Abstract
Cell adhesion molecules (CAMs) of the immunoglobulin superfamily (IgSF) regulate important processes such as cell proliferation, differentiation and morphogenesis. This activity is primarily due to their ability to initiate intracellular signaling cascades at cell–cell contact sites. Junctional adhesion molecule-A (JAM-A) is an IgSF-CAM with a short cytoplasmic tail that has no catalytic activity. Nevertheless, JAM-A is involved in a variety of biological processes. The functional diversity of JAM-A resides to a large part in a C-terminal PDZ domain binding motif which directly interacts with nine different PDZ domain-containing proteins. The molecular promiscuity of its PDZ domain motif allows JAM-A to recruit protein scaffolds to specific sites of cell–cell adhesion and to assemble signaling complexes at those sites. Here, we review the molecular characteristics of JAM-A, including its dimerization, its interaction with scaffolding proteins, and the phosphorylation of its cytoplasmic domain, and we describe how these characteristics translate into diverse biological activities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cells use cell–cell adhesion receptors to sense their local environment allowing them to adapt to changes in the environment. For example, when epithelial cells form new contacts with other epithelial cells, as it occurs during development or during wound healing, they stop proliferation and migration but instead start to develop new cell–cell junctions [1]. When male germ cells migrate across the seminiferous epithelium during meiosis, they sequentially undergo intimate interactions with Sertoli cells, which is required for their development from spermatogonia to spermatids [2]. Or when leukocytes transmigrate across the endothelium at sites of inflammation, they sequentially interact with endothelial cells through distinct adhesion receptors [3]. Rather than just strengthening the physical interaction between the cells, these interactions often serve to trigger intracellular signaling cascades which prime the cells for the next step in the respective chain of events.
Cell–cell adhesion receptors relay the information provided by cell–cell interactions through their cytoplasmic domains which frequently contain specific sequence motifs, such as phosphorylation consensus sites, PDZ domain-binding motifs, FERM domain-binding motifs, or proline-rich motifs. These interact with specific protein–protein interaction domains, such as SH2 domains, PDZ domains, FERM domains, or SH3 domains, respectively, in cytoplasmic proteins [4]. By interacting with cytoplasmic proteins, cell–cell adhesion receptors recruit specific proteins resulting in the assembly of larger protein complexes at sites of cell–cell contacts and the initiation of signaling events at those sites. These interactions can also serve the opposite, i.e., the recruitment of cell adhesion receptors to pre-existing macromolecular complexes. This mechanism can localize the adhesion receptors to specific sites of cell contacts, where these may undergo trans-interactions with other receptors of the opposing cell [5]. Once stabilized through these trans-interactions, the receptors can switch cytoplasmic partners thereby initiating new signaling cascades. Their interaction with cytoplasmic proteins must be considered as a highly dynamic process, which allows the cells to quickly respond to changes in the adhesive state [6].
Junctional adhesion molecule-A (JAM-A) is a member of the JAM family of cell–cell adhesion receptors [7]. JAM-A was originally identified as the receptor of a monoclonal antibody that triggers the activation of platelets [8]. However, JAM-A is expressed by a variety of different cell types including various leukocyte subsets, epithelial cells and endothelial cells, Sertoli cells, hematopoietic stem cells, and cells of the nervous system such as glial cells and neuronal progenitor cells [9]. As expected from the diversity of these cell types and tissues, JAM-A is involved in a variety of physiological processes including the regulation of the epithelial barrier function [10,11,12], the regulation of immune homeostasis and inflammation [13,14,15,16,17,18,19], hemostasis [20,21,22], hematopoiesis [23], angiogenesis [24], and the development of the central nervous system [25]. Interestingly, in some cases like platelet aggregation and hemostasis, JAM-A’s role is to inhibit intracellular signaling pathways [20,21,22] highlighting the diversity of JAM-A functions [9]. The cytoplasmic domain of JAM-A is rather short consisting of only 40 amino acid (AA) residues, which makes it unlikely that the multiple functions can be explained by interactions of the JAM-A cytoplasmic domain with many different proteins through independent regions. However, the cytoplasmic domain contains several potential phosphorylation sites, and, more importantly, it terminates in a PDZ domain binding motif (-SSFLV) [26]. All direct protein interactions hitherto identified are mediated by this short sequence motif. These observations thus suggest that the functional diversity of JAM-A can be explained to a large part by the molecular promiscuity of the C-terminal PDZ domain binding motif. In this review, we describe how this molecular promiscuity translates into functional diversity.
JAM-A: structural organization and functional motifs
JAM-A consists of two immunoglobulin (Ig)-like domains, a transmembrane domain, and a short cytoplasmic tail. Several sequence motifs have been identified to be important for JAM-A functions. These include motifs in the extracellular domain, which regulate the adhesive activity of JAM-A, and motifs in the cytoplasmic tail, which regulate the interactions with cytoplasmic proteins (Fig. 1).
JAM-A: structural motifs and functional interactions. a Principal organization of human JAM-A. The two Ig-like domains are indicated by D1 (membrane-distal, S28–K125, V-type) and D2 (membrane-proximal, P135–R228, C2 type). Disulfide bridges involve C50–C109 (D1) and C153–C212 (D2). The cis-dimerization motif (R59V60E61) and the motif involved in trans-homophilic interaction (N43N44P45) are highlighted in rose. The single N-glycosylation site (N185) is indicated by a symbol (filled circles). The two phosphorylation sites (Y280, S284) are highlighted in red. The type II PDZ domain binding motif (F297L298V299) is highlighted in green. b Cis-dimerization of human JAM-A. Cis-dimerization is mediated by salt bridges (indicated by three magenta dots) between two oppositely charged AA residues (E61···R59, R59···E61). The AA residues involved in cis-dimerization were identified by X-ray crystallography. c Trans-homophilic interaction of human JAM-A. Trans-dimerization is probably mediated predominantly by uncharged, polar residues (indicated by dotted lines) which are localized at the opposite face of the cis-dimerization motif in the D1 Ig domain. The trans-dimerization motif has been identified by structure–function analyses
The extracellular domain of JAM-A
The membrane-distal V-type Ig-like domain (D1 domain) contains two motifs which mediate adhesive interactions in cis and trans. The crystal structure indicates that JAM-A forms cis-dimers, and that the lateral association of two monomers is mediated by salt bridges between two oppositely charged AA residues (mouse JAM-A: R58···E60, E60···R58; human JAM-A: R59···E61, E61···R59; E121···K63, K63···E121) [27, 28]. The cis-dimerization motif of human JAM-A (R59V60E61) is identical in all vertebrates, and a similar motif (consensus R[V,I,L]E) is conserved in JAM-B and JAM-C. Cis-dimer formation is a prerequisite for trans-homophilic interactions, as suggested by the observations that a dimerization-deficient JAM-A mutant (JAM-A/E61R_K63E) is diffusely localized in the membrane but not enriched at cell–cell contacts [29]. The trans-homophilic interaction between two JAM-A cis-dimers on opposing cells is mediated by sequence motifs located at the opposite side of the cis-dimerization motif (Fig. 1b, c). Two motifs were identified by structure–function analyses, N43N44P45 and K97S98V99 [30], which are located within regions that were previously identified to mediate platelet aggregation [31]. Each of the two motifs is necessary for JAM-A enrichment at cell–cell junctions, but only the N43N44P45 motif promotes the clustering of coated beads, suggesting that this motif is the predominant mediator of trans-homophilic binding. N185 seems to contribute to the trans-homophilic interaction of JAM-A as well [32]. This residue, however, is located in the membrane-proximal C2-type Ig-like domain (D2 domain), which is not involved in direct contacts between the two monomers [27, 28]. Since N185 bears N-linked glycans, it is likely that glycosylation of N185 stabilizes the protein in such a way that it more efficiently undergoes trans-homophilic interactions. This is in agreement with previous results obtained using recombinant proteins and biophysical methods [33]. The present data favor the following model: JAM-A forms cis-dimers prior to any adhesive interaction, probably during the passage through the secretory pathway [27]. JAM-A cis-dimers are diffusely localized at the surface of cells as long as cells are not engaged in contact formation. When cells form cell–cell contacts, trans-homophilic interaction of JAM-A dimers followed by lateral clustering results in a network of JAM-A molecules and in strong enrichment of JAM-A at intercellular junctions (Fig. 1b, c). Similar mechanisms of clustering at intercellular junctions have been proposed for nectins and cadherins [34,35,36,37].
The cytoplasmic domain of JAM-A
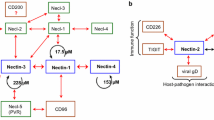
The cytoplasmic domain of JAM-A contains a C-terminal class II PDZ domain binding motif (–S296F297L298V299–COOH) [26] which is conserved in JAM-A of all vertebrate classes. All proteins hitherto identified as direct binding partners interact with JAM-A through this motif, and all direct interaction partners contain at least one PDZ domain. These proteins (summarized in Fig. 2) include afadin [38, 39], zonula occludens protein-1 (ZO-1) [38, 40, 41], ZO-2 [39], calcium/calmodulin-dependent serine protein kinase (CASK) [42], partitioning-defective 3 homolog (PAR-3) [38, 43], multiple PDZ domain protein 1 (MUPP1) [44], protein interacting with C kinase 1 (PICK-1) [45], Rap guanine nucleotide exchange factor 6 (RAPGEF6/PDZ-GEF2) [46], and RAPGEF2/PDZ-GEF1 [39]. Besides cytoplasmic scaffolding proteins, JAM-A interacts with the tetraspanin family member CD9 through its PDZ domain binding motif [47]. Since CD9 does not contain a PDZ domain in its three cytoplasmic domains, the requirement of the PDZ domain motif of JAM-A suggests an indirect interaction with CD9 that is mediated by a cytoplasmic PDZ domain protein. Together, these findings indicate a central role of the PDZ domain binding motif for the biological activities of JAM-A.
JAM-A-interacting proteins. Proteins known to interact with JAM-A through a PDZ domain-dependent interaction. Solid lines indicate direct interactions, broken lines indicate interactions that are mediated by the PDZ domain motif of JAM-A without a proof of being direct. Note that the majority of binding partners of JAM-A are scaffolding proteins containing several protein–protein interaction domains. The PDZ domains mediating the interaction with JAM-A are indicated in gray. aPKC-b aPKC-binding, RA Ras-associating, GuK guanylate kinase, L27 Lin2/Lin7
JAM-A cytoplasmic interactors: PDZ domain-containing scaffolds
All of the scaffolding proteins interacting with JAM-A are expressed by epithelial cells, and some of the proteins even co-localize at the same subcellular compartment. For example, ZO-1, ZO-2, PAR-3, and MUPP1 all co-localize at the tight junctions (TJ) of fully polarized epithelial cells [48]. These interactions, however, probably do not occur simultaneously but exist in a context-dependent manner. For example, some interactions might exist early during cell–cell contact formation, some might occur exclusively in fully polarized cells at the TJs, and some might exist only in migrating cells. In addition, the different interactions most likely occur with different affinities. PDZ domains were originally classified on the basis of COOH-terminal peptide motif binding specificities of their ligands [26, 49]. The two main (and one minor) specificity classes differed primarily in the requirement of specific AA residues at the last three to four positions of their ligands [26, 49]. More recent evidence indicates that PDZ domain interactions with their ligands involve up to seven COOH-terminal AA residues of the ligand, and that 16 specificity classes of PDZ domains exist [50]. Also, protein sequences adjacent to the interacting PDZ domain can influence the interaction with the PDZ domain ligand as well [38, 41]. Finally, binding of other proteins to the scaffold protein can influence the accessibility of its individual PDZ domain for their direct ligands [51]. Therefore, the interaction of JAM-A with a specific partner at a specific subcellular site is most likely influenced by a variety of parameters, including the relative abundance of the binding partner, the nature of the PDZ domain involved in the interaction, and the presence of proteins which allosterically regulate PDZ domain accessibility.
