Abstract
A cancer cell agglutinin isolated from bullfrog (Rana catesbeiana) eggs was named R. catesbeiana sialic acid–binding lectin (cSBL), because its agglutinative activity was inhibited by sialic acid–containing glycoconjugates. The lectin was subsequently found to belong to the vertebrate-secreted ribonuclease family, and cSBL exhibits ribonuclease activity as well as remarkable antitumor effects. Recent studies have shown that cSBL exerts antitumor effects in vitro and in vivo under conditions where cSBL cancer-selective apoptosis-inducing effects and no undesired side effects are observed. Furthermore, cSBL exhibited a preferred synergistic effect with other drugs. Here, we describe the unique antitumor signal revealed in the latest cSBL studies and the possibility of developing a cancer treatment strategy using cSBL.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Advances in modern medicine have reduced the mortality rate of infectious diseases such as HIV and tuberculosis, as well as improving general health worldwide. As of 2021, the development of newly approved RNA vaccines has shown remarkable efficacy in the prevention of new infectious diseases, and modern medical care is still evolving day by day. However, global health is still threatened by diseases such as cancer, heart disease, and cerebrovascular disease (WHO 2020). Despite the remarkable advances in cancer chemotherapy in recent years, for example, immune checkpoint inhibitors (Robert 2020), cancer is still the leading cause of death in many countries. It is estimated that as many as ten million cancer-related deaths occurred in 2020 (Sung et al. 2021). Thus, therapeutic advances in cancer prevention, early detection, and treatment will improve survival and quality of life for many people worldwide.
Sugar chains (glycans) are termed third life chains, coming after nucleic acids and proteins to play essential roles in living bodies via their various biochemical interactions. Such as binding with functional molecules (Reily et al. 2019). The surfaces of cells are covered with innumerable glycans, and the activities of such glycoproteins or glycolipids can be controlled via glycosylation. Cell-surface glycans can significantly alter their sequence structure during cell differentiation, maturation, and activation, and function as discriminating molecules (Marth and Grewal 2008). For example, differentiated cells recognize glycan structures on other cells and cooperate with each other (Varki 2017). Microorganisms such as bacteria and viruses use differences in surface glycans to bind to and infect specific cells (Van Breedam et al. 2014). Furthermore, surface glycan structure also changes significantly in cancer cells (Costa et al. 2020). Changes in glycan structure have been shown to alter the location of cell surface receptors and their susceptibility to ligands, affecting cell proliferation and invasion (Rodrigues et al. 2018). It also affects the cancer-surveillance activity of immune cells (Mereiter et al. 2019). Therefore, glycans are very useful for identifying cells and are thus expected to be utilizable biomarkers for identifying cancer cells. In fact, many biomarkers currently in clinical use recognize glycans (Tikhonov et al. 2020). Hence, glycans are currently attracting much attention, but research on them is somewhat difficult. The structures of sugar chains are complicated and liable to change due to their sensitivity to environmental factors, and can even be decomposed. Furthermore, the structure cannot be directly amplified unlike nucleic acids. Therefore, it is difficult to recognize, analyze, and quantify certain disease-related glycans, and development of tools for recognizing glycans is required for their use in diagnosis and treatment.
Lectins are useful tools for targeting difficult-to-handle glycans. Our laboratory has focused on the identification and analysis of naturally derived lectins for many years, and has purified and reported some lectins from fish and amphibians. Specifically, sialic acid–binding lectin purified from bullfrog (Rana catesbeiana) eggs (cSBL) has potential as a cancer treatment due to its antitumor activity via a unique RNA-degradation mechanism. In this section, we describe in detail the functions of cSBL that have been clarified thus far, including background of the discovery of this lectin and the possibility of its application in cancer treatment.
2 Discovery of cSBL as a Lectin
2.1 Lectins
Lectins are proteins that bind to specific glycan structures and are universally present in species from animals and plants to microorganisms, and have greatly contributed to glycan analysis research. Historically, the first lectin identified is considered to be ricin from castor tree (Ricinus communis) seeds by Stillmark in 1888. Ricin is a plant-derived protein capable of agglutinating red blood cells, and the name hemagglutinins or phytoagglutinins was used at the time (Sharon and Lis 2004). Sumner and Howell reported that concanavalin A, a lectin isolated from jack bean (Canavalia ensiformis), agglutinates cells such as erythrocytes, and this agglutination was inhibited by sucrose, the first demonstration of the sugar specificity of lectins (Sumner and Howell 1936). It was predicted that agglutination was caused by the reaction of concanavalin A with carbohydrates on the cell surface. Since the 1940s, there have been many discoveries of blood type-specific hemagglutinins (Mäkelä 1957; Sharon and Lis 2004). These findings demonstrated the ability of plant agglutinins to distinguish between erythrocytes of different blood types, and thus, Boyd and Shapleigh proposed the name “lectins,” from the Latin legere, meaning to pick out or choose (Boyd and Shapleigh 1954). Subsequently, the term “lectin” was generalized by Sharon et al. to all sugar-specific agglutinins of nonimmune origin, regardless of source and blood group specificity (Sharon and Lis 1972). Goldstein et al. (Goldstein et al. 1980) further defined lectins in detail: “A lectin is a sugar-binding protein or glycoprotein of non-immune origin which agglutinates cells and/or precipitates glycoconjugates. Lectins bear at least two sugar-binding sites, and agglutinate animal and/or plant cells. The specificity of a lectin is usually defined in terms of the monosaccharide(s) or oligosaccharides that inhibit lectin-induced agglutination reactions.” Generally, lectin is used as a general term for substances that have specific binding activity for sugars among proteins or glycoproteins present in living organisms. With the subsequent research into numerous lectins, numerous pivotal biological functions of lectins have been revealed, such as immunomodulating effects (Clement et al. 2010), modulation of the intestinal transport system (Yamamoto et al. 2013), and antimicrobial and insecticidal activities (Dias et al. 2015), among others. Furthermore, many lectins are currently used as important biological and diagnostic tools. Recently, a lectin array technology that acquires sugar chain profiles of various biological samples via their reactivity with multiple lectins has been developed (Hirabayashi et al. 2013), and a new therapeutic approach that uses lectins to target specific cells using lectin-drug conjugates (LDC) has also been reported (Shimomura et al. 2018). Thus, novel research and development using lectins has drawn attention in a wide range of fields, from technological development to diagnosis and medical treatment.
