Abstract
Prokaryotic methylotrophic bacteria are able to consume a number of C1-carbon compounds such as methane, methylamine and methanol, whereas only methanol can be consumed by eukaryotic methylotrophic bacteria as source of carbon and methylamine as a source of nitrogen. The intensive researches explain the beneficial relationship between plants and methylotrophic bacterial communities earlier. Different genera of methylotrophic yeasts such as Candida, Pichia, Torulopsis and Hansenula are able to metabolise C1 corbon compound like formaldehyde and methanol.and a number of genes are involved in the methanol and other substrate utilisation pathways such as AOX (alcohol oxidase), DAS (dihydroxyacetone synthase), FDH (format dehydrogenase) and DAK (dihydroxyacetone kinase). The phylogeny and identification of these methylotrophic yeast strains are done based on either conserved gene sequences or functional gene sequences. The current description involves the genetic diversity of different strains of methylotrophic yeast from various ecosystems, identified at gene level.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
In the environment, abundant reduced C1-compounds are available. Reduced carbon compounds, such as methane and methanol, utilised by methylotrophs, which have the ability to utilise C1-compounds as the sole source of carbon and energy, also appear to be cosmopolitan in nature. There is a remarkable difference between prokaryotic methylotrophs and eukaryotic methylotrophs. Pokaryotic methylotrophs utilize carbon substrate like methanol, methane and methylamine while methanol is used as carbon substrate and methylamine as nitrogen source by eukaryotic methylotrophs. Methylotrophic yeast comprises genera like Pichia, candida and some other related genera close to Pichia i.e. Kuraishia, Ogataea and Komagataella (Yurimoto et al. 2011).
Different lineages of methylotrophic yeast utilising methanol as sole source of carbon and energy were documented and described (de Koning and Harder 1992). The similar metabolic pathway for methanol utilisation was followed by all methylotrophic yeasts, and they composed of several enzymes localised in peroxisomes which proliferate during growth of yeast in methanol (Veenhuis et al. 1992; Yurimoto et al. 2002). The methylotrophic yeast with regulated promoters of methanol oxidising genes is reported to be used in the study of production of recombinant proteins as well as industrial proteins (Ravin et al. 2013). The diversity of methylotrophic yeast involved in glycerol metabolism was also discussed in one of the earlier reports in which methylotrophic strains such as Candida boidinii no. 2201, Hansenula ofunaensis and Hansenula polymorpha DL-1 were identified with specific enzyme activity of glycerol kinase (GK), glycerol dehydrogenase (GDH) and both GK and GDH, respectively (Tani and Yamada 1987).
Methanol is considered as a very recent alternative carbon source that replaces the petroleum and coal (Olah 2005). It is considered that methanol is formed with the combination of CO and H2 or from CO2 with the help of H2 gas. Methane, an abundant natural carbon substrate produces CO and H2. Since methanol is a cheaper substrate, it can be used as feedstock for different biochemical and biotechnological processes. Methane oxidizing bacterial communities are responsible for release of methanol from methane and further triggers the decomposition of lignins and pectins containg methylester and methoxyl groups respectively (Mitsui et al. 2003; Nakagawa et al. 2000). Methylotrophs and methylotrophic yeasts oxodise CO2.
Thus, methylotrophs play indispensable roles in the global carbon cycle between methane and CO2 called “the methane cycle ”. A thorough understanding of the molecular basis of methylotrophy is needed not only to better understand the global methane cycle but also to permit more efficient use of methanol as a renewable carbon source. In the recycling of carbon in the environment, methylotrophic yeast plays a very crucial role. The ability of C. Boidinii to grow over pectin as a substrate shows its methylotrophic metabolism (Nakagawa et al. 2000). The intensive researches explain the beneficial relationship between plants and methylotrophic microbial communities (Kumar et al. 2019; Meena et al. 2012; Verma et al. 2013, 2014, 2015a, b, 2016a, b; Yadav 2009). Moreover, the interaction between plants and methylotrophic yeasts has not been characterised and very less report or documentation is available (Limtong et al. 2005; van der Klei et al. 2006). Therefore, keeping in view the importance of the methylotrophic yeast in the agriculture, industry and environments, the present chapter deals with the impact of methylotrophic yeast in environments and describes recent insights into the molecular basis of yeast methylotrophy.
3.2 Genetic Diversity of Methylotrophic Yeast
Yeasts are well known for their beneficial activity for humankind by their exploitation and application in the food, beverages and in the production of various types of biochemicals. Since they contain a significant content of vitamin B, amino acids and minerals, they can be used as a food supplement also. Moreover, methylotrophic yeast can be used in the gene regulation study in eukaryotes and as biofactories for heterologous and homologous proteins (Cremata and Díaz 1999; Negruţă et al. 2010) (Table 3.1). This group of yeast is able to survive by metabolising monocarbonic compounds such as formaldehyde and methanol (Kaszycki et al. 2001). Methylotrophic yeasts are able to grow on extract of woods and other pectic material especially in fruits and vegetable products (Craveri et al. 1976). The woody materials are the source of methanol because of the presence of metoxi chain in lignin. Figure 3.1 illustrates the diversity of methylotrophic yeast based on genes involved in their metabolism and physiology.
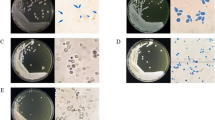
Two methylotrophic yeast strains H. polymorpha and P. pastoris were utilised in the production of heterologous proteins. Apart from this, H. polymorpha is used for studying gene regulation of enzymes associated with abiotic stress tolerance, methanol metabolism, heavy metal resistance and nitrate assimilation. They are widely used in the methanol-contaminated waste water treatment also (Kaszycki and Koloczek 2002; Kaszycki et al. 2001). The methylotrophic yeast has the ability to grow in extreme environment also. In one of the investigations, three novel strains of thermotolerant methylotrophic yeast, which belong to genus Pichia, were reported. The methylotrophic strains were designated as N002, N069 and PT31T. The Pichia strains were isolated from soil (taken from Thailand) enriched with methanol. Thermotolerant yeast is found to grow at 20 °C (minimum temp.) but no limit for the maximum temperature for growth (Arthur and Watson 1976). According to this definition, methylotrophic yeast growing at 20 °C up to a temp. of 37 °C will be called as a thermotolerant methylotrophic yeast (Limtong et al. 2005). The presence of helmet-/hat-shaped ascospores, multilateral budding, ubiquinon Q-7 and negative for Diazonium blue B and urease reactions are the basic characteristics of genus Pichia. They do not have ballistospore and arthrospores also. The phylogenetic analysis based on D1/D2 domain of large subunit rDNA revealed the closeness with Pichia dorogensis. Because of the differences in phenotypic appearance, the above three strains were designated as novel species of Pichia and the name proposed was Pichia thermomethanolica sp. nov. The type strain is PT31T (Limtong et al. 2005).
