Abstract
MicroRNAs (miRNAs) as a class of small noncoding RNA molecules regulate the expression of targeted gene. The dysregulation of microRNAs is reported to be involved in carcinogenesis and tumor progression. Here, we identified miR-140-3p as a downregulated microRNA in most cancer tissues including lung cancer tissues, compared with their normal counterparts. MiR-140-3p was observed to perform its tumor suppressor function via its inhibition on cell growth, migration and invasion but its induction of cell apoptosis. Furthermore, the growth of non-small-cell lung cancer (NSCLC) cells in nude mouse models were suppressed by overexpression of miR-140-3p. ATP8A1 was demonstrated as a novel direct target of miR-140-3p using a luciferase assay. The increased level of intracellular ATP8A1 protein attenuated the inhibitor role of miR-140-3p in the growth and mobility of NSCLC cell. A regulation mechanism of miR-140-3p for the development and progression of NSCLC through downregulating the ATP8A1 expression was first discovered in the present study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the most common cancer all over the world in terms of incidence and mortality [1]. The main types of the disease are small-cell lung cancer (SCLC), and non-small-cell lung cancer (NSCLC). NSCLCs account for 80 % of lung cancer cases [2, 3]. Squamous cell carcinoma, large cell carcinoma, and adenocarcinoma are the most usual subtypes of NSCLC, and they occur in unusual histological variants [4]. NSCLC is relatively insensitive to chemotherapy or radiation, compared with SCLC, and it is usually insidious and produces no symptoms until the disease is well advanced. Due to these characteristics of NSCLC, the prognosis of NSCLC is still unfavorable, with less than 15 % of the patients surviving beyond 5 years [5]. Thus, it is in great need of development and progression for improving the diagnosis, prevention and therapy of this disease.
A microRNA (miRNA) of 20~22 nucleotides in length is a kind of noncoding RNA which regulates gene expression in a post-transcriptional manner by binding to the complementary sequences on target message RNA transcripts [6, 7]. The “seed” region of a microRNA is the most important known determinant for the microRNA to recognize its target message RNA transcripts [8]. The proteins, Drosha and Dicer, are essential to the biogenesis of most miRNAs. After a gene-encoding microRNA is transcribed, the primary miRNA transcript containing a short internal stem-loop structure is recognized and cleaved by Drosha as part of a “microprocessor” complex and processed to a new form of miRNA precursor named pre-microRNA [9, 10]. And then, Dicer binds to the pre-microRNA and cleaves the RNA transcript to a microRNA duplex with one strand as a mature microRNA [11, 12]. The mature microRNA inhibits the expression of its target gene associated with Argonaute proteins within RNA-induced silencing complex (RISC) [13].
Dysregulation of miRNA expression caused by a variety of mechanisms in tumor cells has been described widely and it has been reported to be closely associated with tumor initiation, promotion, and progression via enhancing the expression of key oncogenes or repressing the gene of tumor suppressors [14, 15]. The upregulation of miR-218 resulted in the reduction of cell migration, invasion, proliferation and stem-like qualities by suppressing the expression of the oncogene Bmi1 [16]; miR-195 performed a negative role in the proliferation of glioma cells by downregulating expression of cyclin D1 and cyclin E1 [17]; miR-137 inhibits the growth of gliomas cells by directly targeting Rac1 [18]; miR-16 functions as a tumor suppressor gene in glioma growth and invasiveness through inhibition of BCL2 and the NF-KB1/MMP-9 signaling pathway [19].
The precursor of miR-140, pre-miR-140, was found to locate in one intron of the WW domain containing E3 ubiquitin protein ligase 2 (WWP2) gene and processed to produce two mature microRNAs with different seed sequences [20]. miR-140-5p arises from the 5′ arm of the hairpin existing in precursor while miR-140-3p from the 3′ arm. A regulatory sequence upstream of pre-miR-140 has been discovered in the regulation of miR-140 transcription through the interaction with transcription factors, NFAT3 and SMAD3 in osteoarthritis [21]. Compared with the normal tissues, the expression of miR-140 is downregulated in cell carcinomas including lung cancer cells [22–25]. The outcomes of recent studies have shown that miR-140s not only perform negative roles in tumor growth and metastasis but also suppress the migration and invasion of cancer cells by downregulating the expression of their special target genes [26–28]. In our study, we found that the expression of miR-140-3p was downregulated in NSCLC tissues, compared with their normal counterparts and miR-140-3p performed its negative role in cell growth, migration, and invasion, but positive role in cell apoptosis. We aimed to uncover the regulation mechanism of miR-140-3p for the development and progression of NSCLC through the present study.
Experimental procedures
Tissue sample collection and cell culture
Human NSCLCs and their adjacent normal tissues were obtained from 20 patients. The tissues used for RNA extraction were quick-freezed in liquid nitrogen and stored at −80 °C. Written consent was acquired from each patient who has been informed and this research obtained the agreement by the Medical Ethics and Human Clinical Trial Committee. Two NSCLC cell lines (SPC-A1 and H1299) were purchased from the American Type Culture Collection and cultivated in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Scientific HyClone, Beijing, China) supplemented with 10 % fetal bovine serum, 100 U/mL penicillin and 100 mg/mL streptomycin (Invitrogen, Carlsbad, CA, USA). All cells were incubated in 5 % CO2 humid atmosphere at 37 °C.
