Abstract
Lung carcinoma is the most frequently diagnosed malignant neoplasms and mainly consists of small-cell lung carcinoma (SCLC) and non-small-cell lung carcinoma (NSCLC). Large number of lung carcinoma patients have poor outcomes due to the late diagnosis and the limited therapeutic options. Previous attempts have proved that the evolution of lung carcinoma is a multistep molecular aberration which various genetic or epigenetic alterations may be take part in. Among these molecular aberrations, the inactivation of tumor suppressor gene has been widely observed in all types of carcinoma including lung carcinoma. As a vital inactivated mechanism, DNA methylation of tumor suppressor gene is frequently found in lung cancer. To gain exhaustive comprehension of the carcinogenesis of lung carcinoma, we summarize our current knowledge on DNA methylation of RASSF1 (RAS-Association Domain Family 1) and its clinical significance in lung carcinoma.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
8.1 Introduction
Lung carcinoma is the leading cause of cancer-related death, with an estimated 388,000 deaths in Europe in 2018 [1, 2]. Based upon the data of smoking prevalence from the population-based Adult Health Survey in 2003, the estimated lung cancer mortality was 15.0 and 7.1 per 100,000 among men and women in 2018 [1], respectively. In China, the incidence of lung carcinoma is also high, with the highest mortality rate as compared with other countries [2]. The uptake of tobacco among males and exposure to unventilated cooking fumes among females are the predominant non-genetic risk factors for lung carcinoma [3,4,5,6,7]. The 5-year survival rate of lung carcinoma is very low, especially in Eastern Asia, due to the large proportion of lung carcinoma patients present with advanced metastatic tumors when diagnosed [8, 9].
Lung carcinoma mainly consists of small-cell lung carcinoma (SCLC) as the most aggressive lung carcinoma accounted for about 25% of bronchogenic carcinomas and non-small-cell lung carcinoma (NSCLC) as the most common lung carcinoma subtype for approximately 85% of lung cancer cases [10, 11]. The major histological subtypes of NSCLC are represented by lung adenocarcinoma, squamous cell carcinoma, and large cell carcinoma [12], of which the resection is performed in the early stage and chemo-combination in the late stage, with the mean 5-year survival of 15%. The large number of lung cancer patients have poor outcomes due to the late diagnosis, acquired multidrug resistance, and complex mechanisms [13]. This chapter aims at exploring the comprehensive mechanisms on the carcinogenesis of lung cancer by furthermore understanding DNA methylation of RAS-Association Domain Family 1 (RASSF1) and its clinical significance in lung carcinoma. We pay more specific attention on the potential mechanisms and new specific molecular markers of lung cancer, especially DNA methylation of tumor suppressor genes and inactivated genes in the development of lung carcinoma.
8.2 The RASSF1 Gene
RASSF1 is one of the key tumor-suppressor genes allocated in chromosome 3p21.3 and spans about 11,151 bp [14, 10]. RASSF1 promotes apoptosis, microtubule stability and polymerization, and mitotic progression [15]. The protein encoded by RASSF1 can participate in RAS-related cellular signal pathways and regulate oncogenesis, cell proliferation, differentiation, and apoptosis in a wide variety of cancer types [16]. Eight transcripts, i.e., RASSF1A, B, C, D, E, F, G, and H, are generated by RASSF1 gene and contain a Ras-Association (RA) domain in the carboxyterminal segments, except for RASSF1F-H which is similar to the RAS effector proteins, Raf1. Raf1 is associated with Ras-GTP to activate Ras proteins, suppress cell growth, and promote proapoptotic effects.
