Abstract
Fragment-based drug discovery (FBDD) has become increasingly popular over the last decade. We review here how we have used highly structure-driven fragment-based approaches to complement more traditional lead discovery to tackle high priority targets and those struggling for leads. Combining biomolecular nuclear magnetic resonance (NMR), X-ray crystallography, and molecular modeling with structure-assisted chemistry and innovative biology as an integrated approach for FBDD can solve very difficult problems, as illustrated in this chapter. Here, a successful FBDD campaign is described that has allowed the development of a clinical candidate for BACE-1, a challenging CNS drug target. Crucial to this achievement were the initial identification of a ligand-efficient isothiourea fragment through target-based NMR screening and the determination of its X-ray crystal structure in complex with BACE-1, which revealed an extensive H-bond network with the two active site aspartate residues. This detailed 3D structural information then enabled the design and validation of novel, chemically stable and accessible heterocyclic acylguanidines as aspartic acid protease inhibitor cores. Structure-assisted fragment hit-to-lead optimization yielded iminoheterocyclic BACE-1 inhibitors that possess desirable molecular properties as potential therapeutic agents to test the amyloid hypothesis of Alzheimer’s disease in a clinical setting.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Fragment-based drug discovery (FBDD) is an emerging field in which much lower molecular weight (MW) compounds are screened relative to those in high-throughput screening (HTS) campaigns [1–15]. In theory, fragment-based methods offer the possibility of identifying novel leads with improved pharmaceutical properties and the prospect of tackling less tractable drug targets, and the rationale behind these fragment-based strategies makes intuitive sense. However, optimization of weak-binding fragments into potent leads can be challenging, and fragment-based lead discovery can be difficult in practice. Nevertheless, FBDD has become increasingly popular over the last decade in both the pharmaceutical industry and academia [6]. Both the discovery and advancement of fragment hits are areas of intense research. Although there is still much work to be done to fully exploit the potential of this approach, the increasing number of successful applications that have appeared in the literature [1–15], including the first examples of clinical drug candidates [6, 9, 11] originating from this approach, strongly suggest its viability.
Advantages of fragment-based screening (FBS) over HTS are, first, more efficient sampling due to the smaller chemical space of fragment-sized compounds [16, 17] and, second, a higher probability of fragments possessing good complementarity with the target [18]. Since fragment-based hits are typically weak inhibitors and/or binders (half maximal inhibitory concentration (IC50) and/or the equilibrium dissociation constant (K D) is in the micromolar to millimolar range) due to their low MW, they need to be screened at higher concentrations using suitable detection techniques that can reliably detect weakly interacting compounds, e.g., nuclear magnetic resonance (NMR), surface plasmon resonance (SPR), high concentration functional screening (HCS), or X-ray crystallography. All in all, FBS leads to higher hit rates, and only relatively low numbers of compounds (thousands) need to be screened to identify interesting hits [7], even against challenging targets [12, 19]. However, fragment hits have lower affinities towards the target. As a consequence, more effort has to be spent on optimization to obtain lead compounds with an acceptable affinity and, arguably, structural biology may play a crucial role in accomplishing this goal efficiently [12].
Although fragment hits are simpler, less functionalized compounds [20] than HTS hits, with correspondingly lower potencies, they typically possess good ligand efficiency (LE) [21–28] and ligand lipophilicity efficiency (LLE) [29, 30], especially after some initial analoging or exploratory elaboration. Fragments are therefore highly suitable for optimization into clinical candidates with good drug-like properties. This means that the number of atoms involved in the desired interaction with the drug target is usually high for such fragment hits. Typical HTS hits, on the other hand, tend to be larger and, although having higher potency, contain portions in the molecule that are not directly involved in the desired interaction with the drug target. Therefore, the hit-to-lead optimization process is fundamentally different between fragment hits and typical hits from HTS. Fragment hits need to be extended into nearby binding pockets by increasing their MW to gain potency, whereas the potency of HTS hits often need to be increased without a significant increase of the MW of the initial hit [3]. Strategies have been proposed to guide and evaluate the fragment hit-to-lead optimization process [31–34]. These strategies aim at the efficient optimization of fragment hits while maintaining their generally good physicochemical properties. A recent review suggests, however, that typical medicinal chemistry approaches for lead optimization may fail at accomplishing this task [35], and a larger focus on enthalpy-driven lead optimization may be required [35, 36]. Nevertheless, less complex, polar, low MW hits should serve as better starting points for optimization [37–39] if unfavorable property shifts can be avoided during fragment hit-to-lead optimization.
In our laboratory, we have used a highly structure-driven, iterative FBDD approach composed of fragment-based NMR screening, X-ray crystallography, target-based NMR, computational chemistry, and structure-assisted chemistry. We have thus focused on targets that would be amenable to such a structure-based drug discovery (SBDD) approach, initiating protein production for both NMR and X-ray crystallography early in a project. To focus resources and to maximize impact we have applied this FBDD approach strategically to early targets, high priority targets, and those struggling for leads. Since exploratory chemistry is required for fragment hit-to-lead progression, we also paid special attention to prioritize those internal projects for which chemistry resources would be available to follow up attractive fragment hits. Further emphasis was then given to those fragment hits for which 3D structural data was available to support efficient fragment hit-to-lead progression. As a result, for 73% of the FBDD targets we have followed up fragment-based NMR screening hits through exploratory chemistry and generated 3D structural data of fragment hits when bound to the drug target. This approach has yielded valid lead series in the submicromolar potency range in about one third of those projects.
In this chapter, we first discuss fragment-based NMR screening, then suggest how to progress fragment hits into valid lead series, and finally describe a successful FBDD campaign that yielded a clinical candidate for BACE-1.
2 Fragment-Based NMR Hit Identification
Different FBS techniques have been developed and applied successfully for FBDD, as well documented in the literature (e.g., [15, 40–42]). NMR methods are among the most widely used FBS techniques [40] because they can provide useful information throughout a FBDD campaign. Versatile NMR methods are available to study the interaction of a ligand with its drug target. Such methods can be used for fragment-based NMR screening, the subsequent progression of fragment hits into leads based on structure–activity relationship (SAR) and structural information, and to support different stages in the lead generation process, ranging from hit characterization early in the process to late-stage lead optimization. Techniques can be broadly categorized into target- versus ligand-based NMR methods, depending on whether signals from the drug target or the ligand are detected to characterize the intermolecular interaction. Each of these methods has advantages and limitations and can provide information about the ligand–target interaction at various levels of detail, including the determination of ligand affinities and potencies or their binding site and binding mode when bound to the drug target. NMR experiments can be selected to fit the target size and type, the program status, and the resources that are available. Therefore, different NMR screening and follow-up strategies may be selected for different FBDD campaigns.
2.1 NMR Screening Methods
Target-detected NMR methods (Fig. 1a) have the distinct advantage that they reveal structural information about the ligand binding site and its binding mode with the drug target, can detect site-specific ligand binding over a virtually unlimited affinity range, are very robust and reliable, and can be used to derive ligand affinities for weak fragment hits that are in fast exchange on the NMR time scale (K D > ~10 μM) or for submicromolar hits when combined in a competition format (Table 1). However, they require large amounts of isotope-labeled drug target, necessitating expression of the protein in a host (typically Escherichia coli) that allows high expression yields (> ~1 mg/L) and cost-effective isotope-labeling, and also require knowledge of the 3D structure of the drug target and NMR assignments (or at least a map) of the active site residues to reveal active site binders. Therefore, target-detected NMR approaches are limited to a subset of drug targets (MW < 40–60 kDa) that give quality NMR spectra and do not aggregate at relatively high concentrations (~25–80 μM) in an aqueous NMR buffer.
NMR tools to support fragment hit identification and progression. Lead identification and optimization can broadly be categorized into target- versus ligand-detected methods depending on whether signals from the target or the ligand are detected to monitor binding. (a) Target-detected NMR method: in this case 15N-HSQC, depends on following the movement of cross-peaks as a small molecule is added. If a titration is performed, an NMR-K D can be extracted, as shown in the graph. (b) Ligand-detected STD NMR method: (i) 1D control spectrum of AMP/kinase; (ii) STD spectrum of AMP/kinase; only resonances of atoms that contact the protein are present in the STD spectrum; (iii) STD spectrum of ATP/kinase complex; (iv) STD of ATP/kinase/competitor; the STD signal due to ATP is decreased because ATP is partially displaced from the binding site by the competitor, and new STD signals for the competitor appear, compared to spectrum (iii)
Target-detected NMR screens monitor chemical shift perturbations in the heteronuclear single quantum coherence (HSQC) spectrum of an isotope-labeled protein as a small molecule (or mixture of small molecules) is added [43]. The most commonly used labeling scheme is to uniformly isotope-label the protein with 15N. The HSQC spectrum of a uniformly 15N-labeled protein contains a resonance for almost every amide N–H pair in the protein, and if these resonances have previously been assigned to the primary sequence of the protein, the binding site of the small molecule can be localized to several residues in the protein. If resonance assignments are not available, but there are reference compounds that are known to bind to the target, these reference compounds can be used to “map” residues in the binding site. If these same residues are perturbed during a fragment screen, it is likely that the screened molecule binds at the same site as the reference molecule. Even if no reference compound is available, the pattern of perturbed residues can be used to “bin” small molecules into potentially overlapping binding regions. Finally, for small molecules that are in fast exchange on the chemical shift time-scale, an NMR-K D can be determined by titrating the protein with the small molecule and monitoring the magnitude of the chemical shift perturbations as the concentration of small molecule increases. NMR-K D determination is particularly useful when functional assays for a target have not been developed or are problematic for detecting weak fragment hits.
