Abstract
Prolonged depolarizing pulses that last seconds to minutes cause slow inactivation of Na+ channels, which regulates neuron and myocyte excitability by reducing availability of inward current. In neurons, slow inactivation has been linked to memory of previous excitation and in skeletal muscle it ensures myocytes are able to contract when K+ is elevated. The molecular mechanisms underlying slow inactivation are unclear even though it has been studied for 50+ years. This chapter reviews what is known to date regarding the definition, measurement, and mechanisms of voltage-gated Na+ channel slow inactivation.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Background
1.1 What Is Slow Inactivation?
Slow inactivation of voltage-gated Na+ channels is a gating phenomenon, distinct from fast inactivation, whereby prolonged depolarizing pulses reduce the number of channels available to provide excitatory inward current.
1.2 Physiological Relevance
In neurons and muscle, initiation of action potentials is regulated by the opening and closing of rapidly gated Na+ channels that produce inward current and depolarize the membrane. Thus, slow inactivation, which reduces the number of channels that are available to open, regulates excitability.
A hallmark of slow inactivation is that its onset and recovery take place over time domains that span multiple orders of magnitude, such that channels subjected to longer depolarizing pulses require longer times at negative potential for current to be restored (Narahashi 1964; Toib et al. 1998). Consequently, memory of previous excitation can be encoded through the slow inactivation of the Na+ channels. The lack of a specific time scale over which slow inactivation takes place has been linked to neuronal memory (Toib et al. 1998), which exhibits a similar lack of identifiable time scales (Marom 2010).
In skeletal muscle, slow inactivation prevents contraction when serum K+ is elevated. Patients with mutations that disrupt slow inactivation in the muscle Na+ channel suffer from a condition known as hyperkalemic periodic paralysis (HyperPP) (Bendahhou et al. 2002). As a result, when high levels of extracellular K+ marginally elevate the myocyte membrane potential, Na+ channels that would normally be slow inactivated produce a persistent inward Na+ current that further depolarizes the membrane. This additional depolarization causes a large fraction of Na+ channels to fast inactivate, leading to membrane in-excitability. As a result, when normal neuronal routes of excitation are activated, the muscle is unable to respond (Lengele et al. 2008; Jurkat-Rott et al. 2009; Jurkat-Rott and Lehmann-Horn 2006; George 2005).
Cardiac Na+ channels exhibit much less slow inactivation in comparison to skeletal muscle and neuronal isoforms, showing incomplete inactivation even after very long pulses (O’Reilly et al. 1999). Physiologically, reduced slow inactivation for these channels would be expected, due to the necessity of maintaining cardiac excitability. However, slow inactivation is still relevant to the cardiac rhythm as it is likely to be regulated via the limited population of channels that do inactivate, because the rate at which the cardiac excitation wavefront propagates, known as conduction velocity, is directly proportional to peak Na+ current (Shaw and Rudy 1997). Moreover, the ability of the myocardium to sustain an arrhythmia is known to depend on several factors, including the length of the path the arrhythmia can travel, the excitability of the tissue, and the conduction velocity. Slow inactivation affects two of these parameters, excitability and conduction velocity. Mutations that predispose patients to arrhythmia suggest an antiarrhythmic role for slow inactivation. For example, Vilin et al. showed that a double mutation in the cardiac Na+ channel not only alters rapid gating events but also significantly reduces slow inactivation, positing that the removal of slow inactivation contributes to the propensity of patients to experience an arrhythmia (Vilin et al. 2001a; Vilin and Ruben 2001).
While most channel research is focused on rapid gating events, the slower transitions also play a key physiological role. Moreover, these movements have a degree of permanence that allow them to modulate phenomena that take place over seconds and minutes, such as the memory of a neuron or the sustenance of an arrhythmia. As such, a large number of therapeutically useful small molecules have been designed to target the slow-inactivated states including anesthetics (Balser et al. 1996a), anticonvulsants (Schauf 1987), antiarrhythmics (Sheets et al. 2010), and insecticides (Song et al. 2011).
The physiological potential of slow gating movements was realized early on in the study of Na+ channels and is discussed in the next section.
1.3 Early Results
As early as the 1920s, Woronzow (1924) showed that restoration of excitability in neurons depends on the duration of a pulse to negative potentials, indicating a slow state of the neuron, entered at depolarized potentials, that must be exited before subsequent excitation can take place. Several decades later, Na+ current was found to be responsible for neuronal excitation by Hodgkin and Huxley (1952a), linking it to the Na+ conducting system. As a result, a critical connection was made between the “availability” of the Na+ conductance, which is regulated by inactivation, and excitability. This link between Na+ channel inactivation and excitability was appreciated in some of the first recordings of squid giant axons by Hodgkin and Huxley (1952b) and cardiac Purkinje fibers by Sylvio Weidmann (1955). However, these studies focused on rapid events, such as fast inactivation, that take place over milliseconds. Slow recovery of a refractory neuron is due to another process, slow inactivation, which future results would demonstrate is clearly distinct from fast inactivation.
The detection of multiple inactivation time constants that revealed the existence of slow inactivation was not published until 1964 when Narahashi studied the time dependence of action potential restoration on membrane potential in lobster giant axons, noting different times needed for recovery of inactivation that depended on the magnitude of the depolarizing pulse (Narahashi 1964). Several years later, a study from Adelman and Palti and a series from Chandler and Meves identified time constants of Na+ current inactivation in squid giant axons ranging from hundreds of milliseconds to hundreds of seconds (Adelman and Palti 1969; Chandler and Meves 1970a, b, c, d). The presence of multiple time constants required to describe slow inactivation, spread over several orders of magnitude, indicated an extraordinarily complex process, not easily described by simple, discrete state transitions.