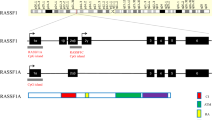
RASSF1A and RASSF1C are two predominant common isoforms and encode an ATM-kinase phosphorylation site and a conserved carboxyterminal SARAH (Sav/RASSF/Hpo) domain as a key component of the Hippo signaling pathway, except for the RA domain. RASSF1A has a diacylglycerol/phorbol ester-binding (DAG) domain containing a central zinc finger which is also known as the protein kinase C conserved domain (C1 domain). RASSF1C variant is shorter than RASSF1A and lacks the amino terminal C1 domain. RASSF1D and E have the RA, SARAH, C1 domains and ATM-kinase phosphorylation site similar to RASSF1A in structure. RASSF1B contains one RA and SARAH domain, respectively. Isoforms F/G and H have a C1 domain and an ATM-kinase phosphorylation site, respectively (Fig. 8.1).
(a) Schematic map of RASSF1 locus. Black boxes indicate exons and open boxes indicate untranslated regions, respectively. Two CpG islands are shown by black lines. The transcription start sites of isoform A and isoform C are indicated with black arrows. (b) The domain structures of polypeptides encoded by RASSF1. C1, DAG/diacylglycerol binding domain (black) putative ATM kinase phosphorylation consensus sequence motif (blue) RA, Ras-association domain (brown) and SARAH, Sav/RASSF/Hpo interaction domain (red)
RASSF1C appears to share many of the biological characteristics of RASSF1A. On basis of the similar structure to Ras effector, RASSF1 gene regulates cell proliferation, differentiation, and apoptosis. RASSF1A functions as a negative regulator of cell proliferation by blocking the cell cycle progression at the level of G1/S-phase [17] and has the dual role in the coordination of p53 and p73 responses [18], while RASSF1C exhibits growth inhibitory potency [19], although there is little known on functions of other variants. RASSF1B, D, and E are found poorly expressed in hemopoietic, cardiac, and pancreatic cells, respectively. RASSF1F, E, D, and G share the same promoter region with RASSF1A, although the biological significance remains unclear.
RASSF2, RASSF3, NORE1, and RASSF6 were identified as the homolog of RASSF1, which share similar Ras-association domain with RASSF1. These genes have SARAH domain and code multiple transcripts. RASSF2 shares a lower homology (29% identity) with RASSF1, while acting as a tumor suppressor gene and undergoing promoter methylation at high frequency similar to RASSF1 [20]. The inactivation of RASSF2 may be associated with tumor progression [20], and RASSF3 suppresses tumor formation through interacting with MDM2 and inducing NSCLC cell apoptosis [21]. NORE1 shares about 50% sequence identities with RASSF1 and has similar pattern of mRNA transcript expression and function as a tumor-suppressor gene [22,23,24]. RASSF6 is found frequently suppressed in several human cancers [20, 25,26,27].
8.3 Inactivation of RASSF1A by DNA Methylation in Lung Carcinoma
The genomic mutational landscape provided evidences that genetic alterations taken part in the tumorigenesis of lung cancer. The epigenetic regulation provides a novel insight in the progression and evolution of lung cancer [28, 29]. Of the epigenetic modifications, DNA methylation mainly occurs in C-G dinucleotide-rich regions, also named CpG islands [30], where the methyl group is added by DNA methyltransferase at the 5-position cytosine and erased by demethylase. DNA methylation mainly occurs at the cytosine‑phosphate‑guanine (CpG) island which locates in promoter region of a gene and regulates the expression of gene, which plays a vital role in genomic imprint erasure, instability of chromatin structure, and X-chromosome inactivation. The aberrant DNA methylation interacts with gene expression in the early stage of human cancers and dynamically during lung carcinogenesis. A lot of methylated genes have been identified in lung carcinoma, including RASSF1, major tumor suppressor 1, fragile histidine triad, methylguanine-DNA methyltransferase, and adenomatosis polyposis coli tumor suppressor.
The loss of heterozygosity (LOH) is the most frequent event during lung tumorigenesis [31], while rarely attributed to somatic mutations, except for one frame-shift and missense mutation identified in nasopharyngeal carcinomas [32]. RASSF1 is inactivated frequently by the hypermethylation of the promoter CpG island in cancers [33,34,35,36,37]. RASSF1 methylation was originally reported in lung cancer and then shown as the common event in cancers [33, 38]. RASSF1A was methylation-inactivated in SCLC, while aberrant methylation of the RASSF1C CpG island promoter was not observed in lung cancer [33].