2.2 Sialic Acid–Binding Lectin from Rana Catesbeiana Oocytes
In our laboratory, research on frog egg agglutinin was started by Kawauchi et al. in the mid-1970s (Kawauchi et al. 1975). Initially, crude egg fractions were used, and samples derived from Japanese brown frog (Rana japonica) and dark-spotted frog (Rana nigromaculata) eggs showed agglutinative activity with rat ascites cancer cells (AH109A) (Yokota et al. 1975). Later, it was found that bullfrog (R. catesbeiana) egg extract, which belongs to the same genus Rana, contains two lectins that show different agglutinative activities, one with human blood group A erythrocytes, and the other with mouse Ehrlich ascites carcinoma cells and AH109A. These two lectins were separated by column chromatography, and their binding specificity, agglutinative activity, and physical and chemical properties were studied. One of the lectins showed preferential agglutination of cancer cells and displayed specific binding to sialyl glycoproteins (Sakakibara et al. 1977), and was consequently designated sialic acid–binding lectin, cSBL. In 1987, Nitta et al. isolated cSBL and characterized its lectin activity (Nitta et al. 1987). cSBL preferentially agglutinates a large variety of tumor cells, but not normal red blood cells, lymphocytes, or fibroblasts. This phenomenon correlates with a higher binding activity of cSBL with tumor cells. The binding reactivity of cSBL was tested in 20 human and animal cancer cells and 10 normal cells, and it was observed that the majority of cancer cells showed high reactivity to cSBL, while normal cells (fibroblasts, lymphocytes, and erythrocytes) exhibited low reactivity (10–50% reactivity compared to S-180 mouse ascites cells used as a standard). Tumor cell agglutination induced by cSBL was inhibited by mucin, fetuin, and keratan sulfate, but not by less-sialylated glycoproteins, such as transferrin. Inhibition by mucin or fetuin was greatly reduced by the desialylation of the glycoproteins with sialidase. Treatment of tumor cells with sialidase reduced cSBL-induced agglutination, and sialidase-dependent reduction of tumor cell agglutination was inhibited by the sialidase inhibitor. However, tumor cell agglutination was not inhibited by chondroitin sulfate or hyaluronic acid. Thus, lectin-dependent tumor cell agglutination was considered to be due to the high density of sialic acid on the cell surface. When the effects of glycosaminoglycans on the agglutinative activity of cSBL were tested, heparin showed strong inhibitory effects, and it was speculated that heparin interfered with the binding of cSBL to cells. Polyamines also inhibit cell agglutination, and it has been proposed that they interact with negatively charged cell surface components, including sialosyl residues (i.e., the lectin-binding region), resulting in inhibition of agglutination. These results suggest that tumor cell agglutination occurs because of the recognition of sialic acid–containing molecules on the cell surface by cSBL. The position of the sialic acid–binding site of cSBL remains unknown, but Irie et al. reported two putative sialic acid–binding sites on the cSBL structure. One is a site near the N-terminus, where sialic acid binds mainly via hydrogen bonds, and the other is a loop consisting of amino acid residues 57–75, where sialic acid binds via hydrophobic interactions. The binding of sialic acid to the loop was thought to be very weak, and the weakly bound sialic acid was considered to be easily washed out by soaking in buffer without sialic acid (Irie et al. 1998). Due to this weak binding to the loop site, and no reports of cSBL-homologous lectins so far, the lectin activity of cSBL remains unclear, especially regarding the structure of the binding target. Nevertheless, the complete amino acid sequence of cSBL was presented by Titani et al. in 1987 (Titani et al. 1987). The 111-residue sequence was determined by peptide sequencing. Initially, this protein was considered to be a new type of lectin, as its sequence was unique and no homology to known protein sequences was found in the database at the time. However, 3 years later, the sequence of a homolog, Rana japonica egg lectin (jSBL), was determined, and a homology search with the updated protein sequence data bank revealed that these amino acid sequences were homologous to those of the pancreatic ribonuclease family (vertebrate-secreted RNase family) (Kamiya et al. 1990). The sequence identities with human angiogenin and pancreatic RNase, members of the vertebrate-secreted RNase family, were 35% and 37%, respectively. Similarities were especially apparent in the four proteins (cSBL, jSBL, human angiogenin, and pancreatic ribonuclease) around His-12, Lys-41, and His-119, the major active site residues conserved in the ribonucleases, and around three of the four disulfide bonds. Thus, research created new insights into cSBL as a ribonuclease.
3 cSBL as an RNase
RNase is a ribonucleic-acid-degrading enzyme that is ubiquitous in all cells, and is a pioneer of enzyme research due to its discovery over 100 years ago and its subsequent thorough study. Enzymatically, it is roughly divided into endo- and exo-types, but there are numerous types present in all species from animals and plants to microorganisms. Since the number of reports on RNases is still increasing, their classification method, including naming, has changed frequently. RNase A, which is reportedly the most studied enzyme in the history of science, is purified from the bovine pancreas. Although a structural family consisting of RNase A and its homologs (more than 100 species) has commonly been called the RNase A superfamily (Beintema et al. 1986, 1997), in recent years, the renaming of this superfamily as the vertebrate-secreted RNase superfamily has been proposed by D’Alessio, because most are secretory proteins produced by vertebrates (D’Alessio 2011). RNases can be classified into the vertebrate-secreted (RNase A), T1, T2, H, L, and P families, among others (Deshpande and Shankar 2002; Fang and Ng 2011). Vertebrate-secreted RNases are relatively small, containing approximately 130 amino acids. They preferentially cleaved pyrimidines at an optimal pH of 7–8. This group of RNases has a characteristic peptide motif (CKXXNTF) in the middle of the sequence and is encoded by a single exon (D’Alessio 2011). They contain two His and one Lys as catalytic active centers and 6–8 cysteines, forming 3–4 disulfide bonds (Cho et al. 2005). Sequence identity varies from 20% to nearly 100%, and their three-dimensional structures are very similar, consisting mainly of three α-helices and four antiparallel β-sheet structures. This structure is also termed kidney-shape, as it is reminiscent of a kidney (Raines 1998).
Immediately after cSBL was found to be a vertebrate-secreted RNase, its RNase activity was investigated (Okabe et al. 1991). The base specificity of cSBL was studied using eight dinucleotide phosphates as substrates and eight nucleotides as inhibitors. The base specificities of the B1 and B2 sites of cSBL were U > C and G > U > A, C, respectively. cSBL was more resistant than RNase A to heat treatment, guanidine-HCl, and pH-induced denaturation; retaining its native conformation up to at least 70 °C at pH 7.5. Agglutination of AH109A cells by cSBL was inhibited by nucleotides, indicating that the agglutination sites are related to the catalytic sites for RNase activity of cSBL. In 1992, Liao et al. purified a pyrimidine-guanine sequence-specific RNase (termed RC-RNase) from R. catesbeiana oocytes, which was found to be identical to cSBL (Liao 1992). Further study clarified that although cSBL lacked a Cys-65–Cys-72 disulfide bond, the locations of the other three disulfide bonds were similar to those of the RNase A family. cSBL was found to exhibit RNase activity and catalytic properties resembling those of bovine RNase A. For example, cSBL hydrolyzed poly-uridylic acid and poly-cytidylic acid and preferred the former (Nitta et al. 1993). Nitta et al. thus proposed the name catalytic lectin (leczyme) as a multifunctional molecule with lectin and enzymatic activity (Nitta et al. 1996). It is also noteworthy that even though members of human vertebrate-secreted RNase were strongly inhibited by the human placental RNase inhibitor, the RNase and tumor cell agglutinative activity of cSBL were not affected by the inhibitor (Nitta et al. 1993). This RNase inhibitor-refractory property has a decisive effect on the antitumor activity of cSBL, described in detail in Sect. 5.
4 cSBL as an Antitumor RNase
4.1 Cytotoxic RNases: Evasion of RNase Inhibitors
To date, many antitumor RNases have been identified in a variety of species, including plants, bacteria, fungi, and animals, across RNase family types. Well-known and representative antitumor RNases mainly belong to the vertebrate-secreted RNase family (D’Alessio 2011). Bovine pancreatic RNase A has been thoroughly studied from all aspects, including physicochemical, enzymatic, biochemical, and molecular biology. Since RNase A exhibits RNA-degrading activity, its antitumor activity was investigated more than half a century ago. Although cytotoxicity was observed at high doses (on the order of mg) (Section et al. 1955; LeDoux 1955), subsequent studies revealed that no significant activity is observed at practically low concentrations (de Lamirande 1961; Roth 1963). The major reason for the restricted antitumor effect of RNase A is the neutralization of its ribonucleolytic activity by the endogenous cytosolic RNase inhibitor.
Human RNase inhibitor is a 450 residues, 49 kDa protein with an isoelectric point of 4.7. It contains leucine-rich repeats, forms strong complexes with mammalian ribonucleases, and plays an important role in controlling the life span of RNA (Vicentini et al. 1990; Kobe and Deisenhofer 1995). It is the major intracellular protein making up approximately 0.1% of the total cytosolic protein and is highly conserved across various mammalian species (Lee and Vallee 1993; D’Alessio and Riordan 1997). RNase inhibitors form a 1:1 noncovalent complex with mammalian vertebrate-secreted RNase family members, and the dissociation constant is of the order of fM, indicating strong affinity (Vicentini et al. 1990; Kobe and Deisenhofer 1996). Therefore, the presence of this inhibitor is inevitably a barrier for RNase-mediated cytotoxicity. Consequently, cytotoxic RNases must satisfy one of the following three conditions: (1) insensitive to the RNase inhibitor (Dickson et al. 2005), (2) present at sufficient levels to saturate the RNase inhibitor thus access RNA (Leich et al. 2007), or (3) binds to other molecules or localizes in organelles where RNase inhibitor is not present, such as the nucleus, so binding to RNase inhibitor is reduced (Gaur et al. 2001; Bosch et al. 2004). Indeed, when RNase inhibitor-sensitive noncytotoxic RNases such as RNase A are injected directly into Xenopus oocytes, which lack strong inhibitors of mammalian RNases, they display cytotoxic activity (Saxena et al. 1991). Currently, onconase, BS-RNase, and PE5 are representatives of known cytotoxic RNases that show antitumor activity while avoiding inhibition by RNase inhibitors.
