Abstract
Programmed death-ligand 1 (PD-L1) has been speculated to play a critical role in suppression of the immune system and it can be upregulated in cancer cells, which may allow cancers to evade the host immune system. MicroRNAs (miRNAs) are small non-coding RNA molecules (containing about 22 nucleotides), that function in RNA silencing and post-transcriptional regulation of gene expression. MiRNAs were found deregulated (upregulated or downregulated) and implicated in cancer development with various roles which depend on their gene target. Using targetscan web server prediction algorithm, we concluded that miR-140-3p is a targeting mirRNA with conserved consequential pairing of target region for PD-L1. Moreover, by reviewing all the available cancer studies in Pub/Medline about miR-140-3p, was found permanently down regulated. Furthermore, in recent immunotherapy related clinical trials in most cancers, evaluated PD-L1, it is found overexpressed. In the near future, in vitro or in vivo studies need to validate whether there is direct correlation between PD-L1 overexpression and miR-140-3p downregulation as targetscan performed algorithm predicted.
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
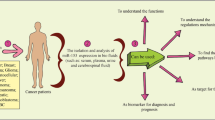
PD-1 (Programmed cell death protein 1) is a key immune-checkpoint receptor expressed by activated T cells, which mediates immunosuppression. PD-1 functions primarily in peripheral tissues, where T cells may encounter the immunosuppressive PD-1 ligands, namely PD-L1 (B7-H1) and PD-L2 (B7-DC), which are expressed by tumor cells, stromal cells, or both [1]. It appears that upregulation of PD-L1 may allow cancers to evade the host’s immune system through the transmission of an immunosuppressing signal from tumor cells. Many recent studies show that PD-L1 expression in tumor is associated with impaired survival [2]. An analysis of 196 tumor specimens from patients with renal cell carcinoma found that high tumor expression of PD-L1 was associated with increased tumor aggressiveness and a 4.5-fold increased risk of death [3]. Therefore, many PD-L1 inhibitors are in development as immuno-oncology therapies, showing good results in clinical trials [4] and consist a new treatment approach against cancer treatment (Fig. 1).
Human cancer immunotherapy with antibodies to the PD-1 and PD-L1 pathway [5]
An effective strategy of preventing immunosuppression by antibodies blocking the PD-1, or its ligand (PD-L1), was shown in phase I trials by inducing a 30–50% response in several cancer types [6]. More recent phase II and III studies took accelerated approval of anti-PD-1 antibodies for metastatic melanoma (MM) [7, 8], non-small cell lung cancer (NSCLC) [9] and renal cell cancer (RCC) [10]. In another very recent phase II study atezolizumab, an anti-PD-L1 antibody was approved for metastatic bladder cancer treatment [11]. The importance of PD-L1 expression in prognosis and in prediction of treatment outcome, promote identifying other molecules modulators or regulators of it’s expression. MicroRNAs (miRNAs) are a class of small, non-coding, endogenous single RNA molecules that play important roles in gene expression through binding to the 3 UTR of the targeted gene mRNA, leading to mRNA cleavage or translational repression [12]. miRNAs are transcribed by RNA polymerase II as part of capped and polyadenylated primary transcripts (pri-miRNAs) that can be either protein-coding or non-coding. The primary transcript is cleaved by the Drosha ribonuclease III enzyme to produce an approximately 70-nt stem-loop precursor miRNA (pre-miRNA), which is further cleaved by the cytoplasmic Dicer ribonuclease to generate the mature miRNA and antisense miRNA star (miRNA*) products. The mature miRNA is incorporated into a RNA-induced silencing complex (RISC), which recognizes targeted mRNAs through imperfect base pairing with the miRNA and most commonly results in translational inhibition or destabilization of the target mRNA (Fig. 2).
Overview of miRNA biogenesis pathway [13]
Numerous studies show that miRNAs participate in various biological processes, such as cell differentiation, cell growth, apoptosis, death and timing development [14]. In the present study we aim to investigate if PD-L1 is a predicted target by microRnas, using the targetscan web server prediction algorithm. The results will be analyzed by reviewing the available publication studies of the target miRnas in various tumors and PD-L1 expression as well.