8.4 Signaling Pathway Involving RASSF1 in Lung Cancer
Among signaling pathways, RASSF1A contributes to the carcinogenesis of lung cancer mainly through Hippo signaling pathways. The Hippo pathway (i.e., Salvador-Warts-Hippo pathway) in a kinase cascade regulates the organ size through regulating cell proliferation, differentiation, and apoptosis [39,40,41]. The core components of the pathway encompass the mammalian sterile 20-like kinase 1 and 2 (MST1 and MST2) and the large tumor suppressor 1 and 2 (LATS1 and LATS2), and cooperate with the adaptor/scaffold proteins, Salvador homolog 1 (hSAV1), and MOB kinase activator 1A and 1B (hMOB1). The downstream effectors of Hippo pathway are two WW domain-containing transcriptional coactivators TAZ and its paralog YAP. Mst1/2 phosphorylates hSAV1 and forms the activated Mst1/2-hSAV1 complex which cooperates with hMOB1 and activates LATS1/2. After then LATS1/2 phosphorylates YAP/TAZ which is prevented from entering to the nucleus. Then the complex with transcriptional enhancer factors (TEADs) is formed and the expression of anti-apoptotic and pro-proliferative genes are activated [42,43,44].
During DNA damage, RASSF1A activated by ATM can induce apoptosis through the interaction of Hippo pathway with MST1/MST2 via the C-terminus to prevent the autophosphorylation of those protein kinases [18, 44, 45]. The components of the Hippo pathway are intimately involved in lung morphogenesis and tumorigenesis [46,47,48]. The abnormal expression of those components is associated with the clinical classification, poor differentiation, metastasis, and poor prognosis and survival in lung cancer [47, 49,50,51,52]. The DNA methylation of promoter results in the inactivation of RASSF1A, RASSF1A-MST1/MST2 complex, and dysfunction of the Hippo pathway. RASSF1A can enhance the transcription of proapoptotic genes through the formation of a complex with YAP and p73 in the nucleus (Fig. 8.2).
A summary of RASSF1A pathways in cancergenesis of lung cancer. RASSF1A regulates cell apoptosis through its interactions with the connector enhancer of KSR (CNK1), the proapoptotic kinase MST1, and the modulator of apoptosis-1 (MAP-1). The CNK1–MST1 complex is also thought to play important role in cell proliferation. RASSF1A can regulate the microtubule network by recruiting effectors of the microtubule-associated protein 1B (MAP1B), C19ORF5, and the Cdc20. RASSF1A also induced G1 and S-phase cell cycle arrest through inhibiting the transcription factor p120E4F (RASSF1A was enhanced by p120E4F) and JNK
RASSF1A may contribute to the carcinogenesis of lung cancer through microtubules and binding of Cdc20 via an N-terminal region. Cdc20 cannot bind with APC and fail to form the complex Cdc20-APC for the spindle assembly checkpoint during mitosis [53]. RASSF1A is required for stabilizing the microtubule. RASSF1A controls the motility and invasion of lung cancer cells through the modulation of tubulin dynamics [54, 55]. The promoter hypermethylation of RASSF1A activates premature APC, following by accelerated cell division, mitotic spindle abnormalities, and chromosome misalignment [53] (Fig. 8.2). The exogenous expression of RASSF1A modulates levels of cyclin D1 and induces cell cycle arrest in lung carcinoma cells [17]. RASSF1A inhibits lung cancer cell growth through reducing the phosphorylation of JNK [56] (Fig. 8.2).
