagapudi V, Suomalainen A (2017) mTORC1 regulates mitochondrial integrated stress response and mitochondrial myopathy progression. Cell Metab 26(2):419-428.e415. https://doi.org/10.1016/j.cmet.2017.07.007
Forsstrom S, Jackson CB, Carroll CJ, Kuronen M, Pirinen E, Pradhan S, Marmyleva A, Auranen M, Kleine IM, Khan NA, Roivainen A, Marjamaki P, Liljenback H, Wang L, Battersby BJ, Richter U, Velagapudi V, Nikkanen J, Euro L, Suomalainen A (2019) Fibroblast growth factor 21 drives dynamics of local and systemic stress responses in mitochondrial myopathy with mtDNA deletions. Cell Metab 30(6):1040-1054.e1047. https://doi.org/10.1016/j.cmet.2019.08.019
Ebert SM, Dyle MC, Kunkel SD, Bullard SA, Bongers KS, Fox DK, Dierdorff JM, Foster ED, Adams CM (2012) Stress-induced skeletal muscle Gadd45a expression reprograms myonuclei and causes muscle atrophy. J Biol Chem 287(33):27290–27301. https://doi.org/10.1074/jbc.M112.374777
Guo X, Aviles G, Liu Y, Tian R, Unger BA, Lin YT, Wiita AP, Xu K, Correia MA, Kampmann M (2020) Mitochondrial stress is relayed to the cytosol by an OMA1-DELE1-HRI pathway. Nature 579(7799):427–432. https://doi.org/10.1038/s41586-020-2078-2
Fessler E, Eckl EM, Schmitt S, Mancilla IA, Meyer-Bender MF, Hanf M, Philippou-Massier J, Krebs S, Zischka H, Jae LT (2020) A pathway coordinated by DELE1 relays mitochondrial stress to the cytosol. Nature 579(7799):433–437. https://doi.org/10.1038/s41586-020-2076-4
Harada T, Iwai A, Miyazaki T (2010) Identification of DELE, a novel DAP3-binding protein which is crucial for death receptor-mediated apoptosis induction. Apoptosis 15(10):1247–1255. https://doi.org/10.1007/s10495-010-0519-3
Fiorese CJ, Schulz AM, Lin YF, Rosin N, Pellegrino MW, Haynes CM (2016) The transcription factor ATF5 mediates a mammalian mitochondrial UPR. Curr Biol CB 26(15):2037–2043. https://doi.org/10.1016/j.cub.2016.06.002
Sorrentino V, Romani M, Mouchiroud L, Beck JS, Zhang H, D’Amico D, Moullan N, Potenza F, Schmid AW, Rietsch S, Counts SE, Auwerx J (2017) Enhancing mitochondrial proteostasis reduces amyloid-beta proteotoxicity. Nature 552(7684):187–193. https://doi.org/10.1038/nature25143
Molenaars M, Janssens GE, Williams EG, Jongejan A, Lan J, Rabot S, Joly F, Moerland PD, Schomakers BV, Lezzerini M, Liu YJ, McCormick MA, Kennedy BK, van Weeghel M, van Kampen AHC, Aebersold R, MacInnes AW, Houtkooper RH (2020) A conserved mito-cytosolic translational balance links two longevity pathways. Cell Metab 31(3):549-563.e547. https://doi.org/10.1016/j.cmet.2020.01.011
Nargund AM, Pellegrino MW, Fiorese CJ, Baker BM, Haynes CM (2012) Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337(6094):587–590. https://doi.org/10.1126/science.1223560
Nargund AM, Fiorese CJ, Pellegrino MW, Deng P, Haynes CM (2015) Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPR(mt). Mol Cell 58(1):123–133. https://doi.org/10.1016/j.molcel.2015.02.008
Palikaras K, Lionaki E, Tavernarakis N (2015) Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521:525–528
Gupte R, Liu Z, Kraus WL (2017) PARPs and ADP-ribosylation: recent advances linking molecular functions to biological outcomes. Genes Dev 31(2):101–126. https://doi.org/10.1101/gad.291518.116
Fang EF, Kassahun H, Croteau DL, Scheibye-Knudsen M, Marosi K, Lu H, Shamanna RA, Kalyanasundaram S, Bollineni RC, Wilson MA, Iser WB, Wollman BN, Morevati M, Li J, Kerr JS, Lu Q, Waltz TB, Tian J, Sinclair DA, Mattson MP, Nilsen H, Bohr VA (2016) NAD+ replenishment improves lifespan and healthspan in ataxia telangiectasia models via mitophagy and DNA repair. Cell Metab 24(4):566–581. https://doi.org/10.1016/j.cmet.2016.09.004
Fang EF, Scheibye-Knudsen M, Brace LE, Kassahun H, SenGupta T, Nilsen H, Mitchell JR, Croteau DL, Bohr VA (2014) Defective mitophagy in XPA via PARP-1 hyperactivation and NAD(+)/SIRT1 reduction. Cell 157(4):882–896. https://doi.org/10.1016/j.cell.2014.03.026
Gerhart-Hines Z, Dominy JE Jr, Blattler SM, Jedrychowski MP, Banks AS, Lim JH, Chim H, Gygi SP, Puigserver P (2011) The cAMP/PKA pathway rapidly activates SIRT1 to promote fatty acid oxidation independently of changes in NAD(+). Mol Cell 44(6):851–863. https://doi.org/10.1016/j.molcel.2011.12.005
Rzymski T, Milani M, Pike L, Buffa F, Mellor HR, Winchester L, Pires I, Hammond E, Ragoussis I, Harris AL (2010) Regulation of autophagy by ATF4 in response to severe hypoxia. Oncogene 29(31):4424–4435. https://doi.org/10.1038/onc.2010.191
Pike LR, Singleton DC, Buffa F, Abramczyk O, Phadwal K, Li JL, Simon AK, Murray JT, Harris AL (2013) Transcriptional upregulation of ULK1 by ATF4 contributes to cancer cell survival. Biochem J 449(2):389–400. https://doi.org/10.1042/bj20120972
Anderson CM, Macleod KF (2019) Autophagy and cancer cell metabolism. Int Rev Cell Mol Biol 347:145–190. https://doi.org/10.1016/bs.ircmb.2019.06.002
Zhou R, Yazdi AS, Menu P, Tschopp J (2011) A role for mitochondria in NLRP3 inflammasome activation. Nature 469(7329):221–225. https://doi.org/10.1038/nature09663
Chen Q, Sun L, Chen ZJ (2016) Regulation and function of the cGAS-STING pathway of cytosolic DNA sensing. Nat Immunol 17(10):1142–1149. https://doi.org/10.1038/ni.3558
McArthur K, Whitehead LW, Heddleston JM, Li L, Padman BS, Oorschot V, Geoghegan ND, Chappaz S, Davidson S, San Chin H, Lane RM, Dramicanin M, Saunders TL, Sugiana C, Lessene R, Osellame LD, Chew TL, Dewson G, Lazarou M, Ramm G, Lessene G, Ryan MT, Rogers KL, van Delft MF, Kile BT (2018) BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science. https://doi.org/10.1126/science.aao6047
Aarreberg LD, Esser-Nobis K, Driscoll C, Shuvarikov A, Roby JA, Gale M Jr (2019) Interleukin-1β induces mtDNA release to activate innate immune signaling via cGAS-STING. Mol Cell 74(4):801-815.e806. https://doi.org/10.1016/j.molcel.2019.02.038
