Abstract
Nitrogen use efficiency (NUE) is an extremely important and challenging area of crop improvement to prevent economic wastage of fertilizers and also to minimize their adverse environmental consequences. Signaling proteins that mediate N response by regulating protein phosphorylation are emerging as useful targets in this regard. This chapter reviews the current understanding of the regulatory role of protein phosphatases in N response, NUE, and the possibilities ahead. Several protein kinases and phosphatases have been found to be involved in all aspects of N response, including uptake, metabolism, sensing, and signaling. Protein phosphatase 2C (PP2C) is involved in the regulation of nitrate transporter NRT1.1 and in the regulation of abscisic acid (ABA) signaling in nitrate sensing. Phosphatases also play regulatory role in photosynthesis and C-metabolism and may be involved in balancing carbon nitrogen cycle. Nitrate metabolism is regulated by phosphatase through target of rapamycin (TOR), and the fine-tuning of nitrate reductase activity through phosphatases is well known. In addition to these, it also explores the phosphatases identified for N response and NUE in different crops. Over all, this chapter provides a foundation in understanding the regulatory role of phosphatases in N response/NUE.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- NUE
- Nitrate sensing
- N response
- Nitrate reductase
- Nitrate transporters
- Phosphatases
- Target of rapamycin
- PP2C
12.1 Introduction
Nitrogen is an essential macronutrient for plant growth and crop yield, as most crop plants barring N-fixing legumes depend on reactive N compounds such as urea, ammonium salts, or nitrates from manures or fertilizers as their main nitrogen source. Fertilizers are the world’s largest anthropogenic source of reactive nitrogen, with annual global consumption at approximately 119.40 million tons and annual growth of 1.4% (FAO 2018). Due to the poor N use efficiency (NUE) of most crops at a global average of <30%, unused reactive N compounds from fertilizers and other sources are a major source of pollution of land, water, and air, affecting human and animal health, biodiversity, and climate change (Sutton and Bleeker 2013; Sutton et al. 2019).The fourth UN Environment Assembly held in March 2019 has adopted a resolution to work toward sustainable nitrogen management (https://sdg.iisd.org/events/fourth-session-of-the-un-environment-assembly-unea-4/). Therefore, the development of N use efficient crops is an extremely important goal, not only for sustainable agriculture but also for environmental sustainability and mitigation of climate change (Raghuram et al. 2007; Raghuram and Sharma 2019).
NUE can be defined in several ways (Raghuram and Sharma 2019), but it is best understood as yield per unit N, or the maximal output with minimal N input. NUE is a complex genetic trait involving several genes and their interactions with the environment, which are yet to be fully characterized (Mandal et al. 2018). NUE involves a combination of processes such as N uptake, retention, assimilation, and remobilization of internal N reserves. Plants possess families of transporters to take up urea, ammonium, and nitrate ions (NO3 −), of which NO3− ions not only acts as nutrient but also play a very important role in signal transduction and regulation of one or more of the above processes (Raghuram et al. 2006; Pathak et al. 2008, 2011; Chakraborty and Raghuram 2011; Mandal et al. 2018; Raghuram and Sharma 2019).
The progress on the precise characterization of the biological avenues for improvement for NUE has been painfully slow, despite some occasional advances on phenotype (Sun et al. 2014; Sharma et al. 2018). Indeed, even though low nutrient stress is known to be an important abiotic stress for crops in developing countries, the interface of N and stress has not been explored in great detail (Jangam et al. 2016). While this has hampered crop improvement by forward genetics, recent advances in genomics and functional genomics of several crops have opened the opportunity for using reverse genetics to address NUE using candidate gene approach. To understand the physiological and molecular basis of N response under normal and low supply of nitrogen, transcriptomic and proteomic studies have identified a large number of differentially expressed genes in crops like rice (Yang et al. 2015;Waqas et al. 2018; Sinha et al. 2018), sorghum (Gelli et al. 2014), maize (Jiang et al. 2018), and barley (Quan et al. 2016, 2019).
Among the multitude of pathways and processes that constitute N response, protein kinases and phosphatases as well as transcription factors and other signaling intermediates regulated by phosphorylation emerged significant. Protein kinases as well as protein phosphatases have been widely studied in rice, Arabidopsis, wheat, maize, etc. (Singh et al. 2010; Xue et al. 2008; Bradai et al. 2018; Trevisan et al. 2011). PP2Cs are a major class of phosphatases and play an important role in stress in plants (Moorhead et al. 2009; Singh et al. 2010, 2015). In this chapter, we discuss the role of protein phosphatases in N-sensing and signaling and their role in carbon and nitrogen metabolism and phosphatases involved in N-response and NUE with enhancement potential for further validation of phosphatases.