Afadin and guanine nucleotide exchange factors RAPGEF2 and RAPGEF6
Afadin (also called AF-6 [52]) is a multi-domain protein and exists in two major isoforms which differ in the presence of a C-terminal F-actin binding region [53] (Fig. 3). Afadin has been described to be localized at adherens junctions (AJ) where it interacts with nectins [53, 54], but also at TJs where it can interact with ZO-1 [55]. Afadin interacts with JAM-A in a direct and PDZ domain-dependent manner [38]. In recently confluent Caco-2 cells, the localization of JAM-A at cell–cell contacts depends on the PDZ domain motif of JAM-A and correlates with the presence of afadin, suggesting a possible function of afadin in the recruitment of JAM-A [38]. This function is supported by the early localization of afadin at primordial, spot-like AJs (pAJs) [56], where it can interact with ZO-1 [55, 57] as well as with α-catenin [58, 59] (Fig. 4). Thus, the interaction of JAM-A with afadin probably helps to immobilize JAM-A at pAJs from where JAM-A promotes cell–cell contact formation through a PAR-3-dependent mechanism (outlined in detail below).
Afadin/AF-6 and guanine nucleotide exchange factors RAPGEF2 and RAPGEF6. a Afadin is a scaffolding protein that contains two Ras-associating domains, one Forkhead-associated domain, one dilute domain, and one PDZ domain. The canonical isoform of afadin contains an F-actin interacting region at its C terminus. b RAPGEF2 (PDZGEF-1) and RAPGEF6 (PDZGEF-2) are guanine nucleotide exchange factors with specificities for Ras and Rap small GTPases. FHA forkhead-associated, RA Ras-associating, RGef Ras-GEF
Dynamic association of JAM-A with scaffolding proteins during epithelial junction formation. a Migrating, non-contacting cells. In migrating cells JAM-A is evenly distributed in the plasma membrane. b Primordial, spot-like adherens junctions. When cells become engaged in cell–cell contact formation, JAM-A is recruited to pAJs through its association with afadin and/or ZO-1. Afadin and ZO-1 probably act in concert in recruiting JAM-A. Trans-homophilic interactions between JAM-A on adjacent cells promotes JAM-A clustering at cell–cell junctions. JAM-A recruits Par3 and thus promotes the localization and activation of the Par3–aPKC–Par6 complex at pAJs. c Maturing junctions. The activity of the Par3–aPKC–Par6 complex promotes the maturation of cell–cell junction and the development of apico-basal polarity. JAM-A phosphorylation by aPKC is required for junction maturation. ZO-1 loses its association with afadin and interacts preferentially with JAM-A. Together with nectins, JAM-A supports the recruitment of MUPP1 and PATJ. d Mature junctions. In mature cell–cell junctions, TJs and AJs are formed and can be structurally distinguished (the TJ area is indicated by close membrane appositions). JAM-A phosphorylation occurs specifically at the TJs, which is mediated by aPKC and antagonized by PP2A. At the TJs, JAM-A is associated with several scaffolding proteins, which link JAM-A to the actin cytoskeleton and which regulate Rap signaling (see text for details). A fraction of non-phosphorylated JAM-A resides at AJs and is likely to be associated wit Afadin
More recent findings suggested a role of the JAM-A–afadin interaction during cell migration. Afadin contains two Ras-associating domains (Fig. 3), the first of which binds Ras family small GTPases like Ras and Rap1 [60]. JAM-A interacts with two guanine nucleotide exchange factors (GEF) for Rap1, RAPGEF6/PDZ-GEF2 [46] and RAPGEF2/PDZ-GEF1 [39]. Assuming that individual monomers can interact independently with distinct cytoplasmic proteins, JAM-A cis dimers could place afadin-bound Rap1 in close proximity to one of its GEFs thus promoting Rap1 activation. Rap1 is known to regulate cell adhesion and migration by acting on cell–cell and cell–matrix adhesion receptors [61, 62]. In line with a regulation of Rap1 activity by JAM-A, its downregulation reduces Rap1 activity and β1 integrin levels, and impairs cell spreading and cell migration [46, 63]. Thus, by interacting with both the Rap1 scaffold afadin and the Rap1 GEF RAPGEF6 JAM-A triggers the activity of this GTPase at sites of cell–cell contacts. This regulatory role of JAM-A might be important during collective cell migration when cell–cell junctions are dynamically remodeled to coordinate the polarization and directed cell migration of individual cells [64].
The association of JAM-A with another Rap GEF, i.e., RAPGEF2/PDZ-GEF1, has been attributed to a role in maintaining the barrier function of epithelial cells [39]. The specific Ras family small GTPase regulated by JAM-A-associated RAPGEF2 is Rap2c. In this case, however, the interaction of JAM-A with afadin seems to be indirect and bridged by ZO-2 [39] (Fig. 4d). RAPGEF2 could be associated not only directly with JAM-A but also indirectly via ZO-2. The regulation of the barrier function by the JAM-A–ZO-2–afadin–RAPGEF2 module is mediated through inhibition of RhoA-mediated actomyosin contractility. Since suppression of actomyosin contractility is also required during collective cell migration [65], the inhibition of RhoA through Rap2c by the JAM-A–ZO-2–afadin–RAPGEF2 module could contribute to collective cell migration as well.
Zonula occludens proteins ZO-1 and ZO-2
ZO-1 and ZO-2 are members of the membrane-associated guanylate kinase (MAGUK) family of scaffolding proteins, which are characterized by one or several PDZ domains that are followed by one SH3 domain and one guanylate kinase (GuK)-like domain [66]. In some cases, additional protein–protein interaction domains such as L27 or WW domains are present, and in a few cases the stereotypical order of the PDZ, SH3 and GuK domains is changed, like in MAGUK with Inverted Domain Structure (MAGI) proteins 1–3. The modular composition of MAGUK proteins of multiple protein–protein interaction domains makes them ideally suited to assemble protein complexes, and they are frequently found at sites of cell–cell contacts such as the TJ or synapses [66]. Importantly, the absence of ZO proteins in epithelial cells results in the absence of TJ strands and a complete loss of the epithelial barrier function, which is attributed to their role in assembling claudins at the TJs to allow their functional interaction and TJ strand formation [67].
Both ZO-1 and ZO-2 interact directly with JAM-A in a PDZ domain-dependent manner [38,39,40,41]. JAM-A interacts with PDZ domain 3 of ZO-1, and this interaction involves parts of the regions flanking the PDZ domain on both sides including the C-terminally localized SH3 domain [38, 41]. JAM-A interacts with PDZ domain 2 of ZO-2 [39], which is surprising since the overall organizations of ZO-1 and ZO-2 are very similar (Fig. 5), with exactly the same distance between the SH3 domain and PDZ domain 3, i.e., 14 AA in the linear sequence.
ZO-1 and ZO-2. ZO-1 and ZO-2 are members of the MAGUK family, which is characterized by the typical arrangement of PDZ domains, SH3 domains and GuK domains. ZO-1 and ZO-2 can directly interact with each other through their PDZ domains 2 (indicated by a double arrow). The region indicated by an α in ZO-1 is absent in a shorter isoform of ZO-1, which is generated by alternative splicing. The domains involved in interactions with JAM-A and with proteins relevant to the biological functions of JAM-A are indicated with arrows. GuK guanylate kinase-like, P proline-rich, SH3 Src homology 3, ZU5 ZO-1 and unc-5, α α-domain
Similar to its interaction with afadin, the interaction of JAM-A with ZO-1 could occur early during cell–cell contact formation. ZO-1 is among the early proteins localized at nascent cell–cell junctions [68] (Fig. 4b). It interacts with α-catenin [69,70,71] and could, therefore, be recruited to E-cadherin-based early contacts through E-cadherin-associated α-catenin [72]. Since its binding to α-catenin is mediated by the GuK domain, PDZ domain 3 would still be free for interacting with JAM-A (Fig. 5). As pointed out above, the JAM-A scaffold afadin also interacts with both ZO-1 and α-catenin, opening the possibility that ZO-1 and afadin act in concert in recruiting JAM-A (Fig. 4b). The interaction of JAM-A with ZO-1 could be in particular important during the step following the formation of pAJs, i.e., the development of TJs separate from AJs. This process is triggered by ZO-1 and requires the interaction of ZO-1 with α-catenin [71]. Before the formation of TJs, ZO-1 is preferentially associated with afadin. When TJs form, ZO-1 dissociates from afadin and interacts with JAM-A [73] (Fig. 4c). These findings suggest that afadin might be more important in recruiting JAM-A to pAJs during early phases of cell–cell junction formation, whereas ZO-1 may be important for recruiting JAM-A to developing TJs. In line with such a role for ZO-1 in regulating JAM-A recruitment and localization at TJs are the observations that pAJs but not TJs do form in the absence of ZO-1, and that JAM-A is lost from cell–cell contacts in the absence of ZO proteins [67, 74].
The interaction of JAM-A with ZO-2 most likely occurs at the TJs, where it serves to recruit Rap2c through afadin to form a functional JAM-A–Rap2c–RAPGEF2 complex (Fig. 3), which suppresses actomyosin contractility [39], as described above. However, given the partial redundancy of ZO-1 and ZO-2 functions [67, 75] and their similar interaction partner profile (Fig. 5), it is possible that the JAM-A–ZO-2 interaction regulates certain aspects of cell–cell contact formation as well.
The interaction of JAM-A with ZO-1 and/or ZO-2 could also serve to link JAM-A with the cytoskeleton. ZO-1 and ZO-2 both contain an F-actin-binding domain in the C-terminal part of the protein (Fig. 5). Both are not only linked to the F-actin cytoskeleton but regulate the assembly and dynamics of the cortical actin cytoskeleton [76]. The observation that JAM-A–ZO-1 complexes can be co-immunoprecipitated preferentially from Triton X-100-insoluble fraction [40] suggests that JAM-A is linked to the F-actin cytoskeleton through ZO-1. The relevance of the association with the actin cytoskeleton is still unknown. Interestingly, in endothelial cells JAM-A redistributes from cell–cell junctions to the apical membrane domain in response to pro-inflammatory stimuli [77,78,79], probably to make JAM-A available at the apical membrane domain for transient interactions with leukocyte-specific αLβ2 integrin [80, 81]. This redistribution occurs via macropinocytosis, a mechanism of endocytosis that requires a dynamic reorganization of the actin cytoskeleton [82]. The association of JAM-A with the actin cytoskeleton through ZO-1 could thus help to internalize and redistribute JAM-A during inflammation.
