Abstract
Next-Generation Sequencing (NGS) has revealed that B chromosomes in several species are enriched in repetitive DNA, mostly satellite DNA (satDNA). This raises the question of whether satDNA is important to B chromosomes for functional reasons or else its abundance on Bs is simply a consequence of properties of B chromosomes such as their dispensability and late replication. Here we review current knowledge in this respect and contextualize it within the frame of practical difficulties to perform this kind of research, the most important being the absence of good full genome sequencing for B-carrying species, which is an essential requisite to ascertain the intragenomic origin of B chromosomes. Our review analysis on 16 species revealed that 38% of them showed B-specific satDNAs whereas only one of them (6%) carried an inter-specifically originated B chromosome. This shows that B-specific satDNA families can eventually evolve in intraspecifically arisen B chromosomes. Finally, the possibility of satDNA accumulation on B chromosomes for functional reasons is exemplified by B chromosomes in rye, as they contain B-specific satDNAs which are transcribed and occupy chromosome locations where they might facilitate the kind of drive shown by this B chromosome during pollen grain mitosis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
Since satellite DNA (satDNA) was uncovered by Kit (1961) through cesium chloride (CsCl) sedimentation analysis, and its repetitive nature was shown by Waring and Britten (1966), its possible presence on supernumerary (B) chromosomes was soon claimed in the grasshopper Myrmeleotettix maculatus (Gibson and Hewitt 1970, 1972). However, the fact that Chilton and McCarthy (1973) did not find differences in the buoyant density distribution in CsCl gradients between 0B and 5B maize plants, and Timmis et al. (1975) reached similar results on B chromosomes in rye, appeared to support the conclusion that DNA of B chromosomes is roughly similar to that of standard (A) chromosomes. Likewise, Klein and Eckhardt (1976) found no significant differences between the buoyant densities or thermal denaturation profiles of B-carrying and B-lacking DNA in the mealy bug. Similarly, Dover (1975) did not find any new highly repetitious DNA family related to the presence of B chromosomes in Aegilops, by using different approaches such as comparisons of the percentage of heterologous associations in DNA/DNA hybridization experiments. Finally, the fact that Dover and Henderson (1976) reanalyzed the M. maculatus case and did not find any satellite to the main band in grasshoppers with 0, 1, or 2 B chromosomes made prevailing the conclusion that A and B chromosomes show considerable similarity in base composition.
Remarkably, 5 years later, G. Dover himself, in collaboration with A. Amos, showed the presence of satellite DNAs shared between A and B chromosomes in tsetse flies (Glossina austeni and G. morsitans morsitans) by means of CsCl gradient density centrifugation and radioactive in situ hybridization. They suggested the first molecular mode of origin for a B chromosome, with important involvement of the Y sex chromosome and satDNA amplification (Amos and Dover 1981). In maize, Peacock et al. (1981) isolated a 185-bp satellite DNA by CsCl gradient centrifugation and cloned it into a plasmid for its chromosomal mapping by in situ hybridization, concluding that this satDNA is present on A chromosomes knob heterochromatin and on the long arm proximal knob of the B chromosome, but at lower copy number. However, the first demonstration of the existence of B-specific satDNA should have to wait until Nur et al. (1988) who showed that the paternal sex ratio B chromosome (PSR) in the wasp Nasonia vitripennis contains at least three B-specific tandem repetitive DNAs (see also Eickbush et al. 1992). B-specific satDNAs were soon found in rye (Secale cereale) (Sandery et al. 1990; Blunden et al. 1993), maize (Zea mays) (Alfenito and Birchler 1993), and the Australian daisy (Brachycome dichromosomatica) (John et al. 1991; Houben et al. 1997), at the same time as satDNAs shared between A and B chromosomes were found in other species, e.g., the grasshopper Eyprepocnemis plorans (López-León et al. 1994, 1995), the fly Drosophila subsilvestris, where the pSsP216 satDNA might have arisen from the dot chromosome (Gutknecht et al. 1995), and the fish Prochilodus lineatus (de Jesus et al. 2003) (see other examples in Camacho 2005).
These observations revealed that B chromosomes at different species may contain a heterogeneous sample of satDNAs, some of them being also located on A chromosomes and others apparently being B-specific. The first hypothesis on the origin of these B-specific satDNAs was suggested by Langdon et al. (2000), who described the de novo creation of satellite repeats, from complex euchromatic sequences, on the rye B chromosome. Therefore, by the end of the twentieth century, it was already clear that satDNA may actually be a major constituent of B chromosomes, in some cases being satDNAs shared with the A chromosomes and, in others, being B-specific satDNAs which might have arisen de novo on the B chromosome itself or else reflecting B chromosome origin through hybridization, as was the case of N. vitripennis (see below).
4.2 SatDNA Is the Prevalent Repetitive DNA on B Chromosomes in Some Species
The arrival of the twenty-first century witnessed the advent of powerful DNA sequencing methods able to yield huge amounts of DNA sequences with low input of time and cost, i.e., the so-called Next-Generation Sequencing (NGS) (Mardis 2008). These NGS technologies were soon applied to the analysis of repetitive DNA thanks to the development of software, such as RepeatExplorer (RE), being able to assemble repetitive DNA from short Illumina sequences (Novák et al. 2013).
The first report on satDNA content for a B chromosome, by means of RE analysis, was performed in rye (Martis et al. 2012) and revealed that these B chromosomes contain a similar proportion of repeats as A chromosomes, but Bs showed an additional massive accumulation of B-specific satDNAs, which were characterized by exceptionally long monomers (0.9–4.0 kb), some of them suggesting chimeric origins. In addition, rye B chromosomes showed accumulation of Bianka Ty1/copia elements and plastid (NUPT)- and mitochondrial (NUMT)-derived sequences. Banaei-Moghaddam et al. (2013) later suggested that the accumulation of some of these repetitive sequences (i.e., mobile elements and satDNA) promoted the pseudogenization of many genes in the B chromosome. Likewise, Klemme et al. (2013) observed that B-enriched satellites were mostly accumulated in the nondisjunction control region of the rye B chromosome.