4.2 Antitumor RNases
4.2.1 Onconase
Onconase (ONC) is a polypeptide containing104 amino acid residues, isolated from leopard frog (Rana pipiens) eggs by Ardelt et al. (1991) belonging to the vertebrate-secreted RNase family (Ardelt et al. 1991). The enzyme activity of ONC is only 1/102 to 1/105 of that of RNase A (Boix et al. 1996). However, the cytotoxicity of ONC is stronger than RNase A, despite its low enzyme activity, due to its mechanism of RNase inhibitor avoidance (Rutkoski and Raines 2008). The antitumor effect of ONC has been verified in many cancer cells, such as hepatoma, colon cancer, and leukemia (Wu et al. 1993; Iordanov et al. 2000). ONC has been studied in clinical trials including patients with malignant mesothelioma, nonsmall cell lung cancer, renal cell cancer, and others (Mikulski et al. 1995; Vogelzang and Stadler 1999; Costanzi et al. 2005). Unfortunately, due to the establishment of other drugs as first-line treatment for malignant mesothelioma (combination of pemetrexed and cisplatin) and the rise of cancer immunotherapy, interest in ONC has diminished, and it has not been approved as a therapeutic drug until now. However, in the previously mentioned clinical trials, ONC was confirmed to prolong the survival period while showing favorable safety, and even now, attempts are being made to use it in combination with other drugs or to utilize an ONC fusion drug (Smolewski et al. 2020).
4.2.2 BS-RNase
BS-RNase, a bovine seminal ribonuclease discovered by Hosokawa and Irie, and Dostal and Matousek in 1972 (Hosokawa and Irie 1971; Dostál and Matoušek 1972; D’Alessio et al. 1972a), is the sole native dimeric member of the vertebrate-secreted RNase family (D’Alessio et al. 1972b; Sorrentino and Libonati 1994). RNase inhibitor binds tightly to monomeric BS-RNase but not to dimeric BS-RNase; thus, it shows remarkable antitumor activity (Murthy and Sirdeshmukh 1992). The cancer-selective antitumor effect of BS-RNase has been confirmed in vitro as well as in vivo in several cancers, including thyroid cancer, melanoma, and seminoma (Poucková et al. 1998; Kotchetkov et al. 2001). Furthermore, Fiorini et al. demonstrated that BS-RNase exhibits strong antiproliferative and proapoptotic effects in pancreatic adenocarcinoma cell lines, and that it triggers Beclin1-mediated autophagy in cancer cells that is ineffective in normal cells (Fiorini et al. 2014). Although it is not completely understood how BS-RNase binds to cancer cells, adsorption to negative charges present on the cell surface has been proposed (Bracale et al. 2002), and BS-RNase has been found to bind to the extracellular matrix, suggesting that the interaction with the extracellular matrix is important for cytotoxicity (Mastronicola et al. 1995; Bracale et al. 2002).
4.2.3 PE5
In humans, the vertebrate-secreted RNase family contains eight members involved in RNA clearance and maturation, and immune defense, while some members have unique physiological effects such as angiogenesis (Koczera et al. 2016). One of the control mechanisms for these functional RNases is RNase inhibitor interference (Dickson et al. 2005). Modification of human RNases has also been attempted to overcome RNase inhibitor obstruction, resulting in cytotoxicity. PE5 is a cytotoxic artificial RNase 1 (human pancreatic RNase, HP-RNase) variant with five mutated residues in the N-terminal domain (Arg4Ala, Lys6Ala, Gln9Glu, Asp16Gly, and Ser17Asn). These mutations result in the RNase having a discontinuous, unconventional nuclear localization signal (NLS) (Rodríguez et al. 2006). The NLS of PE5 is recognized by α-importin, which imports the protein to the nucleus (Bosch et al. 2004). Once inside the nucleus, it degrades nuclear RNA and displays cytotoxicity, although cytoplasmic RNA is unaffected (Tubert et al. 2011). Despite being sensitive to the RNase inhibitor, PE5 demonstrates cytotoxicity against several cancer cell lines (Castro et al. 2011, 2012). PE5 exhibits cytotoxicity via apoptosis associated with p21WAF1/CIP1 pathway induction and the inactivation of JNK in the doxorubicin-resistant ovarian cancer cell line, NCI/ADR-RES. Furthermore, it has cytostatic effects, as evidenced by the increase in the number of cells in S and G2/M cell cycle phases in PE5-treated cells (Castro et al. 2011), and the synergistic effects with doxorubicin were also revealed (Castro et al. 2012). The mechanism observed in the NCI/ADR-RES cells was found to be different from that observed for ONC. The effect of PE5 on NCI/ADR-RES cells has also been studied via transcriptome analysis using microarray technology, revealing that PE5 induces changes in pleiotropic gene expression, especially reducing the expression of genes involved in cancer cell metabolism (Vert et al. 2016). Moreover, studies have been conducted to further modify PE5 to improve its therapeutic potential (Vert et al. 2012) and very recently, one of these variants (NLSPE5) demonstrated cancer-selective cytotoxicity. NLSPE5 reduced migration and invasion of highly invasive breast cancer cells, accompanied by downregulation of N-cadherin expression, and inhibition of cancer stem cell (CSC) development, as well as diminishing the self-renewal capacity of CSCs, suggesting their possible targeting by cytotoxic RNase (Castro et al. 2021).
4.3 cSBL
Since the agglutination activity of cSBL is considered cancer cell-selective and does not interact with human RNase inhibitors, research on cSBL as an anticancer drug candidate has been conducted since its discovery. The substantial antitumor effect of cSBL was first reported by Nitta et al. in 1994 (Nitta et al. 1994a). cSBL significantly inhibited the proliferation of mouse leukemia P388 and L1210 cells in vitro as well as sarcoma 180 and Mep II ascites cells in vivo, while 50 mg/kg injection of cSBL, one-third of the lethal dose for normal mice, did not show any undesired side effects in the in vivo experiment. Meanwhile, the cSBL-resistant P388 cell variant, RC-150, was established (Nitta et al. 1994b). The doubling time, tumorigenicity, and lethality of RC-150 cells were similar to those of parental P388 cells. cSBL agglutinated both P388 and RC-150 cells, and no difference was observed between the sialidase-labile sialic acid levels in RC-150 and P388 cells. The 50% inhibitory concentration of cSBL for growth of P388 cells was approximately 3.1–6.2 μM, whereas concentrations as high as 100 μM were ineffective on the growth of RC-150 cells. Analysis using dansylcadaverine-labeled cSBL showed that the internalization of cSBL was observed in parent P388 cells, but not in resistant RC-150 cells, indicating the importance of not only cell-surface binding but also cell internalization. The addition of benzyl-α-N-acetylgalactosamine (GalNAc) to the culture medium diminished the effect of cSBL, suggesting that the internalization of cSBL may be mediated by O-linked carbohydrate chain(s) of the glycoconjugates (Irie et al. 1998). Thus, the antitumor activity of cSBL is thought to be a result of the coordination between lectin and RNase activity, i.e., recognition of glycoconjugates containing sialic acids on tumor cell surfaces and decomposition of RNA required for cell survival, respectively (Nitta 2001). A few years after the first report of the antitumor effect of cSBL, Liao et al. reported the cancer-selective cytotoxic effect of cSBL: cSBL inhibited the growth of several carcinoma cell lines but did not affect healthy human and mouse fibroblasts (Liao et al. 1996). Subsequently, Hu et al. reported that cSBL was cytotoxic to human hepatoma cell lines at different degrees of differentiation. Interestingly, the cytotoxicity of cSBL in different hepatoma cells correlated with the extent of differentiation, but not the proliferation rate of the cells (Hu et al. 2001a). Wei et al. reported a similar phenomenon in which retinoic acid (RA) or dimethyl sulfoxide (DMSO)-induced differentiation resulted in HL-60 cells becoming resistant to cSBL (Wei et al. 2002). These results indicate that differentiation is a significant factor in the selective cytotoxicity of cSBL. Only a few studies have compared the effects of ONC and cSBL; however, Tang et al. reported that compared with ONC, cSBL harbors more cancer-selective antitumor activity, as ONC was toxic to healthy human HS-68 foreskin fibroblasts, whereas cSBL was not (Tang et al. 2005). Furthermore, Lee et al. showed that cSBL internalization into baby hamster kidney BHK-21 cells was enhanced by Japanese encephalitis virus (JEV) infection, which causes enhanced apoptosis in JEV-infected BHK-21 cells, indicating that cSBL also possesses antiviral activity (Lee et al. 2011). Although there are some unclear or controversial reported mechanisms of cSBL antitumor activity even today, we summarize the signal transduction pathways and related mechanisms of antitumor activity in Sect. 5.