8.5 Clinical Significance of RASSF1 in Lung Carcinoma
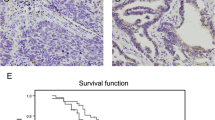
RASSF1 methylation in cancer may serve an important role in clinical utilities, especially in lung cancer. For example, the aberrant RASSF1A methylation may be an ideal biomarker for early diagnostic and prognostic due to the non-invasive, high sensitivity, and high specificity characteristics. It is questionable whether RASSF1A methylation can be a powerful marker for patient prognosis at early stage of lung cancer. RASSF1A exhibited lung cancer-specific methylation pattern, with the hypermethylation level up to 100% in SCLC and 63% in NSCLC [57, 58]. RASSF1A methylation can be detected in body fluids including blood, urine, sputum, and bronchial alveolar lavages [56,57,58,59]. For example, RASSF1A methylation is observed in the blood of patients with NSCLC [59]. The RASSF1A methylation of bronchial aspirates was 21% in patients with lung cancer and smoking and 1% in patients with lung cancer alone, respectively. The methylation level of RASSF1A was associated with the number of cigarette packs and smoking years during the lifetime of patients with lung cancer [60]. The RASSF1 methylation of bronchial washings was found to have diagnostic sensitivity [61], which has the great potential to screen risk populations of patients with lung cancer. DNA methylation of RASSF1A is correlated with poor clinicopathological characteristics in nearly all solid tumors [62], which also includes lung cancer. RASSF1 promoter methylation was found in poorly differentiated tumors [63,64,65], associated with tumor grades, stages, and survival. For example, RASSF1A methylation was associated with patient survival time in lung adenocarcinoma [66]. Decreased survival time was observed in NSCLC patients with RASSF1A methylation, irrespective of whether patients have received adjuvant radio therapy or surgical treatment [58, 64, 67, 68]. On basis of those evidence, RASSF1 and isoforms as disease biomarkers should be furthermore evaluated, since disease biomarkers are expected to have the clear specificity for disease per se, disease stage, phase, severity, duration, or response to therapy [69,70,71,72,73,74,75,76]. Several natural compounds can regulate DNMT activity or expression to re-activate RASSF1A [77]. Peperomin E, as a natural bioactive secolignan polyphenol extracted from the plant peperomia dindygulensis, could demethylate RASSF1A and upregulate the expression of RASSF1A by reducing the level of DNMT1 in lung cancer cells [78].
8.6 Conclusion
Epigenetics changes especially DNA methylation has been proved to take part in the carcinogenesis of cancers. The DNA methylation of the tumor suppressor genes may be exploitable for the biologic and clinical significance of cancers. Overall, as the common tumor suppressor gene of lung cancer, evidence have suggested the DNA methylation of RASSF1 can be an essential potential clinic diagnostic or prognostic marker and may provide new therapeutic strategies for future successful treatment of lung cancer. It will be very interesting to further explore how to develop non-invasive, rapid and less cost detection methods for DNA methylation and to confirm the reliability and sensitivity of DNA methylation.
Abbreviations
- SCLC:
-
Small-cell lung carcinoma
- NSCLC:
-
Non-small-cell lung carcinoma
- RASSF1:
-
RAS-Association Domain Family 1
- TEADs:
-
Transcriptional enhancer factors
- TAZ:
-
WW domain-containing transcriptional coactivators
- YAP:
-
TAZ paralog
- hMOB1:
-
MOB kinase activator 1B
- hSAV1:
-
The adaptor/scaffold proteins Salvador homolog 1
- LATS1 and LATS2:
-
Large tumor suppressor 1 and 2
- MST1 and MST2:
-
Mammalian sterile 20-like kinase 1 and 2
- LOH:
-
Heterozygosity
References
Torre LA, Siegel RL, Jemal A (2016) Lung cancer statistics. Adv Exp Med Biol 893:1–19
Ferlay J, Colombet M, Soerjomataram I et al (2018) Cancer incidence and mortality patterns in Europe: estimates for 40 countries and 25 major cancers in 2018. Eur J Cancer 103:356
Winkler V, Mangolo NJ, Becher H (2015) Lung cancer in South Africa: a forecast to 2025 based on smoking prevalence data. BMJ Open 5:e006993
