Abstract
DNA methylations, including global methylation pattern and specific gene methylation, are associated with pathogenesis and progress of pulmonary fibrosis. This chapter illustrates alteration of DNA methylation in pulmonary fibrosis as a predictive or prognostic factor. Treatment with the DNA methylation inhibitors will be an emerging anti-fibrosis therapy, although we are still in the pre-clinical stage of using epigenetic markers as potential targets for biomarkers and therapeutic interventions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
Idiopathic pulmonary fibrosis (IPF) is a serious form of pulmonary fibrosis, with which patients have the median survival time of about 2–3 years [1]. IPF is also a type of chronic lung disease characterized by a progressive scarring of the lung parenchyma and irreversible decline in lung function with hypoxemia and dyspnea. The prevalence and mortality of pulmonary fibrosis are on the rise with age, especially among people over 50 years old [2]. The incidence of IPF in men is higher than that in women and is more common in smokers [3]. Even after smoking cessation, the status of IPF cannot be improved. The pathogenesis of IPF is not completely clear and the clinical manifestation of IPF is highly variable. However, there are still some recognized potential risk factors such as environmental exposure, microbial agents, or gastroesophageal reflux. Recent studies have shown that gene expression and epigenetic regulation, especially the DNA methylation regulation, play an important role in the development of IPF [4,5,6].
DNA methylation is an inherited epigenetic process, involving the covalent transfer of the c-5 position of the DNA cytosine loop by the catalysis of DNA methyltransferases (DNMTs) [7]. The methylation alters gene function but does not change the sequence. The majority of DNA methylation occurs on the fifth carbon atom of cytosines that precede a guanine nucleotide or CpG sites [8]. DMA methylation is a dynamic and inheritable process. Methylation of CpG island promoters prevents the binding of transcription factors and results in gene silencing and repression. On the contrary, hypo-methylation and demethylation are associated with upregulation of gene expression [9]. DNMTs and methyl-binding proteins (MBPs) are major enzymes to catalyze DNA methylation [10], essential for transcriptional regulation and normal development and related to genomic imprinting, repression of transposable elements, X-chromosome inactivation, carcinogenesis, and aging [7, 11].
Epigenetic changes are associated with numerous diseases including cancers and pulmonary fibrosis, where large hypomethylated blocks of genomes and promoter hypermethylation of classic suppressor genes were found [8]. Studies on DNA methylation analysis confirmed that DNA methylation is common and important in pulmonary fibrosis. And numerous specific genes are involving in pathogenesis, such as Thy-1 (CD90), prostaglandin receptor 2 (PTGER2), cyclo-oxygenase-2 (COX-2), p14ARF, or chemokine IP-10 [12,13,14,15,16]. This chapter will focus on the global genome methylation pattern and targeted DNA methylation status in the pathogenesis of lung fibrosis, and then discuss the potential therapies of methylation inhibitors [17, 18].
4.2 Genome-Wide DNA Methylation in IPF
Methodologies for methylation measurement include next generation high throughput sequencing, whole genome bisulfite sequencing (WGBS), microarray, methylated DNA immunoprecipitation sequencing (Me DIP-Seq), bisulfite genomic sequence (BGS), and methylation-specific PCR (MSP). WGBS, Me DIP-Seq, microarray, and BGS are widely used in genome-wide DNA methylation analysis. For example, the human CpG islands microarray and WGBS were used to detect the alteration of the whole DNA extracted from the lung tissues of patients with or without IPF [15]. The extensive DNA methylation changes were found within CpG islands in IPF lung samples, different from methylation profiles of healthy, although partial methylated areas have many similarities [15]. The DNA methylation and RNA expression changed in lung tissue from IPF using human methylation chip and RNA hybridization chip. Altered DNA methylation is consistent with the mRNA expression of many genes, indicating the importance of DNA methylation in the pathogenesis of IPF [8]. Unfortunately, it is hard to clarify the alternations of DNA methylation within the individual cell type and difference between cell types, since most studies are based on the entire lung tissue.
The genome-wide differences in DNA methylation were detected in fibroblasts isolated from lung tissue of IPF patients, as compared with patients with lung nodules [19]. The methylation differences are mainly concentrated in genes associated with cell proliferation, extracellular matrix generation, potassium channel, and organ organogenesis and corresponded with alteration of gene expression at mRNA and protein levels [19].