RNA isolation
Total RNA containing small RNA was extracted from the tissue using a mirVana ncRNA Isolation Kit according to the manufacturer’s instruction (Ambion, Austin, TX). The RNA concentration was determined according to the measured OD260 value and the RNA purity was evaluated on the basis of OD260/280 ratio detected using NanoDrop 2000c Spectrophotometer (Thermo scientific, Wilmington, USA). Total RNA integrity was assessed using a common method which is to run an aliquot of RNA sample on a non-denaturing agarose gel electrophoresis stained with ethidium bromide. The samples with clearly defined 28S and 18S ribosomal RNA (rRNA) bands observed in the ethidium bromide-staining pattern were used for further analysis.
Transfection of NCSCL cells
pcDNA:ATP8A1 vector ready for transfection were replicated and extracted from Escherichia coli X-blue using DNA Midiprep Kit (Qiagen, Hilden, Germany). Mature miR-140-3p and appropriate negative control mimics were synthesized by AuGCT co. Ltd. The RNA mimics and vector overexpressing ATP8A1 were transfected into NSCLC cells cultured on six-well plate using Lipofectamine 2000 (Invitrogen) following the manual protocol, respectively. Cells were harvested 48 h post transfection for qRT-PCR analyses or Western blot assays.
Bead array-based MicroRNA detection
The expression levels of miR-140-3p among different pairs of cancer tissues and their normal counterparts was performed using the MicroRNA Profiling β-Test Assay Kit for Sentrix Array Matrixes (Illumina, CA). All steps were performed according to the manufacturer’s protocol. In brief, the miRNA microarray assays were started with 500 ng of total RNA. Input RNA was firstly polyadenylated and converted to complementary DNA (cDNA) using a biotinylated oligo-dT primer with a universal PCR sequence at its 5′ end. And then, the modified cDNA was annealed to the miRNA-specific oligo and the cDNA/oligo complexes were then caught by streptavidin-conjugated paramagnetic particles. After the removal of mis-hybridized or non-hybridized oligos by washing, the miRNA-specific oligos was extended by the polymerase added. The PCR amplification of the extended miRNA-specific primers was performed and the labeled PCR products corresponded to miR-140-3p in the RNA samples. The single-stranded PCR products generated by PCR were ready for hybridization. Finally, the signals of hydridization were detected by confocal laser scanning.
Quantitative RT-PCR
Quantitative real-time PCR (qRT-PCR) was performed to determine the expression level of miR-140-3p and ATP8A1 mRNA using iQTM5 Multicolor Real-Time PCR Detection system (Bio-Rad) with Rea-ltime PCR Master Mix (SYBR Green, Toyobo, Osaka, Japan). miR-140-3p and U6 RNA were converted to corresponding complementary DNA (cDNA). The RT reaction was performed under the following conditions: 42 °C for 15 min; 85 °C for 5 s; and then held on 4 °C. The PCR reaction was conducted at 95 °C for 30 s and followed by 40 cycles of 95 °C for 5 s and 60 °C for 34 s in the ABI 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA). The consequences of qRT-PCR were analyzed and shown as relative miRNA expression of CT (threshold cycle) value, which was then converted to fold changes. The reverse transcription of ATP8A1 mRNA was carried out using random primers under the conditions described in the instructional protocol of SuperScript III reverse transcriptase (Takara, Japan). The relative levels of ATP8A1 mRNA were calculated on the basis of the difference between amplification of ATP8A1 and glyceraldehyde 3-phosphate dehydrogenase (GAPDH) mRNA using the 2-Δct method. Each assay was performed independently in triplicate at least three times.
Northern blot
The RNA moleculars with low molecular weight including miR-140-3p was obtained from total RNA using PEG8000/NaCl precipitation. RNAs were separated in a 17.5 % denaturing polyacrylamide gel containing 8-M urea, transferred to nylon membranes and probed with [γ32P] ATP-labeled oligonucleotides. Hybridization signals were detected using a Phosphorimager (Bio-Rad). Synthetic RNA oligonucleotides were loaded as size markers.
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetra-zolium bromide assays
NSCLC cells with the concentration of 104 per well were seeded in a 96-well plate. miRNA mimic, negative control mimic or eukaryotic expression plasmid were transfected into the cell using Lipofectamine 2000 at the next day. The cells were cultured at 37 °C in transfection media for 6 h. And the media was replaced by complete medium containing 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetra-zolium bromide (MTT; final concentration, 250 μg/mL) for the following assays. Plates were incubated for an additional 12–48 h, respectively. The trapped MTT crystals in cells were solubilized in 200-μL DMSO at 37 °C for 15 min. Absorbance was determined in a microtiter plate reader (Molecular Devices, Menlo Park, CA) at 490 nm. All experiments were performed in triplicate.
Wound healing and invasion assays
The wound healing assays were performed as follows. Cells were seeded in six-well plates and cultured to 100 % confluence. Then, a wound in the cell monolayer was produced using a plastic pipette tip. The cells, after being washed with PBS buffer, were cultured for another 48 h. Cell migration toward the wounded area was observed. Wound closure (%) was calculated as the area of migrated cells divided by wounded area before 48 h culture. 105 cells suspended in serum-free media were placed into the upper chamber of an insert precoated with Matrigel (BD Bioscience) and DMEM supplemented with 10 % fetal bovine serum was input into the lower chamber. The cell invasion proceeded for 48 h in assays. The cells remaining on the upper surface of the membrane was removed, whereas the cells which had invaded through the membrane were stained with 20 % methanol and 0.2 % crystal violet, imaged, and counted under a microscope (Olympus, Tokyo, Japan).