12.2 Phosphatases in N Uptake and Primary Nitrate Response
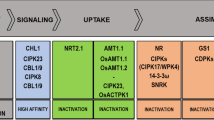
For optimal growth as well as ion balance and homeostasis of plant, adjustment of uptake of ammonium (NH4+) and NO3− is essential as compared to other major ions (Haynes 1990). Phosphorylation and dephosphorylation play major roles in regulating transport of these ions by transporter protein through plasma membrane (Straub et al. 2017). An important target of such regulation is NRT1.1/CHL1/NPF6.3, a dual-affinity nitrate transporter. It works under high as well as low NO3− condition. PP2C plays a major role in dephosphorylation of NRT1.1 by binding to the calcineurin B-like-interacting protein kinase (CIPK). CIPK consists of two domains; one is the conserved N-terminal kinase catalytic domain, and the other is the highly variant C-terminal regulatory domain having NAF/FISL motif and a phosphatase interaction motif (Guo et al. 2001). In response to low NO3 − condition, NO3− binds to a high affinity site, which activates CIPK23 to phosphorylate CHL1 at T101. The phosphorylated CHL1 prevents higher primary nitrate response to low NO3− concentration. At high NO3− concentrations, NO3− binds to a low nitrate affinity site, which dephosphorylates CHL1 at T101 and generates a high primary nitrate response (Ho et al. 2009; Vert and Chory 2009; Straub et al. 2017). In low-affinity phase of (NO3−), CIPK8 is activated by an unknown CBL, which phosphorylates CHL1 at a residue different from T101 (Hu et al. 2009). CIPK23 acts as a negative regulator of high-affinity NO3− response, while it acts as a positive regulator of primary nitrate response in low-affinity phase of NO3− response.
12.3 PP2Cs Are Negative Regulators of ABA Signaling in NO3− Sensing
ABA signaling pathway includes a type 2C protein phosphatase (PP2C; a negative regulator) and a SNF1-related protein kinase 2 (SnRK2; a positive regulator) (Umezawa et al. 2009). ABA can inhibit or stimulate the functioning of the root meristem and modulate root growth depending on its concentration (Cheng et al. 2002). Development of root architecture is strongly regulated by the concentration of NO3− in the root environment, as well as its distribution. If the environment surrounding the root system is uniformly high in NO3−, lateral root growth is generally inhibited (Zhang and Forde 2000; Walch-Liu et al. 2005). NO3− locally stimulates lateral root elongation and in some species initiation, when NO3− presents only in patch (Zhang and Forde 1998). In Arabidopsis, this local stimulation of lateral root elongation by patches of NO3− requires ABA signaling (Signora et al. 2001). In Arabidopsis, PP2C proteins such as ABA-insensitive 1 (ABI1), ABI2, and Hypersensitive to ABA 1 (HAB1) have been found to function in regulation of root development. ABA-induced signaling functions under both biotic and abiotic stresses by interacting with SnRK2s and PYR/PYL/RCARs.
12.4 Phosphatases: Key Players in Carbon and Nitrogen Balance
In addition to the independent utilization of carbon and nitrogen metabolites, their ratio (known as C/N balance) is more important for the regulation of plant growth. The signaling mechanism underlying C/N balance is not clear till date (Sulpice et al. 2013; Lu et al. 2015). In leaves, NO3− and nitrite are reduced to ammonia and then to glutamate (Glu) via the glutamine synthetase (GS)-glutamine-2-oxoglutarate aminotransferase (GOGAT) pathway. Glutamate is a source of C and N for the biosynthesis of most other amino acids (Forde and Lea 2007). GOGAT is found in two isoforms (FdGOGAT form and NADH-GOGAT) and located in the chloroplast in higher plants. Among these, Fd-GOGAT was found to be very active in chloroplast of photosynthetic tissues (Nigro et al. 2014). It plays a very important role in re-assimilation of ammonia released during photorespiration and is potential target to improve NUE (Zeng et al. 2017). Phosphatases and kinases both play very important role in posttranslational modification of chloroplast protein in regulating distribution of light energy between the photosystem I and photosystem II (Michelet et al. 2013; Rochaix 2013; Grabsztunowicz et al. 2017). The PSII core proteins D1 and D2, inner antenna protein CP43, and a minor PSII subunit PsbH are targets for light-dependent Thr phosphorylation catalyzed mainly by the STN8 kinase (Fristedt and Vener 2011), while the PSII core phosphatase is responsible for the dephosphorylation (Samol et al. 2012). Dephosphorylation of light harvesting complex (LHC) by the PPH1/TAP38 (chloroplast protein phosphatase/thylakoid-associated phosphatase of 38 kDa) protein phosphatase results in redistribution of excitation energy toward PSII (Pribil et al. 2010; Shapiguzov et al. 2010).