In a report, whole genome sequencing of a methylotrophic yeast H. polymorpha was performed and total transcripts were analysed from the yeast culture grown in methanol and glucose as well. A total of 9 Mb size of genome was sequenced for Hansenula polymorpha DL1.
In a transcriptome analysis of H. polymorpha under methanol growth condition, 40% genome expression has shown the identified unregulated and abundant gene expression through RNA-seq analysis along with alternate splicing events. From seven chromosomes of H. polymorpha, different proteins of subtelomeric region were identified and the evolutionary relationship established revealed the closeness of H. polymorpha with both non-methylotrophic and methylotrophic yeast Dekkera bruxellensis and Pichia pastoris respectively. In the investigation, phylogenetic analysis indicated the methylotrophic evolutionary pattern in filamentous fungi and yeasts (Ravin et al. 2013).
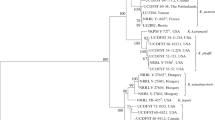
Evolutionary analysis based on methanol-utilising pathway genes evaluated the relatedness of methylotrophic yeasts using NCBI nucleotide database and associated tools. Different genes involved in methanol utilisation pathway such as AOX (alcohol oxidase) , DAS (dihydroxyacetone synthase), FDH (format dehydrogenase) and DAK (dihydroxyacetone kinase) were identified against BLAST searches. Using fast minimum evolution algorithm, phylogenetic tree was prepared to see the evolutionary relatedness. The phylogenetic analysis based on these genes and MEGA software revealed the position of methylotrophic yeasts (Okonechnikov et al. 2012).
Methylotrophic yeast was reported to be a suitable expression system also. To enhance the yields of complex proteins having unnatural amino acids, a recombinant gene expression system was developed in methylotrophic yeast Pichia pastoris . In the investigation, it was emphasised that by modulating aaRS level, the optimization of expression of unnatural amino acids in the methylotrophic host cell was done and better than as reported earlier in Saccharomyces cerevisiae (Young et al. 2012). Earlier, S. cerevisiae was considered to be specific for unnatural amino acids, but in this investigation, it was explained that a mutant of recombinant human serum albumin with p-phenylalanine is expressed efficiently in the methylotrophic yeast system and therefore allows the higher production of complex proteins whose gene expression is difficult in the existing systems (Young et al. 2012).
In a study, the alcohol oxidase activity was analysed in Pichia pastoris on the basis of two genes AOX1 and AOX2 and their expression analysis in the cell (Tschopp et al. 1987). The AOX1 gene expression was observed undetectable when cells are grown in the media with carbon sources other than methanol (Cregg et al. 1989). In a study, the genes encoding polyunsaturated fatty acids are targeted for the identification and phylogenetic analysis of methylotrophic yeast Hansenula polymorpha. In the investigation, gene encoding fatty acid elongase, HpELO1, was identified and characterised. The HpELO1 gene encoding a protein has 319 long amino acids, and it contains 5 different conserved membrane-spanning regions in yeast Elo protein family. The phylogenetic analysis based on amino acid sequences revealed that HpELO1 gene is an orthologue of S. cerevisiae ELO3. ELO3 gene is responsible for the elongation of VLCFAs (very long chain fatty acids) (Fang et al. 2017; Hong et al. 2019; Prasitchoke et al. 2007; Řezanka et al. 2018).
To see the clear classification and taxonomy of Hansenula (Ogataea) polymorpha and other related species, the phylogenetic relatedness was observed based on conserved gene sequences (Suh and Zhou 2010; Yoo et al. 2019). The phylogenetic analysis was done based on ribosomal gene sequences of ITS and D1/D2 region of LSU rRNA gene, and this molecular analysis revealed that most of the O. Polymorpha strains were different phylogenetically from type strain of Pichia angusta (ATCC 14755), Ogataea thermophila was found to be evolutionarily related to O. Polymorpha and two novel strains of Candida (ATCC 26012 and ATCC 58401) were close to O. polymorpha. The character-based method was applied to construct the phylogenetic tree. The maximum likelihood-based and parsimonious trees were constructed by taking sequences of ITS and D1/D2 region of LSU rRNA gene (>1 Kb). The result showed the close evolutionary relatedness of O. angusta and C. parapolymorpha with O. polymorpha along with some significant evolutionary distances in the tree constructed. A very closest matching and relatedness was observed in case of O. thermophila and O. polymorpha with 100% bootstrap value and therefore were grouped in the same clade (Suh and Zhou 2010).
The SSU (small subunit) and LSU (large subunit) ribosomal gene sequence-based study revealed that five different methylotrophic yeast species of Ogataea genus, that is, O. henricii, O. philodendra, O. glucozyma, O. minuta and O. polymorpha were reported earlier but a variety of Ogataea minuta var. nonfermentas was distantly related with genus Pichia (Kurtzman et al. 2008; Naumov et al. 2018; Yamada et al. 1994). Thereafter, O. minuta var. nonfermentas was reclassified separately as O. nonfermentas (Kurtzman and Robnett 2010). The multigene-based identification and phylogenetic analysis illustrated more than 37 species of methylotrophic yeast and recently 67 species of Ogataea genus were reported (Kurtzman 2009; Lu et al. 2017). The multigene analysis involved different types of genes for the study such as LSU rRNA, SSU rRNA, elongation factor EF-1α and mitochondrial SSU rRNA gene. The multigene analysis revealed differentiation among genera such as Pachysolen, Nakazawaea, Ambrosiozyma, Komagataella, Phaffomyces and Ogataea (Kurtzman and Robnett 2010). The pioneer of yeast molecular phylogeny, K.P. Kurtzman, described the phylogenetic classification of yeasts based on multiple genes and evolutionary relationship was observed.
