Abstract
Ferredoxin-dependent glutamate synthase (Fd-GOGAT, EC 1.4.7.1) plays major roles in photorespiration and primary nitrogen assimilation. However, due to no mutant or knockdown lines of OsFd-GOGAT have been reported in rice (Oryza sativa L.), the contribution of OsFd-GOGAT to rice foliar nitrogen metabolism remains little up-to-date. Here, we isolated a rice premature leaf senescence mutant named gogat1, which was reduced in 67% of the total GOGAT enzyme activity in leaves. The gogat1 mutant exhibited chlorosis under natural condition, while showed less extent premature leaf senescence under low light treatment. The gogat1 locus was mapped to a 54.1 kb region on chromosome 7, and the sequencing of OsFd-GOGAT showed one substitution (A to T) at the 3017th nucleotide of the open reading frame, leading to the amino-acid substitution of leucine changed to histidine. The gogat1 mutant showed reduced seed setting rate, while the grain protein content in gogat1 mutant was significantly higher than that in wild type. Meanwhile, during the grain-filling stage, total amino acids in the up three leaves and the upmost internode were increased dramatically. The results in this study suggested that OsFd-GOGAT might participate in nitrogen remobilization during leaf senescence, which provides a potential way to improve nitrogen use efficiency in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Productive agriculture consumes a large amount of expensive nitrogenous fertilizers which have been one of the most expensive nutrients to supply. Lowering fertilizer input and increasing plant nitrogen use efficiency (NUE) are essential for sustainable agriculture (Masclaux-Daubresse et al. 2006; Xu et al. 2012). NUE mainly depends on N uptake, translocation, assimilation, and remobilization. Glutamate synthase (also termed glutamine 2-oxoglutarate aminotransferase, GOGAT), in concert with glutamine synthetase (GS, EC 6.3.1.2), catalyzes the reaction that converts ammonia and 2-oxoglutarate to glutamate. GS/GOGAT cycle is considered to be the primary route of nitrogen assimilation in oxygenic photosynthetic organisms (Suzuki and Knaff 2005; Tripathy et al. 2015). In higher plants, GOGAT occurs in two isoforms (Fd-GOGAT form and NADH-GOGAT form), and both forms are located in chloroplast and/or plastid. NADH-GOGAT is predominantly located in non-photosynthesizing cells. In contrast, Fd-GOGAT enzyme usually performed high activity in the chloroplasts of photosynthetic tissues (Nigro et al. 2014; Suzuki and Knaff 2005; Temple et al. 1998). GS/Fd-GOGAT cycle preforms the predominant role in the assimilation of ammonium derived from photorespiration. Rice plants lose considerable volatile NH3 from their leaves as a consequence of photorespiration (Kumagai et al. 2011a), thus enhancing the reassimilation of photorespiratory ammonia at the grain-filling stage is a potential avenue to improve NUE.
Fd-GOGAT is an iron–sulfur and FMN-containing enzyme with a subunit molecular mass of 130–180 kDa that functions as a monomer (Suzuki and Knaff 2005; van den Heuvel et al. 2004). In Arabidopsis, Fd-GOGAT was encoded by two genes, GLU1 and GLU2, and GLU1 mRNA was expressed at the highest levels in leaves, while GLU2 mRNA was expressed at lower levels in leaves and preferentially accumulated in roots (Coschigano et al. 1998). In rice, immunolocalization studies indicated that there were two different immunogenicity types of Fd-GOGAT, leaf isoform Fd-GOGAT and root isoform Fd-GOGAT (Coschigano et al. 1998; Ishiyama et al. 1998, 2003; Suzuki and Knaff 2005). However, the DNA sequence of root isoform Fd-GOGAT is not found in rice genome database, and the function is still not clear.
Glutamate occupies a central position in amino-acid metabolism in plants. There is evidence that plants are able to maintain the soluble glutamate concentration within fairly narrow limits under most circumstances (Forde and Lea 2007; Ishizaki et al. 2010; Labboun et al. 2009). Fd-GOGAT, catalyzing an essential step in the pathway of glutamate biosynthesis, was suggested to play roles in glutamate and amino-acid metabolism in plant leaves under photorespiratory conditions (Ferrario-Mery et al. 2000; Ishizaki et al. 2010; Potel et al. 2009). GLU1/Fd-GOGAT could be modulated by GNC and CGA1 (chlorophyll biosynthesis genes), suggesting that Fd-GOGAT might also participate in chlorophyll metabolism (Hudson et al. 2011).
Recent study showed that Fd-GOGAT-A was involved in the control of grain protein content (GPC) in durum wheat (Triticum turgidum ssp. durum) (Nigro et al. 2014). GPC in rice not only played an important role in determining the rice grain quality, but also tightly associated with cooking and eating qualities (Lang et al. 2013; Yang et al. 2015). However, GPC is a typical quantitative trait and easily influenced by environmental factors and management practices (Blanco et al. 2012; Cai et al. 2013). It is difficult to achieve the higher GPC rice varieties by the traditional breeding programs. In this study, a mutant of OsFd-GOGAT showed premature leaf senescence in rice (named gogat1) was isolated and characterized. gogat1 mutant exhibited only 33% of the total GOGAT enzyme activity in leaves, while the glutamate content even had higher levels than wild type. The amino-acid pools in the sap of gogat1 mutant leaves and the upmost internode had higher levels than those of wild type, and GPC in gogat1 mutant was 26.4% higher than that in wild type under natural condition, suggesting that OsFd-GOGAT plays an important role in N remobilization from leaves to seeds.