Subsequently much of the early characterization of slow inactivation was accomplished by Bernardo Rudy, who advocated an important physiological role. The first of this series investigated the resistance of slow inactivation to pronase (Rudy 1978), a protease which had recently been used to show that an intracellular peptide must regulate fast inactivation (Armstrong et al. 1973; Rojas and Rudy 1976). Removal of fast inactivation not only left slow inactivation intact but also apparently facilitated the process, Fig. 1a–c. Thus, the ability of slow inactivation to take place even in the absence of fast inactivation showed that it was a distinct gating process, resulting in a putative model allowed slow inactivation to take place from either the open- or fast-inactivated state (Fig. 1d). Later, Rudy and others showed that slow inactivation of Na+ channels, a process orders of magnitude slower than fast inactivation, is responsible for the refractoriness of Myxicola giant axons (Rudy 1975, 1981; Schauf et al. 1976) after long depolarizations, indicating a physiological role for slow gating in regulating neuronal activity. In skeletal muscle, slow inactivation was studied by Ruff et al., who proposed that slow Na+ channel inactivation mediates differences in the excitability of fast and slow twitch skeletal muscle fibers (Ruff et al. 1987, 1988).
Early recording of slow inactivation recovery. Measured by Bernardo Rudy in 1978 in Loligo squid giant axons maintained at 8°C, modified from Rudy (1978). (a) Axons were subjected to depolarization to −8 mV lasting several minutes. Subsequent 5 s pulses to −73 mV allowed the Na+ current to recover from slow inactivation, while short test pulses to +20 mV were used to quantify availability. The first pulse has the smallest magnitude. (b) Same protocol as in panel a, but after treatment with pronase to remove fast inactivation, note lack of a rapid decay in the currents. Slow inactivation of the Na+ conductance still took place, indicating that it is a distinct process. (c) Plot of recovery vs. time in seconds. Full recovery of the Na+ conductance required maintaining negative potentials required 10’s of seconds. Recovery with inactivation removed by pronase, open circles, was more rapid, having a time constant of 12.7 vs. 14.4 s (filled circles) for the untreated axon. (d) A putative model to account for the slow transitions. The ability of slow inactivation to take place in the absence and presence of fast inactivation, suggested a unique state (s) of the channel that could be entered when the channel was open (o) or fast inactivated (h)
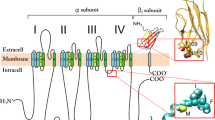
In the 1980s, the first Na+ channel was finally cloned (Tanabe et al. 1984) followed shortly thereafter by mammalian neuronal (Noda et al. 1986a), muscle (Trimmer et al. 1989), and cardiac isoforms (Gellens et al. 1992). The primary structure of the Na+ channel, which could be formed by monomers, was identified as having four domains (DI–DIV), each with six transmembrane segments (S1–S6). The fourth segment (S4) in each domain carried multiple positively charged residues and was identified as the voltage sensor. Subsequently, methods were developed to express channels heterologously (Noda et al. 1986b), isolating the Na+ channels from other channels in native cells and allowing systematic study of the structural basis of gating as well as differences between channel isoforms. The results from this new era are described in Sect. 3. However, it is important to first clarify, in the next section, how slow inactivation is quantified and what differences exist between the different channel isoforms.
2 Measurement and Characterization of Slow Inactivation
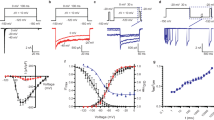
Several steps are required to accurately quantify slow inactivation in Na+ channel expressing cells. First, the time course of recovery from fast inactivation must be measured so that it can be isolated and separated from slow inactivation. Typically Na+ channels recover from fast inactivation after ~30 ms. So, if only fast inactivation is present, the current induced by a second depolarizing pulse after a 30 ms pulse to negative potentials will have the same magnitude as the current from the original pulse. However, if the initial depolarizing pulse is held for longer durations or repeated pulses are applied, the magnitude of the current during the following test pulses will be drastically reduced over time (Fig. 2a), indicating that channels have entered a slow-inactivated state. By applying pulses of different durations or by repeatedly measuring the ionic current after many pulses, the time course of the onset of slow inactivation can be assessed. The recovery kinetics can be measured in analogous fashion by varying the duration of a recovery pulse (Fig. 2b).