In one of the investigations, a total of thirteen methylotrophic yeast strains utilising methanol as a carbon substrate (forming ascospore) were isolated and identified from the sap exudates of a tree Sclerobium sp. from Costa Rica and Brazil. Their characterisation for the identification and phylogenetic study was based on the sequence analysis of D1/D2 large subunit rDNA. The ribosomal gene sequence analysis and neighbour joining method of phylogenetic tree construction revealed their identification as predominant species of Ogataea genus (syn. Pichia) and Candida sp. Later, few new isolates were identified as Ogataea falcaomoraisii (Morais et al. 2004).
Three methylotrophic yeast strains were isolated from leaf and rotten wood samples of temperate forest in Hungary. The strains were found to be nitrate negative and assimilating methanol as a source of carbon. The D1/D2 large subunit rRNA gene sequence analysis grouped these strains in a clade of Ogataea sp. The strains have similar sequences and differ from genetically related and close species Pichia pilisensis. A novel methylotrophic yeast species Ogataea nitratoaversa was proposed to accommodate these nitrate-negative yeast strains. The variation in the D1/D2 and ITS sequences was observed due to several substitutions. Since the investigation does not allow the inclusion of nitrate-negative strains, they were named as Ogataea yamada, maeda and mikata (Péter et al. 2008).
In one of the earlier studies, two novel thermotolerant methylotrophic yeast strains were reported from soil and tree exudates from Thailand. The biochemical and phenotypic characterisation included the nitrate assimilation, hat-shaped ascospore formation, ubiquinone study, multilateral budding, urease reaction and other observations to identify the strains. The sequence analysis of D1/D2 rRNA gene revealed the phylogenetic relatedness between the species. The sequence analysis justifies two different strains PT44T and S051T. The PT44T strain was close to Pichia (Ogataea) dorogensis, whereas S051T strain was closely related to Pichia thermomethanolica with some nucleotide substitutions in the phylogenetic tree constructed. The biochemical, molecular, physiological and phenotypic characterisation of the strains proved them novel strains of genus Ogataea and further proposed with the name of Ogataea chonburiensis sp. nov. and Ogataea nakhonphanomensis sp. nov., respectively. Moreover, thermotolerant Pichia siamensis was renamed as Ogataea siamensis and Pichia thermomethanolica was renamed as Ogataea thermomethanolica in this study (Limtong et al. 2005).
3.3 Genetic Regulation in Yeast Methylotrophy
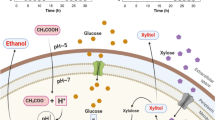
A number of methylotrophic yeast strains are reported and characterised by the identification of C1 carbon substrate-inducible gene expression analysis. For the yeast methylotrophy, different essential enzymes such as AOX and DAS and others are required to carry out formaldehyde oxidation metabolic pathway (Nakagawa et al. 1999; Sakai et al. 1998; Yurimoto et al. 2011). Molecular mechanism of methanol-inducible gene expression is represented during growth on various carbon sources (Fig. 3.2). Genes responsible for carbon substrate metabolism in methylotrophic yeast were investigated in C. boidinii by cloning the genes coding methanol-metabolising enzymes (Yurimoto et al. 2002). Figure 3.2 represents the methanol metabolism in methylotrophic yeasts (Fig. 3.3).
Molecular mechanism of methanol-inducible gene expression. (a) Relative expression levels of H. polymorpha MOX (encoding AOD), C. boidinii AOD1, and C. boidinii DAS1 during growth on various carbon sources. On glucose-containing media, expression is completely repressed. When glucose is completely consumed or cells are shifted to glycerol medium, a derepressed level of expression of the AOD genes is induced (derepression) and the extent of derepression of the AOD genes differs between H. polymorpha and C. boidinii. When cells are grown on methanol, the maximum level of expression of AOD genes is achieved not only by derepression but also by methanol-specific gene activation. The induction of DAS1 on methanol medium is achieved only by methanol-specific gene activation. (b) During growth on glucose, expression of methanol-inducible genes is repressed. When cells are shifted to methanol, initially, a Trm2p-related derepression event occurs followed by a Trm1p-related methanol-specific gene activation. (Adapted from Yurimoto et al. (2011))
Outline of methanol metabolism in methylotrophic yeasts. Enzymes: ADH (MFS), alcohol dehydrogenase (methyl formate-synthesising enzyme); AOD, alcohol oxidase; CTA, catalase; DAK, dihydroxyacetone kinase; DAS, dihydroxyacetone synthase; FDH, formate dehydrogenase; FGH, S-formylglutathione hydrolase; FLD, formaldehyde dehydrogenase; GLR, glutathione reductase; Pmp20, peroxisome membrane protein which has glutathione peroxidase activity. Abbreviations: DHA, dihydroxyacetone; DHAP, dihydroxyacetone phosphate; F6P, fructose 6-phosphate; FBP, fructose 1,6-bisphosphate; GAP, glyceraldehyde 3-phosphate; GS-CH2OH, S-hydroxymethyl glutathione; GS-CHO, S-formylglutathione; GSH, reduced form of glutathione; GSSG, oxidised form of glutathione; RCOOOH, alkyl hydroperoxide; Xu5P, xylulose 5-phosphate. (Adapted from Yurimoto et al. (2011))
For the heterologous protein expression and production, Pichia pastoris strain is generally preferred. In a study, the gene copy number was determined for P. pastoris along with real-time PCR assay for the quantification of integrated expression cassettes (Abad et al. 2010). In yeast methylotrophy, the expression of genes involved in methanol metabolism is regulated by the presence of carbon source. The expression and repression of genes were studied by Ohsawa et al. (2018) in which it was discussed that a significant and maximum gene expression was observed in the presence of methanol, whereas a low level of gene expression was observed in the absence of methanol (derepression). The characterisation and identification of various transcription factors involved in the expression and regulation of methanol-inducible gene expression was done by Ohsawa et al. (2018). Transcription factors KpMit1, leads to the repression of methanol-inducible gene expression or the presence of glucose leads to the repression of methanol-inducible gene expression (Hartner and Glieder 2006; Yamashita et al. 2009). Transcription factors such as KpMit1 in Komagataella phafi (Wang et al. 2016) and KpPrm1 in Candida boidinii (Sahu et al. 2014) along with Hap complex are involved in methanol induction, whereas transcription factors like KpMxr1 and CbTrm2 are involved in derepression (Lin-Cereghino et al. 2006). KpMxr1 and CbTrm2 are homologues to S. cerevisiae Adr1.