The finding of B chromosomes in Drosophila melanogaster (Bauerly et al. 2014) opened the possibility to analyze B chromosomes with the help of all kinds of tools amenable in this model species. These B chromosomes were primarily composed of the AATAT satellite sequence which is also characteristic of autosome 4. Later, Hanlon et al. (2018) reported that this microsatellite showed FISH (fluorescent in situ hybridization) bands on the B, autosome 4, X and Y chromosomes. In addition, these authors found another microsatellite (AAGAT) which only hybridized with the B chromosomes and autosome 4, and suggested the possible origin of B chromosomes from this A chromosome. They also found that D. melanogaster B chromosomes did not carry any known euchromatic sequence and that they are rich in transposable elements and long arrays of short nucleotide repeats, the most abundant being the AAGAT microsatellite. Likewise, Milani and Cabral-de-Mello (2014) suggested that microsatellites are important components of the B chromosome in the grasshopper Abracris flavolineata. These authors later showed, through RE analysis, that this B chromosome shows a 137 bp satDNA (AflaSAT-1) which is shared with A chromosomes (Milani et al. 2017).
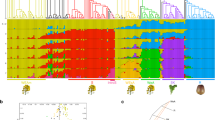
The development of the satMiner protocol, based on reiterative searches for repetitive elements on Illumina reads, by means of RepeatExplorer, allowed us to build the first satellitome in the migratory locust (Ruiz-Ruano et al. 2016). This led to the characterization of tens of different satDNA families constituting a broad satDNA catalog. SatMiner application to Illumina reads obtained from B-carrying and B-lacking genomes in the grasshopper Eumigus monticola uncovered 27 satDNA families whose FISH analysis showed the presence of eight of them on the B chromosome (Ruiz-Ruano et al. 2017). In fact, two of them (EmoSat26-41 and EmoSat27-102) showed FISH bands only on the B chromosome, thus appearing to be B-specific (Fig. 4.1). However, the bioinformatic analysis of abundance indicated that, although extremely scarce, they were also present in the 0B genome. Therefore, the most parsimonious explanation for the differential abundance of these two satDNAs on the B chromosomes is their major amplification on the B chromosome. Another satDNA family (EmoSat11-122) was extremely abundant on the B chromosome whereas it showed a single small FISH band on autosome S8 (Fig. 4.1). Remarkably, this satDNA family was composed of two different subfamilies (named A and B), and a repeat landscape (RL), quantifying genomic abundance at 1% divergence intervals, showed that subfamily B was highly amplified on the B chromosomes whereas subfamily A was highly amplified on the S8 autosome of only one of the two 0B males analyzed (Fig. 4.1). Finally, EmoSat22-12 also showed FISH bands on the B and S8 chromosomes, revealing its amplification on B chromosomes in only one of the two B-carrying males analyzed by satMiner (Fig. 4.1).
Presence of SatDNA on B chromosomes of the grasshopper Eumigus monticola. Two B-lacking (m11_0B and m13_0B, blue lines) and two B-carrying (m11_+B and m13_+B, red lines) males were analyzed by Illumina sequencing (satMiner protocol) and FISH. Two satDNA families (EmoSat26-41 and EmoSat27-102) (upper row) appeared to be B-specific because they yielded large FISH signals on the B chromosomes but no signal on A chromosomes. However, bioinformatic analysis on Illumina reads showed that they are also present on A chromosomes although at very low abundance (note the blue lines, close to the X-axis, in the repeat landscapes). Note that EmoSat26-41 showed diversification into three subfamilies (A–C) amplified on the B chromosome (red lines). The lower row shows two satDNA families showing FISH signals only on the B and S8 chromosomes, thus suggesting the possible B origin from this A chromosome. Note that EmoSat11-122 shows FISH signals across the whole B chromosome area, and its B subfamily is highly abundant on the two B-carrying individuals (most likely on the B chromosome) whereas the A subfamily was abundant on the S8 autosome of the m11_0B male. On the other hand, EmoSat22-12 showed small FISH bands on the B and S8 chromosomes and differential amplification between the two B-carrying males
The same year, Kumke et al. (2016) showed that the 5S rDNA-derived PLsatB satellite DNA found in Plantago lagopus makes up 3.3% of the 1B genotype but only 0.09% of the 0B genotype (see also Dhar et al. 2019). In the mice Apodemus flavicollis and A. peninsulae, Makunin et al. (2018) analyzed B chromosome content by means of low-pass single-chromosome sequencing and found accumulation of repetitive DNA, mainly satDNA and endogenous retroelements. In the plant Aegilops speltoides, however, Wu et al. (2019) have shown that the repetitive fraction of the genome “is mostly composed of LTR-retrotransposons, transposons and satellite repeats, with overall proportions of individual repeat types being similar in the 0B/+B genotypes.”
In the migratory locust, Locusta migratoria, Ruiz-Ruano et al. (2018) performed the quantification of repetitive DNA content of B chromosomes. They found that about 64.1% of the B-lacking genome consists of repetitive DNA, whereas this figure was higher in B-carrying genomes (64.6%) due to B chromosome enrichment in repetitive DNA. Using a subtractive approach, we found that 94.9% of the B chromosome DNA was repetitive. Specifically, 65.2% was satDNA, whereas the most abundant TEs only reached 7.9% for DNA transposons and 7% for LINEs. In addition, several gene families were found on this B chromosome, such as histone genes (2.7%), as reported previously by Teruel et al. (2010), 45S (0.25%) and 5S (0.78%) rRNA genes, snRNA genes (1.3%), especially U2 (1.1%), and tRNA genes (0.7%). Remarkably, about half of the DNA content of the B chromosome corresponded to a single satDNA (LmiSat02-176) whereas all remaining repetitive elements showed abundances lower than 4%. In fact, FISH for this satDNA family yielded a signal occupying the whole B chromosome area (Fig. 4.2). In addition, five other satDNAs showed FISH bands on the B chromosome of different sizes (Fig. 4.2), including the telomeric DNA (LmiSat07-5-tel) and the most abundant satDNA in the L. migratoria genome (LmiSat01-185) which is likely involved in the centromeric function since it is the only satDNA being located on all A chromosomes. On the B chromosome, however, this satDNA family is actually scarce (see Fig. S1b in Ruiz-Ruano et al. 2018). The bioinformatic analysis of satDNA abundance on two 0B and four B-carrying males, displayed two main patterns for their RLs (Fig. 4.3), with five satDNA families showing overabundance in all four B-carrying males (LmiSat02-176, LmiSat04-18, LmiSat09-181, LmiSat10-9, and LmiSat16-278) whereas three other showed overabundance in only two B-carrying males (LmiSat06-185, LmiSat18-210, and LmiSat53-47), suggesting the existence of polymorphic B chromosomes in the population analyzed for an abundance of some satDNA families. In addition, about half of these satDNAs showed RLs with a maximum peak of about 4–5% divergence (likewise LmiSat01-185) whereas the remaining families showed their maximum peak at 0% divergence (in resemblance to LmiSat07-5-tel) (Fig. 4.3). As the telomeric DNA is actively homogenized by the telomerase during each DNA replication, we can infer that the satDNAs displaying their maximum peak at 0% divergence have undergone recent amplification by the addition of many identical repeat units. On the other hand, those families showing their maximum peak at 4–5% showed their last major amplification some time ago so that point mutations have increased divergence for many repeat units added during that amplification. Ruiz-Ruano et al. (2018) called attention to the contrasting difference found between A and B chromosomes in L. migratoria, as TEs comprise 54% of total DNA in A chromosomes but only 19.1% of the B chromosomes. However, satDNA comprises 65.2% of the B chromosome but only 2.4% of A chromosomes. If these B chromosomes are derived from the A chromosomes, it is clear that they have followed a different molecular evolutionary pattern through a marked enrichment in satDNA.