Zhong Z, Liang S, Sanchez-Lopez E, He F, Shalapour S, Lin XJ, Wong J, Ding S, Seki E, Schnabl B, Hevener AL, Greenberg HB, Kisseleva T, Karin M (2018) New mitochondrial DNA synthesis enables NLRP3 inflammasome activation. Nature 560(7717):198–203. https://doi.org/10.1038/s41586-018-0372-z
Shimada K, Crother TR, Karlin J, Dagvadorj J, Chiba N, Chen S, Ramanujan VK, Wolf AJ, Vergnes L, Ojcius DM, Rentsendorj A, Vargas M, Guerrero C, Wang Y, Fitzgerald KA, Underhill DM, Town T, Arditi M (2012) Oxidized mitochondrial DNA activates the NLRP3 inflammasome during apoptosis. Immunity 36(3):401–414. https://doi.org/10.1016/j.immuni.2012.01.009
Misawa T, Takahama M, Kozaki T, Lee H, Zou J, Saitoh T, Akira S (2013) Microtubule-driven spatial arrangement of mitochondria promotes activation of the NLRP3 inflammasome. Nat Immunol 14(5):454–460. https://doi.org/10.1038/ni.2550
Elliott EI, Sutterwala FS (2015) Initiation and perpetuation of NLRP3 inflammasome activation and assembly. Immunol Rev 265(1):35–52. https://doi.org/10.1111/imr.12286
Zhong Z, Umemura A, Sanchez-Lopez E, Liang S, Shalapour S, Wong J, He F, Boassa D, Perkins G, Ali SR, McGeough MD, Ellisman MH, Seki E, Gustafsson AB, Hoffman HM, Diaz-Meco MT, Moscat J, Karin M (2016) NF-kappaB restricts inflammasome activation via elimination of damaged mitochondria. Cell 164(5):896–910. https://doi.org/10.1016/j.cell.2015.12.057
Yu JJ, Nagasu H, Murakami T, Hoang H, Broderick L, Hoffman HM, Horng T (2014) Inflammasome activation leads to caspase-1 dependent mitochondrial damage and block of mitophagy. Proc Natl Acad Sci U S A 111(43):15514–15519
Moore AS, Holzbaur EL (2016) Dynamic recruitment and activation of ALS-associated TBK1 with its target optineurin are required for efficient mitophagy. Proc Natl Acad Sci U S A 113(24):E3349-3358. https://doi.org/10.1073/pnas.1523810113
Richter B, Sliter DA, Herhaus L, Stolz A, Wang C, Beli P, Zaffagnini G, Wild P, Martens S, Wagner SA, Youle RJ, Dikic I (2016) Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria. Proc Natl Acad Sci U S A 113(15):4039–4044. https://doi.org/10.1073/pnas.1523926113
Gui X, Yang H, Li T, Tan X, Shi P, Li M, Du F, Chen ZJ (2019) Autophagy induction via STING trafficking is a primordial function of the cGAS pathway. Nature 567(7747):262–266. https://doi.org/10.1038/s41586-019-1006-9
Gaidt MM, Ebert TS, Chauhan D, Ramshorn K, Pinci F, Zuber S, O’Duill F, Schmid-Burgk JL, Hoss F, Buhmann R, Wittmann G, Latz E, Subklewe M, Hornung V (2017) The DNA inflammasome in human myeloid cells is initiated by a STING-cell death program upstream of NLRP3. Cell 171(5):1110-1124.e1118. https://doi.org/10.1016/j.cell.2017.09.039
Nakahira K, Haspel JA, Rathinam VA, Lee SJ, Dolinay T, Lam HC, Englert JA, Rabinovitch M, Cernadas M, Kim HP, Fitzgerald KA, Ryter SW, Choi AM (2011) Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat Immunol 12(3):222–230. https://doi.org/10.1038/ni.1980
Drake LE, Springer MZ, Poole LP, Kim CJ, Macleod KF (2017) Expanding perspectives on the significance of mitophagy in cancer. Semin Cancer Biol 47:110–124. https://doi.org/10.1016/j.semcancer.2017.04.008
Cesari R, Martin ES, Calin GA, Pentimalli F, Bichi R, McAdams H, Trapasso F, Drusco A, Shimizu M, Masciullo V, D’Andrilli G, Scambia G, Picchio MC, Alder H, Godwin AK, Croce CM (2003) Parkin, a gene implicated in autosomal recessive juvenile parkinsonism, is a candidate tumor suppressor gene on chromosome 6q25-q27. Proc Natl Acad Sci U S A 100(10):5956–5961
Veeriah S, Taylor BS, Meng S, Fang F, Yilmaz E, Vivanco I, Janakiraman M, Schultz N, Hanrahan AJ, Pao W, Ladanyi M, Sander C, Heguy A, Holland EC, Paty PB, Mischel PS, Liau L, Cloughesy TF, Mellinghoff IK, Solit DB, Chan TA (2010) Somatic mutations of the Parkinson’s disease-associated gene PARK2 in glioblastoma and other human malignancies. Nat Genet 42(1):77–82. https://doi.org/10.1038/ng.491
Agnihotri S, Golbourn B, Huang X, Remke M, Younger S, Cairns RA, Chalil A, Smith CA, Krumholtz SL, Mackenzie D, Rakopoulos P, Ramaswamy V, Taccone MS, Mischel PS, Fuller GN, Hawkins C, Stanford WL, Taylor MD, Zadeh G, Rutka JT (2016) PINK1 is a negative regulator of growth and the warburg effect in glioblastoma. Can Res 76(16):4708–4719. https://doi.org/10.1158/0008-5472.can-15-3079
Fujiwara M, Marusawa H, Wang HQ, Iwai A, Ikeuchi K, Imai Y, Kataoka A, Nukina N, Takahashi R, Chiba T (2008) Parkin as a tumor suppressor gene for hepatocellular carcinoma. Oncogene 27:6002–6011
Zhang C, Lin M, Wu R, Wang X, Yang BG, Levine AJ, Hu W, Feng Z (2011) Parkin, a p53 target gene, mediates the role of p53 in glucose metabolism and the Warburg effect. Proc Natl Acad Sci U S A 108(39):16259–16264
Bernardini JP, Lazarou M, Dewson G (2016) Parkin and mitophagy in cancer. Oncogene. https://doi.org/10.1038/onc.2016.302
Kim KY, Stevens MV, Akter MH, Rusk SE, Huang RJ, Cohen A, Noguchi A, Springer D, Bocharov AV, Eggerman TL, Suen DF, Youle RJ, Amar M, Remaley AT, M.N. S, (2011) Parkin is a lipid-responsive regulator of fat uptake in mice and mutant human cells. J Clin Invest 121:3701–3712
Sarraf SA, Sideris DP, Giagtzoglou N, Ni L, Kankel MW, Sen A, Bochicchio LE, Huang CH, Nussenzweig SC, Worley SH, Morton PD, Artavanis-Tsakonas S, Youle RJ, Pickrell AM (2019) PINK1/parkin influences cell cycle by sequestering TBK1 at damaged mitochondria. Inhibiting Mitosis Cell Rep 29(1):225-235.e225. https://doi.org/10.1016/j.celrep.2019.08.085
Gong Y, Zack TI, Morris LG, Lin K, Hukkelhoven E, Raheja R, Tan IL, Turcan S, Veeriah S, Meng S, Viale A, Schumacher SE, Palmedo P, Beroukhim R, Chan TA (2014) Pan-cancer genetic analysis identifies PARK2 as a master regulator of G1/S cyclins. Nat Genet 46(6):588–594. https://doi.org/10.1038/ng.2981