4.4 Benefits of Antitumor RNase in Cancer Treatment
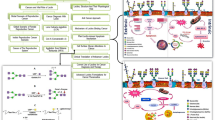
Vertebrate-secreted RNases are expected to be minimally immunogenic because of their compact structure and homology with human pancreatic RNase (Costanzi et al. 2005). They also exhibit high thermal stability and strong resistance to protein denaturants and proteases (Okabe et al. 1991; Notomista et al. 2000; Rosenberg et al. 2001). These features are considered to be one reason for their potent antitumor activity in vivo and provide benefits for commercialization. Furthermore, in current cancer therapies, undesired side effects due to the low cancer cell selectivity of conventional drugs and the emergence of resistance to these drugs have become major issues. Recently, attempts have been made to overcome these issues using molecular-targeted and antibody drugs, but both are still developing fields, and currently, many conventional drugs are used that target DNA. Antitumor RNases as anticancer agents is a new cancer treatment strategy that targets RNA. There are several advantages to this: (1) Cell toxicity of antitumor RNAase has already been demonstrated for many types of cancer. (2) Many cytotoxic RNases have been reported to have high cancer cell selectivity, although the mechanisms have not yet been completely elucidated. (3) They may show synergistic antitumor effects with other drugs in combination treatment. (4) They have no genetic toxicity, unlike conventional DNA-damaging drugs or radiotherapy. (5) There are no reports showing cross-resistance with small molecule compounds, but rather strong cell-death-inducing effects on multidrug-resistant cells. (6) Antitumor RNases can exert pleiotropic effects, because they affect multiple RNA substrates, whereas drugs that target a specific protein have the advantage of being highly specific, but are often inadequate to adapt to the multifactorial complexity of the cancer phenotype, and are often accompanied by the emergence of resistance. There is little information about resistance acquisition in cancer cells against antitumor RNases; thus, it is unclear whether resistance will appear frequently. However, since RNA is targeted, it may be possible to overcome resistance through abnormal gene expression. The advantages of using antitumor RNases for cancer therapy are summarized based on the central dogma in Fig. 1.
Features of therapeutic strategies. Traditional DNA-targeting therapies, including chemotherapy and radiotherapy, are mainly selective for quickly proliferating cells, but are therefore often also toxic to healthy cells. In addition, since they have genotoxicity, it is necessary to pay attention to the development of secondary cancers. In many cases, tolerance is induced through abnormal expression of proteins such as drug efflux pumps. Therapies that target specific molecules, such as antibody therapy and molecular targeted therapy, have the advantage of being nongenotoxic and highly selective. However, the nature of cancer is highly diverse, and targeting only a single protein may be ineffective in some patients. Furthermore, resistance appears due to the acquisition of mutations in target genes and overexpression of abnormal proteins. RNA-targeting therapy is a new approach with several advantages, including nongenotoxicity and pleiotropy. It induces cell death signals by degrading RNA. Although there is little information on resistance and selectivity, it inhibits gene expression, so it may prevent resistance acquisition by abnormal genetic expression, including those for concomitant drugs
5 Antitumor Mechanism of cSBL
5.1 cSBL-Induced Apoptosis
The initial antitumor mechanism of cSBL was thought to be inhibition of protein synthesis via RNA degradation. In fact, reduced incorporation of [3H] leucine was demonstrated in cSBL-treated cells (Nitta et al. 1994a), and treatment with cSBL leads to alterations in signal transduction and intracellular protein kinase cascades, such as decreased intracellular Ca2+ concentration, decreased protein kinase A activity, and increased protein kinase G activity (Nitta et al. 1996). However, subsequent analysis of the antitumor effects revealed that cSBL induced apoptosis. In early experiments with mouse leukemia p388 cells, it was found that cSBL induces apoptotic morphological changes, such as nuclear condensation, the disappearance of microvilli, and biochemical changes, such as caspase-8/−3 activation, phosphatidylserine externalization, and DNA fragmentation. In addition, increased expression of the Fas antigen and tumor necrosis factor (TNF) receptor was observed in cSBL-treated cells (Nitta 2001). Subsequently, cSBL was reported to cause caspase-7 activation in human breast cancer MCF-7 cells that lack caspase-3, and apoptosis is prevented by the overexpression of Bcl-XL in the cells (Hu et al. 2001b).
After 2013, our group reported the validity of cSBL in human leukemia cell lines and the detailed mechanism of cSBL-induced cell death (Tatsuta et al. 2013b). cSBL degrades cellular RNA and manifests significant cytotoxic effects in several types of human leukemia cell lines, including MDR cells, when conventional DNA-damaging clinical agents, such as etoposide and doxorubicin, are not cytotoxic to these cells. cSBL-induced DNA fragmentation was completely blocked by the pan-caspase inhibitor carbobenzoxy-valyl-alanyl-aspartyl- [O-methyl]-fluoromethylketone (z-VAD), suggesting that apoptosis is induced in a caspase-dependent manner. Analysis to identify the cSBL-induced signaling pathway was performed using a combination of specific caspase inhibitors and mitochondrial membrane depolarization–detecting reagents. cSBL-induced mitochondrial depolarization was not diminished by z-VAD, whereas TNF-related apoptosis-inducing ligand (TRAIL)-induced mitochondrial depolarization was completely inhibited by z-VAD. This indicated that the cytotoxicity of cSBL was induced through caspase-dependent apoptosis, whereby mitochondrial perturbation occurs as an upstream event.
Endoplasmic reticulum (ER) stress is also reportedly involved in cSBL-induced apoptosis (Tatsuta et al. 2013a). cSBL induces the endoplasmic unfolded protein response (UPR). Additionally, inhibition of caspase-4, the initiator caspase in ER stress-mediated apoptosis, reduces cSBL-induced apoptosis. In further experiments with specific caspase inhibitors, caspase-9 activation was found to be involved in caspase-4 activation, at least to some extent; however, because cSBL-induced mitochondrial membrane depolarization or UPR is unaffected by caspase-9 or caspase-4 inhibition, respectively, we concluded that mitochondrial perturbations and ER stress occurred independently in cSBL-treated cells. These results suggest that cSBL may induce apoptosis via multiple apoptotic pathways, including ER stress and the mitochondrial pathway. A comparative study using thapsigargin, an inducer of endoplasmic reticulum stress apoptosis, revealed that the mitochondrial pathway contributes significantly to cSBL-induced apoptosis (Tatsuta et al. 2013a).
