Abstract
L1-type proteins are transmembrane cell adhesion molecules with an evolutionary well-conserved protein domain structure of usually six immunoglobulin and five fibronectin type III domains. By engaging in many different protein–protein interactions they are involved in a multitude of molecular functions and are important players during the formation and maintenance of metazoan nervous systems. As a result, mutations in L1-type genes cause a great variety of phenotypes, most of which are neurological in nature. In humans, mutations in the L1CAM gene are responsible for L1 syndrome and other L1-type genes have been implicated in conditions as varied as mental retardation, autism, schizophrenia, multiple sclerosis, and other disorders. Equally, the overexpression of L1-type proteins appears to have deleterious effects in various types of human tumor cells, where they generally contribute to an increase in cell mobility and metastatic potential.
An erratum to this chapter is available at http://dx.doi.org/10.1007/978-1-4614-8090-7_15
An erratum to this chapter can be found at http://dx.doi.org/10.1007/978-1-4614-8090-7_15
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
L1-type proteins are transmembrane cell adhesion molecules (CAMs) and belong to the immunoglobulin superfamily (IgSF) (Moos et al. 1988; Hortsch 1996). Most L1-type proteins contain 13 distinct protein domains, usually six Ig (immunoglobulin) and three to five FN III (fibronectin type III) protein domains (see Fig. 9.1). L1-CAM was not only the first L1-type CAM to be identified, characterized, and cloned, but also yielded its name to the entire gene family (Rathjen and Rutishauser 1984; Rathjen and Schachner 1984; Moos et al. 1988).
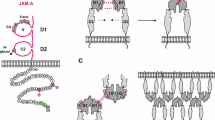
Vertebrate L1-type protein structures. This figure shows the protein domain structure of the four vertebrate L1-type proteins: L1-CAM, Neurofascin, CHL1, and NrCAM. Indicated are several specific protein sequence features, such as conserved integrin-binding RGD motifs, a basic protease target site (KR) in one of the FN III protein domains, the presence of a conserved cysteine residue at the end of the transmembrane segment, which is the target of palmitoylation modification (Ren and Bennett 1998), and the three conserved tyrosine-containing motifs in the cytoplasmic domain. The first two diagrams depict the two putative L1-CAM conformations, extended and horseshoe-shaped, which have been predicted for the L1-CAM ectodomain (Schürmann et al. 2001)
In this review we will focus on pathological mutations in L1-type genes, which have been described not only in humans, but also in a number of different experimental model systems. The phenotypes caused by L1 mutations reveal an amazingly complex picture reflecting a wide range of biological functions that are associated with L1-type proteins in various species and organs. Although our knowledge about the complex biological functionalities of L1-type proteins is still expanding, we will try to provide a timely overview about our current understanding, how this family of adhesive proteins plays crucial roles in the nervous and other organ systems, and how mutations in and also the overexpression of these proteins cause a variety of phenotypes.
2 Structure, Functions, and Genetics of L1-Type CAMs
2.1 The Structure of L1-Type Proteins
All L1-type proteins are predominantly, but not exclusively, expressed in the nervous system and belong to the immunoglobulin superfamily. They share a common arrangement of six amino terminal Ig-protein domains, followed by three to five FN III domains and a single transmembrane segment (Fig. 9.1). In humans, the mature L1-CAM protein has 1,256 amino acids with an extracellular part consisting of six Ig-like domains and five FN III-like domains, a single-pass transmembrane domain and a short cytoplasmic C-terminal tail (Wolff et al. 1988; Kobayashi et al. 1991). Genes encoding proteins with this characteristic domain structure form a unique gene family, now referred to as the L1 family of neural cell adhesion molecules (Hortsch 1996, 2000).
Gene duplication events in various metazoan phyla have resulted in multiple L1-type genes per genome (Mualla et al. 2013), and in most chordate species, including humans, four paralogous L1-type genes have been identified. These are now referred to as L1-CAM, CHL1 (Close Homolog of L1), Neurofascin, and NrCAM (neuron–glia-related cell adhesion molecule) (Fig. 9.1) (Hortsch 2000). In the case of Neurofascin and NrCAM proteins, alternative splicing of the initial transcript is responsible for multiple different protein isoforms (Hassel et al. 1997; Wang et al. 1998). The expression of the alternatively spliced Neurofascin protein isoforms is cell and tissue specific and also developmentally regulated (Hassel et al. 1997; Collinson et al. 1998). In some Neurofascin protein isoforms several of the FN III domains are either deleted or substituted by a PAT domain (Fig. 9.1) (Davis et al. 1993; Volkmer et al. 1992). This Neurofascin protein domain is rich in the amino acids proline, alanine, and threonine (thus termed “PAT”) and appears to be the target of O-linked glycosylation. These Neurofascin splice variants exhibit significant functional differences, not only in their interactions with various extracellular ligands (Volkmer et al. 1992), but also in their cell-specific expression and subcellular localization in neuronal cells (Davis et al. 1996; Zonta et al. 2008).
The Ig domains found in L1-type molecules were originally assigned to the C2 set of Ig-like domains. However, a comparison with other Ig domain proteins revealed that the domains in L1-type proteins belong to a novel structural subset of the Ig superfamily, now referred to as the I set (Harpaz and Chothia 1994; Bateman et al. 1996). Although the homophilic adhesive function of L1-type proteins involves multiple extracellular protein domains, it appears to be centered around the second Ig domain (Zhao et al. 1998). A number of vertebrate L1-type proteins (specifically L1-CAM and Neurofascin) also contain RGD motifs in their ectodomains, which functionally interact with RGD-specific integrins (Ruppert et al. 1995; Montgomery et al. 1996; Felding-Habermann et al. 1997; Yip et al. 1998; Koticha et al. 2005). Based on a general domain homology to the insect Ig domain protein Hemolin, Su et al. (1998) postulated that the 11 extracellular protein domains of L1-CAM exist in two different conformational states, one being extended and the other in a horseshoe shape (Fig. 9.1). Subsequently, structural analyses of the L1-CAM ectodomain gave some support to this notion (He et al. 2009; Schürmann et al. 2001; Wei and Ryu 2012). However, how these two postulated conformational states of the L1-CAM protein correlate with its functional interactions and activities remains unclear.
The size of the cytoplasmic domain in L1-type proteins ranges from 85 to 148 residues with several segments containing characteristic tyrosine-containing amino acid motifs that exhibit the highest degree of sequence conservation throughout the entire L1 gene family (Fig. 9.1). Two of these tyrosine-containing motifs are part of the cytoplasmic Ankyrin-binding site of L1-type proteins (Hortsch et al. 1998a; Zhang et al. 1998). The phosphorylation of the FIGQY motif is downstream of FGFR signaling and abolishes Ankyrin binding (Garver et al. 1997; Jenkins et al. 2001; Whittard et al. 2006). Interestingly, the entire L1 cytoplasmic domain is not required for homophilic L1–L1 interactions to occur (Hortsch et al. 1995; Wong et al. 1995). Nevertheless, extracellular and intracellular interactions involving L1-type proteins often influence and regulate each other (Hortsch et al. 1998a) and Ankyrin binding is important for a number of different L1 functions (Hortsch et al. 2009; Ooashi and Kamiguchi 2009; Guan and Maness 2010; Nakamura et al. 2010; Chen and Hing 2008; Buhusi et al. 2008; Ango et al. 2004; Nishimura et al. 2003).
Most vertebrate L1-type genes contain a well-conserved 12-nucleotide miniexon, which encodes an RSLE amino acid motif in the cytoplasmic L1 protein domain. Only in CHL1 proteins has the presence of an RSLE miniexon not been demonstrated. Due to differential splicing, these amino acids are not included in non-neuronal L1-type molecules (Reid and Hemperly 1992; Takeda et al. 1996). The insertion of this RSLE motif into the cytoplasmic domain of L1-type proteins generates a tyrosine-based signal (YxxL) that results in the sorting of L1-CAM protein to the growth cone and induces the AP-2-mediated endocytosis of L1-CAM and presumably other L1-type paralogous proteins via Clathrin-coated pits (Kamiguchi et al. 1998; Kamiguchi and Lemmon 1998). Non-vertebrate L1-type genes do not contain an RSLE-encoding miniexon. As reported for the Drosophila L1 molecule Neuroglian (Bieber et al. 1989), some of their transcripts undergo rather different, neuron-specific splicing processes (Hortsch et al. 1990), the functional ramifications of which are currently unknown. Therefore, the AP-2-mediated endocytosis of L1-type proteins, which regulates their cell surface expression, appears to be restricted to vertebrate species or non-vertebrate L1-type proteins contain other, yet unidentified, endocytosis-inducing signals.
2.2 The Evolutionary Origin of the L1 Gene Family
Figure 9.2 indicates that genes belonging to the L1 family of CAMs can be identified in most metazoan phyla and probably arose together with the appearance of first primitive nervous systems in evolution about 1,200–1,500 million years ago (Mualla et al. 2013). The genomes of several metazoan phyla, such as arthropods, only contain one L1-type gene, whereas other phyla, including most chordate species, appear to harbor multiple different L1-type genes in their genomes. With the exception of tunicates, all chordate genomes contain at least four L1-type paralogs, which are now known as L1-CAM, CHL1, Neurofascin, and NrCAM (Hortsch 2000). As shown in Fig. 9.1, these paralogous proteins all exhibit the typical L1-type protein structure and other L1 typical features. Consequently, these proteins share many similarities, including six Ig domains, three to five FN III domains, a transmembrane region, and a highly conserved cytoplasmic domain (Hortsch 2000). The existence of four paralogous genes in chordate species is now believed to be the result of two sequential genome-wide duplication events, which occurred during early chordate evolution (Kappen et al. 1989; Schughart et al. 1989). An additional genome-wide duplication event has occurred in the teleost lineage (Amores et al. 1998; Christoffels et al. 2004) resulting in two genes for each vertebrate L1 paralog for a total of eight L1-type genes per genome (Mualla et al. 2013).
Shown is an unrooted phylogenetic tree of the L1 family in several animal phyla. The species highlighted by gray ovals indicate L1-type paralogs that are only found in chordate species. Whereas many sequences were taken from complete open reading frames, which conform to the six Ig plus five FN III consensus L1 protein domain structure, others were translated from partial cDNA and genomic sequences that were downloaded from GenBank (http://www.ncbi.nlm.nih.gov/genbank/) or the JGI Genome Sequence Database (http://www.jgi.doe.gov). Sequences, which cover the region starting with the basic amino acids following the transmembrane segment to the conserved FIGQY motif, were used for a multiple sequence alignment. This alignment was performed using the online version of the MAFFT program (http://align.bmr.kyushu-u.ac.jp/mafft/online/server/). An unrooted phylogenetic tree was constructed using the Promlk and the Drawtree subroutines of the Phylip v3.65 program package (Felsenstein 1981). Species included in the figure represent 11 different metazoan phyla and include Schistosoma japonicum/blood-fluke or bilharzias flatworm (BU799663), Strongylocentrotus purpuratus/California purple sea urchin (XP_784933), Lottia gigantea/giant owl limpet (JGI Genome Sequence Database scaffold_226:99499–99582), Helobdella robusta/Californian leech (JGI Genome Sequence Database scaffold_43: 1086463:1087596, 1086816–1086905 and 1087447–1087596), Caenorhabditis elegans/nematode or roundworm (NP_001033395), Daphnia pulex/water flea (ACJG01004335), Tribolium castaneum/red flour beetle (AAJJ01000894), Drosophila melanogaster/fruit fly (NP_727274), Trichoplax adhaerens/hairy plate or flat animal (XM_002117949), Nematostella vectensis/starlet sea anemone (XM_001637356), Danio rerio/zebrafish (CABZ01014841, CABZ01052948, CABZ01022026, NM_001044805, and XM_002662518), Gallus gallus/chicken (AADN03013871 and AADN03006373), Monodelphis domestica/gray short-tailed opossum (AAFR03023046 and AAFR03020961), and Homo sapiens/human (NP_076493, NP_055905, NP_005001, and NP_006605)
2.3 Biological Functions of L1-Type CAMs
The identified biological functions of the L1-type CAMs cover a wide range and are mostly, but not exclusively, a result of their predominant expression in the nervous system (Hortsch 1996; Wiencken-Barger et al. 2004; Maness and Schachner 2007). Foremost, L1-CAM and other L1-type CAMs have been reported to induce neurite outgrowth (Lemmon et al. 1989; Harper et al. 1994; Hillenbrand et al. 1999; Volkmer et al. 1996; Pruss et al. 2004; Koticha et al. 2005) and to support axon guidance and pathfinding (Hall and Bieber 1997; Wiencken-Barger et al. 2004; Imondi et al. 2007; Demyanenko and Maness 2003); neuronal cell migration (Anderson et al. 2006; Demyanenko et al. 2001; Asou et al. 1992), axonal fasciculation (Wiencken-Barger et al. 2004), and neuronal differentiation (Dihne et al. 2003; Demyanenko et al. 2009; Turner et al. 2009); as well as cell survival (Nishimune et al. 2005; Hulley et al. 1998; Jakovcevski et al. 2009). L1-type proteins also play a prominent role during nervous system regeneration (Becker et al. 2004) and appear to be involved in synapse formation and plasticity (Godenschwege et al. 2006; Saghatelyan et al. 2004; Triana-Baltzer et al. 2006). Several studies also indicate that L1-type CAMs participate in the formation of the myelin sheath that surrounds many axons (Bartsch 2003; Itoh et al. 2005; Wood et al. 1990). These biological functions, which have been attributed to L1-type CAMs, make them promising pharmaceutical agents for aiding regeneration processes after spinal tract and other neuronal injuries. Several studies using animal models support this assertion (Chen et al. 2007; Becker et al. 2004; Roonprapunt et al. 2003; Bernreuther et al. 2006).
In arthropod species, the Neuroglian protein represents the product of the single L1-type gene that is present in the genome of species belonging to this phylum. Neuroglian has an additional essential function in stabilizing epithelial integrity as they are part of septate junction complexes (Genova and Fehon 2003; Laval et al. 2008; Wei et al. 2004; Faivre-Sarrailh et al. 2004; Banerjee et al. 2006). Although in vertebrate species the sealing function of epithelial septate junctions has been replaced by tight junctions, two L1-type proteins, Neurofascin and NrCAM, are still components of paranodal septate junctions where they stabilize the molecular architecture of nodes of Ranvier in myelinated nerves (Charles et al. 2002; Jenkins and Bennett 2002; Sherman et al. 2005; Koticha et al. 2006; Davis et al. 1996; Hortsch and Margolis 2003). In addition, L1-CAM is also expressed in many vertebrate epithelia outside the nervous system (Nolte et al. 1999). The physiological role of L1-CAM expression in these epithelia remains unknown, but the addition of anti-L1 antibodies to kidney organ cultures indicates a role in kidney branching morphogenesis (Debiec et al. 1998). Maddaluno et al. (2009) published another interesting finding about the function of L1-type proteins in epithelia. They reported that L1-CAM regulates the transendothelial trafficking of dendritic cells in mice.