Measuring the onset of and recovery from slow inactivation. Figures modified from Vilin and Ruben (2001). (a) Quantifying onset of inactivation over time at different potentials shown on the left. As in the protocol below, after holding at −150 mV for 30 s, pulses of varying duration were applied to different potentials to induce slow inactivation. After a short pulse of 20 ms to −150 mV that allowed recovery from fast inactivation, a third test pulse was applied to evaluate how much current remained. The peak current from this pulse is plotted above at different voltages and durations for the induction pulse. (b) A protocol to measure recovery from slow inactivation is shown below. A pulse to 0 mV for 30 s was used to induce slow inactivation. The interpulse to different potentials with varying durations allowed slow inactivation to recover. Since at some potentials fast inactivation was still present, a short 20 ms pulse to −150 mV was used to remove it. The final test pulse revealed the amount that the slow inactivation had recovered. As in panel a, the peak current during this pulse for different voltages and interpulse durations is plotted in the above figure
The magnitude of slow inactivation is spread over several orders of magnitude. Typically, slow inactivation refers to a process that occurs over seconds to minutes. Channel inactivation that occurs in time domains between that of fast inactivation (several ms) and slow inactivation is called intermediate inactivation and has been linked to the Na+ channel C-terminus (Cormier et al. 2002; Tateyama et al. 2004; Mantegazza et al. 2001). Ultraslow inactivation occurs at scales longer than that of slow inactivation and requires on the order of 10’s of minutes (Szendroedi et al. 2007; Sandtner et al. 2004; Hilber et al. 2002). Still, given the potential for overlap between different types of inactivation, their temperature dependence, and several orders of magnitude covered by the slow inactivation time domain, assessing which event is being observed from one study to the next can be confounding. Thus, difficulty in isolating a single gating motion associated with slow inactivation is one reason why studies have proven to be quite challenging.
The slow inactivation phenotype varies depending on the channel isoform. In terms of inactivation onset, it abolishes 90 % of Na+ current after a 60 s pulse in the skeletal muscle isoform in comparison to 40 % in the cardiac isoform (Richmond et al. 1998). The time constant for inactivation onset is also ~6× faster, the voltage dependence steeper, and the recovery slower in the muscle isoform, compared to the cardiac version (O’Reilly et al. 1999). The neuronal isoform (NaV1.2) is similar to the muscle isoform, showing 90 % completeness after 100 s with time constants that resembles the muscle isoform (Toib et al. 1998).
3 Structural Basis of Slow Inactivation
Given the complexity of slow inactivation kinetics, it is unsurprising that many channel locales have been implicated in its generation. These include the channel pore, both intracellular and extracellular facing sides, the channel voltage-sensing domains, and the intracellular loops. Unraveling how all of these different channel motifs interact to cause slow inactivation is a difficult task and there is still much to be done to gain a better understanding of the protein motions that underlie slow Na+ channel gating. We begin with the role of the channel pore.
3.1 The Role of the Na+ Channel Pore in Slow Inactivation
Many lines of evidence point to involvement of the channel pore in mediating slow inactivation. Perhaps most directly, decreasing the concentration of Na+ ions available to flow through the pore impairs slow inactivation on the seconds scale (Townsend and Horn 1997). Additionally, tetrodotoxin (TTX), which blocks the channel pore has been recently shown to inhibit slow inactivation in the time domains of seconds (Capes et al. 2012) and minutes (Silva and Goldstein 2013a). The ability of an extracellular pore blocker to inhibit slow inactivation is consistent with previously published mutations near the outer pore that have also been shown to modulate slow inactivation. These mutations tend to reside near the Na+ selectivity filter such as DI mutations—W402C (Balser et al. 1996b) and W402A (Kambouris et al. 1998), DII—V787C (O’Reilly et al. 2001), DIII—F1236C (Ong et al. 2000), and DIV—V1583C (Vedantham and Cannon 2000) in the rat skeletal muscle isoform (Fig. 3a). Other residues that are involved with slow inactivation are found directly above the selectivity filter in a ring of four negative charges, one charge per domain, which is thought to draw positively charged Na+ ions near to the filter. In these experiments, when channels were slow inactivated, double cysteine mutants were not effectively modified by redox catalyst Cu(II) (1,10-phenanthroline)3, implying that slow inactivation renders the sites inaccessible (Xiong et al. 2003).
Mutations proximal to the channel pore modulate slow inactivation. (a) Mutations found to affect slow inactivation near the extracellular facing pore are in DI, W402C (Balser et al. 1996b) and W402A (Kambouris et al. 1998); DII, V787C (O’Reilly et al. 2001); DIII, F1236C (Ong et al. 2000); and DIV, V1583C (Vedantham and Cannon 2000) (Squares). Several mutations associated with HyperPP, linked to a destabilization of the slow-inactivated state, are localized to the inner pore. In DII, L689I (Bendahhou et al. 2002), I693T (Hayward et al. 1999), and T704M (Cummins and Sigworth 1996; Brancati et al. 2003); DIII, P1158S (Webb and Cannon 2008); and DIV, M1592V (Hayward et al. 1999) (Circles). (a) An example of a HyperPP mutation, rT689M (homologous to human T704M) that does not significantly affect channel fast kinetics after a single pulse (top), but has substantial consequences for slow inactivation (bottom), which was measured in response to prolonged depolarization. Figure modified from Cummins and Sigworth (1996)
Additional residues near the outer pore were found by comparing differences between the cardiac and muscle isoforms. The first of these studies swapped entire domains and identified DI and DII as key (O’Reilly et al. 1999). Next, chimeras were created in which the loops between the S5 and S6 segments in each channel domain in the muscle channel were replaced with the cardiac version and conferred the cardiac channel phenotype, consistent with the above results (Vilin et al. 1999). Further study identified a single residue in the DII S5–S6 P-loop that could confer the cardiac phenotype in the muscle channel when replaced with the cardiac residue and conversely impart the muscle phenotype in the cardiac isoform when the muscle residue was transplanted (Vilin et al. 2001b).