Lee SB, Kim JJ, Nam HJ, Gao B, Yin P, Qin B, Yi SY, Ham H, Evans D, Kim SH, Zhang J, Deng M, Liu T, Zhang H, Billadeau DD, Wang L, Giaime E, Shen J, Pang YP, Jen J, van Deursen JM, Lou Z (2015) Parkin regulates mitosis and genomic stability through Cdc20/Cdh1. Mol Cell 60(1):21–34. https://doi.org/10.1016/j.molcel.2015.08.011
Pillai S, Nguyen J, Johnson J, Haura E, Coppola D, Chellappan S (2015) Tank binding kinase 1 is a centrosome-associated kinase necessary for microtubule dynamics and mitosis. Nat Commun 6:10072. https://doi.org/10.1038/ncomms10072
Villa E, Marchetti S, Ricci JE (2018) No parkin zone: mitophagy without parkin. Trends Cell Biol 28(11):882–895. https://doi.org/10.1016/j.tcb.2018.07.004
Sowter HM, Ferguson M, Pym C, Watson P, Fox SB, Han C, Harris AL (2003) Expression of the cell death genes BNip3 and Nix in ductal carcinoma in situ of the breast; correlation of BNip3 levels with necrosis and grade. JPath 201:573–580
Okami J, Simeone DM, Logsdon CD (2004) Silencing of the hypoxia-inducible cell death protein BNIP3 in pancreatic cancer. Cancer Res 64(15):5338–5346
Erkan M, Kleef J, Esposito I, Giese T, Ketterer K, Buchler MW, Giese NA, Friess H (2005) Loss of BNIP3 expression is a late event in pancreatic cancer contributing to chemoresistance and worsened prognosis. Oncogene 24:4421–4432
Murai M, Toyota M, Suzuki H, Satoh A, Sasaki Y, Akino K, Ueno M, Takahashi F, Kusano M, Mita H, Yanagihara K, Endo T, Hinoda Y, Tokino T, Imai K (2005) Aberrant methylation and silencing of the BNIP3 gene in colorectal and gastric cancer. Clin Cancer Res 11(3):1021–1027
Akada M, Crnogorac-Jurcevic T, Lattimore S, Mahon P, Lopes R, Sunamura M, Matsuno S, Lemoine NR (2005) Intrinsic chemoresistance to gemcitabine is associated with decreased expression of BNIP3 in pancreatic cancer. Clin Cancer Res 11(8):3094–3101
Humpton TJ, Alagesan B, DeNicola GM, Lu D, Yordanov GN, Leonhardt CS, Yao MA, Alagesan P, Zaatari MN, Park Y, Skepper JN, Macleod KF, Perez-Mancera PA, Murphy MP, Evan GI, Vousden KH, Tuveson DA (2019) Oncogenic KRAS induces NIX-mediated mitophagy to promote pancreatic cancer. Cancer Discov 9(9):1268–1287. https://doi.org/10.1158/2159-8290.cd-18-1409
Beroukhim R, Mermel CH, Porter D, Wei G, Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J, Urashima M, Mc Henry KT, Pinchback RM, Ligon AH, Cho YJ, Haery L, Greulich H, Reich M, Winckler W, Lawrence MS, Weir BA, Tanaka KE, Chiang DY, Bass AJ, Loo A, Hoffman C, Prensner J, Liefeld T, Gao Q, Yecies D, Signoretti S, Maher E, Kaye FJ, Sasaki H, Tepper JE, Fletcher J, Tabernero J, Baselga J, Tsao MS, Demichelis F, Rubin MA, Janne PA, Daly MJ, Nucera C, Levine RL, Ebert BL, Gabriel S, Rustgi AK, Antonescu CR, Ladanyi M, Letai A, Garraway LA, Loda M, Beer DG, True LD, Okamoto A, Pomeroy SL, Singer S, Golub TR, Lander ES, Getz G, Sellers WR, Meyerson M (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463:899–905
Koop EA, van Laar T, Weger RA, van der Wall E, van Diest PJ (2009) Expression of BNIP3 in invasive breast cancer: correlations with the hypoxic response and clinicopathological features. BMC Cancer 9:175–182
Chourasia AH, Tracy K, Frankenberger C, Boland ML, Sharifi MN, Drake LE, Sachleben JR, Asara JM, Locasale JW, Karczmar GS, Macleod KF (2015) Mitophagy defects arising from BNip3 loss promote mammary tumor progression to metastasis. EMBO Rep 16(9):1145–1163. https://doi.org/10.15252/embr.201540759
Montagner M, Enzo E, Forcato M, Zanconato F, Parenti A, Rampazzo E, Basso G, Leo G, Rosato A, Bicciato S, Cordenonsi M, Piccolo S (2012) SHARP1 suppresses breast cancer metastasis by promoting degradation of hypoxia-inducible factors. Nature 487(7407):380–384. https://doi.org/10.1038/nature11207
Guha M, Srinivasan S, Raman P, Jiang Y, Kaufman BA, Taylor D, Dong D, Chakrabarti R, Picard M, Carstens RP, Kijima Y, Feldman M (1864) Avadhani NG (2018) Aggressive triple negative breast cancers have unique molecular signature on the basis of mitochondrial genetic and functional defects. Biochim Biophys Acta Mol Basis Dis 4:1060–1071. https://doi.org/10.1016/j.bbadis.2018.01.002
Lee J, Yesilkanal AE, Wynne JP, Frankenberger C, Liu J, Yan J, Elbaz M, Rabe DC, Rustandy FD, Tiwari P, Grossman EA, Hart PC, Kang C, Sanderson SM, Andrade J, Nomura DK, Bonini MG, Locasale JW, Rosner MR (2019) Effective breast cancer combination therapy targeting BACH1 and mitochondrial metabolism. Nature 568(7751):254–258. https://doi.org/10.1038/s41586-019-1005-x
Manka D, Spicer Z, Millhorn DE (2005) Bcl-2/Adenovirus E1B 19 kDa interacting protein-3 knockdown enables growth of breast cancer metastases in the lung, liver and bone. Cancer Res 65:11689–11693
Labuschagne CF, Cheung EC, Blagih J, Domart MC, Vousden KH (2019) Cell clustering promotes a metabolic switch that supports metastatic colonization. Cell Metab 30(4):720-734.e725. https://doi.org/10.1016/j.cmet.2019.07.014
Smith AG, Macleod KF (2019) Autophagy, cancer stem cells and drug resistance. J Pathol 247(5):708–718. https://doi.org/10.1002/path.5222
Liu K, Lee J, Kim JY, Wang L, Tian Y, Chan ST, Cho C, Machida K, Chen D, Ou JJ (2017) Mitophagy controls the activities of tumor suppressor p53 to regulate hepatic cancer stem cells. Mol Cell 68(2):281–292. https://doi.org/10.1016/j.molcel.2017.09.022
Katajisto P, Dohla J, Chaffer CL, Pentinmikko N, Marjanovic N, Iqbal S, Zoncu R, Chen W, Weinberg RA, Sabatini DM (2015) Stem cells. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness. Science 348(6232):340–343. https://doi.org/10.1126/science.1260384
Ho TT, Warr MR, Adelman ER, Lansinger OM, Flach J, Verovskaya EV, Figueroa ME, Passegue E (2017) Autophagy maintains the metabolism and function of young and old stem cells. Nature 543(7644):205–210. https://doi.org/10.1038/nature21388