2 Method
In bioinformatics, TargetScan is a web server that predicts biological targets of microRNAs by searching for the presence of sites that match the seed region of each miRNA. Performing targetscan, results showed PD-L1 as predictive target for miR-140-3p (Fig. 3).
The prediction targetscan method indicates that miR 140-3p.2 predicted target PD-L1(CD274) with conserved consequential pairing of target region and miRna, in position 43–49, near the stop codon end of the 3-UTR, a region with high efficacy for repression [15] (Fig. 4).
The prediction targetscan method, indicate that miR 140-3p.2 is the only miRna that predicted target PD-L1(CD274) with conserved consequential pairing of target region and miRna, in this position. Searching in Pubmed/Medline database, all available publications about: ‘miR-140-3p and cancer’, in almost all studies’ results that referred to cells or tissues samples analysed, miR140-3p was found down regulated and higher levels of miR 140-3p expression, favored tumor’s deprivation. In a study analyzing miR-140-3p in breast cancer cell lines, were found evidence that suggest there is functional synergy between the canonical hsa-miR-140-3p and the newly identified 5’isomiR-140-3p in suppressing growth and progression of breast cancer by simultaneously targeting genes related to differentiation, proliferation and migration. In patients with higher levels there was a trend for better survival [16]. In others studies, miR-140-3p expression level was lower in lung cancer tissues compared to adjacent normal lung cancer tissue and after miR-140-3p was upregulated in A549 or H1299 cells, cell proliferation, invasion and migration was notably attenuated [17, 18]. Moreover, analyzing miRNA expression signatures in angioimmunoblastic T-cell lymphomas (AITLs) of 760 microRNAs in 30 nodal AITL, miR-140-3p, was found to be downregulated to a significant level [19]. The same results found in multiple myeloma cells study, demonstrating miR-140-3p being repressed [20].
In contrast, plasma levels of miR-140-3p were significantly higher in the papillary thyroid carcinoma (PTC) patients than in those with benign nodules or the healthy controls [21]. This is not supported by others studies referred to circulated miRnas in tumors. In a study assessing circulating microRna in blood in five different tumors, miR-140-3p was a potential biomarker, among those selected to distinguish patients with cancer or other diseases from healthy controls. Significantly miR-140-3p was downregulated in all five tumors samples (lung, ovarian, colon, nervous system, hematologic) [22]. Furthermore, from the rest of available studies evaluating miR-140-3p in cancer, published in pubmed database, the conclusion is the same. There was a clear miR-140-3p underexpression in all different tumor samples detected in these studies (cutaneous squamous cell carcinoma, basal cell carcinoma, colon adenocarcinoma, oral squamous cell carcinoma, ovarian carcinoma, lung squamous cell carcinoma, multiple myeloma) [23–29].
Moreover, the expression of PD-l1 in various tumor types has been widely explored in context to predict the most effective immunotherapy for the disease. In studies evaluating PD-L1 expression, as a predictive biomarker in immunotherapy, solid tumors with most increased expression were in concordance with tumor types in which miR-140-3p was found downregulated before [30] (Fig. 5).
Table 1
3 Discussion
The association of miR-140-3p with his target gene PD-L1, as was predicted by targetscan prediction algorithm, is more complex and unclear than it seems. In a study at chordomas, miR-140-3p was found upregulated and correlated with recurrence, tumor invasion and worse recurrence-free survival [31]. However, at a recent study analyzing PD-L1 overexpression in tumor infiltrating lymphocytes, in spinal. In spinal chordoma samples, there was no association between PD-L1 with local recurrence-free survival (LRFS) or overall survival (OS), but instead with favorable prognosis [32]. Certainly, its apoptotic role is more elucidated in preclinical or clinical studies that used miR-140-3p, in fields other than in oncology. Preclinically, accessing miR-140-3p with others apoptotic miRnas in myocardial infarction post transplantation with skeletal myoblast in rats, there was apparently increased expression of this miRna, starting apoptosis [33]. Clinically, targeting synovial fibroblast by intraarticular delivery of miRNAs 140-3p and 140-5p can ameliorate autoimmune arthritis [34]. These findings indicate that miR-140-3p has an apparent role in apoptosis, which might implicate additionally PD-L1 regulation. All the above studies suggest that miR-140-3p functions through apoptosis network but the only way to assure that it is inversely correlated or even downregulate PD-L1 expression, are in vivo studies. In a previous study exploring miRNA profiling of normal mammary stem cells and cancer stem-like cells (CSC) from DCIS (ductal carcinoma in situ) tumors, revealed that miR-140 is significantly downregulated in cancer stem-like cells compared with normal stem cells [35]. Moreover, in ER-negative/basal-like DCIS, a cancer stem cell fenotype, restoration of miR-140-3p reduced tumor growth in vivo. Considering that PD-L1 in basal like breast cancer cells is significantly upregulated compare to others breast cancer cells [36] there is an obvious indirect correlation with miR-140-3p downregulation.