HSQC of uniformly 15N-labeled protein can work well up to about 40–60 kDa. For protein targets larger than this, spectral overlap becomes a major problem, and methods that simplify the spectrum and improve the signal-to-noise ratio are needed. HSQC of proteins in which methyl groups are labeled with 13C has been used to simplify spectra while still providing good coverage of the target [44]. This isotope-labeling scheme also has the advantage of yielding a favorable threefold sensitivity increase. In order to further simplify the HSQC spectrum of a large protein, amino-acid-type-selective (AATS) labeling can be used with either 15N-labeling or 13C-labeling of methyl carbons. In AATS labeling, the labeling is confined to either a single amino acid type (i.e., Phe) or a small group of amino acids types (i.e., Ile, Leu, Val). Choice of which amino acid types to label is based on the presence of an amino acid type in the binding site (if the binding site is known) and/or the distribution of the amino acid in the primary sequence of the protein. Not every protein is amenable to the labeling schemes required for target-based fragment screens or may not produce quality NMR spectral data. In these cases, ligand-based fragment screens may be employed.
Ligand-detected NMR methods (Fig. 1b, Table 1) can be applied much more broadly than target-detected fragment screens because they require about 1–10% the amount of drug target, do not require isotope-labeling, and have no upper MW size limitation (in fact they work better on large proteins). Although some details about the ligand binding epitope can be obtained, ligand-detected NMR methods do not reveal the ligand binding site on the drug target. Ligand-based screens rely on monitoring the change in some NMR parameter of the ligand upon its binding to the protein. One of the most useful of these NMR methods is saturation transfer difference (STD) spectroscopy [45], and its variant, competition-STD (c-STD) spectroscopy [46, 47]. If spins anywhere in the protein are saturated, the saturation will quickly spread throughout the protein by spin diffusion, and will be transferred to a ligand if it has a long-enough residence time in the binding site. If the ligand has a fast-enough off-rate, the bound-state saturation will be observed on the free state of the ligand, with its narrow resonances. In practice, the STD experiment works well for the range 0.1 μM < K D < 1 mM, with protein concentration of 0.5–5.0 μM and ligand present in 50- to 500-fold molar excess.
The presence of signal in the STD spectrum of a ligand–protein complex must be interpreted in the broadest possible sense: there might be relatively tight binding at one binding site, weak binding at multiple sites, or some combination of the two. If there is a reference ligand with known binding site, c-STD may be used to localize the binding site of a screened molecule. Competition-STD is a two-part experiment. First, the STD spectrum of the reference molecule is obtained. Next, the competitor is added, and the STD spectrum of the ternary mixture (reference molecule, competitor molecule, protein) is obtained. If both molecules are competing for the same binding site, the STD signal of the reference molecule will decrease. The magnitude of the decrease can be used to estimate the affinity of the competitor if the affinity of the reference is known and the two molecules are strictly competitive with each other for the same binding site [48]. Since c-STD can help rule out weak, nonspecific binding, it is a highly valuable addition if well-characterized reference molecules are known for the target.
Finally, substrate-based functional NMR assays can be used to derive the percentage inhibition or IC50 values [49]. In our experience, functional NMR assays can also reveal valuable details about the mode of action of modulators, since the substrate, the product, and the ligand can be monitored in simple one-dimensional (1D) 1H NMR spectra.
From the previous discussion it becomes clear that depending on the knowledge and characteristics of a drug target, an appropriate NMR screening method needs to be selected for any given FBDD campaign. Moreover, suitable NMR methods can be selected to derive ligand affinity or potency to assist SAR development (Table 2).
2.2 Library Considerations
Fragment-based approaches probe chemical space more efficiently than HTS approaches, are less dependent on legacy compound collections, and can provide hits for challenging targets. One of the great advantages of NMR-based methods is the ability to reliably identify weak binders with K Ds in the low millimolar range, while still obtaining useful structural information about their potential binding site(s). With this affinity cut-off, screening a library of 1,000–2,000 fragments will result in multiple hits for most targets. The selection of these 1,000–2,000 compounds for an NMR-based screening library can be crucial to the success of the endeavor, and details of this important topic have been described in a number of publications (e.g., [50–54]). Candidate molecules are filtered to ensure favorable physicochemical properties and a lack of reactive functional groups. Issues of “chemical diversity” versus “drug likeness” must be balanced. Library members might be synthetic cores for which chemically elaborated back-up libraries are readily available for fast SAR. The chosen fragment screening method may to some degree also influence the design of such a fragment library [53]. If 3D structural information is available for the drug target, virtual screening may be employed to select focused FBS libraries to increase hit rates [15]. Several companies nowadays sell FBS libraries as part of their business since FBDD has become increasingly popular over the last decade.
Once candidate library members have been chosen, they must be validated by experiment. For each library member, the chemical structure is verified, the purity of the sample determined, and aqueous solubility measured. In addition, the fragments should be tested for their potential to aggregate at the high screening concentrations used for fragment-based NMR screening [51]. DMSO-d6 stock solutions of the library must be plated and stored in a way that minimizes freeze/thaw cycles and exposure to atmospheric water. In order to facilitate the identification of hits in ligand-detected NMR methods, library members are plated so that each screening cluster is “chemical shift encoded”; that is, within each cluster there are no degenerate chemical shifts between cluster members. Target-detected NMR screening methods do not have such requirements, but fragments in an “active” cluster must be deconvoluted to identify hit(s).
3 Fragment Hit-to-Lead Progression
3.1 Fragment Hit Validation and Initial SAR Development
Cross-validation of NMR results with information yielded by other biophysical, biochemical, and cell-based assays can be crucial to the progression of a fragment hit. Access to other assay methods is especially important when STD is used as the NMR screening method because STD reveals no information on the ligand binding site and is more susceptible to unrecognized nonspecific binding. Results from biophysical methods such as SPR, thermal denaturation, and isothermal calorimetry (ITC), in addition to X-ray crystallography and structure-based NMR studies, can be used to validate NMR hits. If available, biochemical and cell-biological functional assays are valuable tools for probing the interaction of a fragment hit with its target.
Even before project chemists become actively involved, SAR can be quickly progressed by testing obvious analogs of the initial fragment hit from readily available commercial or internal sources, which may include “expansion” libraries that have been prepared on the basis of members of the screening library. The value of a chemotype or structural motif becomes clearer if a series of molecules has been studied, and some initial SAR is seen. On the basis of results from the first round of analoging, project chemists will usually have ideas for further SAR development. It is important that the “iteration time” between submission of new compounds for testing and the reporting of test results back to the project team be as short as possible to maintain project momentum.
3.2 Evaluation of Binding Site and Binding Mode
Target-based NMR methods can often provide this crucial information, especially if site-specific assignments are available from the literature or can be obtained internally, and the 3D structure of the drug target is known. The detailed binding mode of a fragment hit by NMR can, however, only be obtained for smaller targets with MWs up to about 20–30 kDa, and requires significant resources. Thus, X-ray crystallography becomes the method of choice for determining the detailed binding mode of a fragment hit.
The preferred binding mode within a chemical series can change even within the same binding site as substituents are changed, thus confusing SAR development. In these cases, knowledge of the detailed binding modes of key members within a lead series is crucial for efficient fragment hit progression.
3.3 Ligand Efficiency Indices to Guide Fragment Hit Selection and Progression
Traditionally, affinity/potency has been the primary factor for hit selection and optimization. However, there is a strong correlation between increased MW and improved affinity/potency. Moreover, lead optimization typically yields bigger and more lipophilic compounds [30]. However, almost all absorption, distribution, metabolism, excretion, and toxicity (ADMET) parameters deteriorate with either increasing MW and/or partition coefficient (logP) [55] and good physicochemical properties help to reduce the attrition rate in late stage clinical trials [56]. Therefore, selecting appropriate hits with a good balance of MW and lipophilicity, and monitoring this balance in addition to affinity/potency during hit optimization, have been recognized as important factors for successful drug discovery.