The inner side of the pore has also been implicated in slow inactivation with several mutations that cause HyperPP and inhibit slow inactivation colocalized to this location (Fig. 3a). Several of these mutations are found in DII, particular in the linker between the charged S4 and the pore-forming S5 segments. Biophysical characterization of these mutants shows insignificant to moderate consequences for the fast gating of the channel, while slow inactivation is measurably impaired (Bendahhou et al. 2002; Cummins and Sigworth 1996; Brancati et al. 2003; Webb and Cannon 2008; Hayward et al. 1999) (Fig. 3b).
3.2 A Link Between Fast Inactivation and Slow Inactivation
While the molecular basis of slow inactivation of sodium channels has remained difficult to pin down, fast inactivation is readily perturbed by intracellular modification of the channel. While early approaches employed pronase to cleave the intracellular sodium channel loops, the availability and identification of the IFM inactivation motif in the loop that connects DIII and DIV, the DIII–DIV linker, allowed researchers to directly target fast inactivation via site-directed mutagenesis without perturbing the rest of the channel (West et al. 1992). One of the first results showed that if IFM is changed to ICM, a construct that still inactivates the modification of the introduced cysteine by methane sulfanilides, which block fast inactivation, is relatively unaffected by slow inactivation (Vedantham and Cannon 1998).
As suggested by Bernardo Rudy after applying pronase to squid axons, slow inactivation is facilitated by the removal of fast inactivation in several Na+ channel isoforms. Thus, Featherstone et al. showed a greater rate of entry into the slow-inactivated state and a more complete inactivation in the skeletal muscle isoform (Featherstone et al. 1996) when the IFM mutation was mutated to QQQ (Fig. 4a). Similarly, Richmond et al. found enhanced slow inactivation in the cardiac isoform (Richmond et al. 1998) with fast inactivation removed (Fig. 4b). The result in the cardiac isoform is particularly stunning because of the lack of inactivation with fast inactivation present in comparison to the skeletal muscle isoform. More recently, the acidic residues that are adjacent to the IFM motif, which also modulate fast inactivation, were found to interact with slow inactivation (McCollum et al. 2003). Neutralization of acidic residues on either side of the IFM motif altered the voltage dependence of slow inactivation as well as its kinetics.
Removal of fast inactivation potentiates slow inactivation. 60 s pulses to varying potentials are applied, followed by a short pulse to −150 mV and a test pulse as in Fig. 2. Mutations of the IFM motif to QQQ ablate fast inactivation. (a) Removing fast inactivation in the skeletal muscle (Skm) isoform allows slow inactivation to occur with a steeper voltage dependence and to a higher degree of completeness. Figure modified from Featherstone et al. (1996). (b) The cardiac isoform, which is normally resistant to slow inactivation, shows nearly complete inactivation as well with fast inactivation. Figure modified from Richmond et al. (1998)
3.3 Voltage Sensing and Slow Inactivation
Voltage sensing in Na+ channels arises chiefly from outward displacement of the positively S4 voltage sensors in each domain by depolarization of the membrane potential. Given that slow inactivation is voltage dependent, it would not be surprising if were somehow linked to the voltage sensor motion. The first suggestion of this link came from early studies of the gating current by Bezanilla, Taylor, and Fernandez, which showed that, like the ionic current, it could inactivate. Precisely, after a long pulse to depolarized potentials, less gating current was observed after a test pulse that followed a short pulse to negative potentials (Bezanilla et al. 1982). A further correlation between the gating charge from a depolarizing pulse and slow inactivation was shown by Ruben et al. (1992). Surprisingly, the off-gating charge that resulted from a return to negative potential was not correlated. With the advent of routine site-directed mutagenesis, Kontis and Goldin (1997) neutralized charges on the S4 segments of each of the four Na+ channel domains to probe the dependence of steady-state inactivation on the voltage sensor. Each sensor contained at least one charge that significantly shifted the voltage dependence of steady-state inactivation.
The recent development of voltage clamp fluorometry has provided a means to assess the structural basis of Na+ channel gating (Cha et al. 1999; Chanda and Bezanilla 2002). In this method, a fluorophore is conjugated to cysteines that are placed in the extracellular S3–S4 linker in different constructs corresponding to each of the four domains (DI–DIV). Then, when depolarizing pulses are applied, fluorescence changes directly reflect the position of the sensor. Silva and Goldstein applied voltage clamp fluorometry to evaluate which sensors undergo slow immobilization and whether their kinetics correlate to slow inactivation (Silva and Goldstein 2013a) (Fig. 5a). Initial recordings showed that all four sensors are immobilized by long pulses at depolarized potentials (Fig. 5b). However, kinetics of fluorescence immobilization, which should correlate to the inactivation of the gating current, was significantly different. However, the gating current was recorded in the presence of TTX, which may have altered slow inactivation. Additional fluorescence recording with TTX showed that the immobilization of sensors in domains I and II was impaired by the toxin, as was slow inactivation. Recent work by Capes et al. (2012) also showed interaction between block by TTX and the DIV sensor. Thus, the extracellular pore, which was previously shown to interact with the slow-inactivated state, also communicates with the sensors in DI, DII, and DIV.