4 Conclusions
MiR140-3p, evaluated in many tissue cancers, has been found downregulated. Moreover, his target gene PD-L1, with high clinical importance in most cancer, is overexpressed. In view of the fact that, miR-140-3p is corroborated as apoptotic related miRna, and its ability in pairing conserved with his target PD-L1 in a unique way (near the stop codon end of the 3-UTR) as targetscan performed, makes warrant the confirmation with further in vivo or in vitro experiments as a direct PD-L1 regulator.
References
Pardoll, D.M. 2012. The blockade of immune checkpoints in cancer immunotherapy. Nature Reviews Cancer 12(4): 252–264.
Pyo, J., G. Kang, and J. Kim. 2016. Prognostic role of pd-l1 in malignant solid tumors: A meta-analysis. The International Journal of Biological Markers 2:32(1): e68–e74.
Thompson, R.H., M.D. Gillett, J.C. Cheville, C.M. Lohse, H. Dong, W.S. Webster, K.G. Krejci, J.R. Lobo, S. Sengupta, L. Chen et al. 2004. Costimulatory b7-h1 in renal cell carcinoma patients: Indicator of tumor aggressiveness and potential therapeutic target. Proceedings of the National Academy of Sciences of the United States of America 101(49): 17174–17179.
Velcheti, V., K.A. Schalper, D.E. Carvajal, V.K. Anagnostou, K.N. Syrigos, M. Sznol, R.S. Herbst, S.N. Gettinger, L. Chen, and D.L. Rimm. 2014. Programmed death ligand-1 expression in non-small cell lung cancer. Laboratory Investigation 94(1): 107–116.
Ohaegbulam, K.C., A. Assal, E. Lazar-Molnar, Y. Yao, and X. Zang. 2015. Human cancer immunotherapy with antibodies to the pd-1 and pd-l1 pathway. Trends in Molecular Medicine 21(1): 24–33.
Topalian, S.L., F.S. Hodi, J.R. Brahmer, S.N. Gettinger, D.C. Smith, D.F. McDermott, J.D. Powderly, R.D. Carvajal, J.A. Sosman, M.B. Atkins et al. 2012. Safety, activity, and immune correlates of anti–pd-1 antibody in cancer. New England Journal of Medicine 366(26), 2443–2454.
Robert, C., G.V. Long, B. Brady, C. Dutriaux, M. Maio, L. Mortier, J.C. Hassel, P. Rutkowski, C. McNeil, E. Kalinka-Warzocha et al. 2015. Nivolumab in previously untreated melanoma without braf mutation. New England Journal of Medicine 372(4): 320–330.
Robert, C., J. Schachter, G.V. Long, A. Arance, J.J. Grob, L. Mortier, A. Daud, M.S. Carlino, C. McNeil, M. Lotem et al. 2015. Pembrolizumab versus ipilimumab in advanced melanoma. New England Journal of Medicine 372(26): 2521–2532.
Garon, E.B., N.A. Rizvi, R. Hui, N. Leighl, A.S. Balmanoukian, J.P. Eder, A. Patnaik, C. Aggarwal, M. Gubens, L. Horn et al. 2015. Pembrolizumab for the treatment of non–small-cell lung cancer. New England Journal of Medicine 372(21): 2018–2028.