RNase activity has also been shown to be required for cSBL cytotoxicity; however, it remains unclear how cSBL-induced RNA degradation leads to apoptosis. In our study conducted on human leukemia (Tatsuta et al. 2013b), malignant mesothelioma (Tatsuta et al. 2014b), and breast cancer cell lines (Kariya et al. 2016; Tatsuta et al. 2018a), JNK and p38 mitogen-activated protein kinases (MAPK) were strongly activated by cSBL treatment, suggesting that these proteins may be involved in cSBL-induced apoptosis. p38 inhibitors and knockdown of p38 attenuates cSBL-induced cell death (Kariya et al. 2016), indicating that RNA degradation by cSBL activates these stress kinases, which leads to apoptosis. However, similarly to inhibition of caspase activation by z-VAD, the inhibition of p38 MAPK did not completely rescue cSBL-induced cell death. Thus, it is possible that cSBL might be involved in a p38- and caspase-independent cell death-inducing pathway, or when the action of stress kinases and caspases is prevented, RNA breakdown by cSBL may lead to cell death via a different signal.
5.2 In Vivo Antitumor Activity of cSBL
The in vivo antitumor effect of cSBL was initially investigated using mouse cell lines. The survival of mice inoculated with cSBL-treated sarcoma-180 cells was prolonged compared to untreated controls (Nitta et al. 1994a). After inoculation with sarcoma-180 cells (1 × 106 cells/200 μL), the control group had 0% survival at 16 days, whereas the group treated with cSBL (1 mg) had 100% survival at 45 days. Intraperitoneal administration of cSBL to mice with sarcoma-180 and Mep II ascites cancer was also performed, and again, continuous administration of cSBL for 10 days achieved 100% survival within the experimental period (45 days) without any undesired side effects. Although there are only a few in vivo studies that evaluate human cells, Chen et al. investigated the inhibitory ability of cSBL on the growth of glioblastoma DBTRG cells in a mouse model (Chen et al. 2015). Single subcutaneous administration of cSBL on the opposite side of the lower abdominal region to the tumor significantly inhibited DBTRG cell growth compared to the control under conditions where no changes in body weight were observed. Furthermore, we recently established malignant mesothelioma xenografts with H2452 and MSTO cells (Tatsuta et al. 2018b). In in vivo studies with these grafts on the effect of cSBL, no obvious toxicities or body weight changes were observed during the experimental period; however, in both types of xenografts, significant tumor growth suppression was observed after twice-weekly cSBL (2.5 mg/kg) intratumoral injection for 4 weeks. The effect of cSBL was also superior to that of the current therapeutic agent, pemetrexed, administered by a previously reported method based on the maximum tolerated dose (Tonkinson et al. 1999; Kawabata et al. 2014). In H2452 xenografted groups, cSBL showed a tumor-suppressive effect earlier than in the pemetrexed-treated group, and antitumor effects of pemetrexed were not observed in the MSTO xenografts.
5.3 Cancer Selectivity
The antitumor effect of cSBL has been examined in several different cancer cell lines and primary or immortalized nonmalignant cells derived from normal tissues (Tatsuta et al. 2014c). In general, cSBL suppresses the growth of various types of cancer cells (carcinoma [cervical, hepatocellular, oral, and breast], sarcoma, glioma, mesothelioma, leukemia [T-cell, promyelotic, and erythro], and lymphoma), but not healthy cells (fibroblasts, melanocytes, keratinocytes, mesothelial cells, and mammary epithelial cells). The sensitivities of some cancer cell lines are contradictory between reports, but such contradictions may be caused by different experimental conditions. There is still no clear evidence for why cSBL is selective for cancer cells, but some hypotheses have been proposed. The binding of cSBL is selective to cancer cells and abolished by sialic acid–containing molecules, but not by monosialic acids (Nitta et al. 1987). Since changes in the glycans on cancer cell surfaces can occur during tumorigenesis (Dennis et al. 1999; Hakomori 2002), and the anion content (such as sialic acid) on cancer cells may increase (Wang et al. 2009), the particular sialic acid–containing structure recognized by cSBL may only exist on cancer cell membranes. It has also been shown that cSBL accumulates in the peripheral region of the nucleus after being taken up by cancer cells, although the detailed mechanism is not clear (Kariya et al. 2016). There may be differences in incorporation into cells, intracellular transportation, and/or localization between cancer cells and healthy cells. Furthermore, there are various factors that affect the selective cytotoxicity. As mentioned in Sect. 4.3, it has been suggested that differentiation status (Hu et al. 2001a; Wei et al. 2002) or JEV infection (Lee et al. 2011) are significant factors in the selective cytotoxicity. These may influence the effect of cSBL through changes such as the binding and uptake of cSBL. Tseng et al. suggested that the estrogen receptor is also an essential factor that enables cSBL cytotoxicity, since cSBL selectively induced cell death in estrogen-receptor-positive breast cancer cell lines (MCF7 and ZR-75-1), but not in estrogen-receptor-negative breast cancer cell lines (MDA-MB-231 and ZR-75-30) in their experiment. However, our recent comprehensive investigation focusing on breast cancer cell lines with different phenotypes, such as estrogen receptor-positive, progesterone receptor (PgR)-positive, human epidermal growth factor receptor type 2 (HER2)-positive, and triple-negative, revealed that cSBL induces apoptosis in all cancer cell lines tested, including estrogen-receptor-negative cells, even though it was less effective to healthy cells (Kariya et al. 2016; Tatsuta et al. 2018a). Therefore, we concluded that the estrogen receptor is not a significant factor in cSBL selectivity. Investigation of breast cancer revealed other aspects of the effect of cSBL on the expression of cancer-related proteins such as epidermal growth factor receptor (EGFR), which we discuss in Sect. 5.4.
5.4 Effect of cSBL in Cancer-Related Molecules
By investigating breast cancers, we found that cSBL decreases the expression levels of estrogen receptor, progesterone receptor (PgR), and human EGFR type 2 (HER2) (Tatsuta et al. 2018a). Furthermore, treatment with cSBL resulted in decreased protein levels of all ErbB proteins in each breast cancer cell line tested in the experiment (six cell lines), including EGFR in triple-negative MDA-MB231 and MDA-MB-468 cells. These results are interesting, because therapies targeting the hormone receptor or ErbB family could be novel efficient breast cancer treatments (Masoud and Pagès 2017). The cause of estrogen receptor, PgR, and ErbB family downregulation is currently under investigation. Degradation by proteasomes and/or stabilization by heat shock proteins (HSPs) is implicated in the turnover of these molecules. cSBL causes strong activation of p38, and it has also been reported to change the localization of HSP70 and HSC70 (Ogawa et al. 2014; Tatsuta et al. 2014a). We speculate that the reduced expression levels of estrogen receptor, PgR, and Erb B family in cSBL-treated cells may be associated with p38 activation and/or HSPs.
Most recently, we aimed to identify the key molecules whose expression was affected by cSBL to further understand the mechanism of action of cSBL using microarrays. Since cSBL degrades intracellular RNA, it is difficult to analyze gene expression in cSBL-treated cells. We exposed malignant mesothelioma H28 cells to low concentrations of cSBL for extended periods of time to obtain cloned cell lines with altered cell properties. Two clones were selected, and the RNA obtained from them was used for transcriptome analysis by microarray (Tatsuta et al. 2021). Microarray analysis detected 927 differentially expressed genes (DEGs). Bioinformatic analysis of these DEGs indicated that there were significant pleiotropic changes in the expression profiles of several genes, including multiple genes involved in metabolic pathways in cSBL-resistant cells. Interestingly, all DEGs involved in metabolic pathways were related to lipid and carbohydrate metabolism. At present, the significance of this change in carbohydrate and lipid metabolism is unknown, but these findings are novel insights into cSBL, and it may inform new methodologies of anticancer strategies using cSBL. In particular, glucose metabolism and the importance of the Warburg effect in cancer has been refocused on (Liberti and Locasale 2016). Regarding the DEGs, the expressions of some members of the aldo-keto reductase (AKR) family and the ATP-binding cassette (ABC) transporter superfamily were markedly downregulated. Among these, it was particularly interesting that cSBL action reduced the level of AKR1B10, which has been reported as a candidate biomarker for malignant pleural mesothelioma prognosis. These findings reveal novel aspects of the effect of cSBL, which may contribute to the development of new therapeutic strategies for malignant mesothelioma.