Potential role for the voltage sensors in slow inactivation. Previously developed constructs used to fluorescently track the position of the S4 segment in each Na+ channel domain (DI–DIV) were labeled and fluorescence changes were tracked simultaneously with changes in ionic current. Adapted from references Silva and Goldstein (2013a, b) and Cummins and Sigworth (1996). (a) Left: Applying repetitive pulses to induce slow inactivation of the ionic current, which can be probed by repetitions of a triple pulse protocol from a holding potential of −100 mV: a 5 ms test pulse to +45 mV is used to measure peak current (phase a); then slow inactivation is induced by a 500 ms pulse at −100, −75, −45, −15, 15, or 45 mV (phase b); and a 30 ms pulse at −100 mV employed (phase c) to allow for recovery from fast inactivation before the next test pulse. Sample traces are shown for cycle 1, 2, 80, and 320 and demonstrate progressive decrease in peak current at +45 mV. Center: The same protocol can also be used to assess slow kinetics of gating currents in the presence of TTX. Right: Tracking the voltage sensor immobilization from a construct with the DI-S4 labeled. (b) Voltage dependence of ionic current slow inactivation is shown at the left. On the right fluorescence showing that the S4 segments from all four domains undergo immobilization over a similar voltage range and time domain. (c) Introducing the HyperPP mutation L689I caused a loss in a slow component of inactivation that takes place over 10’s of seconds (left, arrow). Immobilization of the sensors in DI and DII was dramatically reduced, with the sensor in DII most impacted. The immobilization of the sensors in DIII and DIV was much less affected
The identification of disease-linked mutations that impairs slow inactivation suggested a means to perturb both the voltage sensor movements and slow inactivation to test whether there could be a correlation (Silva and Goldstein 2013b). In response, the L689I mutation, which resides on the S4–S5 linker in DII was introduced into each fluorescence construct to evaluate its effect on voltage sensor immobilization. The greatest effects of L689I were observed in DII immobilization, which was nearly abolished, with lesser consequences seen in the immobilization of the DIII and DIV sensors (Fig. 5c). Close examination of the L689I slow inactivation kinetics shows that inactivation that takes place in the 10 s time domain is most affected, which would imply a close link between DII sensor immobilization and this component of slow inactivation.
3.4 Modulation of Slow Inactivation by the β1 Subunit
In native cells, sodium channels are formed by the co-assembly of α and β subunits, which modify expression and channel gating (Isom et al. 1992). In muscle and cardiac Na+ channels, the β1 subunit is present, at least to some extent in native cells (Fozzard and Hanck 1996). Based on frog oocyte experiments, a role for the β1 subunit in regulating fast inactivation was assumed for muscle channels (Ji et al. 1994). However, more recent experiments in mammalian cells have shown that fast Na+ channel kinetics are not dramatically affected, while slow inactivation remains intact (Webb et al. 2009). Quantitatively, the muscle isoform expressed without the β1 subunit inactivates to 60 % completion, in comparison to when the β1 subunit is present it inactivates to 90 %. The cardiac isoform is unaffected and consistently shows 40 % inactivation (Vilin et al. 1999).
Conclusions
After many decades of study, myriad questions still remain regarding the molecular underpinnings of slow inactivation. First among these is in regard to the molecular underpinnings. As listed above, many single-point mutations near the outer channel pore have been found to impair slow inactivation, suggesting strong similarity to C-type inactivation in K+ channels, which has been linked to pore collapse (Choi et al. 1991; Hoshi et al. 1991; Kiss et al. 1999; Kiss and Korn 1998). Recently, crystal structures of the C-type-inactivated state have provided convincing evidence of its molecular mechanism (Cuello et al. 2010) and within the past 2 years several tetrameric bacterial Na+ channel structures have been published (Zhang et al. 2012; Payandeh et al. 2011, 2012; McCusker et al. 2012). As it becomes tractable to obtain crystallography data from four-domain monomeric Na+ channels, we can expect questions regarding the slow-inactivated state of the Na+ channel to be better resolved.
Even with many mutations identified in the extracellular pore that affect slow inactivation, colocalization of many HyperPP mutations to the inner pore, interaction with the intracellular fast inactivation gate and voltage sensors, suggests that C-type inactivation does fully encapsulate slow inactivation of Na+ channels. Some argue that the stretching of slow inactivation over many time domains as well as the ability of mutations over the span of the entire channel to perturb the process, make identification of discrete structurally based transitions futile effort. Instead they propose that slow inactivation should thought of as a gating event that is composed of infinitely many small transitions described not as the combination of many single exponentials, but by the more generalized stretch exponential in which discrete structural states of the channel are not defined (Marom 2010; Soudry et al. 2010).
Others may be tempted to speculate as to the structural basis of the different components of slow inactivation based on the time domains over which many interactions with the process occur. Over the fastest domains described by slow inactivation (~1 s), DIV has arisen several times, first in the experiments of Vedantham and Cannon who observed that V1583C modification by methane sulfonimides takes place over milliseconds to seconds (Vedantham and Cannon 2000), then by Mitrovic et al. (2000), who examined slow inactivation over the 5 s time domain, and finally by Silva and Goldstein who showed that DIV immobilization is fastest of all the domains (Silva and Goldstein 2013a) (Fig. 5b).
Over the 10 s time domain, two HyperPP mutations that are localized to DII, L689I (Silva and Goldstein 2013b) (Fig. 5c), and T704M (Melamed-Frank and Marom 1999) remove slow inactivation. Furthermore, skeletal muscle inactivation that takes place over this time domain is impaired by replacing a residue in the DII P-loop with the cardiac residue (Vilin et al. 2001b) or by swapping the entire DI or DII domains (O’Reilly et al. 1999). Finally, limited data observing TTX interaction with slow inactivation shows an inhibition of the 10 s time domain and inhibition of immobilization of DI and DII S4’s (Silva and Goldstein 2013a), which is consistent with experiments that localized its binding site to DI and DII (Fozzard and Lipkind 2010).