Ito K, Turcotte R, Cui J, Zimmerman SE, Pinho S, Mizoguchi T, Arai F, Runnels JM, Alt C, Teruya-Feldstein J, Mar JC, Singh R, Suda T, Lin CP, Frenette PS, Ito K (2016) Self-renewal of a purified Tie2+ hematopoietic stem cell population relies on mitochondrial clearance. Science 354(6316):1156–1160. https://doi.org/10.1126/science.aaf5530
Vannini N, Girotra M, Naveiras O, Nikitin G, Campos V, Giger S, Roch A, Auwerx J, Lutolf MP (2016) Specification of haematopoietic stem cell fate via modulation of mitochondrial activity. Nat Commun 7:13125. https://doi.org/10.1038/ncomms13125
Vannini N, Campos V, Girotra M, Trachsel V, Rojas-Sutterlin S, Tratwal J, Ragusa S, Stefanidis E, Ryu D, Rainer PY, Nikitin G, Giger S, Li TY, Semilietof A, Oggier A, Yersin Y, Tauzin L, Pirinen E, Cheng WC, Ratajczak J, Canto C, Ehrbar M, Sizzano F, Petrova TV, Vanhecke D, Zhang L, Romero P, Nahimana A, Cherix S, Duchosal MA, Ho PC, Deplancke B, Coukos G, Auwerx J, Lutolf MP, Naveiras O (2019) The NAD-booster nicotinamide riboside potently stimulates hematopoiesis through increased mitochondrial clearance. Cell Stem Cell 24(3):405-418.e407. https://doi.org/10.1016/j.stem.2019.02.012
Li J, Agarwal E, Bertolini I, Seo JH, Caino MC, Ghosh JC, Kossenkov AV, Liu Q, Tang HY, Goldman AR, Languino LR, Speicher DW, Altieri DC (2020) The mitophagy effector FUNDC1 controls mitochondrial reprogramming and cellular plasticity in cancer cells. Sci Signal. https://doi.org/10.1126/scisignal.aaz8240
Wu L, Zhang D, Zhou L, Pei Y, Zhuang Y, Cui W, Chen J (2019) FUN14 domain-containing 1 promotes breast cancer proliferation and migration by activating calcium-NFATC1-BMI1 axis. EBioMedicine 41:384–394. https://doi.org/10.1016/j.ebiom.2019.02.032
Cleaver JE (2005) Cancer in xeroderma pigmentosum and related disorders of DNA repair. Nat Rev Cancer 5(7):564–573. https://doi.org/10.1038/nrc1652
Kee Y, D’Andrea AD (2012) Molecular pathogenesis and clinical management of Fanconi anemia. J Clin Investig 122(11):3799–3806. https://doi.org/10.1172/jci58321
Shiloh Y, Ziv Y (2013) The ATM protein kinase: regulating the cellular response to genotoxic stress, and more. Nat Rev Mol Cell Biol 14(4):197–210
Sumpter R Jr, Sirasanagandla S, Fernandez AF, Wei Y, Dong X, Franco L, Zou Z, Marchal C, Lee MY, Clapp DW, Hanenberg H, Levine B (2016) Fanconi anemia proteins function in mitophagy and immunity. Cell 165(4):867–881. https://doi.org/10.1016/j.cell.2016.04.006
Orvedahl A, Sumpter R Jr, Xiao G, Ng A, Zou Z, Tang Y, Narimatsu M, Gilpin C, Sun Q, Roth M, Forst CV, Wrana JL, Zhang YE, Luby-Phelps K, Xavier RJ, Xie Y, Levine B (2011) Image-based genome-wide siRNA screen identifies selective autophagy factors. Nature 480(7375):113–117. https://doi.org/10.1038/nature10546
de Vries Y, Lwiwski N, Levitus M, Kuyt B, Israels SJ, Arwert F, Zwaan M, Greenberg CR, Alter BP, Joenje H, Meijers-Heijboer H (2012) A dutch fanconi anemia FANCC founder mutation in Canadian manitoba mennonites. Anemia 2012:865170. https://doi.org/10.1155/2012/865170
Ambrose M, Goldstine JV, Gatti RA (2007) Intrinsic mitochondrial dysfunction in ATM-deficient lymphoblastoid cells. Hum Mol Genet 16(18):2154–2164. https://doi.org/10.1093/hmg/ddm166
Eaton JS, Lin ZP, Sartorelli AC, Bonawitz ND, Shadel GS (2007) Ataxia-telangiectasia mutated kinase regulates ribonucleotide reductase and mitochondrial homeostasis. J Clin Investig 117(9):2723–2734. https://doi.org/10.1172/jci31604
Valentin-Vega YA, Maclean KH, Tait-Mulder J, Milasta S, Steeves M, Dorsey FC, Cleveland JL, Green DR, Kastan MB (2012) Mitochondrial dysfunction in ataxia-telangiectasia. Blood 119(6):1490–1500
Gomes LC, Di Benedetto G, Scorrano L (2011) During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat Cell Biol 13(5):589–598. https://doi.org/10.1038/ncb2220
Rambold AS, Kostelecky B, Elia N, Lippincott-Schwartz J (2011) Tubular network formation protects mitochondrial from autophagosomal degradation during nutrient starvation. Proc Natl Acad Sci U S A 108(25):10190–10195
Yamada T, Murata D, Adachi Y, Itoh K, Kameoka S, Igarashi A, Kato T, Araki Y, Huganir RL, Dawson TM, Yanagawa T, Okamoto K, Iijima M, Sesaki H (2018) Mitochondrial stasis reveals p62-mediated ubiquitination in parkin-independent mitophagy and mitigates nonalcoholic fatty liver disease. Cell Metab 28(4):588-604.e585. https://doi.org/10.1016/j.cmet.2018.06.014
Mishra P, Chan DC (2014) Mitochondrial dynamics and inheritance during cell division, development and disease. Nat Rev Mol Cell Biol 15(10):634–646. https://doi.org/10.1038/nrm3877
Otera H, Wang C, Cleland MM, Setoguchi K, Yokota S, Youle RJ, Mihara K (2010) Mff is an essential factor for mitochondrial recruitment of Drp1 during mitochondrial fission in mammalian cells. J Cell Biol 191(6):1141–1158. https://doi.org/10.1083/jcb.201007152
Wong YC, Ysselstein D, Krainc D (2018) Mitochondria-lysosome contacts regulate mitochondrial fission via RAB7 GTP hydrolysis. Nature 554(7692):382–386. https://doi.org/10.1038/nature25486
Yamano K, Fogel AI, Wang C, van der Bliek AM, Youle RJ (2014) Mitochondrial Rab GAPs govern autophagosome biogenesis during mitophagy. eLife 3:e01612. https://doi.org/10.7554/eLife.01612
Shen Q, Yamano K, Head BP, Kawajiri S, Cheung JT, Wang C, Cho JH, Hattori N, Youle RJ, van der Bliek AM (2014) Mutations in Fis1 disrupt orderly disposal of defective mitochondria. Mol Biol Cell 25(1):145–159. https://doi.org/10.1091/mbc.E13-09-0525
Cho HM, Ryu JR, Jo Y, Seo TW, Choi YN, Kim JH, Chung JM, Cho B, Kang HC, Yu SW, Yoo SJ, Kim H, Sun W (2019) Drp1-Zip1 interaction regulates mitochondrial quality surveillance system. Mol Cell 73(2):364-376.e368. https://doi.org/10.1016/j.molcel.2018.11.009
Lee YK, Lee HY, Hanna RA, Gustafsson AB (2011) Mitochondrial autophagy by Bnip3 involves Drp1-mediated mitochondrial fission and recruitment of Parkin in cardiac myocytes. Am j Physiol Heart Circ Physiol 301(5):H1924-1931