4.3 IPF Specificity of Thy-1 DNA Methylation
Several specific genes were considered as IPF-specific and their DNA hypermethylation is consistent with the downregulated expression, such as Thy-1, COX-2, PTGER2, p14ARF, and chemokine IP-10 [13,14,15,16, 20, 21]. The reduction in the expression of those genes can directly induce the initiation of fibro-genesis, activation of fibroblast proliferation, and resistance to apoptosis [1]. Of those, Thy-1 cell surface antigen (Thy-1) is also known as CD90, a 25–37 kDa glycoprotein, localizing to lipid rafts and on the external leaflet of the lipid bilayer [22]. The activation of Thy-1 promotes T cell activation and affects multiple non-immunologic biological processes, such as cellular adhesion, migration, cell death, wound healing, neurite outgrowth, tumor repression, and fibrosis. Thy-1 as a highly conserved molecule has two membrane-bound and soluble forms and the biological role of Thy-1 dependent upon cell type and tissue specificity [23]. Thy-1 is often used as a marker for cell types and has a crucial effect on cell biology, of which the dysregulation is related to fibrotic diseases and malignancy [23]. Thy-1 located in chromosome 9 in mice and chromosome 11q22.3 in human are both initially expressed in the form of 161 a.a pro-form and have different post-transcriptional modifications [24]. Two different proteins encoded from the alleles differ only in position 89, of which one is arginine and the other is glycine. Thy-1 in human has only one allele for thymine, and the first 19 a.a pro-form positions the signal to targets the endoplasmic reticulum (ER) [25]. Thy-1 has two isoforms in mice: Thy-1.2 in Bal/c mice and Thy-1.1 in AKR mice have a glutamine and an arginine at the position 89, respectively. Genetic characteristics of Thy-1 genes are similar among human, mouse, and rat [26]. Human Thy-1 contains four exons, of which exon 1 (Fig. 4.1a, b) produce two mRNA splicing variants after transcription and exon 2 contains the translation starting site, exon 3 encodes the amino acids 7–106, and exon 4 is mainly responsible for the C-terminal end and poly-A tail [27] (Fig. 4.1).
Thy-1 gene structure. Exons 1a and 1b encode two distinct alternative spliced mRNA; exon 3 for the mature protein, and the 50-end of exon 4 for the trans-membrane sequence. Portions of the gene encoding for the mature Thy-1 protein are marked as light gray orthogons. Dark gray orthogons complete the exons
Thy-1 participates in a number of signaling cascades and acts as a universal signal modulator in proliferation, survival, cell adhesion, and cytokine/growth factor responses [23]. Thy-1 undergoes signal transduction in non-immunologic cells by integrins, growth factors, cytokines, and protein tyrosine kinases. The roles of those signaling cascades mainly focus on cell proliferation, apoptosis, cellular adhesion, and migration. Thy-1 interacts with itself, adaptors, scaffolds, or signaling molecules, such as reggies-1/2, Src family of C-terminal Src kinase (Csk)-binding protein (CBP) and protein tyrosine kinases (SFK), in the cell membrane of several cell types to convey signals to the cell interior. Thy-1 is an important component of protein complexes, to initiate cell signaling from rafts (Fig. 4.2). In addition, Thy-1 interacts with other receptors at the plasma membrane such as αVβ5 integrin in fibroblasts [28]. Thy-1(−) fibroblasts move faster and migrate more efficiently in wound healing than Thy-1(+) ones [28]. A mechanism to regulate fibroblast migration is involved in SFK and Rho GTPase activation [27]. It is proposed that Thy-1 expression regulates Src and FAK kinase activation, as well as phosphorylation of p190RhoGAP by increasing RhoA-GTP levels, to stress fiber and focal adhesion formation [29]. Decreased migration of Thy-1(+) fibroblast subpopulations may occur as the consequence of a complex Thy-1-triggered signaling process, in addition to passive Thy-1-to-matrix adhesion [27]. It implies Thy-1-dependent roles in fibroblast-matrix adhesion and migration.
Signaling induced by Thy-1. Thy-1 binds to its ligand (R) and undergoes molecular clustering at the plasma membrane. Thy-1 interacts with itself, with adaptors, scaffolds, or signaling molecules, such as reggies-1/2, Src family of C-terminal Src kinase (Csk)-binding protein (CBP) and protein tyrosine kinases (SFK), in the cell membrane of several cell types to convey signals to the cell interior
The loss of Thy-1 expression in lung fibroblasts correlates with many aspects of the fibrogenic phenotype including proliferation [25]. The proliferated myofibroblasts in the fibroblast foci were found Thy-1 negative in IPF, rather than in the normal fibroblasts [30]. Thy-1 can not only regulate the expression of myogenic gene, promote myofibroblastic differentiation, but also determine the survival of lung fibroblasts. Yan Y. Sanders et al. [20] demonstrated that Thy(−) fibroblasts proliferated in myofibroblastic foci, inhibiting the myofibroblast differentiation of fibroblasts, which was restored by DNA methyltransferase inhibitors. The epigenetic downregulation of Thy-1 occurred in cell transformation and clinical malignant tumor [20]. Rat lung fibroblasts without Thy-1 on the surface, low expression of myogenic genes and low protein levels of sarcomeric myosin, α-SMA, and MyoD, had high responses to pro-myofibroblastic stimuli including TGF-β [30].
Loss of Thy-1 expression appears to be associated with the differentiation of myofibroblasts both in mouse bleomycin model and IPF patients [31]. The relation between Thy-1 and myofibroblasts phenotype seems to be tissue-specific and dependent. Loss of Thy-1 expression also resulted in the hypermethylation of the Thy-1 promoter in IPF Samples and was restored through demethylation, similar between human and rat lung fibroblasts [20].