In a recent investigation, the function of a gene encoding Hac1 transcription factor was characterised in thermotolerant methylotrophic yeast Ogataea thermomethanolica TBRC656 (OtHAC1). Hac1 generally triggers the unfolded protein response pathway in yeasts. Under the characterisation study of this gene, the comparative proteomic analysis was done between OtHAC1 mutant and wild-type Ogataea strain. About 400–500 proteins were detected and gene encoding Hac1 annotated different functions involved in transcription, translation, oxidative stress and secretary pathway. Subsequently, two different novel OtHAC1-dependent proteins, viz. Iml1 and Npr2, were also identified which are responsible for the regulation of autophagy. This research on methylotrophic yeast therefore revealed the regulation of different metabolic pathways or processes through OtHAC1 gene expression in thermotolerant Ogataea thermomethanolica TBRC656 (Phithakrotchanakoon et al. 2018).
In Pichia pastoris , the regulation of AOX gene expression is done through repression/derepression mechanism along with induction mechanism and mostly this gene regulation resembles the regulation of GAL1 gene expression in Saccharomyces cerevisiae. The rich level of methanol in the media facilitates the high rate of transcription in case of methylotrophic yeast and the repression of gene regulation in the presence of glucose (a repressing carbon substrate) was not seen unlike GAL1 gene in case of Saccharomyces cerevisiae. The rate of gene expression was found directly proportional to the presence of methanol in the medium (Tschopp et al. 1987).
The methylotrophic yeast Ogataea thermomethanolica TBRC656 is a well-known host cell for the heterologous protein expression. In this thermotolerant methylotrophic yeast, maltase gene (mal) promoter-based new gene expression system was developed. The gene expression of xylanase and phytase was found to be enhanced many fold when Ogataea thermomethanolica TBRC656 was supplemented with sucrose in media. The increase in fold expression was due to activation of OtMal promoter gene as compared to constitutive OtGAP promoter gene. The presence of sucrose in the media also activates the more expression of OtMal promoter gene as compared to methanol-inducible OtAOX promoter. This enhances the enzyme activity by increasing higher gene expression of reporter genes coding xylanase and phytase. Therefore, it was suggested that methylotrophic yeast in the presence of sucrose as source of carbon substrate can be utilised for the production of heterologous proteins at large scale (Puseenam et al. 2018).
3.4 Methylotrophic Yeast and Impact to the Environments
The methylotrophic communities have been found to be applicable in diverse potential biotechnological applications (Fig. 3.4). The methylotrophic communities have been reported from diverse habitats including plant microbiomes as rhizospheric (Verma et al. 2013, 2014; Yadav 2017), endophytic (Rana et al. 2018; Verma et al. 2015a, 2016b) and epiphytic (Verma et al. 2015b, 2016a) and natural habitats as well as from different extreme environments of acidic, alkaline, drought, low temperature (Yadav 2015; Yadav et al. 2015a, b, 2016, 2019c), salinities and radiations (Yadav and Saxena 2018; Yadav et al. 2015c; Yadav and Yadav 2018). The methylotrophic communities having potential and efficient multifarious plant growth-promoting attributes have been used for plant growth promotion, crop yield, and soil health for sustainable agriculture (Biswas et al. 2018; Yadav et al. 2017; Yadav 2009). Along with agricultural application, the methylotrophic communities have been reported to use in different processes in medical, industrial and pharmaceutical sectors as well as in environment for sustainable future (Rastegari et al. 2019; Yadav et al. 2019a, b).
Biotechnological application of methylotrophic yeast. (Adapted with permission from Kuroda and Ueda (2011))
The genetically engineered thermotolerant methylotrophic yeast strains are reported to have properties of bioremediation (Kour et al. 2019; Suman et al. 2016; Yadav et al. 2018). Particularly chromate bioremediation was observed in Hansenula polymorpha which triggers the many fold enhanced gene expression of FCb2 gene coding flavocytochrome b2 enzyme as compared to parental strain (Smutok et al. 2011). The flavocytochrome b2 enzyme is known to be specific for L-lactate. In the presence of L-lactate, the enzyme flavocytochrome b2 leads to the reduction of chromate by living cells. Pichia pastoris is reported to be involved in the degradation of azo dyes and anthraquinone dyes and bioremediation of different xenobiotic compounds (Colao et al. 2006). In the production of fungal laccase, the expression of lccI gene was done in Pichia pastoris from cDNA synthesised from the white rot fungus Trametes togi.
Methylotrophic yeast isolated from oil-contaminated soil has distinct enzymatic activity and identified as Trichosporon cutaneum. Later, the identification was confirmed by ribosomal DNA-based molecular characterisation. In the isolate, methanol assimilation was found and can use formaldehyde also as a source of carbon substrate along with other carbon substrates like ethanol, glycerol, glucose and other petroleum derivatives. The microorganism was proved as an extremophile. In the isolate, different enzymatic activities were observed such as formaldehyde reductase, unspecific aldehyde dehydrogenase and formaldehyde dehydrogenase activity. Therefore, the biochemical, metabolic and physiological characteristics of methylotrophic isolate explore the new possibilities in the field of environmental biotechnology (Kaszycki et al. 2006).
Pichia pastoris is well-known yeast used in the production of animal contaminant-free hydroxylated collagen. The enzyme-based and molecular methods are utilised in the production and selection of triple-transformed Pichia pastoris strain, useful in the expression of P4H (prolyl 4-hydroxylase) tetramer obtained from marine sponge Chondrosia reniformis along with a hydroxylated collagen from the same animal (Pozzolini et al. 2015).
The environmental pollutants like household, industrial wastes and oil spills cost a lot to the sustainability of environment, and therefore some strains of methylotrophic yeast were found to be a better alternative for the bioremediation of these potent pollutants. The restoration of ecological balance is achieved by diminishing the level of environmental pollution. Methylotrophic yeast are able to degrade a number of xenobiotic compounds as a source of carbon. Methylotrophic strains such as Rhodosporidium, Pichia, Trichosporon, Rhodotorula and Yarrowia are able to degrade xenobiotic compounds like phenol, aromatic compounds and polar compounds. The degradation intensity decreases from n-alkanes to polar and aromatic carbon substrates (Csutak et al. 2010).