Equally, the expression of L1-CAM protein in mammalian leukocytes still remains a mystery (Hubbe et al. 1993; Kowitz et al. 1992; Ebeling et al. 1996) and a topic of speculation (Kadmon et al. 1998). It has been suggested that L1-CAM functions as a co-stimulatory molecule during T-cell activation (Balaian et al. 2000). Another publication reports that a monoclonal antibody specific for L1-CAM disrupts the normal remodeling of lymph node reticular matrix during an immune response in vivo (Di Sciullo et al. 1998). Interestingly, the L1-type protein Neuroglian is also expressed in the moth Manduca sexta (tobacco hornworm) plasmatocytes where it contributes to the primitive innate immune functions, which these cells carry out by encapsulating foreign material (Williams 2009; Zhuang et al. 2007; Nardi et al. 2006).
2.4 L1 Syndrome: A Wide Spectrum of Phenotypes
The human L1CAM gene is located on the X-chromosome at Xq28 (Dietrich et al. 1992). It consists of 29 exons with the first exon of 125 base pairs being part of the 5′ untranslated region (Kallunki et al. 1997). Similar to other L1-type genes, the human L1CAM gene is primarily expressed in the nervous system and encodes a protein of 1,257 amino acids, comprising a signal peptide of 19 amino acids and a final processed product of 1,238 amino acids. In non-neural tissue, alternative RNA splicing generates mRNA molecules, which lack exons 2 and 27 of the gene (Jouet et al. 1995; De Angelis et al. 2001) and results in a protein that is 9 amino acids shorter.
Mutations in the human L1CAM gene manifest themselves in a wide range of dysfunctions that are usually neurological in origin and appearance. Therefore, the phenotype caused by mutations in the L1CAM gene was originally described as four distinct neurological disorders, namely, X-linked hydrocephalus, which is caused by a stenosis of the aqueduct of Sylvius (HSAS) (Rosenthal et al. 1992; Jouet et al. 1994; Bickers and Adams 1949; Finckh et al. 2000; Gu et al. 1996; Fransen et al. 1997), MASA (mental retardation, aphasia, shuffling gait, and adducted thumbs) syndrome (Winter et al. 1989; Schrander-Stumpel et al. 1990; Fryns et al. 1991), X-linked complicated hereditary spastic paraplegia type 1 (SPG1), and X-linked complicated corpus callosum agenesis (X-linked ACC) (Kaplan 1983). The allelic nature of these disorders remained unrecognized until the pathological mutations were identified as affecting the same gene (Fryns et al. 1991; Jouet et al. 1994; Fransen et al. 1994; Vits et al. 1994). Also the term CRASH syndrome (corpus callosum hypoplasia, retardation, adducted thumbs, spastic paraplegia, and hydrocephalus syndrome) has been proposed as a collective name for these disorders (Fransen et al. 1995). Now these terms are usually summarized under the name L1 syndrome (Panicker et al. 2003).
L1 syndrome is an X-linked recessive disorder with an incidence of 1:30,000 newborn males and is caused by mutations in the L1CAM gene. Well over 200 different pathogenic L1CAM mutations have been identified and reported in the literature. 130 were reviewed by Weller and Gartner (2001). Since then, many additional L1-CAM mutations were reported (Simonati et al. 2006; Senat et al. 2001; Sztriha et al. 2002; Silan et al. 2005; Felsenstein 1981; Okamoto et al. 2004; Hübner et al. 2004; Moya et al. 2002; Panayi et al. 2005; Tegay et al. 2007; Knops et al. 2008; Griseri et al. 2009; Nakakimura et al. 2008; Wilson et al. 2009; Kanemura et al. 2006; Piccione et al. 2010; Rodriguez Criado et al. 2003; Rehnberg et al. 2011) and another 52 new L1CAM mutations were recently reported in an online L1-CAM mutations databank (Vos and Hofstra 2010). As mentioned above, the individual phenotype associated with different mutations in the L1CAM gene varies considerably, but usually includes various degrees of mental retardation.
2.4.1 The Diversity of Pathogenic L1-CAM Mutations
The mutations affecting the L1-CAM protein have been subdivided into four different classes (Fransen et al. 1998b). Class I mutations lead to a truncation and thereby to a complete absence of L1-CAM protein. These include frameshift mutations (small deletions or insertions) or point mutations resulting in a premature stop codon (nonsense mutations). Class II includes missense mutations resulting in an amino acid substitution in the extracellular part of the L1-CAM protein. Class III includes any mutation in the L1-CAM cytoplasmic domain. Class IV mutations comprise extracellular mutations that result in an aberrant splicing of the L1-CAM pre-mRNA.
The different types of L1-CAM mutations generally correlate with the severity of the observed phenotype. Class 1 mutations in the extracellular part of L1-CAM cause a more severe phenotype; Class 2 extracellular missense mutations, which affect amino acids located on the surface of protein, usually cause a milder phenotype than those affecting amino acids, which are predicted to be buried in the core of an L1-CAM protein domain (Fransen et al. 1998b; Bateman et al. 1996). Mutations affecting the cytoplasmic domain of L1-CAM generally result in a milder phenotype than extracellular mutations (Fransen et al. 1998b; Yamasaki et al. 1997).
2.4.2 L1-CAM Missense Mutations and Functional Defects
In the past, a large number of L1-CAM missense mutations have been published that cause a wide spectrum of neurological abnormalities, including mental retardation, hydrocephalus, shuffling gait, and agenesis of corpus callosum (Finckh et al. 2000; Gu et al. 1996; Rosenthal et al. 1992; Fransen et al. 1995; Jouet et al. 1994; Vits et al. 1994, 1998; Ruiz et al. 1995). Interestingly, almost every family with an identified L1-CAM mutation has its own individual mutation. The pathogenic potential of L1-CAM missense mutations varies considerably and depends on the exact location of the affected residue and the type of amino acid exchange involved. Therefore, these L1-CAM mutations have also been classified as disease-causing, likely disease-causing, likely non-disease-causing polymorphisms, or as unknown (Vos et al. 2010).
L1-CAM missense mutations have been identified in all extracellular protein domains, as well as the cytoplasmic region of the L1-CAM protein (Fig. 9.3). Specifically, disease-causing mutations have been reported in Ig1, Ig2, Ig3, Ig4, and Ig5 and Fn1, Fn2, and Fn5 protein domains of extracellular region of the L1-CAM and also in the cytoplasmic domain. In addition, likely disease-causing mutations have been identified in the L1-CAM Ig6 and Fn1 protein domains. A more complete list of currently known and previously published human L1-CAM mutations can be accessed at the “L1CAM Mutation Database,” which is being maintained by the University Medical Center Groningen at http://www.l1cammutationdatabase.info (Vos and Hofstra 2010).
Selection of human pathogenic L1-CAM missense mutations. This figure depicts the position of several pathogenic L1-CAM mutations that have been determined to cause L1 syndrome in humans and that have been experimentally analyzed in various in vitro and in vivo assay systems for their ability to perform specific L1-CAM-associated functions (De Angelis et al. 1999; Nagaraj et al. 2009; Rünker et al. 2003; Michelson et al. 2002; Needham et al. 2001; Moulding et al. 2000; Zhao and Siu 1996; Godenschwege et al. 2006)
However, the severity of the phenotype and the clinical features vary between different L1-CAM mutations. Certainly the location of the mutated amino acid residue and the affected protein domain (Ig, FN III, or cytoplasmic) do influence and partially determine the severity of the phenotype. This has been analyzed for the severity of L1-CAM-related hydrocephalus and the mortality rate caused by individual L1-CAM mutations (Yamasaki et al. 1997; Bertolin et al. 2010). In general, mutations in the extracellular part of L1-CAM that led to truncation or absence of L1-CAM protein cause a most severe phenotype. In contrast, L1-CAM mutations affecting the cytoplasmic region result in a milder phenotype (Fransen et al. 1998b; Yamasaki et al. 1997). In addition, extracellular mutations affecting amino acids situated on the surface of a domain cause a milder phenotype than those affecting amino acids situated in the core of the protein domains (Bateman et al. 1996; Fransen et al. 1998b). Equally, L1-CAM mutations interfering with homophilic and/or heterophilic protein–protein interactions usually cause more significant neuronal dysfunctions (De Angelis et al. 2002). In addition, the variance of the phenotype between affected siblings, who share an identical L1-CAM mutation, suggests a strong epigenetic influence on the expression of specific phenotypic aspects. Using an L1-CAM knockout mouse model, Tapanes-Castillo et al. (2009) identified such a modifier locus for X-linked hydrocephalus on mouse chromosome 5.
2.5 Molecular Mechanisms by Which L1-CAM Mutations Cause Neurological Dysfunctions
The phenotypic diversity that is observed for L1-CAM mutations is well documented and it has been speculated that the multitude of L1-CAM interactions with itself and its various binding partners may play an important role in this heterogeneity. Many extracellular L1-CAM mutations have been demonstrated to disrupt L1-CAM’s homophilic interaction and have a reduced ability to stimulate neurite outgrowth in vitro (De Angelis et al. 1999; Zhao and Siu 1996) (Fig. 9.3). As L1-CAM exerts many of its physiological functions by its strong homophilic adhesive ability, the relative contribution of homophilic versus heterophilic L1-CAM interactions to the phenotypic expression of mutational defects remains an interesting question. Regardless, it appears reasonable to assume that alterations in ligand-specific L1-CAM binding properties represent one central pathological mechanism, which is at play in L1 syndrome (De Angelis et al. 2002).
As evidenced by two studies that used a transgenic mouse model expressing L1-6D, which lacks the sixth Ig domain of L1-CAM and as a result has no homophilic and RGD-dependent integrin interactions, the homophilic adhesive function of L1-CAM is involved in some phenotypic aspects of L1 syndrome, specifically hydrocephalus. However, its severity is also influenced by other genetic modifiers (Tapanes-Castillo et al. 2009; Itoh et al. 2004). Based on their observations these authors hypothesize that a co-receptor for L1-CAM-mediated neurite outgrowth is involved and some pathogenic mutations affect neurite outgrowth or branching by disrupting the interaction with this co-receptor.
Not surprisingly, some pathogenic L1-CAM mutations also disrupt some of L1-CAM’s known heterophilic interactions. For example, many pathological L1-CAM mutations influence the heterophilic interaction with TAX-1/Axonin-1 (De Angelis et al. 1999, 2002) (Fig. 9.3). Similarly, the functional interaction between L1-CAM and EGFR (epidermal growth factor receptor) interaction is impaired by some L1-CAM missense mutations (Nagaraj et al. 2009). In the cytoplasmic domain, two pathogenic L1-CAM mutations (S1124L and Y1229H, Fig. 9.3) affect the binding of the L1-CAM cytoplasmic domain to the Spectrin–Actin cytoskeleton and abolish endocytosis of L1-CAM (Buhusi et al. 2008; Needham et al. 2001). It appears reasonable to hypothesize that other heterophilic L1-CAM interactions, such as the binding of Neuropilin-1 that partially mediates L1-CAM’s axon guidance function (Soker et al. 1998; Castellani et al. 2000) and the interaction with RGD-specific integrins that leads to cell migration and myelination (Haney et al. 1999; Mechtersheimer et al. 2001), are equally changed by some pathogenic L1-CAM mutations. Finally, some mutations appear to influence the tertiary structure of the L1-CAM protein, which in turn disrupts protein–protein interactions involving L1-CAM indirectly (Cheng and Lemmon 2004).
Another mechanism by which mutations influence or abrogate L1-CAM function is their effect on the cell surface expression of the L1-CAM protein (De Angelis et al. 1999, 2002; Michelson et al. 2002; Moulding et al. 2000). Particularly, the pathological missense mutations I179S and Y194C affect L1-CAM’s neurite outgrowth inducing capability by decreasing the cell surface localization of the mutant L1-CAM protein (Michelson et al. 2002) (Fig. 9.3). The missense mutation C264Y in the extracellular domain of L1-CAM, which is known to cause HSAS (hydrocephalus with stenosis of aqueduct of Sylvius) in affected humans, impairs cellular L1-CAM protein trafficking (Rünker et al. 2003) (Fig. 9.3). Transfection studies in vitro demonstrate that this mutant L1-CAM protein is not expressed at the cell surface, but instead is located intracellularly, most likely within the endoplasmic reticulum. This was further confirmed by an in vivo analysis using a transgenic mouse line that expresses the C264Y mutant L1-CAM protein (Rünker et al. 2003).
There are also multiple lines of evidence that pathogenic L1-CAM mutations alter interactions with the intracellular Actin–Spectrin membrane skeleton and the cytoplasmic machinery that regulates L1-CAM-mediated axonal branching. The L1-CAM cytoplasmic domain (L1CD) is involved in axonal branching and the interaction between L1-CAM and the Actin cytoskeleton is also critical for this activity (Cheng et al. 2005a). Some extracellular mutations, such as I219T and W1036L (Fig. 9.3), alter the interaction between L1-CAM and the Actin cytoskeleton by changing L1-CAM’s conformation. This observation might simply be a reflection of extracellular adhesive events regulating the binding of Ankyrin to the cytoplasmic L1 domain (Dubreuil et al. 1996; Hortsch et al. 1998b). Ankyrin acts as the linker between the L1 cytoplasmic domain and the Actin–Spectrin network. Alternatively, Cheng et al. speculated that axonal branching is regulated by L1-CAM interacting extracellularly in cis with an unknown co-receptor. The pathogenic mutations I219T and W1036L may disrupt this interaction. The S542P mutation (Fig. 9.3), which exhibits a reduction in both L1-CAM protein surface expression and homophilic adhesion, also elicits a decrease in the number of axonal branching (Cheng and Lemmon 2004).
Some pathogenic L1-CAM mutations have also been reported to affect L1-CAM-associated signaling processes. Using in vitro as well as in vivo Drosophila assay systems, we demonstrated that two pathogenic human L1-CAM mutations (E309K and Y1070C, Fig. 9.3), which exhibit wild-type levels of homophilic adhesion, have a reduced ability to induce L1-CAM-dependent EGFR signaling in vitro and are unable to rescue L1 loss-of-function conditions in vivo (Nagaraj et al. 2009).
Pathogenic L1-CAM mutations have various effects on the L1-CAM protein. Many missense L1-CAM mutations are predicted to distort the structure of individual domains and as a result to affect the intracellular processing of the L1-CAM protein and potentially reduce its cell surface expression. A recent study by Bertolin et al. (2010) demonstrated that some frameshift mutations and all nonsense mutations result in truncated L1-CAM proteins, which have carboxy termini in different extracellular L1-CAM protein domains. The observed neurological dysfunctions that are associated with these L1-CAM mutations give support to the notion that the severity of the L1 syndrome phenotype correlates with the severity of the molecular effect of the individual mutation and is also dependent on the epigenetic context into which a particular mutation is placed (Yamasaki et al. 1997).
2.6 Mutational Defects of L1-Type Genes in Various Model Animal Systems
2.6.1 Drosophila melanogaster
L1-type genes and proteins have been identified in many other metazoan species (Mualla et al. 2013). Drosophila Neuroglian (Nrg) was the second L1-type protein identified and described (Bieber et al. 1989). Coincidentally, the neuroglian (nrg) gene is localized on the fly’s X-chromosome at cytological location 7F1. Hall and Bieber (Hall and Bieber 1997) have analyzed and described three mutant lines. All are late embryonic lethal mutations that alter or abolish Drosophila Neuroglian expression during development. The nrg 1 mutation is a protein null allele that is caused by an inversion with breakpoints at chromosomal locations 6E-7F1 the proximal breakpoint residing in the nrg transcription unit (Bieber et al. 1989; Hall and Bieber 1997). The mutation nrg 2 represents a hypomorphic mutation with markedly reduced expression levels of both Neuroglian protein isoforms. The mutation nrg3 is temperature sensitive and also represents a late embryonic lethal allele when raised at a nonpermissive temperature (Hall and Bieber 1997). At a nonpermissive temperature the Nrg protein in homozygous nrg 3-mutant animals is mislocalized inside Nrg expressing cells and is not transported to the cell surface.