4.4 IPF Specificity of COX-2 DNA Methylation
Cyclooxygenases (COXs) are a 67–72 kDa integral membrane protein, are located on the nuclear membrane and the endoplasmic reticulum (ER), and contain three isoforms [32]. COX-1 is expressed constitutively like “housekeeping” enzyme associated with homeostasis, COX-2 is the inducible form and is upregulated in both inflammation and cancer, and COX-3 is expressed in spinal cord and brain although its functions remain unclear [33]. Cyclooxygenase-2 (COX-2) is referred to prostaglandin endoperoxide synthase (PTGS)I as a key enzyme that catalyzes the conversion of arachidonic acid (AA) to prostaglandins (PGs) [34]. COX-2 plays a crucial role in some pathophysiological processes, including angiogenesis, inflammation, tumorigenesis, and tumor drug resistance, and becomes a new target for cancer treatment [35]. In solid tumors such as colorectal cancer, prostate cancer, breast cancer, and most recently hematological malignancies, COX-2 mainly functions as a regulator of cell proliferation and apoptosis [33]. The activation and overexpression of COX-2 were found in tumor cells related to tumor progression and aggressiveness [36]. COX-2 expression could be induced by anticancer chemoradiotherapy, resulting in drug resistance [36]. The inhibition of COX-2 was proposed as an attractive new strategy for cancer treatment in patients [37]. Non-steroidal anti-inflammatory drugs (NSAIDs), broad spectrum COX-2-inhibitors, or COX-2-specific inhibitors were found to have side-effects, such as myocardial infarction [36]. The development of new anti-COX-2 drugs with less side-effects seems particularly urgent [34, 38].
COX-1 gene is located on chromosome 9 (9q32-9q33.3), nearly 40 kilobase (kb) pairs, containing 11 exons and its mRNA is 2.8 kb. COX-2 is located on chromosome 1 (1q25.2-25.3), containing ten exons approximately 8.3 kb and transcript about 4.5 kb [39]. In the flanking region of COX-2, there are 50 bps of the regulation area of gene transcription, containing a TATA box and a few putative transcription-factor binding sites of NF-IL-6, NF-κB, and a TGF-β response element, which demonstrates a complex combination of the factors associated with COX-2 gene regulation [40]. Single nucleotide polymorphism (SNP) in the gene promoter affects transcription of COX-2 gene. The most frequently functional polymorphisms of COX-2 gene, _765G>C (rs20417) and _1195G>A (rs689466), are correlated with inflammatory disorders, such as chronic periodontitis [41], inflammatory bowel diseases, and subclinical atherosclerosis [41]. This is probably because those gene polymorphisms may alter the function of COX-2 by regulation of COX-2 expression and affect the synthesis of prostaglandins in the pathogenesis of inflammatory diseases [42].
Prostaglandin E2 (PGE2), the major catalyzed product of COX-2, plays a key role in the tumorigenesis of colorectal cancer [43]. The COX-2/PGE2-JAK2/STAT3 signaling pathway may be the drug target for berberine to mediate the effect on metastasis and invasiveness of cancer. The berberine reduced COX-2/PGE2 levels, inhibited JAK2/STAT3 activation, decreased expression of downstream target genes MMP-2/-9, and caused less metastasis and invasiveness in cancer [44] (Fig. 4.3). PGE2 is associated with occurrence of malignant tumors and plays a beneficial role in lung fibrotic diseases. This is partially due to the function of PGE2 to limit the proliferation of lung fibroblasts and to inhibit myofibroblast differentiation, migration, and collagen secretion. Figure 4.4 diagrams the homeostatic and anti-fibrotic behavior of PGE2 signaling pathway in fibroblasts and lung epithelial cells (AECs) [45].
COX-2/PGE2-JAK2/STAT3 signaling pathway. PGE2, the main catalyzed product of COX-2 from arachidonic acid, could bind to the EP receptor on the cell membrane, thereby activating the JAK2, followed by the phosphorylating of STAT3 in the Tyr705 site. Berberine inhibits invasion and metastasis of colorectal cancer cells via COX-2/PGE2 mediated JAK2/STAT3 signaling pathway
The expression of COX-2 was downregulated in IPF and upregulated in COPD as well as in IPF and sclerosis [46, 47]. COX-2 downregulation and reduced PGE2 production are related to myofibroblasts in the development and progression of IPF [48]. The downregulation of COX-2 could reduce PGE2 and induce the continuous proliferation of fibroblasts, which is considered as a new viewpoint in the pathogenesis of IPF [49]. Lung fibroblasts derived from IPF patients were unable to induce PGE2 synthesis, even if stimulated by proinflammatory cytokines and LPS, probably due to the abnormal expression of COX-2 [45, 50]. In patients with IPF, the PGE2 level of bronchoalveolar lavage fluid was significantly lower than that of normal individuals, which is because PGE2 could reduce the proliferation of fibroblast and collagen aggregation by inhibiting COX-2-dominated synthesis and promotion of degradation, beneficial for inhibiting pulmonary fibrosis [51].