Wang L, Yu C, Liu Y et al (2016) Lung cancer mortality trends in China from 1988 to 2013: new challenges and opportunities for the government. Int J Environ Res Public Health 13:1052
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30
Hong S, Mok Y, Jeon C et al (2016) Tuberculosis, smoking and risk for lung cancer incidence and mortality. Int J Cancer 139:2447–2455
Raaschou-Nielsen O, Andersen ZJ, Beelen R et al (2013) Air pollution and lung cancer incidence in 17 European cohorts: prospective analyses from the European Study of Cohorts for Air Pollution Effects (ESCAPE). Lancet Oncol 14:813–822
Mirtavoos-Mahyari H, Ghafouri-Fard S, Khosravi A, Motevaseli E, Esfahani-Monfared Z, Seifi S, Salimi B et al (2019) Circulating free DNA concentration as a marker of disease recurrence and metastatic potential in lung cancer. Clin Transl Med 8(1):14. https://doi.org/10.1186/s40169-019-0229-6
Phan TT, Tran BT, Nguyen ST, Ho TT, Hang Nguyen HT, Le VT, Le AT (2019) EGFR plasma mutation in prediction models for resistance with EGFR TKI and survival of non-small cell lung cancer. Clin Transl Med 8(1):4. https://doi.org/10.1186/s40169-019-0219-8
Virmani AK, Gazdar AF (2003) Tumor suppressor genes in lung cancer. Methods Mol Biol 222:97–115
Devarakonda S, Morgensztern D, Govindan R (2015) Genomic alterations in lung adenocarcinoma. Lancet Oncol 16:e342–e351
The World Health Organization (1982) Histological typing of lung tumours. Neoplasma 29:111–123
Solyanik GI (2010) Multifactorial nature of tumor drug resistance. Exp Oncol 32:181–185
Hesson LB, Cooper WN, Latif F (2007) Evaluation of the 3p21.3 tumour-suppressor gene cluster. Oncogene 26:7283–7301
Vos MD, Ellis CA, Bell A et al (2000) Ras uses the novel tumor suppressor RASSF1 as an effector to mediate apoptosis. J Biol Chem 275:35669–35672
Zhou Y, Zhang X, Klibanski A (2014) Genetic and epigenetic mutations of tumor suppressive genes in sporadic pituitary adenoma. Mol Cell Endocrinol 386:16–33
Shivakumar L, Minna J, Sakamaki T et al (2002) The RASSF1A tumor suppressor blocks cell cycle progression and inhibits cyclin D1 accumulation. Mol Cell Biol 22:4309–4318
Hamilton G, Yee KS, Scrace S et al (2009) ATM regulates a RASSF1A-dependent DNA damage response. Curr Biol 19:2020–2025
Li J, Wang F, Protopopov A et al (2004) Inactivation of RASSF1C during in vivo tumor growth identifies it as a tumor suppressor gene. Oncogene 23:5941–5949
Guo W, Dong Z, Guo Y et al (2016) Decreased expression and frequent promoter hypermethylation of RASSF2 and RASSF6 correlate with malignant progression and poor prognosis of gastric cardia adenocarcinoma. Mol Carcinog 55:1655–1666
Fukatsu A, Ishiguro F, Tanaka I et al (2014) RASSF3 downregulation increases malignant phenotypes of non-small cell lung cancer. Lung Cancer 83:23–29
Hesson L, Dallol A, Minna JD et al (2003) NORE1A, a homologue of RASSF1A tumour suppressor gene is inactivated in human cancers. Oncogene 22:947–954
Aoyama Y, Avruch J, Zhang XF (2004) Nore1 inhibits tumor cell growth independent of Ras or the MST1/2 kinases. Oncogene 23:3426–3433
Vos MD, Martinez A, Ellis CA et al (2003) The pro-apoptotic Ras effector Nore1 may serve as a Ras-regulated tumor suppressor in the lung. J Biol Chem 278:21938–21943
Liang YY, Deng XB, Zeng LS et al (2018) RASSF6-mediated inhibition of Mcl-1 through JNK activation improves the anti-tumor effects of sorafenib in renal cell carcinoma. Cancer Lett 432:75–83
Mi Y, Zhang D, Jiang W et al (2017) miR-181a-5p promotes the progression of gastric cancer via RASSF6-mediated MAPK signalling activation. Cancer Lett 389:11–22
Chen E, Yang F, He H et al (2016) Decreased level of RASSF6 in sporadic colorectal cancer and its anti-tumor effects both in vitro and in vivo. Oncotarget 7:19813–19823
Ma H, Chen X, Hu H et al (2018) Hypermethylation of MDFI promoter with NSCLC is specific for females, non-smokers and people younger than 65. Oncol Lett 15:9017–9024
Grasse S, Lienhard M, Frese S et al (2018) Epigenomic profiling of non-small cell lung cancer xenografts uncover LRP12 DNA methylation as predictive biomarker for carboplatin resistance. Genome Med 10:55
Toh TB, Lim JJ, Chow EK-H (2019) Epigenetics of hepatocellular carcinoma. Clin Transl Med 8(1):13. https://doi.org/10.1186/s40169-019-0230-0.