FISH pattern for four SatDNA families in B-carrying embryos of the migratory locust (Locusta migratoria). Two of them (upper row) showed large bands which, in LmiSat02-176, occupy the whole B chromosome area, and this satDNA represented about half of the whole B chromosome DNA. No B-specific satDNAs were found in this species, as they were also present on A chromosomes, on exclusively pericentromeric locations for LmiSat02-176 but pericentromeric and interstitial locations for LmiSat06-181. The lower row shows two satDNA families yielding FISH bands on the S9 and B chromosomes, and also on S11 in the case of LmiSat04-18 (note that this cell is haploid)
Repeat landscapes for eight satDNA families showing overabundance on the B chromosome of Locusta migratoria, compared with the patterns shown by the centromeric (LmiSat01-185) and telomeric (LmiSat07-5-tel) satDNAs (upper row). Note that the left column shows satDNA families showing their maximum peak of abundance at 4–5% divergence, in resemblance to the centromeric satDNA, suggesting that they have shown a certain degree of degeneration since their last major amplification. The only exception was LmiSat10-9 whose peak was placed at 10% divergence, presumably due to faster degeneration of its extremely short repeat units (9 bp). Within the left column, note that LmiSat02-176, LmiSat04-18, and LmiSat10-9 showed overabundance (in respect to B-lacking males) in the four B-carrying males (red lines), whereas LmiSat06-185 showed this fact only in two out of the four B-carrying males analyzed, suggesting the existence of B chromosome polymorphism for the abundance of some satDNAs. The right column, however, shows satDNA families showing curves resembling the telomeric DNA pattern, i.e., with the maximum peak at 0% divergence. The only exception was LmiSat53-47 where the curves for B-carrying males showed a peak at about 3% divergence, indicating some degeneration on the B chromosome copies. These 0% peaks suggest recent amplification of these satDNA families on the B chromosome
In the same vein, Benetta et al. (2020) have recently shown that 89.80% of the PSR chromosome in Nasonia vitripennis is composed of repetitive DNA, the most abundant being complex satellites belonging to four main families (70.32%), three of which were B-specific (PSR2, PSR18, and PSR22) and the other was shared with the A chromosomes (NV79). However, only 13.97% of B-sequences corresponded to TEs. In summary, the information on the species mentioned in this section, reveals that six out of the 16 species where chromosome location was analyzed (Table 4.1), showed B-specific satDNA families (i.e., 38%). In the case of N. vitripennis, the presence of B-specific satDNA is explained by the interspecific origin of its B chromosome (McAllister and Werren 1997), but the B chromosomes in the other five species appeared to have originated intraspecifically, so that B-specific satDNAs were most likely originated in them by means of differential amplification on the B chromosomes, as explained above for EmoSat26-41 and EmoSat27-102.
4.3 SatDNA as Marker of B Chromosome Origin
During the pre-NGS times, B chromosome origin was mainly delimited between intra- or interspecific origins (see some examples in Table S1 in Ruiz-Ruano et al. 2017). Assuming that B chromosomes are most likely derived from A chromosomes, the intragenomic origin of B chromosomes is more amenable for analysis in the case of intraspecifically arisen B chromosomes, although not without serious difficulty. In the grasshopper E. plorans, we found evidence for contradictory hypotheses on the intragenomic origin of B chromosomes, as we first inferred that B chromosomes in Spanish populations most likely derived from the X chromosome because the order of a satDNA and rDNA in respect to the centromere was only coincident on the pericentromeric region of B and X chromosomes (López-León et al. 1994). We later observed that B chromosomes from Moroccan populations supported the former conclusion, but those found in populations from Daghestan (North Caucasus, Russia) were most likely derived from the smallest autosome, which was the only A chromosome carrying the three markers (5S and 45S rDNA, and the 180-bp satDNA) found on Caucasian B chromosomes (Cabrero et al. 2003). Later, sequence comparison for ITS rDNA sequences obtained from B chromosomes and several A chromosomes (X, M8, S9, S10, and S11), and for the 180-bp satDNA obtained from the X, B, and S11 chromosomes, through chromosome microdissection in spermatocytes from males collected at the Torrox population (Spain), indicated that B chromosome sequences showed higher similarity with those coming from the smallest autosome (S11) than with those from the X chromosome. This gave support to the hypothesis of B origin from the S11 autosome also in Spanish populations (Teruel et al. 2014).
In Locusta migratoria, the exclusive presence of genes for H3 and H4 histones on the B chromosome and the M8 autosome, indicated by FISH analysis, provided evidence for B chromosome origin from this A chromosome (Teruel et al. 2010). However, our FISH analysis of 58 satDNA families, previously found in this species (Ruiz-Ruano et al. 2017), on A and B chromosomes revealed that autosome S9 was the only A chromosome carrying all six satDNAs visualized on the B chromosome (Ruiz-Ruano et al. 2018), with one of them being exclusive of S9 and B (Fig. 4.2), on which basis we concluded that both M8 and S9 chromosomes could have contributed to B chromosome origin in this species.