Motzer, R.J., B. Escudier, D.F. McDermott, S. George, H.J. Hammers, S. Srinivas, S.S. Tykodi, J.A. Sosman, G. Procopio, E.R. Plimack et al.: Nivolumab versus everolimus in advanced renal-cell carcinoma. New England Journal of Medicine 373(19): 1803–1813.
Hoffman-Censits, J.H., P. Grivas, M.S. Van Der Heijden, R. Dreicer, Y. Loriot, M. Retz, N.J. Vogelzang, J.L. Perez-Gracia, A. Rezazadeh, S. Bracarda et al. 2016. Imvigor 210, a phase ii trial of atezolizumab (mpdl3280a) in platinum-treated locally advanced or metastatic urothelial carcinoma (muc). In ASCO Annual Meeting Proceedings, vol. 34, 355.
Bartel, D.P. 2004. Micrornas: Genomics, biogenesis, mechanism, and function. Cell 116(2): 281–297.
Lin, S., and R.I. Gregory. 2015. Microrna biogenesis pathways in cancer. Nature Reviews Cancer 15(6): 321–333.
Ambros, V. 2004. The functions of animal micrornas. Nature 431(7006): 350–355.
Grimson, A., K.K.H. Farh, W.K. Johnston, P. Garrett-Engele, L.P., Lim, and D.P. Bartel. 2007. Microrna targeting specificity in mammals: Determinants beyond seed pairing. Molecular Cell 27(1): 91–105.
Salem, O., N. Erdem, J. Jung, E. Münstermann, A. Wörner, H. Wilhelm, S. Wiemann, and C. Körner. 2016. The highly expressed 5-isomir of hsa-mir-140-3p contributes to the tumor-suppressive effects of mir-140 by reducing breast cancer proliferation and migration. BMC Genomics 17(1): 566.
Kong, X.M., G.H. Zhang, Y.K. Huo, X.H. Zhao, D.W. Cao, S.F. Guo, A.M. Li, and X.R. Zhang. 2015. Microrna-140-3p inhibits proliferation, migration and invasion of lung cancer cells by targeting atp6ap2. International Journal of Clinical and Experimental Pathology 8(10): 12845.
Dong, W., C. Yao, X. Teng, J. Chai, X. Yang, and B. Li. 2016. Mir-140-3p suppressed cell growth and invasion by downregulating the expression of atp8a1 in non-small cell lung cancer. Tumor Biology 37(3): 2973–2985.
Reddemann, K., D. Gola, A. Schillert, J. Knief, C. Kuempers, J. Ribbat-Idel, S. Ber, J. Schemme, V. Bernard, N. Gebauer et al. 2015. Dysregulation of micrornas in angioimmunoblastic t-cell lymphoma. Anticancer Research 35(4): 2055–2061.
Yuan, L., G.C.F. Chan, K.L. Fung, and C.S. Chim, Rankl expression in myeloma cells is regulated by a network involving rankl promoter methylation, dnmt1, microrna and tnfα in the microenvironment. Biochimica et Biophysica Acta (BBA)-Molecular Cell Research 1843(9): 1834–1838.
Li, M., Q. Song, H. Li, Y. Lou, and L. Wang. 2015. Circulating mir-25-3p and mir-451a may be potential biomarkers for the diagnosis of papillary thyroid carcinoma. PloS One 10(7): e0132403.
Taguchi, Y., and Y. Murakami. 2013. Principal component analysis based feature extraction approach to identify circulating microrna biomarkers. PloS One 8(6): e66714.
Sand, M., M. Skrygan, D. Georgas, D. Sand, S.A., Hahn, T. Gambichler, P. Altmeyer, and F.G. Bechara. 2012. Microarray analysis of microrna expression in cutaneous squamous cell carcinoma. Journal of Dermatological Science 68(3): 119–126.
Sand, M., M. Skrygan, D. Sand, D. Georgas, S. Hahn, T. Gambichler, P. Altmeyer, and F. Bechara. 2012. Expression of micrornas in basal cell carcinoma. British Journal of Dermatology 167(4): 847–855.
Piepoli, A., F. Tavano, M. Copetti, T. Mazza, O. Palumbo, A. Panza, F.F. Di Mola, V. Pazienza, G. Mazzoccoli, G. Biscaglia et al. 2012. Mirna expression profiles identify drivers in colorectal and pancreatic cancers. PloS One 7(3): e33663.