Materials and methods
Plant materials
The gogat1 mutant was derived from cultivar Zhenong 34 (ssp. indica) by mutagenesis with ethyl methanesulfonate (EMS). Two F2 populations, derived from crossing gogat1 with Zhenong 34 (wild type) and Zhenongda 104 (ssp. japonica), were used for the genetic analysis and molecular mapping of gogat1, respectively. All the plants were grown in the paddy field of Zhejiang University in Hangzhou, China.
Rice plants for light treatment were grown in field condition before treatment. When the rice plants were at the heading stage, plants grown under field condition (natural light, NL) were used as controls, whereas plants grown under a shadow net (about 50% intensity of natural light) throughout the heading to maturity stage were used as the low light (LL) treatment.
Genetic analysis and map-based cloning of OsFd-GOGAT
The segregative ratio of mutant phenotype in the F2 population from the cross of gogat1/Zhenong 34 was analyzed using Microsoft Excel.
The OsFd-GOGAT locus was primarily mapped to a region closely linked to InDel marker R7M37 and SSR marker RM172 on the long arm of chromosome 7 using 14 F2 plants of gogat1/Zhenongda 104. For fine mapping the OsFd-GOGAT gene, another 1231 F2 mutants were used to narrow the OsFd-GOGAT locus to 54.1 kb between two InDel markers I7-153 and I7-159. Mutation in gogat1 mutant was determined by PCR amplification and sequence analysis.
Histochemistry
Histochemistry assays for cell death and reactive oxygen species (ROS) accumulation were conducted as previously described by Zhou et al. (2013). Briefly, for trypan blue staining, samples were submerged in lactic acid–phenol–trypan blue solution [2.5 mg/mL trypan blue, 25% (w/v) lactic acid, 23% water-saturated phenol, and 25% glycerol in H2O] at 70 °C, infiltrated for 20 min, heated in boiling water for 2 min, and then cooled for 1.5 h. Samples were destained in chloral hydrate solution (2.5 g/mL in H2O). For superoxide determination, leaf samples were immersed in 0.5 mg/mL nitro blue tetrazolium (NBT) in 10 mM potassium phosphate buffer (pH 7.8) for 16 h in darkness. For H2O2 detection, leaf samples were immersed in 1 mg/mL 3,3′-diaminobenzidine (DAB) containing 10 mM MES (pH 6.5) for 18 h in darkness. Both reactions were stopped by transferring to 90% ethanol and held at 70 °C until chlorophyll was completely removed.
RNA isolation and quantitative RT-PCR analysis
Total RNAs were extracted using Trizol reagent according to the manufacturer’s protocol (Invitrogen, USA). 1 μg RNase-free DNase I treated total RNA was used to synthesize the first-strand cDNA in a reaction volume of 20 μL (Takara, Japan). Quantitative RT-PCR was performed using the SYBR Premix ExTaq™ (Tli RNaseH Plus) (Takara, Japan). The rice actin gene was used as an internal control for the quantitative RT-PCR analysis. PCR was carried out with the two-step protocol as follows: activation at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 5 s, and annealing/extension at 60 °C for 30 s.
Enzyme extraction and assays
Glutamate synthase was extracted and assayed as described by Wallsgrove et al. (1982). Briefly, glutamate synthase was extracted by grinding leaves in a pre-cooled pestle and mortar with buffer A [100 mM potassium phosphate pH 7.5, 100 mM KCl, 5 mM ethylene diamine tetraacetic acid (EDTA), 0.1% (v/v) 2-mercaptoethanol, 10% (v/v) glycerol, and 0.1% (v/v) triton X-100] at a tissue to volume ratio of 0.25. The homogenate was centrifuged 10,000g at 4 °C for 20 min. 200 μL supernatant was used in a 1 mL reaction volume mixture containing 4 mM 2-oxoglutarate, 1 mM l-glutamine, 50 mM potassium phosphate (pH 7.5), and 100 μg methyl viologen plus 100 μL sodium dithionite solution (25 mg/mL in 0.3 M NaHCO3). The reaction was started by the addition of dithionite incubated at 30 °C for 10 min and stopped at boiled water for 90 s. Glutamate content was determined by amino-acid auto-analyzer (L-8900, Hitachi). GOGAT enzyme activities were estimated by the amount of l-glutamate after terminating the reaction.