Cell polarity protein PAR-3
PAR-3 is a highly conserved cell polarity protein which forms a functional complex with PAR-6 and aPKC, the PAR–aPKC complex [83, 84] (Fig. 6). This complex regulates various aspects of cell polarity including the cortical polarization of the C. elegans zygote, axon formation in developing neurons, front-to-rear polarity in migrating epithelial cells, and apico-basal polarity in epithelial cells [85]. The regulation of apico-basal polarity in epithelial cells by the PAR–aPKC complex is mediated by mutual antagonistic interactions between the PAR–aPKC complex and PAR-1 kinase. In polarized epithelia, PAR-3 phosphorylation by membrane-associated PAR-1 and PAR-1 phosphorylation by membrane-associated aPKC result in cytoplasmic sequestration of PAR-3 and PAR-1, respectively, through a protein 14-3-3-dependent mechanism [86,87,88,89]. This biochemical mechanism allows the formation of distinct membrane domains without a physical intramembrane barrier.
PAR-3. PAR-3 is a highly conserved polarity protein with three conserved regions (CR). The CR1 is required for self-oligomerization and is involved in targeting PAR-3 to the TJs. The CR2 contains three PDZ domains which interact with various integral membrane proteins including JAM-A, nectins and VE-cadherin (PDZ1 and PDZ3) as well as with PAR-6, phosphoinositide lipids and PTEN. CR3 (AA 818–832) contains a sequence motif that interacts directly with the catalytic domain of aPKC and which is phosphorylated by aPKC at Ser827. The region interacting with aPKC (AA 712–936) is shaded in red. The C-terminal region contains four coiled-coil motifs which are involved in the interaction with TIAM1. CC coiled coil, CR conserved region, PI phosphoinositide
JAM-A directly interacts with PDZ domain 1 of PAR-3 [43, 90] (Fig. 6). The JAM-A–PAR-3 interaction most likely occurs at pAJs. As pointed out above, JAM-A is localized at pAJs together with E-cadherin, afadin, ZO-1 and α-catenin. The PAR–aPKC complex is not present at pAJs [90,91,92]. Shortly after the formation of pAJs, the inactive PAR–aPKC complex is recruited to pAJs by binding to JAM-A (Fig. 4). The local activity of Rho family small GTPases results in aPKC activation and the dissociation of the PAR-6–aPKC module from JAM-A-bound and Ser827-phosphorylated PAR-3 [93, 94]. Through the phosphorylation of PAR-1 [86, 87], Numb [95], Lgl [96] and JAM-A [92] (see below in more detail), aPKC promotes the maturation of cell–cell junctions and apico-basal membrane polarity [97]. In the absence of aPKC activity, pAJs still form but do not develop further into TJ and AJs [91, 97]. The interaction of PAR-3 with JAM-A thus serves to precisely localize the PAR–aPKC complex to pAJs to regulate its activation at the correct subcellular site.
In addition to its role in recruiting the PAR–aPKC complex, JAM-A could also be involved in its activation. As pointed out before, the cytoplasmic PAR–aPKC complex is inactive and requires the interaction of Rho family small GTPases like Rac1 and Cdc42 with PAR-6 for activation [98, 99]. Recent observations indicate that JAM-A can activate Cdc42 during mitosis [100], suggesting that JAM-A might cooperate with other cell–cell adhesion receptors such as E-cadherin [101,102,103] and Nectins [104] in activating Cdc42 and/or Rac1 at pAJs and thus in the activation of the PAR–aPKC complex at pAJs.
The multi-PDZ domain scaffolding proteins MUPP1 and PATJ
MUPP1 and PATJ are scaffolding proteins with multiple PDZ domains (Fig. 7). Both are localized at TJ of polarized epithelial cells [44, 105]. They contain a L27 domain at their N terminus which mediates the interaction with PALS1 [105]. PATJ is part of the Crumbs3 (CRB3) complex, which consists of the transmembrane protein Crumbs3 (CRB3) and the two scaffolding proteins Pals1 and PATJ. The Crumbs complex is required for the development of apico-basal polarity in epithelial cells both in invertebrates and vertebrates [106, 107].
MUPP1 and PATJ. MUPP1 and PATJ are structural paralogues with a highly similar organization. Both are scaffold proteins that consist almost exclusively of PDZ domains. PDZ domains in MUPP1 which are absent in PATJ are indicated by red asterisks. Note that two PDZ domains in MUPP1 which are not conserved in PATJ interact with JAM-A and CAR. CAR coxsackie and adenovirus receptor, Cld claudin, Nec nectin
JAM-A interacts with both MUPP1 and PATJ in a PDZ domain-dependent manner [44, 108]. The interaction with MUPP1 can be mediated through both PDZ domain 3 and 9, the interaction with PATJ is mediated through PDZ domain 3 [44, 108]. Since PDZ domain 9 of MUPP1 is not conserved in PATJ [108], the interaction is probably predominantly mediated by PDZ domain 3. Both MUPP1 and PATJ interact with several other proteins including integral membrane proteins like nectins, claudins, and the JAM-A-related coxsackie and adenovirus receptor (CAR) [108, 109], and with other scaffolding proteins localized at TJs, like PALS1 and ZO-3 [105, 110] (Fig. 7).
The functions of the interactions of JAM-A with MUPP1 and PATJ have not been characterized in detail. During junction formation, MUPP1 and PATJ appear later than JAM-A at nascent cell–cell contact sites, and JAM-A- or nectin-1/-2-binding-deficient mutants of MUPP1 and PATJ do not efficiently accumulate at developing cell–cell junctions [108]. These findings suggest that JAM-A cooperates with nectins in recruiting MUPP1 and PATJ at early phases during cell–cell contact formation (Fig. 4). The absence of PATJ in epithelial cells abrogates the development of functional TJs affecting both the paracellular diffusion of ions and the intramembrane diffusion barrier of proteins [108, 111, 112], which is most likely due to its role as part of the CRB3–PALS1–PATJ complex and the connection of this complex with the PAR–aPKC complex [113]. The absence of MUPP1, surprisingly, does not impair the development of functional TJs [108], suggesting that MUPP1 may only have an auxiliary role in cell–cell contact and TJ formation of epithelial cells and may be more important in assembling protein complexes at other subcellular sites known to be enriched in large protein assemblies such as synapses in neurons [114, 115]. The functional relevance of the association of JAM-A with MUPP1 and PATJ in epithelial cell–cell contact and TJ formation is still largely unexplored.
The MAGUK protein CASK
Calcium/calmodulin-dependent serine protein kinase (CASK) is a member of the MAGUK family of scaffolding proteins [66] (Fig. 8) with a central role in the assembly of protein complexes at synapses in neurons [116]. In epithelial cells, CASK localizes to the basolateral surface [117], where it interacts with the PDZ domain protein LIN-7 and with the MAGUK family member Dlg [118]. The absence of CASK in epithelial cells disturbs the basolateral localization of LIN7 and of the Dlg–Scribble polarity complex without affecting apico-basal polarity, suggesting that CASK may have redundant functions in polarized epithelia [118].
CASK. CASK is a member of the MAGUK family of scaffolding proteins. In addition to the domains characterizing the MAGUK family, it contains two Lin2/Lin7 domains and one N-terminal Ser/Thr kinase domain which exerts catalytic activity in the absence of divalent cations such as Mg2+. 4.1 protein 4.1, DLG1 disks large homolog 1, GuK guanylate kinase, L27 Lin2/Lin7, LIN abnormal cell lineage protein, PMCA4B plasma membrane Ca2+ transporting ATPase isoform 4B, Synd-2 Syndecan-1, VELI1 vertebrate Lin7 homolog 1
JAM-A directly interacts with CASK through its PDZ domain motif and the CASK PDZ domain [40]. During cell–cell contact formation induced by Ca2+-depletion/repletion (Ca2+-switch), CASK appears significantly later than JAM-A and other scaffolding proteins associating with JAM-A, such as afadin or ZO-1 [40]. CASK might, therefore, interact with JAM-A only in fully polarized epithelial cells, which might involve a fraction of JAM-A molecules that are localized basally to the TJs [92, 119]. The functional relevance of the JAM-A–CASK interaction in epithelial cells is still unclear. A JAM-A–CASK interaction has also been identified in sperm [120]. CASK interacts with JAM-A as well as with PMCA4B [121], a ten transmembrane domain-containing Ca2+ efflux pump, through its PDZ domain, and these interactions occur in the sperm flagellum. The absence of JAM-A increases CASK binding to PMCA4B, which inhibits PMCA4B activity, and as a consequence results in unbalanced intracellular Ca2+ levels and impaired sperm motility [120, 122]. The interaction of JAM-A with CASK in sperms thus serves to sequester CASK away from PMCA4B to allow functional activity of its Ca2+ efflux pump.
The scaffolding protein PICK1
Protein interacting with C kinase 1 (PICK1) is a scaffolding protein containing one PDZ domain and one Bin/amphiphysin/Rvs (BAR) domain which mediates homo-dimerization [123] (Fig. 9). Its PDZ domain can interact with integral membrane proteins, cytoplasmic proteins and with membrane-associated phosphoinositide lipids. More than 40 ligands of different classes for this PDZ domain have been identified [123], indicating that the PDZ domain of PICK1 is highly promiscuous. PICK1 is strongly expressed in the brain, where it is involved in synaptic plasticity by regulating the trafficking of glutamate receptors [124]. In epithelial cells PICK1 localizes to the basolateral membrane compartment, where it interacts with ErbB2 and EphB1 [125, 126] and regulates the stability of cell–cell junctions [126]. PICK1 also interacts with several Nectin family members as well as with JAM-A, JAM-B, and JAM-C [45].
PICK1. PICK1 is a scaffolding protein with a PDZ domain and a BAR domain. The PDZ domain is highly promiscuous and interacts with integral membrane proteins (shown below the PDZ domain), cytoplasmic proteins and phosphoinositide lipids (above the PDZ domain). The bar domain mediates homodimerization as well a phosphoinositide lipid binding. BAR Bin/amphiphysin/Rvs, PI phosphoinositide
The interaction of PICK1 with JAM-A is most likely direct and PDZ domain dependent [45]. Since PICK1 localizes to the basolateral membrane compartment in polarized epithelial cells, the interaction with JAM-A most likely occurs at the basal part of cell–cell contacts and thus involves the fraction of JAM-A molecules that is localized basally to the TJs. Apart from the observations that JAM-A exists with PICK1 in a protein complex as evidenced by co-immunoprecipitation [45], no additional information on the JAM-A–PICK1 interaction is available. The functional relevance of this interaction is, therefore, unclear.
JAM-A lateral interactors: tetraspanins and integrins
CD9 is a member of the tetraspanin (Tspn) family which contains 33 members in mammals [127]. Tspns are type II proteins characterized by two extracellular loops, four transmembrane domains, and three cytoplasmic domains (Fig. 10). Some structural features distinguish Tspns from other proteins with four transmembrane domains like claudins or MarvelD. These include a number of AA residues which are conserved in Tspns but not other proteins. These comprise in particular amino acids present in the small intracellular loop as well as the CCG motif and two additional Cys residues in the large extracellular loop, which are involved in the formation of disulfide bonds [128] (Fig. 10). One of the most distinctive functional features of Tspns is their ability to form lateral associations with partner proteins both through their extracellular domains, which mediate interactions in particular integrins and Ig superfamily members, and through their transmembrane domains, which mediate Tspn–Tspn associations. These properties combined with their ability to bind specific cytoplasmic proteins through their cytoplasmic domains highlight the principal function of Tspns, i.e., the formation of a platform for proteins to allow their physical and functional interaction [129].