Neuroglian is also expressed in two protein isoforms, one being restricted to neuronal cells (Nrg180) and the other to non-neuronal cells (Nrg167) (Hortsch et al. 1990). However, the differential cell-specific splicing of the nrg transcript in Drosophila differs considerably in its effect on the resulting L1 protein isoforms from that described for L1-CAM in vertebrates (De Angelis et al. 2001; Miura et al. 1991). Although the Neuroglian protein is predominantly expressed in the nervous system throughout the life cycle of the fly (Bieber et al. 1989), the non-neuronal Nrg167 isoform also exhibits a high level of expression in most, if not all, Drosophila epithelia. In arthropod epithelia Neuroglian is an essential part of septate junction protein complexes and thereby stabilizes epithelial integrity (Genova and Fehon 2003; Laval et al. 2008; Wei et al. 2004; Faivre-Sarrailh et al. 2004; Banerjee et al. 2006).
The lack of Neuroglian expression in vivo causes abnormalities in embryonic motor neuron projections of both the intersegmental and the segmental nerves (Hall and Bieber 1997). Temperature shift experiments using homozygous nrg3-mutant animals also revealed axonal pathfinding defects in the larval ocellar sensory system, which are mediated by EGFR and FGFR (fibroblast growth factor receptor) signaling (Garcia-Alonso et al. 2000; Kristiansen et al. 2005). Similarly, Neuroglian appears to mediate sensory axon advances in the Drosophila embryonic nervous system (Martin et al. 2008). Neuroglian is also an important player during the development of the adult mushroom body where it controls axonogenesis, axon bundling, axon branching, and guidance through signaling mechanisms that are different from the ocellar sensory system (Goossens et al. 2011). However, the physiological role of the fly’s sole L1-type protein is not restricted to neuronal cells and to axonal pathfinding. It is also involved in dendritic arborization (Yamamoto et al. 2006) and in the proper differentiation of certain glial cells (Chen and Hing 2008). In addition, by using a S213L Neuroglian missense mutation, Godenschwege et al. uncovered an essential role of this Drosophila L1-type protein at the pupal giant synapse, which is independent of Neuroglian’s role in axonal pathfinding (Godenschwege et al. 2006). A different Neuroglian missense mutation (G92R, designated as ibx for icebox) not only causes a nonlethal central brain morphology phenotype, but when homozygous in female flies results in a specific deficiency in female mating behavior (Carhan et al. 2005). Male flies show no behavioral defects, nor are other female behaviors visibly affected.
Although the Drosophila Neuroglian protein only exhibits a moderate level of amino acid identity when compared with mouse or human L1-CAM (Bieber et al. 1989; Zhao and Hortsch 1998), all major L1 features, including the characteristic L1 protein domain structure and a highly conserved intracellular Ankyrin binding domain, are preserved in the fly ortholog. In contrast to vertebrate species, arthropod genomes have only one L1-type gene in their genome. However, the similarities between human L1-CAM and Drosophila Neuroglian extend to several functional aspects. The expression of human L1-CAM in transgenic flies rescues some of nrg loss-of-function axonal pathfinding defects (Kristiansen et al. 2005). Similarly, human L1-CAM expression rescues a central nervous system synaptic phenotype in the fly that is caused by the lack of Neuroglian protein (Godenschwege et al. 2006). This surprising functional conservation between two members of the L1 gene family that are separated by more than 600 million years of evolution has made it possible to analyze pathogenic human L1-CAM proteins under in vivo conditions for specific functional aspects (Godenschwege et al. 2006; Nagaraj et al. 2009).
2.6.2 Caenorhabditis elegans
The nematode Caenorhabditis elegans genome encodes two L1-type genes, sax-7/lad-1 and lad-2 (Chen et al. 2001; Wang et al. 2008), which appear to serve different, nonoverlapping functions (Chen and Zhou 2010). The SAX-7/LAD-1 protein has all the hallmarks of L1-type proteins (Chen et al. 2001) and when mutated or deleted causes a pleiotropic phenotype, which includes embryonic and gonadal malformations (Chen et al. 2001), the misorganization of ganglia and abnormal positioning of neuronal cells in the adult (Sasakura et al. 2005), as well as embryonic lethality, inappropriate axon trajectories, and uncoordinated movements (Wang et al. 2005). In contrast, the LAD-2 protein, which is expressed in only a subset of nematode neurons, has a traditional L1-type ectodomain, but is truncated at its cytoplasmic tail and is missing the Ankyrin-binding domain (Wang et al. 2008). Animals that are mutant for the lad-2 gene primarily exhibit axonal pathfinding defects. These defects are caused when the LAD-2 protein is unable to fulfill its function as a co-receptor together with PLX-2/Plexin to bind MAB-20/Sema 2 protein (Wang et al. 2008).
2.6.3 Mus musculus
In mice, knockouts of the L1CAM gene (L1-KO mice) exhibit a phenotype similar to that observed in humans with L1 syndrome. These phenotypic aspects include a reduced corticospinal tract, abnormal pyramidal decussation, decreased axonal association with non-myelinating Schwann cells, ventricular dilation, and hypoplasia of the cerebellar vermis (Itoh et al. 2004; Dahme et al. 1997; Fransen et al. 1998a; Demyanenko et al. 1999, 2001). Using a co-culture in vitro assay with cells isolated from an L1-CAM-deficient mouse line, Castellani et al. revealed a role for L1-CAM in the Sema3A signaling pathway of axonal guidance (Castellani et al. 2000). Their finding suggests that some L1-CAM mutations may also disrupt Sema3A’s chemorepulsive signaling activity in the growth cone (Castellani et al. 2000). This situation is reminiscent of the phenotype that has been observed for the L1-type protein LAD-2 in nematodes (Wang et al. 2008).
2.7 L1-CAM Expression in Cancer Cells: The Multidimensional Nature of L1-CAM Function
2.7.1 Expression of L1-CAM in Various Human Tumors
As described above, L1-CAM and other vertebrate L1-type proteins are primarily expressed in the nervous system (Rathjen and Schachner 1984; Sanes et al. 1986). Not surprisingly, L1-CAM expression has been reported in a number of human tumors of neuroectodermal origin, specifically gliomas and neuroblastomas (Figarella-Branger et al. 1990; Izumoto et al. 1996; Tsuzuki et al. 1998). L1-CAM expression in gliomas correlates with increased tumor invasion (Izumoto et al. 1996) and the downregulation of L1-CAM expression decreases glioma growth in a mouse model (Bao et al. 2008). Whereas L1-CAM expression in adult gliomas is associated with reduced survival, the opposite was reported for pediatric gliomas (Wachowiak et al. 2007). This suggests that L1-CAM expression has different effects in different types of neuroectodermal tumors.
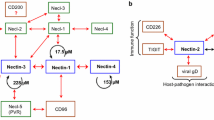
Some recent publications have reported an aberrant expression of L1-CAM in several non-neuronal types of human cancer (Gavert et al. 2008). For example, L1-CAM protein is often expressed during advanced stages of colon cancer development (Raveh et al. 2009; Gavert et al. 2005). In addition, L1-CAM was found in certain ovarian cancers (Zecchini et al. 2008), renal cell carcinoma (Allory et al. 2005), human cutaneous malignant melanoma (Thies et al. 2002; Fogel et al. 2003; Linnemann et al. 1989), pancreatic adenocarcinomas (Sebens Müerköster et al. 2007), breast cancers (Valladares et al. 2006; Shtutman et al. 2006; Gutwein et al. 2000), and various lung cancer cell lines (Katayama et al. 1997). This suggests that the expression of L1-type proteins might play an important role in the development and progression of different types of cancers. The study by Gavert et al. also demonstrates that when transfected into LS174T human colon carcinoma cells, L1-CAM expression results in an increased growth rate and cell motility and in an enhancement of the cells’ tumorigenic capacity (Gavert et al. 2008). When injected into the spleen of nude mice, L1-CAM-transfected LS174T cells gained the ability to form liver metastases (Gavert et al. 2007). In colon cancer tissue samples, L1-CAM expression is correlated with higher levels of nuclear β-catenin and these cells are exclusively localized at the invasive front of the tumor tissue (Gavert et al. 2005). The authors provide ample evidence that in human colon carcinoma cells, L1-CAM is a target of aberrantly activated β-catenin–TCF signaling and increases their metastatic potential (Gavert et al. 2008).
Similar to colon carcinoma, L1-CAM has been detected at the invasive front of epithelial ovarian carcinoma tissue cells and its expression is associated with a poor clinical prognosis and increased levels of metastasis (Zecchini et al. 2008). Also in renal cell carcinoma (RCC), the presence of L1-CAM is linked to a propensity to develop metastasis, and when coupled with the loss of Cyclin D1 expression, L1-CAM was defined as an independent prognostic factor for metastasis occurrence in a multivariate analysis (Allory et al. 2005).
Other studies show that L1-CAM is also expressed in a significant portion of various histological subtypes of human cutaneous malignant melanomas. Again it is a good predictor for metastatic ability and indicates a poor prognosis in melanoma patients (Thies et al. 2002; Fogel et al. 2003; Linnemann et al. 1989). Furthermore, the downregulation of L1-CAM expression reduces the migration and invasiveness of metastatic B16 cells in vitro (Meier et al. 2006). A gene array analysis of malignant melanoma tissues demonstrated that L1-CAM RNA is expressed at more than tenfold higher levels when compared with benign lesions or normal skin samples (Talantov et al. 2005). L1-CAM expression has also been reported in neuroendocrine carcinomas of the skin (Deichmann et al. 2003).
For pancreatic adenocarcinomas, one study reported that L1-CAM was expressed in 80 % (16 of 20 samples) of all tissue sections analyzed (Sebens Müerköster et al. 2007). However, other studies reported L1-CAM expression to be less common in pancreatic adenocarcinoma (2 of 111 samples) (Kaifi et al. 2006a) and 5 out of 63 pancreatic neuroendocrine tumors (Kaifi et al. 2006b). Again L1-CAM expression in pancreatic adenocarcinomas is a valid indicator for a poor clinical prognosis (Tsutsumi et al. 2011; Chen et al. 2011). L1-CAM is also detectable in some breast cancers (Valladares et al. 2006; Shtutman et al. 2006; Gutwein et al. 2000). In MCF7 breast cancer cells, L1-CAM expression disrupts E-cadherin-containing adherens junctions and increases cell scattering and motility (Shtutman et al. 2006). This correlates with the observation that L1-CAM expression is more abundant in metastatic breast cancer cells and lower in non-metastatic breast cancer cells (Valladares et al. 2006).
L1-CAM and Neurofascin proteins both contain an RGD motif in their sixth Ig or third FN III domain, respectively, both of which have been shown to interact with RGD-specific integrins (Ruppert et al. 1995; Montgomery et al. 1996; Felding-Habermann et al. 1997; Blaess et al. 1998). The RGD motif in the sixth L1-CAM Ig domain appears to increase the incident of metastasis formation in some tumor cells expressing L1-CAM. An investigation by Duczmal et al. showed that the L1-CAM-mediated migration of human MED-B1 tumor cells is RGD dependent and can be blocked by αγβ3 integrin-specific antibodies (Duczmal et al. 1997). The evidence that RGD motifs in Neurofascins and L1-CAMs can serve as ligands for integrins has led researchers to argue for a functional significance of this interaction in L1-mediated tumor progression (Duczmal et al. 1997; Montgomery et al. 1996).
2.7.2 L1-Type Protein Expression Affects Signaling Pathways in Cancer Cells
L1-CAM and other vertebrate L1-type proteins play an active role in cancer development by affecting different signaling pathways and thereby contributing directly to tumor progression. Several groups have demonstrated that in ovarian cancer cells L1-CAM-dependent ERK activation is associated with increased FGFR, EGFR, and hepatocyte growth factor receptor (HGFR) activity (Zecchini et al. 2008; Stoeck et al. 2007; Novak-Hofer et al. 2008). These and a number of other studies suggest that L1-CAM plays a role in carcinogenesis by its ability to activate the ERK signaling pathway (Gast et al. 2008; Schaefer et al. 1999; Silletti et al. 2004; Schmid et al. 2000). L1-CAM itself is also phosphorylated by Erk2 and interacts with different components of the ERK pathway (Silletti et al. 2004; Schmid et al. 2000). This ultimately results in the expression of proteins that contribute to cell motility and cell invasion. In addition, L1-CAM-mediated ERK activation was shown to involve Src (Gast et al. 2008; Silletti et al. 2004). The reported association of the L1-CAM cytoplasmic domain with RamBPM also indicates a direct link between L1-CAM expression and the MAPK/ERK signaling pathway (Cheng et al. 2005b).
The co-expression of ADAM10 (A Disintegrin and Metalloproteinase domain-containing protein 10) and L1-CAM in invasive colon cancer tumor cell indicates that the proteolytic processing of L1-CAM may have a role in invasive tumor development (Gavert et al. 2007). The L1-CAM protein is often cleaved by MMPs and the shedded L1-CAM ectodomain can interact with integrins, RTKs, or L1-CAM on the surface of the same or of neighboring cells (Gavert et al. 2007). Similarly, the ectodomain of the Nr-CAM protein is cleaved by matrix metalloproteinases, which results in an enhancement of cell motility, proliferation, ERK and AKT activation, and ultimately oncogenesis (Conacci-Sorrell et al. 2005).
3 Mutant Phenotypes Associated with Vertebrate L1-CAM Paralogs
3.1 Neurofascin
Neurofascin is one of three vertebrate L1-CAM paralogs and is also primarily expressed in the nervous system (Rathjen et al. 1987). The multiple protein isoforms that are expressed from the neurofascin gene vary in the number of their FN III extracellular domains (Hassel et al. 1997). Some Neurofascin protein isoforms also contain an unusual PAT domain, which is usually positioned close to the transmembrane segment (see Fig. 9.1) (Volkmer et al. 1992).
Two major Neurofascin isoforms, Nfasc155 and Nfasc186, are expressed at nodes of Ranvier in myelinated axons (Tait et al. 2000). The neuronal isoform Nfasc186 is required for the clustering of voltage-gated Na+ channels at nodes of Ranvier (Howell et al. 2006). It thereby controls rapid impulse conduction in these axons (Zonta et al. 2008). In contrast, Nfasc155 is a glia cell-specific protein isoform and is required for the correct assembly of paranodal junctions (Sherman et al. 2005). In neurofascin null mice, neither paranodal adhesion junctions nor nodal complexes are formed (Sherman et al. 2005). This demonstrates the essential function of these two major Neurofascin protein isoforms for the formation of these structures. Nfasc155 null mutant mice exhibit severe ataxia, motor paralysis, and death before the third postnatal week (Pillai et al. 2009). In the absence of glia cell-specific Nfasc155, paranodal axonal junctions fail to form, axonal domains do not segregate, and myelinated axons undergo degeneration. Furthermore, in vivo deletion of Neurofascin Ig domains 5 and 6 reveals a requirement for specific Neurofascin protein domains in myelinated axons (Thaxton et al. 2010).