Ribulose-1,5-bisphosphate carboxylase-oxygenase (Rubisco) plays a central role in CO2 assimilation of all photosynthetic organisms. It catalyzes the photosynthetic CO2 fixation by the carboxylation of ribulose-1,5-bisphosphate (RuBP) in C3 photosynthesis. Rubisco is a slow enzyme, and its large amount (up to 25% of leaf N and 50% of leaf-soluble protein) is needed for photosynthesis (Parry et al. 2013; Whitney et al. 2015). In a study, it has been shown that in antisense plants by decreasing Rubisco content up to 15–20%, nitrogen demand reduced by 10% without negatively affecting photosynthetic carbon fixation (Sitt and Schulze 1994; Parry et al. 2013). Thus, increasing CO2 fixing capacity by changing Rubisco and/or Rubisco activase function, i.e., more carbon fixed for same amount of leaf protein, would increase nitrogen use efficiency. Rubisco has been reported as a target of reversible phosphorylation in many plant species (Wang et al. 2014; Roitinger et al. 2015). Two subunits of Rubisco, RBCL (Rubisco large chain gene) and RBCS (Rubisco small chain gene), contain multiple phosphorylation sites (Cao et al. 2011; Wang et al. 2014). Phosphorylation of the highly conserved RBCL residues Ser208, Thr246, Tyr239, and Thr330, located in the close proximity to RuBP binding site, might affect the catalytic activity of the enzyme (Hodges et al. 2013). Dephosphorylation of RBCL has been found to dissociate Rubisco holocomplex and decreased activity of the enzyme by affecting the interaction between Rubisco and Rubisco activase (RA) (Chen et al. 2011; Hodges et al. 2013).
12.5 PP2A-TOR in Regulation of Nitrate Metabolism
Target of rapamycin (TOR) is a serine/threonine kinase and is highly conserved among all eukaryotes. TOR is activated by both nitrogen and carbon metabolites and promotes cell division, mRNA translation, and repressing nutrient remobilization through autophagy (Dobrenel et al. 2016). It is a master regulator of growth and development including transcription, ribosome biogenesis, autophagy, nutrition recycling, and cellular metabolism in plants (Bakshi et al. 2019). The plant TOR complex consists of TOR, Raptor, and LST8. The TOR complex promotes cell growth and translation through the regulation of S6K and its substrate ribosomal protein S6(RPS6). It has been reported that TAP46 and its associated phosphatases (PP2Ac and PP2Ac-like subunits) play a critical role in mediating TOR signaling, leading to the promotion of protein translation, the repression of autophagy, and nitrate metabolism (Fig. 12.1). TAP42-phosphatase complex associates with TOR complex 1 (TORC1), under normal condition, while in nutrient deprivation condition or rapamycin treatment, TAP42-phosphatase complex dissociates and is released to cytosol. Interaction of TAP42 with PP2A or PP2A-like phosphatases modulates the activity of downstream effectors and regulates transcription and translation of starvation-responsive genes (Ahn et al. 2011). In Arabidopsis transgenic plants, overexpression of TAP46 leads to upregulation of genes related to nitrogen metabolism and nitrate assimilation and also increased TOR activity, whereas TAP46-silenced plants exhibited a reduction in N-assimilating enzymes (Ahn et al. 2015).