5.5 Synergistic Effects with Other Drugs
It has been reported that cSBL has a highly synergistic antitumor effect with other reagents. Combination treatment of cSBL with interferon γ (IFN-γ) enhances apoptosis in MCF-7 and SK-Hep-1 cells (Hue et al. 2001a; Tang et al. 2005). Hu et al. proposed that binding and entry of cSBL may be facilitated by IFN-γ, resulting in severe cell death. Tang et al. found that synergistic cytotoxicity with IFN-γ was not observed in HL-60 cells, and suggested that this is because of the differentiation status of HL-60. As cSBL is thought to be ineffective to the differentiated cells, differentiation by IFN-γ could make HL-60 cells insusceptible in that case.
Combination treatment of cSBL with TRAIL induced synergistic apoptosis. The synergistic effects were caused by the amplification of an apoptotic signal, which enhanced the truncation of Bid, a proapoptotic member of the Bcl-2 family, and caspase activation, resulting in drastic mitochondrial perturbations (Tatsuta et al. 2014b). Furthermore, the combined effect of cSBL was evaluated with conventional drugs for malignant mesothelioma, pemetrexed and cisplatin. The combination of cSBL and pemetrexed exhibited a strong synergistic effect that was comparable or even superior to the standard regimen of pemetrexed and cisplatin. The cytostatic effect of pemetrexed and the cytotoxic effect of cSBL cooperated without any repulsion, while the effects of pemetrexed and cisplatin on p21 expression were counteractive when used in combination (Satoh et al. 2017a). We further analyzed the combined effect of cSBL using a larger number of drugs, including EGFR-tyrosine kinase inhibitors. In a multiple drug combination study, pemetrexed + cSBL and pemetrexed+cisplatin+cSBL were more effective than the other regimens in terms of antiproliferative and synergistic effects. The pemetrexed + cSBL+TRAIL combination also showed strong synergy by playing additive roles without interference; thus, these combinations may serve as a rational regimen with antiproliferative mechanisms (Satoh et al. 2017b).
Additionally, as cSBL reduces the expression of EGFR, PgR, and estrogen receptors, combining cSBL with drugs that target the signaling of these receptors may provide favorable synergistic effects. Furthermore, since cSBL suppresses the expression of AKR1B10 and some ABC transporters, which are related to malignancy and resistance of cancers, multidrug combinations including cSBL may enhance the effects of other drugs and prevent drug resistance.
6 Conclusions and Future Problems
cSBL is a multifunctional protein with both lectin and ribonuclease activities, showing cancer cell-selective antitumor effects via a unique mechanism on various cancer cells, including multidrug-resistant cells. Furthermore, it has favorable properties as an anticancer drug, such as minimal immunogenicity and high stability in blood flow because of its compact structure, homology to human analogs, and resistance to heat, denaturants, and proteases. cSBL also has synergistic effects in combination with other drugs and has the potential to suppress the expression of molecules involved in cancer cell malignancy. A schematic illustration describing the antitumor effect of cSBL, including the signal induced by cSBL, and its valuable features for developing new therapeutic strategies are summarized in Fig. 2. The use of cSBL opens up new possibilities for treatment strategies in addition to currently limited chemotherapies. It is necessary in future research to further elucidate the mechanism of antitumor action, especially the factors of cancer cell selectivity of cSBL, as well as in vivo safety and pharmacokinetic studies for clinical application. Furthermore, as with other protein-based drugs, the development of stronger cSBL variants and cSBL fusion drugs using genetic modification techniques may lead to new therapies. We hope that a more detailed understanding of the antitumor mechanism of cSBL will contribute to the development of cancer therapeutics in the future.
Schematic illustration of antitumor effects of cSBL. cSBL somehow binds (or adsorbs) to the cell membrane. Once cSBL internalizes into cells, it localizes near the nucleus and degrades RNA. The RNA cleavage induces pleiotropic signals including apoptosis signaling. There are some favorable features of its antitumor activity, such as synergistic effect with other drugs and downregulation of cancer-related molecules
References
Ardelt W, Mikulski SM, Shogen K (1991) Amino acid sequence of an anti-tumor protein from Rana pipiens oocytes and early embryos. Homology to pancreatic ribonucleases. J Biol Chem 266:245–251. https://doi.org/10.1016/S0021-9258(18)52427-3
Beintema JJ, Fitch WM, Carsana A (1986) Molecular evolution of pancreatic-type ribonucleases. Mol Biol Evol 3:262–275. https://doi.org/10.1093/oxfordjournals.molbev.a040393
Beintema JJ, Breukelman HJ, Carsana A, Furiat A (1997) 8 - evolution of vertebrate ribonucleases: ribonuclease A superfamily. In: D’Alessio G, Riordan JF (eds) Ribonucleases. Academic Press, New York, pp 245–269
Boix E, Wu Y, Vasandani VM et al (1996) Role of the N terminus in RNase A homologues: differences in catalytic activity, ribonuclease inhibitor interaction and cytotoxicity. J Mol Biol 257:992–1007. https://doi.org/10.1006/jmbi.1996.0218
Bosch M, Benito A, Ribó M et al (2004) A nuclear localization sequence endows human pancreatic ribonuclease with cytotoxic activity. Biochemistry 43:2167–2177. https://doi.org/10.1021/bi035729+
Boyd WC, Shapleigh E (1954) Specific precipitating activity of plant agglutinins (lectins). Science 119:419. https://doi.org/10.1126/science.119.3091.419
Bracale A, Spalletti-Cernia D, Mastronicola M et al (2002) Essential stations in the intracellular pathway of cytotoxic bovine seminal ribonuclease. Biochem J 362:553–560. https://doi.org/10.1042/bj3620553
Castro J, Ribó M, Navarro S et al (2011) A human ribonuclease induces apoptosis associated with p21WAF1/CIP1 induction and JNK inactivation. BMC Cancer 11:9. https://doi.org/10.1186/1471-2407-11-9
Castro J, Ribó M, Puig T et al (2012) A cytotoxic ribonuclease reduces the expression level of P-glycoprotein in multidrug-resistant cell lines. Investig New Drugs 30:880–888. https://doi.org/10.1007/s10637-011-9636-2
Castro J, Tornillo G, Ceada G et al (2021) A nuclear-directed ribonuclease variant targets cancer stem cells and inhibits migration and invasion of breast cancer cells. Cancers (Basel) 13(17):4350. https://doi.org/10.3390/cancers13174350
Chen JN, Yiang GT, Lin YF et al (2015) Rana catesbeiana ribonuclease induces cell apoptosis via the caspase-9/−3 signaling pathway in human glioblastoma DBTRG, GBM8901 and GBM8401 cell lines. Oncol Lett 9:2471–2476. https://doi.org/10.3892/ol.2015.3117
Cho S, Beintema JJ, Zhang J (2005) The ribonuclease A superfamily of mammals and birds: identifying new members and tracing evolutionary histories. Genomics 85:208–220. https://doi.org/10.1016/j.ygeno.2004.10.008