The longest slow inactivation time domain was not affected by L689I or T704M and persists in the cardiac Na+ channel. However, this process is enhanced by the removal of fast inactivation, which has been shown to interact with the fast inactivation gate in the DIII S4–S5 linker in both the cardiac and skeletal muscle isoforms (Goldin 2003; Smith and Goldin 1997). Furthermore, sensor immobilization in DIII, observed via fluorescence, contained a significant slow component (Fig. 5b), suggesting a contribution to this slowest component.
In sum, a compelling working hypothesis arises for those that insist on assigning a structural basis for the different components of slow inactivation. In this hypothesis, DIV accounts for the fastest component (1–5 s), DI and DII are responsible for the intermediate component 10–30 s, and DIII and fast inactivation are linked to the slowest component in the time domain of minutes.
References
Adelman WJ Jr, Palti Y (1969) The effects of external potassium and long duration voltage conditioning on the amplitude of sodium currents in the giant axon of the squid, Loligo pealei. J Gen Physiol 54(5):589–606
Armstrong CM, Bezanilla F, Rojas E (1973) Destruction of sodium conductance inactivation in squid axons perfused with pronase. J Gen Physiol 62(4):375–391
Balser JR et al (1996a) Local anesthetics as effectors of allosteric gating. Lidocaine effects on inactivation-deficient rat skeletal muscle Na channels. J Clin Invest 98(12):2874–2886
Balser JR et al (1996b) External pore residue mediates slow inactivation in mu 1 rat skeletal muscle sodium channels. J Physiol 494(Pt 2):431–442
Bendahhou S et al (2002) Impairment of slow inactivation as a common mechanism for periodic paralysis in DIIS4-S5. Neurology 58(8):1266–1272
Bezanilla F, Taylor RE, Fernandez JM (1982) Distribution and kinetics of membrane dielectric polarization. 1. Long-term inactivation of gating currents. J Gen Physiol 79(1):21–40
Brancati F et al (2003) Severe infantile hyperkalaemic periodic paralysis and paramyotonia congenita: broadening the clinical spectrum associated with the T704M mutation in SCN4A. J Neurol Neurosurg Psychiatry 74(9):1339–1341
Capes DL et al (2012) Gating transitions in the selectivity filter region of a sodium channel are coupled to the domain IV voltage sensor. Proc Natl Acad Sci U S A 109(7):2648–2653
Cha A et al (1999) Voltage sensors in domains III and IV, but not I and II, are immobilized by Na + channel fast inactivation. Neuron 22(1):73–87
Chanda B, Bezanilla F (2002) Tracking voltage-dependent conformational changes in skeletal muscle sodium channel during activation. J Gen Physiol 120(5):629–645
Chandler WK, Meves H (1970a) Slow changes in membrane permeability and long-lasting action potentials in axons perfused with fluoride solutions. J Physiol 211(3):707–728
Chandler WK, Meves H (1970b) Rate constants associated with changes in sodium conductance in axons perfused with sodium fluoride. J Physiol 211(3):679–705
Chandler WK, Meves H (1970c) Evidence for two types of sodium conductance in axons perfused with sodium fluoride solution. J Physiol 211(3):653–678
Chandler WK, Meves H (1970d) Sodium and potassium currents in squid axons perfused with fluoride solutions. J Physiol 211(3):623–652
Choi KL, Aldrich RW, Yellen G (1991) Tetraethylammonium blockade distinguishes two inactivation mechanisms in voltage-activated K + channels. Proc Natl Acad Sci U S A 88(12):5092–5095
Cormier JW et al (2002) Secondary structure of the human cardiac Na + channel C terminus: evidence for a role of helical structures in modulation of channel inactivation. J Biol Chem 277(11):9233–9241
Cuello LG et al (2010) Structural mechanism of C-type inactivation in K(+) channels. Nature 466(7303):203–208
Cummins TR, Sigworth FJ (1996) Impaired slow inactivation in mutant sodium channels. Biophys J 71(1):227–236
Featherstone DE, Richmond JE, Ruben PC (1996) Interaction between fast and slow inactivation in Skm1 sodium channels. Biophys J 71(6):3098–3109
Fozzard HA, Hanck DA (1996) Structure and function of voltage-dependent sodium channels: comparison of brain II and cardiac isoforms. Physiol Rev 76(3):887–926
Fozzard HA, Lipkind GM (2010) The tetrodotoxin binding site is within the outer vestibule of the sodium channel. Mar Drugs 8(2):219–234
Gellens ME et al (1992) Primary structure and functional expression of the human cardiac tetrodotoxin-insensitive voltage-dependent sodium channel. Proc Natl Acad Sci U S A 89(2):554–558
George AL Jr (2005) Inherited disorders of voltage-gated sodium channels. J Clin Invest 115(8):1990–1999
Goldin AL (2003) Mechanisms of sodium channel inactivation. Curr Opin Neurobiol 13(3):284–290
Hayward LJ, Sandoval GM, Cannon SC (1999) Defective slow inactivation of sodium channels contributes to familial periodic paralysis. Neurology 52(7):1447–1453
Hilber K et al (2002) Interaction between fast and ultra-slow inactivation in the voltage-gated sodium channel. Does the inactivation gate stabilize the channel structure? J Biol Chem 277(40):37105–37115