COX-2 was downregulated in lung tissue from patients with IPF [15, 52]. By upregulation of DNMT3a expression, PGE2 increases the gene-specific DNA methylation of lung fibroblasts, such as MGMT gene and IGFBP2 gene [53].
The transcriptional regulatory factor c8orf4 for COX-2 was demethylated via 5-AZAdc, a DNA methylation inhibitor to reverse decreased level of COX-2 mRNA in a dose-dependent pattern [15, 53]. C8orf4 regulates the expression of COX-2 in lung fibroblasts by binding of the proximal promoter by the hypermethylation of the transcription regulator as an indirect epigenetic mechanism to regulate COX-2 expression and COX-2 derived PGE2 synthesis in pulmonary fibrosis [15].
4.5 p14ARF and Function
The p14ARF protein as a tumor suppressor protein is an alternate reading frame protein (ARF) encoded by CDKN2A gene. ARF is a 14 kDa, 132 a.a protein named p14ARF in human, and a 19 kDa, 169 a.a protein named p19ARF in mice [54]. P14ARF is a cell cycle regulation protein to block the cell cycle in the G1 and G2 phases and inhibit the growth of abnormal cells by activating p53 indirectly [55]. p14ARF protein binds to and interferes with the Mdm2 protein, a p53 negative-regulator, and then stabilizes and activates p53 pathway [54, 56]. The role of p14ARF in carcinogenesis was evidenced by the finding that ARF-null mice have a high tendency to induce tumors, e.g., carcinomas, gliomas, lymphomas, and sarcomas, leading to death early in life [57].
The INK4a–ARF locus (CDKN2A in humans) on chromosome 9p21 encodes two structure-similar tumor suppressor proteins with different functions, p14ARF (p19ARF in the mouse) and p16INK4a to indirectly control the activities of p53 and the retinoblastoma protein (RB) transcription factor, respectively [58]. p14ARF and INK4a mRNA consist of 3 exons of which exons 2 and 3 are the same with two different exon 1 transcripts (α and β) [59, 60]. Although p14ARF has an unrelated structure, it can also cause cell cycle arrest in G1 and G2 phase [61]. P14ARF gene as a tumor suppressor gene plays an important role in the progression and pathogenesis of tumor, since it is usually mutated or deleted [62, 63].
The dysfunction of the p14ARF-Mdm2-p53 pathway, also known as p53 pathway, is one of the most important signals of cancer pathogenesis. The p14ARF in the p53 pathway binds with Mdm2 in the nucleolus, resulting in the inability of Mdm2 to degrade p53 [64, 65] (Fig. 4.5). The activity of Mdm2 can be inhibited by p14ARF, to indirectly block the degradation of p53. When p53 is activated, the consequences of the ARF-p53 binding depend on the cell cycle state [66]. P14ARF controls the expression of p53, and then activated p53 secondarily regulates the expression of p14ARF by negative feedback [67]. Overexpression of p14ARF in the nucleus contributes to the loss of shuttling ability of Mdm2 and induces p53 mutations [68]. This pathway is inactivated by p14ARF deletion, p53 mutation, or amplification of Mdm2, which is complex and interactive but common and important.
The p53/p14ARF signaling pathway is often downregulated in patients with colorectal cancer, and p14ARF is highly methylated in the early stages of colorectal cancer [69]. The methylation of p14ARF may have predictive value for early colorectal cancer patients, but not as a prognostic factor. The target drug for p14ARF demethylation may be a new direction for the development of new colorectal cancer drugs [69]. The p14ARF gene can be inactivated in many cancers, due to deletion, promoter hypermethylation, or mutations [69]. In the evolution of oligodendrogliomas, the hypermethylation-resulted aberrant p14ARF expression and the deletions of p14ARF/p16INK4a are associated with the progression to anaplastic oligodendroglioma [70, 71]. Studies on the methylation status of the p14ARF promoter suggested that p14ARF can be a useful biomarker for the pathological TNM stage, prognosis, and clinical outcome of cancer patients [72]. Homozygous deletion of the p14ARF gene loci was detected in multiple carcinomas and was associated with tumorigenesis. DNA methylation can regulate p14ARF mRNA levels, and the methylation status of p14ARF is related to the occurrence of primary liver cancer and TNM staging [73]. The promoter methylation status of p14ARF in fibroblasts isolated from IPF and normal lung demonstrated that hypermethylated p14ARF occurred in half of the IPF fibroblasts and was correlated with the decreased expression of the gene and protein as well as increased resistance to apoptosis [16].
Hypermethylation and downregulated expression of PTGER2 also play an important role in the development of IPF. Levels of DNA hypermethylation were higher in fibroblasts isolated from mice and human lungs with pulmonary fibrosis, leading to a decrease in EP2 expression level and PGE2 resistance [14]. Therapies with DNA methylation inhibitors (e.g., 5-Aza-2′-deoxycytidine and zebularine) reversed the reduced mRNA and protein expression of EP2, and restored PGE2 activities in fibrotic fibroblasts. Those results indicate that DNA hypermethylation play the decisive role in the downregulation of PTGER2 expression and subsequent PGE2 resistance. The enhancement of Akt signal transduction may be a new mechanism of the promotion of DNA hypermethylation in the formation of lung fibrosis [14].