Determination of rice grain protein content and amino acids in leaves and seeds
The seeds were dehusked, and then the milled rice samples were ground to 100-mesh for the determination of GPC. The rice GPC was determined by the Kjeldahl method (Sáez-Plaza et al. 2013). For free amino acids detection, 200 mg fresh leaves were grinded with 1 mL 8% sulfosalicylic acid. The homogenate was shaken for 1 h, and then centrifuged at 10,000g for 10 min. The supernatant was purified with a 0.45 μm filter, and a 20 μL sample was injected into the amino-acid analyzer (L-8900, Hitachi) for cation-exchange chromatography separation. Free amino acids were detected by spectrophotometric analysis with ninhydrin reagent. Data were analyzed by the software EZChrom Elite for Hitachi AAA.
Statistical analysis
All the determinations were performed in at least three independent experiments. Statistical differences were analyzed by the analysis of variance using the Microsoft Excel. The mean were compared by the least significant difference (LSD) test. Standard deviation (SD) was calculated and shown in the figures and tables.
Results
Phenotype characterization of gogat1
There was no obvious phenotypic difference between gogat1 mutant and wild-type Zhenong 34 before five-leaf stage. The premature leaf senescence of gogat1 was initiated from the bottom leaves about 40 days after germination, and all the functional leaves exhibited significantly accelerated senescence from heading time under natural condition. All the up four leaves showed rust-colored premature leaf senescence at late filling stage, while most leaves of wild type were still green at the same stage (Fig. 1a, b). The survey of agronomic traits showed that the plant height of gogat1 mutant was shorter, and the tillering number and seed setting rate were significantly decreased (Fig. 1c). However, the 1000-grain weight did not show visibly difference (Table 1).
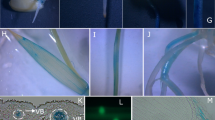
Phenotype characterization of gogat1. a Phenotypes of the gogat1 mutant at 10 days after flowering (DAF), Scale bars 10 cm. b Morphology of the four leaves from the top. FL flag leaf, 2L penultimate leaf, 3L antepenultimate leaf, 4L the fourth leaf from the top. c The panicle comparison of gogat1 mutant and wild-type plant at mature stage. Scale bars 2.5 cm. d Chlorophyll content of flag leaves in gogat1 mutant and wild-type plant after flowering. Values are mean ± SD of three biological replicates; FW fresh weight. e Analysis of histological staining of gogat1 mutant and the wild type. f Expression of senescence-associated genes (OsDOS, Osl85, and OsNAP) and chlorophyll degradation-related genes (SGR, OsNYC3, and OsRCCR1) in gogat1 mutant and wild-type plant. The gray bar represents wild-type plants and the white bar represents gogat1 mutants. Values are mean ± SD of three biological replicates; **P ≤ 0.01; Student’s t test
After flowering, the chlorophyll content of gogat1 was dramatically degraded compared with wild-type plants. At 30 DAF (days after flowering), the chlorophyll content in the flag leaves of gogat1 mutant was only 20.1% of that in wild-type plants (Fig. 1d). Synchronously, the chlorophyll degradation-related genes (CDGs), SGR (Jiang et al. 2007), OsNYC3 (Morita et al. 2009), and OsRCCR (Tang et al. 2011), in gogat1 flag leaves at 10 DAF were expressed with higher levels than those in wild type (Fig. 1f).
Typan blue staining is a histochemical indicator of irreversible membrane damage or cell death. The results showed that more deep blue color necrosis sites were accumulated in gogat1 mutant. When NBT was used as an indicator of O2 − accumulation, more blue formazan precipitates were formed in gogat1 leaves. DAB was used to detect the accumulation of H2O2; widespread intense brown staining was found in the leaves of gogat1 (Fig. 1e). In accordance with the histochemistry assay, senescence down-regulate gene, OsDOS (Kong et al. 2006), was expressed at lower level, while the mRNA levels of two other senescence up-regulate genes, OsNAP (Liang et al. 2014) and Osl85 (Lee et al. 2001), were accumulated more in gogat1 than those in wild type (Fig. 1f).
Map-based cloning of OsFd-GOGAT
The F1 plants derived from the cross between gogat1 and Zhenong 34 or Zhenongda 104 both exhibited the wild-type phenotype. The F2 segregation of gogat1/Zhenong34 was 168 wild-type plants to 52 mutant-type plants, which fitted the ratio of 3:1 (χ 2 = 0.15 < χ 20.05 = 3.84). These results demonstrated that the gogat1 mutant trait was controlled by a single recessive nuclear gene.
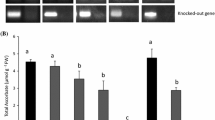
To elucidate the premature leaf senescence phenotype of mutant from molecular genetics, map-based cloning was performed with the F2 population derived from gogat1/Zhenongda 104. The OsFd-GOGAT locus was primarily mapped to a region closely linked to InDel marker R7M37 and SSR marker RM172 on the long arm of chromosome 7 (Fig. 2a), and was subsequently fine mapped at an interval of 54.1 kb between InDel markers I7-153 and I7-159 (Fig. 2b, c). According to Rice Genome Annotation Project (http://rice.plantbiology.msu.edu/), five putative genes were annotated in this region (Fig. 2d). Among these genes, LOC_Os07g46460, encoding a ferredoxin-dependent glutamate synthase, was specially focused on in current study. To identify the mutation of gogat1, the locus LOC_Os07g46460 was amplified from gogat1 mutant and wild-type plants by PCR and then sequenced. The genome DNA sequencing result showed that the gogat1 mutant had an A-to-T substitution (+3017) in the open reading frame (ORF) of LOC_Os07g46460 (Fig. 2e), which could be confirmed by the cDNA sequencing. These results indicated that LOC_Os07g46460 was responsible for the mutation of gogat1.