Hodgkin AL, Huxley AF (1952a) A quantitative description of membrane current and its application to conduction and excitation in nerve. J Physiol 117(4):500–544
Hodgkin AL, Huxley AF (1952b) The dual effect of membrane potential on sodium conductance in the giant axon of Loligo. J Physiol 116(4):497–506
Hoshi T, Zagotta WN, Aldrich RW (1991) Two types of inactivation in Shaker K + channels: effects of alterations in the carboxy-terminal region. Neuron 7(4):547–556
Isom LL et al (1992) Primary structure and functional expression of the beta 1 subunit of the rat brain sodium channel. Science 256(5058):839–842
Ji S et al (1994) Voltage-dependent regulation of modal gating in the rat SkM1 sodium channel expressed in Xenopus oocytes. J Gen Physiol 104(4):625–643
Jurkat-Rott K, Lehmann-Horn F (2006) Paroxysmal muscle weakness: the familial periodic paralyses. J Neurol 253(11):1391–1398
Jurkat-Rott K et al (2009) K + -dependent paradoxical membrane depolarization and Na + overload, major and reversible contributors to weakness by ion channel leaks. Proc Natl Acad Sci U S A 106(10):4036–4041
Kambouris NG et al (1998) Mechanistic link between lidocaine block and inactivation probed by outer pore mutations in the rat micro1 skeletal muscle sodium channel. J Physiol 512(Pt 3):693–705
Kiss L, Korn SJ (1998) Modulation of C-type inactivation by K + at the potassium channel selectivity filter. Biophys J 74(4):1840–1849
Kiss L, LoTurco J, Korn SJ (1999) Contribution of the selectivity filter to inactivation in potassium channels. Biophys J 76(1 Pt 1):253–263
Kontis KJ, Goldin AL (1997) Sodium channel inactivation is altered by substitution of voltage sensor positive charges. J Gen Physiol 110(4):403–413
Lengele JP, Belge H, Devuyst O (2008) Periodic paralyses: when channels go wrong. Nephrol Dial Transplant 23(4):1098–1101
Mantegazza M et al (2001) Role of the C-terminal domain in inactivation of brain and cardiac sodium channels. Proc Natl Acad Sci U S A 98(26):15348–15353
Marom S (2010) Neural timescales or lack thereof. Prog Neurobiol 90(1):16–28
McCollum IJ et al (2003) Negatively charged residues adjacent to IFM motif in the DIII-DIV linker of hNa(V)1.4 differentially affect slow inactivation. FEBS Lett 552(2–3):163–169
McCusker EC et al (2012) Structure of a bacterial voltage-gated sodium channel pore reveals mechanisms of opening and closing. Nat Commun 3:1102
Melamed-Frank M, Marom S (1999) A global defect in scaling relationship between electrical activity and availability of muscle sodium channels in hyperkalemic periodic paralysis. Pflugers Arch 438(2):213–217
Mitrovic N, George AL Jr, Horn R (2000) Role of domain 4 in sodium channel slow inactivation. J Gen Physiol 115(6):707–718
Narahashi T (1964) Restoration of action potential by anodal polarization in lobster giant axons. J Cell Physiol 64:73–96
Noda M et al (1986a) Existence of distinct sodium channel messenger RNAs in rat brain. Nature 320(6058):188–192
Noda M et al (1986b) Expression of functional sodium channels from cloned cDNA. Nature 322(6082):826–828
Ong BH, Tomaselli GF, Balser JR (2000) A structural rearrangement in the sodium channel pore linked to slow inactivation and use dependence. J Gen Physiol 116(5):653–662
O’Reilly JP et al (1999) Comparison of slow inactivation in human heart and rat skeletal muscle Na + channel chimaeras. J Physiol 515(Pt 1):61–73
O’Reilly JP, Wang SY, Wang GK (2001) Residue-specific effects on slow inactivation at V787 in D2-S6 of Na(v)1.4 sodium channels. Biophys J 81(4):2100–2111
Payandeh J et al (2011) The crystal structure of a voltage-gated sodium channel. Nature 475(7356):353–358
Payandeh J et al (2012) Crystal structure of a voltage-gated sodium channel in two potentially inactivated states. Nature 486(7401):135–139
Richmond JE et al (1998) Slow inactivation in human cardiac sodium channels. Biophys J 74(6):2945–2952
Rojas E, Rudy B (1976) Destruction of the sodium conductance inactivation by a specific protease in perfused nerve fibres from Loligo. J Physiol 262(2):501–531
Ruben PC, Starkus JG, Rayner MD (1992) Steady-state availability of sodium channels. Interactions between activation and slow inactivation. Biophys J 61(4):941–955
Rudy B (1975) Proceedings: slow recovery of the inactivation of sodium conductance in myxicola giant axons. J Physiol 249(1):22P–24P
Rudy B (1978) Slow inactivation of the sodium conductance in squid giant axons. Pronase resistance J Physiol 283:1–21
Rudy B (1981) Inactivation in myxicola giant axons responsible for slow and accumulative adaptation phenomena. J Physiol 312:531–549
Ruff RL, Simoncini L, Stuhmer W (1987) Comparison between slow sodium channel inactivation in rat slow- and fast-twitch muscle. J Physiol 383:339–348
Ruff RL, Simoncini L, Stuhmer W (1988) Slow sodium channel inactivation in mammalian muscle: a possible role in regulating excitability. Muscle Nerve 11(5):502–510
Sandtner W et al (2004) Lidocaine: a foot in the door of the inner vestibule prevents ultra-slow inactivation of a voltage-gated sodium channel. Mol Pharmacol 66(3):648–657