In cases of multiple sclerosis, a disruption of Neurofascin localization at nodes of Ranvier often appears to precede subsequent demyelination (Howell et al. 2006; Lonigro and Devaux 2009). Both Nfasc186 and Nfasc155 proteins have been found in areas of inflammation, demyelination, and remyelination in postmortem brains of multiple sclerosis patients (Howell et al. 2006). Mathey et al. identified Neurofascin as an autoimmune target in patients with multiple sclerosis (Mathey et al. 2007). This suggests a direct involvement of Neurofascin in immune-mediated axonal injury. The alteration of oligodendrocyte Nfasc155 expression that accompanies inflammation and demyelination processes indicates a chronic disruption of the axon–glia cell interaction. This will eventually result in the destruction of the Nfasc186/Na+ v nodal complexes (Howell et al. 2006). Recently, the involvement in the progression of multiple sclerosis of two newly described protein isoforms of Nfasc155 has been analyzed in more detail (Pomicter et al. 2010). Using conditional knockout mice the authors show that Nfasc155 high and Nfasc155 low are exclusively expressed by oligodendrocytes within the CNS. The timing and expression levels of these two Nfasc155 isoforms are distinctly regulated. Nfasc155 low is incapable of preserving paranodal structures, thus indicating that Nfasc155 high is required for paranodal stability. Comparisons between Nfasc155 high and Nfasc155 low in human samples revealed significant alterations in multiple sclerosis plaques (Pomicter et al. 2010).
3.2 NrCAM
NrCAM (neuron–glia-related cell adhesion molecule) was the third vertebrate L1-type gene/protein to be identified and like its paralogs is primarily expressed in the nervous system (Grumet et al. 1991; Grumet 1997). In mice, lack of NrCAM has been implicated in the formation of lens cataracts (More et al. 2001). It is also involved in the formation of nodes of Ranvier (Custer et al. 2003) and in addiction vulnerability (Ishiguro et al. 2006). In humans, NrCAM has been associated with autism (Sakurai et al. 2006; Bonora et al. 2005) and is overexpressed in papillary thyroid carcinomas (Gorka et al. 2007).
The absence of NrCAM causes the formation of cataracts in murine lenses (More et al. 2001). In NrCAM-deficient mice, the authors observed a general disorganization of lens fibers with ensuing cellular disintegration and an accumulation of cellular debris. This mirrors the phenotype found in Ankyrin-B-deficient mice and points to an important interaction between NrCAM and Ankyrin-B in lens fiber cells (More et al. 2001). Similar to Neurofascin, NrCAM is also expressed at nodes of Ranvier and is implicated in node formation and maintenance (Custer et al. 2003). Na+ channels and Ankyrin G sequestration at developing nodes is delayed in NrCAM null mutant mice.
The genetic mapping of a locus involved in substance abuse vulnerabilities to mouse chromosome 7 identified a positive linkage with several NrCAM haplotypes (Ishiguro et al. 2006). Differential gene display identified the NrCAM gene as a drug-regulated gene that is expressed in neurons linked to reward and memory (Ishiguro et al. 2006). NrCAM knockout mice exhibit reduced opiate- and stimulant-conditional preferences. These observations suggest that in humans NrCAM may also be a drug-regulated gene, whose variants are likely linked to vulnerabilities in drug addiction and reward. In another study, wild-type, heterozygous, and NrCAM null mice were tested for a cognitive and behavioral phenotype (Matzel et al. 2008). These different genotypes were assessed using five different learning tasks (such as Lashley maze, odor discrimination, passive avoidance, spatial water maze, and fear conditioning). NrCAM null mutant mice are viable, have normal body weight, and exhibit normal levels of general activity. However, they display an increased propensity to enter stressful areas of novel environments, exhibit higher sensitivity to pain (hot), and are more sensitive to the aversive effects of foot shock. This behavioral phenotype suggests that NrCAM might play a central role in the regulation of general cognitive abilities and might serve a critical function in regulating impulsivity as well as susceptibly to drug abuse and addiction. In humans, two genetic linkage studies have identified the NrCAM gene as a potential candidate to be associated with autism susceptibility and with substance abuse (Sakurai et al. 2006; Bonora et al. 2005). Together with the results obtained from the mouse models, which are cited above, this indicates that NrCAM has an important function in the formation and/or maintenance of the brain’s reward circuitry.
Like L1-CAM and Neurofascin, NrCAM is also overexpressed in at least one type of human cancer, specifically papillary thyroid carcinomas (PTCs) (Gorka et al. 2007). The level of NrCAM mRNA and protein overexpression in tumor tissues appears to be independent of the primary tumor stage (PT) or its size. How NrCAM induction and upregulation might potentially influence the pathogenesis and the behavior of papillary thyroid cancer cells still remains to be evaluated.
3.3 CHL1
The Close Homologue of L1, CHL1 (or CALL for Cell Adhesion L1-Like), is the fourth vertebrate L1-type paralog (Holm et al. 1996) and is located on human chromosome 3p26.1 (Wei et al. 1998). CHL1 protein promotes neurite outgrowth (Holm et al. 1996; Hillenbrand et al. 1999), and the gene expression of CHL1 and L1CAM in the mouse and rat nervous system shows overlapping but distinct patterns in neuronal and glia cell populations (Hillenbrand et al. 1999). In contrast to the other three vertebrate L1 paralogs, relatively little is known about CHL1’s physiological and molecular functions. Nevertheless, the CHL1 gene appears to be associated with several interesting genetic conditions and phenotypes.
In cortical slices from CHL1 knockout mice the migration of cortical neurons proceeds at a slower rate of radial migration and migratory cells accumulate in the intermediate and ventricular/subventricular zones (Demyanenko et al. 2004). In neocortical areas, especially in the visual and somatosensory cortex, CHL1 appears to regulate neuronal connectivity (Demyanenko et al. 2004). CHL1 also has a role in regulating the uncoating of Clathrin-coated synaptic vesicles (Leshchyns’ka et al. 2006). CHL1 deficiency or disruption of the CHL1/Hsc70 complex results in an accumulation of abnormally high levels of Clathrin-coated synaptic vesicles. These observed abnormalities of Clathrin-dependent synaptic vesicle recycling have the potential to cause or to contribute to brain malfunctions in humans and mice that carry mutations in their CHL1 gene.
Nikonenko et al. (2006) reported an enhanced perisomatic inhibition and impaired long-term potentiation in the CA1 region of juvenile CHL1-deficient mice. These authors analyzed the functional role of CHL1 in the synaptic transmission in the CA1 region hippocampus comparing juvenile CHL1-deficient and wild-type mice. The inhibitory postsynaptic currents evoked in pyramidal cells by minimal stimulation of perisomatically projecting interneurons were increased in mice lacking CHL1 when compared with wild-type littermates. Also, the long-term potentiation (LTP) at CA3–CA1 excitatory synapses was reduced under physiological conditions in CHL1-deficient mice. A quantitative immunohistochemical analysis revealed that CA1 interneurons usually express CHL1 protein. This suggests that CHL1 is important for the regulation of inhibitory synaptic transmission in this and potentially other interneuron populations. This observed enhancement of inhibitory transmission in CHL1-deficient mice contrasts with the previous finding of a reduced inhibition in L1-CAM-deficient mice and illustrates a functional difference between these two paralogous L1-type adhesion molecules (Nikonenko et al. 2006).
Mutations in the murine CHL1 gene have been found to alter the connectivity and morphology of several brain regions (Heyden et al. 2008). In addition, CHL1 acts in a gene dosage-dependent manner to control murine brain development and to influence behavior and cognitive abilities. Mice deficient for CHL1 display alterations in emotional reactivity and motor coordination (Pratte et al. 2003). These mice also display signs of decreased stress and a modification of exploratory behavior and show impairments in a Rotarod test. However, they were able to move as fast as control mice in a T-maze test. The observed changes have been attributed to an attention deficit. CHL1-deficient mice have normal learning abilities, but exhibit a widespread impairment in working memory duration (Kolata et al. 2008). Montag-Saliaz et al. demonstrated that the absence of CHL1 in mice results in aberrant hippocampal mossy fiber and olfactory nerve projections, which might explain the reduced reactivity towards novel environments that is exhibited by these mice (Montag-Sallaz et al. 2003). Together, these observations that have been made using CHL1-deficient mouse models suggest an important role for CHL1 in short-term memory retention in the adult brain.
Similar to the other three vertebrate L1-type CAMs, mutations in the CHL1 gene and protein have been implicated in several human disease conditions. CHL1 has been identified as a prime candidate gene for an autosomal form of mental retardation and a translocation breakpoint in intron five of the CHL1 gene at 46,Y, t(X;3)(p22.1;p26.3) was described in a man with nonspecific mental retardation (Frints et al. 2003). In addition, a haploinsufficiency for the CHL1 gene has been reported in a mentally retarded patient with 3p-syndrome (Angeloni et al. 1999). A missense mutation (Leu17Phe) in the signal peptide of CHL1 exhibits a positive association with the occurrence of schizophrenia in a group of 282 Japanese patients (Sakurai et al. 2002). This association of CHL1 with schizophrenia was later confirmed by a second study of 560 schizophrenia cases and 576 controls in a Han Chinese population (Chen et al. 2005). These two reports indicate that CHL1 might somehow be involved in the etiology of schizophrenia.
4 Conclusions
L1-type genes and their protein products appeared rather early during metazoan evolution when the first primitive neuronal nets became part of animal body plans. From the beginning, the overall L1-type protein domain structure, a predominant expression of L1-type genes in neural cells, and many characteristic protein–protein interactions (both homo- as well as heterophilic) and molecular functions have been remarkably well conserved. However, gene duplication events have allowed for some variations in the L1 protein structure and for the development of novel molecular interactions and physiological functions (Mualla et al. 2013). Vertebrate species usually contain four L1-type genes in their genome, designated as L1-CAM, Neurofascin, NrCAM, and CHL1. Loss-of-function conditions have been studied in several animal model systems and usually include a broad range of neurological dysfunctions. In humans, mutations in the L1CAM gene have been analyzed in great detail as they cause an X-linked recessive disorder, now known as L1 syndrome. Well over 200 different L1-CAM mutations have been identified in individual families. The severity of the phenotype usually correlates with the impact of the specific mutation on the structure of the L1-CAM protein and its interactions with other proteins. However, epigenetic effects also contribute to a significant variability of specific phenotypic aspects.
Though L1-type genes are predominantly expressed in the nervous system, their protein products have also been identified in a number of other tissues, specifically leukocytes and epithelia. Their physiological functions in these non-neural tissues still remain poorly understood. Overexpression of L1-type proteins has been reported in a wide range of different human tumors, where they affect different signaling pathways and contribute to tumor progression and metastasis. As our understanding of the multifaceted normal physiological role of these important proteins is still incomplete and as both loss-of-function and gain-of-function conditions for L1-type genes/proteins cause clinically relevant disorders, the published findings paint a complex and sometimes confusing picture. Certainly, more research is needed for a more complete insight into the physiological role of L1-type CAMs and how they are implicated into various pathological processes.