Many fragment hits found through NMR screening will have weak affinity and will require substantial modification to become viable leads. As fragment hits are different from traditional HTS hits, a process tailored for fragment hit progression is required. Several LE indices have been proposed for guiding this process [34]. Thus, weak binders identified by fragment-based NMR screening might be good starting points for lead generation if they exhibit good LE and good LLE. LE and related indices estimate the efficiency of a binding interaction with respect to the number of non-hydrogen atoms and is a way of normalizing the binding energy by the size of the molecule [21–23]. Because LE cannot be evaluated independently of the molecular size [24], scaled LE scores have been proposed to enable a size-independent comparison of ligands [25–28]. LLE is a measure of the minimally acceptable lipophilicity per unit of in vitro potency: LLE(Leeson) = pIC50−cLogP (computed partition coefficient) [29] or a normalized LLE(Astex) = 0.111−(−1.36 × LLE(Leeson)/number of heavy atoms) can be used for practical reasons for fragment hits [57]. Chemists will have the freedom to elaborate low MW, high LE hits before reaching unacceptable limits of MW and complexity, which often lead to compounds that exhibit unacceptable solubility, absorption, and permeability properties. Similarly, fragments with good LLE provide the opportunity to increase lipophilicity during lead optimization without reaching an unfavorable physical profile for the drug candidate. However, LLE does not include LE and vice versa. Since there is a significant predisposition towards improving potency simply by adding lipophilicity, LELP = logP/LE was proposed as a useful function to depict the price of LE paid in logP [30].
Affinity, or binding energy, comprises two components: enthalpy and entropy. It has recently been proposed that there are advantages to starting with enthalpically driven leads [35, 58, 59], in which binding arises from specific molecular interactions such as hydrogen bonds, salt bridges, and van der Waals interactions. In contrast, entropically driven binding generally arises from nonspecific hydrophobic interactions. ITC is the tool of choice for determining the relative contributions of entropy and enthalpy to binding affinity [60]. The information from ITC is best interpreted in conjunction with a detailed structural model of the binding interaction (usually from X-ray crystallography) and provides a strong starting point for optimization of a lead series. The relative balance of entropy and enthalpy will, of course, change as optimization progresses, but thermodynamic analysis and detailed structure models can go a long way towards explaining unexpected SAR and in providing guidance on where to focus synthetic efforts. Thus, an enthalpic efficiency (EE) and a specific EE were proposed as additional tools for selecting compounds in lead discovery and for aiding lead optimization [36].
The tractability of a fragment hit for chemical elaboration is judged by project chemists, who have the expert knowledge needed to assess the possibilities for elaboration of a fragment hit with substituents, or recasting of a chemotype into an isostere. Chemists also assess the fragment hit for potential chemical novelty, especially important if the target has been extensively studied by other groups. Close interaction with project chemists is crucial to the success of a project. In the early stage of a project, a core structure that can easily be derivatized is advantageous for fragment hit progression.
Structural information about the binding mode of a fragment hit can be crucial for efficient hit-to-lead optimization, as discussed above. Therefore, we prefer to apply this FBDD approach to high-priority targets and drug targets for which X-ray or NMR structures can be obtained. Whenever possible, with this approach we like to provide the chemist with low MW, high LE, and high LLE compounds for which we know their binding mode to the drug target, thereby providing chemists with more room for optimizing pharmokinetic (PK)/ADMET properties during the lead optimization process. Thus, a structure-focused FBDD approach can produce leads for very challenging targets where other methods may fail (see BACE example below).
Follow-up strategies for fragment hits strongly depend on the nature and characteristics of the drug target and the fragment hits. For more challenging targets, structural data is crucial for efficient fragment hit-to-lead optimization, whereas for other targets with deep, well-defined active sites this may not necessarily be the case. In the latter case, high-concentration biochemical screening of libraries that contain “lead-like” compounds [39] may be more efficient than a structure-based NMR fragment screen, especially if a robust functional assay can be developed. High-concentration biochemical screens have the distinct advantage that they already provide a functional readout for the fragment hit, and the hit-to-lead process follows a traditional progression path. However, HCS of fragment libraries could be prone to larger numbers of “false positives,” and orthogonal biophysical methods might become important for pruning fragment hit lists.
Although tethering/linking fragments that bind to proximal binding sites can in principle yield high-affinity linked molecules, this approach is often not very practical due to difficulties in finding proximal binders, knowing their detailed binding mode, and due to limitations in linker chemistry and optimization [61]. Thus, expanding or growing initial fragment hits into more potent leads has become much more common than tethering for FBDD. FBDD approaches may also become very useful in better understanding the contributions of individual components of an existing lead [62], or for improving an existing lead by “fragment hopping” [63].
4 Structure-Based FBDD Approach Applied to BACE-1
4.1 BACE-1 as a Drug Target for Alzheimer’s Disease
Alzheimer’s disease (AD) is a progressive, ultimately fatal neurodegenerative disease that gives patients an average life expectancy of 7–10 years after diagnosis [64]. It is the leading cause of dementia in the elderly population, causing gradual loss of mental and physical function. In addition to the devastating physical and emotional impact of AD on patients and their families, all patients at an advanced stage of the disease will inevitably need long-term care, which places a huge social and economic burden on their families and society [65]. In the USA alone, there are currently four million AD patients, with an additional eight million subjects diagnosed with mild cognitive impairment (MCI), of whom many will progress to AD [66]. This number is expected to quadruple in the next three decades unless therapies that impact the underlying pathophysiology of AD can be identified.
Current therapies for AD, comprising acetylcholine esterase inhibitors and an NMDA receptor antagonist, offer only symptomatic relief by compensating for the neuronal and synaptic losses in AD patients through prolonging activation of the remaining neuronal network [67]. These therapies offer patients transient improvements in cognition and daily living functions, but do not halt disease progression. Thus, there are enormous unmet medical needs for the AD population.
The pathological hallmarks observed in the brains of AD patients are the extracellular amyloid plaques, mainly composed of an amyloid-β peptide with 42 amino acids in length (Aβ42), and the intracellular neurofibrillary tangles of hyperphosphorylated tau protein. According to the amyloid hypothesis [68–73], the prevailing theory in the field, the underlying cause of AD is the aggregation and deposition of Aβ42 in the brain due to its overproduction and/or diminished clearance. This hypothesis is supported by strong genetic, histopathological, and clinical evidence. All early-onset familial AD is identified by genetic mutations in amyloid precursor protein (APP) or presenilins (PS1 and PS2) that result in increased Aβ peptide production. Down’s syndrome patients, who have an extra copy of chromosome 21 containing the APP gene, or individuals who have a duplication of only a portion of chromosome 21 that contains the APP gene, produce more Aβ peptides and develop early-onset AD [74, 75]. One Down’s individual [76], whose extra copy of the portion of chromosome 21 lacked the APP gene, did not develop AD. Among all β-amyloid species, Aβ42 is most prone to aggregation and most cytotoxic in vitro [77–80]. Lastly, active immunization against Aβ peptides reduced amyloid load in animal models [81, 82] and was associated with cognitive improvement for AD patients who developed robust anti-Aβ titers in human clinical trials [83–87].
Aβ peptides, ranging from 37 to 42 amino acids in length, differ from each other at the C-terminus. They are produced as minor products (5–10%) of the metabolism of the membrane-bound APP via two consecutive cleavages: first by β-site APP cleaving enzyme (BACE-1, also known as β-secretase or memapsin-2) [88–91], followed by γ-secretase, in competition with the major pathway (90–95%) of non-amyloidogenic processing of APP by α-secretase. There are two BACE isoforms, with BACE-1 mainly expressed in the central nervous system (CNS) and responsible for Aβ peptide production. BACE-2 cleaves APP at a different site to the BACE-1 cleavage and is mainly expressed in the periphery [92, 93]. BACE-1 knockout (KO) mice are normal, do not produce Aβ peptides, and have few overt phenotypes [94–97]. Crossing BACE-1 KO with transgenic mice that overproduce human APP eliminates Aβ production and amyloid plaque formation and rescues memory dysfunction [98]. These data suggest that Aβ peptide inhibition through small molecule BACE-1 inhibitors is highly promising as a disease-modifying therapy that may halt or even reverse the progression of AD. Therefore, BACE-1 has been a high priority therapeutic target for the treatment of AD throughout the pharmaceutical industry over the last decade.
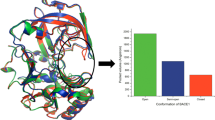
BACE-1 is a membrane-anchored aspartic acid protease that is localized to the acidic compartments of endosomes and lysosomes in the CNS and has an optimal enzymatic activity at around pH 5. As a consequence, a BACE-1 inhibitor needs to be able to cross the blood–brain barrier and to have a significant non-protein bound fraction in order to reach the active site of the enzyme. This makes traditional aspartic protease inhibitors, which typically are large and peptidic, unsuitable as BACE-1 inhibitors. Moreover, the BACE-1 active site is extended, shallow and hydrophilic (Fig. 2) [99]. Therefore, the development of potent, selective, orally active, and brain penetrant low MW compounds has been a big challenge for the pharmaceutical industry [101, 102].