Wistuba II, Behrens C, Virmani AK et al (2000) High resolution chromosome 3p allelotyping of human lung cancer and preneoplastic/preinvasive bronchial epithelium reveals multiple, discontinuous sites of 3p allele loss and three regions of frequent breakpoints. Cancer Res 60:1949–1960
Lo KW, Kwong J, Hui AB et al (2001) High frequency of promoter hypermethylation of RASSF1A in nasopharyngeal carcinoma. Cancer Res 61:3877–3881
Dammann R, Li C, Yoon JH et al (2000) Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3. Nat Genet 25:315–319
Byun DS, Lee MG, Chae KS et al (2001) Frequent epigenetic inactivation of RASSF1A by aberrant promoter hypermethylation in human gastric adenocarcinoma. Cancer Res 61:7034–7038
Kuroki T, Trapasso F, Yendamuri S et al (2003) Allele loss and promoter hypermethylation of VHL, RAR-beta, RASSF1A, and FHIT tumor suppressor genes on chromosome 3p in esophageal squamous cell carcinoma. Cancer Res 63:3724–3728
Lee MG, Kim HY, Byun DS et al (2001) Frequent epigenetic inactivation of RASSF1A in human bladder carcinoma. Cancer Res 61:6688–6692
Wong N, Li L, Tsang K et al (2002) Frequent loss of chromosome 3p and hypermethylation of RASSF1A in cholangiocarcinoma. J Hepatol 37:633–639
Dammann R, Takahashi T, Pfeifer GP (2001) The CpG island of the novel tumor suppressor gene RASSF1A is intensely methylated in primary small cell lung carcinomas. Oncogene 20:3563–3567
Yu FX, Zhao B, Guan KL (2015) Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell 163:811–828
Grusche FA, Degoutin JL, Richardson HE et al (2011) The Salvador/Warts/Hippo pathway controls regenerative tissue growth in Drosophila melanogaster. Dev Biol 350:255–266
Dobrokhotov O, Samsonov M, Sokabe M, Hirata H (2018) Mechanoregulation and pathology of YAP/TAZ via Hippo and non-Hippo mechanisms. Clin Transl Med 7(1):23. https://doi.org/10.1186/s40169-018-0202-9
Hong JH, Hwang ES, McManus MT et al (2005) TAZ, a transcriptional modulator of mesenchymal stem cell differentiation. Science 309:1074–1078
Bora-Singhal N, Nguyen J, Schaal C et al (2015) YAP1 regulates OCT4 activity and SOX2 expression to facilitate self-renewal and vascular mimicry of stem-like cells. Stem Cells 33:1705–1718
Lehmann W, Mossmann D, Kleemann J et al (2016) ZEB1 turns into a transcriptional activator by interacting with YAP1 in aggressive cancer types. Nat Commun 7:10498
Pefani DE, Latusek R, Pires I et al (2014) RASSF1A-LATS1 signalling stabilizes replication forks by restricting CDK2-mediated phosphorylation of BRCA2. Nat Cell Biol 16:962–971. 1–8
Lange AW, Sridharan A, Xu Y et al (2015) Hippo/Yap signaling controls epithelial progenitor cell proliferation and differentiation in the embryonic and adult lung. J Mol Cell Biol 7:35–47
Lau AN, Curtis SJ, Fillmore CM et al (2014) Tumor-propagating cells and Yap/Taz activity contribute to lung tumor progression and metastasis. EMBO J 33:468–481
Luo SY, Sit KY, Sihoe AD et al (2014) Aberrant large tumor suppressor 2 (LATS2) gene expression correlates with EGFR mutation and survival in lung adenocarcinomas. Lung Cancer 85:282–292