4.6 Conclusion and Prospective
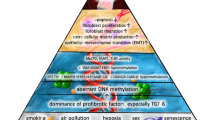
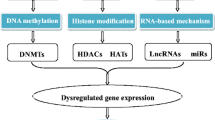
DNA methylation is one of mechanisms by which the epigenetic regulation plays a crucial role in lung fibrosis, cancer, and chronic diseases. Global methylation pattern and specific gene methylation status as an important regulatory factor contribute to the development of pulmonary fibrosis. DNA methylation of associated genes is associated with the occurrence and progression of pulmonary fibrosis and change the phenotype and destiny of fibroblasts through the regulation of cell activation, differentiation, and balance of fibrotic and anti-fibrotic gene expressions.
Methylation patterns and severities of the promoter regions of Thy-1, COX-2, p14ARF, and PTGER2 genes should be considered as disease-specific biomarkers to predict the occurrence and development of IPF. The intracellular mechanisms and heterogeneity of DNA methylation in the regulation of signal pathway activities should be investigated by single-cell DNA and RNA sequencing [74,75,76]. The promoter methylation of the target genes can contribute to the pathogenesis and development of pulmonary fibrosis through multiple signal pathways, which should be furthermore identified and validated with advanced biotechnologies [77,78,79,80,81].
References
Sundarakrishnan A, Chen Y, Black LD, Aldridge BB, Kaplan DL (2018) Engineered cell and tissue models of pulmonary fibrosis. Adv Drug Deliv Rev 129:78
King TE Jr, Pardo A, Selman M (2011) Idiopathic pulmonary fibrosis. Lancet 378(9807):1949–1961
Rangarajan S, Locy ML, Luckhardt TR, Thannickal VJ (2016) Targeted therapy for idiopathic pulmonary fibrosis: where to now? Drugs 76(3):291–300
Macneal K, Schwartz DA (2012) The genetic and environmental causes of pulmonary fibrosis. Proc Am Thorac Soc 9(3):120–125
Ziobro R, Henry B, Edwards MJ, Lentsch AB, Gulbins E (2013) Ceramide mediates lung fibrosis in cystic fibrosis. Biochem Biophys Res Commun 434(4):705–709
Wolters PJ, Collard HR, Jones KD (2014) Pathogenesis of idiopathic pulmonary fibrosis. Ann Rev Pathol Mech Dis 9:157–179
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38(1):23–38
Sanders YY, Ambalavanan N, Halloran B, Zhang X, Liu H, Crossman DK et al (2012) Altered DNA methylation profile in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 186(6):525–535
Marchal C, Miotto B (2015) Emerging concept in DNA methylation: role of transcription factors in shaping DNA methylation patterns. J Cell Physiol 230(4):743–751
Du Q, Luu PL, Stirzaker C, Clark SJ (2015) Methyl-CpG-binding domain proteins: readers of the epigenome. Epigenomics 7(6):1051–1073
Gendrel AV, Heard E (2014) Noncoding RNAs and epigenetic mechanisms during X-chromosome inactivation. Annu Rev Cell Dev Biol 30:561–580
Rabinovich EI, Kapetanaki MG, Steinfeld I, Gibson KF, Pandit KV, Yu G et al (2012) Global methylation patterns in idiopathic pulmonary fibrosis. PLoS One 7(4):e33770
Robinson CM, Neary R, Levendale A, Watson CJ, Baugh JA (2012) Hypoxia-induced DNA hypermethylation in human pulmonary fibroblasts is associated with Thy-1 promoter methylation and the development of a pro-fibrotic phenotype. Respir Res 13(1):74
Huang SK, Fisher AS, Scruggs AM, White ES, Hogaboam CM, Richardson BC et al (2010) Hypermethylation of PTGER2 confers prostaglandin E2 resistance in fibrotic fibroblasts from humans and mice. Am J Pathol 177(5):2245–2255
Evans IC, Barnes JL, Garner IM, Pearce DR, Maher TM, Shiwen X et al (2016) Epigenetic regulation of cyclooxygenase-2 by methylation of c8orf4 in pulmonary fibrosis. Clin Sci 130(8):575–586
Cisneros J, Hagood J, Checa M, Ortiz-Quintero B, Negreros M, Herrera I et al (2012) Hypermethylation-mediated silencing of p14ARF in fibroblasts from idiopathic pulmonary fibrosis. Am J Phys Lung Cell Mol Phys 303(4):L295–L303
Zhang X, Hu M, Lyu X, Li C, Thannickal VJ, Sanders YY (2017) DNA methylation regulated gene expression in organ fibrosis. Biochim Biophys Acta Mol basis Dis 1863(9):2389–2397