Map-based cloning of OsFd-GOGAT locus. a Primary mapping of gogat1 between the markers I7-92 and I7-60 on chromosome 7 based on 84 F2 mutant individuals. b and c The OsFd-GOGAT gene was fine mapped to an interval of 54.1 kb region by 1231 F2 mutant individuals. d Five genes were annotated in the 54.1 kb candidate region, and LOC_Os07g46460 was suspected to be OsFd-GOGAT. e Structure of OsFd-GOGAT and the mutation locus
Analysis of OsFd-GOGAT and orthologous proteins
The coding DNA fragment of OsFd-GOGAT is 15,525-bp with 33 exons and 32 introns. The full-length cDNA of OsFd-GOGAT (GeneBank Accession No: AB024716.1) was 5803-bp in length containing a 5`-UTR (411-bp), a coding sequence (4848-bp), and a 3`-UTR (544-bp). The CDs was predicted to encode a polypeptide of 1615 amino acids with calculated molecular mass of 175 kDa. In the gogat1 mutant, the A-to-T substitution was located in the 20th exon, leading to the amino-acid substitution of leucine to histidine.
The orthologous proteins were detected in the National Center for Biotechnology Information (NCBI) database from Arabidopsis thaliana, Medicago truncatula, Zea mays, Spinacia oleracea, Cyanidium caldarium, Synechocystis sp. PCC 6803, and Paenibacillus sp. Aloe-1. These proteins had 45–94% amino-acid sequence identity, containing glutamine amidotransferase domain (GAT domain), FMN-binding domain, glutamate synthase central domain, and C-terminal domain (Fig. S1). Orthologs of OsFd-GOGAT have been detected in higher plants, algae, and cyanobacteria as the phylogenetic tree (Fig. S2). The Leucine-to-Histidine change was located in the FMN-binding domain which was suggested that a reduced ferredoxin (Fd) binds to the domain to transfer an electron for 2-iminoglutarate reduction (van den Heuvel et al. 2004). The three-dimensional protein structure prediction OsFd-GOGAT model of gogat1 mutant showed that a fragment of α-helix was missing compared with wild type in the FMN-binding domain by Swiss-Model server (Biasini et al. 2014) (Fig. S3).
Expression pattern of OsFd-GOGAT
The mRNA levels of OsFd-GOGAT in different organs were detected to identify the expression pattern of OsFd-GOGAT at heading stage. The result showed that the OsFd-GOGAT was mainly expressed in leaf, while performed lower levels in panicle, stem, and root. It was notable that OsFd-GOGAT transcript levels in leaves were declining dramatically when the leaves become senescent (Fig. 3a).
Expression pattern of OsFd-GOGAT and the detection of total GOGAT activities and the expression of other GOGAT genes (OsNADH-GOGAT1 and OsNADH-GOGAT2) at heading stage. a qRT-PCR analysis showing OsFd-GOGAT expression in leaf, root, stem, and panicle. FL flag leaf, 2L penultimate leaf, 3L antepenultimate leaf, 4L the fourth leaf from the top. b Determination of total GOGAT activities in the flag leaf of gogat1 mutant and wild type. c and d qRT-PCR detection of the expression of OsNADH-GOGAT1 and OsNADH-GOGAT2, respectively. The gray bar in b–d represents wild-type plants, and the black bar in b–d represents gogat1 mutants. Values are mean ± SD of three biological replicates; **P ≤ 0.01; Student’s t test
Detection of total GOGAT activities and the expression of other GOGAT genes
As Fd-GOGAT accounts for up to 96% of the total GOGAT activity in leaves (Coschigano et al. 1998), the total GOGAT enzyme activity was determined to evaluate the Fd-GOGAT enzyme activity. The result exhibited that gogat1 mutant only performed 33% leaf GOGAT enzyme activity compared with that in wild type (Fig. 3b).
Besides OsFd-GOGAT, the GOGAT gene family has another two NADH-dependent members (OsNADH-GOGAT1 and OsNADH-GOGAT2) in rice (Tamura et al. 2010). We further detected the expression of OsNADH-GOGAT1 and OsNADH-GOGAT2, and both genes expressed significantly lower levels in the flag leave of gogat1 mutant (Fig. 3c, d), indicating that NADH-dependent-type GOGATs were not able to compensate for OsFd-GOGAT function.
Determination of free amino-acid pool in leaves and the upmost internode
To evaluate the effect of reduced OsFd-GOGAT enzyme activity in amino-acid metabolism, 18 out of the 20 protein-forming ones in the flag leaves and upmost internode were determined. As the results showed in Table 2, most of the amino acids in both flag leaves and upmost internode of gogat1 mutant had different levels in comparison with those in wild type. Amino-acid pool in both organs of gogat1 mutant had higher levels than that in wild type. In the sap of flag leaf, glutamine contents of gogat1 mutant accumulated about 12-fold as much as those of wild type, and contributed the most to the increased amino acid. In the sap of the upmost internode, the amino-acid pool of gogat1 mutant was almost eight times as much as that of wild type and asparagine accounted 61.4% of the total amino acid.