BACE-1 characteristics. The overall fold of BACE-1 is typical for an aspartic acid protease, consisting of an N- and C-terminal lobe with the substrate binding site located in a crevice between the two lobes [99, 100]. A flexible hairpin, called the flap (Yellow see-through surface), partially covers the active site of BACE-1 and can adopt many different conformations as a result of inhibitor binding. In the center of the active site are the two aspartic acid residues (orange and inset) that are involved in the enzymatic reaction
Many of the early drug discovery efforts focused on the development of transition state peptidomimetics that were known from the aspartic acid protease field [99, 103]. Although this approach yielded highly potent and selective BACE-1 inhibitors, the resulting compounds lacked in vivo efficacy probably due to their large MW and suboptimal PK properties. We review here how we have used a highly structure-driven approach, consisting of the integrated application of target-detected fragment-based NMR screening, X-ray crystallography, structure-based design and structure-assisted chemistry together with innovative biology, to develop a first-in-class clinical candidate as a potential proof-of-concept for the inhibition of BACE-1 in AD [104, 105]. Recently, several other fragment-derived BACE-1 inhibitors have also been described [106–113].
4.2 Fragment Hit Identification
We developed an efficient protocol for the large scale production of a fully processed soluble version of 15N-labeled BACE-1 for fragment-based NMR screening and X-ray crystallography in which the pre- and pro-sequences are autocatalytically removed within about 3 days at room temperature or 18 days at 4 °C at protein concentrations of ~5–10 mg/mL [114]. This refolding protocol from inclusion bodies yielded around 40 mg BACE-1/L cell paste. We used NMR to monitor structural details of the autocatalytic conversion, which revealed a major structural rearrangement in the N-terminal lobe from a partially disordered to a well-folded conformation suggesting that the pro-sequence may assist the proper folding of the protein. Once the protein was completely folded, we could recycle it multiple times for fragment-based NMR screening.
We screened over 10,000 fragments of a custom-built fragment library [7] at high concentrations (100 μM–1 mM each) in cocktails of 12 to identify active-site-directed hits by 15N-HSQC NMR [104]. About half of these fragments were strictly rule-of-three compliant [20], whereas a large majority followed “reduced complexity” rules (MW < 350, cLogP ≤ 2.2, H-bond donor ≤ 3, H-bond acceptor ≤ 8, rotatable bonds ≤ 6, heavy atom count ≤ 22) [39]. At first, we did not have protein NMR resonance assignments for BACE-1. In order to not delay fragment-based NMR screening, we initially identified peaks of active site residues of BACE-1 by binding peptide inhibitors known from the literature and then screening for fragments that showed chemical shift perturbations of some of those peaks. Eventually, we obtained sequence-specific NMR resonance assignments for BACE-1, which then allowed us to study ligand binding in more structural detail [115]. Overall we identified nine distinct chemical classes of active site binders to BACE-1 in the 30 μM to 3 mM K D range, as determined by NMR titration experiments (Fig. 3a).
BACE-1 fragment hit identification and fragment hit-to-lead progression. (a) Fragment-based NMR screening hits for BACE-1. Nine classes of BACE-1 active-site-directed NMR hits were identified by screening 10,000 compounds from a customized NMR fragment library by 15N-HSQC NMR. (b) Isothiourea fragment hit identification and optimization by NMR and X-ray crystallography. (c) Search for heterocyclic isothiourea isosteres. (d) 2-Aminopyridines and related heterocyclic isothiourea isosteres were identified through directed fragment-based NMR screening. (e) Structure-based design of prototype iminohydantoins yielded attractive starting points for the development of novel low MW BACE-1 inhibitors. See text for details
Among our initial fragment hits were several amidine-containing chemotypes, including the isothiourea 1 (Fig. 3b). We then tested over 200 analogs by NMR to derive initial SAR and discovered isothiourea 2, which showed an NMR-K D of 15 μM (LE = 0.39) and weak activity in an enzymatic assay. The NMR chemical shift perturbation data suggested that compound 2 binds to the two active site aspartates and extends into the S3 pocket while leaving the flap untouched in its “open” apo-conformation. Subsequently, the co-crystal structure of compound 2 with BACE-1 revealed details about how the isothiourea moiety forms an extensive H-bond donor acceptor array with the two active site aspartates and places the chloro-phenyl ring into the S1 pocket and extends deep into the shallow S3 pocket through the butyl-ether group. From that point on, this fragment was used in an X-ray soaking system to solve the X-ray structures of over 1,000 BACE-1 inhibitors that followed in this project.
When we discovered this NMR fragment hit several years ago, this type of hydrogen-bond network to the two active site aspartates was unprecedented in the aspartic acid protease field. Unfortunately, potential hydrolytic instability of the isothiourea moiety of compound 2 renders it unsuitable for drug development. Therefore, we started an extensive search for heterocyclic isothiourea isosteres that would be pharmaceutically attractive with an appropriate basicity (pK a range 6–10) to maintain the crucial H-bonding network with the two active site aspartates while limiting the number of H-bond donating groups and have molecular properties compatible with brain penetration. We pursued two approaches (Fig. 3c). In the first approach, we carried out focused NMR screens to identify heterocyclic structures including 2-aminoimidazoles and 2-aminopyridines to bind into the active site of BACE-1 [104]. In the second approach, we designed cyclic acylguanidines, including iminohydantoins and iminopyrimidinones [105].
4.3 Focused Search for Pharmaceutically Attractive Isothiourea Isosteres
While our general NMR fragment screening was still in progress, we initiated focused directed NMR screens of heterocyclic isothiourea isosteres that were available from our corporate library. During this process, we identified several heterocyclic cores as active site BACE-1 binders, which included 2-aminopyridines, 2-aminoimidazoles, 2-aminobenzimidazoles, 2-aminotriazines, and benzoamidines, whereas other related cores were not identified as hits (Fig. 3d). In the 2-aminopyridine series, we discovered compound 3, which bound to the two active site aspartates with an NMR-K D of 32 μM (LE = 0.39) as judged by the NMR chemical shift perturbation data. Compound 3 thus had LE [21] and fit quality (FQ) [25, 26] values similar to those of compound 2. Its LLE [29] was, however, significantly reduced due to its increased hydrophobicity. Interestingly, the X-ray crystal structure of this fragment in complex with BACE-1 revealed the same H-bonding network as previously seen for compound 2. Only a few months into the FBDD campaign, compound 3 provided the first attractive starting point for chemical elaboration. Exploratory chemistry on the 2-aminopyridine series was initiated. Small chemical libraries based on the 2-aminopyridine-phenethyl core were built to explore this chemotype. Several analogs with activities in the micromolar range were identified, and crystal structures for some of these suggested the synthesis of 3,6-disubstituted 2-aminopyridine, which yielded the first submicromolar inhibitors in this series. However, the planar nature of the 2-aminopyridine core and difficulties in synthesizing 3,6-disubstituted analogs prevented the easy development of more potent BACE-1 inhibitors with lead-like properties in this series.
In an alternate approach, novel cyclic acylguanidine active-site-binding cores such as iminohydantoin and iminopyrimidinone were conceptualized (Fig. 3c) in which the crucial aspartate-binding amidino motif, common to fragment-based NMR screening hits 2 and 3 and of similar weak basicity, is conserved. It was suggested that disubstitution at C5 (iminohydantoin) or C5 and C6 (iminopyrimidinone) would simultaneously provide direct access to both prime and non-prime binding sites adjacent to the catalytic aspartate residues, with substitution on the second ring nitrogen providing a further handle for accessing binding pockets adjacent to the active site. To test this hypothesis, the prototype iminohydantoin (compound 4) and its N1-analog were designed and synthesized. The 3-chlorobenzyl substituent was predicted to bind in the S1 pocket, in analogy to 2 and 3. We were delighted to find that iminohydantoin 4 bound to BACE-1 with an NMR-K D of 200 μM, whereas no binding was observed for its N1-analog. Despite its weak binding activity, compound 4 showed promising LE and LLE values for fragment hit progression. An X-ray structure of 4 in complex with BACE-1 confirmed that 4 bound as predicted (Fig. 3e). We then tested several related N3 and N1 analogs. We consistently found by NMR that the N3-, but not the N1-prototype iminohydantoins bound into the active site of BACE-1. About a year into the FBDD approach, we had now discovered a very attractive novel core structure that was chemically stable, had a pK a compatible with CNS penetration, and provided ample opportunities to extend the molecule into nearby substrate binding pockets using well-known hydantoin chemistry.