Tetraspanin CD9 and αVβ3 integrin. a Tetraspanins consist of four transmembrane domains, three cytoplasmic domains which include the N and C termini and a small loop, and two extracellular loops, a small extracellular loop (SEL) and a large extracellular loop (LEL). The CCG motif and two additional Cys residues in the LEL (C152C153G154, C167 and C181 in CD9) are involved in SS bond formation and are the most distinctive structural feature of tetraspanins. b A ternary JAM-A–CD9–αVβ3 integrin complex. CD9 serves as scaffold to link JAM-A to αVβ3 integrin. JAM-A interacts with αVβ3 integrin most likely through a PDZ domain protein (blue) that has not been identified yet. The interaction between αVβ3 integrin and CD9 most likely involves the extracellular domains of the proteins (indicated by a dotted line double arrow). SEL small extracellular loop, LEL large extracellular loop
JAM-A interacts with tetraspanin CD9 in platelets and in endothelial cells [47, 130]. The interaction with CD9 requires the PDZ domain motif of JAM-A [47]. CD9 does not contain a PDZ domain in its cytoplasmic domains to which JAM-A could bind, but contains a C-terminal type III PDZ domain motif (–R225E226M227V228–COOH) [131] suggesting that the interaction is mediated by a cytoplasmic PDZ domain protein. The interaction between JAM-A and CD9 is stable in the presence of detergents like Triton X-100, which disrupts Tspn–Tspn interactions [132], strongly suggesting that the putative PDZ domain protein directly connects JAM-A with CD9.
In endothelial cells, the interaction of JAM-A with CD9 most likely serves to assemble a ternary complex consisting of JAM-A, CD9 and αVβ3 integrin, which regulates p44/42 MAPK activation in response to bFGF signaling [47, 133]. How JAM-A, which dissociates from CD9 and αVβ3 integrin in response to bFGF [47, 133], mediates this activity is still unclear. In the absence of JAM-A or CD9, cells fail to respond to bFGF [24, 47] suggesting that an intact ternary JAM-A–CD9–αVβ3 integrin complex and the release of JAM-A is necessary for bFGF to mount a proper MAPK activation. One potential mechanism would be that the release of JAM-A, which is predominantly present as a monomer in the complex [47], allows the formation of a functional and signaling-active JAM-A dimer. A similar mechanism has been described for the JAM-A-related protein JAML [134].
JAM-A phosphorylation
The cytoplasmic domain of human JAM-A contains two tyrosine residues and eleven serine or threonine residues (Fig. 11). Mass spectrometry analyses identified phosphorylation of Y280 in different cell types including lung cancer cell lines, epithelial cells, and endothelial cells [135,136,137], as well as phosphorylations of S284, S287 and S296 in HeLa cells, HEK293 cells, liver cells, and platelets [138,139,140,141]. Two of these phosphorylations, i.e., Y280 and S284, have so far been identified to be functionally important. Y280 is phosphorylated in endothelial cells and in platelets by as yet unidentified kinases [21, 133]. S284 is phosphorylated by a conventional PKC isoform (PKCα) in platelets [142], and by an atypical PKC isoform (aPKCζ) in epithelial cells [92] (Fig. 11).
Cytoplasmic domain of JAM-A. a Phosphorylation sites in the cytoplasmic domain of JAM-A. Experimentally identified phosphorylation sites are colored. Kinases identified to mediate phosphorylations are indicated. b Sequence conservation of the cytoplasmic domain of JAM-A in vertebrates. Dr, Danio rerio; Gg, Gallus gallus; Hs, Homo sapiens; Mm, Mus musculus; Ps, Pelodiscus sinensis (Chinese softshell turtle); Rn, Rattus norvegicus; TM, transmembrane; Xl, Xenopus laevis
JAM-A phosphorylation at Tyr280
The evidence suggesting a role for Tyr280 phosphorylation has been obtained from ectopic expression studies. As pointed out before, bFGF fails to mount a p44/42 MAPK response in endothelial cells which lack JAM-A expression [47, 133]. Both a JAM-A mutant that lacks the cytoplasmic domain and a JAM-A mutant that lacks Tyr280 (JAM-A/Y280F) act in a dominant-negative way in this assay suggesting that phosphorylation of Tyr280 is involved in JAM-A’s signaling activity during bFGF-mediated p44/42 MAPK activation in endothelial cells (as described above).
More direct evidence for JAM-A Tyr280 phosphorylation and in particular its functional role has been obtained in platelets, in which JAM-A is associated with αIIbβ3 integrin [21, 130]. As analyzed using P-Tyr-specific antibodies, αIIbβ3 integrin-associated JAM-A is tyrosine phosphorylated in the absence of stimulation [21]. Tyr-phosphorylated JAM-A binds c-Src kinase (Csk), a kinase that negatively regulates the activity of c-Src [143]. Since the cytoplasmic tail of JAM-A contains only two tyrosine residues (Fig. 11), and since experimental evidence supporting tyrosine phosphorylation exists only for Tyr280 [135,136,137], it is likely that Csk binds to the P-Tyr280 residue of JAM-A, possibly through its SH2 domain [144]. Platelet stimulation with agonists such as fibrinogen or thrombin results in reduced Tyr phosphorylation of JAM-A and its dissociation from αIIbβ3 integrin [21, 22]. Interestingly, in resting platelets as well as in fibrinogen-stimulated platelets JAM-A is associated with the tyrosine-protein phosphatase non-receptor type 1 (PTPN1), and specific inhibition of PTPN1 prevents fibrinogen-stimulated JAM-A dephosphorylation [22], strongly suggesting that PTPN1 is responsible for JAM-A dephosphorylation in response to platelet activation. These observations thus suggest that JAM-A tyrosine phosphorylation in platelets prevents premature platelet activation in the absence of agonists by linking Csk to αIIbβ3 integrin-associated c-Src [145]. In line with this model, the absence of JAM-A in mice results in platelet hyperreactivity, enhanced thrombosis and atherosclerosis [19, 22, 146].
JAM-A phosphorylation at Ser284
In platelets, Ser284 (Ser285 in mice, Fig. 11) of JAM-A is phosphorylated by PKCα (or other conventional PKC isoforms) in response to agonists such as thrombin or collagen [142]. Phosphorylation is rapid and peaks 1 min after thrombin stimulation. The kinetics of Ser284 phosphorylation correlates with the accumulation of JAM-A as well as the translocation of PKC isoforms to sites of cell–cell contact in platelet aggregates [142], suggesting that Ser284 phosphorylation occurs at platelet–platelet contact sites. The specific function of JAM-A Ser284 phosphorylation in platelets is not known yet.
In epithelial cells, Ser284 of JAM-A is phosphorylated by aPKCζ [92]. This phosphorylation occurs early during cell–cell contact formation and correlates with the localization of the PAR–aPKC complex at pAJs. Given the function of JAM-A in recruiting PAR-3 to pAJs [90], this suggests that JAM-A serves both as positional cue for the localization of the PAR–aPKC complex and as substrate after aPKC has been activated at pAJs. Ser284 phosphorylation is important for junctional maturation and TJ formation as indicated by the delay in cell–cell contact formation and impaired barrier formation, respectively, in cells expressing a JAM-A S285A mutant. In fully polarized epithelial cells, Ser284 phosphorylation of JAM-A is restricted to TJ, suggesting that continuous Ser284 phosphorylation is required for the maintenance of the epithelial barrier [92]. JAM-A Ser284 phosphorylation at TJs is antagonized by PP2A, a phosphatase localized at TJs. Interestingly, PP2A also directly interacts and dephosphorylates aPKC as well as other TJ proteins, and ectopic expression of its catalytic subunit results in TJ disassembly and increased paracellular permeability [147, 148]. These observations suggest that JAM-A phosphorylation at Ser284 is balanced by the aPKC–PP2A module. It is, therefore, possible that JAM-A phosphorylation not only regulates TJ formation, but that JAM-A phosphorylation is altered during pathological conditions which are associated with a loss of the epithelial barrier function, such as epithelial-to-mesenchymal transition (EMT) [149], hypoxia [150], or the presence of pathogens [151, 152].
Phosphorylation of JAM-A at Ser284 has recently been described to be involved in the function of JAM-A as mechanosensor in endothelial cells. When mechanical forces are applied on JAM-A, or, alternatively, when endothelial cells are exposed to shear stress, the cells respond by increasing the activity of RhoA [153], suggesting that JAM-A acts as a force sensor which transmits mechanical force-triggered signals to RhoA. The increased RhoA activity elicited by either of the two mechanical stimulations depends on phosphorylation of JAM-A at Ser284. Interestingly, the force-on-JAM-A-triggered JAM-A Ser284 phosphorylation is preceded by activation of aPKCζ, and force-on-JAM-A-triggered RhoA activation is abolished after inhibition of aPKCζ [153]. These observations not only illustrate a novel role of JAM-A as mechanosensor in endothelial cells but also highlight the importance of aPKCζ-mediated Ser284 phosphorylation during this process.
Conclusions
JAM-A is a cell–cell adhesion molecule which is expressed by a variety of cell types and which has adopted a variety of physiological functions in vertebrates [9]. In light of these pleiotropic functions the structural organization of JAM-A appears rather simple. Its first Ig domain contains two motifs which regulate cis-dimerization and trans-homophilic interaction. Its cytoplasmic tail with only 40 AA residues contains four phosphorylation sites and a C-terminal PDZ domain binding motif. From the four sites of phosphorylation that have been experimentally identified so far, evidence for a functional relevance of these phosphorylations exists for two, i.e., Tyr280 and Ser284. Kinases that are responsible for the phosphorylations have only been identified for Ser284. Therefore, it will be important to identify the kinases responsible for the three other sites of phosphorylation. With respect to Tyr280, which is a possible binding site for Csk, it is tempting to speculate that its phosphorylation in platelets is mediated by c-Src, as a mechanism of phosphorylation by c-Src to generate binding sites for Csk has been described in endothelial cells for VE-cadherin Tyr685 [154]. The C-terminal PDZ domain binding motif is surprisingly promiscuous. At least eight different proteins directly interact with JAM-A through this motif. This molecular promiscuity explains in part the functional pleiotropy of JAM-A. Many interactions have not been studied in enough detail to understand their functional relevance. It is also possible that phosphorylations influence the binding specificity of the PDZ domain binding motif thereby contributing to its promiscuity. In particular, Ser296 at position minus 3 from the COOH terminus (the COOH-terminal Val299 being defined as position 0 in the PDZ domain motif nomenclature [26] (Fig. 1), which has been identified to be phosphorylated in HEK293T cells by mass spectrometry [141], is likely to contribute to the interaction of JAM-A with PDZ domains [50]. It is to be expected that more molecular mechanisms which regulate the physiological functions of JAM-A will be revealed in the future.