Genetically engineered methylotrophic yeast Hansenula polymorpha is known for the development of lactate-selective microbial biosensor. This thermotolerant yeast was utilised for the overproduction of lactate:cytochrome c-oxidoreductase enzyme system [FC b(2)] by overexpression of the CYB2 gene encoding FC b(2). The strong alcohol oxidase promoter of H. polymorpha controls the gene expression of HpCYB2 gene, and it was transformed into the host methylotrophic yeast strain H. polymorpha C-105 (gcr1 catX) in the absence of catalase activity and with glucose repression. Using a cathodic electrodeposition polymer, the cells are immobilised over graphite base. The redox mediator phenazine methosulphate used with this reacts with FC b (2) inside the cell in the presence of L-lactate. A higher Km value is observed in a biosensor based on recombinant methylotrophic yeast with a higher linear range towards lactate (Smutok et al. 2011).
Now it is very clear that methylotrophic yeast can be exploited as a suitable microbial culture used in the heavy-load wastewater treatment process. The pure monoculture of yeast can be utilised as a biofilter for the treatment of concentrated wastewater. The dilution of contaminants is done through degradation of wastes generated as a result of several technological processes. In earlier reports, it was illustrated that up to 10 g/l of formaldehyde can be diluted with the help of these biofilters. Moreover, methylotrophic yeast can be useful in the bioremediation of soil contaminated with oil (Kaszycki et al. 2006). Since methylotrophic yeast is well known for the production of heterologous proteins, they can change the product or compounds after processing due to genetic changes in their genome. Yeast cells do not contain pyogenes, pathogens, and viral inclusions and therefore can be used in the production of therapeutic administration. Earlier it was assumed that mainly S. cerevisiae was utilised for the production of pharmaceutical proteins but later methylotrophic yeast Hansenula polymorpha was used widely in modern genetics for the production of pharmaceuticals. Hansenula polymorpha possess especial advantageous characteristics as a host cell for the production of pharmaceuticals proteins. The pharmaceutical protein production system based on methylotrophic yeast Hansenula polymorpha has been established for different vaccines, serum proteins and other important therapeutics. In future, different products based on H. Polymorpha will be introduced to market after preclinical and clinical trials (Gellissen and Melber 1996).
3.5 Conclusion and Future Prospects
The molecular-level analysis enhances our understanding of methylotrophic yeast structure and function. The gene-level diversity and their study for the phylogenetic relatedness helped us to understand the establishment of methylotrophic yeast with the natural ecosystems. The methylotrophic yeasts are used in the research area and in biotechnological applications, one of the most important being the production of heterologous proteins of a large industrial and medical importance. The biotechnological production of heterologous proteins is reported from methylotrophic yeast Pichia pastoris which is renamed as Komagataella pastoris after recent taxonomy. The increase of SCP (Single Cell Protein) requirements or the remediations of the polluted systems by making use of natural alternatives represent important reasons for the necessity of characterising the methylotrophic yeasts. Moreover, further sophisticated and intensive research in the field of yeast methylotrophy and its molecular basis will explore and reveal new insights of physiological functions along with its importance in the natural ecosystem.
References
Abad S, Kitz K, Hörmann A, Schreiner U, Hartner FS, Glieder A (2010) Real-time PCR-based determination of gene copy numbers in Pichia pastoris. Biotechnol J 5:413–420
Ahmad M, Hirz M, Pichler H, Schwab H (2014) Protein expression in Pichia pastoris: recent achievements and perspectives for heterologous protein production. Appl Microbiol Biotechnol 98:5301–5317
Arthur H, Watson K (1976) Thermal adaptation in yeast: growth temperatures, membrane lipid, and cytochrome composition of psychrophilic, mesophilic, and thermophilic yeasts. J Bacteriol 128:56–68
Biswas S, Kundu D, Mazumdar S, Saha A, Majumdar B, Ghorai A, Ghosh D, Yadav A, Saxena A (2018) Study on the activity and diversity of bacteria in a New Gangetic alluvial soil (Eutrocrept) under rice-wheat-jute cropping system. J Environ Biol 39:379–386
Cereghino JL, Cregg JM (2000) Heterologous protein expression in the methylotrophic yeast Pichia pastoris. FEMS Microbiol Rev 24:45–66
Colao MC, Lupino S, Garzillo AM, Buonocore V, Ruzzi M (2006) Heterologous expression of lcc1 gene from Trametes trogii in Pichia pastoris and characterization of the recombinant enzyme. Microb Cell Factories 5:31
Craveri R, Cavazzoni V, Sarra P, Succi G, Molteni L, Cardini G, Di Fiore L (1976) Taxonomical examination and characterization of a methanol-utilizing yeast. Antonie Van Leeuwenhoek 42:533–540
Cregg JM, Madden K, Barringer K, Thill G, Stillman C (1989) Functional characterization of the two alcohol oxidase genes from the yeast Pichia pastoris. Mol Cell Biol 9:1316–1323
Cremata JA, Díaz JM (1999) Conventional and non-conventional yeasts in modern biotechnology. Biotecnol Apl 16:117–125
Csutak O, Stoica I, Ghindea R, Tanase A-M, Vassu T (2010) Insights on yeast bioremediation processes. Rom Biotechnol Lett 15:5066–5071