Clement F, Pramod SN, Venkatesh YP (2010) Identity of the immunomodulatory proteins from garlic (Allium sativum) with the major garlic lectins or agglutinins. Int Immunopharmacol 10:316–324. https://doi.org/10.1016/j.intimp.2009.12.002
Costa AF, Campos D, Reis CA, Gomes C (2020) Targeting glycosylation: a new road for cancer drug discovery. Trends Cancer 6:757–766. https://doi.org/10.1016/j.trecan.2020.04.002
Costanzi J, Sidransky D, Navon A, Goldsweig H (2005) Ribonucleases as a novel pro-apoptotic anticancer strategy: review of the preclinical and clinical data for ranpirnase. Cancer Investig 23:643–650. https://doi.org/10.1080/07357900500283143
D’Alessio G (2011) The superfamily of vertebrate-secreted ribonucleases. In: Ribonucleases. Springer, Berlin, pp 1–34
D’Alessio G, Riordan JF (1997) Ribonucleases: structures and functions. Elsevier Science
D’Alessio G, Floridi A, De Prisco R et al (1972a) Bull semen ribonucleases. 1. Purification and physico-chemical properties of the major component. Eur J Biochem 26:153–161. https://doi.org/10.1111/j.1432-1033.1972.tb01751.x
D’Alessio G, Parente A, Guida C, Leone E (1972b) Dimeric structure of seminal ribonuclease. FEBS Lett 27:285–288. https://doi.org/10.1016/0014-5793(72)80642-2
de Lamirande G (1961) Action of deoxyribonuclease and ribonuclease on the growth of Ehrlich ascites carcinoma in mice. Nature 192:52–54. https://doi.org/10.1038/192052a0
Dennis JW, Granovsky M, Warren CE (1999) Glycoprotein glycosylation and cancer progression. Biochim Biophys Acta 1473:21–34. https://doi.org/10.1016/s0304-4165(99)00167-1
Deshpande RA, Shankar V (2002) Ribonucleases from T2 family. Crit Rev Microbiol 28:79–122. https://doi.org/10.1080/1040-840291046704
Dias RD, Machado LDS, Migliolo L, Franco OL (2015) Insights into animal and plant lectins with antimicrobial activities. Molecules 20:519–541. https://doi.org/10.3390/molecules20010519
Dickson KA, Haigis MC, Raines RT (2005) Ribonuclease inhibitor: structure and function. Prog Nucleic Acid Res Mol Biol 80:349–374. https://doi.org/10.1016/S0079-6603(05)80009-1
Dostál J, Matoušek J (1972) Purification of aspermatogenic substance in bull seminal vesicle fluid. Reproduction 31:273–274. https://doi.org/10.1530/jrf.0.0310273
Fang EF, Ng TB (2011) Ribonucleases of different origins with a wide spectrum of medicinal applications. Biochim Biophys Acta Rev Cancer 1815:65–74. https://doi.org/10.1016/j.bbcan.2010.09.001
Fiorini C, Gotte G, Donnarumma F et al (2014) Bovine seminal ribonuclease triggers Beclin1-mediated autophagic cell death in pancreatic cancer cells. Biochim Biophys Acta, Mol Cell Res 1843:976–984. https://doi.org/10.1016/j.bbamcr.2014.01.025
Gaur D, Swaminathan S, Batra JK (2001) Interaction of human pancreatic ribonuclease with human ribonuclease inhibitor: generation of inhibitor-resistant cytotoxic variants. J Biol Chem 276:24978–24984. https://doi.org/10.1074/jbc.M102440200
Goldstein I, Hughes R, Monsigny M et al (1980) What should be called a lectin? Nature 285:66. https://doi.org/10.1038/285066b0
Hakomori S (2002) Glycosylation defining cancer malignancy: new wine in an old bottle. Proc Natl Acad Sci 99:10231–10233. https://doi.org/10.1073/pnas.172380699
Hirabayashi J, Yamada M, Kuno A, Tateno H (2013) Lectin microarrays: concept, principle and applications. Chem Soc Rev 42:4443–4458. https://doi.org/10.1039/c3cs35419a
Hosokawa S, Irie M (1971) Purification and properties of seminal vesicle ribonucleases. J Biochem 69:683–697. https://doi.org/10.1093/oxfordjournals.jbchem.a129518
Hu C-CACA, Lee Y-HH, Tang C-HAHA et al (2001a) Synergistic cytotoxicity of Rana catesbeiana ribonuclease and IFN-γ, on hepatoma cells. Biochem Biophys Res Commun 280:1229–1236. https://doi.org/10.1006/bbrc.2001.4272
Hu CCA, Tang CHA, Wang JJ (2001b) Caspase activation in response to cytotoxic Rana catesbeiana ribonuclease in MCF-7 cells. FEBS Lett 503:65–68. https://doi.org/10.1016/S0014-5793(01)02691-6
Iordanov MS, Ryabinina OP, Wong J et al (2000) Molecular determinants of apoptosis induced by the cytotoxic ribonuclease onconase: evidence for cytotoxic mechanisms different from inhibition of protein synthesis. Cancer Res 60:1983–1994
Irie M, Nitta K, Nonaka T (1998) Biochemistry of frog ribonucleases. Cell Mol Life Sci 54:775–784. https://doi.org/10.1007/s000180050206
Kamiya Y, Oyama F, Oyama R et al (1990) Amino acid sequence of a lectin from japanese frog (Rana japonica) eggs. J Biochem 108:139–143. https://doi.org/10.1093/oxfordjournals.jbchem.a123153
Kariya Y, Tatsuta T, Sugawara S et al (2016) RNase activity of sialic acid-binding lectin from bullfrog eggs drives antitumor effect via the activation of p38 MAPK to caspase-3/7 signaling pathway in human breast cancer cells. Int J Oncol 49:1334–1342. https://doi.org/10.3892/ijo.2016.3656
Kawabata S, Chiang C-T, Tsurutani J et al (2014) Rapamycin downregulates thymidylate synthase and potentiates the activity of pemetrexed in non-small cell lung cancer. Oncotarget 5:1062–1070. https://doi.org/10.18632/oncotarget.1760
Kawauchi H, Sakakibara F, Watanabe K (1975) Agglutinins of frog eggs: a new class of proteins causing preferential agglutination of tumor cells. Experientia 31:364–365. https://doi.org/10.1007/BF01922588
Kobe B, Deisenhofer J (1995) A structural basis of the interactions between leucine-rich repeats and protein ligands. Nature 374:183–186. https://doi.org/10.1038/374183a0
Kobe B, Deisenhofer J (1996) Mechanism of ribonuclease inhibition by ribonuclease inhibitor protein based on the crystal structure of its complex with ribonuclease A. J Mol Biol 264:1028–1043. https://doi.org/10.1006/jmbi.1996.0694
Koczera P, Martin L, Marx G, Schuerholz T (2016) The ribonuclease a superfamily in humans: canonical RNases as the buttress of innate immunity. Int J Mol Sci 17:1278. https://doi.org/10.3390/ijms17081278
Kotchetkov R, Cinatl J, Krivtchik AA et al (2001) Selective activity of BS-RNase against anaplastic thyroid cancer. Anticancer Res 21:1035–1042
LeDoux L (1955) Action of ribonuclease on certain ascites tumours. Nature 175:258–259. https://doi.org/10.1038/175258b0
Lee FS, Vallee BL (1993) Structure and action of mammalian ribonuclease (angiogenin) inhibitor. Prog Nucleic Acid Res Mol Biol 44:1–30. https://doi.org/10.1016/s0079-6603(08)60215-9
Lee YH, Wei CW, Wang JJ, Chiou CT (2011) Rana catesbeiana ribonuclease inhibits Japanese encephalitis virus (JEV) replication and enhances apoptosis of JEV-infected BHK-21 cells. Antivir Res 89:193–198. https://doi.org/10.1016/j.antiviral.2011.01.002
Leich F, Stöhr N, Rietz A et al (2007) Endocytotic internalization as a crucial factor for the cytotoxicity of ribonucleases. J Biol Chem 282:27640–27646. https://doi.org/10.1074/jbc.M702240200
Liao YD (1992) A pyrimidine-guanine sequence-specific ribonuclease from Rana catesbeiana (bullfrog) oocytes. Nucleic Acids Res 20:1371–1377. https://doi.org/10.1093/nar/20.6.1371
Liao Y-D, Huang H-C, Chan H-J, Kuo S-J (1996) Large-scale preparation of a ribonuclease from Rana catesbeiana (bullfrog) oocytes and characterization of its specific cytotoxic activity against tumor cells. Protein Expr Purif 7:194–202. https://doi.org/10.1006/prep.1996.0027