Spagnolo P, Sverzellati N, Rossi G, Cavazza A, Tzouvelekis A, Crestani B et al (2015) Idiopathic pulmonary fibrosis: an update. Ann Med 47(1):15–27
Huang SK, Scruggs AM, McEachin RC, White ES, Peters-Golden M (2014) Lung fibroblasts from patients with idiopathic pulmonary fibrosis exhibit genome-wide differences in DNA methylation compared to fibroblasts from nonfibrotic lung. PLoS One 9(9):e107055
Sanders YY, Pardo A, Selman M, Nuovo GJ, Tollefsbol TO, Siegal GP et al (2008) Thy-1 promoter hypermethylation: a novel epigenetic pathogenic mechanism in pulmonary fibrosis. Am J Respir Cell Mol Biol 39(5):610–618
Roman J, Mutsaers SE (2018) Epigenetic control of CXCL10: regulating the counterregulator in idiopathic pulmonary fibrosis. Am J Respir Cell Mol Biol 58:419
Rege TA, Hagood JS (2006) Thy-1 as a regulator of cell-cell and cell-matrix interactions in axon regeneration, apoptosis, adhesion, migration, cancer, and fibrosis. FASEB J 20(8):1045–1054
Hagood JS (2019) Thy-1 as an integrator of diverse extracellular signals. Front Cell Dev Biol 7:26
Abd-Elmotelb M (2012) Thy-1 (CD90) Expression in dental pulp stem cells: Guy’s, King’s and St. Thomas’s School of Dentistry
Sauzay C, Voutetakis K, Chatziioannou A, Chevet E, Avril T (2019) CD90/Thy-1, a cancer-associated cell surface signaling molecule. Front Cell Dev Biol 7:66
Bradley JE, Ramirez G, Hagood JS (2009) Roles and regulation of Thy-1, a context-dependent modulator of cell phenotype. Biofactors 35(3):258–265
Herrera-Molina R, Valdivia A, Kong M, Alvarez A, Cardenas A, Quest AF et al (2013) Thy-1-interacting molecules and cellular signaling in cis and trans. Int Rev Cell Mol Biol 305:163–216
Herrera-Molina R, Valdivia A, Kong M, Alvarez A, Cárdenas A, Quest AF et al (2013) Thy-1-interacting molecules and cellular signaling in cis and trans. Int Rev Cell Mol Biol 305:163–216
Rege TA, Pallero MA, Gomez C, Grenett HE, Murphy-Ullrich JE, Hagood JS (2006) Thy-1, via its GPI anchor, modulates Src family kinase and focal adhesion kinase phosphorylation and subcellular localization, and fibroblast migration, in response to thrombospondin-1/hep I. Exp Cell Res 312(19):3752–3767
Sanders YY, Kumbla P, Hagood JS (2007) Enhanced myofibroblastic differentiation and survival in Thy-1(-) lung fibroblasts. Am J Respir Cell Mol Biol 36(2):226–235
Nicola T, Hagood JS, James ML, Macewen MW, Williams TA, Hewitt MM et al (2009) Loss of Thy-1 inhibits alveolar development in the newborn mouse lung. Am J Phys Lung Cell Mol Phys 296(5):L738–L750
Yu T, Lao X, Zheng H (2016) Influencing COX-2 activity by COX related pathways in inflammation and cancer. Mini-Rev Med Chem 16(15):1230–1243
Sobolewski C, Cerella C, Dicato M, Ghibelli L, Diederich M (2010) The role of cyclooxygenase-2 in cell proliferation and cell death in human malignancies. Int J Cell Biol 2010:215158
Liao X, Wang W, Fan C, Yang N, Zhao J, Zhang Y et al (2017) Prokaryotic expression, purification and characterization of human cyclooxygenase-2. Int J Mol Med 40(1):75–82
Ghosh N, Chaki R, Mandal V, Mandal SC (2010) COX-2 as a target for cancer chemotherapy. Pharmacol Rep 62(2):233–244
Khan Z, Khan N, Tiwari RP, Sah NK, Prasad GB, Bisen PS (2011) Biology of Cox-2: an application in cancer therapeutics. Curr Drug Targets 12(7):1082–1093
Gulyas M, Mattsson JSM, Lindgren A, Ek L, Lamberg Lundström K, Behndig A et al (2018) COX-2 expression and effects of celecoxib in addition to standard chemotherapy in advanced non-small cell lung cancer. Acta Oncol 57(2):244–250
Zarghi A, Arfaei S (2011) Selective COX-2 inhibitors: a review of their structure-activity relationships. Iran J Pharmaceut Res 10(4):655–683
Tay A, Squire JA, Goldberg H, Skorecki K (1994) Assignment of the human prostaglandin-endoperoxide synthase 2 (PTGS2) gene to 1q25 by fluorescence in situ hybridization. Genomics 23(3):718–719
Park GY, Christman JW (2006) Involvement of cyclooxygenase-2 and prostaglandins in the molecular pathogenesis of inflammatory lung diseases. Am J Phys Lung Cell Mol Phys 290(5):L797–L805