References
Allory Y, Matsuoka Y, Bazille C, Christensen EI, Ronco P, Debiec H (2005) The L1 cell adhesion molecule is induced in renal cancer cells and correlates with metastasis in clear cell carcinomas. Clin Cancer Res 11(3):1190–1197
Amores A, Force A, Yan YL, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang YL, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish hox clusters and vertebrate genome evolution. Science 282(5394):1711–1714
Anderson RB, Turner KN, Nikonenko AG, Hemperly J, Schachner M, Young HM (2006) The cell adhesion molecule l1 is required for chain migration of neural crest cells in the developing mouse gut. Gastroenterology 130(4):1221–1232. doi:10.1053/j.gastro.2006.01.002, S0016-5085(06)00003-5 [pii]
Angeloni D, Lindor NM, Pack S, Latif F, Wei MH, Lerman MI (1999) CALL gene is haploinsufficient in a 3p-syndrome patient. Am J Med Genet 86(5):482–485
Ango F, di Cristo G, Higashiyama H, Bennett V, Wu P, Huang ZJ (2004) Ankyrin-based subcellular gradient of neurofascin, an immunoglobulin family protein, directs GABAergic innervation at purkinje axon initial segment. Cell 119(2):257–272
Asou H, Miura M, Kobayashi M, Uyemura K, Itoh K (1992) Cell adhesion molecule L1 guides cell migration in primary reaggregation cultures of mouse cerebellar cells. Neurosci Lett 144(1–2):221–224
Balaian LB, Moehler T, Montgomery AM (2000) The human neural cell adhesion molecule L1 functions as a costimulatory molecule in T cell activation. Eur J Immunol 30(3):938–943
Banerjee S, Pillai AM, Paik R, Li J, Bhat MA (2006) Axonal ensheathment and septate junction formation in the peripheral nervous system of Drosophila. J Neurosci 26(12):3319–3329
Bao S, Wu Q, Li Z, Sathornsumetee S, Wang H, McLendon RE, Hjelmeland AB, Rich JN (2008) Targeting cancer stem cells through L1CAM suppresses glioma growth. Cancer Res 68(15):6043–6048. doi:10.1158/0008-5472.CAN-08-1079, 68/15/6043 [pii]
Bartsch U (2003) Neural CAMs and their role in the development and organization of myelin sheaths. Front Biosci 8:D477–D490
Bateman A, Jouet M, MacFarlane J, Du JS, Kenwrick S, Chothia C (1996) Outline structure of the human L1 cell adhesion molecule and the sites where mutations cause neurological disorders. EMBO J 15(22):6050–6059
Becker CG, Lieberoth BC, Morellini F, Feldner J, Becker T, Schachner M (2004) L1.1 is involved in spinal cord regeneration in adult zebrafish. J Neurosci 24(36):7837–7842
Bernreuther C, Dihne M, Johann V, Schiefer J, Cui Y, Hargus G, Schmid JS, Xu J, Kosinski CM, Schachner M (2006) Neural cell adhesion molecule L1-transfected embryonic stem cells promote functional recovery after excitotoxic lesion of the mouse striatum. J Neurosci 26(45):11532–11539
Bertolin C, Boaretto F, Barbon G, Salviati L, Lapi E, Divizia MT, Garavelli L, Occhi G, Vazza G, Mostacciuolo ML (2010) Novel mutations in the L1CAM gene support the complexity of L1 syndrome. J Neurol Sci 294(1–2):124–126. doi:10.1016/j.jns.2010.03.030, S0022-510X(10)00156-5 [pii]
Bickers DS, Adams RD (1949) Hereditary stenosis of the aqueduct of Sylvius as a casue of congenital hydrocephalus. Brain 72:246–262
Bieber AJ, Snow PM, Hortsch M, Patel NH, Jacobs JR, Traquina ZR, Schilling J, Goodman CS (1989) Drosophila neuroglian: a member of the immunoglobulin superfamily with extensive homology to the vertebrate neural adhesion molecule L1. Cell 59(3):447–460
Blaess S, Kammerer RA, Hall H (1998) Structural analysis of the sixth immunoglobulin-like domain of mouse neural cell adhesion molecule L1 and its interactions with alpha(v)beta3, alpha(IIb)beta3, and alpha5beta1 integrins. J Neurochem 71(6):2615–2625
Bonora E, Lamb JA, Barnby G, Sykes N, Moberly T, Beyer KS, Klauck SM, Poustka F, Bacchelli E, Blasi F, Maestrini E, Battaglia A, Haracopos D, Pedersen L, Isager T, Eriksen G, Viskum B, Sorensen EU, Brondum-Nielsen K, Cotterill R, Engeland H, Jonge M, Kemner C, Steggehuis K, Scherpenisse M, Rutter M, Bolton PF, Parr JR, Poustka A, Bailey AJ, Monaco AP (2005) Mutation screening and association analysis of six candidate genes for autism on chromosome 7q. Eur J Hum Genet 13(2):198–207. doi:10.1038/sj.ejhg.5201315, 5201315 [pii]
Buhusi M, Schlatter MC, Demyanenko GP, Thresher R, Maness PF (2008) L1 interaction with ankyrin regulates mediolateral topography in the retinocollicular projection. J Neurosci 28(1):177–188
Carhan A, Allen F, Armstrong JD, Hortsch M, Goodwin SF, O’Dell KM (2005) Female receptivity phenotype of icebox mutants caused by a mutation in the L1-type cell adhesion molecule neuroglian. Genes Brain Behav 4(8):449–465
Castellani V, Chedotal A, Schachner M, Faivre-Sarrailh C, Rougon G (2000) Analysis of the L1-deficient mouse phenotype reveals cross-talk between Sema3A and L1 signaling pathways in axonal guidance. Neuron 27(2):237–249
Charles P, Tait S, Faivre-Sarrailh C, Barbin G, Gunn-Moore F, Denisenko-Nehrbass N, Guennoc AM, Girault JA, Brophy PJ, Lubetzki C (2002) Neurofascin is a glial receptor for the paranodin/Caspr-contactin axonal complex at the axoglial junction. Curr Biol 12(3):217–220
Chen W, Hing H (2008) The L1-CAM, neuroglian, functions in glial cells for Drosophila antennal lobe development. Dev Neurobiol 68(8):1029–1045. doi:10.1002/dneu.20644
Chen L, Zhou S (2010) “CRASH”ing with the worm: insights into L1CAM functions and mechanisms. Dev Dyn 239(5):1490–1501. doi:10.1002/dvdy.22269
Chen L, Ong B, Bennett V (2001) LAD-1, the Caenorhabditis elegans L1CAM homologue, participates in embryonic and gonadal morphogenesis and is a substrate for fibroblast growth factor receptor pathway-dependent phosphotyrosine-based signaling. J Cell Biol 154(4):841–856
Chen QY, Chen Q, Feng GY, Lindpaintner K, Chen Y, Sun X, Chen Z, Gao Z, Tang J, He L (2005) Case-control association study of the close homologue of L1 (CHL1) gene and schizophrenia in the Chinese population. Schizophr Res 73(2–3):269–274
Chen J, Wu J, Apostolova I, Skup M, Irintchev A, Kugler S, Schachner M (2007) Adeno-associated virus-mediated L1 expression promotes functional recovery after spinal cord injury. Brain 130(Pt 4):954–969. doi:10.1093/brain/awm049, 130/4/954 [pii]
Chen MM, Lee CY, Leland HA, Silletti S (2011) Modification of the L1-CAM carboxy-terminus in pancreatic adenocarcinoma cells. Tumour Biol 32(2):347–357. doi:10.1007/s13277-010-0127-4
Cheng L, Lemmon V (2004) Pathological missense mutations of neural cell adhesion molecule L1 affect neurite outgrowth and branching on an L1 substrate. Mol Cell Neurosci 27(4):522–530
Cheng L, Itoh K, Lemmon V (2005a) L1-mediated branching is regulated by two ezrin-radixin-moesin (ERM)-binding sites, the RSLE region and a novel juxtamembrane ERM-binding region. J Neurosci 25(2):395–403
Cheng L, Lemmon S, Lemmon V (2005b) RanBPM is an L1-interacting protein that regulates L1-mediated mitogen-activated protein kinase activation. J Neurochem 94(4):1102–1110. doi:10.1111/j.1471-4159.2005.03254.x, JNC3254 [pii]
Christoffels A, Koh EG, Chia JM, Brenner S, Aparicio S, Venkatesh B (2004) Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Mol Biol Evol 21(6):1146–1151. doi:10.1093/molbev/msh114msh114 [pii]
Collinson JM, Marshall D, Gillespie CS, Brophy PJ (1998) Transient expression of neurofascin by oligodendrocytes at the onset of myelinogenesis: implications for mechanisms of axon-glial interaction. Glia 23(1):11–23
Conacci-Sorrell M, Kaplan A, Raveh S, Gavert N, Sakurai T, Ben-Ze’ev A (2005) The shed ectodomain of Nr-CAM stimulates cell proliferation and motility, and confers cell transformation. Cancer Res 65(24):11605–11612
Custer AW, Kazarinova-Noyes K, Sakurai T, Xu X, Simon W, Grumet M, Shrager P (2003) The role of the ankyrin-binding protein NrCAM in node of Ranvier formation. J Neurosci 23(31):10032–10039
Dahme M, Bartsch U, Martini R, Anliker B, Schachner M, Mantei N (1997) Disruption of the mouse L1 gene leads to malformations of the nervous system. Nat Genet 17(3):346–349
Davis JQ, McLaughlin T, Bennett V (1993) Ankyrin-binding proteins related to nervous system cell adhesion molecules: candidates to provide transmembrane and intercellular connections in adult brain. J Cell Biol 121(1):121–133
Davis JQ, Lambert S, Bennett V (1996) Molecular composition of the node of Ranvier: Identification of ankyrin-binding cell adhesion molecules neurofascin (mucin+/third FNIII domain-) and NrCAM at nodal axon segments. J Cell Biol 135(5):1355–1367
De Angelis E, MacFarlane J, Du JS, Yeo G, Hicks R, Rathjen FG, Kenwrick S, Brümmendorf T (1999) Pathological missense mutations of neural cell adhesion molecule L1 affect homophilic and heterophilic binding activities. EMBO J 18(17):4744–4753
De Angelis E, Brümmendorf T, Cheng L, Lemmon V, Kenwrick S (2001) Alternative use of a mini exon of the L1 gene affects L1 binding to neural ligands. J Biol Chem 276(35):32738–32742
De Angelis E, Watkins A, Schäfer M, Brümmendorf T, Kenwrick S (2002) Disease-associated mutations in L1 CAM interfere with ligand interactions and cell-surface expression. Hum Mol Genet 11(1):1–12
Debiec H, Christensen EI, Ronco PM (1998) The cell adhesion molecule L1 is developmentally regulated in the renal epithelium and is involved in kidney branching morphogenesis. J Cell Biol 143(7):2067–2079
Deichmann M, Kurzen H, Egner U, Altevogt P, Hartschuh W (2003) Adhesion molecules CD171 (L1CAM) and CD24 are expressed by primary neuroendocrine carcinomas of the skin (Merkel cell carcinomas). J Cutan Pathol 30(6):363–368
Demyanenko GP, Maness PF (2003) The L1 cell adhesion molecule is essential for topographic mapping of retinal axons. J Neurosci 23(2):530–538
Demyanenko GP, Tsai AY, Maness PF (1999) Abnormalities in neuronal process extension, hippocampal development, and the ventricular system of L1 knockout mice. J Neurosci 19(12):4907–4920
Demyanenko GP, Shibata Y, Maness PF (2001) Altered distribution of dopaminergic neurons in the brain of L1 null mice. Brain Res Dev Brain Res 126(1):21–30
Demyanenko GP, Schachner M, Anton E, Schmid R, Feng G, Sanes J, Maness PF (2004) Close homolog of L1 modulates area-specific neuronal positioning and dendrite orientation in the cerebral cortex. Neuron 44(3):423–437
Demyanenko GP, Halberstadt AI, Rao RS, Maness PF (2009) CHL1 cooperates with PAK1-3 to regulate morphological differentiation of embryonic cortical neurons. Neuroscience 165(1):107–115
Di Sciullo G, Donahue T, Schachner M, Bogen SA (1998) L1 antibodies block lymph node fibroblastic reticular matrix remodeling in vivo. J Exp Med 187(12):1953–1963
Dietrich A, Korn B, Poustka A (1992) Completion of the physical map of Xq28: the location of the gene for L1CAM on the human X chromosome. Mamm Genome 3(3):168–172
Dihne M, Bernreuther C, Sibbe M, Paulus W, Schachner M (2003) A new role for the cell adhesion molecule L1 in neural precursor cell proliferation, differentiation, and transmitter-specific subtype generation. J Neurosci 23(16):6638–6650
Dubreuil RR, MacVicar G, Dissanayake S, Liu C, Homer D, Hortsch M (1996) Neuroglian-mediated cell adhesion induces assembly of the membrane skeleton at cell contact sites. J Cell Biol 133(3):647–655
Duczmal A, Schollhammer S, Katich S, Ebeling O, Schwartz-Albiez R, Altevogt P (1997) The L1 adhesion molecule supports alpha v beta 3-mediated migration of human tumor cells and activated T lymphocytes. Biochem Biophys Res Commun 232(1):236–239
Ebeling O, Duczmal A, Aigner S, Geiger C, Schöllhammer S, Kemshead JT, Möller P, Schwartz-Albiez R, Altevogt P (1996) L1 adhesion molecule on human lymphocytes and monocytes: expression and involvement in binding to alpha v beta 3 integrin. Eur J Immunol 26(10):2508–2516
Faivre-Sarrailh C, Banerjee S, Li J, Hortsch M, Laval M, Bhat MA (2004) Drosophila contactin, a homolog of vertebrate contactin, is required for septate junction organization and paracellular barrier function. Development 131(20):4931–4942
Felding-Habermann B, Silletti S, Mei F, Siu CH, Yip PM, Brooks PC, Cheresh DA, O’Toole TE, Ginsberg MH, Montgomery AM (1997) A single immunoglobulin-like domain of the human neural cell adhesion molecule L1 supports adhesion by multiple vascular and platelet integrins. J Cell Biol 139(6):1567–1581
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17(6):368–376
Figarella-Branger DF, Durbec PL, Rougon GN (1990) Differential spectrum of expression of neural cell adhesion molecule isoforms and L1 adhesion molecules on human neuroectodermal tumors. Cancer Res 50(19):6364–6370
Finckh U, Schroder J, Ressler B, Veske A, Gal A (2000) Spectrum and detection rate of L1CAM mutations in isolated and familial cases with clinically suspected L1-disease. Am J Med Genet 92(1):40–46
Fogel M, Mechtersheimer S, Huszar M, Smirnov A, Abu-Dahi A, Tilgen W, Reichrath J, Georg T, Altevogt P, Gutwein P (2003) L1 adhesion molecule (CD 171) in development and progression of human malignant melanoma. Cancer Lett 189(2):237–247
Fransen E, Schrander-Stumpel C, Vits L, Coucke P, Van Camp G, Willems PJ (1994) X-linked hydrocephalus and MASA syndrome present in one family are due to a single missense mutation in exon 28 of the L1CAM gene. Hum Mol Genet 3(12):2255–2256
Fransen E, Lemmon V, Van Camp G, Vits L, Coucke P, Willems PJ (1995) CRASH syndrome: clinical spectrum of corpus callosum hypoplasia, retardation, adducted thumbs, spastic paraparesis and hydrocephalus due to mutations in one single gene, L1. Eur J Hum Genet 3(5):273–284
Fransen E, Van Camp G, Vits L, Willems PJ (1997) L1-associated diseases: clinical geneticists divide, molecular geneticists unite. Hum Mol Genet 6(10):1625–1632
