Abstract
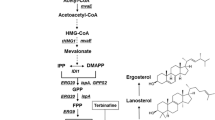
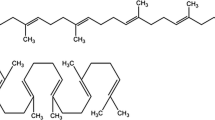
Squalene is a precursor in the eukaryotic sterol biosynthesis. It is a valuable compound with several human health-related applications. Since the traditional natural resources of squalene are limited, alternatives for the production of squalene on industrial scale have been intensively explored during past years. The yeast Saccharomyces cerevisiae represents an attractive option due to elaborated techniques of genetic and metabolic engineering that can be applied to improve squalene yields. We discuss in this chapter some theoretical aspects of genetic manipulations of the ergosterol biosynthesis pathway aimed at increased squalene production and describe analytical methods for squalene purification and determination of its content in yeast cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Spanova M, Daum G (2011) Squalene—biochemistry, molecular biology, process biotechnology, and applications. Eur J Lipid Sci Technol 113:1299–1320
Chang MH, Kim HJ, Jahng KY, Hong SC (2008) The isolation and characterization of Pseudozyma sp. JCC 207, a novel producer of squalene. Appl Microbiol Biotechnol 78:963–972
Daum G, Lees N, Bard M, Dickson R (1998) Biochemistry, cell biology and molecular biology of lipids of Saccharomyces cerevisiae. Yeast 14:1471–1510
Basson ME, Thorsness M, Rine J (1986) Saccharomyces cerevisiae contains 2 functional genes encoding 3-hydroxy-3-methylglutaryl coenzyme-A reductase. Proc Natl Acad Sci U S A 83:5563–5567
Burg JS, Espenshade PJ (2011) Regulation of HMG-CoA reductase in mammals and yeast. Prog Lipid Res 50:403–410
Wright R, Basson M, D'Ari L, Rine J (1988) Increased amounts of HMG-CoA reductase induce “karmellae”: a proliferation of stacked membrane pairs surrounding the yeast nucleus. J Cell Biol 107:101–114
Donald KA, Hampton RY, Fritz IB (1997) Effects of overproduction of the catalytic domain of 3-hydroxy-3-methylglutaryl coenzyme a reductase on squalene synthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 63:3341–3344
Polakowski T, Stahl U, Lang C (1998) Overexpression of a cytosolic hydroxymethylglutaryl-CoA reductase leads to squalene accumulation in yeast. Appl Microbiol Biotechnol 49:66–71
Asadollahi MA, Maury J, Schalk M, Clark A, Nielsen J (2010) Enhancement of farnesyl diphosphate pool as direct precursor of sesquiterpenes through metabolic engineering of the mevalonate pathway in Saccharomyces cerevisiae. Biotechnol Bioeng 106:86–96
Jandrositz A, Turnowsky F, Hogenauer G (1991) The gene encoding squalene epoxidase from Saccharomyces cerevisiae: cloning and characterization. Gene 107:155–160
Ryder NS (1992) Terbinafine: mode of action and properties of the squalene epoxidase inhibition. Br J Dermatol 126(Suppl):2–7
Naziri E, Mantzouridou F, Tsimidou MZ (2011) Enhanced squalene production by wild-type Saccharomyces cerevisiae strains using safe chemical means. J Agric Food Chem 59:9980–9989
Ta MT, Kapterian TS, Fei W, Du X, Brown AJ, Dawes IW, Yang H (2012) Accumulation of squalene is associated with the clustering of lipid droplets. FEBS J 279:4231–4244
Garaiová M, Zambojová V, Šimová Z, Griač P, Hapala I (2014) Squalene epoxidase as a target for manipulation of squalene levels in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 14:1567–1364
Drozdíková E, Garaiová M, Csáky Z, Obernauerová M, Hapala I (2015) Production of squalene by lactose‐fermenting yeast Kluyveromyces lactis with reduced squalene epoxidase activity. Lett Appl Microbiol 61:77–84
Klobučníková V, Kohút P, Leber R, Fuchsbichler S, Schweighofer N, Turnowsky F, Hapala I (2003) Terbinafine resistance in a pleiotropic yeast mutant is caused by a single point mutation in the ERG1 gene. Biochem Biophys Res Commun 309:666–671
Leber R, Fuchsbichler S, Klobučníková V, Schweighofer N, Pitters E, Wohlfarter K, Lederer M, Landl K, Hapala I, Turnowsky F (2003) Molecular mechanism of resistance to terbinafine in Saccharomyces cerevisiae. Antimicrob Agents Chemother 47:3890–3900
Ruckenstuhl C, Lang S, Poschenel A, Eidenberger A, Baral PK, Kohut P, Hapala I, Gruber K, Turnowsky F (2007) Characterization of squalene epoxidase of Saccharomyces cerevisiae applying terbinafine sensitive variants. Antimicrob Agents Chemother 51:275–284
Germann M, Gallo C, Donahue T, Shirzadi R, Stukey J, Lang S, Ruckenstuhl C, Oliaro-Bosso S, McDonough V, Turnowsky F, Balliano G, Nickels JT Jr (2005) Characterizing sterol defect suppressors uncovers a novel transcriptional signaling pathway regulating zymosterol biosynthesis. J Biol Chem 280:35904–35913
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Gietz RD, Schiestl RH, Willems AR, Woods RA (1995) Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11:355–360
Strathern JN, Higgins DR (1991) Recovery of plasmids from yeast into Escherichia coli: shuttle vectors. Methods Enzymol 194:319–329
Spanova M, Czabany T, Zellnig G, Leitner E, Hapala I, Daum G (2010) Effect of lipid particle biogenesis on the subcellular distribution of squalene in the yeast Saccharomyces cerevisiae. J Biol Chem 285:6127–6133
Acknowledgement
This work was supported by the Slovak Research and Development Agency grant APVV-0785-11 and VEGA 2/0185/14.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Valachovič, M., Hapala, I. (2017). Biosynthetic Approaches to Squalene Production: The Case of Yeast. In: Fox, C. (eds) Vaccine Adjuvants. Methods in Molecular Biology, vol 1494. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6445-1_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6445-1_7
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6443-7
Online ISBN: 978-1-4939-6445-1
eBook Packages: Springer Protocols