In rye, NGS analysis allowed the identification of several B-specific repeats, mostly being satDNA (Martis et al. 2012; Klemme et al. 2013). This revealed that rye Bs showed higher ancestry from the 3RS and 7R standard (A) chromosomes, with subsequent accumulation of repeats and genic fragments from other A chromosomes and insertions of organellar DNA (Martis et al. 2012). Remarkably, accumulation of B chromosome-enriched tandem repeats was mainly found in the nondisjunction control region of the B (Klemme et al. 2013), involved in B chromosome drive (Langdon et al. 2000).
In the fish Moenkhausia sanctafilomenae, Utsunomia et al. (2016) found two types of B chromosomes both containing the same tandem repeat DNA sequences (18S rDNA, H3 histone genes, and the MS3 and MS7 satDNAs), all of which were together only in the paracentromeric region of autosome pair no. 6, suggesting that the B chromosomes derived from this A chromosome.
The FISH analysis of the full satellitome in the grasshopper Eumigus monticola (Ruiz-Ruano et al. 2017) revealed the presence of two satDNA families being informative for B origin in this species. These were EmoSat11-122 and EmoSat22-12, which only showed FISH bands on the S8 autosome and the B chromosome (Fig. 4.1). As S8 carries an interstitial FISH band for H3 histone genes which is not apparent on the B chromosome, Ruiz-Ruano et al. (2017) suggested the possible origin of this B chromosome from the proximal third of the S8 autosome, thus excluding the H3 cluster. In addition, two other satDNA families (EmoSat26-41 and EmoSat27-102) showed FISH bands only on the B chromosome, and sequence analysis provided evidence that these two satDNAs were actually present, at very low abundance, in the 0B libraries, suggesting that intraspecifically arisen B chromosomes can harbor satDNAs apparently being B-specific at cytogenetic level but not at the genomic level. In contrast, we did not find any B-specific satDNA in L. migratoria (Ruiz-Ruano et al. 2018), and Milani et al. (2018) also failed to find B-specific satDNAs in three other grasshopper species, although satDNA location on A and B chromosomes suggested that B chromosomes might have arisen, in all three cases, from one of the three shortest autosomes.
In the fish Characidium gomesi, satellitome analysis was also useful to get insights on the intragenomic origin of B chromosomes. Chromosome painting analysis suggested that B chromosomes in this species most likely derived from sex chromosomes (Pansonato-Alves et al. 2014), and subsequent FISH analysis of 18 satDNA families and the comparison of DNA sequences, obtained through chromosome microdissection, supported this hypothesis (Serrano-Freitas et al. 2020).
All these cases point to specific A chromosomes that could have been ancestors of B chromosomes that arisen intraspecifically. However, a recent analysis of B chromosome content for protein-coding genes is revealing that B chromosome origin appears to be multichromosomal, as claimed, for instance, in rye (Martis et al. 2012), the fish Astatotilapia latifasciata (Valente et al. 2014), and the plant Aegilops speltoides (Ruban et al. 2020). On this basis, the results obtained only from satDNA location should be interpreted with caution. Of course, we agree with Ruban et al. (2017) that conclusions from satDNA location remain provisional, given the dynamic behavior of satDNA repeats, as it is expected that synteny similarity for protein-coding genes would be more reliable for inferring A chromosome contribution to B chromosome origin. Unfortunately, none of the three cases above are exempt from problems, as the multichromosomal origin was not concluded using the same species genome as a reference, which dilutes the synteny advantage, and they used NGS sequences obtained either from microdissected B chromosomes, whose reliability is low for single-copy DNA, or from flow-sorted B chromosomes where minimal contamination with A chromosomes would unavoidably yield the multichromosomal pattern of B chromosome origin.
In fact, very few B-carrying species have their standard genome sequenced. One of these species is Locusta migratoria, but its genome has two serious problems since it has not yet reached the chromosome level (Wang et al. 2014) and was obtained from a B-carrying individual with the consequent interference of B-chromosome sequences for genome assembly (see Ruiz-Ruano et al. 2018, 2019). The B-carrying species with the best-sequenced genome is N. vitripennis, but a recent analysis of DNA content of its PSR chromosome has shown that it consists primarily of three complex repeats (70.32%) and other sequences that are undetectable in the standard genome and, in some cases, have strong similarity with genes from other organisms (Benetta et al. 2020), a logical expectation for B chromosomes of interspecific origin.
The origin of small supernumerary marker chromosomes (sSMCs) in humans, however, is easier to infer due to the high quality of the human genome and their youth as extra chromosomes, compared with B chromosomes, as this makes it easier inferring their A chromosome ancestry because they still have not had time for undergoing many changes in sequence or structure (Fuster et al. 2004). Recently, Makunin et al. (2018) performed the low-pass sequencing of a human sSMC derived from the pericentromeric region of chromosome 15 long arm. Interestingly, they could map the two breakpoints that yielded this extra chromosome and both were located within alphoid satDNA. They suggested that “sSMCs might correspond to an early stage of B chromosome evolution, and acquisition of drive mechanism for more efficient transmission could in principle transform those elements into true B chromosomes.” However, in our opinion, sSMCs (and any other kind of extra chromosome, included the so-called proto-Bs) lacking drive from their very origin would most likely be eliminated before they could gain drive and thus reach the polymorphism status in natural populations, and this kind of evolutionary models assuming that drive can be obtained a posteriori are all flawed (for instance, see Martis et al. 2012). In the case of sSMCs in humans, their deleterious effects make it even more unlikely their conversion into B chromosomes. In fact, there are no examples of polymorphic sSMCs in human populations beyond spontaneous cases of independently arisen ones (Fuster et al. 2004).
Evidence for possible differential geographical patterns of chromosomal location for B chromosome content of repetitive DNA in E. plorans was found by López-León et al. (2008), whose FISH analysis revealed that B chromosomes from Daghestan, Armenia, Turkey, and Greece were mostly composed of rDNA, whereas those from Spain and Morocco contained about similar amounts of rDNA and a 180-bp satDNA, the latter being actually scarce in eastern Bs. The development of a sequence characterized amplified region (SCAR) marker located at intergenic regions of 45S rDNA provided additional support to the intraspecific origin of B chromosomes in E. plorans, as its DNA sequence was identical in B chromosome variants from several localities from Spain and Morocco, and it was highly similar in B chromosome variants from Greece and Armenia. The scarce sequence variation observed between such distant populations suggested either a functional constraint or, most likely, a recent and unique origin for B chromosomes in this species (Muñoz-Pajares et al. 2011). The later finding that the widespread geographical distribution of the B1 variant makes it the best candidate for being the ancestor B chromosome in the whole western Mediterranean region (Cabrero et al. 2014) is consistent with a recent origin of B chromosomes in this species.