4.4 Fragment Hit-to-Lead Progression
During fragment hit-to-lead progression we quickly identified a second binding mode of the iminohydantoin core in the active site of BACE-1 using X-ray crystallography. This is represented by compound 5, in which an extensive ligand–BACE-1 H-bonding network is maintained, but the iminohydantoin core is flipped in the active site (Fig. 4a). This observation turned out to be highly significant because this mode proved to be the preferred binding mode as lead optimization evolved. NMR chemical shift perturbation data could be used to quickly categorize ligands with respect to these two binding modes (Fig. 4b). Simple changes in the substituents could not only cause the iminohydantoin core to flip, but also to tilt or slightly shift in the binding pocket while maintaining an extensive H-bond network with the two active site aspartates. Therefore, X-ray structural data was crucial for medicinal chemists to understand otherwise confusing SAR (Fig. 4b).
Iminohydantoin fragment hit progression. (a) A second binding mode of the iminohydantoin core in the active site of BACE-1 was revealed by X-ray crystallography. (b) Simple changes in the substituents could cause the iminohydantoin core to flip, tilt, or shift in the active site while maintaining an extensive H-bond network with the two aspartates, thus structural data simplified SAR development. (c) Structure-based design of the first series of submicromolar iminohydantoin BACE-1 inhibitors. (d) Truncated N-methyl iminohydantoins provided a more direct way to build toward S3 through a contiguous hydrophobic patch from S1 through S3 into S3sp. (e) Truncated N-methyl iminohydantoins showed improved LE, with compound 12 showing excellent lead-like properties. The X-ray crystal structures of BACE-1 in complex with compound 11 and 12 showed relatively open flap conformations, with flap residue Tyr71 (shown in cyan in the “closed” flap, peptidomimetic inhibitor conformation [99]) displaced by one of the phenyls at the 5-position of compound 12 [105]. (f) Iterative structure-assisted chemistry was able to improve ligand efficiency indices during fragment hit-to-lead optimization and lead optimization in the iminohydantoin series. See text for details
It was important to demonstrate quickly that we can produce potent iminohydantoin BACE-1 inhibitors that had submicromolar IC50s in the enzymatic assay. The binding mode of iminohydantoin 7 (Fig. 4b) suggested that cyclohexylmethyl and cyclohexylethyl extensions into the respective hydrophobic S1 and S2′ pockets should achieve this goal (Fig. 4c). The resulting iminohydantoin 8 was in fact the first submicromolar inhibitor in this series. Its crystal structure confirmed the underlying structure-based design and suggested that a further increase in potency should be possible by introducing a cyclic urea with the propyl extension in the proper (S)-configuration. Again, the ensuing iminohydantoin 9 bound to BACE-1 as expected and showed an increased potency in the enzymatic assay. Isolation of the single stereoisomer with 4(S)/4(R) configuration yielded compound 10 with a cellular IC50 in the submicromolar range (Fig. 4d). However, the resultant compounds became non-leadlike with significantly reduced LE (despite an improved FQ), increased cLogP (yielding a very poor LLE), poor rat PK, and no selectivity over the aspartic acid protease Cathepsin D. Molecular modeling suggested that we should be able to truncate compound 10 to a N-methyl and extend the iminohydantoin core deep into the S3 subpocket (S3sp) more directly through a contiguous hydrophobic patch without adding as much MW to the iminohydantoin core. It was good to see that the truncated N-methyl iminohydantoin analog (compound 11) showed much higher LE than compound 10 (while maintaining a good FQ) and only a three- to fourfold loss in cellular 70–75% decerase in potency (Fig. 4d). Knowing that we needed to develop a CNS drug, we then reduced the number of rotatable bonds of compound 11 and designed the rigid, compact 5,5′-diphenyl iminohydantoin core structure (compound 12) which, despite being only weakly active, now showed excellent lead-like properties with good LE, much better LLE, and an overall favorable profile with respect to cellular potency, selectivity, rat PK, and brain penetration (Fig. 4e). Thus, compound 12 was the superior choice for lead optimization.
The X-ray structure of BACE-1 in complex with compound 11 (Fig. 4e) revealed a relatively open flap conformation, with the aliphatic chain of the inhibitor projecting towards S2′ in close proximity to a pocket that we termed F′. In X-ray structures of peptidomimetic inhibitors bound to BACE-1 [99], the F′ pocket is occupied by flap Tyr71 but is vacated in X-ray crystal structures of iminohydantoin 11 and related compounds. Incorporation of 5-phenyl substitution to exploit occupancy of F′ was a key design concept that resulted in identification of 5,5′-diphenyl-iminohydantoin 12, in which one of the C5 phenyl substituents now occupied the unique F′ pocket, again yielding a more “open” flap conformation (Fig. 4e).
During iminohydantoin fragment hit-to-lead optimization, which involved an iterative process of molecular modeling, structure-assisted synthesis, and functional and structural evaluation, the LE between the initial fragment hit and the optimized fragment lead was increased significantly from 0.24 to 0.37 kcal/mol/heavy atom (Fig. 4f), yielding a corresponding increase in FQ. Due to a significant reduction in cLogP, compound 12 also showed a much improved LLE as compared to the initial iminohydantoin fragment hit. Thus, the primary goal during lead optimization was to increase potency and selectivity of the iminohydantoin lead series while maintaining good LE and molecular properties that would be compatible with brain penetration.
4.5 Iminohydantoins: S1–S3 Occupancy
The truncated N-methyl iminohydantoins (compounds 11 and 12) showed much higher LE than compound 10, and provided opportunities to build into the S3 pocket more directly without increasing the MW of the iminohydantoins as much as in the earlier series, which was extended at the N1-position towards the S2 pocket. Compound 12 possesses a diphenyl substitution at C5, with one of the C5 phenyl substituents occupying a unique binding pocket designated F′ that is normally filled by the enzyme flap Tyr residue in the closed-flap enzyme conformation (Fig. 5). Thus, compound 12 offered several opportunities to extend the iminohydantoin core from the C5 position into the surrounding S1–S3 and S2′ substrate binding pockets. Based on the X-ray structure of compound 12 in complex with BACE-1, molecular modeling suggested that we could extend the phenyl in the S1 pocket at the meta-position toward the S3 pocket. We then tested this hypothesis by synthesizing analogs that probed different extensions at this meta-position (Fig. 5). SAR revealed that a phenyl extension is tolerated and that small hydrophobic substituents at the 3-position of this distal phenyl improved the enzymatic K I by about an order of magnitude, yielding several submicromolar BACE-1 inhibitors. The X-ray structure of the diphenyl-iminohydantoin with a 3-pyridine extension (compound 13) in complex with BACE-1 exhibited an H-bond to a bound water molecule in the S3 subpocket and could explain SAR that showed a preference of the 3-pyridine over the 4-pyridine analog. It could explain additional SAR that revealed a preference of substitutions at the 3- over the 4-position at the distal phenyl in the S3 pocket. Substitutions at the 3-position presumably could reach deep into the S3 subpocket by replacing this nonstructural bound water molecule in the S3 subpocket.
Despite this structural knowledge it still turned out to be challenging to significantly improve the potency of the iminohydantoin series with respect to cellular potency and PK properties. However, by use of structure-assisted SAR development the team was ultimately able to develop BACE-1 inhibitors with high affinity, selectivity, and excellent PK properties to achieve brain penetration and CNS efficacy in vivo [116].
5 Conclusion and Perspectives
We have used a highly structure-driven approach composed of fragment-based NMR screening, X-ray crystallography, and structure-assisted chemistry to develop a first-in-class clinical candidate as a potential proof-of-concept for the inhibition of BACE-1 in AD. Crucial to this achievement was the initial identification of a ligand-efficient isothiourea fragment and its X-ray crystal structure, which revealed an extensive H-bond network with the two active site aspartates. This interaction was unprecedented in the aspartic acid protease field when we discovered it several years ago. This detailed 3D structural information then enabled the design and validation of novel, chemically stable and accessible heterocyclic acylguanidines as aspartic acid protease inhibitor cores. Lead optimization guided by structure-based design afforded unique, low MW, high affinity, selective iminopyrimidinones as BACE-1 inhibitors in which the hydrophobic interactions in the S1, S3, and S3sp pockets were optimized to achieve excellent cellular potency. The resulting leads were conformationally restricted with few rotatable bonds, which contribute to their high LE indices. These iminoheterocyclic BACE-1 inhibitors possess desirable molecular properties as potential therapeutic agents to test the amyloid hypothesis in a clinical setting. Optimized iminopyrimidinones have shown high oral bioavailability, good CNS penetration, and robust reductions of cerebrospinal fluid and brain Aβ in animal models.
Combining biomolecular NMR, X-ray crystallography, and molecular modeling with structure-assisted chemistry and innovative biology as an integrated approach for FBDD can solve very difficult problems, as illustrated in this chapter. BACE-1 has been a challenging CNS target for small molecule drug discovery, where more conventional lead generation approaches had failed despite extensive efforts for over a decade. However, none of the components mentioned above would have been successful if applied in isolation. Therefore, the future for FBDD looks bright as long as an appropriate infrastructure can be provided for this technology to tackle appropriate problems in drug discovery.