Discussion
Large amount of ammonium is generated from a wide range of physiological process in leaf, such as photorespiration, nitrate reduction, protein turnover, and lignin biosynthesis (Kumagai et al. 2011b). During photorespiration, ammonium is derived from the conversion of glycine to serine, and the photorespiratory ammonium can be 10–20-fold as much as the primary nitrate reduction in leaves (Keys et al. 1978; Somerville and Ogren 1980). Efficient recapture of the photorespiratory ammonium is essential for survival of the plant (Coschigano et al. 1998; Kendall et al. 1986; Krapp 2015; Potel et al. 2009). The Fd-GOGAT mutations or antisense lines with decreased Fd-GOGAT enzyme activity became chlorotic and even died when grew under air conditions, while grew normal as wild type under high CO2 conditions (Coschigano et al. 1998; Ferrario-Mery et al. 2000; Somerville and Ogren 1980). In this study, we obtained a premature leaf senescence mutant in rice named gogat1, which was reduced in 67% of the total GOGAT enzyme activity in leaves. By map-based cloning, an A-to-T substitution was identified at the 3017th nucleotide of LOC_Os07g46460 which encodes a ferredoxin-dependent glutamate synthase. In accordance with the altered Fd-GOGAT lines, gogat1 mutant grew under LL treatment showed less degree leaf senescence than that under NL condition (Fig. 4). These results proved that LOC_Os07g46460 is the allele of OsFd-GOGAT.
In rice, the GOGAT gene family has two NADH-dependent types (OsNADH-GOGAT1 and OsNADH-GOGAT2) and one Fd-dependent type (OsFd-GOGAT) (Tamura et al. 2010). OsNADH-GOGAT1, mainly expressed in roots, was important in primary N assimilation. OsNADH-GOGAT2 was mainly expressed in leaves, and the mutants had reduced spikelet number per panicle (Yamaya1 and Kusano 2014). Expression of OsNADH-GOGAT1 and OsNADH-GOGAT2 in gogat1 mutant both exhibited lower levels than that in wild type (Fig. 3c, d), indicating that the NADH-dependent GOGATs have distinct functions and NADH-dependent-type GOGATs could not compensate for OsFd-GOGAT function.
The photorespiratory ammonium originates in glutamate and is eventually reassimilated to glutamate (Igarashi et al. 2003; Ishizaki et al. 2010). Glutamate, playing a central role in plant nitrogen metabolism, can be synthesized and metabolized by a number of different pathways. The soluble concentration of glutamate is able to be maintained within stable levels under most circumstances (Forde and Lea 2007). In the Fd-GOGAT mutant lines of Arabidopsis and barely, an accumulation of glutamine and reduction of glutamate was showed after transferring from high CO2 condition to normal air (Blackwell et al. 1988; Leegood et al. 1995; Potel et al. 2009). The antisense lines containing only 15% Fd-GOGAT activity in tobacco showed similar results (Ferrario-Mery et al. 2000). However, gogat1 mutant performed 33% of wild-type OsFd-GOGAT enzyme activity, the glutamate content of gogat1 mutant even had higher levels than that of wild type (Table 2). There should be some other pathways to replenish the glutamate pool. Glutamate dehydrogenase (GDH) can alternatively incorporate ammonium to glutamate in response to high levels of ammonium (Skopelitis et al. 2006), and the mRNA levels of OsGDH1 and OsGDH2 in gogat1 mutant showed significantly higher levels than those in wild type (Fig. 5). Total free amino acid in the leaves of gogat1 mutant was higher than that of wild type (Fig. 6a), while soluble protein content was lower with respect to wild type (Fig. 6b). As glutamate is mainly synthesized from glutamine via GOGAT or from amino acids via aminotransferase (Forde and Lea 2007), the results suggested that the replenished glutamate might derive from the hydrolysis of proteins and transamination reaction from other amino acids.