Kashatus JA, Nascimento A, Myers LJ, Sher A, Byrne FL, Hoehn KL, Counter CM, Kashatus DF (2015) Erk2 phosphorylation of Drp1 promotes mitochondrial fission and MAPK-driven tumor growth. Mol Cell 57(3):537–551. https://doi.org/10.1016/j.molcel.2015.01.002
Serasinghe MN, Wieder SY, Renault TT, Elkholi R, Asciolla JJ, Yao JL, Jabado O, Hoehn K, Kageyama Y, Sesaki H, Chipuk JE (2015) Mitochondrial division is requisite to RAS-induced transformation and targeted by oncogenic MAPK pathway inhibitors. Mol Cell 57(3):521–536. https://doi.org/10.1016/j.molcel.2015.01.003
Wieder SY, Serasinghe MN, Sung JC, Choi DC, Birge MB, Yao JL, Bernstein E, Celebi JT, Chipuk JE (2015) Activation of the mitochondrial fragmentation protein DRP1 correlates with BRAF(V600E) melanoma. J Invest Dermatol 135(10):2544–2547. https://doi.org/10.1038/jid.2015.196
Trotta AP, Gelles JD, Serasinghe MN, Loi P, Arbiser JL, Chipuk JE (2017) Disruption of mitochondrial electron transport chain function potentiates the pro-apoptotic effects of MAPK inhibition. J Biol Chem 292(28):11727–11739. https://doi.org/10.1074/jbc.M117.786442
Nagdas S, Kashatus JA, Nascimento A, Hussain SS, Trainor RE, Pollock SR, Adair SJ, Michaels AD, Sesaki H, Stelow EB, Bauer TW, Kashatus DF (2019) Drp1 promotes KRas-driven metabolic changes to drive pancreatic tumor growth. Cell Rep 28(7):1845-1859.e1845. https://doi.org/10.1016/j.celrep.2019.07.031
Kinsey CG, Camolotto SA, Boespflug AM, Guillen KP, Foth M, Truong A, Schuman SS, Shea JE, Seipp MT, Yap JT, Burrell LD, Lum DH, Whisenant JR, Gilcrease GW 3rd, Cavalieri CC, Rehbein KM, Cutler SL, Affolter KE, Welm AL, Welm BE, Scaife CL, Snyder EL, McMahon M (2019) Protective autophagy elicited by RAF–>MEK–>ERK inhibition suggests a treatment strategy for RAS-driven cancers. Nat Med 25(4):620–627. https://doi.org/10.1038/s41591-019-0367-9
Bryant KL, Stalnecker CA, Zeitouni D, Klomp JE, Peng S, Tikunov AP, Gunda V, Pierobon M, Waters AM, George SD, Tomar G, Papke B, Hobbs GA, Yan L, Hayes TK, Diehl JN, Goode GD, Chaika NV, Wang Y, Zhang GF, Witkiewicz AK, Knudsen ES, PetricoinSingh EFPK, Macdonald JM, Tran NL, Lyssiotis CA, Ying H, Kimmelman AC, Cox AD, Der CJ (2019) Combination of ERK and autophagy inhibition as a treatment approach for pancreatic cancer. Nat Med 25(4):628–640. https://doi.org/10.1038/s41591-019-0368-8
Kalas W, Swiderek E, Rapak A, Kopij M, Rak JW, Strzadala L (2011) H-ras up-regulates expression of BNIP3. Anticancer Res 31(9):2869–2875
Ishikawa K, Takenaga K, Akimoto M, Koshikawa N, Yaaguchi A, Imanishi H, Nakada K, Honma Y, Hayashi JI (2008) ROS-generating mitochondrial DNA mutations can regulate tumor cell metastasis. Science 320:661–664
Porporato PE, Payen VL, Perez-Escuredo J, De Saedeleer CJ, Danhier P, Copetti T, Dhup S, Tardy M, Vazeille T, Bouzin C, Feron O, Michiels C, Gallez B, Sonveaux P (2014) A mitochondrial switch promotes tumor metastasis. Cell Rep 8(3):754–766. https://doi.org/10.1016/j.celrep.2014.06.043
Tan A, Baty JW, Dong LF, Bezawork-Geleta A, Endaya B, Goodwin J, Bajzikova M, Kovarova J, Peterka M, Yan B, Pesdar EA, Sobol M, Filimonenko A, Stuart S, Vondrusova M, Kluckova K, Sachaphibulkij K, Rohlena J, Hozak P, Truksa J, Eccles D, Haupt LM, Griffiths LR, Neuzil J, Berridge MV (2015) Mitochondrial genome acquisition restores respiratory function and tumorigenic potential of cancer cells without mitochondrial DNA. Cell Metab 21:81–94
Dong LF, Kovarova J, Bajzikova M, Bezawork-Geleta A, Svec D, Endaya B, Sachaphibulkij K, Coelho AR, Sebkova N, Ruzickova A, Tan AS, Kluckova K, Judasova K, Zamecnikova K, Rychtarcikova Z, Gopalan V, Andera L, Sobol M, Yan B, Pattnaik B, Bhatraju N, Truksa J, Stopka P, Hozak P, Lam AK, Sedlacek R, Oliveira PJ, Kubista M, Agrawal A, Dvorakova-Hortova K, Rohlena J, Berridge MV, Neuzil J (2017) Horizontal transfer of whole mitochondria restores tumorigenic potential in mitochondrial DNA-deficient cancer cells. eLife. https://doi.org/10.7554/eLife.22187
Cheung EC, DeNicola GM, Nixon C, Blyth K, Labuschagne CF, Tuveson DA, Vousden KH (2020) Dynamic ROS control by TIGAR regulates the initiation and progression of pancreatic cancer. Cancer Cell 37(2):168-182.e164. https://doi.org/10.1016/j.ccell.2019.12.012
Radisky DC, Levy E, Littlepage LE, Liu H, Nelson CM, Fata JE, Leake D, Godden EL, Albertson DG, Nieto MA, Werb Z, Bissell MJ (2005) Rac1b and reactive oxygen species mediate MMP3-induced EMT and genomic instability. Nature 436:123–127
Yang MH, Wu MZ, Chiou SH, Chen PM, Chang SY, Liu CJ, Teng SC, Wu KJ (2008) Direct regulation of TWIST by HIF-1alpha promotes metastasis. Nat Cell Biol 10(3):295–305. https://doi.org/10.1038/ncb1691
Holmstrom KM, Finkel T (2014) Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat Rev Mol Cell Biol 15:411–421
Sabharwal SS, Schumacker PT (2014) Mitochondrial ROS in cancer: initiators, amplifiers or an Achilles’ heel? Nat Rev Cancer 14(11):709–721. https://doi.org/10.1038/nrc3803
Piskounova E, Agathocleous M, Murphy MM, Hu Z, Huddlestun SE, Zhao Z, Leitch AM, Johnson TM, DeBerardinis RJ, Morrison SJ (2015) Oxidative stress inhibits distant metastasis by human melanoma cells. Nature 527(7577):186–191. https://doi.org/10.1038/nature15726
Le Gal K, Ibrahim MX, Wiel C, Sayin VI, Akula MK, Karlsson C, Dalin MG, Akyurek LM, Lindahl P, Nilsson J, Bergo MO (2015) Antioxidants can increase melanoma metastasis in mice. Sci Transl Med 7(308):308. https://doi.org/10.1126/scitranslmed.aad3740
Kenny TC, Gomez ML, Germain D (2019) Mitohormesis, UPR(mt), and the complexity of mitochondrial DNA landscapes in cancer. Can Res. https://doi.org/10.1158/0008-5472.can-19-1395
Kenny TC, Craig AJ, Villanueva A, Germain D (2019) Mitohormesis primes tumor invasion and metastasis. Cell Rep 27(8):2292-2303.e2296. https://doi.org/10.1016/j.celrep.2019.04.095