Colony formation assays
For colony formation assays, the cells were harvested and seeded at a density of 200 cells per well in 12-well plates and incubated at 37 °C and 5 % CO2 in a humidified incubator for 2 weeks. During colony growth, the culture medium was replaced every 3 days. The colony number in each well was counted and calculated.
Xenografted tumor model
Four pairs of the BALB/c nude mice were inoculated subcutaneously NSCLC cells of 3.0 × 106 infected with miR-140-3p mimics or negative control mimics in the right scapulas of nude mice. On day 25, animals were killed and tumors were photographed and weighed after excised.
Flow-cytometric analysis of apoptosis
SPC-A1 and H1299 cells transiently transfected with miR-140-3p or negative control mimics were harvested at 48 h after transfection by trypsinization. The cells, after being double stained with FITC-Annexin V and propidium iodide (PI) double following the instructional manual of Annexin V-FITC Apoptosis Detection Kit, were analyzed with a flow cytometry (FACScan; BD Biosciences) equipped with a CellQuest software (BD Biosciences). The relative ratio of early apoptotic cells transfected with miR-140-3p mimics were compared with that of the cells with negative control mimics. All of the samples were assayed in triplicate.
Bioinformatics methods
The miRNA targets predicted by computer-aided algorithms were obtained from miRanda (http://www.microrna.org/microrna/home.do), miRDB (http://mirdb.org/miRDB/), miRWalk (http://www.umm.uni-heidelberg.de/apps/zmf/mirwalk/), and targetscan (http://www.targetscan.org).
Plasmid constructs and luciferase assay
The NSCLC cells cultured in a 48-well plate were cotransfected with miR-140-3p mimics, firefly luciferase reporter comprising wild type or mutant 3′UTR of ATP8A1 gene, and a Renilla luciferase vector as control (Promega, Madison, WI, USA) according to the manual protocols described in the instruction of Lipofectamine 2000. Cells were harvested 48 h post transfection, and the luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA).
Western blot
The extraction of total protein from cells was achieved using RIPA lysis buffer supplemented with protease inhibitors cocktail (Roche) and PMSF (Roche). Protein concentration was measured using BCA protein assay (Pierce) performed by NanoDrop 2000c Spectrophotometer (Thermo scientific, Wilmington, USA). The total protein extractions were separated by 12 % SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred to NC membranes. The ATP8A1 protein was probed with specific antibodies obtained from Proteintech Co. Ltd. after the blot blocked with 5 % non-fat milk. The gray intensities of protein bands were quantified using ImageJ software (http://rsb.info.nih.gov/ij/). GAPDH antibody was purchased from sigma.
Immunohistochemistry assays
The sections of NSCLC cancer tissues and matched normal ones were mechanically deparaffinized and incubated in Target Retrieval Solution (Dako, Carpinteria, CA, USA) at 95 °C for 40 min. After the endogenous peroxidase activity was blocked with methanol containing 3 % hydrogen peroxide (Dako) for 30 min, the tissue sections were incubated with primary antibody of ATP8A1 at 4 °C overnight. The sections were then incubated in secondary antibody at 37 °C for 60 min. Subsequently, the sections were counterstained with hematoxylin for 1 min. The evaluation of immunohistochemical (IHC) staining was performed as described previously [29].
Statistical analysis
Data are presented as the mean ± standard deviation (SD). Differences were considered to be statistically significant at P < 0.05 in the results acquired from student’s t test.
Results
The downregulation of miR-140-3p expression in human NSCLC tissues
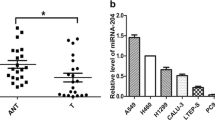
To investigate the roles of miR-140-3p in human lung cancer development, miRNA Map2.0 was used for the analysis of miR-140-3p in diverse normal tissues and tumor tissues, including lung cancer., the results in Fig. 1a shows miR-140-3p was downregulated in tumor human lung compared with normal human lung, which is in line with the results reported previously [22, 25]. The relative expression level of miR-140-3p in normal lung tissue is 1.25 times higher than that in tumor tissue (Fig. 1b). To further confirm the downregulated expression of miR-140-3p in lung carcinoma, we measured the relative expression levels of miR-140-3p normalized to U6 RNA in 20 pairs of cancer issues and adjacent normal ones using real-time RT-PCR analysis. The results showed that expression levels of miR-140-3p in 19 out of 20 tumor issues were significantly lower than the levels of the microRNA in matched normal ones (Fig. 1b). Northern blotting assays were performed on a series of 20 tumor and normal tissue pairs to validate the expression levels of miR-140-3p in samples. As the Fig. 1c shows, the results of northern blotting assays paralleled the data acquired from real-time RT-PCR assays.