Schematic representation of TOR signaling in plants in response to nutrients (Modified from Ahn et al. 2011)
12.6 Protein Phosphatases: Fine-Tuning of Nitrate Reductase
Phosphatases are not only passive partners of kinases, rather they play a dynamic regulatory role in several metabolic processes. They activate or deactivate themselves through posttranslational modification (Heidari et al. 2011). Posttranslational regulation of nitrate reductase (NR) activity through phosphorylation/dephosphorylation by PP2A (protein phosphatase 2A), CDPK (calcium-dependent protein kinase), and SNRK (SNF/sucrose nonfermenting-related kinase) is light dependent (Raghuram and Sopory 1995; Huber et al. 1996). Activation of NR is triggered by photosynthesis, but the signaling cascades from chloroplasts to cytosol, where NR is located, are not clear. Okadaic acid and microcystin-LR are known inhibitors of the protein phosphatase families including protein phosphatase 1 (PP1), PP2A, PP4, PP5, PP6, and PPP-Kelch, and they prevent light activation of NR in plant system (Lillo et al. 1996; Ali et al. 2007). PP2C also plays a major regulatory role in different stress signalings in plants (Moorhead et al. 2007; Singh et al. 2015). Dephosphorylation of NR is essential for activation of NR.14-3-3 which is highly conserved family of proteins found in all eukaryotes. Depending on the developmental and environmental requirements, 14-3-3 activity could direct carbon either into sucrose and storage carbohydrate synthesis or, via inactivation of carbon metabolism and activation of nitrogen assimilation, divert carbon skeletons into the synthesis of amino acids. Plants could use the various possible combinations of 14-3-3 homo- and heterodimers for fine-tuning (Comparot et al. 2003). In a study in rice, dephosphorylation of SnRK and 14-3-3 by PP2C led to the activation of nitrate reductase for increasing NUE and assimilating efficiency of plants (Waqas et al. 2018). In extract of spinach leaf, okadaic acid and microcystin were found to inactivate NR activity, but inhibitor 2 (known inhibitor of PP1 family) showed no effect on NR. Mammalian PP2A activates NR in vitro, and it further suggested the role of other phosphatases including PP2A in this process (Mackintosh 1992). BSL1, BSL2, and BSL3 are expressed in leaf and can be considered as potential candidate genes for dephosphorylating NR (Mora-García et al. 2004). Heidari et al. (2011) showed that by knocking down of all three subunits of PP2A in Arabidopsis, NR activity was negatively regulated. This provides a confirmation of involvement of PP2A in NR activation in light-dark transition in plants.
12.7 Phosphatases Identified in N Response/NUE
Global gene expression and comparative analysis of genotypes contrasting for NUE allow the enrichment of candidate genes for NUE. Considerable literatures are now available on transcriptomics, proteomics, as well as on quantitative trait loci (QTL) affecting N response and NUE (Sinha et al. 2018). However, only limited studies explored the regulatory role of different classes of phosphatases in N responses and NUE (Waqas et al. 2018; Sinha et al. 2018; Xiong et al. 2019).
A compilation of the different phosphatases reported to be involved in plant N response and/or NUE is provided in Table 12.1. In a study conducted by Sinha et al. (2018), two contrasting rice genotypes IR64 and Nagina22 were used for genome-wide transcriptome analysis under optimal and chronic starvation of nitrogen from 15-day-old root and shoot tissues. Two genes, LOC_Os01g71420 and LOC_Os10g25430 belonging to Ser/Thr phosphatase family, are reported as N-responsive genes.In transgenic line, over expression of PP2C9TL has been reported to improve NUE due to higher activity of NR by downregulation of SnRK and 14-3-3 (Waqas et al. 2018). The higher activity of NR leads to more production of nitric oxide, which then increases N uptake by enhancing the lateral root development. Besides NR, phosphoenolpyruvate carboxylase (PEPC) was also found to be downregulated due to overexpression of PP2C, which dephosphorylates PEPC. Upregulation of PSII and Rubisco increases photosynthetic rate under low N (Waqas et al. 2018). PP2C68 and lipid phosphatase 2 have been reported as upregulated N-responsive phosphatases, while PP2C39 and PP2C46 have been reported to be downregulated in root under N starvation condition (Hsieh et al. 2018). Purple acid phosphatase 1 and serine threonine phosphatase have been found to be nitrate-responsive gene under low nitrate condition in maize (Jiang et al. 2018). Two nitrogen-responsive genes (trehalose-6-phosphate, T6P) in barley have been found to be downregulated under low N condition (Fataftah et al. 2018). These genes along with other genes reverse reaction from pyruvate back to phosphoenolpyruvate during N starvation as an efficient pathway for the remobilization of N sources. In wheat, Traes_6BS_143FEF476 encoding bifunctional protein-serine/threonine kinase/phosphatase was found to be a candidate gene providing tolerance to low N (Xiong et al. 2019).
12.8 Conclusions and Future Prospects
Understanding the genetic basis of tolerance to low N in crops is important for the development of NUE improvement strategies. Protein phosphatases are emerging as an important gene family among the candidate genes/families identified from different plants, as they regulate N uptake, assimilation, and remobilization. While functional genomic approaches have provided several leads in this regard, their potential in the improvement of NUE needs further validation through mutant/transgenic lines, before using them as targets in crop breeding programs for NUE. In the meantime, integration of all the available information at the molecular, physiological, and genetic level will be useful to mine SNPs through genome-wide association studies (GWAS) and marker development using candidate gene approach. We can expect to see some developments in this direction in the years ahead.