Abbreviations
- AA:
-
Amino acid
- AJ:
-
Adherens junctions
- aPKC:
-
Atypical protein kinase C
- BAR:
-
Bin/amphiphysin/RVS
- CAR:
-
Coxsackie and adenovirus receptor
- CASK:
-
Calcium/calmodulin-dependent serine protein kinase
- Cdc42:
-
Cell division cycle 42
- Csk:
-
C-Src kinase
- EMT:
-
Epithelial-to-mesenchymal transition
- FERM:
-
4.1 protein and ERM
- GEF:
-
Guanine nucleotide exchange factor
- IgSF:
-
Immunoglobulin superfamily
- JAM:
-
Junctional adhesion molecule
- LGL:
-
Lethal(2) giant larvae protein homolog
- LIN:
-
Abnormal cell lineage protein
- MAPK:
-
Mitogen-activated protein kinase
- MUPP1:
-
Multiple PDZ domain protein 1
- pAJs:
-
Primordial, spot-like adherens junctions
- Pals1:
-
Protein associated with Lin-7
- PAR:
-
Partitioning defective
- PATJ:
-
Pals1-associated tight junction protein
- PICK1:
-
Protein interacting with C kinase 1
- PMCA4B:
-
Plasma membrane calcium-transporting ATPase 4
- PDZ:
-
PSD95–Discs large–ZO-1
- RAC1:
-
Ras-related C3 botulinum toxin substrate 1
- RAPGEF:
-
Rap guanine nucleotide exchange factor
- SH:
-
Src homology
- Src:
-
Sarcoma
- TJ:
-
Tight junctions
- ZO:
-
Zonula occludens
References
Yeaman C, Grindstaff KK, Nelson WJ (1999) New perspectives on mechanisms involved in generating epithelial cell polarity. Physiol Rev 79(1):73–98
Cheng CY, Mruk DD (2002) Cell junction dynamics in the testis: sertoli-germ cell interactions and male contraceptive development. Physiol Rev 82(4):825–874. https://doi.org/10.1152/physrev.00009.2002
Vestweber D (2015) How leukocytes cross the vascular endothelium. Nat Rev Immunol 15(11):692–704. https://doi.org/10.1038/nri3908
Pawson T, Nash P (2003) Assembly of cell regulatory systems through protein interaction domains. Science 300(5618):445–452
Famulski JK, Trivedi N, Howell D, Yang Y, Tong Y, Gilbertson R, Solecki DJ (2010) Siah regulation of Pard3A controls neuronal cell adhesion during germinal zone exit. Science 330(6012):1834–1838
Takeichi M (2014) Dynamic contacts: rearranging adherens junctions to drive epithelial remodelling. Nat Rev Mol Cell Biol 15(6):397–410. https://doi.org/10.1038/nrm3802
Martin-Padura I, Lostaglio S, Schneemann M, Williams L, Romano M, Fruscella P, Panzeri C, Stoppacciaro A, Ruco L, Villa A, Simmons D, Dejana E (1998) Junctional adhesion molecule, a novel member of the immunoglobulin superfamily that distributes at intercellular junctions and modulates monocyte transmigration. J Cell Biol 142(1):117–127
Kornecki E, Walkowiak B, Naik UP, Ehrlich YH (1990) Activation of human platelets by a stimulatory monoclonal antibody. J Biol Chem 265(17):10042–10048
Ebnet K (2017) Junctional adhesion molecules (JAMs): cell adhesion receptors with pleiotropic functions in cell physiology and development. Physiol Rev 97(4):1529–1554. https://doi.org/10.1152/physrev.00004.2017
Laukoetter MG, Nava P, Lee WY, Severson EA, Capaldo CT, Babbin BA, Williams IR, Koval M, Peatman E, Campbell JA, Dermody TS, Nusrat A, Parkos CA (2007) JAM-A regulates permeability and inflammation in the intestine in vivo. J Exp Med 204(13):3067–3076
Mitchell LA, Ward C, Kwon M, Mitchell PO, Quintero DA, Nusrat A, Parkos CA, Koval M (2015) Junctional adhesion molecule A promotes epithelial tight junction assembly to augment lung barrier function. Am J Pathol 185(2):372–386. https://doi.org/10.1016/j.ajpath.2014.10.010
Khounlotham M, Kim W, Peatman E, Nava P, Medina-Contreras O, Addis C, Koch S, Fournier B, Nusrat A, Denning TL, Parkos CA (2012) Compromised intestinal epithelial barrier induces adaptive immune compensation that protects from colitis. Immunity 37(3):563–573. https://doi.org/10.1016/j.immuni.2012.06.017
Del Maschio A, De Luigi A, Martin-Padura I, Brockhaus M, Bartfai T, Fruscella P, Adorini L, Martino G, Furlan R, De Simoni MG, Dejana E (1999) Leukocyte recruitment in the cerebrospinal fluid of mice with experimental meningitis is inhibited by an antibody to Junctional Adhesion Molecule (JAM). J Exp Med 190(9):1351–1356
Cera MR, Del Prete A, Vecchi A, Corada M, Martin-Padura I, Motoike T, Tonetti P, Bazzoni G, Vermi W, Gentili F, Bernasconi S, Sato TN, Mantovani A, Dejana E (2004) Increased DC trafficking to lymph nodes and contact hypersensitivity in junctional adhesion molecule-A-deficient mice. J Clin Investig 114(5):729–738
Corada M, Chimenti S, Cera MR, Vinci M, Salio M, Fiordaliso F, De Angelis N, Villa A, Bossi M, Staszewsky LI, Vecchi A, Parazzoli D, Motoike T, Latini R, Dejana E (2005) Junctional adhesion molecule-A-deficient polymorphonuclear cells show reduced diapedesis in peritonitis and heart ischemia–reperfusion injury. Proc Natl Acad Sci USA 102(30):10634–10639
Khandoga A, Kessler JS, Meissner H, Hanschen M, Corada M, Motoike T, Enders G, Dejana E, Krombach F (2005) Junctional adhesion molecule-A deficiency increases hepatic ischemia–reperfusion injury despite reduction of neutrophil transendothelial migration. Blood 106(2):725–733
Vetrano S, Rescigno M, Rosaria Cera M, Correale C, Rumio C, Doni A, Fantini M, Sturm A, Borroni E, Repici A, Locati M, Malesci A, Dejana E, Danese S (2008) Unique role of junctional adhesion molecule-A in maintaining mucosal homeostasis in inflammatory bowel disease. Gastroenterology 135(1):173–184
Cera MR, Fabbri M, Molendini C, Corada M, Orsenigo F, Rehberg M, Reichel CA, Krombach F, Pardi R, Dejana E (2009) JAM-A promotes neutrophil chemotaxis by controlling integrin internalization and recycling. J Cell Sci 122(Pt 2):268–277
Lakshmi SP, Reddy AT, Naik MU, Naik UP, Reddy RC (2012) Effects of JAM-A deficiency or blocking antibodies on neutrophil migration and lung injury in a murine model of ALI. Am J Physiol Lung Cell Mol Physiol 303(9):L758–L766. https://doi.org/10.1152/ajplung.00107.2012
Naik MU, Stalker TJ, Brass LF, Naik UP (2012) JAM-A protects from thrombosis by suppressing integrin alphaIIbbeta3-dependent outside-in signaling in platelets. Blood 119(14):3352–3360
Naik MU, Caplan JL, Naik UP (2014) Junctional adhesion molecule-A suppresses platelet integrin alphaIIbbeta3 signaling by recruiting Csk to the integrin-c-Src complex. Blood 123(9):1393–1402. https://doi.org/10.1182/blood-2013-04-496232
Karshovska E, Zhao Z, Blanchet X, Schmitt MM, Bidzhekov K, Soehnlein O, von Hundelshausen P, Mattheij NJ, Cosemans JM, Megens RT, Koeppel TA, Schober A, Hackeng TM, Weber C, Koenen RR (2015) Hyperreactivity of junctional adhesion molecule a-deficient platelets accelerates atherosclerosis in hyperlipidemic mice. Circ Res 116(4):587–599. https://doi.org/10.1161/CIRCRESAHA.116.304035
Kobayashi I, Kobayashi-Sun J, Kim AD, Pouget C, Fujita N, Suda T, Traver D (2014) Jam1a-Jam2a interactions regulate haematopoietic stem cell fate through Notch signalling. Nature 512(7514):319–323. https://doi.org/10.1038/nature13623
Cooke VG, Naik MU, Naik UP (2006) Fibroblast growth factor-2 failed to induce angiogenesis in junctional adhesion molecule-A-deficient mice. Arterioscler Thromb Vasc Biol 26(9):2005–2011
Fededa JP, Esk C, Mierzwa B, Stanyte R, Yuan S, Zheng H, Ebnet K, Yan W, Knoblich JA, Gerlich DW (2016) MicroRNA-34/449 controls mitotic spindle orientation during mammalian cortex development. EMBO J 35(22):2386–2398. https://doi.org/10.15252/embj.201694056
Songyang Z, Fanning AS, Fu C, Xu J, Marfatia SM, Chisti AH, Crompton A, Chan AC, Anderson JM, Cantley LC (1997) Recognition of unique carboxy-terminal motifs by distinct PDZ domains. Science 275:73–77
Kostrewa D, Brockhaus M, D’Arcy A, Dale GE, Nelboeck P, Schmid G, Mueller F, Bazzoni G, Dejana E, Bartfai T, Winkler FK, Hennig M (2001) X-ray structure of junctional adhesion molecule: structural basis for homophilic adhesion via a novel dimerization motif. EMBO J 20(16):4391–4398
Prota AE, Campbell JA, Schelling P, Forrest JC, Watson MJ, Peters TR, Aurrand-Lions M, Imhof BA, Dermody TS, Stehle T (2003) Crystal structure of human junctional adhesion molecule 1: implications for reovirus binding. Proc Natl Acad Sci USA 100(9):5366–5371
Mandell KJ, McCall IC, Parkos CA (2004) Involvement of the junctional adhesion molecule-1 (JAM1) homodimer interface in regulation of epithelial barrier function. J Biol Chem 279(16):16254–16262
Monteiro AC, Luissint AC, Sumagin R, Lai C, Vielmuth F, Wolf MF, Laur O, Reiss K, Spindler V, Stehle T, Dermody TS, Nusrat A, Parkos CA (2014) Trans-dimerization of JAM-A regulates Rap2 and is mediated by a domain that is distinct from the cis-dimerization interface. Mol Biol Cell 25(10):1574–1585. https://doi.org/10.1091/mbc.E14-01-0018
Babinska A, Kedees MH, Athar H, Sobocki T, Sobocka MB, Ahmed T, Ehrlich YH, Hussain MM, Kornecki E (2002) Two regions of the human platelet F11-receptor (F11R) are critical for platelet aggregation, potentiation and adhesion. Thromb Haemost 87(4):712–721
Scott DW, Tolbert CE, Graham DM, Wittchen E, Bear JE, Burridge K (2015) N-glycosylation controls the function of junctional adhesion molecule-A. Mol Biol Cell 26(18):3205–3214. https://doi.org/10.1091/mbc.E14-12-1604
Wojcikiewicz EP, Koenen RR, Fraemohs L, Minkiewicz J, Azad H, Weber C, Moy VT (2009) LFA-1 binding destabilizes the JAM-A homophilic interaction during leukocyte transmigration. Biophys J 96(1):285–293