Fransen E, D’Hooge R, Van Camp G, Verhoye M, Sijbers J, Reyniers E, Soriano P, Kamiguchi H, Willemsen R, Koekkoek SK, De Zeeuw CI, De Deyn PP, Van der Linden A, Lemmon V, Kooy RF, Willems PJ (1998a) L1 knockout mice show dilated ventricles, vermis hypoplasia and impaired exploration patterns. Hum Mol Genet 7(6):999–1009
Fransen E, Van Camp G, D’Hooge R, Vits L, Willems PJ (1998b) Genotype-phenotype correlation in L1 associated diseases. J Med Genet 35(5):399–404
Frints SG, Marynen P, Hartmann D, Fryns JP, Steyaert J, Schachner M, Rolf B, Craessaerts K, Snellinx A, Hollanders K, D’Hooge R, De Deyn PP, Froyen G (2003) CALL interrupted in a patient with non-specific mental retardation: gene dosage-dependent alteration of murine brain development and behavior. Hum Mol Genet 12(13):1463–1474
Fryns JP, Spaepen A, Cassiman JJ, van den Berghe H (1991) X linked complicated spastic paraplegia, MASA syndrome, and X linked hydrocephalus owing to congenital stenosis of the aqueduct of Sylvius: variable expression of the same mutation at Xq28. J Med Genet 28(6):429–431
Garcia-Alonso L, Romani S, Jimenez F (2000) The EGF and FGF receptors mediate neuroglian function to control growth cone decisions during sensory axon guidance in Drosophila. Neuron 28(3):741–752
Garver TD, Ren Q, Tuvia S, Bennett V (1997) Tyrosine phosphorylation at a site highly conserved in the L1 family of cell adhesion molecules abolishes ankyrin binding and increases lateral mobility of neurofascin. J Cell Biol 137(3):703–714
Gast D, Riedle S, Issa Y, Pfeifer M, Beckhove P, Sanderson MP, Arlt M, Moldenhauer G, Fogel M, Kruger A, Altevogt P (2008) The cytoplasmic part of L1-CAM controls growth and gene expression in human tumors that is reversed by therapeutic antibodies. Oncogene 27(9):1281–1289. doi:10.1038/sj.onc.1210747, 1210747 [pii]
Gavert N, Conacci-Sorrell M, Gast D, Schneider A, Altevogt P, Brabletz T, Ben-Ze’ev A (2005) L1, a novel target of beta-catenin signaling, transforms cells and is expressed at the invasive front of colon cancers. J Cell Biol 168(4):633–642
Gavert N, Sheffer M, Raveh S, Spaderna S, Shtutman M, Brabletz T, Barany F, Paty P, Notterman D, Domany E, Ben-Ze’ev A (2007) Expression of L1-CAM and ADAM10 in human colon cancer cells induces metastasis. Cancer Res 67(16):7703–7712
Gavert N, Ben-Shmuel A, Raveh S, Ben-Ze’ev A (2008) L1-CAM in cancerous tissues. Expert Opin Biol Ther 8(11):1749–1757. doi:10.1517/14712598.8.11.1749
Genova JL, Fehon RG (2003) Neuroglian, gliotactin, and the Na+/K+ ATPase are essential for septate junction function in Drosophila. J Cell Biol 161(5):979–989
Godenschwege TA, Kristiansen LV, Uthaman SB, Hortsch M, Murphey RK (2006) A conserved role for Drosophila Neuroglian and human L1-CAM in central-synapse formation. Curr Biol 16(1):12–23
Goossens T, Kang YY, Wuytens G, Zimmermann P, Callaerts-Vegh Z, Pollarolo G, Islam R, Hortsch M, Callaerts P (2011) The Drosophila L1CAM homolog Neuroglian signals through distinct pathways to control different aspects of mushroom body axon development. Development 138(8):1595–1605. doi:10.1242/dev.052787, dev.052787 [pii]
Gorka B, Skubis-Zegadlo J, Mikula M, Bardadin K, Paliczka E, Czarnocka B (2007) NrCAM, a neuronal system cell-adhesion molecule, is induced in papillary thyroid carcinomas. Br J Cancer 97(4):531–538
Griseri P, Vos Y, Giorda R, Gimelli S, Beri S, Santamaria G, Mognato G, Hofstra RM, Gimelli G, Ceccherini I (2009) Complex pathogenesis of Hirschsprung’s disease in a patient with hydrocephalus, vesico-ureteral reflux and a balanced translocation t(3;17)(p12;q11). Eur J Hum Genet 17(4):483–490. doi:10.1038/ejhg.2008.191, ejhg2008191 [pii]
Grumet M (1997) Nr-CAM: a cell adhesion molecule with ligand and receptor functions. Cell Tissue Res 290(2):423–428
Grumet M, Mauro V, Burgoon MP, Edelman GM, Cunningham BA (1991) Structure of a new nervous system glycoprotein, Nr-CAM, and relationship to subgroups of neural cell adhesion molecules. J Cell Biol 113(6):1399–1412
Gu SM, Orth U, Veske A, Enders H, Klünder K, Schlösser M, Engel W, Schwinger E, Gal A (1996) Five novel mutations in the L1CAM gene in families with X linked hydrocephalus. J Med Genet 33(2):103–106
Guan H, Maness PF (2010) Perisomatic GABAergic innervation in prefrontal cortex is regulated by ankyrin interaction with the L1 cell adhesion molecule. Cereb Cortex 20(11):2684–2693
Gutwein P, Oleszewski M, Mechtersheimer S, Agmon-Levin N, Krauss K, Altevogt P (2000) Role of Src kinases in the ADAM-mediated release of L1 adhesion molecule from human tumor cells. J Biol Chem 275(20):15490–15497
Hall SG, Bieber AJ (1997) Mutations in the Drosophila neuroglian cell adhesion molecule affect motor neuron pathfinding and peripheral nervous system patterning. J Neurobiol 32(3):325–340
Haney CA, Sahenk Z, Li C, Lemmon VP, Roder J, Trapp BD (1999) Heterophilic binding of L1 on unmyelinated sensory axons mediates Schwann cell adhesion and is required for axonal survival. J Cell Biol 146(5):1173–1184
Harpaz Y, Chothia C (1994) Many of the immunoglobulin superfamily domains in cell adhesion molecules and surface receptors belong to a new structural set which is close to that containing variable domains. J Mol Biol 238(4):528–539
Harper SJ, Bolsover SR, Walsh FS, Doherty P (1994) Neurite outgrowth stimulated by L1 requires calcium influx into neurons but is not associated with changes in steady state levels of calcium in growth cones. Cell Adhes Commun 2(5):441–453
Hassel B, Rathjen FG, Volkmer H (1997) Organization of the neurofascin gene and analysis of developmentally regulated alternative splicing. J Biol Chem 272(45):28742–28749
He Y, Jensen GJ, Bjorkman PJ (2009) Cryo-electron tomography of homophilic adhesion mediated by the neural cell adhesion molecule L1. Structure 17(3):460–471. doi:10.1016/j.str.2009.01.009, S0969-2126(09)00073-2 [pii]
Heyden A, Angenstein F, Sallaz M, Seidenbecher C, Montag D (2008) Abnormal axonal guidance and brain anatomy in mouse mutants for the cell recognition molecules close homolog of L1 and NgCAM-related cell adhesion molecule. Neuroscience 155(1):221–233. doi:10.1016/j.neuroscience.2008.04.080, S0306-4522(08)00645-3 [pii]
Hillenbrand R, Molthagen M, Montag D, Schachner M (1999) The close homologue of the neural adhesion molecule L1 (CHL1): patterns of expression and promotion of neurite outgrowth by heterophilic interactions. Eur J Neurosci 11(3):813–826
Holm J, Hillenbrand R, Steuber V, Bartsch U, Moos M, Lubbert H, Montag D, Schachner M (1996) Structural features of a close homologue of L1 (CHL1) in the mouse: a new member of the L1 family of neural recognition molecules. Eur J Neurosci 8(8):1613–1629
Hortsch M (1996) The L1 family of neural cell adhesion molecules: old proteins performing new tricks. Neuron 17(4):587–593
Hortsch M (2000) Structural and functional evolution of the L1-family: are four adhesion molecules better than one? Mol Cell Neurosci 15(1):1–10
Hortsch M, Margolis B (2003) Septate and paranodal junctions: kissing cousins. Trends Cell Biol 13(11):557–561
Hortsch M, Bieber AJ, Patel NH, Goodman CS (1990) Differential splicing generates a nervous system-specific form of Drosophila neuroglian. Neuron 4(5):697–709
Hortsch M, Wang YM, Marikar Y, Bieber AJ (1995) The cytoplasmic domain of the Drosophila cell adhesion molecule Neuroglian is not essential for its homophilic adhesive properties in S2 cells. J Biol Chem 270(32):18809–18817
Hortsch M, Homer D, Malhotra JD, Chang S, Frankel J, Jefford G, Dubreuil RR (1998a) Structural requirements for “outside-in” and “inside-out” signaling by Drosophila neuroglian, a member of the L1 family of cell adhesion molecules. J Cell Biol 142(1):251–261
Hortsch M, O’Shea KS, Zhao G, Kim F, Vallejo Y, Dubreuil RR (1998b) A conserved role for L1 as a transmembrane link between neuronal adhesion and membrane cytoskeleton assembly. Cell Adhes Commun 5(1):61–73
Hortsch M, Nagaraj K, Godenschwege TA (2009) The interaction between L1-type proteins and ankyrins – a master switch for L1-type CAM function. Cell Mol Biol Lett 14:57–69
Howell OW, Palser A, Polito A, Melrose S, Zonta B, Scheiermann C, Vora AJ, Brophy PJ, Reynolds R (2006) Disruption of neurofascin localization reveals early changes preceding demyelination and remyelination in multiple sclerosis. Brain 129(Pt 12):3173–3185. doi:10.1093/brain/awl290, awl290 [pii]
Hubbe M, Kowitz A, Schirrmacher V, Schachner M, Altevogt P (1993) L1 adhesion molecule on mouse leukocytes: regulation and involvement in endothelial cell binding. Eur J Immunol 23(11):2927–2931
Hübner CA, Utermann B, Tinschert S, Kruger G, Ressler B, Steglich C, Schinzel A, Gal A (2004) Intronic mutations in the L1CAM gene may cause X-linked hydrocephalus by aberrant splicing. Hum Mutat 23(5):526. doi:10.1002/humu.9242
Hulley P, Schachner M, Lubbert H (1998) L1 neural cell adhesion molecule is a survival factor for fetal dopaminergic neurons. J Neurosci Res 53(2):129–134
Imondi R, Jevince AR, Helms AW, Johnson JE, Kaprielian Z (2007) Mis-expression of L1 on pre-crossing spinal commissural axons disrupts pathfinding at the ventral midline. Mol Cell Neurosci 36(4):462–471. doi:10.1016/j.mcn.2007.08.003, S1044-7431(07)00179-0 [pii]
Ishiguro H, Liu QR, Gong JP, Hall FS, Ujike H, Morales M, Sakurai T, Grumet M, Uhl GR (2006) NrCAM in addiction vulnerability: positional cloning, drug-regulation, haplotype-specific expression, and altered drug reward in knockout mice. Neuropsychopharmacology 31(3):572–584. doi:10.1038/sj.npp.1300855, 1300855 [pii]
Itoh K, Cheng L, Kamei Y, Fushiki S, Kamiguchi H, Gutwein P, Stoeck A, Arnold B, Altevogt P, Lemmon V (2004) Brain development in mice lacking L1-L1 homophilic adhesion. J Cell Biol 165(1):145–154
Itoh K, Fushiki S, Kamiguchi H, Arnold B, Altevogt P, Lemmon V (2005) Disrupted Schwann cell-axon interactions in peripheral nerves of mice with altered L1-integrin interactions. Mol Cell Neurosci 30(1):131–136
Izumoto S, Ohnishi T, Arita N, Hiraga S, Taki T, Hayakawa T (1996) Gene expression of neural cell adhesion molecule L1 in malignant gliomas and biological significance of L1 in glioma invasion. Cancer Res 56(6):1440–1444
Jakovcevski I, Siering J, Hargus G, Karl N, Hoelters L, Djogo N, Yin S, Zecevic N, Schachner M, Irintchev A (2009) Close homologue of adhesion molecule L1 promotes survival of Purkinje and granule cells and granule cell migration during murine cerebellar development. J Comp Neurol 513(5):496–510. doi:10.1002/cne.21981
Jenkins SM, Bennett V (2002) Developing nodes of Ranvier are defined by ankyrin-G clustering and are independent of paranodal axoglial adhesion. Proc Natl Acad Sci USA 99(4):2303–2308
Jenkins SM, Kizhatil K, Kramarcy NR, Sen A, Sealock R, Bennett V (2001) FIGQY phosphorylation defines discrete populations of L1 cell adhesion molecules at sites of cell-cell contact and in migrating neurons. J Cell Sci 114(Pt 21):3823–3835
Jouet M, Rosenthal A, Armstrong G, MacFarlane J, Stevenson R, Paterson J, Metzenberg A, Ionasescu V, Temple K, Kenwrick S (1994) X-linked spastic paraplegia (SPG1), MASA syndrome and X-linked hydrocephalus result from mutations in the L1 gene. Nat Genet 7(3):402–407
Jouet M, Rosenthal A, Kenwrick S (1995) Exon 2 of the gene for neural cell adhesion molecule L1 is alternatively spliced in B cells. Mol Brain Res 30(2):378–380
Kadmon G, Montgomery AM, Altevogt P (1998) L1 makes immunological progress by expanding its relations. Dev Immunol 6(3–4):205–213
Kaifi JT, Heidtmann S, Schurr PG, Reichelt U, Mann O, Yekebas EF, Wachowiak R, Strate T, Schachner M, Izbicki JR (2006a) Absence of L1 in pancreatic masses distinguishes adenocarcinomas from poorly differentiated neuroendocrine carcinomas. Anticancer Res 26(2A):1167–1170
Kaifi JT, Zinnkann U, Yekebas EF, Schurr PG, Reichelt U, Wachowiak R, Fiegel HC, Petri S, Schachner M, Izbicki JR (2006b) L1 is a potential marker for poorly-differentiated pancreatic neuroendocrine carcinoma. World J Gastroenterol 12(1):94–98
Kallunki P, Edelman GM, Jones FS (1997) Tissue-specific expression of the L1 cell adhesion molecule is modulated by the neural restrictive silencer element. J Cell Biol 138(6):1343–1354
Kamiguchi H, Lemmon V (1998) A neuronal form of the cell adhesion molecule L1 contains a tyrosine-based signal required for sorting to the axonal growth cone. J Neurosci 18(10):3749–3756
Kamiguchi H, Long KE, Pendergast M, Schaefer AW, Rapoport I, Kirchhausen T, Lemmon V (1998) The neural cell adhesion molecule L1 interacts with the AP-2 adaptor and is endocytosed via the clathrin-mediated pathway. J Neurosci 18(14):5311–5321
Kanemura Y, Okamoto N, Sakamoto H, Shofuda T, Kamiguchi H, Yamasaki M (2006) Molecular mechanisms and neuroimaging criteria for severe L1 syndrome with X-linked hydrocephalus. J Neurosurg 105(5 Suppl):403–412. doi:10.3171/ped.2006.105.5.403
Kaplan P (1983) X linked recessive inheritance of agenesis of the corpus callosum. J Med Genet 20(2):122–124
Kappen C, Schughart K, Ruddle FH (1989) Two steps in the evolution of Antennapedia-class vertebrate homeobox genes. Proc Natl Acad Sci USA 86(14):5459–5463
Katayama M, Iwamatsu A, Masutani H, Furuke K, Takeda K, Wada H, Masuda T, Ishii K (1997) Expression of neural cell adhesion molecule L1 in human lung cancer cell lines. Cell Struct Funct 22(5):511–516
Knops NB, Bos KK, Kerstjens M, van Dael K, Vos YJ (2008) Nephrogenic diabetes insipidus in a patient with L1 syndrome: a new report of a contiguous gene deletion syndrome including L1CAM and AVPR2. Am J Med Genet A 146A(14):1853–1858. doi:10.1002/ajmg.a.32386