Total free amino-acid and soluble protein contents in the up three leaves of gogat1 mutant and wild type under natural condition at late grain-filling stage. a Total free amino-acid contents in gogat1 mutant and wild type under nature condition. b Soluble protein contents in gogat1 mutant and wild type under natural condition. FL flag leaf, 2L penultimate leaf, 3L antepenultimate leaf. The gray bar represents wild-type plants, and the black bar represents gogat1 mutants. Values are mean ± SD of three biological replicates; *P ≤ 0.05, **P ≤ 0.01; Student’s t test
GS facilitates glutamate and ammonium to yield glutamine, which is a key step in N recycling in senescent organs. Glutamine contributes the most to the increased amino-acid pool in the flag leaf of gogat1 mutant (Table 2), suggesting that the photorespiratory ammonium was primarily detoxified to glutamine. GS is represented by the cytosolic form (GS1) and plastidic form (GS2). There are three GS1 genes (OsGS1;1, OsGS1;2, and OsGS1;3) and one GS2 gene (OsGS2) in rice genome. OsGS1;1 was mainly expressed in vascular tissues of mature leaf blades (Yamaya and Kusano 2014), and GS2 and Fd-GOGAT in chloroplasts are responsible for the reassimilation of ammonia released during photorespiration (Kendall et al. 1986). However, the detection of OsGS2 expression in the flag leaf at late filling stage showed its mRNA accumulated at very low levels (Fig. 5). GS2 is located in chloroplasts which exhibited the first symptoms of deterioration during leaf senescence (Masclaux-Daubresse et al. 2010). As the flag leaf of gogat1 mutant showed severe premature senescence at grain-filling stage, the dismantling of chloroplasts might inhibit the transcription of OsGS2. By contrast, OsGS1;1 transcript level in gogat1 mutant was sharply increased (Fig. 5), indicating that OsGS1;1 performs a crucial function in assimilating ammonia in senescent leaves.
During seed production, nitrogen uptake is negatively regulated or even totally inhibited in some cases, and therefore, the strong N demand from grain filling has to be supported from N recycling from leaf, stem, and root (Masclaux-Daubresse et al. 2010). N remobilization in plants is a very complex metabolic process and has important function in plant productivity, because it recycles organic N to young leaves and storage organs. In rice, around 80% of the total N of the panicle derives from remobilization through the phloem from senescent organs (Tabuchi et al. 2007). The GPC of gogat1 mutant was 26.4% higher than that of wild type (Fig. 7), and the decreased seed setting rate could lead to higher N accumulation per grain. In the sap of the upmost internode, total amino-acid content in gogat1 mutant was almost eightfold as much as that in wild type (Table 2). Besides decreased seed setting rate, the soaring amino-acid content might contribute to the increased GPC in the gogat1 mutant.
Glutamine and asparagine have higher N/C ratio and can be used as long-range transport and storage compounds. The major forms of nitrogen in the phloem sap of rice are glutamine and asparagine (Hiroaki and Chino 1990). Glutamine accounted for about 40% of total amino acid in the sap of wild type upmost internode, while asparagine constituted 61.4% of the total amino in gogat1 mutant (Table 2). Asparagine is catalyzed by asparagine synthetase (AS) from aspartate using either the Gln-amide group or ammonium, indicating that AS could compensate the reduced GS-dependent ammonium assimilatory activity in certain situations and the excessive ammonium during photorespiration could be incorporated into asparagine serving as an additional detoxification molecule (Masclaux-Daubresse et al. 2006, 2010). The observation of OsAS1 and OsAS2 transcript levels in the leaves of gogat1 mutant showed that these genes had significantly higher levels than those in the leaves of wild type (Fig. 5). The rapid senescence of goagat1 leaves might induce the expression of AS genes, leading to more asparagine generation and reallocation to the sap of phloem.
N remobilization is associated with leaf senescence and is crucial for seed filling and yield. However, crop breeding has to cope with the dilemma that delayed senescence could not only lead to higher yields but also to a decrease in N remobilization efficiency (NRE) and NUE as such leaves usually maintain high N content. In contrast, rapid senescence increases N remobilization from the vegetative parts and thus results in relatively higher NRE and GPC (Masclaux-Daubresse et al. 2010; Xu et al. 2012). The mutation of OsFd-GOGAT led to severe leaf senescence during grain-filling stage, thus resulting in relatively higher NRE and GPC. However, large amount of N might volatilize through photorespiratory pathways, and the yield of gogat1 mutant reduced significantly as a consequence of rapid leaf senescence.
In conclusion, OsFd-GOGAT played a predominant role in the recapture of the photorespiratory ammonium. The mutation of OsFd-GOGAT led to a premature leaf senescence phenotype and facilitated N remobilization which resulted in higher GPC in gogat1 mutant. Further functional analysis of OsFd-GOGAT and enhancing the reassimilation of photorespiratory ammonium at grain-filling stage will provide a new insight to improved NUE.