de Koning W, Harder W (1992) Methanol-utilizing yeasts. In: Murrell JC, Dalton H (eds) Methane and methanol utilizers. Springer US, Boston, pp 207–244. https://doi.org/10.1007/978-1-4899-2338-7_7
Fang Z, Chen Z, Wang S, Shi P, Shen Y, Zhang Y, Xiao J, Huang Z (2017) Overexpression of OLE1 enhances cytoplasmic membrane stability and confers resistance to cadmium in Saccharomyces cerevisiae. Appl Environ Microbiol 83:e02319–e02316
Gellissen G, Melber K (1996) Methylotrophic yeast hansenula polymorpha as production organism for recombinant pharmaceuticals. Arzneimittelforschung 46:943–948
Hartner FS, Glieder A (2006) Regulation of methanol utilisation pathway genes in yeasts. Microb Cell Factories 5:39
Hong J, Park S-H, Kim S, Kim S-W, Hahn J-S (2019) Efficient production of lycopene in Saccharomyces cerevisiae by enzyme engineering and increasing membrane flexibility and NAPDH production. Appl Microbiol Biotechnol 103:211–223
Kaszycki P, Koloczek H (2002) Biodegradation of formaldehyde and its derivatives in industrial wastewater with methylotrophic yeast Hansenula polymorpha and with the yeast-bioaugmented activated sludge. Biodegradation 13:91–99
Kaszycki P, Tyszka M, Malec P, Kołoczek H (2001) Formaldehyde and methanol biodegradation with the methylotrophic yeast Hansenula polymorpha. An application to real wastewater treatment. Biodegradation 12:169–177
Kaszycki P, Czechowska K, Petryszak P, Miedzobrodzki J, Pawlik B, Koloczek H (2006) Methylotrophic extremophilic yeast Trichosporon sp.: a soil-derived isolate with potential applications in environmental biotechnology. Acta Biochim Pol 53:463
Kour D, Rana KL, Yadav N, Yadav AN, Singh J, Rastegari AA, Saxena AK (2019) Agriculturally and industrially important fungi: current developments and potential biotechnological applications. In: Yadav AN, Singh S, Mishra S, Gupta A (eds) Recent advancement in white biotechnology through fungi, Volume 2: perspective for value-added products and environments. Springer International Publishing, Cham, pp 1–64. https://doi.org/10.1007/978-3-030-14846-1_1
Kumar M, Kour D, Yadav AN, Saxena R, Rai PK, Jyoti A, Tomar RS (2019) Biodiversity of methylotrophic microbial communities and their potential role in mitigation of abiotic stresses in plants. Biologia 74:287–308
Kurtzman CP (2009) Biotechnological strains of Komagataella (Pichia) pastoris are Komagataellaphaffii as determined from multigene sequence analysis. J Ind Microbiol Biotechnol 36:1435
Kurtzman CP, Robnett CJ (2010) Systematics of methanol assimilating yeasts and neighboring taxa from multigene sequence analysis and the proposal of Peterozyma gen. nov., a new member of the Saccharomycetales. FEMS Yeast Res 10:353–361
Kurtzman CP, Robnett CJ, Basehoar-Powers E (2008) Phylogenetic relationships among species of Pichia, Issatchenkia and Williopsis determined from multigene sequence analysis, and the proposal of Barnettozyma gen. nov., Lindnera gen. nov. and Wickerhamomyces gen. nov. FEMS Yeast Res 8:939–954
Kuroda K, Ueda M (2011) Cell surface engineering of yeast for applications in white biotechnology. Biotechnol Lett 33:1–9
Leão-Helder AN, Krikken AM, Van der Klei IJ, Kiel JA, Veenhuis M (2003) Transcriptional down-regulation of peroxisome numbers affects selective peroxisome degradation in Hansenula polymorpha. J Biol Chem 278:40749–40756
Limtong S, Srisuk N, Yongmanitchai W, Yurimoto H, Nakase T, Kato N (2005) Pichia thermomethanolica sp. nov., a novel thermotolerant, methylotrophic yeast isolated in Thailand. Int J Syst Evol Microbiol 55:2225–2229. https://doi.org/10.1099/ijs.0.63712-0
Limtong S, Srisuk N, Yongmanitchai W, Yurimoto H, Nakase T (2008) Ogataea chonburiensis sp. nov. and Ogataea nakhonphanomensis sp. nov., thermotolerant, methylotrophic yeast species isolated in Thailand, and transfer of Pichia siamensis and Pichia thermomethanolica to the genus Ogataea. Int J Syst Evol Microbiol 58:302–307
Lin-Cereghino GP, Godfrey L, Bernard J, Johnson S, Khuongsathiene S, Tolstorukov I, Yan M, Lin-Cereghino J, Veenhuis M, Subramani S (2006) Mxr1p, a key regulator of the methanol utilization pathway and peroxisomal genes in Pichia pastoris. Mol Cell Biol 26:883–897
Lu Y-F, Wang M, Zheng J, Hui F-L (2017) Ogataea neixiangensis sp. nov. and Ogataea paraovalis fa, sp. nov., two methanol-assimilating yeast species isolated from rotting wood. Int J Syst Evol Microbiol 67:3038–3042
Meena KK, Kumar M, Kalyuzhnaya MG, Yandigeri MS, Singh DP, Saxena AK, Arora DK (2012) Epiphytic pink-pigmented methylotrophic bacteria enhance germination and seedling growth of wheat (Triticum aestivum) by producing phytohormone. Antonie Van Leeuwenhoek 101:777–786
Mitsui R, Kusano Y, Yurimoto H, Sakai Y, Kato N, Tanaka M (2003) Formaldehyde fixation contributes to detoxification for growth of a nonmethylotroph, Burkholderia cepacia TM1, on vanillic acid. Appl Environ Microbiol 69:6128–6132
Morais PB, Teixeira LC, Bowles JM, Lachance M-A, Rosa CA (2004) Ogataea falcaomoraisii sp. nov., a sporogenous methylotrophic yeast from tree exudates. FEMS Yeast Res 5:81–85
Nakagawa T, Mukaiyama H, Yurimoto H, Sakai Y, Kato N (1999) Alcohol oxidase hybrid oligomers formed in vivo and in vitro. Yeast 15:1223–1230
Nakagawa T, Miyaji T, Yurimoto H, Sakai Y, Kato N, Tomizuka N (2000) A methylotrophic pathway participates in pectin utilization by Candida boidinii. Appl Environ Microbiol 66:4253–4257
Nakase T, Imanishi Y, Ninomiya S, Takashima M (2010) Candida rishirensis sp. nov., a novel methylotrophic anamorphic yeast species isolated from soil on Rishiri Island in Japan. J Gen Appl Microbiol 56:169–173