Pertz O, Bozic D, Koch AW, Fauser C, Brancaccio A, Engel J (1999) A new crystal structure, Ca2+ dependence and mutational analysis reveal molecular details of E-cadherin homoassociation. EMBO J 18(7):1738–1747
Narita H, Yamamoto Y, Suzuki M, Miyazaki N, Yoshida A, Kawai K, Iwasaki K, Nakagawa A, Takai Y, Sakisaka T (2011) Crystal Structure of the cis-Dimer of Nectin-1: implications for the architecture of cell–cell junctions. J Biol Chem 286(14):12659–12669
Harrison OJ, Jin X, Hong S, Bahna F, Ahlsen G, Brasch J, Wu Y, Vendome J, Felsovalyi K, Hampton CM, Troyanovsky RB, Ben-Shaul A, Frank J, Troyanovsky SM, Shapiro L, Honig B (2011) The extracellular architecture of adherens junctions revealed by crystal structures of type I cadherins. Structure 19(2):244–256. https://doi.org/10.1016/j.str.2010.11.016
Harrison OJ, Vendome J, Brasch J, Jin X, Hong S, Katsamba PS, Ahlsen G, Troyanovsky RB, Troyanovsky SM, Honig B, Shapiro L (2012) Nectin ectodomain structures reveal a canonical adhesive interface. Nat Struct Mol Biol 19(9):906–915. https://doi.org/10.1038/nsmb.2366
Ebnet K, Schulz CU, Meyer Zu Brickwedde MK, Pendl GG, Vestweber D (2000) Junctional adhesion molecule interacts with the PDZ domain-containing proteins AF-6 and ZO-1. J Biol Chem 275(36):27979–27988. https://doi.org/10.1074/jbc.m002363200
Monteiro AC, Sumagin R, Rankin CR, Leoni G, Mina MJ, Reiter DM, Stehle T, Dermody TS, Schaefer SA, Hall RA, Nusrat A, Parkos CA (2013) JAM-A associates with ZO-2, afadin, and PDZ-GEF1 to activate Rap2c and regulate epithelial barrier function. Mol Biol Cell 24(18):2849–2860. https://doi.org/10.1091/mbc.E13-06-0298
Bazzoni G, Martinez-Estrada OM, Orsenigo F, Cordenonsi M, Citi S, Dejana E (2000) Interaction of junctional adhesion molecule with the tight junction components ZO-1, cingulin, and occludin. J Biol Chem 275(27):20520–20526
Nomme J, Fanning AS, Caffrey M, Lye MF, Anderson JM, Lavie A (2011) The Src homology 3 domain is required for junctional adhesion molecule binding to the third PDZ domain of the scaffolding protein ZO-1. J Biol Chem 286(50):43352–43360
Martinez-Estrada OM, Villa A, Breviario F, Orsenigo F, Dejana E, Bazzoni G (2001) Association of junctional adhesion molecule with calcium/calmodulin-dependent serine protein kinase (CASK/LIN-2) in human epithelial Caco-2 cells. J Biol Chem 276(12):9291–9296
Itoh M, Sasaki H, Furuse M, Ozaki H, Kita T, Tsukita S (2001) Junctional adhesion molecule (JAM) binds to PAR-3: a possible mechanism for the recruitment of PAR-3 to tight junctions. J Cell Biol 154(3):491–498
Hamazaki Y, Itoh M, Sasaki H, Furuse M, Tsukita S (2002) Multi-PDZ domain protein 1 (MUPP1) is concentrated at tight junctions through its possible interaction with claudin-1 and junctional adhesion molecule. J Biol Chem 277(1):455–461
Reymond N, Garrido-Urbani S, Borg JP, Dubreuil P, Lopez M (2005) PICK-1: a scaffold protein that interacts with Nectins and JAMs at cell junctions. FEBS Lett 579(10):2243–2249
Severson EA, Lee WY, Capaldo CT, Nusrat A, Parkos CA (2009) Junctional adhesion molecule A interacts with Afadin and PDZ-GEF2 to activate Rap1A, regulate beta1 integrin levels, and enhance cell migration. Mol Biol Cell 20(7):1916–1925
Peddibhotla SS, Brinkmann BF, Kummer D, Tuncay H, Nakayama M, Adams RH, Gerke V, Ebnet K (2013) Tetraspanin CD9 links junctional adhesion molecule-A to alphavbeta3 integrin to mediate basic fibroblast growth factor-specific angiogenic signaling. Mol Biol Cell 24(7):933–944. https://doi.org/10.1091/mbc.E12-06-0481
Ebnet K (2008) Organization of multiprotein complexes at cell-cell junctions. Histochem Cell Biol 130(1):1–20
Stricker NL, Christopherson KS, Yi BA, Schatz PJ, Raab RW, Dawes G, Bassett DE Jr, Bredt DS, Li M (1997) PDZ domain of neuronal nitric oxide synthase recognizes novel C-terminal peptide sequences. Nat Biotechnol 15(4):336–342. https://doi.org/10.1038/nbt0497-336
Tonikian R, Zhang Y, Sazinsky SL, Currell B, Yeh JH, Reva B, Held HA, Appleton BA, Evangelista M, Wu Y, Xin X, Chan AC, Seshagiri S, Lasky LA, Sander C, Boone C, Bader GD, Sidhu SS (2008) A specificity map for the PDZ domain family. PLoS Biol 6(9):e239. https://doi.org/10.1371/journal.pbio.0060239
Morales FC, Takahashi Y, Momin S, Adams H, Chen X, Georgescu MM (2007) NHERF1/EBP50 head-to-tail intramolecular interaction masks association with PDZ domain ligands. Mol Cell Biol 27(7):2527–2537. https://doi.org/10.1128/MCB.01372-06
Prasad R, Gu Y, Alder H, Nakamura T, Canaani O, Saito H, Huebner K, Gale RP, Nowell PC, Kuriyama K et al (1993) Cloning of the ALL-1 fusion partner, the AF-6 gene, involved in acute myeloid leukemias with the t(6;11) chromosome translocation. Cancer Res 53(23):5624–5628
Mandai K, Nakanishi H, Satoh A, Obaishi H, Wada M, Nishioka H, Itoh M, Mizoguchi A, Aoki T, Fujimoto T, Matsuda Y, Tsukita S, Takai Y (1997) Afadin: a novel actin filament-binding protein with one PDZ domain localized at cadherin-based cell-to-cell adherens junction [published erratum appears in J Cell Biol 1997 Nov 17;139(4):1060]. J Cell Biol 139(2):517–528
Takahashi K, Nakanishi H, Miyahara M, Mandai K, Satoh K, Satoh A, Nishioka H, Aoki J, Nomoto A, Mizoguchi A, Takai Y (1999) Nectin/PRR: an immunoglobulin-like cell adhesion molecule recruited to cadherin-based adherens junctions through interaction with Afadin, a PDZ domain-containing protein. J Cell Biol 145(3):539–549
Yamamoto T, Harada N, Kano K, Taya S, Canaani E, Matsuura Y, Mizoguchi A, Ide C, Kaibuchi K (1997) The Ras target AF-6 interacts with ZO-1 and serves as a peripheral component of tight junctions in epithelial cells. J Cell Biol 139(3):785–795
Asakura T, Nakanishi H, Sakisaka T, Takahashi K, Mandai K, Nishimura M, Sasaki T, Takai Y (1999) Similar and differential behaviour between the nectin-afadin-ponsin and cadherin-catenin systems during the formation and disruption of the polarized junctional alignment in epithelial cells. Genes Cells 4(10):573–581
Yokoyama S, Tachibana K, Nakanishi H, Yamamoto Y, Irie K, Mandai K, Nagafuchi A, Monden M, Takai Y (2001) alpha-catenin-independent recruitment of ZO-1 to nectin-based cell–cell adhesion sites through afadin. Mol Biol Cell 12(6):1595–1609
Tachibana K, Nakanishi H, Mandai K, Ozaki K, Ikeda W, Yamamoto Y, Nagafuchi A, Tsukita S, Takai Y (2000) Two cell adhesion molecules, nectin and cadherin, interact through their cytoplasmic domain-associated proteins. J Cell Biol 150(5):1161–1176
Pokutta S, Drees F, Takai Y, Nelson WJ, Weis WI (2002) Biochemical and structural definition of the l-afadin- and actin-binding sites of alpha-catenin. J Biol Chem 277(21):18868–18874
Boettner B, Govek EE, Cross J, Van Aelst L (2000) The junctional multidomain protein AF-6 is a binding partner of the Rap1A GTPase and associates with the actin cytoskeletal regulator profilin. Proc Natl Acad Sci USA 97(16):9064–9069
Kooistra MR, Dube N, Bos JL (2007) Rap1: a key regulator in cell–cell junction formation. J Cell Sci 120(Pt 1):17–22. https://doi.org/10.1242/jcs.03306
Boettner B, Van Aelst L (2009) Control of cell adhesion dynamics by Rap1 signaling. Curr Opin Cell Biol 21(5):684–693. https://doi.org/10.1016/j.ceb.2009.06.004
Mandell KJ, Babbin BA, Nusrat A, Parkos CA (2005) Junctional adhesion molecule 1 regulates epithelial cell morphology through effects on beta1 integrins and Rap1 activity. J Biol Chem 280(12):11665–11674
Friedl P, Mayor R (2017) Tuning collective cell migration by cell–cell junction regulation. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a029199
Hidalgo-Carcedo C, Hooper S, Chaudhry SI, Williamson P, Harrington K, Leitinger B, Sahai E (2011) Collective cell migration requires suppression of actomyosin at cell–cell contacts mediated by DDR1 and the cell polarity regulators Par3 and Par6. Nat Cell Biol 13(1):49–58. https://doi.org/10.1038/ncb2133
Funke L, Dakoji S, Bredt DS (2005) Membrane-associated guanylate kinases regulate adhesion and plasticity at cell junctions. Annu Rev Biochem 74:219–245
Umeda K, Ikenouchi J, Katahira-Tayama S, Furuse K, Sasaki H, Nakayama M, Matsui T, Tsukita S, Furuse M (2006) ZO-1 and ZO-2 independently determine where claudins are polymerized in tight-junction strand formation. Cell 126(4):741–754
Yonemura S, Itoh M, Nagafuchi A, Tsukita S (1995) Cell-to-cell adherens junction formation and actin filament organization: similarities and differences between non-polarized fibroblasts and polarized epithelial cells. J Cell Sci 108(Pt 1):127–142
Rajasekaran AK, Hojo M, Huima T, Rodriguez-Boulan E (1996) Catenins and zonula occludens-1 form a complex during early stages in the assembly of tight junctions. J Cell Biol 132(3):451–463
Itoh M, Nagafuchi A, Moroi S, Tsukita S (1997) Involvement of ZO-1 in cadherin-based cell adhesion through its direct binding to alpha catenin and actin filaments. J Cell Biol 138(1):181–192
Maiers JL, Peng X, Fanning AS, DeMali KA (2013) ZO-1 recruitment to alpha-catenin–a novel mechanism for coupling the assembly of tight junctions to adherens junctions. J Cell Sci 126(Pt 17):3904–3915. https://doi.org/10.1242/jcs.126565