References
Biasini M, Bienert S, Waterhouse A et al (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258. doi:10.1093/nar/gku340
Blackwell RD, Murray AJS, Lea PJ, Kendall AC, Hall NP, Turner JC, Wallsgrove RM (1988) The value of mutants unable to carry out photorespiration. Photosynth Res 16:155–176. doi:10.1007/Bf00039491
Blanco A, Mangini G, Giancaspro A et al (2012) Relationships between grain protein content and grain yield components through quantitative trait locus analyses in a recombinant inbred line population derived from two elite durum wheat cultivars. Mol Breed 30:79–92. doi:10.1007/s11032-011-9600-z
Cai SG, Yu G, Chen XH, Huang YC, Jiang XG, Zhang GP, Jin XL (2013) Grain protein content variation and its association analysis in barley. BMC Plant Biol 13:35. doi:10.1186/1471-2229-13-35
Coschigano KT, Melo-Oliveira R, Lim J, Coruzzi GM (1998) Arabidopsis gls mutants and distinct Fd-GOGAT genes: implications for photorespiration and primary nitrogen assimilation. Plant Cell 10:741–752. doi:10.1105/Tpc.10.5.741
Ferrario-Mery S, Suzuki A, Kunz C, Valadier MH, Roux Y, Hirel B, Foyer CH (2000) Modulation of amino acid metabolism in transformed tobacco plants deficient in Fd-GOGAT. Plant Soil 221:67–79. doi:10.1023/A:1004715208478
Forde BG, Lea PJ (2007) Glutamate in plants: metabolism, regulation, and signalling. J Exp Bot 58:2339–2358. doi:10.1093/jxb/erm121
Hiroaki H, Chino M (1990) Chemical composition of phloem sap from the upper most internode of the rice plant. Plant Cell Physiol 31:247–251
Hudson D, Guevara D, Yaish MW, Hannam C, Long N, Clarke JD, Bi YM, Rothstein SJ (2011) GNC and CGA1 modulate chlorophyll biosynthesis and glutamate synthase (GLU1/Fd-GOGAT) expression in Arabidopsis. PLoS One 6:e26765. doi:10.1371/journal.pone.0026765
Igarashi D, Miwa T, Seki M et al (2003) Identification of photorespiratory glutamate: glyoxylate aminotransferase (GGAT) gene in Arabidopsis. Plant J 33:975–987. doi:10.1046/j.1365-313X.2003.01688.x
Ishiyama K, Hayakawa T, Yamaya T (1998) Expression of NADH-dependent glutamate synthase protein in the epidermis and exodermis of rice roots in response to the supply of ammonium ions. Planta 204:288–294. doi:10.1007/s004250050258
Ishiyama K, Kojima S, Takahashi H, Hayakawa T, Yamaya T (2003) Cell type distinct accumulations of mRNA and protein for NADH-dependent glutamate synthase in rice roots in response to the supply of NH4 +. Plant Physiol Biochem 41:643–647. doi:10.1016/S0981-9428(03)00078-0
Ishizaki T, Ohsumi C, Totsuka K, Igarashi D (2010) Analysis of glutamate homeostasis by overexpression of Fd-GOGAT gene in Arabidopsis thaliana. Amino Acids 38:943–950. doi:10.1007/s00726-009-0303-2
Jiang H, Li M, Liang N et al (2007) Molecular cloning and function analysis of the stay green gene in rice. Plant J 52:197–209. doi:10.1111/j.1365-313X.2007.003221.x
Kendall AC, Wallsgrove RM, Hall NP, Turner JC, Lea PJ (1986) Carbon and nitrogen metabolism in barley (Hordeum vulgare L.) mutants lacking ferredoxin-dependent glutamate synthase. Planta 168:316–323. doi:10.1007/BF00392355
Keys AJ, Bird IF, Cornelius MJ, Lea PJ, Wallsgrove RM, Miflin BJ (1978) Photorespiratory nitrogen cycle. Nature 275:741–743. doi:10.1038/275741a0
Kong ZS, Li MN, Yang WQ, Xu WY, Xue YB (2006) A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice. Plant Physiol 141:1376–1388. doi:10.1104/pp.106.082941
Krapp A (2015) Plant nitrogen assimilation and its regulation: a complex puzzle with missing pieces. Curr Opin Plant Biol 25:115–122. doi:10.1016/j.pbi.2015.05.010
Kumagai E, Araki T, Hamaoka N, Ueno O (2011a) Ammonia emission from rice leaves in relation photorespiration and genotypic differences in glutamine synthetase activity. Ann Bot 108:1381–1386. doi:10.1093/aob/mcr245
Kumagai E, Araki T, Ueno O (2011b) Ammonia emission from leaves of different rice (Oryza sativa L.) cultivars. Plant Prod Sci 14:249–253
Labboun S, Tercé-Laforgue T, Roscher A et al (2009) Resolving the role of plant glutamate dehydrogenase. I. in vivo real time nuclear magnetic resonance spectroscopy experiments. Plant Cell Physiol 50:1761–1773. doi:10.1093/pcp/pcp118
Lang GH, Kagiya Y, Ohnishi-Kameyama M, Kitta K (2013) Evaluation of extraction solutions for biochemical analyses of the proteins in rice grains. Biosci Biotechnol Biochem 77:126–131. doi:10.1271/bbb.120617
Lee RH, Wang CH, Huang LT, Chen SCG (2001) Leaf senescence in rice plants: cloning and characterization of senescence up-regulated genes. J Exp Bot 52:1117–1121. doi:10.1093/jexbot/52.358.1117
Leegood RC, Lea PJ, Adcock MD, Hausler RE (1995) The regulation and control of photorespiration. J Exp Bot 46:1397–1414
Liang C, Wang Y, Zhu Y et al (2014) OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. P Natl Acad Sci USA 111:10013–10018. doi:10.1073/pnas.1321568111