Analysis of miR-140-3p in NSCLC tissues and matched normal tissues. a Detecting the relative expression level of mature miR-140-3p in 16 pairs of tumor issues and adjacent normal tissues using bead-based microarray. The pseudocolor scale bar presented transcript levels of mature miR-140-3p in different tissue samples. The determined level of miR-140-3p in NSCLC tissues and their normal counterparts were indicated with arrows. The quantitative analysis of miR-140-3p level in NSCLC tissues and matched normal ones was listed on the right. b Measuring the relative miR-140-3p level normalized to U6 RNA in 20 pairs of lung cancer tissues and their corresponding tissues using qRT-PCR. Error bars indicate standard deviation (SD). c The qualitative analysis of relative expression level of miR-140-3p standardized with U6 RNA using northern blot
Negative effect of miR-140-3p on cell viability and growth
To study the effect of miR-140-3p on the cell growth, the mature miR-140 and their corresponding negative control mimics were transiently transfected into SPC-A1 or H1299 cells. The levels of miR-140-3p in SPC-A1 transfected with miR-140-3p mimics increased by 8.2-fold while that in H1299 with the microRNA mimics by 7.8-fold, when compared with the miR-140-3p levels in tumor cells transfected with their negative control mimics (Fig. 2a). The cell viabilities of the lung cancer cells, SPC-A1 and H1299, were decreased by 0.6- and 0.7-fold after 48 h post-transfection of miR-140-3p respectively, compared with that of the tumor cells transfected with negative control mimics according to the results of MTT assays (Fig. 2b, c). Similar to the consequences of MTT assays, the data obtained from colony formation assays showed that the colony numbers of the two cell lines treated with miR-140-3p decreased to 36 % of the numbers of the cells with negative control mimics for SPC-1A and 51 % for H1299 cells approximately, indicating miR-140-3p performs a negative function in the long-term growth of NSCLC cells (Fig. 2d). Moreover, the influence of upregulated level of miR-140-3p on tumor growth was investigated using in vivo assays. It is apparent that the volumes of the tumors derived from the cells transfected with miR-140-3p were smaller than those originated from cancer cells with negative control mimics (n = 4 animals per group). At 25 days post-injection, the mean weight of the tumors with transfected miR-140-3p was about 38 % of that of the tumors with negative control mimics (Fig. 2e). Thus, we suggested that the transfection of miR-140-3p into lung cancer cells can inhibit cell growth both in vitro and in vivo.
Effect of miR-140-3p on cell viability and growth. a Checking the relative level of mature miR-140-3p expressed in SPC-A1 and H1299 cells transfected with miR-140-3p mimics or control mimics. b, c The viabilities of SPC-A1 and H1299 cells after the transfection of miR-140-3p or control mimics were determined using MTT assays. Data represent the mean ± SD from three independent experiments. d Colony-forming growth assays revealed the proliferation capacity of SPC-A1 and H1299 transfected with miR-140-3p or control mimics. e The ability of NSCLC cell with miR-140-3p or control mimics in growth in vivo was determined after the injection of transfected cell into nude mice. Error bars indicate SD. *P < 0.05
The inhibition of cell migration and invasion but induction of cell apoptosis by miR-140-3p
A cancer cell has a capacity to spread through the tissues relying on its ability to undergo migration and invasion. Although the cellular and molecular mechanism of migration and invasion is relatively poorly understood so far, microRNAs have been discovered to play important roles in the invasion and metastasis in lung cancer [30, 31]. To test whether miR-140-3p had an impact on the NSCLC cell migration and invasion, we carried out wound healing assays and transwell invasion assays. As Fig. 3a shows, the closure rate of SPC-A1 cells transfected with miR-140-3p mimics decreased by 0.37-fold while that of H1299 cells with the microRNA mimics by 0.34-fold, compared with the rate of tumor cells with control mimics. Similarly, the abilities in invasion of SPC-A1 reduced by 0.64-fold while that of H1299 by 0.71-fold after transfection with miR-140-3p mimics, when compared with the invasion abilities of the cancer cells transfected with negative control mimics (Fig. 3b). As described previously, apoptosis is a key process in cancer development and progression [32, 33]. It is interesting to know if miR-140-3p expression should affect the apoptosis of NSCLC cell. The flow cytometry analysis revealed that the apoptosis rate of SPC-A1 cells transfected with miR-140-3p increased by 0.44-fold while that of H1299 cell with the microRNA mimics by 0.51-fold, compared with the apoptosis rate in control groups. These data indicated that miR-140-3p functioned as a tumor suppressor that inhibited the growth, migratory and invasive phenotype of NSCLC cells but enhanced their programmed cell death (Fig. 3d).