In some cases, B chromosomes showing highly similar size and morphology have been found in closely related species, and the high-throughput analysis of satDNA has revealed interesting insights on B chromosome origin. This was the case of several fish species of the genus Astyanax, where Silva et al. (2016) analyzed the large metacentric B chromosomes in A. bockmanni, A. paranae, and A. fasciatus, by means of (1) chromosome painting, (2) FISH for 18S rDNA, the H1 histone genes, the As51 satDNA and the (AC)15 microsatellite, and (3) ITS rDNA sequence comparison between genomic DNA from B-lacking individuals and DNA obtained from the metacentric B chromosomes in the two latter species. Whereas approaches 1 and 3 suggested the common origin of B chromosomes at different species, approach 2 failed to do it. Subsequent analysis of the satellitome in A. paranae revealed the presence of 45 satDNA families, 35 of which were analyzed by FISH in A. paranae, A. fasciatus, and A. bockmanni, showing that most satDNA families were shared between the three species and showed highly similar patterns on their B chromosomes (Silva et al. 2017). The exceptions were two B-specific satDNAs in A. paranae (ApaSat44-21 and ApaSat20-18), the former not being observed on A or B chromosomes of the two other species and the latter showing FISH bands on them. The symmetric location of many satDNAs on both B chromosome arms demonstrated the isochromosome nature of these large metacentric B chromosomes, and their high similarity in satDNA content and location gave additional support to the common origin hypothesis for these B chromosomes. Recently, the analysis of gene content in the large metacentric B chromosomes of these three species plus A. scabripinnis, by means of the genome and transcriptome sequencing and qPCR, has revealed that the Bs in the four species showed such high similarity in gene content that cannot be explained by chance, thus giving stronger support to the common origin hypothesis (Silva et al. 2021).
4.4 Function of satDNA for B Chromosomes
If satDNA accumulates into B chromosomes becoming the most abundant DNA type in them, a pertinent question is whether it plays an important function for B chromosomes. In principle, we should not expect that a possible function would have nothing to do with being transcribed, as satDNA typically belong to the noncoding fraction of repetitive. Alternatively, satDNA might accumulate in B chromosomes because they are dispensable and thus tolerate the burden of carrying high amounts of useless DNA, as long as this burden can be faced by the host genome which makes the machinery for DNA replication. It is conceivable that the late replication which characterizes B chromosomes (Fox et al. 1974) might facilitate replication errors leading to satDNA accumulation on B chromosomes. In addition, the dispensability of B chromosomes may make them be prone to these failures in DNA replication, e.g., unequal crossovers, leading to the accumulation of satDNA (and other tandem repeats) on them.
A possible functional role of satDNA was suggested for rye B chromosomes, after finding that the E3900 and D1100 B-specific satDNAs are transcriptionally active in the subterminal domain of the B chromosome, which acts as the nondisjunction control region, with the B-transcripts possibly functioning as structural or catalytic RNA (Carchilan et al. 2007). Recently, Gómez-Aldecoa (2021) has found another B-specific satDNA (ScCL11-1), which is interspersed with E3900 in the nondisjunction control region and is the only satDNA being also located on the pericentromeric region of the B chromosome, where persistent cohesion maintains the two B chromatids together for migration to the generative pole during pollen grain mitosis. Banaei-Moghaddam et al. (2012) suggested the possibility that B-derived RNAs could act as guide molecules to direct protein complexes to specific genomic loci, such as the B pericentromeric regions. However, these authors noticed that it is not known whether the B transcripts act directly or indirectly on B nondisjunction, and suggested the possibility that some protein-coding genes located in the rye B chromosome (Martis et al. 2012) might also play a role in nondisjunction control.
The case of B rye is thus suggestive for a possible function of a specific satDNA to increase B chromosome viability in natural populations through transcription to yield noncoding RNAs facilitating B drive, with or without interaction with proteins also coded by B chromosome themselves, thus interfering with the normal course of cell division regulation. In N. vitripennis, the PSR chromosome expresses a unique set of small RNAs derived from several satDNAs (Li et al. 2017) and contains a gene named haploidizer, which appears to be involved in the sex conversion which this B chromosome drive is based on (Benetta et al. 2020). This B chromosome system is thus the best positioned in the race of demonstrating the molecular basis of B chromosome drive, even though many details are still unknown.
4.5 Future Directions
The extreme scarcity of quality reference genomes of B-carrying species, at the chromosome level, is a serious handicap to investigate the A chromosome ancestry of B chromosomes. Meanwhile, satellitome analysis constitutes an excellent tool to get some insights in this respect in non-model species. In the case of intraspecifically arisen B chromosomes, the best tool would be gene content and the syntenic resemblance between A and B chromosomes, but it needs previous obtaining of high-quality sequenced genomes of B-lacking individuals in the same B-carrying species, a goal that has not yet been reached for any intraspecifically originated B chromosome.
Regarding a possible function of satDNA for B chromosomes, a first indication could be its transcription, as recent research has shown that both B chromosomes (Huang et al. 2016; Ma et al. 2016; Navarro-Domínguez et al. 2017; Kinsella et al. 2019) and satDNA (Menon and Meller 2012; Ugarkovic 2005; Usakin et al. 2007) are not transcriptionally inert. In fact, transcription of B-specific satDNAs in rye and jewel wasp B chromosomes is suggestive of their possible implication in interfering cell division in favor of the B chromosome (see above). However, for an excellent discussion on the possible functional role of B chromosome transcripts, see Benetta et al. (2019).