Schauf CL (1987) Zonisamide enhances slow sodium inactivation in Myxicola. Brain Res 413(1):185–188
Schauf CL, Pencek TL, Davis FA (1976) Slow sodium inactivation in Myxicola axons. Evidence for a second inactive state. Biophys J 16(7):771–778
Shaw RM, Rudy Y (1997) Ionic mechanisms of propagation in cardiac tissue. Roles of the sodium and L-type calcium currents during reduced excitability and decreased gap junction coupling. Circ Res 81(5):727–741
Sheets MF et al (2010) Sodium channel molecular conformations and antiarrhythmic drug affinity. Trends Cardiovasc Med 20(1):16–21
Silva JR, Goldstein SA (2013a) Voltage-sensor movements describe slow inactivation of voltage-gated sodium channels I: wild-type skeletal muscle Na(V)1.4. J Gen Physiol 141(3):309–321
Silva JR, Goldstein SA (2013b) Voltage-sensor movements describe slow inactivation of voltage-gated sodium channels II: a periodic paralysis mutation in Na(V)1.4 (L689I). J Gen Physiol 141(3):323–334
Smith MR, Goldin AL (1997) Interaction between the sodium channel inactivation linker and domain III S4-S5. Biophys J 73(4):1885–1895
Song W et al (2011) Analysis of the action of lidocaine on insect sodium channels. Insect Biochem Mol Biol 41(1):36–41
Soudry D, Meir R (2010) History-dependent dynamics in a generic model of ion channels—an analytic study. Front Comput Neurosci 4. pii: 3. doi:10.3389/fncom.2010.00003. http://www.ncbi.nlm.nih.gov/pubmed/20725633
Szendroedi J et al (2007) Speeding the recovery from ultraslow inactivation of voltage-gated Na + channels by metal ion binding to the selectivity filter: a foot-on-the-door? Biophys J 93(12):4209–4224
Tanabe T et al (1984) Primary structure of beta subunit precursor of calf muscle acetylcholine receptor deduced from cDNA sequence. Eur J Biochem 144(1):11–17
Tateyama M et al (2004) Structural effects of an LQT-3 mutation on heart Na + channel gating. Biophys J 86(3):1843–1851
Toib A, Lyakhov V, Marom S (1998) Interaction between duration of activity and time course of recovery from slow inactivation in mammalian brain Na + channels. J Neurosci 18(5):1893–1903
Townsend C, Horn R (1997) Effect of alkali metal cations on slow inactivation of cardiac Na + channels. J Gen Physiol 110(1):23–33
Trimmer JS et al (1989) Primary structure and functional expression of a mammalian skeletal muscle sodium channel. Neuron 3(1):33–49
Vedantham V, Cannon SC (1998) Slow inactivation does not affect movement of the fast inactivation gate in voltage-gated Na + channels. J Gen Physiol 111(1):83–93
Vedantham V, Cannon SC (2000) Rapid and slow voltage-dependent conformational changes in segment IVS6 of voltage-gated Na(+) channels. Biophys J 78(6):2943–2958
Vilin YY, Ruben PC (2001) Slow inactivation in voltage-gated sodium channels: molecular substrates and contributions to channelopathies. Cell Biochem Biophys 35(2):171–190
Vilin YY et al (1999) Structural determinants of slow inactivation in human cardiac and skeletal muscle sodium channels. Biophys J 77(3):1384–1393
Vilin YY, Fujimoto E, Ruben PC (2001a) A novel mechanism associated with idiopathic ventricular fibrillation (IVF) mutations R1232W and T1620M in human cardiac sodium channels. Pflugers Arch 442(2):204–211
Vilin YY, Fujimoto E, Ruben PC (2001b) A single residue differentiates between human cardiac and skeletal muscle Na + channel slow inactivation. Biophys J 80(5):2221–2230
Webb J, Cannon SC (2008) Cold-induced defects of sodium channel gating in atypical periodic paralysis plus myotonia. Neurology 70(10):755–761
Webb J, Wu FF, Cannon SC (2009) Slow inactivation of the NaV1.4 sodium channel in mammalian cells is impeded by co-expression of the beta1 subunit. Pflugers Arch 457(6):1253–1263
Weidmann S (1955) The effect of the cardiac membrane potential on the rapid availability of the sodium-carrying system. J Physiol 127(1):213–224
West JW et al (1992) A cluster of hydrophobic amino acid residues required for fast Na(+)-channel inactivation. Proc Natl Acad Sci U S A 89(22):10910–10914
Woronzow DS (1924) Über die Einwirkung des konstanten Stromes auf den mit Wasser, Zuckerlösung, Alkali- und Erdalkalichloridlösungen behandelten Nerven. Pflüger’s Archiv für die gesamte Physiologie des Menschen und der Tiere 203(1):300–318
Xiong W et al (2003) Molecular motions of the outer ring of charge of the sodium channel: do they couple to slow inactivation? J Gen Physiol 122(3):323–332
Zhang X et al (2012) Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature 486(7401):130–134
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Silva, J. (2014). Slow Inactivation of Na+ Channels. In: Ruben, P. (eds) Voltage Gated Sodium Channels. Handbook of Experimental Pharmacology, vol 221. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-41588-3_3
Download citation
DOI: https://doi.org/10.1007/978-3-642-41588-3_3
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-41587-6
Online ISBN: 978-3-642-41588-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)