Impact of miR-140-3p on cell migration and invasion in vitro. a Wound healing assays were used to investigate the migratory ability of NSCLC cells transfected with miR-140-3p or control mimics. b Transwell assays were performed to uncover the invasive capacities of NSCLC cells with miR-140-3p or control mimics. c The apoptosis rates of SPC-A1 and H1299 after transfection with miR-140-3p or control mimics were calculated out through the data analysis of flow cytometry. Error bars indicate SD. *P < 0.05
ATP8A1 identified as a direct target of miR-140-3p in NSCLC cell
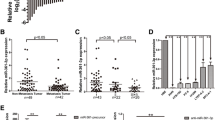
To uncover the molecular mechanism by which miR-140-3p performs a suppressive role in tumor development, four cited programs, miRanda, miRDB, miRWalk, and Targetscan, were exploited to predicate the potential targets of the miR-140-3p in NSCLC cells. Nine genes were considered as the candidates after our analysis using all programs (Fig. 4a). ATP8A1 were reported to participate in the transportation of exposure phosphatidyl-serine which is a sign of early apoptosis back to the inner layer [34, 35]. Recently, the complex of ATP8A1 and CDC50A is described to be related to the cell migration [36]. Thus, ATP8A1 gene was selected for further analysis. The wild-type 3′ untranslated region of ATP8A1 which was supposed to be bound by miR-140-3p and its mutant without putative binding sites of miR-140-3p were fused directly downstream of the firefly luciferase gene (pLuc), respectively. The complementary sequences between miR-140-3p and the wild-type 3′ untranslated region of ATP8A1 or its mutant were described in Fig. 4b. Two constructed vectors, Luc-ATP8A1-3′UTR-WT and Luc-ATP8A1-3′UTR-MUT, were cotransfected with miR-140-3p mimics into SPC-A1 or H1299 cells, respectively. The luciferase activity of reporter protein encoded by the gene locating at Luc-ATP8A1-3′UTR-WT decreased by 0.54-fold in SPC-A1 cells and by 0.61-fold in H1299 cells, respectively, compared with that of the reporter protein encoded by the gene lying in Luc-ATP8A1-3′UTR-MUT. However, there were no obvious differences in luciferase activities of reporter protein encoded by the gene situated at Luc-ATP8A1-3′UTR-WT in tumor cells when the vectors cotransfected with the control mimics (Fig. 4c). These data indicated that miR-140-3p inhibited the luciferase activity of reporter protein by directly binding to their putative binding sites which were mutated in the research. The results of further studies on the protein levels using western blot show that the relative level of ATP8A1 normalized to GAPDH decreased by 0.72-fold in SPC-A1 and by 0.59-fold in H1299 cells transfected with miR-140-3p mimics, compared with the level in NSCLC cells with the control mimics (Fig. 4d). To test if miR-140-3p could regulate ATP8A1 expression at mRNA level, the mRNA level of ATP8A1 was monitored in NSCLC cells transfected with miR-140-3p or the negative control mimics. Quantitative real-time PCR analysis revealed the mRNA level of ATP8A1 decreased by 0.67-fold in SPC-A1 and by 0.59-fold in H1299 cells, compared with the corresponding mRNA level in NSCLC cells with control mimics (Fig. 4e). Consequently, the increased level of miR-140-3p in NSCLC cell had a significant silencing impact on endogenous ATP8A1 expression both at the mRNA and protein level.
The ATP8A1 gene directly targeted by miR-140-3p. a The potential targeted genes were listed according to the predication by all four algorithms. The gene to be further analyzed, ATP8A1 gene, was circled. b The 3′ untranslated region (UTR) fragments of ATP8A1 gene and its mutant as indicated with two arrows, respectively. The putative base pair region was labeled with vertical lines between the words representing the bases. c The relative firefly activities generated from the luciferase reporter plasmids containing wild-type or mutant ATP8A1 3′UTR in NSCLC cells with miR-140-3p or control mimics were determined using the dual luciferase assay. d Western blot analyses of ATP8A1 protein level following treatment of SPCA1 cells and H1299 cells with miR-140-3p or control mimics were performed. e qRT-PCR analyses of ATP8A1 mRNA level following treatment of SPCA1 cells and H1299 cells with miR-140-3p or control mimics were performed. GAPDH was used as control. Error bars indicate SD. *P < 0.05
The impact of miR-140-3p on the NSCLC cell growth and invasion reversed by ATP8A1 overexpression
To determine whether the increased level of ATP8A1 protein could reverse the tumor-suppressive effects of miR-140-3p, the vector overexpressing ATP8A1, pCDNA3.0::ATP8A1, was cotransfected with miR-140-3p mimics into SPC-A1 and H1299 cells, respectively. The level of miR-140-3p in cell contransfected with the microRNA and pCDNA3.0::ATP8A1 decreased by 0.13-fold in SPC-A1 and by 0.15-fold in H1299 cell, compared with the miR-140-3p level in cells just transfected with miR-140 mimics, but increased by 3.8-fold in SPC-A1 and by 4.7-fold in H1299 cells, compared with the microRNA level in cells just with negative control mimics (Fig. 5a). The relative expression level of ATP8A1 protein in NSCLC cells cotransfected with miR-140-3p mimics and plasmids overexpressing ATP8A1 were close to those in NSCLC cells with control mimics but significantly higher than those in cells transfected only with miR-140-3p mimics (Fig. 5b). The colony number of SPC-A1 cells increased by 1.1-fold after ATP8A1 overexpressing while that of H1299 cells by 0.81-fold, compared with the colony number of the NSCLC cells only with miR-140-3p mimics (Fig. 5c, d). Likewise, the percentage of invading cells increased by 1.71-fold in SPC-A1 and by 1.91-fold in H1299 cells contransfected with miR-140-3p mimics and pCDNA3.0::ATP8A1, compared with that in NSCLC cells after tranfected only with miR-140-3p mimics (Fig. 5e, f). These data indicated that upregulation of ATP8A1 expression could offset the negative impact of miR-140-3p on NSCLC cell growth and invasion partially.