Kobayashi M, Miura M, Asou H, Uyemura K (1991) Molecular cloning of cell adhesion molecule L1 from human nervous tissue: a comparison of the primary sequences of L1 molecules of different origin. Biochim Biophys Acta 1090(2):238–240
Kolata S, Wu J, Light K, Schachner M, Matzel LD (2008) Impaired working memory duration but normal learning abilities found in mice that are conditionally deficient in the close homolog of L1. J Neurosci 28(50):13505–13510. doi:10.1523/JNEUROSCI.2127-08.2008, 28/50/13505 [pii]
Koticha D, Babiarz J, Kane-Goldsmith N, Jacob J, Raju K, Grumet M (2005) Cell adhesion and neurite outgrowth are promoted by neurofascin NF155 and inhibited by NF186. Mol Cell Neurosci 30(1):137–148
Koticha D, Maurel P, Zanazzi G, Kane-Goldsmith N, Basak S, Babiarz J, Salzer J, Grumet M (2006) Neurofascin interactions play a critical role in clustering sodium channels, ankyrin G and beta IV spectrin at peripheral nodes of Ranvier. Dev Biol 293(1):1–12
Kowitz A, Kadmon G, Eckert M, Schirrmacher V, Schachner M, Altevogt P (1992) Expression and function of the neural cell adhesion molecule L1 in mouse leukocytes. Eur J Immunol 22(5):1199–1205
Kristiansen LV, Velasquez E, Romani S, Baars S, Berezin V, Bock E, Hortsch M, Garcia-Alonso L (2005) Genetic analysis of an overlapping functional requirement for L1- and NCAM-type proteins during sensory axon guidance in Drosophila. Mol Cell Neurosci 28(1):141–152
Laval M, Bel C, Faivre-Sarrailh C (2008) The lateral mobility of cell adhesion molecules is highly restricted at septate junctions in Drosophila. BMC Cell Biol 9:38. doi:10.1186/1471-2121-9-38, 1471-2121-9-38 [pii]
Lemmon V, Farr KL, Lagenaur C (1989) L1-mediated axon outgrowth occurs via a homophilic binding mechanism. Neuron 2(6):1597–1603
Leshchyns’ka I, Sytnyk V, Richter M, Andreyeva A, Puchkov D, Schachner M (2006) The adhesion molecule CHL1 regulates uncoating of clathrin-coated synaptic vesicles. Neuron 52(6):1011–1025. doi:10.1016/j.neuron.2006.10.020, S0896-6273(06)00820-8 [pii]
Linnemann D, Raz A, Bock E (1989) Differential expression of cell adhesion molecules in variants of K1735 melanoma cells differing in metastatic capacity. Int J Cancer 43(4):709–712
Lonigro A, Devaux JJ (2009) Disruption of neurofascin and gliomedin at nodes of Ranvier precedes demyelination in experimental allergic neuritis. Brain 132(Pt 1):260–273
Maddaluno L, Verbrugge SE, Martinoli C, Matteoli G, Chiavelli A, Zeng Y, Williams ED, Rescigno M, Cavallaro U (2009) The adhesion molecule L1 regulates transendothelial migration and trafficking of dendritic cells. J Exp Med 206(3):623–635. doi:10.1084/jem.20081211, jem.20081211 [pii]
Maness PF, Schachner M (2007) Neural recognition molecules of the immunoglobulin superfamily: signaling transducers of axon guidance and neuronal migration. Nat Neurosci 10(1):19–26. doi:10.1038/nn1827, nn1827 [pii]
Martin V, Mrkusich E, Steinel MC, Rice J, Merritt DJ, Whitington PM (2008) The L1-type cell adhesion molecule Neuroglian is necessary for maintenance of sensory axon advance in the Drosophila embryo. Neural Dev 3:10. doi:10.1186/1749-8104-3-10, 1749-8104-3-10 [pii]
Mathey EK, Derfuss T, Storch MK, Williams KR, Hales K, Woolley DR, Al-Hayani A, Davies SN, Rasband MN, Olsson T, Moldenhauer A, Velhin S, Hohlfeld R, Meinl E, Linington C (2007) Neurofascin as a novel target for autoantibody-mediated axonal injury. J Exp Med 204(10):2363–2372
Matzel LD, Babiarz J, Townsend DA, Grossman HC, Grumet M (2008) Neuronal cell adhesion molecule deletion induces a cognitive and behavioral phenotype reflective of impulsivity. Genes Brain Behav 7(4):470–480. doi:10.1111/j.1601-183X.2007.00382.x, GBB382 [pii]
Mechtersheimer S, Gutwein P, Agmon-Levin N, Stoeck A, Oleszewski M, Riedle S, Fogel M, Lemmon V, Altevogt P (2001) Ectodomain shedding of L1 adhesion molecule promotes cell migration by autocrine binding to integrins. J Cell Biol 155(4):661–673
Meier F, Busch S, Gast D, Goppert A, Altevogt P, Maczey E, Riedle S, Garbe C, Schittek B (2006) The adhesion molecule L1 (CD171) promotes melanoma progression. Int J Cancer 119(3):549–555. doi:10.1002/ijc.21880
Michelson P, Hartwig C, Schachner M, Gal A, Veske A, Finckh U (2002) Missense mutations in the extracellular domain of the human neural cell adhesion molecule L1 reduce neurite outgrowth of murine cerebellar neurons. Hum Mutat 20(6):481–482. doi:10.1002/humu.9096
Miura M, Kobayashi M, Asou H, Uyemura K (1991) Molecular cloning of cDNA encoding the rat neural cell adhesion molecule L1. Two L1 isoforms in the cytoplasmic region are produced by differential splicing. FEBS Lett 289(1):91–95
Montag-Sallaz M, Baarke A, Montag D (2003) Aberrant neuronal connectivity in CHL1-deficient mice is associated with altered information processing-related immediate early gene expression. J Neurobiol 57(1):67–80. doi:10.1002/neu.10254
Montgomery AMP, Becker JC, Siu CH, Lemmon VP, Cheresh DA, Pancook JD, Zhao XN, Reisfeld RA (1996) Human neural cell-adhesion molecule L1 and rat homolog NILE are ligands for integrin alpha(V)beta(3). J Cell Biol 132(3):475–485
Moos M, Tacke R, Scherer H, Teplow D, Früh K, Schachner M (1988) Neural adhesion molecule L1 as a member of the immunoglobulin superfamily with binding domains similar to fibronectin. Nature 334(6184):701–703
More MI, Kirsch FP, Rathjen FG (2001) Targeted ablation of NrCAM or ankyrin-B results in disorganized lens fibers leading to cataract formation. J Cell Biol 154(1):187–196
Moulding HD, Martuza RL, Rabkin SD (2000) Clinical mutations in the L1 neural cell adhesion molecule affect cell-surface expression. J Neurosci 20(15):5696–5702
Moya GE, Michaelis RC, Holloway LW, Sanchez JM (2002) Prenatal diagnosis of L1 cell adhesion molecule mutations. Capabilities and limitations. Fetal Diagn Ther 17(2):115–119, fdt17115 [pii]
Mualla R, Nagaraj K, Hortsch M (2013) A phylogenetic analysis of the L1 family of neural cell adhesion molecules. Neurochem Res 38:1196–1207
Nagaraj K, Kristiansen LV, Skrzynski A, Castiella C, Garcia-Alonso L, Hortsch M (2009) Pathogenic human L1-CAM mutations reduce the adhesion-dependent activation of EGFR. Hum Mol Genet 18(20):3822–3831
Nakakimura S, Sasaki F, Okada T, Arisue A, Cho K, Yoshino M, Kanemura Y, Yamasaki M, Todo S (2008) Hirschsprung’s disease, acrocallosal syndrome, and congenital hydrocephalus: report of 2 patients and literature review. J Pediatr Surg 43(5):E13–E17. doi:10.1016/j.jpedsurg.2007.12.069, S0022-3468(08)00002-X [pii]
Nakamura Y, Lee S, Haddox CL, Weaver EJ, Lemmon VP (2010) Role of the cytoplasmic domain of the L1 cell adhesion molecule in brain development. J Comp Neurol 518(7):1113–1132. doi:10.1002/cne.22267
Nardi JB, Pilas B, Bee CM, Zhuang S, Garsha K, Kanost MR (2006) Neuroglian-positive plasmatocytes of Manduca sexta and the initiation of hemocyte attachment to foreign surfaces. Dev Comp Immunol 30(5):447–462
Needham LK, Thelen K, Maness PF (2001) Cytoplasmic domain mutations of the L1 cell adhesion molecule reduce L1-ankyrin interactions. J Neurosci 21(5):1490–1500
Nikonenko AG, Sun M, Lepsveridze E, Apostolova I, Petrova I, Irintchev A, Dityatev A, Schachner M (2006) Enhanced perisomatic inhibition and impaired long-term potentiation in the CA1 region of juvenile CHL1-deficient mice. Eur J Neurosci 23(7):1839–1852
Nishimune H, Bernreuther C, Carroll P, Chen S, Schachner M, Henderson CE (2005) Neural adhesion molecules L1 and CHL1 are survival factors for motoneurons. J Neurosci Res 80(5):593–599
Nishimura K, Yoshihara F, Tojima T, Ooashi N, Yoon W, Mikoshiba K, Bennett V, Kamiguchi H (2003) L1-dependent neuritogenesis involves ankyrinB that mediates L1-CAM coupling with retrograde actin flow. J Cell Biol 163(5):1077–1088
Nolte C, Moos M, Schachner M (1999) Immunolocalization of the neural cell adhesion molecule L1 in epithelia of rodents. Cell Tissue Res 298(2):261–273
Novak-Hofer I, Cohrs S, Grunberg J, Friedli A, Schlatter MC, Pfeifer M, Altevogt P, Schubiger PA (2008) Antibodies directed against L1-CAM synergize with Genistein in inhibiting growth and survival pathways in SKOV3ip human ovarian cancer cells. Cancer Lett 261(2):193–204. doi:10.1016/j.canlet.2007.11.012, S0304-3835(07)00558-7 [pii]
Okamoto N, Del Maestro R, Valero R, Monros E, Poo P, Kanemura Y, Yamasaki M (2004) Hydrocephalus and Hirschsprung’s disease with a mutation of L1CAM. J Hum Genet 49(6):334–337. doi:10.1007/s10038-004-0153-4
Ooashi N, Kamiguchi H (2009) The cell adhesion molecule L1 controls growth cone navigation via ankyrin(B)-dependent modulation of cyclic AMP. Neurosci Res 63(3):224–226. doi:10.1016/j.neures.2008.11.009, S0168-0102(08)00294-0 [pii]
Panayi M, Gokhale D, Mansour S, Elles R (2005) Prenatal diagnosis in a family with X-linked hydrocephalus. Prenat Diagn 25(10):930–933. doi:10.1002/pd.1228
Panicker AK, Buhusi M, Thelen K, Maness PF (2003) Cellular signalling mechanisms of neural cell adhesion molecules. Front Biosci 8:D900–D911
Piccione M, Matina F, Fichera M, Lo Giudice M, Damiani G, Jakil MC, Corsello G (2010) A novel L1CAM mutation in a fetus detected by prenatal diagnosis. Eur J Pediatr 169(4):415–419. doi:10.1007/s00431-009-1037-6
Pillai AM, Thaxton C, Pribisko AL, Cheng JG, Dupree JL, Bhat MA (2009) Spatiotemporal ablation of myelinating glia-specific neurofascin (Nfasc NF155) in mice reveals gradual loss of paranodal axoglial junctions and concomitant disorganization of axonal domains. J Neurosci Res 87(8):1773–1793
Pomicter AD, Shroff SM, Fuss B, Sato-Bigbee C, Brophy PJ, Rasband MN, Bhat MA, Dupree JL (2010) Novel forms of neurofascin 155 in the central nervous system: alterations in paranodal disruption models and multiple sclerosis. Brain 133(Pt 2):389–405. doi:10.1093/brain/awp341, awp341 [pii]
Pratte M, Rougon G, Schachner M, Jamon M (2003) Mice deficient for the close homologue of the neural adhesion cell L1 (CHL1) display alterations in emotional reactivity and motor coordination. Behav Brain Res 147(1–2):31–39
Pruss T, Niere M, Kranz EU, Volkmer H (2004) Homophilic interactions of chick neurofascin in trans are important for neurite induction. Eur J Neurosci 20(11):3184–3188. doi:10.1111/j. 1460-9568.2004.03773.x, EJN3773 [pii]
Rathjen FG, Rutishauser U (1984) Comparison of two cell surface molecules involved in neural cell adhesion. EMBO J 3(2):461–465
Rathjen FG, Schachner M (1984) Immunocytological and biochemical characterization of a new neuronal cell surface component (L1 antigen) which is involved in cell adhesion. EMBO J 3(1):1–10
Rathjen FG, Wolff JM, Chang S, Bonhoeffer F, Raper JA (1987) Neurofascin: a novel chick cell-surface glycoprotein involved in neurite-neurite interactions. Cell 51(5):841–849
Raveh S, Gavert N, Ben-Ze’ev A (2009) L1 cell adhesion molecule (L1CAM) in invasive tumors. Cancer Lett 282(2):137–145. doi:10.1016/j.canlet.2008.12.021, S0304-3835(08)00969-5 [pii]
Rehnberg M, Jonasson J, Gunnarsson C (2011) Novel L1CAM splice site mutation in a young male with L1 syndrome. Am J Med Genet A 155A(2):439–441. doi:10.1002/ajmg.a.33803
Reid RA, Hemperly JJ (1992) Variants of human L1 cell adhesion molecule arise through alternate splicing of RNA. J Mol Neurosci 3(3):127–135
Ren Q, Bennett V (1998) Palmitoylation of neurofascin at a site in the membrane-spanning domain highly conserved among the L1 family of cell adhesion molecules. J Neurochem 70(5):1839–1849
Rodriguez Criado G, Perez Aytes A, Martinez F, Vos YJ, Verlind E, Gonzalez-Meneses Lopez A, de Terreros G, Sanchez I, Schrander-Stumpel C (2003) X-linked hydrocephalus: another two families with an L1 mutation. Genet Couns 14(1):57–65
Roonprapunt C, Huang W, Grill R, Friedlander D, Grumet M, Chen S, Schachner M, Young W (2003) Soluble cell adhesion molecule L1-Fc promotes locomotor recovery in rats after spinal cord injury. J Neurotrauma 20(9):871–882
Rosenthal A, Jouet M, Kenwrick S (1992) Aberrant splicing of neural cell adhesion molecule L1 mRNA in a family with X-linked hydrocephalus. Nat Genet 2(2):107–112
Ruiz JC, Cuppens H, Legius E, Fryns JP, Glover T, Marynen P, Cassiman JJ (1995) Mutations in L1-CAM in two families with X linked complicated spastic paraplegia, MASA syndrome, and HSAS. J Med Genet 32(7):549–552
Rünker AE, Bartsch U, Nave KA, Schachner M (2003) The C264Y missense mutation in the extracellular domain of L1 impairs protein trafficking in vitro and in vivo. J Neurosci 23(1):277–286
Ruppert M, Aigner S, Hubbe M, Yagita H, Altevogt P (1995) The L1 adhesion molecule is a cellular ligand for VLA-5. J Cell Biol 131:1881–1891
Saghatelyan AK, Nikonenko AG, Sun M, Rolf B, Putthoff P, Kutsche M, Bartsch U, Dityatev A, Schachner M (2004) Reduced GABAergic transmission and number of hippocampal perisomatic inhibitory synapses in juvenile mice deficient in the neural cell adhesion molecule L1. Mol Cell Neurosci 26(1):191–203
Sakurai K, Migita O, Toru M, Arinami T (2002) An association between a missense polymorphism in the close homologue of L1 (CHL1, CALL) gene and schizophrenia. Mol Psychiatry 7(4):412–415
Sakurai T, Ramoz N, Reichert JG, Corwin TE, Kryzak L, Smith CJ, Silverman JM, Hollander E, Buxbaum JD (2006) Association analysis of the NrCAM gene in autism and in subsets of families with severe obsessive-compulsive or self-stimulatory behaviors. Psychiatr Genet 16(6):251–257. doi:10.1097/01.ypg.0000242196.81891.c900041444-200612000-00011 [pii]
Sanes JR, Schachner M, Covault J (1986) Expression of several adhesive macromolecules (N-CAM, L1, J1, NILE, uvomorulin, laminin, fibronectin, and a heparan sulfate proteoglycan) in embryonic, adult, and denervated adult skeletal muscle. J Cell Biol 102(2):420–431