References
Ahn CS, Ahn HK, Pai HS (2015) Overexpression of the PP2A regulatory subunit Tap46 leads to enhanced plant growth through stimulation of the TOR signaling pathway. J Exp Bot 66:827–840
Ahn CS, Han JA, Lee HS, Lee S, Pai HS (2011) The PP2A regulatory subunit Tap46, a component of the TOR signaling pathway, modulates growth and metabolism in plants. Plant Cell 23:185–209
Ali A, Sivakami S, Raghuram N (2007) Regulation of activity and transcript levels of NR in rice (Oryza sativa): roles of protein kinase and G-proteins. Plant Sci 172:406–413
Bakshi A, Moin M, Madhav MS, Kirti PB (2019) Target of rapamycin, a master regulator of multiple signaling pathways and a potential candidate gene for crop improvement. Plant Biol 21:190–205
Bradai M, Mahjoubi H, Chini A, Chaboute M-E, Hanin M, Ebel C (2018) Genome wide identification of wheat and Brachypodium type one protein phosphatases and functional characterization of durum wheat TdPP1a. PLoS One 13:e0191272
Cao X, Gao Y, Wang Y, Li CM, Zhao YB, Han ZH, Zhang XZ (2011) Differential expression and modification of proteins during ontogenesis in Malus domestica. Proteomics 11(24):4688–4701
Chakraborty N, Raghuram N (2011) Nitrate sensing and signaling in genomewide plant N response. In: Jain V, Anandakumar P (eds) Nitrogen use efficiency in plants. New India Publishing Agency, New Delhi, pp 45–62
Chen X, Zhang W, Zhang B, Zhou J, Wang Y, Yang Q, Ke Y, He H (2011) Phosphoproteins regulated by heat stress in rice leaves. Proteome Sci 9:37
Cheng WH, Endo A, Zhou L, Penney J, Chen HC, Arroyo A, Leon P, Nambara E, Asami T, Seo M, Koshiba T, Sheen J (2002) A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell 14:2723–2743
Comparot S, Lingiah G, Martin T (2003) Function and specificity of 14-3-3 proteins in the regulation of carbohydrate and nitrogen metabolism. J Exp Bot 54:595–604
Dobrenel T, Caldana C, Hanson J, Robaglia C, Vincentz M, Veit B, Meyer C (2016) TOR signaling and nutrient sensing. Annu Rev Plant Biol 67:261–285
FAO (2018) World Fertilizer Trends and Outlook to 2018. FAO, Rome, p 66
Fataftah N, Mohr C, Hajirezaei MR, Wirén NV, Humbeck K (2018) Changes in nitrogen availability lead to a reprogramming of pyruvate metabolism. BMC Plant Biol 18:77
Forde BG, Lea PJ (2007) Glutamate in plants: metabolism, regulation, and signaling. J Exp Bot 58:2339–2358
Fristedt R, Vener AV (2011) High light-induced disassembly of photosystem II super complexes in Arabidopsis requires STN7-dependent phosphorylation of CP29. PLoS One 6(9):e24565
Gelli M, Duo Y, Konda AR, Zhang C, Holding D, Dweikat I (2014) Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genom 15(1):179
Grabsztunowicz M, Koskela MM, Mulo P (2017) Post-translational modifications in regulation of chloroplast function: recent advances. Front Plant Sci 8:240
Guo Y, Halfter U, Ishitani M, Zhu JK (2001) Molecular characterization of functional domains in the protein kinase SOS2 that is required for plant salt tolerance. Plant Cell 13:1383–1400
Haynes RJ (1990) Active ion uptake and maintenance of cation-anion balance: a critical examination of their role in regulating rhizosphere pH. Plant Soil 126:247–264
Heidari B, Matre P, Nemie-Feyissa D, Meyer C, Rognli OA, Moller SG, Lillo C (2011) Protein phosphatase 2A B55 and a regulatory subunits interact with nitrate reductase and are essential for nitrate reductase activation. Plant Physiol 156:165–172
Ho CH, Lin SH, Hu HC, Tsay YF (2009) CHL1 functions as a nitrate sensor in plants. Cell 138:1184–1194
Hodges M, Jossier M, Boex-Fontvieille E, Tcherkez G (2013) Protein phosphorylation and photorespiration. Plant Biol 15:694–706
Hsieh PH, Kan CC, Wu HY, Yang HC, Hsieh MH (2018) Early molecular events associated with nitrogen deficiency in rice seedling roots. Sci Rep 8:12207
