Abstract
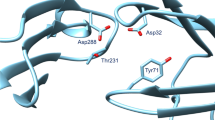
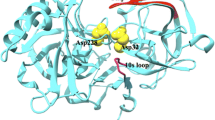
Computational docking and scoring techniques have revolutionized structural bioinformatics by providing unprecedented insights on key aspects of ligand-receptor interaction. Docking is used for optimizing known drugs and for identifying novel binders by predicting their binding mode and affinity. AutoDock and AutoDockTools are free of charge techniques that have been extensively cited in the literature as essential tools in structure-based drug design. Moreover, these methods are fast enough to permit virtual screening of ligand libraries containing tens of thousands of compounds. However using Autodock requires some knowledge in programming which creates a limitation for biologists and makes them prone for commercial applications. Here, we selected a relevant target involved in the progression of Alzheimer disease and provided a fully reproducible docking protocol. This example will show how docking techniques would be an important asset to identify new BACE1 inhibitors. The following friendly user tutorial targets both undergraduate and graduate students, allowing them to understand docking as a computational tool for structure-based drug design.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Forli S, Huey R, Pique ME, Sanner MF, Goodsell DS, Olson AJ (2016) Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat Protoc 11:905–919
Siddiquee K, Zhang S, Guida WC, Blaskovich MA, Greedy B, Lawrence HR, Yip ML, Jove R, McLaughlin MM, Lawrence NJ, Sebti SM, Turkson J (2007) Selective chemical probe inhibitor of Stat3, identified through structure-based virtual screening, induces antitumor activity. Proc Natl Acad Sci U S A 104:7391–7396
Li C, Xu L, Wolan DW, Wilson IA, Olson AJ (2004) Virtual screening of human 5-aminoimidazole-4-carboxamide ribonucleotide transformylase against the NCI diversity set by use of AutoDock to identify novel nonfolate inhibitors. J Med Chem 47:6681–6690
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Javaid FZ, Brenton J, Guo L, Cordeiro MF (2016) Visual and ocular manifestations of Alzheimer's disease and their use as biomarkers for diagnosis and progression. Front Neurol 7:55
Yan R, Vassar R (2014) Targeting the beta secretase BACE1 for Alzheimer's disease therapy. Lancet Neurol 13:319–329
Chen JJ, Liu Q, Yuan C, Gore V, Lopez P, Ma V, Amegadzie A, Qian W, Judd TC, Minatti AE, Brown J, Cheng Y, Xue M, Zhong W, Dineen TA, Epstein O, Human J, Kreiman C, Marx I, Weiss MM, Hitchcock SA, Powers TS, Chen K, Wen PH, Whittington DA, Cheng AC, Bartberger MD, Hickman D, Werner JA, Vargas HM, Everds NE, Vonderfecht SL, Dunn RT 2nd, Wood S, Fremeau RT Jr, White RD, Patel VF (2015) Development of 2-aminooxazoline 3-azaxanthenes as orally efficacious beta-secretase inhibitors for the potential treatment of Alzheimer's disease. Bioorg Med Chem Lett 25:767–774
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
El-Hachem, N., Haibe-Kains, B., Khalil, A., Kobeissy, F.H., Nemer, G. (2017). AutoDock and AutoDockTools for Protein-Ligand Docking: Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1(BACE1) as a Case Study. In: Kobeissy, F., Stevens, Jr., S. (eds) Neuroproteomics. Methods in Molecular Biology, vol 1598. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6952-4_20
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6952-4_20
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6950-0
Online ISBN: 978-1-4939-6952-4
eBook Packages: Springer Protocols