Highly interesting prospects for B chromosome research have resulted from recent results in mice, where Akera et al. (2017) found that oocyte spindle asymmetry depends on CDC42 signaling inducing microtubule tyrosination, and thus selfish meiotic drivers could exploit this asymmetry to bias their transmission. Likewise, Iwata-Otsubo et al. (2017) showed that “centromeres with more satellite repeats house more nucleosomes that confer centromere identity, containing the histone H3 variant CENP-A, and bias their segregation to the egg relative to centromeres with fewer repeats,” and suggested that “amplified repetitive sequences act as selfish elements by promoting expansion of CENP-A chromatin and increased transmission through the female germline.” Finally, Akera et al. (2019) showed that drive depends on slowing meiotic progression, and suggested that “selfish centromeres can be suppressed by regulating meiotic timing.” These findings suggest the possibility that satDNA accumulation on B chromosome centromere, along with the transcription of some protein-coding genes harbored by B chromosomes with putative functions to slow meiosis progression, could play a role in B chromosome drive. The possibility to focus B chromosome research on these molecular aspects is thus served, at least in some species.
References
Akera T, Chmátal L, Trimm E et al (2017) Spindle asymmetry drives non-Mendelian chromosome segregation. Science 358(6363):668–672. https://doi.org/10.1126/science.aan0092
Akera T, Trimm E, Lampson MA (2019) Molecular strategies of meiotic cheating by selfish centromeres. Cell 178(5):1132–1144. https://doi.org/10.1016/j.cell.2019.07.001
Alfenito MR, Birchler JA (1993) Molecular characterization of a maize B chromosome centric sequence. Genetics 135(2):589–597. https://www.genetics.org/content/135/2/589.long
Amos A, Dover G (1981) The distribution of repetitive DNAs between regular and supernumerary chromosomes in species of Glossina (tsetse): a two-step process in the origin of supernumeraries. Chromosoma 81(5):673–690. https://doi.org/10.1007/BF00329579
Banaei-Moghaddam AM, Schubert V, Kumke K et al (2012) Nondisjunction in favor of a chromosome: the mechanism of rye B chromosome drive during pollen mitosis. Plant Cell 24(10):4124–4134. https://doi.org/10.1105/tpc.112.105270
Banaei-Moghaddam AM, Meier K, Karimi-Ashtiyani R, Houben A (2013) Formation and expression of pseudogenes on the B chromosome of rye. Plant Cell 25(7):2536–2544. https://doi.org/10.1105/tpc.113.111856
Bauerly E, Hughes SE, Vietti DR et al (2014) Discovery of supernumerary B chromosomes in Drosophila melanogaster. Genetics 196(4):1007–1016. https://doi.org/10.1534/genetics.113.160556
Benetta DE, Akbari OS, Ferree PM (2019) Sequence expression of supernumerary B chromosomes: function or fluff? Genes 10(2):123. https://doi.org/10.3390/genes10020123
Benetta DE, Antoshechkin I, Yang T et al (2020) Genome elimination mediated by gene expression from a selfish chromosome. Sci Adv 6(14):eaaz9808. https://doi.org/10.1126/sciadv.aaz9808
Blunden R, Wilkes TJ, Forster JW et al (1993) Identification of the E3900 family, a second family of rye B chromosome specific repeated sequences. Genome 36(4):706–711. https://doi.org/10.1139/g93-095
Cabrero J, Bakkali M, Bugrov A et al (2003) Multiregional origin of B chromosomes in the grasshopper Eyprepocnemis plorans. Chromosoma 112(4):207–211. https://doi.org/10.1007/s00412-003-0264-2
Cabrero J, López-León MD, Ruíz-Estévez M et al (2014) B1 was the ancestor B chromosome variant in the western Mediterranean area in the grasshopper Eyprepocnemis plorans. Cytogenet Genome Res 142(1):54–58. https://doi.org/10.1159/000356052
Camacho JPM (2005) B chromosomes. In: Gregory TR (ed) The evolution of the genome. Elsevier, San Diego, pp 223–286. https://doi.org/10.1016/B978-012301463-4/50006-1
Carchilan M, Delgado M, Ribeiro T et al (2007) Transcriptionally active heterochromatin in rye B chromosomes. Plant Cell 19(6):1738–1749. https://doi.org/10.1105/tpc.106.046946
Chilton MD, McCarthy BJ (1973) DNA from maize with and without B chromosomes: a comparative study. Genetics 74(4):605–614. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1212976/
de Jesus CM, Galetti PM, Valentini SR, Moreira-Filho O (2003) Molecular characterization and chromosomal localization of two families of satellite DNA in Prochilodus lineatus (Pisces, Prochilodontidae), a species with B chromosomes. Genetica 118(1):25–32. https://doi.org/10.1023/A:1022986816648
Dhar M, Kour J, Kaul S (2019) Origin, behaviour, and transmission of B chromosome with special reference to Plantago lagopus. Genes 10(2):152. https://doi.org/10.3390/genes10020152
Dover GA (1975) The heterogeneity of B-chromosome DNA: no evidence for a B-chromosome specific repetitive DNA correlated with B-chromosome effects on meiotic pairing in the Triticinae. Chromosoma 53(2):153–173
Dover GA, Henderson SA (1976) No detectable satellite DNA in supernumerary chromosomes of the grasshopper Myrmeleotettix. Nature 259(5538):57–59. https://doi.org/10.1038/260170a0
Eickbush DG, Eickbush TH, Werren JH (1992) Molecular characterization of repetitive DNA sequences from a B chromosome. Chromosoma 101(9):575–583. https://doi.org/10.1007/bf00660317
Fox DP, Hewitt GM, Hall DJ (1974) DNA replication and RNA transcription of euchromatic and heterochromatic chromosome regions during grasshopper meiosis. Chromosoma 45(1):43–62. https://doi.org/10.1007/BF00283829
Fuster C, Rigola MA, Egozcue J (2004) Human supernumeraries: are they B chromosomes? Cytogenet Genome Res 106(2–4):165–172. https://doi.org/10.1159/000079283
Gibson I, Hewitt G (1970) Isolation of DNA from B chromosomes in grasshoppers. Nature 225(5227):67–68. https://doi.org/10.1038/225067a0