Piranda DN, Festa-Vasconcellos JS, Amaral LM, Bergmann A, Vianna-Jorge R (2010) Polymorphisms in regulatory regions of cyclooxygenase-2 gene and breast cancer risk in Brazilians: a case-control study. BMC Cancer 10:613
Andersen V, Nimmo E, Krarup HB, Drummond H, Christensen J, Ho GT et al (2011) Cyclooxygenase-2 (COX-2) polymorphisms and risk of inflammatory bowel disease in a Scottish and Danish case-control study. Inflamm Bowel Dis 17(4):937–946
Mizuno R, Kawada K (2019) Prostaglandin E2/EP signaling in the tumor microenvironment of colorectal cancer. Int J Mol Sci 20(24):E6254
Liu X, Ji Q, Ye N, Sui H, Zhou L, Zhu H et al (2015) Berberine inhibits invasion and metastasis of colorectal cancer cells via COX-2/PGE2 mediated JAK2/STAT3 signaling pathway. PLoS One 10(5):e0123478
Bozyk PD, Moore BB (2011) Prostaglandin E2 and the pathogenesis of pulmonary fibrosis. Am J Respir Cell Mol Biol 45(3):445–452
Coward WR, Feghali-Bostwick CA, Jenkins G, Knox AJ, Pang L (2014) A central role for G9a and EZH2 in the epigenetic silencing of cyclooxygenase-2 in idiopathic pulmonary fibrosis. FASEB J 28(7):3183–3196
Cheng J, Dackor RT, Bradbury JA, Li H, DeGraff LM, Hong LK et al (2016) Contribution of alveolar type II cell-derived cyclooxygenase-2 to basal airway function, lung inflammation, and lung fibrosis. FASEB J 30(1):160–173
Gabasa M, Royo D, Molina-Molina M, Roca-Ferrer J, Pujols L, Picado C et al (2013) Lung myofibroblasts are characterized by down-regulated cyclooxygenase-2 and its main metabolite, prostaglandin E2. PLoS One 8(6):e65445
Maher TM, Evans IC, Bottoms SE, Mercer PF, Thorley AJ, Nicholson AG et al (2010) Diminished prostaglandin E2 contributes to the apoptosis paradox in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 182(1):73–82
Wilborn J, Crofford LJ, Burdick MD, Kunkel SL, Strieter RM, Peters-Golden M (1995) Cultured lung fibroblasts isolated from patients with idiopathic pulmonary fibrosis have a diminished capacity to synthesize prostaglandin E2 and to express cyclooxygenase-2. J Clin Invest 95(4):1861–1868
McAnulty RJ, Hernandez-Rodriguez NA, Mutsaers SE, Coker RK, Laurent GJ (1997) Indomethacin suppresses the anti-proliferative effects of transforming growth factor-beta isoforms on fibroblast cell cultures. Biochem J 321(Pt 3):639–643
Xaubet A, Fu W, Li M (2010) A haplotype of cyclooxygenase-2 gene is associated with idiopathic pulmonary fibrosis. Sarcoid Vasc Diff Lung Dis 27(2):121–130
Huang SK, Scruggs AM, Donaghy J, McEachin RC, Fisher AS, Richardson BC et al (2012) Prostaglandin E2 increases fibroblast gene-specific and global DNA methylation via increased DNA methyltransferase expression. FASEB J 26(9):3703–3714
Sherr CJ (2006) Divorcing ARF and p53: an unsettled case. Nat Rev Cancer 6(9):663–673
Silva J, Silva JM, Dominguez G, Garcia JM, Cantos B, Rodriguez R et al (2003) Concomitant expression of p16INK4a and p14ARF in primary breast cancer and analysis of inactivation mechanisms. J Pathol 199(3):289–297
Cortot AB, Younes M, Martel-Planche G, Guibert B, Isaac S, Souquet PJ et al (2014) Mutation of TP53 and alteration of p14(arf) expression in EGFR- and KRAS-mutated lung adenocarcinomas. Clin Lung Cancer 15(2):124–130
Stott FJ, Bates S, James MC, McConnell BB, Starborg M, Brookes S et al (1998) The alternative product from the human CDKN2A locus, p14ARF, participates in a regulatory feedback loop with p53 and MDM2. EMBO J 17(17):5001–5014
Zhang Y, Hyle J, Wright S, Shao Y, Zhao X, Zhang H et al (2019) A cis-element within the ARF locus mediates repression of p16INK4A expression via long-range chromatin interactions. Proc Natl Acad Sci
Inoue K, Fry EA (2018) Tumor suppression by the EGR1, DMP1, ARF, p53, and PTEN Network. Cancer Investig 36(9-10):520–536
López F, Sampedro T, Llorente JL, Hermsen M, Álvarez-Marcos C (2017) Alterations of p14 ARF, p15 INK4b, and p16 INK4a genes in primary laryngeal squamous cell carcinoma. Pathol Oncol Res 23(1):63–71
Bayramov B, Gunes S, Buyukalpelli R, Aydın O, Henkel R (2018) Promoter methylation analysis of CDH1 and p14ARF genes in patients with urothelial bladder cancer. OncoTarget Ther 11:4189