Chourasia AH, Macleod KF (2015) Tumor suppressor functions of BNIP3 and mitophagy. Autophagy 11(10):1937–1938. https://doi.org/10.1080/15548627.2015.1085136
Pedanou VE, Gobeil S, Tabaries S, Simone TM, Zhu LJ, Siegel PM, Green MR (2016) The histone H3K9 demethylase KDM3A promotes anoikis by transcriptionally activating pro-apoptotic genes BNIP3 and BNIP3L. eLife. https://doi.org/10.7554/eLife.16844
Maes H, Van Eygen S, Krysko DV, Vandenabeele P, Nys K, Rillaerts K, Garg AD, Verfaillie T, Agostinis P (2014) BNIP3 supports melanoma cell migration and vasculogenic mimicry by orchestrating the actin cytoskeleton. Cell Death Dis 5(3):e1127. https://doi.org/10.1038/cddis.2014.94
Zhang J, Zhang C, Jiang X, Li L, Zhang D, Tang D, Yan T, Zhang Q, Yuan H, Jia J, Hu J, Zhang J, Huang Y (2019) Involvement of autophagy in hypoxia-BNIP3 signaling to promote epidermal keratinocyte migration. Cell Death Dis 10(3):234. https://doi.org/10.1038/s41419-019-1473-9
Zhang N, Yang X, Yuan F, Zhang L, Wang Y, Wang L, Mao Z, Luo J, Zhang H, Zhu WG, Zhao Y (2018) Increased amino acid uptake supports autophagy-deficient cell survival upon glutamine deprivation. Cell Rep 23(10):3006–3020. https://doi.org/10.1016/j.celrep.2018.05.006
Towers CG, Fitzwalter BE, Regan D, Goodspeed A, Morgan MJ, Liu CW, Gustafson DL, Thorburn A (2019) Cancer cells upregulate NRF2 signaling to adapt to autophagy inhibition. Dev Cell 50(6):690-703.e696. https://doi.org/10.1016/j.devcel.2019.07.010
Lau A, Wang XJ, Zhao F, Villeneuve NF, Wu T, Jiang T, Sun Z, White E, Zhang DD (2010) A noncanonical mechanism of Nrf2 activation by autophagy deficiency: direct interaction between Keap1 and p62. Mol Cell Biol 30(13):3275–3285
Fearon KC, Glass DJ, Guttridge DC (2012) Cancer cachexia: mediators, signaling, and metabolic pathways. Cell Metab 16(2):153–166. https://doi.org/10.1016/j.cmet.2012.06.011
Biswas AK, Acharyya S (2020) Understanding cachexia in the context of metastatic progression. Nat Rev Cancer 20(5):274–284. https://doi.org/10.1038/s41568-020-0251-4
Porporato PE (2016) Understanding cachexia as a cancer metabolism syndrome. Oncogenesis 5:e200. https://doi.org/10.1038/oncsis.2016.3
Petruzzelli M, Wagner EF (2016) Mechanisms of metabolic dysfunction in cancer-associated cachexia. Genes Dev 30(5):489–501. https://doi.org/10.1101/gad.276733.115
Tsoli M, Robertson G (2013) Cancer cachexia: malignant inflammation, tumorkines, and metabolic mayhem. Trends Endocrinol Metab TEM 24(4):174–183. https://doi.org/10.1016/j.tem.2012.10.006
Stephens NA, Skipworth RJ, Gallagher IJ, Greig CA, Guttridge DC, Ross JA, Fearon KC (2015) Evaluating potential biomarkers of cachexia and survival in skeletal muscle of upper gastrointestinal cancer patients. J Cachexia Sarcopenia Muscle 6(1):53–61. https://doi.org/10.1002/jcsm.12005
Julienne CM, Dumas JF, Goupille C, Pinault M, Berri C, Collin A, Tesseraud S, Couet C, Servais S (2012) Cancer cachexia is associated with a decrease in skeletal muscle mitochondrial oxidative capacities without alteration of ATP production efficiency. J Cachexia Sarcopenia Muscle 3(4):265–275. https://doi.org/10.1007/s13539-012-0071-9
Antunes D, Padrao AI, Maciel E, Santinha D, Oliveira P, Vitorino R, Moreira-Goncalves D, Colaco B, Pires MJ, Nunes C, Santos LL, Amado F, Duarte JA, Domingues MR (2014) Molecular insights into mitochondrial dysfunction in cancer-related muscle wasting. Biochem Biophys Acta 6:896–905. https://doi.org/10.1016/j.bbalip.2014.03.004
Amaravadi R, Kimmelman A, Debnath J (2019) Targeting autophagy in cancer: recent advances, future directions. Cancer Discov Under review
Levy JMM, Towers CG, Thorburn A (2017) Targeting autophagy in cancer. Nat Rev Cancer 17(9):528–542. https://doi.org/10.1038/nrc.2017.53
Briceno E, Reyes S, Sotelo J (2003) Therapy of glioblastoma multiforme improved by the antimutagenic chloroquine. Neurosurg Focus 14(2):e3
Sotelo J, Briceno E, Lopez-Gonzalez MA (2006) Adding chloroquine to conventional treatment for glioblastoma multiforme: a randomized, double-blind, placebo-controlled trial. Ann Intern Med 144(5):337–343
Pascolo S (2016) Time to use a dose of Chloroquine as an adjuvant to anti-cancer chemotherapies. Eur J Pharmacol 771:139–144. https://doi.org/10.1016/j.ejphar.2015.12.017
Elliott IA, Dann AM, Xu S, Kim SS, Abt ER, Kim W, Poddar S, Moore A, Zhou L, Williams JL, Capri JR, Ghukasyan R, Matsumura C, Tucker DA, Armstrong WR, Cabebe AE, Wu N, Li L, Le TM, Radu CG, Donahue TR (2019) Lysosome inhibition sensitizes pancreatic cancer to replication stress by aspartate depletion. Proc Natl Acad Sci U S A 116(14):6842–6847. https://doi.org/10.1073/pnas.1812410116
Gao M, Yi J, Zhu J, Minikes AM, Monian P, Thompson CB, Jiang X (2019) Role of mitochondria in ferroptosis. Mol Cell 73(2):354-363.e353. https://doi.org/10.1016/j.molcel.2018.10.042
Georgakopoulos ND, Wells G, Campanella M (2017) The pharmacological regulation of cellular mitophagy. Nat Chem Biol 13(2):136–146. https://doi.org/10.1038/nchembio.2287
Chourasia AH, Boland ML, Macleod KF (2015) Mitophagy and cancer. Cancer Metab 3:4. https://doi.org/10.1186/s40170-015-0130-8
Rao VA, Klein SR, Bonar SJ, Zielonka J, Mizuno N, Dickey JS, Keller PW, Joseph J, Kalyanaraman B, Shacter E (2010) The antioxidant transcription factor Nrf2 negatively regulates autophagy and growth arrest induced by the anticancer redox agent mitoquinone. J Biol Chem 285(45):34447–34459. https://doi.org/10.1074/jbc.M110.133579
Meyer N, Zielke S, Michaelis JB, Linder B, Warnsmann V, Rakel S, Osiewacz HD, Fulda S, Mittelbronn M, Münch C, Behrends C, Kögel D (2018) AT 101 induces early mitochondrial dysfunction and HMOX1 (heme oxygenase 1) to trigger mitophagic cell death in glioma cells. Autophagy 14(10):1693–1709. https://doi.org/10.1080/15548627.2018.1476812