Abbreviations
- AD:
-
Alzheimer’s disease
- ADMET:
-
Absorption distribution, metabolism, excretion, and toxicity
- APP:
-
Amyloid precursor protein
- Aβ:
-
Amyloid beta peptides ranging from 37 to 42 amino acids in length
- BACE-1:
-
β-site APP cleaving enzyme
- cLogP:
-
Computed partition coefficient of a compound
- CNS:
-
Central nervous system
- c-STD:
-
Competition saturation transfer difference
- FBDD:
-
Fragment-based drug discovery
- FBS:
-
Fragment-based screening
- FQ:
-
Fit quality
- HCS:
-
High concentration functional screening
- HSQC:
-
Heteronuclear single quantum coherence
- HTS:
-
High-throughput screening
- IC50 :
-
Half maximal inhibitory concentration
- ITC:
-
Isothermal calorimetry
- K D :
-
Equilibrium dissociation constant
- K I :
-
Equilibrium inhibition constant
- LE:
-
Ligand efficiency
- LLE:
-
Ligand lipophilicity efficiency
- MW:
-
Molecular weight
- NMR:
-
Nuclear magnetic resonance
- PK:
-
Pharmacokinetics
- SAR:
-
Structure–activity relationship
- SPR:
-
Surface plasmon resonance
- STD:
-
Saturation transfer difference
References
Jahnke W, Erlanson DA (eds) (2006) Fragment-based approaches in drug discovery. In: Mannhold R, Kubinyi H, Folkers G (series eds) Methods and principles in medicinal chemistry, Vol 34. Wiley-VCH, Weinheim, Germany
Zartler ER, Shapiro MJ (eds) (2008) Fragment-based drug discovery: a practical approach. Wiley, Chichester, UK
Rees DC, Congreve M, Murray CW, Carr R (2004) Fragment-based lead discovery. Nat Rev Drug Discov 3:660–672
Erlanson DA, McDowell RS, O’Brien T (2004) Fragment-based drug discovery. J Med Chem 47:3463–3482
Erlanson DA (2006) Fragment-based lead discovery: a chemical update. Curr Opin Biotechnol 17:643–652
Hajduk PJ, Greer J (2007) A decade of fragment-based drug design: strategic advances and lessons learned. Nat Rev Drug Discov 6:211–219
Wyss DF, Eaton HL (2007) Fragment-based approaches to lead discovery. Front Drug Des Discov 3:171–202
Alex AA, Flocco MM (2007) Fragment-based drug discovery: what has it achieved so far? Curr Top Med Chem 7:1544–1567
Congreve M, Chessari G, Tisi D, Woodhead AJ (2008) Recent developments in fragment-based drug discovery. J Med Chem 51:3661–3680
de Kloe GE, Bailey D, Leurs R, de Esch IJP (2009) Transforming fragments into candidates: small becomes big in medicinal chemistry. Drug Discov Today 14:630–646
Chessari G, Woodhead AJ (2009) From fragment to clinical candidate-a historical perspective. Drug Discov Today 14:668–675
Murray CW, Rees DC (2009) The rise of fragment-based drug discovery. Nat Chem 1:187–192
Schulz MN, Hubbard RE (2009) Recent progress in fragment-based lead discovery. Curr Opin Pharm 9:615–621
Murray CW, Blundell TL (2010) Structural biology in fragment-based drug design. Curr Opin Struct Biol 20:497–507
Gozalbes R, Carbajo RJ, Pineda-Lucena A (2010) Contributions of computational chemistry and biophysical techniques to fragment-based drug discovery. Curr Med Chem 17:1769–1794
Fink T, Bruggesser H, Reymond JL (2005) Virtual exploration of the small-molecule chemical universe below 160 Daltons. Angew Chem Int Ed Engl 44:1504–1508
Fink T, Reymond JL (2007) Virtual exploration of the chemical universe up to 11 atoms of C, N, O, F: assembly of 26.4 million structures (110.9 million stereoisomers) and analysis for new ring systems, stereochemistry, physicochemical properties, compound classes, and drug discovery. J Chem Inf Model 47:342–353
Hann MM, Leach RL, Harper G (2001) Molecular complexity and its impact on the probability of finding leads for drug discovery. J Chem Inf Comput Sci 41:856–864
Coyne AG, Scott DE, Abell C (2010) Drugging challenging targets using fragment-based approaches. Curr Opin Chem Biol 14:299–307
Congreve M, Carr R, Murray CW, Jhoti H (2003) A ‘rule of three’ for fragment-based lead discovery? Drug Discov Today 8:876–877
Hopkins AL, Groom CR, Alex A (2004) Ligand efficiency: a useful metric for lead selection. Drug Discov Today 9:430–431
Abad-Zapatero C, Metz JT (2005) Ligand efficiency indices as guideposts for drug discovery. Drug Discov Today 10:464–469
Verdonk ML, Rees DC (2008) Group efficiency: a guideline for hits-to-leads chemistry. ChemMedChem 3:1179–1180
Reynolds CH, Bembenek SD, Tounge BA (2007) The role of molecular size in ligand efficiency. Bioorg Med Chem Lett 42:4258–4261
Reynolds CH, Tounge BA, Bembenek SD (2008) Ligand binding efficiency: trends, physical basis, and implications. J Med Chem 51:2432–2438
Bembenek SD, Tounge BA, Reynolds CH (2009) Ligand efficiency and fragment-based drug discovery. Drug Discov Today 14:278–283
Nissink JWM (2009) Simple size-independent measure of ligand efficiency. J Chem Inf Model 49:1617–1622
Orita M, Ohno K, Niimi T (2009) Two ‘golden ratio’ indices in fragment-based drug discovery. Drug Discov Today 14:321–328
Leeson PD, Springthorpe B (2007) The influence of drug-like concepts on decision-making in medicinal chemistry. Nat Rev Drug Discov 6:881–890
Keseru GM, Makara GM (2009) The influence of lead discovery strategies on the properties of drug candidates. Nat Rev Drug Discov 8:203–212
Hajduk PJ (2006) Fragment-based drug design: how big is too big? J Med Chem 49:6972–6976
Perola E (2010) An analysis of the binding efficiencies of drugs and their leads in successful drug discovery programs. J Med Chem 53:2986–2997
Abad-Zapatero C, Perisic O, Wass J, Bento AP, Overington J, Al-Lazikani B, Johnson ME (2010) Ligand efficiency indices for an effective mapping of chemico-biological space: the concept of an atlas-like representation. Drug Discov Today 15:804–811
Schultes S, de Graaf C, Haaksma EEJ, de Esch IJP, Leurs R, Kramer O (2010) Ligand efficiency as a guide in fragment hit selection and optimization. Drug Discov Today Technol 7:e157–e162
Ferenczy GG, Keseru GM (2010) Thermodynamics guided lead discovery and optimization. Drug Discov Today 15:919–932
Ladbury JE, Klebe G, Freire E (2010) Adding calorimetric data to decision making in lead discovery: a hot tip. Nat Rev Drug Discov 9:23–27
Teague SJ et al (1999) The design of lead like combinatorial libraries. Angew Chem Int Ed Engl 38:3743–3748
Oprea TI et al (2001) Is there a difference between leads and drugs? A historical perspective. J Chem Inf Comput Sci 41:1308–1315
Hann MM, Oprea TI (2004) Pursuing the leadlikeness concept in pharmaceutical research. Curr Opin Chem Biol 8:255–263
Dalvit C (2009) NMR methods in fragment screening: theory and a comparison with other biophysical techniques. Drug Discov Today 14:1051–1057
Retra K, Irth H, Muijlwijk-Koezen JE (2010) Surface plasmon resonance biosensor analysis as a useful tool in FBDD. Drug Discov Today Technol 7:e181–e187
Jhoti H, Cleasby A, Verdonk M, Williams G (2007) Fragment-based screening using X-ray crystallography and NMR spectroscopy. Curr Opin Chem Biol 11:485–493
Shuker SB, Hajduk PJ, Meadows RP, Fesik SW (1996) Discovering high affinity ligands for proteins: SAR by NMR. Science 274:1531–1534
Hajduk PJ, Augeri DJ, Mack J, Mendoza R, Yang J, Betz SF, Fesik SW (2000) NMR-based screening of proteins containing 13 C-labeled methyl groups. J Am Chem Soc 122:7898–7904
Mayer M, Meyer B (2001) Group epitope mapping by saturation transfer difference NMR to identify segments of a ligand in direct contact with a protein receptor. J Am Chem Soc 123:6108–6117