Hu HC, Wang YY, Tsay YF (2009) AtCIPK8, a CBL-interacting protein kinase, regulates the low-affinity phase of the primary nitrate response. Plant J 57:264–278
Huber SC, Bachmann M, Huber JL (1996) Post-translation regulation of nitrate reductase activity: a role for Ca2+ and 14-3-3 proteins. Trends Plant Sci 1:432–438
Jangam AP, Pathak RR, Raghuram N (2016) Microarray analysis of Rice d1 (RGA1) mutant reveals the potential role of G-protein alpha subunit in regulating multiple abiotic stresses such as drought, salinity, heat, and old. Front Plant Sci 7:11
Jiang L, Ball G, Hodgman C, Coules A, Zhao H, Lu C (2018) Analysis of gene regulatory networks of maize in response to nitrogen. Genes 9:1–20
Leran S, Edel KH, Pervent M, Hashimoto K, Corratge-Faillie C, Offenborn JN, Tillard P, Gojon A, Kudla J, Lacombe B (2015) Nitrate sensing and uptake in Arabidopsis are enhanced by ABI2, a phosphatase inactivated by the stress hormone abscisic acid. Sci Signal 8:ra43
Lillo C, Smith LH, Nimmo HG, Wilkins MB (1996) Regulation of nitrate reductase and phosphoenolpyruvate carboxylase activities in barley leaf protoplasts. Planta 200:181–185
Liu R, Zhang H, Zhao P, Zhang Z, Liang W, Tian Z, Zheng Y (2011) Mining of candidate maize genes for nitrogen use efficiency by integrating gene expression and QTL data. Plant Mol Biol Rep 30:297–308
Lu Y, Sasaki Y, Li X, Mori IC, Matsuura T, Hirayama T, Sato T, Yamaguchi J (2015) ABI1 regulates carbon/nitrogen-nutrient signal transduction independent of ABA biosynthesis and canonical ABA signaling pathways in Arabidopsis. J Exp Bot 66:2763–2771
Mackintosh C (1992) Regulation of spinach-leaf nitrate reductase by reversible phosphorylation. Biochim Biophys Acta 1137:121–126
Mandal VK, Sharma N, Raghuram N (2018) Molecular targets for improvement of crop nitrogen use efficiency: current and emerging options. In: Shrawat A, Zayed A, Lightfoot DA (eds) Engineering nitrogen utilization in crop plants. Elsevier, New York, NY
Michelet L, Zaffagnini M, Morisse S, Sparla F, Pérez-Pérez ME, Francia F, Danon A, Marchand CH, Fermani S, Trost P, Lemaire SD (2013) Redox regulation of the calvin-benson cycle: something old, something new. Front Plant Sci 4:470
Moorhead GB, Trinkle-Mulcahy L, Ulke-Lemee A (2007) Emerging roles of nuclear protein phosphatases. Nat Rev Mol Cell Biol 8:234–244
Moorhead GBG, De Wever V, Templeton G, Kerk D (2009) Evolution of protein phosphatases in plants and animals. Biochem J 417:401–409
Mora-García S, Vert G, Yin Y, Caño-Delgado A, Cheong H, Chory J (2004) Nuclear protein phosphatases with Kelch-repeat domains modulate the response to brassinosteroids in Arabidopsis. Genes Dev 18:448–460
Nigro D, Blanco A, Anderson OD, Gadaleta A (2014) Characterization of ferredoxin-dependent glutamine-oxoglutarate amidotransferase (Fd-GOGAT) genes and their relationship with grain protein content QTL in wheat. PLoS One 9:e103869
Parry MAJ, Andralojc PJ, Scales JC, Salvucci ME, Carmo-Silva AE, Alonso H, Whitney SM (2013) Rubisco activity and regulation as targets for crop improvement. J Exp Bot 64:717–730
Pathak RR, Ahmad A, Lochab S, Raghuram N (2008) Molecular physiology of plant N-use efficiency and biotechnological options for its enhancement. Curr Sci 94(11):1394–1403
Pathak RR, Lochab S, Raghuram N (2011) Improving nitrogen use efficiency. In: Comprehensive biotechnology, vol 4, 2nd edn. Elsevier, Oxford, pp 209–218
Plavcova L, Hacke UG, Almeida-Rodriguez AM, Li E, Douglas CJ (2013) Gene expression patterns underlying changes in xylem structure and function in response to increased nitrogen availability in hybrid poplar. Plant Cell Environ 36:186–199
Pribil M, Pesaresi P, Hertle A, Barbato R, Leister D (2010) Role of plastid protein phosphatase TAP38 in LHCII dephosphorylation and thylakoid electron flow. PLoS Biol 8:e100028810