The inhibition of cell growth and invasion caused by miR-140-3p reversed by the overexpression of ATP8A1 in NSCLC cells. a The relative expression levels of miR-140-3p were detected in the NSCLC cells transfected with miR-140-3p or control mimics, and those cotransfected with miR-140-3p and pcDNA3.0::ATP8A1 using qRT-PCR. b The relative protein levels of ATP8A1 were determined in the NSCLC cells transfected with miR-140-3p or control mimics, and those cotransfected with miR-140-3p and pcDNA3.0::ATP8A1 using western blot. c The colonigenic capacities of the NSCLC cells with miR-140-3p or control mimics, and those cotransfected with miR-140-3p and pcDNA3.0::ATP8A1 were described using colony formation assays. d The invasion abilities of the NSCLC cells with miR-140-3p or control mimics, and those cotransfected with miR-140-3p and pcDNA3.0::ATP8A1 were presented using colony formation assays. Experiments were performed in triplicate. Error bars indicate SD. *P < 0.05
The upregulation of ATP8A1 expression in human NSCLC tissues
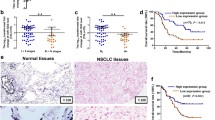
Since the downregulated expression level of miR-140-3p was observed in NSCLC cell as described above, it was reasonable that the expression of ATP8A1 might be enhanced in lung cancer tissues compared with adjacent normal lung tissues. To verify this hypothesis, we checked the relative levels of ATP8A1 mRNA in 20 pairs of cancer tissues and adjacent normal ones using real-time RT-PCR. The data analysis revealed that the mRNA levels in 18 cancer tissues were obviously higher than those in their adjacent normal lung tissues (Fig. 6a). Furthermore, determination of ATP8A1 protein levels in 20 pairs of NSCLC tissues and their normal counterparts using western blot showed that the expression levels of ATP8A1 proteins were significantly upregulated in NSCLC cells (Fig. 6b). Consistent with the results above, the fluorescence of ATP8A1 proteins immunohistochemical stained in lung cancer tissues were distinctly stronger than that in adjacent normal lung tissues (Fig. 6c). All these data indicated that the expression of ATP8A1 was enhanced in the lung cancer tissues.
Analysis of ATP8A1 in NSCLC tissues and matched normal tissues. a Determining the relative mRNA level of ATP8A1 normalized to GAPDH mRNA in 20 pairs of lung cancer tissues and their matched normal counterparts using qRT-PCR. Error bars indicate standard deviation (SD). b The relative quantitative analysis of the protein levels of ATP8A1 in 20 pairs of lung cancer tissues and their adjacent normal ones standardized with GAPDH protein using western blot. c The qualitative analysis of ATP8A1 proteins in NSCLC tissue and the corresponding normal tissue was performed using immunostaining. *P < 0.05
Discussion
Variations in the expression of microRNAs have been identified in lung cancer tissues with respect to the normal tissue counterparts. The expression of microRNAs including miR-21, -191, -210, -155, -205, -17-3p, -214, -212, -106a, -192, -197, -146, -203, and -150 were reported to be upregulated in lung cancer but the levels of expressed microRNAs involving miR -126*, -143, -224, -126, -30a-5p, -149, -198, -145, -95, -218, -124a, -27b, -32, -29b -2, -220, -33, -101-1, -125a, and -124a-3 were decreased [37]. The decreased expression of microRNAs miR-15a and -21 has been noted in two common types of non-small cell lung cancer, squamous cell carcinomas and adenocarcinomas [38]. In similar studies, the levels of five microRNAs including miR-210, -182, -486-5p, -30a, and -140-3p were successfully detected to distinguish squamous cell carcinomas from normal lung tissues and miR-31 functions as a promoter in cancer development by repressing the expression of tumor suppressor DICER1. And downregulations of miR-107, miR-185, and let-7a and upregulation of miR-31 expressed in lung cancer tissues and cancer cell lines have been found and overexpressions of miR-107 and miR-185 play made a negative effect on cell proliferation through causing G1 cell cycle phase arrest in cancer cells [39]. The alterations of microRNA expressions in tumors can be attributed to a variety of mechanisms, including deletions, amplification, or mutations of microRNA loci [40–42]. In addition, the expression of microRNAs can be affected by epigenetic changes or by dysregulation of transcription factors [43, 44].
Estrogen receptor α signaling regulates tumor-initiating cells by suppressing the expression of miR-140 which functions as an inhibitor in the expression of the transcription factor SOX2 in breast cancer [45]. It has been recently reported that miR-140-5p had a negative impact on the development and progression of hepatocellular carcinoma by inhibiting transforming growth factor β (TGF-β) and mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) signaling and suppressed the migration and invasion of human tongue cancer cells by downregulating the expressions of LAMC1, HDAC7, and PAX6 targeted by the microRNA directly but upregulating the expression of ERBB4 due in part to the suppression of ADAM10 by this microRNA(26, 28). MiR-140 was characterized as a tumor suppressor because of its inhibiting the proliferation, migration and invasion of the cells by downregulating the IGF1R expression (27).
In the present study, we found that the relative levels of miR-140-3p in most lung cancer tissues (19 out of 20) were significantly higher than those in matched normal ones (Fig. 1b). The increased level of miR-140-3p in NSCLC cells leads to the decrease of cell growth either in vitro or in vivo (Fig. 2d, e). The migration and invasion of tumor cells were repressed following the transfection with miR-140-3p (Fig. 3a, b). However, miR-140-3p performs a positive role in the apoptosis of NSCLC cells (Fig. 3c). All these evidences indicated that miR-140-3p worked as a tumor suppressor in non-small cell lung cancer which was consistent with previous reports on miR-140. ATP8A1 was identified as a novel target of miR-140-3p in NSCLC cells through our study on the mechanism of the inhibition.