Wang YS, Liu D, Wyss DF (2004) Competition STD NMR for the detection of high-affinity ligands and NMR-based screening. Magn Reson Chem 42:485–499
McCoy MA, Senior MM, Wyss DF (2005) Screening of protein kinases by ATP-STD NMR spectroscopy. J Am Chem Soc 127:7978–7979
Dalvit C (2008) Theoretical analysis of the competition ligand-based NMR experiments and selected applications to fragment screening and binding constant measurements. Concepts Magn Reson A 32A:341–372
Dalvit C, Ardini E, Flocco M, Fogliatto GP, Mongelli N, Veronesi M (2003) A general NMR method for rapid, efficient, and reliable biochemical screening. J Am Chem Soc 125:14620–14625
Jacoby E, Davies J, Blommers MJ (2003) Design of small molecule libraries for NMR screening and other applications in drug discovery. Curr Top Med Chem 3:11–23
Baurin N, Aboul-Ela F, Barril X, Davis B, Drysdale M, Dymock B, Finch H, Fromont C, Richardson C, Simmonite H, Hubbard RE (2004) Design and characterization of libraries of molecular fragments for use in NMR screening against protein targets. J Chem Inf Comput Sci 44:2157–2166
Blomberg N, Cosgrove DA, Kenny PW, Kolmodin K (2009) Design of compound libraries for fragment screening. J Comput Aided Mol Des 23:513–525
Chen IJ, Hubbard RE (2009) Lessons for fragment library design: analysis of output from multiple screening campaigns. J Comput Aided Mol Des 23:603–620
Boyd SM, de Kloe GE (2010) Fragment library design: efficiently hunting drugs in chemical space. Drug Discov Today Technol 7:e173–e180
Gleeson MP (2008) Generation of a set of simple, interpretable ADMET rules of thumb. J Med Chem 51:817–834
Wenlock MC, Austin RP, Barton P, Davis AM, Leeson PD (2003) A comparison of physicochemical property profiles of development and marketed oral drugs. J Med Chem 46:1250–1256
Mortenson PN, Murray CW (2009) Ligand lipophilicity efficiency – assessing lipophilicity of fragments and early hits. Presented at RSC Fragments 2009, Astra Zeneca Alderley Park, UK, 4–5 March 2009, Poster 9
Freire E (2008) Do enthalpy and entropy distinguish first in class from best in class? Drug Discov Today 13:869–874
Scott AD, Phillips C, Alex A, Flocco M, Bent A, Randall A, O’Brien R, Damian L, Jones LH (2009) Thermodynamic optimisation in drug discovery: a case study using carbonic anhydrase inhibitors. ChemMedChem 4:1985–1989
Ward HJ, Holdgate GA (2001) Isothermal titration calorimetry in drug discovery. Prog Med Chem 38:309–376
Chung S, Parker JB, Bianchet M, Amzel LM, Stivers JT (2009) Impact of linker strain and flexibility in the design of a fragment-based inhibitor. Nat Chem Biol 5:407–413
Barelier S, Pons J, Marcillat O, Lancelin J-M, Krimm I (2010) Fragment-based deconstruction of Bcl-xL inhibitors. J Med Chem 53:2577–2588
Ji H, Li H, Martasek P, Roman LJ, Poulos TL, Silverman RB (2009) Discovery of highly potent and selective inhibitors of neuronal nitric oxide synthase by fragment hopping. J Med Chem 52:779–797
Levy-Lehad E, Wijsman E, Nemens E, Anderson A, Goddard KA, Weber JL (1995) A familial Alzheimer’s disease locus on chromosome 1. Science 269:970–973
Melnikova I (2007) Therapies for Alzheimer’s disease. Nat Rev Drug Discov 6:341–342
Mount C, Downton C (2006) Alzheimer disease: progress or profit? Nat Med 12:780–784
Moreira PI, Zhu X, Nunomura A, Smith MA, Perry G (2006) Therapeutic options in Alzheimer’s disease. Expert Rev Neurother 6:897–910
Hardy J, Selkoe DJ (2002) The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 297:353–356
Haass C, Selkoe DJ (2007) Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer’s amyloid β-peptide. Nat Rev Mol Cell Biol 8:101–112
Korczyn AD (2008) The amyloid cascade hypothesis. Alzheimers Dement 4:176–178
Hardy J (2006) Has the amyloid cascade hypothesis for Alzheimer’s been proved? Curr Alzheimer Res 3:71–73
Selkoe D (2001) Alzheimer’s disease: genes, proteins, and therapy. Physiol Rev 81:741–766
Archer HA, Edison P, Brooks DJ, Barnes J, Frost C, Yeatman T (2006) Amyloid load and cerebral atrophy in Alzheimer’s disease: a 11 C-BIP positron emission tomography study. Ann Neurol 60:145–147
Olson MI, Shaw CM (1969) Presenile dementia and Alzheimer’s disease in mongolism. Brain 92:147–156
Mann DMA (1998) Alzheimer’s disease and Down’s syndrome. Histopathology 13:125–137
Prasher VP, Farrer MJ, Kessling AM, Fisher EMC, West RJ, Barber SPC, Butler AC (1998) Molecular mapping of IIv Alzheimer-type dementia in Down’s syndrome. Ann Neurol 43:380–383
Pike CJ, Walencewicz AJ, Glabe CG, Cotman CW (1991) In vitro aging of amyloid-beta protein causes peptide aggregation and neurotoxicity. Brain Res 573:311–314
Lorenzo A, Yankner B (1994) Beta-amyloid neurotoxicity requires fibril formation and is inhibited by congo red. Proc Natl Acad Sci USA 91:12243–12247
Iversen LL, Mortishire-Sith RJ, Pollack SJ, Shearman MS (1995) The toxicity in vitro of beta-amyloid protein. Biochem J 311:1–16
Tsai J, Grutzendler J, Duff K, Gan W (2004) Fibrillar amyloid deposition leads to local synaptic abnormalities and breakage of neuronal branches. Nat Neurosci 7:1181–1183
Mor F, Monsonego A (2006) Immunization therapy in Alzheimer’s disease. Expert Rev Neurother 6:653–659
Qu B, Boyer PJ, Johnston SA, Hynan LS, Rosenberg RN (2006) Aβ42 gene vaccination reduces brain amyloid plaque burden in transgenic mice. J Neurol Sci 244:151–158
Bayer AJ, Bullock R, Jones RW, Wilkinson D, Paterson KR, Jenkins L (2005) Evaluation of the safety and immunogenicity of syntheticAβ42 (AN1792) in patients with AD. Neurology 64:94–101
Gilman S, Koller M, Black RS, Jenkins L, Griffith SG, Fox NC (2005) Clinical effects of Aβ immunization (AN1792) in patients with AD in an interrupted trial. Neurology 64:1553–1562
Hock C, Konietzko U, Streffer JR, Tracy J, Signorell A, Muller-Tilmanns B (2003) Antibodies against b-amyloid slow cognitive decline in Alzheimer’s disease. Neuron 38:547–554
Fox NC, Black RS, Gilman S, Rossor MN, Griffith SG, Jenkins L (2005) Effects of Ab immunization (AN1792) on MRI measures of cerebral volume in Alzheimer disease. Neurology 64:1563–1572
DaSilva KA, Aubert I, McLaurin J (2006) Vaccine development for Alzheimer’s disease. Curr Pharm Des 12:4283–4293
Hussain I, Powell D, Howlett DR, Tew DG, Meek TD, Chapman C, Gloger IS, Murphy KE, Southan CD, Ryan DM, Smith TS, Simmons DL, Walsh FS, Dingwall C, Christie G (1999) Identification of a novel aspartic protease (Asp 2) as beta-secretase. Mol Cell Neurosci 14:419–427
Sinha S, Anderson JP, Barbour R, Basi GS, Caccavello R, Davis D, Doan M, Dovey HF, Frigon N, Hong J, Jacobson-Croak K, Jewett N, Keim P, Knops J, Lieberburg I, Power M, Tan H, Tatsuno G, Tung J, Schenk D (1999) Purification and cloning of amyloid precursor protein beta secretase from human brain. Nature 402:537–540
Vassar R, Bennett BD, Babu-Khan S, Kahn S, Mendiaz EA, Denis P, Teplow DB, Ross S, Amarante P, Loeloff R, Luo Y, Fisher S, Fuller J, Edenson S, Lile J, Jarosinski MA, Biere AL, Curran E, Burgess T, Louis JC (1999) Beta-secretase cleavage of Alzheimer’s amyloid precursor protein by the transmembrane aspartic protease BACE. Science 286:735–741
Yan R, Bienkowski MJ, Shuck ME, Miao H, Tory MC, Pauley AM, Brashier JR, Stratman NC, Mathews WR, Buhl AE, Carter DB, Tomasselli AG, Parodi LA, Heinrikson RL, Gurney ME (1999) Membrane-anchored aspartyl protease with Alzheimer’s disease beta-secretase activity. Nature 402:533–537