Georgakopoulos ND, Frison M, Alvarez MS, Bertrand H, Wells G, Campanella M (2017) Reversible Keap1 inhibitors are preferential pharmacological tools to modulate cellular mitophagy. Sci Rep 7(1):10303. https://doi.org/10.1038/s41598-017-07679-7
Heo JM, Ordureau A, Paulo JA, Rinehart J, Harper J (2015) The PINK1-PARKIN mitochondrial ubiquitylation pathway drives a program of OPTN/NDP52 recruitment and TBK1 activation to promote mitophagy. Mol Cell 60:7–20
Wong YC, Holzbaur EL (2014) Optineurin is an autophagy receptor for damaged mitochondria in parkin-mediated mitophagy that is disrupted by an ALS-linked mutation. Proc Natl Acad Sci USA 111(42):E4439-4448. https://doi.org/10.1073/pnas.1405752111
Li L, Shen C, Nakamura E, Ando K, Signoretti S, Beroukhim R, Cowley GS, Lizotte P, Liberzon E, Bair S, Root DE, Tamayo P, Tsherniak A, Cheng SC, Tabak B, Jacobsen A, Hakimi AA, Schultz N, Ciriello G, Sander C, Hsieh JJ, Kaelin WGJ (2013) SQSTM1 is a pathogenic target of 5q copy number gains in kidney cancer. Cancer Cell 24(6):738–750
Moscat J, Karin M, Diaz-Meco MT (2016) p62 in cancer: signaling adaptor beyond autophagy. Cell 167(3):606–609. https://doi.org/10.1016/j.cell.2016.09.030
Hsieh CH, Shaltouki A, Gonzalez AE, Bettencourt da Cruz A, Burbulla LF, St Lawrence E, Schule B, Krainc D, Palmer TD, Wang X (2016) Functional impairment in miro degradation and mitophagy is a shared feature in familial and sporadic Parkinson’s disease. Cell Stem Cell 19(6):709–724. https://doi.org/10.1016/j.stem.2016.08.002
Sugiura A, Nagashima S, Tokuyama T, Amo T, Matsuki Y, Ishido S, Kudo Y, McBride HM, Fukuda T, Matsushita N, Inatome R, Yanagi S (2013) MITOL regulates endoplasmic reticulum-mitochondria contacts via Mitofusin2. Mol Cell 51(1):20–34. https://doi.org/10.1016/j.molcel.2013.04.023
Szargel R, Shani V, Abd Elghani F, Mekies LN, Liani E, Rott R, Engelender S (2016) The PINK1, synphilin-1 and SIAH-1 complex constitutes a novel mitophagy pathway. Hum Mol Genet 25(16):3476–3490. https://doi.org/10.1093/hmg/ddw189
Anding AL, Wang C, Chang TK, Sliter DA, Powers CM, Hofmann K, Youle RJ, Baehrecke EH (2018) Vps13D encodes a ubiquitin-binding protein that is required for the regulation of mitochondrial size and clearance. Curr Biol CB 28(2):287-295.e286. https://doi.org/10.1016/j.cub.2017.11.064
Peterson TR, Sengupta SS, Harris TE, Carmack AE, Kang SA, Balderas E, Guertin DA, Madden KL, Carpenter AE, Finck BN, Sabatini DM (2011) mTOR complex 1 regulates lipin 1 localization to control the SREBP pathway. Cell 146(3):408–420. https://doi.org/10.1016/j.cell.2011.06.034
Winter RN, Kramer A, Borkowski A, Kyprianou N (2001) Loss of caspase-1 and caspase-3 protein expression in human prostate cancer. Can Res 61(3):1227–1232
Yang YM, Ramadani M, Huang YT (2003) Overexpression of Caspase-1 in adenocarcinoma of pancreas and chronic pancreatitis. World J Gastroenterol 9(12):2828–2831. https://doi.org/10.3748/wjg.v9.i12.2828
Feng Q, Li P, Salamanca C, Huntsman D, Leung PC, Auersperg N (2005) Caspase-1alpha is down-regulated in human ovarian cancer cells and the overexpression of caspase-1alpha induces apoptosis. Can Res 65(19):8591–8596. https://doi.org/10.1158/0008-5472.can-05-0239
Zaki MH, Vogel P, Body-Malapel M, Lamkanfi M, Kanneganti TD (2010) IL-18 production downstream of the Nlrp3 inflammasome confers protection against colorectal tumor formation. J Immunol 185(8):4912–4920. https://doi.org/10.4049/jimmunol.1002046
Daley D, Mani VR, Mohan N, Akkad N, Pandian G, Savadkar S, Lee KB, Torres-Hernandez A, Aykut B, Diskin B, Wang W, Farooq MS, Mahmud AI, Werba G, Morales EJ, Lall S, Wadowski BJ, Rubin AG, Berman ME, Narayanan R, Hundeyin M, Miller G (2017) NLRP3 signaling drives macrophage-induced adaptive immune suppression in pancreatic carcinoma. J Exp Med 214(6):1711–1724. https://doi.org/10.1084/jem.20161707
Vanpouille-Box C, Demaria S, Formenti SC, Galluzzi L (2018) Cytosolic DNA sensing in organismal tumor control. Cancer Cell 34(3):361–378. https://doi.org/10.1016/j.ccell.2018.05.013
Rzymski T, Milani M, Singleton DC, Harris AL (2009) Role of ATF4 in regulation of autophagy and resistance to drugs and hypoxia. Cell Cycle 8(23):3838–3847. https://doi.org/10.4161/cc.8.23.10086
Dey S, Sayers CM, Verginadis II, Lehman SL, Cheng Y, Cerniglia GJ, Tuttle SW, Feldman MD, Zhang PJ, Fuchs SY, Diehl JA, Koumenis C (2015) ATF4-dependent induction of heme oxygenase 1 prevents anoikis and promotes metastasis. J Clin Investig 125(7):2592–2608. https://doi.org/10.1172/jci78031
Zhao J, Brault JJ, Schild A, Cao P, Sandri M, Schiaffino S, Lecker SH, Goldberg AL (2007) FoxO3 coordinately activates protein degradation by tthe autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab 6:472–483
Zhao Y, Yang J, Liao W, Liu X, Zhang H, Wang S, Wang D, Feng J, Yu L, Zhu WG (2010) Cytosolic FoxO1 is essential for the induction of autophagy and tumour suppressor activity. Nat Cell Biol 12(7):665–675. https://doi.org/10.1038/ncb2069
Warr MR, Binnewies M, Flach J, Reynaud D, Garg T, Malhotra R, Debnath J, Passegue E (2013) FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494(7437):323–327. https://doi.org/10.1038/nature11895
Fitzwalter BE, Towers CG, Sullivan KD, Andrysik Z, Hoh M, Ludwig M, O’Prey J, Ryan KM, Espinosa JM, Morgan MJ, Thorburn A (2018) Autophagy inhibition mediates apoptosis sensitization in cancer therapy by relieving FOXO3a turnover. Dev Cell 44(5):555-565.e553. https://doi.org/10.1016/j.devcel.2018.02.014
Keith B, Simon MC (2007) Hypoxia-inducible factors, stem cells and cancer. Cell 129:465–472
Semenza GL (2016) Hypoxia-inducible factors: coupling glucose metabolism and redox regulation with induction of the breast cancer stem cell phenotype. Embo J. https://doi.org/10.15252/embj.201695204