Kotake Y, Naemura M, Murasaki C, Inoue Y, Okamoto H (2015) Transcriptional regulation of the p16 tumor suppressor gene. Anticancer Res 35(8):4397–4401
Flaherty K, Jones D, Yussuf S, Davis S, Huselid E, Wang W et al (2015) Conditional mouse and zebrafish models of INK4-mediated tumor suppression reveal ARF-independent regulation of cellular senescence. Cancer Res 75:Abstract nr 1264
Werner LR, Huang S, Francis DM, Armstrong EA, Ma F, Li C et al (2015) Small molecule inhibition of MDM2–p53 interaction augments radiation response in human tumors. Mol Cancer Ther 14(9):1994–2003
Fazal L, Ahn M, Bevan L, Buck I, Castro J, Chessari G et al (2018) Development of a potent class of small molecule inhibitors of the MDM2-p53 protein-protein interaction. Cancer Res 78:Abstract nr 1652
Yaddanapudi SCS (2016) The human ARF tumor suppressor regulates drosha nucleolar localization and rRNA processing activity: Washington University in St. Louis
Joerger AC, Fersht AR (2016) The p53 pathway: origins, inactivation in cancer, and emerging therapeutic approaches. Annu Rev Biochem 85:375–404
Cui L, Zhou F, Chen C, Wang CC (2019) Overexpression of CCDC69 activates p14 ARF/MDM2/p53 pathway and confers cisplatin sensitivity. J Ovar Res 12(1):4
Chaar I, Amara S, Elamine OE, Khiari M, Ounissi D, Khalfallah T et al (2014) Biological significance of promoter hypermethylation of p14/ARF gene: relationships to p53 mutational status in Tunisian population with colorectal carcinoma. Tumour Biol 35(2):1439–1449
Delmonico L, dos Santos MA, Franco MF, Esteves EB, Scherrer L, de Moura Gallo CV et al (2015) CDKN2A (p14ARF/p16INK4a) and ATM promoter methylation in patients with impalpable breast lesions. Hum Pathol 46(10):1540–1547
Tripathi A, Sharma R, Kejriwal N, Ambasta RK, Kumar P (2016) Epigenetic post transcriptional mutation in neuro-oncology. In: Epigenetic advancements in cancer. Springer, New York, NY, pp 177–205
Mansouri A, Karamchandani J, Das S (2017) Molecular genetics of secondary glioblastoma. In: Glioblastoma. Codon Publications, Brisbane, QLD
Zhang H, Nie W, Huang F (2015) The correlation relationship between P14ARF gene DNA methylation and primary liver cancer. Med Sci Monit 21:3077
Song D, Yang D, Powell CA, Wang X (2019) Cell-cell communication: old mystery and new opportunity. Cell Biol Toxicol 35(2):89–93
Wang W, Gao D, Wang X (2018) Can single-cell RNA sequencing crack the mystery of cells? Cell Biol Toxicol 34(1):1–6
Zeng Y, Chen X, Gao H, Wang X (2018) An artificial intelligent single cell is part of the cell dream world. Cell Biol Toxicol 34(4):247–249
Mirtavoos-Mahyari H, Ghafouri-Fard S, Khosravi A, Motevaseli E, Esfahani-Monfared Z, Seifi S et al (2019) Circulating free DNA concentration as a marker of disease recurrence and metastatic potential in lung cancer. Clin Transl Med 8(1):14
El Khoury D, Fayjaloun S, Nassar M, Sahakian J, Aad PY (2019) Updates on the effect of mycotoxins on male reproductive efficiency in mammals. Toxins (Basel) 11(9):E515
Sanchez A, Kuras M, Murillo JR, Pla I, Pawlowski K, Szasz AM et al (2019) Novel functional proteins coded by the human genome discovered in metastases of melanoma patients. Cell Biol Toxicol
Wang W, Hou J, Zheng N, Wang X, Zhang J (2019) Keeping our eyes on CRISPR: the “Atlas” of gene editing. Cell Biol Toxicol 35(4):285–288
Marcell Szasz A, Malm J, Rezeli M, Sugihara Y, Betancourt LH, Rivas D et al (2019) Challenging the heterogeneity of disease presentation in malignant melanoma-impact on patient treatment. Cell Biol Toxicol 35(1):1–14
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Zhou, S., Wang, X., Gao, H., Zeng, Y. (2020). DNA Methylation in Pulmonary Fibrosis. In: Yu, B., Zhang, J., Zeng, Y., Li, L., Wang, X. (eds) Single-cell Sequencing and Methylation. Advances in Experimental Medicine and Biology, vol 1255. Springer, Singapore. https://doi.org/10.1007/978-981-15-4494-1_4
Download citation
DOI: https://doi.org/10.1007/978-981-15-4494-1_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4493-4
Online ISBN: 978-981-15-4494-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)