Naumov GI, Naumova ES, Lee C-F (2017) Ogataea haglerorum sp. nov., a novel member of the species complex, Ogataea (Hansenula) polymorpha. Int J Syst Evol Microbiol 67:2465–2469
Naumov G, Shalamitskiy MY, Naumova E, Lee C-F (2018) Phylogenetics, biogeography, and ecology of methylotrophic yeasts of the heterogeneous genus Ogataea: achivements and prospects. Microbiology 87:443–452
Negruţă O, Csutak O, Stoica I, Rusu E, Vassu T (2010) Methylotrophic yeasts: diversity and methanol metabolism. Rom Biotechnol Lett 15:5369–5375
Ohsawa S, Nishida S, Oku M, Sakai Y, Yurimoto H (2018) Ethanol represses the expression of methanol-inducible genes via acetyl-CoA synthesis in the yeast Komagataella phaffii. Sci Rep 8:18051
Okonechnikov K, Golosova O, Fursov M, Team U (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167
Olah GA (2005) Beyond oil and gas: the methanol economy. Angew Chem Int Ed 44:2636–2639
Péter G, Tornai-Lehoczki J, Dlauchy D (2008) Ogataea nitratoaversa sp. nov., a methylotrophic yeast species from temperate forest habitats. Antonie Van Leeuwenhoek 94:217
Phithakrotchanakoon C, Puseenam A, Phaonakrop N, Roytrakul S, Tanapongpipat S, Roongsawang N (2018) Hac1 function revealed by the protein expression profile of a OtHAC1 mutant of thermotolerant methylotrophic yeast Ogataea thermomethanolica. Mol Biol Rep 45:1311–1319
Pozzolini M, Scarfì S, Mussino F, Salis A, Damonte G, Benatti U, Giovine M (2015) Pichia pastoris production of a prolyl 4-hydroxylase derived from Chondrosia reniformis sponge: a new biotechnological tool for the recombinant production of marine collagen. J Biotechnol 208:28–36
Prasitchoke P, Kaneko Y, Bamba T, Fukusaki E, Kobayashi A, Harashima S (2007) Identification and characterization of a very long-chain fatty acid elongase gene in the methylotrophic yeast, Hansenula polymorpha. Gene 391:16–25
Puseenam A, Kocharin K, Tanapongpipat S, Eurwilaichitr L, Ingsriswang S, Roongsawang N (2018) A novel sucrose-based expression system for heterologous proteins expression in thermotolerant methylotrophic yeast Ogataea thermomethanolica. FEMS Microbiol Lett 365:238
Rana KL, Kour D, Yadav AN (2018) Endophytic microbiomes: biodiversity, ecological significance and biotechnological applications. Res J Biotechnol 14:1–30
Rastegari AA, Yadav AN, Gupta A (2019) Prospects of renewable bioprocessing in future energy systems. Springer International Publishing, Cham
Ravin NV, Eldarov MA, Kadnikov VV, Beletsky AV, Schneider J, Mardanova ES, Smekalova EM, Zvereva MI, Dontsova OA, Mardanov AV (2013) Genome sequence and analysis of methylotrophic yeast Hansenula polymorpha DL1. BMC Genomics 14:837
Řezanka T, Lukavský J, Vítová M, Nedbalová L, Sigler K (2018) Lipidomic analysis of Botryococcus (Trebouxiophyceae, Chlorophyta)-identification of lipid classes containing very long chain fatty acids by offline two-dimensional LC-tandem MS. Phytochemistry 148:29–38
Sahu U, Rao KK, Rangarajan PN (2014) Trm1p, a Zn (II) 2Cys6-type transcription factor, is essential for the transcriptional activation of genes of methanol utilization pathway, in Pichia pastoris. Biochem Biophys Res Commun 451:158–164
Sakai Y, Nakagawa T, Shimase M, Kato N (1998) Regulation and physiological role of theDAS1 gene, encoding dihydroxyacetone synthase, in the methylotrophic yeast Candida boidinii. J Bacteriol 180:5885–5890
Smutok O, Broda D, Smutok H, Dmytruk K, Gonchar M (2011) Chromate-reducing activity of Hansenula polymorpha recombinant cells over-producing flavocytochrome b2. Chemosphere 83:449–454
Suh S-O, Zhou JJ (2010) Methylotrophic yeasts near Ogataea (Hansenula) polymorpha: a proposal of Ogataea angusta comb. nov. and Candida parapolymorpha sp. nov. FEMS Yeast Res 10:631–638
Suman A, Yadav AN, Verma P (2016) Endophytic microbes in crops: diversity and beneficial impact for sustainable agriculture. In: Singh D, Abhilash P, Prabha R (eds) Microbial inoculants in sustainable agricultural productivity, research perspectives. Springer-Verlag, New Delhi, pp 117–143. https://doi.org/10.1007/978-81-322-2647-5_7
Tani Y, Yamada K (1987) Diversity in glycerol metabolism of methylotrophic yeasts. FEMS Microbiol Lett 40:151–153
Tschopp JF, Brust PF, Cregg JM, Stillman CA, Gingeras TR (1987) Expression of the lacZ gene from two methanol-regulated promoters in Pichia pastoris. Nucleic Acids Res 15:3859–3876
van der Klei IJ, Yurimoto H, Sakai Y, Veenhuis M (2006) The significance of peroxisomes in methanol metabolism in methylotrophic yeast. Biochim Biophys Acta 1763:1453–1462
Veenhuis M, Van Der Klei I, Titorenko V, Harder W (1992) Hansenula polymorpha: an attractive model organism for molecular studies of peroxisome biogenesis and function. FEMS Microbiol Lett 100:393–403
Verma P, Yadav AN, Kazy SK, Saxena AK, Suman A (2013) Elucidating the diversity and plant growth promoting attributes of wheat (Triticum aestivum) associated acidotolerant bacteria from southern hills zone of India. Natl J Life Sci 10:219–227
Verma P, Yadav AN, Kazy SK, Saxena AK, Suman A (2014) Evaluating the diversity and phylogeny of plant growth promoting bacteria associated with wheat (Triticum aestivum) growing in central zone of India. Int J Curr Microbiol App Sci 3:432–447
Verma P, Yadav AN, Khannam KS, Panjiar N, Kumar S, Saxena AK, Suman A (2015a) Assessment of genetic diversity and plant growth promoting attributes of psychrotolerant bacteria allied with wheat (Triticum aestivum) from the northern hills zone of India. Ann Microbiol 65:1885–1899