Masclaux-Daubresse C, Reisdorf-Cren M, Pageau K et al (2006) Glutamine synthetase-glutamate synthase pathway and glutamate dehydrogenase play distinct roles in the sink-source nitrogen cycle in tobacco. Plant Physiol 140:444–456
Masclaux-Daubresse C, Daniel-Vedele F, Dechorgnat J, Chardon F, Gaufichon L, Suzuki A (2010) Nitrogen uptake, assimilation and remobilization in plants: challenges for sustainable and productive agriculture. Ann Bot 105:1141–1157. doi:10.1093/aob/mcq028
Morita R, Sato Y, Masuda Y, Nishimura M, Kusaba M (2009) Defect in non-yellow coloring 3, an α/β hydrolase-fold family protein, causes a stay-green phenotype during leaf senescence in rice. Plant J 59:940–952. doi:10.1111/j.1365-313X.2009.03919.x
Nigro D, Blanco A, Anderson OD, Gadaleta A (2014) Characterization of ferredoxin-dependent glutamine-oxoglutarate amidotransferase (Fd-GOGAT) genes and their relationship with grain Protein content QTL in wheat. PLoS One 9:e103869. doi:10.1371/journal.pone.0103869
Potel F, Potel F, Valadier MH, Ferrario-Mery S et al (2009) Assimilation of excess ammonium into amino acids and nitrogen translocation in Arabidopsis thaliana—roles of glutamate synthases and carbamoylphosphate synthetase in leaves. FEBS J 276:4061–4076. doi:10.1111/j.1742-4658.2009.07114.x
Sáez-Plaza P, Navas MJ, Wybraniec S, Michałowski T, Asuero AG (2013) An overview of the Kjeldahl method of nitrogen determination. Part II. Sample preparation, working scale, instrumental finish, and quality control. Crit Rev Anal Chem 43:224–272
Skopelitis DS, Paranychianakis NV, Paschalidis KA et al (2006) Abiotic stress generates ROS that signal expression of anionic glutamate dehydrogenases to form glutamate for proline synthesis in tobacco and grapevine. Plant Cell 18:2767–2781
Somerville CR, Ogren WL (1980) Inhibition of photosynthesis in arabidopsis mutants lacking leaf glutamate synthase activity. Nature 286:257–259. doi:10.1038/286257a0
Suzuki A, Knaff DB (2005) Glutamate synthase: structural, mechanistic and regulatory properties, and role in the amino acid metabolism. Photosynth Res 83:191–217. doi:10.1007/s11120-004-3478-0
Tabuchi M, Abiko T, Yamaya T (2007) Assimilation of ammonium ions and reutilization of nitrogen in rice (Oryza sativa L.). J Exp Bot 58:2319–2327. doi:10.1093/jxb/erm016
Tamura W, Hidaka Y, Tabuchi M, Kojima S, Hayakawa T, Sato T, Obara M, Kojima M, Sakakibara Yamaya T (2010) Reverse genetics approach to characterize a function of NADH-glutamate synthase1 in rice plants. Amino Acids 39:1003–1012. doi:10.1007/s00726-010-0531-5
Temple SJ, Vance CP, Gantt JS (1998) Glutamate synthase and nitrogen assimilation. Trends Plant Sci 3:51–56. doi:10.1016/S1360-1385(97)01159-X
Tripathy JN, Hirasawa M, Sutton RB et al (2015) A loop unique to ferredoxin-dependent glutamate synthases is not absolutely essential for ferredoxin-dependent catalytic activity. Photosynth Res 123:129–139. doi:10.1007/s11120-014-0044-2
van den Heuvel RHH, Curti B, Vanoni MA, Mattevi A (2004) Glutamate synthase: a fascinating pathway from l-glutamine to l-glutamate. Cell Mol Life Sci 61:669–681. doi:10.1007/s00018-003-3316-0
Wallsgrove RM, Lea PJ, Miflin BJ (1982) The development of NAD(P)H-dependent and ferredoxin-dependent glutamate synthase in greening barley and pea leaves. Planta 154:473–476. doi:10.1007/Bf01267816
Xu GH, Fan XR, Miller AJ (2012) Plant nitrogen assimilation and use efficiency. Annu Rev Plant Biol 63:153–182
Yang YH, Guo M, Li R et al (2015) Identification of quantitative trait loci responsible for rice grain protein content using chromosome segment substitution lines and fine mapping of qPC-1 in rice (Oryza sativa L.). Mol Breed 35:1–9. doi:10.1007/S11032-015-0328-Z
Zhou QY, Yu Q, Wang Z et al (2013) Knockdown of GDCH gene reveals reactive oxygen species-induced leaf senescence in rice. Plant Cell Environ 36:1476–1489. doi:10.1111/pce.12078
Acknowledgements
This work was supported by the National Science and Technology Support Program (2011BAD35B02), Science and Technology Office of Zhejiang Province (2012C12901-2), the Ministry of Education and Bureau of Foreign Experts of China (Grant B14027), and the Program for Innovative Research Team in University (IRT1185).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Communicated by S. Hohmann.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zeng, DD., Qin, R., Li, M. et al. The ferredoxin-dependent glutamate synthase (OsFd-GOGAT) participates in leaf senescence and the nitrogen remobilization in rice. Mol Genet Genomics 292, 385–395 (2017). https://doi.org/10.1007/s00438-016-1275-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-016-1275-z