Quan X, Zeng J, Chen G, Zhang G (2019) Transcriptomic analysis reveals adaptive strategies to chronic low nitrogen in Tibetan wild barley. BMC Plant Biol 19:68
Quan XY, Zeng JB, Ye LZ, Chen G, Han ZG, Shah JM, Zhang G (2016) Transcriptome profiling analysis for two Tibetan wild barley genotypes in responses to low nitrogen. BMC Plant Biol 16:30
Raghuram N, Pathak RR, Sharma P (2006) Signaling and the molecular aspects of N-use efficiency in higher plants. In: Singh RP, Jaiwal PK (eds) Biotechnological approaches to improve nitrogen use efficiency in plants. Studium Press LLC, Houston, TX, pp 19–40
Raghuram N, Sachdev MS, Abrol YP (2007) Towards an integrative understanding of reactive nitrogen. In: Abrol YP, Raghuram N, Sachdev MS (eds) Agricultural nitrogen use and its environmental implications. IK International, Delhi, pp 1–6
Raghuram N, Sharma N (2019) Improving crop nitrogen use efficiency. In: Moo-Young M (ed) Comprehensive biotechnology, Vol. 4. Elsevier, Pergamon, pp 211–220
Raghuram N, Sopory SK (1995) Evidence for some common signal transduction events for opposite regulation of nitrate reductase and phytochrome-I gene expression by light. Plant Mol Biol 29:25–35
Rochaix JD (2013) Redox regulation of thylakoid protein kinases and photosynthetic gene expression. Antioxid Redox Signal 18:2184–2201
Roitinger E, Hofer M, Kocher T, Pichler P, Novatchkova M, Yang J, Schlogelhofer P, Mechtler K (2015) Quantitative phosphoproteomics of the ataxia telangiectasia-mutated (ATM) and ataxia telangiectasia-mutated and rad3-related (ATR) dependent DNA damage response in Arabidopsis thaliana. Mol Cell Proteomics 14:556–571
Samol I, Shapiguzov A, Ingelsson B, Fucile G, Crevecoeur M, Vener AV, Rochaix JD, Goldschmidt-Clermont M (2012) Identification of a photosystem II phosphatase involved in light acclimation in Arabidopsis. Plant Cell 24:2596–2609
Shapiguzov A, Ingelsson B, Samol I, Andres C, Kessler F, Rochaix JD, Vener AV, Goldschmidt-Clermont M (2010) The PPH1 phophatase is specifically involved in LHCII dephosphorylation and state transitions in Arabidopsis. Proc Natl Acad Sci U S A 107(10):4782–4787
Sharma N, Sinha VB, Gupta N, Rajpal S, Kuchi S, Sitaramam V, Parsad R, Raghuram N (2018) Phenotyping for nitrogen use efficiency (NUE): Rice genotypes differ in N-responsive germination, oxygen consumption, seed urease activities, root growth, crop duration and yield at low N. Front Plant Sci 9:1452
Signora L, De Smet I, Foyer CH, Zhang H (2001) ABA plays a central role in mediating the regulatory effects of nitrate on root branching in Arabidopsis. Plant J 28:655–662
Singh A, Giri J, Kapoor S, Tyagi AK, Pandey GK (2010) Protein phosphatase complement in rice: genome-wide identification and transcriptional analysis under abiotic stress conditions and reproductive development. BMC Genomics 11:435
Singh A, Jha SK, Bagri J, Pandey GK (2015) ABA inducible rice protein phosphatase 2 C confers ABA insensitivity and abiotic stress tolerance in Arabidopsis. PLoS One 10:e0125168
Sinha SK, Sevanthi VAM, Chaudhary S, Tyagi P, Venkadesan S, Rani M, Mandal PK (2018) Transcriptome analysis of two rice varieties contrasting for nitrogen use efficiency under chronic N starvation reveals differences in chloroplast and starch metabolism-related genes. Genes 11:E206
Sitt M, Schulze ED (1994) Does Rubisco control the rate of photosynthesis and plant growth? An exercise in molecular ecophysiology. Plant Cell Environ 17:465–487
Straub T, Ludewig U, Neuhauser B (2017) The kinase CIPK23 inhibits ammonium transport in Arabidopsis thaliana. Plant Cell 29:409–422
Sulpice R, Nikoloski Z, Tschoep H, Antonio C, Kleessen S, Larhlimi A, Selbig J, Ishihara H, Gibon Y, Fernie AR, Stitt M (2013) Impact of the carbon and nitrogen supply on relationships and connectivity between metabolism and biomass in a broad panel of Arabidopsis accessions. Plant Physiol 162:347–363