Dhanasekaran SM, Balbin OA, Chen G et al (2014) Transcriptome meta-analysis of lung cancer reveals recurrent aberrations in NRG1 and Hippo pathway genes. Nat Commun 5:5893
Su LL, Ma WX, Yuan JF et al (2012) Expression of Yes-associated protein in non-small cell lung cancer and its relationship with clinical pathological factors. Chin Med J 125:4003–4008
Kim JM, Kang DW, Long LZ et al (2011) Differential expression of Yes-associated protein is correlated with expression of cell cycle markers and pathologic TNM staging in non-small-cell lung carcinoma. Hum Pathol 42:315–323
Noguchi S, Saito A, Horie M et al (2014) An integrative analysis of the tumorigenic role of TAZ in human non-small cell lung cancer. Clin Cancer Res 20:4660–4672
Song MS, Song SJ, Ayad NG et al (2004) The tumour suppressor RASSF1A regulates mitosis by inhibiting the APC-Cdc20 complex. Nat Cell Biol 6:129–137
Liu L, Tommasi S, Lee DH et al (2003) Control of microtubule stability by the RASSF1A tumor suppressor. Oncogene 22:8125–8136
Dallol A, Agathanggelou A, Tommasi S et al (2005) Involvement of the RASSF1A tumor suppressor gene in controlling cell migration. Cancer Res 65:7653–7659
Whang YM, Kim YH, Kim JS et al (2005) RASSF1A suppresses the c-Jun-NH2-kinase pathway and inhibits cell cycle progression. Cancer Res 65:3682–3690
Jimenez AP, Traum A, Boettger T et al (2017) The tumor suppressor RASSF1A induces the YAP1 target gene ANKRD1 that is epigenetically inactivated in human cancers and inhibits tumor growth. Oncotarget 8:88437–88452
Burbee DG, Forgacs E, Zochbauer-Muller S et al (2001) Epigenetic inactivation of RASSF1A in lung and breast cancers and malignant phenotype suppression. J Natl Cancer Inst 93:691–699
Ramirez JL, Taron M, Balana C et al (2003) Serum DNA as a tool for cancer patient management. Rocz Akad Med Bialymst 48:34–41
Schmiemann V, Bocking A, Kazimirek M et al (2005) Methylation assay for the diagnosis of lung cancer on bronchial aspirates: a cohort study. Clin Cancer Res 11:7728–7734
van der Drift MA, Prinsen CFM, Knuiman GJ et al (2012) Diagnosing peripheral lung cancer: the additional value of the Ras-association domain family 1A gene methylation and Kirsten rat sarcoma 2 viral oncogene homolog mutation analyses in washings in nondiagnostic bronchoscopy. Chest 141:169–175
Grawenda AM, O’Neill E (2015) Clinical utility of RASSF1A methylation in human malignancies. Br J Cancer 113:372–381
Drilon A, Sugita H, Sima CS et al (2014) A prospective study of tumor suppressor gene methylation as a prognostic biomarker in surgically resected stage I to IIIA non-small-cell lung cancers. J Thorac Oncol 9:1272–1277
Wang J, Lee JJ, Wang L et al (2004) Value of p16INK4a and RASSF1A promoter hypermethylation in prognosis of patients with resectable non-small cell lung cancer. Clin Cancer Res 10:6119–6125
Pelosi G, Fumagalli C, Trubia M et al (2010) Dual role of RASSF1 as a tumor suppressor and an oncogene in neuroendocrine tumors of the lung. Anticancer Res 30:4269–4281
Tomizawa Y, Kohno T, Kondo H et al (2002) Clinicopathological significance of epigenetic inactivation of RASSF1A at 3p21.3 in stage I lung adenocarcinoma. Clin Cancer Res 8:2362–2368