Gibson I, Hewitt G (1972) Interpopulation variation in the satellite DNA from grasshoppers with B-chromosomes. Chromosoma 38(2):121–138. https://doi.org/10.1007/BF00326190
Gómez-Aldecoa F (2021) Análisis estructural y funcional del cromosoma B de centeno. Doctoral dissertation, Universidad Complutense de Madrid
Gutknecht L, Sperlich D, Bachmann L (1995) A species specific satellite DNA family of Drosophila subsilvestris appearing predominantly in B chromosomes. Chromosoma 103(8):539–544. https://doi.org/10.1007/BF00355318
Hanlon S, Miller DE, Eche S, Hawley RS (2018) Origin, composition and structure of the supernumerary B chromosome of Drosophila. Genetics 210(4):1197–1212. https://doi.org/10.1534/genetics.118.300904
Houben A, Leach CR, Verlin D et al (1997) A repetitive DNA sequence common to the different B chromosomes of the genus Brachycome. Chromosoma 106(8):513–519. https://doi.org/10.1007/s004120050273
Huang W, Du Y, Zhao X, Jin W (2016) B chromosome contains active genes and impacts the transcription of A chromosomes in maize (Zea mays L.). BMC Plant Biol 16(1):88. https://doi.org/10.1186/s12870-016-0775-7
Iwata-Otsubo A, Dawicki-McKenna JM, Akera T et al (2017) Expanded satellite repeats amplify a discrete CENP-A nucleosome assembly site on chromosomes that drive in female meiosis. Curr Biol 27(15):2365–2373. https://doi.org/10.1016/j.cub.2017.06.069
John UP, Leach CR, Timmis JN (1991) A sequence specific to B chromosomes of Brachycome dichrosomatica. Genome 34:739–744. https://doi.org/10.1159/000444873
Kinsella CM, Ruiz-Ruano FJ, Dion-Côté AM et al (2019) Programmed DNA elimination of germline development genes in songbirds. Nat Commun 10:5468. https://doi.org/10.1038/s41467-019-13427-4
Kit S (1961) Equilibrium sedimentation in density gradients of DNA preparations from animal tissues. J Mol Biol 3(6):711–716. https://doi.org/10.1016/S0022-2836(61)80075-2
Klein AS, Eckhardt RA (1976) The DNAs of the a and B chromosomes of the mealy bug, Pseudococcus obscurus. Chromosoma 57(4):333–340
Klemme S, Banaei-Moghaddam AM, Macas J et al (2013) High-copy sequences reveal distinct evolution of the rye B chromosome. New Phytol 199(2):550–558. https://doi.org/10.1111/nph.12289
Kumke K, Macas J, Fuchs J et al (2016) Plantago lagopus B chromosome is enriched in 5S rDNA-derived satellite DNA. Cytogenet Genome Res 148(1):68–73. https://doi.org/10.1159/000444873
Langdon T, Seago C, Jones RN et al (2000) De novo evolution of satellite DNA on the rye B chromosome. Genetics 154(2):869–884. https://doi.org/10.1093/genetics/154.2.869
Li Y, Jing XA, Aldrich JC et al (2017) Unique sequence organization and small RNA expression of a “selfish” B chromosome. Chromosoma 126(6):753–768. https://doi.org/10.1007/s00412-017-0641-x
López-León M, Neves N, Schwarzacher T et al (1994) Possible origin of a B chromosome deduced from its DNA composition using double FISH technique. Chromosom Res 2(2):87–92. https://doi.org/10.1007/BF01553487
López-León MD, Vázquez P, Hewitt G, Camacho JPM (1995) Cloning and sequence analysis of an extremely homogeneous tandemly repeated DNA in the grasshopper Eyprepocnemis plorans. Heredity 75(4):370–375. https://doi.org/10.1038/hdy.1995.148
López-León MD, Cabrero J, Dzyubenko VV et al (2008) Differences in ribosomal DNA distribution on A and B chromosomes between eastern and western populations of the grasshopper Eyprepocnemis plorans plorans. Cytogenet Genome Res 121(3–4):260–265. https://doi.org/10.1159/000138894
Ma W, Gabriel TS, Martis MM et al (2016) Rye B chromosomes encode a functional Argonaute-like protein with in vitro slicer activities similar to its A chromosome paralog. New Phytol 213(2):916–928. https://doi.org/10.1111/nph.14110
Makunin AI, Rajičić M, Karamysheva TV et al (2018) Low-pass single-chromosome sequencing of human small supernumerary marker chromosomes (sSMCs) and Apodemus B chromosomes. Chromosoma 127(3):301–311. https://doi.org/10.1007/s00412-018-0662-0
Mardis ER (2008) Next-generation DNA sequencing methods. Annu Rev Genomics Hum Genet 9:387–402. https://doi.org/10.1146/annurev.genom.9.081307.164359ç
Martis MM, Klemme S, Banaei-Moghaddam AM et al (2012) Selfish supernumerary chromosome reveals its origin as a mosaic of host genome and organellar sequences. Proc Natl Acad Sci USA 109(33):13343–13346. https://doi.org/10.1073/pnas.1204237109
McAllister BF, Werren JH (1997) Hybrid origin of a B chromosome (PSR) in the parasitic wasp Nasonia vitripennis. Chromosoma 106(4):243–253. https://doi.org/10.1007/s004120050245
Menon DU, Meller VH (2012) A role for siRNA in X-chromosome dosage compensation in Drosophila melanogaster. Genetics 191(3):1023–1028
Milani D, Cabral-de-Mello DC (2014) Microsatellite organization in the grasshopper Abracris flavolineata (Orthoptera: Acrididae) revealed by FISH mapping: remarkable spreading in the A and B chromosomes. PLoS One 9(5):e97956. https://doi.org/10.1371/journal.pone.0097956
Milani D, Ramos É, Loreto V et al (2017) The satellite DNA AflaSAT-1 in the A and B chromosomes of the grasshopper Abracris flavolineata. BMC Genet 18:81. https://doi.org/10.1186/s12863-017-0548-9
Milani D, Bardella V, Ferretti A et al (2018) Satellite DNAs unveil clues about the ancestry and composition of B chromosomes in three grasshopper species. Genes 9(11):523. https://doi.org/10.3390/genes9110523
Muñoz-Pajares AJ, Martínez-Rodríguez L, Teruel M et al (2011) A single, recent origin of the accessory B chromosome of the grasshopper Eyprepocnemis plorans. Genetics 187(3):853–863. https://doi.org/10.1534/genetics.110.122713
Navarro-Domínguez B, Ruiz-Ruano FJ, Cabrero J et al (2017) Protein-coding genes in B chromosomes of the grasshopper Eyprepocnemis plorans. Sci Rep 7:45200. https://doi.org/10.1038/srep45200