ATP8A1 has been described as an aminophospholipid translocase (APLT) or flippase which is responsible for the translocation of phosphatidyl-serine (PS) and phosphatidylethanolamine (PE) across lipid bilayers [46]. The deficiency of ATP8A1 can result in the exposure of phosphatidyl-serine on the surface of the cells [47]. The collapse of membrane lipid asymmetry with phosphatidyl-serine externalization is the hallmark of cells undergoing apoptosis[48, 49]. Our study shows that miR-140-3p inhibited the expression of ATP8A1 at both mRNA and protein levels by binding directly to its 3′UTR depending on the base pair region between its seed sequence and UTR (Fig. 4b–e). Transporting the exposed PS and PE back to the inner layer may be frustrated caused by the disrupted expression of ATP8A1. Thus, more apoptosis cells after the transfection with miR-140-3p were detected using flow cytometry (Fig. 3c). Macrophage can recognize externalized PS on cells using a receptor to remove apoptotic cells [50]. The smaller volume of the tumors derived from the NSCLC cells with high level of miR-140-3p may be partially due to the phagocyte when the tumors growing in nude mice (Fig. 2e). As reported, depletion of ATP8A1 proteins inhibited the cell migration because of a severe defect in the formation of membrane ruffles (38). MiR-140-3p resulted in the low mobility of the NSCLC cells by downregulating the protein level of ATP8A1 (Fig. 3a, b). And the suggestion was further supported by the following experimental result that overexpression of ATP8A1 reversed the inhibition of NSCLC cell invasion (Fig. 5e, f). As shown in Fig. 5c, d, ATP8A1 overexpression was able to cancel out negative impacts on the growth of NSCLC cells caused by miR-140-3p in part. That may suggest that miR-140-3p should suppress the growth of NSCLC cell via the pathways other than inhibiting ATP8A1 expression. In conclusion, we have revealed that miR-140-3p acted as a tumor suppressor in non-small cell lung cancer cells by downregulating the expression of ATP8A1 to inhibit the cell growth, migration and invasion, but to promote the apoptosis of the tumor cells.
References
Siegel R, Ward E, Brawley O, Jemal A. Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA: A Cancer J Clini. 2011;61:212–36.
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA: A Cancer J Clin. 2011;61:69–90.
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA: A Cancer J Clin. 2012;62:10–29.
Asmis TR, Ding K, Seymour L, Shepherd FA, Leighl NB, Winton TL, et al. Age and comorbidity as independent prognostic factors in the treatment of non small-cell lung cancer: A review of national cancer institute of canada clinical trials group trials. J Clin Oncol: Off J Ame Soc Clin Oncol. 2008;26:54–9.
Jemal A, Murray T, Ward E, Samuels A, Tiwari RC, Ghafoor A, et al. Cancer statistics, 2005. CA: A Cancer J Clin. 2005;55:10–30.
Bartel DP. Micrornas: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Ambros V. The functions of animal micrornas. Nature. 2004;431:350–5.
Zanetti KA, Haznadar M, Welsh JA, et al. 3′-UTR and functional secretor haplotypes in mannose-binding lectin 2 are associated with increased colon cancer risk in African Americans. Cancer Res. 2012;72(6):1467–77.
Kirigin FF et al. Dynamic microRNA gene transcription and processing during T cell development. J Immunol. 2012;188:3257–67.
Zeng Y, Yi R, Cullen BR. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 2005;24:138–48.
Macrae IJ et al. Structural basis for double-stranded RNA processing by Dicer. Science. 2006;311:195–8.
Lau PW, Guiley KZ, De N, Potter CS, Carragher B, MacRae IJ. The molecular architecture of human Dicer. Nat Struct Mol Biol. 2012;19:436–40.
Pasquinelli AE. MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet. 2012;13:271–82.
Calin GA, Croce CM. Microrna signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10:704–14.
Tu Y et al. MicroRNA-218 inhibits glioma invasion, migration, proliferation, and cancer stem-like cell self-renewal by targeting the polycomb group gene Bmi1. Cancer Res. 2013;73(19):6046–55.
Hui W et al. MicroRNA-195 inhibits the proliferation of human glioma cells by directly targeting cyclin D1 and cyclin E1. PLoS One. 2013;8(1), e54932.
Guan S et al. Overexpressed miRNA-137 inhibits human glioma cells growth by targeting Rac1. Cancer Biother Radiopharm. 2013;28(4):327–34.
Yang TQ et al. MicroRNA-16 inhibits glioma cell growth and invasion through suppression of BCL2 and the nuclear factor-kappaB1/MMP9 signaling pathway. Cancer Sci. 2014;105(3):265–71.
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A. Identification of mammalian microrna host genes and transcription units. Genome Res. 2004;14:1902–10.
Tardif G, Pelletier JP, Fahmi H, Hum D, Zhang Y, Kapoor M, et al. Nfat3 and tgf-beta/smad3 regulate the expression of mir-140 in osteoarthritis. Arthritis Res Ther. 2013;15:R197.
Sand M, Skrygan M, Sand D, Georgas D, Hahn SA, Gambichler T, et al. Expression of micrornas in basal cell carcinoma. Br J Dermatol. 2012;167:847–55.
Iorio MV, Visone R, Di Leva G, Donati V, Petrocca F, Casalini P, et al. Microrna signatures in human ovarian cancer. Cancer Res. 2007;67:8699–707.
Tan X, Qin W, Zhang L, Hang J, Li B, Zhang C, et al. A 5-microrna signature for lung squamous cell carcinoma diagnosis and hsa-mir-31 for prognosis. Clin Cancer Res: Off J Am Assoc Cancer Res. 2011;17:6802–11.