Verma P, Yadav AN, Shukla L, Saxena AK, Suman A (2015b) Alleviation of cold stress in wheat seedlings by Bacillus amyloliquefaciens IARI-HHS2-30, an endophytic psychrotolerant K-solubilizing bacterium from NW Indian Himalayas. Natl J Life Sci 12:105–110
Verma P, Yadav AN, Khannam KS, Kumar S, Saxena AK, Suman A (2016a) Molecular diversity and multifarious plant growth promoting attributes of Bacilli associated with wheat (Triticum aestivum L.) rhizosphere from six diverse agro-ecological zones of India. J Basic Microbiol 56:44–58
Verma P, Yadav AN, Khannam KS, Mishra S, Kumar S, Saxena AK, Suman A (2016b) Appraisal of diversity and functional attributes of thermotolerant wheat associated bacteria from the peninsular zone of India. Saudi J Biol Sci. https://doi.org/10.1016/j.sjbs.2016.01.042
Wang X, Wang Q, Wang J, Bai P, Shi L, Shen W, Zhou M, Zhou X, Zhang Y, Cai M (2016) Mit1 transcription factor mediates methanol signaling and regulates the alcohol oxidase 1 (AOX1) promoter in Pichia pastoris. J Biol Chem 291:6245–6261
Yadav AN (2009) Studies of Methylotrophic Community from the phyllosphere and rhizosphere of tropical crop plants. M.Sc. Thesis, Bundelkhand University, pp 66, https://doi.org/10.13140/2.1.5099.0888
Yadav AN (2015) Bacterial diversity of cold deserts and mining of genes for low temperature tolerance. Ph.D. Thesis, IARI, New Delhi/BIT, Ranchi pp 234, https://doi.org/10.13140/RG.2.1.2948.1283/2
Yadav AN (2017) Agriculturally important microbiomes: biodiversity and multifarious pgp attributes for amelioration of diverse abiotic stresses in crops for sustainable agriculture. Biomed J Sci Tech Res 1:1–4
Yadav AN, Saxena AK (2018) Biodiversity and biotechnological applications of halophilic microbes for sustainable agriculture. J Appl Biol Biotechnol 6:1–8
Yadav AN, Yadav N (2018) Stress-adaptive microbes for plant growth promotion and alleviation of drought stress in plants. Acta Sci Agric 2:85–88
Yadav AN, Sachan SG, Verma P, Saxena AK (2015a) Prospecting cold deserts of north western Himalayas for microbial diversity and plant growth promoting attributes. J Biosci Bioeng 119:683–693
Yadav AN, Sachan SG, Verma P, Tyagi SP, Kaushik R, Saxena AK (2015b) Culturable diversity and functional annotation of psychrotrophic bacteria from cold desert of Leh Ladakh (India). World J Microbiol Biotechnol 31:95–108
Yadav AN, Verma P, Kumar M, Pal KK, Dey R, Gupta A, Padaria JC, Gujar GT, Kumar S, Suman A, Prasanna R, Saxena AK (2015c) Diversity and phylogenetic profiling of niche-specific Bacilli from extreme environments of India. Ann Microbiol 65:611–629
Yadav AN, Sachan SG, Verma P, Saxena AK (2016) Bioprospecting of plant growth promoting psychrotrophic Bacilli from cold desert of north western Indian Himalayas. Indian J Exp Biol 54:142–150
Yadav A, Verma P, Kumar R, Kumar V, Kumar K (2017) Current applications and future prospects of eco-friendly microbes. EU Voice 3:21–22
Yadav AN, Verma P, Kumar V, Sangwan P, Mishra S, Panjiar N, Gupta VK, Saxena AK (2018) Biodiversity of the genus Penicillium in different habitats. In: Gupta VK, Rodriguez-Couto S (eds) New and future developments in microbial biotechnology and bioengineering, Penicillium system properties and applications. Elsevier, Amsterdam, pp 3–18. https://doi.org/10.1016/B978-0-444-63501-3.00001-6
Yadav AN, Mishra S, Singh S, Gupta A (2019a) Recent advancement in white biotechnology through fungi Volume 1: diversity and enzymes perspectives. Springer International Publishing, Cham
Yadav AN, Mishra S, Singh S, Gupta A (2019b) Recent advancement in white biotechnology through fungi. Volume 2: perspective for value-added products and environments. Springer International Publishing, Cham
Yadav AN, Yadav N, Sachan SG, Saxena AK (2019c) Biodiversity of psychrotrophic microbes and their biotechnological applications. J Appl Biol Biotechnol. Online first
Yamada Y, Maeda K, Mikata K (1994) The phylogenetic relationships of the hat-shaped ascospore-forming, nitrate-assimilating Pichia species, formerly classified in the genus Hansenula Sydow et Sydow, based on the partial sequences of 18S and 26S ribosomal RNAs (Saccharomycetaceae): the proposals of three new genera, Ogataea, Kuraishia, and Nakazawaea. Biosci Biotechnol Biochem 58:1245–1257
Yamashita S, Yurimoto H, Murakami D, Yoshikawa M, Oku M, Sakai Y (2009) Lag-phase autophagy in the methylotrophic yeast Pichia pastoris. Genes Cells 14:861–870
Yoo SJ, Moon HY, Kang HA (2019) Screening and selection of production strains: secretory protein expression and analysis in Hansenula polymorpha. In: Gasser B, Mattanovich D (eds) Recombinant protein production in yeast. Springer New York, New York, pp 133–151
Young EM, Comer AD, Huang H, Alper HS (2012) A molecular transporter engineering approach to improving xylose catabolism in Saccharomyces cerevisiae. Metab Eng 14:401–411
Yurimoto H, Sakai Y, Kato N (2002) Methanol metabolism. In: Gellissen G (ed) Hansenula polymorpha. https://doi.org/10.1002/3527602356.ch5
Yurimoto H, Oku M, Sakai Y (2011) Yeast methylotrophy: metabolism, gene regulation and peroxisome homeostasis. Int J Microbiol 2011:101298
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kumar, M. et al. (2019). Genetic Diversity of Methylotrophic Yeast and Their Impact on Environments. In: Yadav, A., Singh, S., Mishra, S., Gupta, A. (eds) Recent Advancement in White Biotechnology Through Fungi. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-030-25506-0_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-25506-0_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-25505-3
Online ISBN: 978-3-030-25506-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)