Novák P, Neumann P, Pech J et al (2013) RepeatExplorer: a galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 29(6):792–793. https://doi.org/10.1093/bioinformatics/btt054
Nur U, Werren JH, Eickbush DG et al (1988) A “selfish” B chromosome that enhances its transmission by eliminating the paternal genome. Science 240(4851):512–514. https://doi.org/10.1126/science.3358129
Pansonato-Alves JC, Serrano ÉA, Utsunomia R et al (2014) Single origin of sex chromosomes and multiple origins of B chromosomes in fish genus Characidium. PLoS One 9(9):e107169. https://doi.org/10.1371/journal.pone.0107169
Peacock WJ, Dennis ES, Rhoades M, Pryor AJ (1981) Highly repeated DNA sequence limited to knob heterochromatin in maize. Proc Natl Acad Sci USA 78(7):4490–4494. https://doi.org/10.1073/pnas.78.7.4490
Ruban A, Schmutzer T, Scholz U, Houben A (2017) How next-generation sequencing has aided our understanding of the sequence composition and origin of B chromosomes. Genes 8(11):294. https://doi.org/10.3390/genes8110294
Ruban A, Schmutzer T, Wu DD et al (2020) Supernumerary B chromosomes of Aegilops speltoides undergo precise elimination in roots early in embryo development. Nat Commun 11:2764. https://doi.org/10.1038/s41467-020-16594-x
Ruiz-Ruano FJ, López-León MD, Cabrero J, Camacho JPM (2016) High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci Rep 6:28333. https://doi.org/10.1038/srep28333
Ruiz-Ruano FJ, Cabrero J, López-León MD, Camacho JPM (2017) Satellite DNA content illuminates the ancestry of a supernumerary (B) chromosome. Chromosoma 26(4):487–500. https://doi.org/10.1007/s00412-016-0611-8
Ruiz-Ruano FJ, Cabrero J, López-León MD et al (2018) Quantitative sequence characterization for repetitive DNA content in the supernumerary chromosome of the migratory locust. Chromosoma 127(1):45–57. https://doi.org/10.1007/s00412-017-0644-7
Ruiz-Ruano FJ, Navarro-Domínguez B, López-León MD et al (2019) Evolutionary success of a parasitic B chromosome rests on gene content. bioRxiv 683417. https://doi.org/10.1101/683417
Sandery MJ, Forster JW, Blunden R, Jones RN (1990) Identification of a family of repeated sequences on the rye B chromosome. Genome 33(1985):908–913. https://doi.org/10.1139/g90-137
Serrano-Freitas ÉA, Silva DMZA, Ruiz-Ruano FJ et al (2020) Satellite DNA content of B chromosomes in the characid fish Characidium gomesi supports their origin from sex chromosomes. Mol Genet Genomics 295(1):195–207. https://doi.org/10.1007/s00438-019-01615-2
Silva DMZA, Daniel SN, Camacho JPM et al (2016) Origin of B chromosomes in the genus Astyanax (Characiformes, Characidae) and the limits of chromosome painting. Mol Genet Genomics 291(3):1407–1418. https://doi.org/10.1007/s00438-016-1195-y
Silva DMZA, Utsunomia R, Ruiz-Ruano FJ et al (2017) High-throughput analysis unveils a highly shared satellite DNA library among three species of fish genus Astyanax. Sci Rep 7:12726. https://doi.org/10.1038/s41598-017-12939-7
Silva DMZA, Ruiz-Ruano FJ, Utsunomia R et al (2021) Long-term persistence of supernumerary B chromosomes in multiple species of Astyanax fish. BMC Biol 19:52. https://doi.org/10.1186/s12915-021-00991-9
Teruel M, Cabrero J, Perfectti F, Camacho JPM (2010) B chromosome ancestry revealed by histone genes in the migratory locust. Chromosoma 119(2):217–225. https://doi.org/10.1007/s00412-009-0251-3
Teruel M, Ruiz-Ruano FJ, Marchal JA et al (2014) Disparate molecular evolution of two types of repetitive DNAs in the genome of the grasshopper Eyprepocnemis plorans. Heredity 112(5):531–542. https://doi.org/10.1038/hdy.2013.135
Timmis JN, Ingle J, Sinclair J, Jones N (1975) The genomic quality of Rye B chromosomes. J Exp Bot 26(3):367–378. https://doi.org/10.1093/jxb/26.3.367
Ugarkovic D (2005) Functional elements residing within satellite DNAs. EMBO Rep 6:1035–1039. https://doi.org/10.1038/sj.embor.7400558
Usakin L, Abad J, Vagin VV et al (2007) Transcription of the 1.688 satellite DNA family is under the control of RNA interference machinery in Drosophila melanogaster ovaries. Genetics 176:1343–1349. https://doi.org/10.1534/genetics.107.071720
Utsunomia R, Silva DMZA, Ruiz-Ruano FJ et al (2016) Uncovering the ancestry of B chromosomes in Moenkhausia sanctaefilomenae (Teleostei, Characidae). PLoS One 11(3):e0150573. https://doi.org/10.1371/journal.pone.0150573
Valente GT, Conte MA, Fantinatti BEA et al (2014) Origin and evolution of B chromosomes in the cichlid fish Astatotilapia latifasciata based on integrated genomic analyses. Mol Biol Evol 31(8):2061–2072. https://doi.org/10.1093/molbev/msu148
Wang X, Fang X, Yang P et al (2014) The locust genome provides insight into swarm formation and long-distance flight. Nat Commun 5:2957. https://doi.org/10.1038/ncomms3957
Waring M, Britten RJ (1966) Nucleotide sequence repetition: a rapidly reassociating fraction of mouse DNA. Science 154(3750):791–794. https://doi.org/10.1126/science.154.3750.791
Wu DD, Ruban A, Fuchs J et al (2019) Nondisjunction and unequal spindle organization accompany the drive of Aegilops speltoides B chromosomes. New Phytol 223(3):1340–1352. https://doi.org/10.1111/nph.15875
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Camacho, J.P.M., Ruiz-Ruano, F.J., López-León, M.D., Cabrero, J. (2021). Satellite DNA Is an Inseparable Fellow Traveler of B Chromosomes. In: Ugarković, Ð. (eds) Satellite DNAs in Physiology and Evolution. Progress in Molecular and Subcellular Biology, vol 60. Springer, Cham. https://doi.org/10.1007/978-3-030-74889-0_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-74889-0_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-74888-3
Online ISBN: 978-3-030-74889-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)