Kim DH, Kim JS, Ji YI et al (2003) Hypermethylation of RASSF1A promoter is associated with the age at starting smoking and a poor prognosis in primary non-small cell lung cancer. Cancer Res 63:3743–3746
Wang J, Wang B, Chen X et al (2011) The prognostic value of RASSF1A promoter hypermethylation in non-small cell lung carcinoma: a systematic review and meta-analysis. Carcinogenesis 32:411–416
Wang X (2018) Clinical trans-omics: an integration of clinical phenomes with molecular multiomics. Cell Biol Toxicol 34(3):163–166. https://doi.org/10.1007/s10565-018-9431-3
Qiao T, Wang X (2019) A new light of proteomics in cell biology and toxicology. Cell Biol Toxicol 35(4):289–291. https://doi.org/10.1007/s10565-019-09492-6
Wu D, Cheng Y, Wang X (2019) CSGT group. Definition of clinical gene tests. Cell Biol Toxicol 35(2):83–87. https://doi.org/10.1007/s10565-019-09464-w
Song D, Yang D, Powell CA, Wang X (2019) Cell-cell communication: old mystery and new opportunity. Cell Biol Toxicol 35(2):89–93. https://doi.org/10.1007/s10565-019-09470-y
Chirshev E, Oberg KC, Ioffe YJ, Unternaehrer JJ (2019) Let-7 as biomarker, prognostic indicator, and therapy for precision medicine in cancer. Clin Transl Med 8(1):24. https://doi.org/10.1186/s40169-019-0240-y
Malm J, Sugihara Y, Szasz M, Kwon HJ, Lindberg H, Appelqvist R, Marko-Varga G (2018) Biobank integration of large-scale clinical and histopathology melanoma studies within the European Cancer Moonshot Lund Center. Clin Transl Med 7(1):28. https://doi.org/10.1186/s40169-018-0203-8
Omrani S, Taheri M, Omrani MD, Arsang-Jang S, Ghafouri-Fard S (2019) The effect of omega-3 fatty acids on clinical and paraclinical features of intractable epileptic patients: a triple blind randomized clinical trial. Clin Transl Med 8(3):3. https://doi.org/10.1186/s40169-019-0220-2
Qi X, Yu C, Wang Y, Lin Y, Shen B (2019) Network vulnerability-based and knowledge-guided identification of microRNA biomarkers indicating platinum resistance in high-grade serous ovarian cancer. Clin Transl Med 8(1):28. https://doi.org/10.1186/s40169-019-0245-6
Fang M, Chen D, Yang CS (2007) Dietary polyphenols may affect DNA methylation. J Nutr 137:223S–228S
Wang XZ, Cheng Y, Wang KL et al (2016) Peperomin E reactivates silenced tumor suppressor genes in lung cancer cells by inhibition of DNA methyltransferase. Cancer Sci 107:1506–1519
Acknowledgement
This work is supported by the project of “Henan province clinical biomedical big data” by Department of science and technology of Henan Province (CXJD2019002).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Zhang, T., Li, Y., Zhang, H., Wang, X., Liu, X., Li, L. (2020). The Role of RASSF1 Methylation in Lung Carcinoma. In: Yu, B., Zhang, J., Zeng, Y., Li, L., Wang, X. (eds) Single-cell Sequencing and Methylation. Advances in Experimental Medicine and Biology, vol 1255. Springer, Singapore. https://doi.org/10.1007/978-981-15-4494-1_8
Download citation
DOI: https://doi.org/10.1007/978-981-15-4494-1_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4493-4
Online ISBN: 978-981-15-4494-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)