Farzan M, Schnitzler CE, Vasilieva N, Leung D, Choe H (2000) BACE2, a beta-secretase homolog, cleaves at the beta site and within the amyloid-beta region of the amyloid-beta precursor protein. Proc Natl Acad Sci USA 97:9712–9717
Saunders AJ, Kim T-M, Tanzi RE (1999) BACE maps to chromosome 11 and a BACE homolog, BACE2, reside in the obligate Down syndrome region of chromosome 21. Science 286:1255a
Luo Y, Bolon B, Kahn S, Bennett BD, Babu-Khan S, Denis P, Fan W, Kha H, Zhang J, Gong Y, Martin L, Louis JC, Yan Q, Richards WG, Citron M, Vassar R (2001) Mice deficient in BACE1, the Alzheimer’s beta secretase, have normal phenotype and abolished beta-amyloid generation. Nat Neurosci 4:231–232
Luo Y, Bolon B, Damore MA, Fitzpatrick D, Liu H, Zhang J, Yan Q, Vassar R, Citron M (2003) BACE1 (beta secretase) knockout mice do not acquire compensatory gene expression changes or develop neural lesions over time. Neurobiol Dis 14:81–88
Willem M, Garratt AN, Novak B, Citron M, Kaufmann S, Rittger A, DeStrooper B, Saftig P, Birchmeier C, Haass C (2006) Control of peripheral nerve myelination by the β-secretase BACE1. Science 314:664–666
Wang H, Song L, Laird F, Wong PC, Lee H-K (2008) BACE1 knock-outs display deficits in activity-dependent potentiation of synaptic transmission at mossy fiber to CA3 synapses in the hippocampus. J Neurosci 28:8677–8681
Ohno M, Sametsky EA, Younkin LH, Oakley H, Younkin SG, Citron M, Vassar R, Disterhoft JF (2004) BACE1 deficiency rescues memory deficits and cholinergic dysfunction in a mouse model of Alzheimer’s disease. Neuron 41:27–33
Ghosh AK, Shin D, Downs D, Koelsch G, Lin X, Ermolieff J, Tang J (2000) Design of potent inhibitors for human brain memapsin 2 (β-secretase). J Am Chem Soc 122:3522–3523
Hong L, Koelsch G, Lin X, Wu S, Terzyan S, Ghosh AK, Zhang XC, Tang J (2000) Structure of the protease domain of memapsin 2 (β-secretase) complexed with inhibitor. Science 290:150–153
Stachel SJ (2009) Progress toward the development of a viable BACE-1 inhibitor. Drug Dev Res 70:101–110
Durham TB, Shepherd TA (2006) Progress toward the discovery and development of efficacious BACE inhibitors. Curr Opin Drug Discov Dev 9:776–791
Maillard MC, Hom RK, Benson TE, Moon JB, Mamo S, Bienkowski M, Tomasselli AG, Woods DD, Prince DB, Paddock DJ, Emmons TL, Tucker JA, Dappen MS, Brogley L, Thorsett ED, Jewett N, Sinha S, Varghese J (2007) Design, synthesis, and crystal structure of hydroxyethyl secondary amine-based peptidomimetic inhibitors of human beta -secretase. J Med Chem 50:776–781
Wang YS, Strickland C, Voigt JH, Kennedy ME, Beyer BM, Senior MM, Smith EM, Nechuta TL, Madison VS, Czarniecki M, McKittrick BA, Stamford AW, Parker EM, Hunter JC, Greenlee WJ, Wyss DF (2010) Application of fragment-based NMR screening, X-ray crystallography, structure-based design, and focused chemical library design to identify novel μM leads for the development of nM BACE-1 (β-site APP cleaving enzyme 1) inhibitors. J Med Chem 53:942–950
Zhu Z, Sun ZY, Ye Y, Voigt J, Strickland C, Smith EM, Cumming J, Wang L, Wong J, Wang YS, Wyss DF, Chen X, Kuvelkar R, Kennedy ME, Favreau L, Parker E, McKittrick BA, Stamford A, Czarniecki M, Greenlee W, Hunter JC (2010) Discovery of cyclic acylguanidines as highly potent and selective β-site amyloid cleaving enzyme (BACE) inhibitors: part I – inhibitor design and validation. J Med Chem 53:951–965
Geschwindner S, Olsson LL, Albert JS, Deinum J, Edwards PD, de Beer T, Folmer RH (2007) Discovery of a novel warhead against β-secretase through fragment-based lead generation. J Med Chem 50:5903–5911
Kuglstatter A, Stahl M, Peters JU, Huber W, Stihle M, Schlatter D, Benz J, Ruf A, Roth D, Enderle T, Hennig M (2008) Tyramine fragment binding to BACE-1. Bioorg Med Chem Lett 18:1304–1307
Murray CW, Callaghan O, Chessari G, Cleasby A, Congreve M, Frederickson M, Hartshorn MJ, McMenamin R, Patel S, Wallis N (2007) Application of fragment screening by X-ray crystallography to β-secretase. J Med Chem 50:1116–1123
Congreve M, Aharony D, Albert J, Callaghan O, Campbell J, Carr RA, Chessari G, Cowan S, Edwards PD, Frederickson M, McMenamin R, Murray CW, Patel S, Wallis N (2007) Application of fragment screening by X-ray crystallography to the discovery of aminopyridines as inhibitors of β-secretase. J Med Chem 50:1124–1132
Edwards PD, Albert JS, Sylvester M, Aharony D, Andisik D, Callaghan O, Campbell JB, Carr RA, Chessari G, Congreve M, Frederickson M, Folmer RH, Geschwindner S, Koether G, Kolmodin K, Krumrine J, Mauger RC, Murray CW, Olsson LL, Patel S, Spear N, Tian G (2007) Application of fragment-based lead generation to the discovery of novel, cyclic amidine β-secretase inhibitors with nanomolar potency, cellular activity, and high ligand efficiency. J Med Chem 50:5912–5925
Yang W, Fucini RV, Fahr BT, Randal M, Lind KE, Lam MB, Lu W, Lu Y, Cary DR, Romanowski MJ, Colussi D, Pietrak B, Allison TJ, Munshi SK, Penny DM, Pham P, Sun J, Thomas AE, Wilkinson JM, Jacobs JW, McDowell RS, Ballinger MD (2009) Fragment-based discovery of nonpeptidic BACE-1 inhibitors using tethering. Biochemistry 48:4488–4496
Godemann R, Madden J, Krämer J, Smith M, Fritz U, Hesterkamp T, Barker J, Höppner S, Hallett D, Cesura A, Ebneth A, Kemp J (2009) Fragment-based discovery of BACE1 inhibitors using functional assays. Biochemistry 48:10743–10751
Madden J, Dod JR, Godemann R, Kraemer J, Smith M, Biniszkiewicz M, Hallett DJ, Barker J, Dyekjaer JD, Hesterkamp T (2010) Fragment-based discovery and optimization of BACE1 inhibitors. Bioorg Med Chem Lett 20:5329–5333
Wang Y-S, Beyer BM, Senior MM, Wyss DF (2005) Characterization of autocatalytic conversion of precursor BACE1 by heteronuclear NMR spectroscopy. Biochemistry 44:16594–16601
Liu D, Wang Y-S, Gesell JJ, Wilson E, Beyer BM, Wyss DF (2004) Backbone resonance assignments of the 45.3 kDa catalytic domain of human BACE1. J Biomol NMR 29:425–426
Stamford A (2010) Discovery of small molecule, orally active and brain penetrant BACE1 inhibitors. Paper presented at 239th ACS National Meeting, San Francisco, CA, 21–25 March 2010
Acknowledgments
We would like to thank Mark McCoy and Jennifer Gesell for their many successful contributions to fragment-based NMR screening and FBDD, and Zhong-Yue Sun, Matthew E. Kennedy, Brian M. Beyer, Mary M. Senior, Elizabeth M. Smith, Terry L. Nechuta, Yuanzan Ye, Jared Cumming, Lingyan Wang, Jesse Wong, Xia Chen, Reshma Kuvelkar, Leonard Favreau, Vincent S. Madison, Michael Czarniecki, Brian A. McKittrick, Eric M. Parker, John C. Hunter, and William J. Greenlee for their invaluable contributions to the BACE-1 work and for their exceptional teamwork, as well as many other colleagues who have contributed to the success of this project over the years.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin-Heidelberg
About this chapter
Cite this chapter
Wyss, D.F. et al. (2011). Combining NMR and X-ray Crystallography in Fragment-Based Drug Discovery: Discovery of Highly Potent and Selective BACE-1 Inhibitors. In: Davies, T., Hyvönen, M. (eds) Fragment-Based Drug Discovery and X-Ray Crystallography. Topics in Current Chemistry, vol 317. Springer, Berlin, Heidelberg. https://doi.org/10.1007/128_2011_183
Download citation
DOI: https://doi.org/10.1007/128_2011_183
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-27539-5
Online ISBN: 978-3-642-27540-1
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)