Sasakura H, Inada H, Kuhara A, Fusaoka E, Takemoto D, Takeuchi K, Mori I (2005) Maintenance of neuronal positions in organized ganglia by SAX-7, a Caenorhabditis elegans homologue of L1. EMBO J 24(7):1477–1488
Schaefer AW, Kamiguchi H, Wong EV, Beach CM, Landreth G, Lemmon V (1999) Activation of the MAPK signal cascade by the neural cell adhesion molecule L1 requires L1 internalization. J Biol Chem 274(53):37965–37973
Schmid RS, Pruitt WM, Maness PF (2000) A MAP kinase-signaling pathway mediates neurite outgrowth on L1 and requires Src-dependent endocytosis. J Neurosci 20(11):4177–4188
Schrander-Stumpel C, Legius E, Fryns JP, Cassiman JJ (1990) MASA syndrome: new clinical features and linkage analysis using DNA probes. J Med Genet 27(11):688–692
Schughart K, Kappen C, Ruddle FH (1989) Duplication of large genomic regions during the evolution of vertebrate homeobox genes. Proc Natl Acad Sci USA 86(18):7067–7071
Schürmann G, Haspel J, Grumet M, Erickson HP (2001) Cell adhesion molecule L1 in folded (horseshoe) and extended conformations. Mol Biol Cell 12(6):1765–1773
Sebens Müerköster S, Werbing V, Sipos B, Debus MA, Witt M, Grossmann M, Leisner D, Kötteritzsch J, Kappes H, Kloppel G, Altevogt P, Folsch UR, Schäfer H (2007) Drug-induced expression of the cellular adhesion molecule L1CAM confers anti-apoptotic protection and chemoresistance in pancreatic ductal adenocarcinoma cells. Oncogene 26(19):2759–2768. doi:10.1038/sj.onc.1210076, 1210076 [pii]
Senat MV, Bernard JP, Delezoide A, Saugier-Veber P, Hillion Y, Roume J, Ville Y (2001) Prenatal diagnosis of hydrocephalus-stenosis of the aqueduct of Sylvius by ultrasound in the first trimester of pregnancy. Report of two cases. Prenat Diagn 21(13):1129–1132. doi:10.1002/pd.184 [pii]
Sherman DL, Tait S, Melrose S, Johnson R, Zonta B, Court FA, Macklin WB, Meek S, Smith AJ, Cottrell DF, Brophy PJ (2005) Neurofascins are required to establish axonal domains for saltatory conduction. Neuron 48(5):737–742
Shtutman M, Levina E, Ohouo P, Baig M, Roninson IB (2006) Cell adhesion molecule L1 disrupts E-cadherin-containing adherens junctions and increases scattering and motility of MCF7 breast carcinoma cells. Cancer Res 66(23):11370–11380
Silan F, Ozdemir I, Lissens W (2005) A novel L1CAM mutation with L1 spectrum disorders. Prenat Diagn 25(1):57–59. doi:10.1002/pd.978
Silletti S, Yebra M, Perez B, Cirulli V, McMahon M, Montgomery AM (2004) Extracellular signal-regulated kinase (ERK)-dependent gene expression contributes to L1 cell adhesion molecule-dependent motility and invasion. J Biol Chem 279(28):28880–28888
Simonati A, Boaretto F, Vettori A, Dabrilli P, Criscuolo L, Rizzuto N, Mostacciuolo ML (2006) A novel missense mutation in the L1CAM gene in a boy with L1 disease. Neurol Sci 27(2):114–117. doi:10.1007/s10072-006-0610-2
Soker S, Takashima S, Miao HQ, Neufeld G, Klagsbrun M (1998) Neuropilin-1 is expressed by endothelial and tumor cells as an isoform-specific receptor for vascular endothelial growth factor. Cell 92(6):735–745, S0092-8674(00)81402-6 [pii]
Stoeck A, Gast D, Sanderson MP, Issa Y, Gutwein P, Altevogt P (2007) L1-CAM in a membrane-bound or soluble form augments protection from apoptosis in ovarian carcinoma cells. Gynecol Oncol 104(2):461–469. doi:10.1016/j.ygyno.2006.08.038, S0090-8258(06)00683-4 [pii]
Su XD, Gastinel LN, Vaughn DE, Faye I, Poon P, Bjorkman PJ (1998) Crystal structure of hemolin: a horseshoe shape with implications for homophilic adhesion. Science 281(5379):991–995
Sztriha L, Vos YJ, Verlind E, Johansen J, Berg B (2002) X-linked hydrocephalus: a novel missense mutation in the L1CAM gene. Pediatr Neurol 27(4):293–296, S088789940200440X [pii]
Tait S, Gunn-Moore F, Collinson JM, Huang J, Lubetzki C, Pedraza L, Sherman DL, Colman DR, Brophy PJ (2000) An oligodendrocyte cell adhesion molecule at the site of assembly of the paranodal axo-glial junction. J Cell Biol 150(3):657–666
Takeda Y, Asou H, Murakami Y, Miura M, Kobayashi M, Uyemura K (1996) A nonneuronal isoform of cell adhesion molecule L1: tissue-specific expression and functional analysis. J Neurochem 66(6):2338–2349
Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J, Atkins D, Wang Y (2005) Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res 11(20):7234–7242. doi:10.1158/1078-0432.CCR-05-0683, 11/20/7234 [pii]
Tapanes-Castillo A, Weaver EJ, Smith RP, Kamei Y, Caspary T, Hamilton-Nelson KL, Slifer SH, Martin ER, Bixby JL, Lemmon VP (2009) A modifier locus on chromosome 5 contributes to L1 cell adhesion molecule X-linked hydrocephalus in mice. Neurogenetics 11(1):53–71. doi:10.1007/s10048-009-0203-3
Tegay DH, Lane AH, Roohi J, Hatchwell E (2007) Contiguous gene deletion involving L1CAM and AVPR2 causes X-linked hydrocephalus with nephrogenic diabetes insipidus. Am J Med Genet A 143(6):594–598. doi:10.1002/ajmg.a.31536
Thaxton C, Pillai AM, Pribisko AL, Labasque M, Dupree JL, Faivre-Sarrailh C, Bhat MA (2010) In vivo deletion of immunoglobulin domains 5 and 6 in neurofascin (nfasc) reveals domain-specific requirements in myelinated axons. J Neurosci 30(14):4868–4876
Thies A, Schachner M, Moll I, Berger J, Schulze HJ, Brunner G, Schumacher U (2002) Overexpression of the cell adhesion molecule L1 is associated with metastasis in cutaneous malignant melanoma. Eur J Cancer 38(13):1708–1716
Triana-Baltzer GB, Liu Z, Berg DK (2006) Pre- and postsynaptic actions of L1-CAM in nicotinic pathways. Mol Cell Neurosci 33(2):214–226
Tsutsumi S, Morohashi S, Kudo Y, Akasaka H, Ogasawara H, Ono M, Takasugi K, Ishido K, Hakamada K, Kijima H (2011) L1 Cell adhesion molecule (L1CAM) expression at the cancer invasive front is a novel prognostic marker of pancreatic ductal adenocarcinoma. J Surg Oncol 103(7):669–673. doi:10.1002/jso.21880
Tsuzuki T, Izumoto S, Ohnishi T, Hiraga S, Arita N, Hayakawa T (1998) Neural cell adhesion molecule L1 in gliomas: correlation with TGF-beta and p53. J Clin Pathol 51(1):13–17
Turner KN, Schachner M, Anderson RB (2009) Cell adhesion molecule L1 affects the rate of differentiation of enteric neurons in the developing gut. Dev Dyn 238(3):708–715. doi:10.1002/dvdy.21861
Valladares A, Hernandez NG, Gomez FS, Curiel-Quezada E, Madrigal-Bujaidar E, Vergara MD, Martinez MS, Arenas Aranda DJ (2006) Genetic expression profiles and chromosomal alterations in sporadic breast cancer in Mexican women. Cancer Genet Cytogenet 170(2):147–151. doi:10.1016/j.cancergencyto.2006.06.002, S0165-4608(06)00368-2 [pii]
Vits L, Van Camp G, Coucke P, Fransen E, De Boulle K, Reyniers E, Korn B, Poustka A, Wilson G, Schrander-Stumpel C, Winter RM, Schwartz C, Willems PJ (1994) MASA syndrome is due to mutations in the neural cell adhesion gene L1CAM. Nat Genet 7(3):408–413
Vits L, Chitayat D, Van Camp G, Holden JJ, Fransen E, Willems PJ (1998) Evidence for somatic and germline mosaicism in CRASH syndrome. Hum Mutat Suppl 1:S284–S287
Volkmer H, Hassel B, Wolff JM, Frank R, Rathjen FG (1992) Structure of the axonal surface recognition molecule neurofascin and its relationship to a neural subgroup of the immunoglobulin superfamily. J Cell Biol 118(1):149–161
Volkmer H, Leuschner R, Zacharias U, Rathjen FG (1996) Neurofascin induces neurites by heterophilic interactions with axonal NrCAM while NrCAM requires F11 on the axonal surface to extend neurites. J Cell Biol 135(4):1059–1069
Vos YJ, Hofstra RM (2010) An updated and upgraded L1CAM mutation database. Hum Mutat 31(1):E1102–E1109. doi:10.1002/humu.21172
Vos YJ, de Walle HE, Bos KK, Stegeman JA, Ten Berge AM, Bruining M, van Maarle MC, Elting MW, den Hollander NS, Hamel B, Fortuna AM, Sunde LE, Stolte-Dijkstra I, Schrander-Stumpel CT, Hofstra RM (2010) Genotype-phenotype correlations in L1 syndrome: a guide for genetic counselling and mutation analysis. J Med Genet 47(3):169–175. doi:10.1136/jmg.2009.071688, jmg.2009.071688 [pii]
Wachowiak R, Fiegel HC, Kaifi JT, Quaas A, Krickhahn A, Schurr PG, Erttmann R, Schachner M, Kluth D, Sauter G, Izbicki JR (2007) L1 is associated with favorable outcome in neuroblastomas in contrast to adult tumors. Ann Surg Oncol 14(12):3575–3580. doi:10.1245/s10434-007-9608-0
Wang B, Williams H, Du JS, Terrett J, Kenwrick S (1998) Alternative splicing of human NrCAM in neural and nonneural tissues. Mol Cell Neurosci 10(5–6):287–295
Wang X, Kweon J, Larson S, Chen L (2005) A role for the C. elegans L1CAM homologue lad-1/sax-7 in maintaining tissue attachment. Dev Biol 284(2):273–291
Wang X, Zhang W, Cheever T, Schwarz V, Opperman K, Hutter H, Koepp D, Chen L (2008) The C. elegans L1CAM homologue LAD-2 functions as a coreceptor in MAB-20/Sema2 mediated axon guidance. J Cell Biol 180(1):233–246
Wei CH, Ryu SE (2012) Homophilic interaction of the L1 family of cell adhesion molecules. Exp Mol Med 44(7):413–423. doi:10.3858/emm.2012.44.7.050, emm.2012.44.050 [pii]
Wei MH, Karavanova I, Ivanov SV, Popescu NC, Keck CL, Pack S, Eisen JA, Lerman MI (1998) In silico-initiated cloning and molecular characterization of a novel human member of the L1 gene family of neural cell adhesion molecules. Hum Genet 103(3):355–364
Wei J, Hortsch M, Goode S (2004) Neuroglian stabilizes epithelial structure during Drosophila oogenesis. Dev Dyn 230(4):800–808
Weller S, Gartner J (2001) Genetic and clinical aspects of X-linked hydrocephalus (L1 disease): mutations in the L1CAM gene. Hum Mutat 18(1):1–12
Whittard JD, Sakurai T, Cassella MR, Gazdoiu M, Felsenfeld DP (2006) MAP kinase pathway-dependent phosphorylation of the L1-CAM ankyrin-binding site regulates neuronal growth. Mol Biol Cell 17(6):2696–2706
Wiencken-Barger AE, Mavity-Hudson J, Bartsch U, Schachner M, Casagrande VA (2004) The role of L1 in axon pathfinding and fasciculation. Cereb Cortex 14(2):121–131
Williams MJ (2009) The Drosophila cell adhesion molecule Neuroglian regulates Lissencephaly-1 localisation in circulating immunosurveillance cells. BMC Immunol 10:17. doi:10.1186/1471-2172-10-17, 1471-2172-10-17 [pii]
Wilson PL, Kattman BB, Mulvihill JJ, Li S, Wilkins J, Wagner AF, Goodman JR (2009) Prenatal identification of a novel R937P L1CAM missense mutation. Genet Test Mol Biomarkers 13(4):515–519. doi:10.1089/gtmb.2009.0017
Winter RM, Davies KE, Bell MV, Huson SM, Patterson MN (1989) MASA syndrome: further clinical delineation and chromosomal localisation. Hum Genet 82(4):367–370
Wolff JM, Frank R, Mujoo K, Spiro RC, Reisfeld RA, Rathjen FG (1988) A human brain glycoprotein related to the mouse cell adhesion molecule L1. J Biol Chem 263(24):11943–11947
Wong EV, Cheng GH, Payne HR, Lemmon V (1995) The cytoplasmic domain of the cell-adhesion molecule L1 is not required for homophilic adhesion. Neurosci Lett 200(3):155–158
Wood PM, Schachner M, Bunge RP (1990) Inhibition of Schwann cell myelination in vitro by antibody to the L1 adhesion molecule. J Neurosci 10(11):3635–3645
Yamamoto M, Ueda R, Takahashi K, Saigo K, Uemura T (2006) Control of axonal sprouting and dendrite branching by the nrg-ank complex at the neuron-glia interface. Curr Biol 16(16):1678–1683
Yamasaki M, Thompson P, Lemmon V (1997) CRASH syndrome: mutations in L1CAM correlate with severity of the disease. Neuropediatrics 28(3):175–178
Yip PM, Zhao X, Montgomery AM, Siu CH (1998) The Arg-Gly-Asp motif in the cell adhesion molecule L1 promotes neurite outgrowth via interaction with the alphavbeta3 integrin. Mol Biol Cell 9(2):277–290
Zecchini S, Bianchi M, Colombo N, Fasani R, Goisis G, Casadio C, Viale G, Liu J, Herlyn M, Godwin AK, Nuciforo PG, Cavallaro U (2008) The differential role of L1 in ovarian carcinoma and normal ovarian surface epithelium. Cancer Res 68(4):1110–1118. doi:10.1158/0008-5472.CAN-07-2897, 68/4/1110 [pii]
Zhang X, Davis JQ, Carpenter S, Bennett V (1998) Structural requirements for association of neurofascin with ankyrin. J Biol Chem 273(46):30785–30794
Zhao G, Hortsch M (1998) The analysis of genomic structures in the L1 family of cell adhesion molecules provides no evidence for exon shuffling events after the separation of arthropod and chordate lineages. Gene 215(1):47–55
Zhao X, Siu CH (1996) Differential effects of two hydrocephalus/MASA syndrome-related mutations on the homophilic binding and neuritogenic activities of the cell adhesion molecule L1. J Biol Chem 271(12):6563–6566
Zhao X, Yip PM, Siu CH (1998) Identification of a homophilic binding site in immunoglobulin-like domain 2 of the cell adhesion molecule L1. J Neurochem 71(3):960–971
Zhuang S, Kelo L, Nardi JB, Kanost MR (2007) Neuroglian on hemocyte surfaces is involved in homophilic and heterophilic interactions of the innate immune system of Manduca sexta. Dev Comp Immunol 31(11):1159–1167
Zonta B, Tait S, Melrose S, Anderson H, Harroch S, Higginson J, Sherman DL, Brophy PJ (2008) Glial and neuronal isoforms of Neurofascin have distinct roles in the assembly of nodes of Ranvier in the central nervous system. J Cell Biol 181(7):1169–1177. doi:10.1083/jcb.200712154, jcb.200712154 [pii]
Compliance with Ethics Requirements
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Nagaraj, K., Mualla, R., Hortsch, M. (2014). The L1 Family of Cell Adhesion Molecules: A Sickening Number of Mutations and Protein Functions. In: Berezin, V., Walmod, P. (eds) Cell Adhesion Molecules. Advances in Neurobiology, vol 8. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-8090-7_9
Download citation
DOI: https://doi.org/10.1007/978-1-4614-8090-7_9
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-8089-1
Online ISBN: 978-1-4614-8090-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)