Perera RM, Stoykova S, Nicolay BN, Ross KN, Fitamant J, Boukhali M, Lengrand J, Deshpande V, Selig MK, Ferrone CR, Settleman J, Stephanopoulos G, Dyson NJ, Zoncu R, Ramaswamy S, Haas W, Bardeesy N (2015) Transcriptional control of the autophagy-lysosome system in pancreatic cancer. Nature 524:361–365
Strappazzon F, Nazio F, Corrado M, Cianfanelli V, Romagnoli A, Fimia GM, Campello S, Nardacci R, Piacentini M, Campanella M, Cecconi F (2015) AMBRA1 is able to induce mitophagy via LC3 binding, regardless of PARKIN and p62/SQSTM1. Cell Death Differ 22(3):419–432. https://doi.org/10.1038/cdd.2014.139
Falasca L, Torino F, Marconi M, Costantini M, Pompeo V, Sentinelli S, De Salvo L, Patrizio M, Padula C, Gallucci M, Piacentini M, Malorni W (2015) AMBRA1 and SQSTM1 expression pattern in prostate cancer. Apoptosis 20(12):1577–1586. https://doi.org/10.1007/s10495-015-1176-3
Strappazzon F, Cecconi F (2015) AMBRA1-induced mitophagy: a new mechanism to cope with cancer? Mol Cell Oncol 2(2):e975647. https://doi.org/10.4161/23723556.2014.975647
Cianfanelli V, Fuoco C, Lorente M, Salazar M, Quondamatteo F, Gherardini PF, De Zio D, Nazio F, Antonioli M, D’Orazio M, Skobo T, Bordi M, Rohde M, Dalla Valle L, Helmer-Citterich M, Gretzmeier C, Dengjel J, Fimia GM, Piacentini M, Di Bartolomeo S, Velasco G, Cecconi F (2015) AMBRA1 links autophagy to cell proliferation and tumorigenesis by promoting c-Myc dephosphorylation and degradation. Nat Cell Biol 17(1):20–30. https://doi.org/10.1038/ncb3072
Schoenherr C, Byron A, Sandilands E, Paliashvili K, Baillie GS, Garcia-Munoz A, Valacca C, Cecconi F, Serrels B, Frame MC (2017) Ambra1 spatially regulates Src activity and Src/FAK-mediated cancer cell invasion via trafficking networks. eLife. https://doi.org/10.7554/eLife.23172
Park S, Choi SG, Yoo SM, Son JH, Jung YK (2014) Choline dehydrogenase interacts with SQSTM1/p62 to recruit LC3 and stimulate mitophagy. Autophagy 10(11):1906–1920. https://doi.org/10.4161/auto.32177
Karbowski M, Jeong SY, Youle RJ (2004) Endophilin B1 is required for the maintenance of mitochondrial morphology. J Cell Biol 166(7):1027–1039. https://doi.org/10.1083/jcb.200407046
Takahashi Y, Hori T, Cooper TK, Liao J, Desai N, Serfass JM, Young MM, Park S, Izu Y, Wang HG (2013) Bif-1 haploinsufficiency promotes chromosomal instability and accelerates Myc-driven lymphomagenesis via suppression of mitophagy. Blood 121(9):1622–1632. https://doi.org/10.1182/blood-2012-10-459826
Bhujabal Z, Birgisdottir AB, Sjottem E, Brenne HB, Overvatn A, Habisov S, Kirkin V, Lamark T, Johansen T (2017) FKBP8 recruits LC3A to mediate Parkin-independent mitophagy. EMBO Rep 18(6):947–961. https://doi.org/10.15252/embr.201643147
Shlevkov E, Kramer T, Schapansky J, LaVoie MJ, Schwarz TL (2016) Miro phosphorylation sites regulate Parkin recruitment and mitochondrial motility. Proc Natl Acad Sci U S A 113(41):E6097-e6106. https://doi.org/10.1073/pnas.1612283113
Le Guerroue F, Eck F, Jung J, Starzetz T, Mittelbronn M, Kaulich M, Behrends C (2017) Autophagosomal content profiling reveals an LC3C-dependent piecemeal mitophagy pathway. Mol Cell 68(4):786-796.e786. https://doi.org/10.1016/j.molcel.2017.10.029
D’Amico D, Mottis A, Potenza F, Sorrentino V, Li H, Romani M, Lemos V, Schoonjans K, Zamboni N, Knott G, Schneider BL, Auwerx J (2019) The RNA-binding protein PUM2 impairs mitochondrial dynamics and mitophagy during aging. Mol Cell 73(4):775-787.e710. https://doi.org/10.1016/j.molcel.2018.11.034
Schneider AT, Gautheron J, Feoktistova M, Roderburg C, Loosen SH, Roy S, Benz F, Schemmer P, Büchler MW, Nachbur U, Neumann UP, Tolba R, Luedde M, Zucman-Rossi J, Panayotova-Dimitrova D, Leverkus M, Preisinger C, Tacke F, Trautwein C, Longerich T, Vucur M, Luedde T (2017) RIPK1 suppresses a TRAF2-dependent pathway to liver cancer. Cancer Cell 31(1):94–109. https://doi.org/10.1016/j.ccell.2016.11.009
Hawk MA, Gorsuch CL, Fagan P, Lee C, Kim SE, Hamann JC, Mason JA, Weigel KJ, Tsegaye MA, Shen L, Shuff S, Zuo J, Hu S, Jiang L, Chapman S, Leevy WM, DeBerardinis RJ, Overholtzer M, Schafer ZT (2018) RIPK1-mediated induction of mitophagy compromises the viability of extracellular-matrix-detached cells. Nat Cell Biol 20(3):272–284. https://doi.org/10.1038/s41556-018-0034-2
Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N, Oomori H, Noda T, Haraguchi T, Hiraoka Y, Amano A, Yoshimori T (2013) Autophagosomes form at ER-mitochondria contact sites. Nature 495(7441):389–393. https://doi.org/10.1038/nature11910
Arasaki K, Nagashima H, Kurosawa Y, Kimura H, Nishida N, Dohmae N, Yamamoto A, Yanagi S, Wakana Y, Inoue H, Tagaya M (2018) MAP1B-LC1 prevents autophagosome formation by linking syntaxin 17 to microtubules. EMBO Rep. https://doi.org/10.15252/embr.201745584
Sugo M, Kimura H, Arasaki K, Amemiya T, Hirota N, Dohmae N, Imai Y, Inoshita T, Shiba-Fukushima K, Hattori N, Cheng J, Fujimoto T, Wakana Y, Inoue H, Tagaya M (2018) Syntaxin 17 regulates the localization and function of PGAM5 in mitochondrial division and mitophagy. Embo J. https://doi.org/10.15252/embj.201798899
Xian H, Yang Q, Xiao L, Shen HM, Liou YC (2019) STX17 dynamically regulated by Fis1 induces mitophagy via hierarchical macroautophagic mechanism. Nat Commun 10(1):2059. https://doi.org/10.1038/s41467-019-10096-1
Kumar S, Gu Y, Abudu YP, Bruun JA, Jain A, Farzam F, Mudd M, Anonsen JH, Rusten TE, Kasof G, Ktistakis N, Lidke KA, Johansen T, Deretic V (2019) Phosphorylation of syntaxin 17 by TBK1 controls autophagy initiation. Dev Cell 49(1):130–144. https://doi.org/10.1016/j.devcel.2019.01.027
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Poole, L.P., Macleod, K.F. Mitophagy in tumorigenesis and metastasis. Cell. Mol. Life Sci. 78, 3817–3851 (2021). https://doi.org/10.1007/s00018-021-03774-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-021-03774-1