Song B, Wang Y, Xi Y, Kudo K, Bruheim S, Botchkina GI, et al. Mechanism of chemoresistance mediated by mir-140 in human osteosarcoma and colon cancer cells. Oncogene. 2009;28:4065–74.
Kai Y, Peng W, Ling W, Jiebing H, Zhuan B. Reciprocal effects between microrna-140-5p and adam10 suppress migration and invasion of human tongue cancer cells. Biochem Biophys Res Commun. 2014;448:308–14.
Yang H, Fang F, Chang R, Yang L. Microrna-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor beta receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013;58:205–17.
Yuan Y, Shen Y, Xue L, Fan H. Mir-140 suppresses tumor growth and metastasis of non-small cell lung cancer by targeting insulin-like growth factor 1 receptor. PLoS One. 2013;8, e73604.
Usui A, Hoshino I, Akutsu Y, Sakata H, Nishimori T, Murakami K, et al. The molecular role of fra-1 and its prognostic significance in human esophageal squamous cell carcinoma. Cancer. 2012;118:3387–96.
Hu H, Li S, Liu J, Ni B. Microrna-193b modulates proliferation, migration, and invasion of non-small cell lung cancer cells. Acta Biochim Biophys Sin. 2012;44:424–30.
Chen LT, Xu SD, Xu H, Zhang JF, Ning JF, Wang SF. Microrna-378 is associated with non-small cell lung cancer brain metastasis by promoting cell migration, invasion and tumor angiogenesis. Med Oncol. 2012;29:1673–80.
Pai LM, Barcelo G, Schupbach T. D-cbl, a negative regulator of the EGFR pathway, is required for dorsoventral patterning in Drosophila oogenesis. Cell. 2000;103:51–61.
Ashkenazi A. Targeting death and decoy receptors of the tumour-necrosis factor superfamily. Nat Rev Cancer. 2002;2:420–30.
Soupene E, Kuypers FA. Identification of an erythroid atp-dependent aminophospholipid transporter. Br J Haematol. 2006;133:436–8.
Soupene E, Kemaladewi DU, Kuypers FA. Atp8a1 activity and phosphatidylserine transbilayer movement. J Recept Ligand Channel Res. 2008;1:1–10.
Kato U, Inadome H, Yamamoto M, Emoto K, Kobayashi T, Umeda M. Role for phospholipid flippase complex of ATP8A1 and CDC50A proteins in cell migration. J Biol Chem. 2013;288:4922–34.
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, et al. Unique microrna molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006;9:189–98.
Bandi N, Zbinden S, Gugger M, Arnold M, Kocher V, Hasan L, et al. Mir-15a and mir-16 are implicated in cell cycle regulation in a rb-dependent manner and are frequently deleted or down-regulated in non-small cell lung cancer. Cancer Res. 2009;69:5553–9.
Takahashi Y, Forrest AR, Maeno E, Hashimoto T, Daub CO, Yasuda J. Mir-107 and mir-185 can induce cell cycle arrest in human non small cell lung cancer cell lines. PLoS One. 2009;4, e6677.
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, et al. A microrna signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med. 2005;353:1793–801.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, et al. Frequent deletions and down-regulation of micro- rna genes mir15 and mir16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A. 2002;99:15524–9.
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, et al. A microrna expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A. 2006;103:2257–61.
Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, et al. Specific activation of microrna-127 with downregulation of the proto-oncogene bcl6 by chromatin-modifying drugs in human cancer cells. Cancer Cell. 2006;9:435–43.
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. C-myc-regulated micrornas modulate e2f1 expression. Nature. 2005;435:839–43.
Zhang Y, Eades G, Yao Y, Li Q, Zhou Q. Estrogen receptor alpha signaling regulates breast tumor-initiating cells by down-regulating mir-140 which targets the transcription factor sox2. J Biol Chem. 2012;287:41514–22.
Daleke DL, Lyles JV. Identification and purification of aminophospholipid flippases. Biochim Biophys Acta. 2000;1486:108–27.
Levano K, Punia V, Raghunath M, Debata PR, Curcio GM, Mogha A, et al. Atp8a1 deficiency is associated with phosphatidylserine externalization in hippocampus and delayed hippocampus-dependent learning. J Neurochem. 2012;120:302–13.
Fadok VA, de Cathelineau A, Daleke DL, Henson PM, Bratton DL. Loss of phospholipid asymmetry and surface exposure of phosphatidylserine is required for phagocytosis of apoptotic cells by macrophages and fibroblasts. J Biol Chem. 2001;276:1071–7.
Martin SJ, Reutelingsperger CP, McGahon AJ, Rader JA, van Schie RC, LaFace DM, et al. Early redistribution of plasma membrane phosphatidylserine is a general feature of apoptosis regardless of the initiating stimulus: inhibition by overexpression of bcl-2 and abl. J Exp Med. 1995;182:1545–56.
Fadok VA, Bratton DL, Rose DM, Pearson A, Ezekewitz RA, Henson PM. A receptor for phosphatidylserine-specific clearance of apoptotic cells. Nature. 2000;405:85–90.
Acknowledgments
This work was support by The development of science and technology plan projects of Chinese medicine in Shandong, No. 2013-209.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dong, W., Yao, C., Teng, X. et al. MiR-140-3p suppressed cell growth and invasion by downregulating the expression of ATP8A1 in non-small cell lung cancer. Tumor Biol. 37, 2973–2985 (2016). https://doi.org/10.1007/s13277-015-3452-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3452-9