Ando-Akatsuka Y, Yonemura S, Itoh M, Furuse M, Tsukita S (1999) Differential behavior of E-cadherin and occludin in their colocalization with ZO-1 during the establishment of epithelial cell polarity. J Cell Physiol 179(2):115–125
Ooshio T, Kobayashi R, Ikeda W, Miyata M, Fukumoto Y, Matsuzawa N, Ogita H, Takai Y (2010) Involvement of the interaction of afadin with ZO-1 in the formation of tight junctions in Madin–Darby canine kidney cells. J Biol Chem 285(7):5003–5012. https://doi.org/10.1074/jbc.M109.043760
Umeda K, Matsui T, Nakayama M, Furuse K, Sasaki H, Furuse M, Tsukita S (2004) Establishment and characterization of cultured epithelial cells lacking expression of ZO-1. J Biol Chem 279(43):44785–44794
Ikenouchi J, Umeda K, Tsukita S, Furuse M (2007) Requirement of ZO-1 for the formation of belt-like adherens junctions during epithelial cell polarization. J Cell Biol 176(6):779–786. https://doi.org/10.1083/jcb.200612080
Fanning AS, Anderson JM (2009) Zonula occludens-1 and -2 are cytosolic scaffolds that regulate the assembly of cellular junctions. Ann N Y Acad Sci 1165:113–120. https://doi.org/10.1111/j.1749-6632.2009.04440.x
Ozaki H, Ishii K, Horiuchi H, Arai H, Kawamoto T, Okawa K, Iwamatsu A, Kita T (1999) Cutting edge: combined treatment of TNF-alpha and IFN-gamma causes redistribution of junctional adhesion molecule in human endothelial cells. J Immunol 163(2):553–557
McKenzie JA, Ridley AJ (2007) Roles of Rho/ROCK and MLCK in TNF-alpha-induced changes in endothelial morphology and permeability. J Cell Physiol 213(1):221–228. https://doi.org/10.1002/jcp.21114
Stamatovic SM, Sladojevic N, Keep RF, Andjelkovic AV (2012) Relocalization of junctional adhesion molecule A during inflammatory stimulation of brain endothelial cells. Mol Cell Biol 32(17):3414–3427. https://doi.org/10.1128/MCB.06678-11
Ostermann G, Weber KS, Zernecke A, Schroder A, Weber C (2002) JAM-1 is a ligand of the beta(2) integrin LFA-1 involved in transendothelial migration of leukocytes. Nat Immunol 3(2):151–158
Fraemohs L, Koenen RR, Ostermann G, Heinemann B, Weber C (2004) The functional interaction of the beta2 integrin lymphocyte function-associated antigen-1 with junctional adhesion molecule-A is mediated by the I domain. J Immunol 173(10):6259–6264
Doherty GJ, McMahon HT (2009) Mechanisms of endocytosis. Annu Rev Biochem 78:857–902. https://doi.org/10.1146/annurev.biochem.78.081307.110540
Macara IG (2004) Parsing the polarity code. Nat Rev Mol Cell Biol 5(3):220–231
Suzuki A, Ohno S (2006) The PAR-aPKC system: lessons in polarity. J Cell Sci 119(Pt 6):979–987
Campanale JP, Sun TY, Montell DJ (2017) Development and dynamics of cell polarity at a glance. J Cell Sci 130(7):1201–1207. https://doi.org/10.1242/jcs.188599
Suzuki A, Hirata M, Kamimura K, Maniwa R, Yamanaka T, Mizuno K, Kishikawa M, Hirose H, Amano Y, Izumi N, Miwa Y, Ohno S (2004) aPKC acts upstream of PAR-1b in both the establishment and maintenance of mammalian epithelial polarity. Curr Biol 14(16):1425–1435
Hurov JB, Watkins JL, Piwnica-Worms H (2004) Atypical PKC phosphorylates PAR-1 kinases to regulate localization and activity. Curr Biol 14(8):736–741
Benton R, St Johnston D (2003) Drosophila PAR-1 and 14-3-3 inhibit Bazooka/PAR-3 to establish complementary cortical domains in polarized cells. Cell 115(6):691–704
Morais-de-Sa E, Mirouse V, St Johnston D (2010) aPKC phosphorylation of Bazooka defines the apical/lateral border in Drosophila epithelial cells. Cell 141(3):509–523
Ebnet K, Suzuki A, Horikoshi Y, Hirose T, Meyer Zu Brickwedde MK, Ohno S, Vestweber D (2001) The cell polarity protein ASIP/PAR-3 directly associates with junctional adhesion molecule (JAM). EMBO J 20(14):3738–3748
Suzuki A, Ishiyama C, Hashiba K, Shimizu M, Ebnet K, Ohno S (2002) aPKC kinase activity is required for the asymmetric differentiation of the premature junctional complex during epithelial cell polarization. J Cell Sci 115(Pt 18):3565–3573
Iden S, Misselwitz S, Peddibhotla SS, Tuncay H, Rehder D, Gerke V, Robenek H, Suzuki A, Ebnet K (2012) aPKC phosphorylates JAM-A at Ser285 to promote cell contact maturation and tight junction formation. J Cell Biol 196(5):623–639. https://doi.org/10.1083/jcb.201104143
Yamanaka T, Horikoshi Y, Suzuki A, Sugiyama Y, Kitamura K, Maniwa R, Nagai Y, Yamashita A, Hirose T, Ishikawa H, Ohno S (2001) Par-6 regulates aPKC activity in a novel way and mediates cell–cell contact-induced formation of epithelial junctional complex. Genes Cells 6:721–731
Nagai-Tamai Y, Mizuno K, Hirose T, Suzuki A, Ohno S (2002) Regulated protein–protein interaction between aPKC and PAR-3 plays an essential role in the polarization of epithelial cells. Genes Cells 7(11):1161–1171
Wang Z, Sandiford S, Wu C, Li SS (2009) Numb regulates cell–cell adhesion and polarity in response to tyrosine kinase signalling. EMBO J 28(16):2360–2373
Yamanaka T, Horikoshi Y, Sugiyama Y, Ishiyama C, Suzuki A, Hirose T, Iwamatsu A, Shinohara A, Ohno S (2003) Mammalian Lgl forms a protein complex with PAR-6 and aPKC independently of PAR-3 to regulate epithelial cell polarity. Curr Biol 13(9):734–743
Suzuki A, Yamanaka T, Hirose T, Manabe N, Mizuno K, Shimizu M, Akimoto K, Izumi Y, Ohnishi T, Ohno S (2001) Atypical protein kinase C is involved in the evolutionary conserved PAR protein complex and plays a critical role in establishing epithelia-specific junctional structures. J Cell Biol 152(6):1183–1196
Joberty G, Petersen C, Gao L, Macara IG (2000) The cell-polarity protein Par6 links Par3 and atypical protein kinase C to Cdc42. Nat Cell Biol 2(8):531–539
Lin D, Edwards AS, Fawcett JP, Mbamalu G, Scott JD, Pawson T (2000) A mammalian PAR-3-PAR-6 complex implicated in Cdc42/Rac1 and aPKC signalling and cell polarity. Nat Cell Biol 2(8):540–547
Tuncay H, Brinkmann BF, Steinbacher T, Schurmann A, Gerke V, Iden S, Ebnet K (2015) JAM-A regulates cortical dynein localization through Cdc42 to control planar spindle orientation during mitosis. Nat Commun 6:8128. https://doi.org/10.1038/ncomms9128
Nakagawa M, Fukata M, Yamaga M, Itoh N, Kaibuchi K (2001) Recruitment and activation of Rac1 by the formation of E-cadherin-mediated cell–cell adhesion sites. J Cell Sci 114(Pt 10):1829–1838
Goodwin M, Kovacs EM, Thoreson MA, Reynolds AB, Yap AS (2003) Minimal mutation of the cytoplasmic tail inhibits the ability of E-cadherin to activate Rac but not phosphatidylinositol 3-kinase: direct evidence of a role for cadherin-activated Rac signaling in adhesion and contact formation. J Biol Chem 278(23):20533–20539
Yamada S, Nelson WJ (2007) Localized zones of Rho and Rac activities drive initiation and expansion of epithelial cell–cell adhesion. J Cell Biol 178(3):517–527
Fukuhara T, Shimizu K, Kawakatsu T, Fukuyama T, Minami Y, Honda T, Hoshino T, Yamada T, Ogita H, Okada M, Takai Y (2004) Activation of Cdc42 by trans interactions of the cell adhesion molecules nectins through c-Src and Cdc42-GEF FRG. J Cell Biol 166(3):393–405
Roh MH, Makarova O, Liu CJ, Shin K, Lee S, Laurinec S, Goyal M, Wiggins R, Margolis B (2002) The Maguk protein, Pals1, functions as an adapter, linking mammalian homologues of Crumbs and Discs Lost. J Cell Biol 157(1):161–172
Bulgakova NA, Knust E (2009) The Crumbs complex: from epithelial-cell polarity to retinal degeneration. J Cell Sci 122(Pt 15):2587–2596. https://doi.org/10.1242/jcs.023648
Margolis B (2017) The Crumbs3 polarity protein. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a027961
Adachi M, Hamazaki Y, Kobayashi Y, Itoh M, Tsukita S, Furuse M, Tsukita S (2009) Similar and distinct properties of MUPP1 and Patj, two homologous PDZ domain-containing tight-junction proteins. Mol Cell Biol 29(9):2372–2389
Coyne CB, Voelker T, Pichla SL, Bergelson JM (2004) The coxsackievirus and adenovirus receptor interacts with the multi-PDZ domain protein-1 (MUPP-1) within the tight junction. J Biol Chem 279(46):48079–48084
Roh MH, Liu CJ, Laurinec S, Margolis B (2002) The carboxyl terminus of zona occludens-3 binds and recruits a mammalian homologue of discs lost to tight junctions. J Biol Chem 277(30):27501–27509
Shin K, Straight S, Margolis B (2005) PATJ regulates tight junction formation and polarity in mammalian epithelial cells. J Cell Biol 168(5):705–711
Michel D, Arsanto JP, Massey-Harroche D, Beclin C, Wijnholds J, Le Bivic A (2005) PATJ connects and stabilizes apical and lateral components of tight junctions in human intestinal cells. J Cell Sci 118(Pt 17):4049–4057
Hurd TW, Gao L, Roh MH, Macara IG, Margolis B (2003) Direct interaction of two polarity complexes implicated in epithelial tight junction assembly. Nat Cell Biol 5(2):137–142
Krapivinsky G, Medina I, Krapivinsky L, Gapon S, Clapham DE (2004) SynGAP-MUPP1-CaMKII synaptic complexes regulate p38 MAP kinase activity and NMDA receptor-dependent synaptic AMPA receptor potentiation. Neuron 43(4):563–574. https://doi.org/10.1016/j.neuron.2004.08.003
Baumgart S, Jansen F, Bintig W, Kalbe B, Herrmann C, Klumpers F, Koster SD, Scholz P, Rasche S, Dooley R, Metzler-Nolte N, Spehr M, Hatt H, Neuhaus EM (2014) The scaffold protein MUPP1 regulates odorant-mediated signaling in olfactory sensory neurons. J Cell Sci 127(Pt 11):2518–2527. https://doi.org/10.1242/jcs.144220