Sun H, Qian Q, Wu K, Luo J, Wang S, Zhang C, Ma Y, Liu Q, Huang X, Yuan Q, Han R, Zhao M, Dong G, Guo L, Zhu X, Gou Z, Wang W, Wu Y, Lin H, Fu X (2014) Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet 46(6):652–656
Sutton M, Raghuram N, Adhya TK (2019) The nitrogen fix: from nitrogen cycle pollution to nitrogen circular economy in emerging issues of environmental concern, Frontiers 2018/19, pp 52–64
Sutton MA, Bleeker A (2013) The shape of nitrogen to come. Nature 494:435–437
Trevisan S, Manoli A, Begheldo M, Nonis A, Enna M, Vaccaro S, Caporale G, Ruperti B, Quaggiotti S (2011) Transcriptome analysis reveals coordinated spatiotemporal regulation of hemoglobin and nitrate reductase in response to nitrate in maize roots. New Phytol 192:338–352
Umezawa T, Sugiyama N, Mizoguchi M, Hayashi S, Myouga F, Yamaguchi-Shinozaki K, Ishihama Y, Hirayama T, Shinozaki K (2009) Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc Natl Acad Sci U S A 106:17588–17593
Vert G, Chory J (2009) A toggle switch in plant nitrate uptake. Cell 138:1064–1066
Walch-Liu P, Filleur S, Gan Y, Forde BG (2005) Signaling mechanisms integrating root and shoot responses to changes in the nitrogen supply. Photosynth Res 83:239–250
Wang Y, Wang Y, Zhao YB, Chen DM, Han ZH, Zhang XZ (2014) Protein phosphorylation differs significantly among ontogenetic phases in malus seedlings. Proteome Sci 12:31
Waqas M, Feng S, Amjad H, Letuma P, Zhan W, Li Z, Fang C, Arafat Y, Khan MU, Tayyab M, Lin W (2018) Protein phosphatase (PP2C9) induces protein expression differentially to mediate nitrogen utilization efficiency in rice under nitrogen-deficient condition. Int J Mol Sci 19:2827
Whitney SM, Birch R, Kelso C, Beck JL, Kapralov MV (2015) Improving recombinant rubisco biogenesis, plant photosynthesis and growth by co-expressing its ancillary RAF1 chaperone. Proc Natl Acad Sci U S A 112:3564–3569
Xiong H, Guo H, Zhou C, Guo X, Xie Y, Zhao L, Gu J, Zhao S, Ding Y, Liu L (2019) A combined association mapping and t-test analysis of SNP loci and candidate genes involving in resistance to low nitrogen traits by a wheat mutant population. PLoS One 14(1):e0211492
Xue T, Wang D, Zhang S, Ehlting J, Ni F, Jakab S, ZhengC Zhong Y (2008) Genome-wide and expression analysis of protein phosphatase 2C in rice and Arabidopsis. BMC Genom 9:550
Yang W, Yoon J, Choi H, Fan Y, Chen R, An G (2015) Transcriptome analysis of nitrogen-starvation-responsive genes in rice. BMC Plant Biol 15:31
Zeng DD, Qin R, Li M, Alamin M, Jin XL, Liu Y, Shi CH (2017) The ferredoxin-dependent glutamate synthase (OsFd-GOGAT) participates in leaf senescence and the nitrogen remobilization in rice. Mol Genet Genomics 292(2):385–395
Zhang H, Forde B (2000) Regulation of Arabidopsis root development by nitrate availability. J Exp Bot 51:51–59
Zhang H, Forde BG (1998) An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture. Science 279:407–409
Acknowledgments
This work was supported in part by research grant to N.R. and fellowship to S.K. from the Indo-UK Virtual Centre on Nitrogen Efficiency of Whole cropping Systems (NEWS) BT/IN/UK-VNC/44/NR/ 2015-16 and UKRI GCRF South Asian Nitrogen Hub (NE/S009019/1).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kumari, S., Raghuram, N. (2020). Protein Phosphatases in N Response and NUE in Crops. In: Pandey, G.K. (eds) Protein Phosphatases and Stress Management in Plants. Springer, Cham. https://doi.org/10.1007/978-3-030-48733-1_12
Download citation
DOI: https://doi.org/10.1007/978-3-030-48733-1_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-48732-4
Online ISBN: 978-3-030-48733-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)