Serrano, N.A., C. Xu, Y. Liu, P. Wang, W. Fan, M.P. Upton, J.R. Houck, P. Lohavanichbutr, M. Kao, L.P. Zhao et al. 2012. Integrative analysis in oral squamous cell carcinoma reveals dna copy number–associated mirnas dysregulating target genes. Otolaryngology–Head and Neck Surgery 0194599812442490.
Miles, G.D., M. Seiler, L. Rodriguez, G. Rajagopal, and B. Bhanot. 2012. Identifying microrna/mrna dysregulations in ovarian cancer. BMC Research Notes 5(1): 1.
Tan, X., W. Qin, L. Zhang, J. Hang, B. Li, C. Zhang, J. Wan, F. Zhou, K. Shao, Y. Sun et al. 2011. A 5-microrna signature for lung squamous cell carcinoma diagnosis and hsa-mir-31 for prognosis. Clinical Cancer Research 17(21): 6802–6811.
Lionetti, M., M. Biasiolo, L. Agnelli, K. Todoerti, L. Mosca, S. Fabris, G. Sales, G.L., Deliliers, S., Bicciato, L. Lombardi et al. 2009. Identification of microrna expression patterns and definition of a microrna/mrna regulatory network in distinct molecular groups of multiple myeloma. Blood 114(25): e20–e26.
Patel, S.P., and R. Kurzrock. 2015. Pd-l1 expression as a predictive biomarker in cancer immunotherapy. Molecular Cancer Therapeutics 14(4): 847–856.
Zou, M.X., W. Huang, X.B. Wang, G.H., Lv, J. Li, and Y.W. Deng. Identification of mir-140-3p as a marker associated with poor prognosis in spinal chordoma. International Journal of Clinical and Experimental Pathology 7(8): 4877.
Zou, M.X., A.B. Peng, G.H., Lv, X.B. Wang, J. Li, X.L. She, and Y. Jiang. 2016. Expression of programmed death-1 ligand (pd-l1) in tumor-infiltrating lymphocytes is associated with favorable spinal chordoma prognosis. American Journal of Translational Research 8(7): 3274.
Liu, Q., G.Q. Du, Z.T. Zhu, C. Zhang, X.W. Sun, J.J. Liu, X. Li, Y.S. Wang, and W.J. Du. 2015. Identification of apoptosis-related micrornas and their target genes in myocardial infarction post-transplantation with skeletal myoblasts. Journal of Translational Medicine 13(1): 1.
Peng, J.S., S.Y. Chen, C.L. Wu, H.E. Chong, Y.C. Ding, A.L. Shiau, and C.R. Wang. 2016. Amelioration of experimental autoimmune arthritis through targeting of synovial fibroblasts by intraarticular delivery of micrornas 140-3p and 140-5p. Arthritis & Rheumatology 68(2): 370–381.
Li, Q., Y. Yao, G. Eades, Z. Liu, Y. Zhang, and Q. Zhou. 2014. Downregulation of mir-140 promotes cancer stem cell formation in basal-like early stage breast cancer. Oncogene 33(20): 2589–2600.
Ali, H., S.E. Glont, F. Blows, E. Provenzano, S.J. Dawson, B. Liu, L. Hiller, J. Dunn, C. Poole, S. Bowden et al. 2015. Pd-l1 protein expression in breast cancer is rare, enriched in basal-like tumours and associated with infiltrating lymphocytes. Annals of Oncology 26(7): 1488–1493.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this paper
Cite this paper
Kapodistrias, N., Bobori, C., Theocharopoulou, G. (2017). MiR-140-3p Downregulation in Association with PDL-1 Overexpression in Many Cancers: A Review from the Literature Using Predictive Bioinformatics Tools. In: Vlamos, P. (eds) GeNeDis 2016. Advances in Experimental Medicine and Biology, vol 988. Springer, Cham. https://doi.org/10.1007/978-3-319-56246-9_18
Download citation
DOI: https://doi.org/10.1007/978-3-319-56246-9_18
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-56245-2
Online ISBN: 978-3-319-56246-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)