Butz S, Okamoto M, Sudhof TC (1998) A tripartite protein complex with the potential to couple synaptic vesicle exocytosis to cell adhesion in brain. Cell 94(6):773–782
Nix SL, Chishti AH, Anderson JM, Walther Z (2000) hCASK and hDlg associate in epithelia, and their src homology 3 and guanylate kinase domains participate in both intramolecular and intermolecular interactions. J Biol Chem 275(52):41192–41200. https://doi.org/10.1074/jbc.M002078200
Lozovatsky L, Abayasekara N, Piawah S, Walther Z (2009) CASK deletion in intestinal epithelia causes mislocalization of LIN7C and the DLG1/Scrib polarity complex without affecting cell polarity. Mol Biol Cell 20(21):4489–4499. https://doi.org/10.1091/mbc.E09-04-0280
Liu Y, Nusrat A, Schnell FJ, Reaves TA, Walsh S, Pochet M, Parkos CA (2000) Human junction adhesion molecule regulates tight junction resealing in epithelia. J Cell Sci 113(Pt 13):2363–2374
Aravindan RG, Fomin VP, Naik UP, Modelski MJ, Naik MU, Galileo DS, Duncan RL, Martin-Deleon PA (2012) CASK interacts with PMCA4b and JAM-A on the mouse sperm flagellum to regulate Ca(2+) homeostasis and motility. J Cell Physiol 227(8):3138–3150
Schuh K, Uldrijan S, Gambaryan S, Roethlein N, Neyses L (2003) Interaction of the plasma membrane Ca2+ pump 4b/CI with the Ca2+/calmodulin-dependent membrane-associated kinase CASK. J Biol Chem 278(11):9778–9783. https://doi.org/10.1074/jbc.M212507200
Shao M, Ghosh A, Cooke VG, Naik UP, Martin-DeLeon PA (2008) JAM-A is present in mammalian spermatozoa where it is essential for normal motility. Dev Biol 313(1):246–255
Erlendsson S, Madsen KL (2015) Membrane binding and modulation of the PDZ domain of PICK1. Membranes (Basel) 5(4):597–615. https://doi.org/10.3390/membranes5040597
Hanley JG (2006) Molecular mechanisms for regulation of AMPAR trafficking by PICK1. Biochem Soc Trans 34(Pt 5):931–935. https://doi.org/10.1042/BST0340931
Jaulin-Bastard F, Saito H, Le Bivic A, Ollendorff V, Marchetto S, Birnbaum D, Borg JP (2001) The ERBB2/HER2 receptor differentially interacts with ERBIN and PICK1 PSD-95/DLG/ZO-1 domain proteins. J Biol Chem 276(18):15256–15263. https://doi.org/10.1074/jbc.M010032200
Son J, Park MS, Park I, Lee HK, Lee SH, Kang B, Min BH, Ryoo J, Lee S, Bae JS, Kim SH, Park MJ, Lee HS (2014) Pick1 modulates ephrinB1-induced junctional disassembly through an association with ephrinB1. Biochem Biophys Res Commun 450(1):659–665. https://doi.org/10.1016/j.bbrc.2014.06.027
Charrin S, Jouannet S, Boucheix C, Rubinstein E (2014) Tetraspanins at a glance. J Cell Sci 127(Pt 17):3641–3648. https://doi.org/10.1242/jcs.154906
Hemler ME (2001) Specific tetraspanin functions. J Cell Biol 155(7):1103–1107. https://doi.org/10.1083/jcb.200108061
Levy S, Shoham T (2005) Protein–protein interactions in the tetraspanin web. Physiology (Bethesda, Md) 20:218–224
Sobocka MB, Sobocki T, Babinska A, Hartwig JH, Li M, Ehrlich YH, Kornecki E (2004) Signaling pathways of the F11 receptor (F11R; a.k.a. JAM-1, JAM-A) in human platelets: F11R dimerization, phosphorylation and complex formation with the integrin GPIIIa. J Recept Signal Transduct Res 24(1–2):85–105
Nourry C, Grant SG, Borg JP (2003) PDZ domain proteins: plug and play! Sci STKE 2003(179):RE7. https://doi.org/10.1126/stke.2003.179.re7
Stipp CS, Kolesnikova TV, Hemler ME (2003) Functional domains in tetraspanin proteins. Trends Biochem Sci 28(2):106–112
Naik MU, Mousa SA, Parkos CA, Naik UP (2003) Signaling through JAM-1 and {alpha}v{beta}3 is required for the angiogenic action of bFGF: dissociation of the JAM-1 and {alpha}v{beta}3 complex. Blood 102(6):2108–2114
Luissint AC, Lutz PG, Calderwood DA, Couraud PO, Bourdoulous S (2008) JAM-L-mediated leukocyte adhesion to endothelial cells is regulated in cis by alpha4beta1 integrin activation. J Cell Biol 183(6):1159–1173
Rikova K, Guo A, Zeng Q, Possemato A, Yu J, Haack H, Nardone J, Lee K, Reeves C, Li Y, Hu Y, Tan Z, Stokes M, Sullivan L, Mitchell J, Wetzel R, Macneill J, Ren JM, Yuan J, Bakalarski CE, Villen J, Kornhauser JM, Smith B, Li D, Zhou X, Gygi SP, Gu TL, Polakiewicz RD, Rush J, Comb MJ (2007) Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131(6):1190–1203
Guo A, Villen J, Kornhauser J, Lee KA, Stokes MP, Rikova K, Possemato A, Nardone J, Innocenti G, Wetzel R, Wang Y, MacNeill J, Mitchell J, Gygi SP, Rush J, Polakiewicz RD, Comb MJ (2008) Signaling networks assembled by oncogenic EGFR and c-Met. Proc Natl Acad Sci USA 105(2):692–697
Heibeck TH, Ding SJ, Opresko LK, Zhao R, Schepmoes AA, Yang F, Tolmachev AV, Monroe ME, Camp DG 2nd, Smith RD, Wiley HS, Qian WJ (2009) An extensive survey of tyrosine phosphorylation revealing new sites in human mammary epithelial cells. J Proteome Res 8(8):3852–3861
Villen J, Beausoleil SA, Gerber SA, Gygi SP (2007) Large-scale phosphorylation analysis of mouse liver. Proc Natl Acad Sci USA 104(5):1488–1493. https://doi.org/10.1073/pnas.0609836104
Dephoure N, Zhou C, Villen J, Beausoleil SA, Bakalarski CE, Elledge SJ, Gygi SP (2008) A quantitative atlas of mitotic phosphorylation. Proc Natl Acad Sci USA 105(31):10762–10767
Zahedi RP, Lewandrowski U, Wiesner J, Wortelkamp S, Moebius J, Schutz C, Walter U, Gambaryan S, Sickmann A (2008) Phosphoproteome of resting human platelets. J Proteome Res 7(2):526–534
Gauci S, Helbig AO, Slijper M, Krijgsveld J, Heck AJ, Mohammed S (2009) Lys-N and trypsin cover complementary parts of the phosphoproteome in a refined SCX-based approach. Anal Chem 81(11):4493–4501
Ozaki H, Ishii K, Arai H, Horiuchi H, Kawamoto T, Suzuki H, Kita T (2000) Junctional adhesion molecule (JAM) is phosphorylated by protein kinase C upon platelet activation. Biochem Biophys Res Commun 276(3):873–878
Okada M, Nada S, Yamanashi Y, Yamamoto T, Nakagawa H (1991) CSK: a protein-tyrosine kinase involved in regulation of src family kinases. J Biol Chem 266(36):24249–24252
Songyang Z, Shoelson SE, McGlade J, Olivier P, Pawson T, Bustelo XR, Barbacid M, Sabe H, Hanafusa H, Yi T et al (1994) Specific motifs recognized by the SH2 domains of Csk, 3BP2, fps/fes, GRB-2, HCP, SHC, Syk, and Vav. Mol Cell Biol 14(4):2777–2785
Obergfell A, Eto K, Mocsai A, Buensuceso C, Moores SL, Brugge JS, Lowell CA, Shattil SJ (2002) Coordinate interactions of Csk, Src, and Syk kinases with [alpha]IIb[beta]3 initiate integrin signaling to the cytoskeleton. J Cell Biol 157(2):265–275. https://doi.org/10.1083/jcb.200112113
Zhao Z, Vajen T, Karshovska E, Dickhout A, Schmitt MM, Megens RT, von Hundelshausen P, Koeppel TA, Hackeng TM, Weber C, Koenen RR (2017) Deletion of junctional adhesion molecule A from platelets increases early-stage neointima formation after wire injury in hyperlipidemic mice. J Cell Mol Med. https://doi.org/10.1111/jcmm.13083
Nunbhakdi-Craig V, Machleidt T, Ogris E, Bellotto D, White CL 3rd, Sontag E (2002) Protein phosphatase 2A associates with and regulates atypical PKC and the epithelial tight junction complex. J Cell Biol 158(5):967–978
Seth A, Sheth P, Elias BC, Rao R (2007) Protein phosphatases 2A and 1 interact with occludin and negatively regulate the assembly of tight junctions in the CACO-2 cell monolayer. J Biol Chem 282(15):11487–11498
Ozdamar B, Bose R, Barrios-Rodiles M, Wang HR, Zhang Y, Wrana JL (2005) Regulation of the polarity protein Par6 by TGFbeta receptors controls epithelial cell plasticity. Science 307(5715):1603–1609
Caraballo JC, Yshii C, Butti ML, Westphal W, Borcherding JA, Allamargot C, Comellas AP (2011) Hypoxia increases transepithelial electrical conductance and reduces occludin at the plasma membrane in alveolar epithelial cells via PKC-zeta and PP2A pathway. Am J Physiol Lung Cell Mol Physiol 300(4):L569–L578. https://doi.org/10.1152/ajplung.00109.2010
Walters RW, Freimuth P, Moninger TO, Ganske I, Zabner J, Welsh MJ (2002) Adenovirus fiber disrupts CAR-mediated intercellular adhesion allowing virus escape. Cell 110(6):789–799
Coureuil M, Mikaty G, Miller F, Lecuyer H, Bernard C, Bourdoulous S, Dumenil G, Mege RM, Weksler BB, Romero IA, Couraud PO, Nassif X (2009) Meningococcal type IV pili recruit the polarity complex to cross the brain endothelium. Science 325(5936):83–87. https://doi.org/10.1126/science.1173196
Scott DW, Tolbert CE, Burridge K (2016) Tension on JAM-A activates RhoA via GEF-H1 and p115 RhoGEF. Mol Biol Cell 27(9):1420–1430. https://doi.org/10.1091/mbc.E15-12-0833
Baumeister U, Funke R, Ebnet K, Vorschmitt H, Koch S, Vestweber D (2005) Association of Csk to VE-cadherin and inhibition of cell proliferation. EMBO J 24(9):1686–1695
Acknowledgements
We wish to thank Volker Gerke for continuous support. This work is supported by grants from the German Research Foundation to K.E. (EB 160/4-2, EB 160/5-1, EXC-1003 FF-2016-01) and from the Medical Faculty of the University Münster to K.E. (IZKF Eb2/020/14).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare to have no conflict of interest.
Rights and permissions
About this article
Cite this article
Steinbacher, T., Kummer, D. & Ebnet, K. Junctional adhesion molecule-A: functional diversity through molecular promiscuity. Cell. Mol. Life Sci. 75, 1393–1409 (2018). https://doi.org/10.1007/s00018-017-2729-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2729-0