Liberti MV, Locasale JW (2016) The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci 41:211–218. https://doi.org/10.1016/j.tibs.2015.12.001
Mäkelä O (1957) Studies in hemagglutinins of leguminosae seeds. Ann Med Exp Biol Fenn 35:1–133
Marth JD, Grewal PK (2008) Mammalian glycosylation in immunity. Nat Rev Immunol 8:874–887. https://doi.org/10.1038/nri2417
Masoud V, Pagès G (2017) Targeted therapies in breast cancer: new challenges to fight against resistance. World J Clin Oncol 8:120–134. https://doi.org/10.5306/wjco.v8.i2.120
Mastronicola MR, Piccoli R, D’Alessio G (1995) Key extracellular and intracellular steps in the antitumor action of seminal ribonuclease. Eur J Biochem 230:242–249. https://doi.org/10.1111/j.1432-1033.1995.tb20557.x
Mereiter S, Balmaña M, Campos D et al (2019) Glycosylation in the era of cancer-targeted therapy: where are we heading? Cancer Cell 36:6–16. https://doi.org/10.1016/j.ccell.2019.06.006
Mikulski S, Chun H, Mittelman A et al (1995) Relationship between response rate and median survival in patients with advanced nonsmall cell lung-cancer - comparison of onconase(r) with other anticancer agents. Int J Oncol 6:889–897. https://doi.org/10.3892/ijo.6.4.889
Murthy BS, Sirdeshmukh R (1992) Sensitivity of monomeric and dimeric forms of bovine seminal ribonuclease to human placental ribonuclease inhibitor. Biochem J 281(2):343–348. https://doi.org/10.1042/bj2810343
Nitta K (2001) Leczyme. Methods Enzymol 341:368–374
Nitta K, Takayanagi G, Kawauchi H, Hakomori S (1987) Isolation and characterization of Rana catesbeiana lectin and demonstration of the lectin-binding glycoprotein of rodent and human tumor cell membranes. Cancer Res 47:4877–4883
Nitta K, Oyama F, Oyama R et al (1993) Ribonuclease activity of sialic acid-binding lectin from Rana catesbeiana eggs. Glycobiology 3:37–45. https://doi.org/10.1093/glycob/3.1.37
Nitta K, Ozaki K, Ishikawa M et al (1994a) Inhibition of cell proliferation by Rana catesbeiana and Rana japonica lectins belonging to the ribonuclease superfamily. Cancer Res 54:920–927
Nitta K, Ozaki K, Tsukamoto Y et al (1994b) Characterization of a Rana catesbeiana lectin-resistant mutant of leukemia P388 cells. Cancer Res 54:928–934. https://doi.org/10.3892/ijo.9.1.19
Nitta K, Ozaki K, Tsukamoto Y et al (1996) Catalytic lectin (leczyme) from bullfrog (Rana catesbeiana) eggs. Int J Oncol 9:19–23. https://doi.org/10.3892/ijo.9.1.19
Notomista E, Catanzano F, Graziano G et al (2000) Onconase: an unusually stable protein. Biochemistry 39:8711–8718. https://doi.org/10.1021/bi000415x
Ogawa Y, Sugawara S, Tatsuta T et al (2014) Sialyl-glycoconjugates in cholesterol-rich microdomains of P388 cells are the triggers for apoptosis induced by Rana catesbeiana oocyte ribonuclease. Glycoconj J 31:171–184. https://doi.org/10.1007/s10719-013-9513-7
Okabe Y, Katayama N, Iwama M et al (1991) Comparative base specificity, stability, and lectin activity of two lectins from eggs of Rana catesbeiana and R. japonica and liver ribonuclease from R. catesbeiana. J Biochem 109:786–790. https://doi.org/10.1093/oxfordjournals.jbchem.a123457
Poucková P, Soucek J, Jelínek J et al (1998) Antitumor action of bovine seminal ribonuclease. Cytostatic effect on human melanoma and mouse seminoma. Neoplasma 45:30–34
Raines RT (1998) Ribonuclease A. Chem Rev 98:1045–1065. https://doi.org/10.1021/cr960427h
Reily C, Stewart TJ, Renfrow MB, Novak J (2019) Glycosylation in health and disease. Nat Rev Nephrol 15:346–366. https://doi.org/10.1038/s41581-019-0129-4
Robert C (2020) A decade of immune-checkpoint inhibitors in cancer therapy. Nat Commun 11:10–12. https://doi.org/10.1038/s41467-020-17670-y
Rodrigues JG, Balmaña M, Macedo JA et al (2018) Glycosylation in cancer: selected roles in tumour progression, immune modulation and metastasis. Cell Immunol 333:46–57. https://doi.org/10.1016/j.cellimm.2018.03.007
Rodríguez M, Benito A, Tubert P et al (2006) A cytotoxic ribonuclease variant with a discontinuous nuclear localization signal constituted by basic residues scattered over three areas of the molecule. J Mol Biol 360:548–557. https://doi.org/10.1016/j.jmb.2006.05.048
Rosenberg HF, Zhang J, Di LY, Dyer KD (2001) Rapid diversification of RNase A superfamily ribonucleases from the bullfrog, Rana catesbeiana. J Mol Evol 53:31–38. https://doi.org/10.1007/s002390010188
Roth JS (1963) Ribonuclease activity and cancer: a review. Cancer Res 23:657–666
Rutkoski TJ, Raines RT (2008) Evasion of ribonuclease inhibitor as a determinant of ribonuclease cytotoxicity. Curr Pharm Biotechnol 9:185–199. https://doi.org/10.2174/138920108784567344
Sakakibara F, Takayanagi G, Ise H, Kawauchi H (1977) Isolation of two agglutinins with different biological properties from the eggs of Rana catesbiana (author’s transl). Yakugaku Zasshi 97:855–862. https://doi.org/10.1248/yakushi1947.97.8_855
Satoh T, Tatsuta T, Sugawara S et al (2017a) Synergistic anti-tumor effect of bullfrog sialic acid-binding lectin and pemetrexed in malignant mesothelioma. Oncotarget 8:42466–42477. https://doi.org/10.18632/oncotarget.17198
Satoh T, Tatsuta T, Sugawara S et al (2017b) Combinatorial treatment using Leczyme as a new candidate of anti-cancer drug for malignant mesothelioma cell lines. 東北医科薬科大学研究誌 = J Tohoku Med Pharm Univ 64:65–76
Saxena SKK, Rybak SMM, Winkler G et al (1991) Comparison of RNases and toxins upon injection into xenopus oocytes. J Biol Chem 266:21208–21214. https://doi.org/10.1016/s0021-9258(18)54842-0
Section C, Ledoux L, Section C (1955) Action of ribonuclease on two solid tumours in vivo. Nature 176:36–37. https://doi.org/10.1038/176036a0
Sharon N, Lis H (1972) Lectins: cell-agglutinating and sugar-specific proteins. Science 177:949–959. https://doi.org/10.1126/science.177.4053.949
Sharon N, Lis H (2004) History of lectins: from hemagglutinins to biological recognition molecules. Glycobiology 14:53–62. https://doi.org/10.1093/glycob/cwh122
Shimomura O, Oda T, Tateno H et al (2018) A novel therapeutic strategy for pancreatic cancer: targeting cell surface glycan using rBC2LC-N lectin-drug conjugate (LDC). Mol Cancer Ther 17:183–195. https://doi.org/10.1158/1535-7163.MCT-17-0232
Smolewski P, Kowalik M, Witkowska M, Smolewski P (2020) Ribonucleases and immunoribonucleases as potential modalities of anticancer therapy. Drug Drug Abus:1–8. https://doi.org/10.31487/j.dda.2020.01.04
Sorrentino S, Libonati M (1994) Human pancreatic-type and nonpancreatic-type ribonucleases: a direct side-by-side comparison of their catalytic properties. Arch Biochem Biophys 312:340–348. https://doi.org/10.1006/abbi.1994.1318
Sumner JB, Howell SF (1936) Identification of hemagglutinin of Jack Bean with concanavalin A. J Bacteriol 32:227–237. https://doi.org/10.1128/jb.32.2.227-237.1936
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Tang C-HA, Hu C-CA, Wei C-W, Wang J-J (2005) Synergism of Rana catesbeiana ribonuclease and IFN-gamma triggers distinct death machineries in different human cancer cells. FEBS Lett 579:265–270. https://doi.org/10.1016/j.febslet.2004.11.086
Tatsuta T, Hosono M, Miura Y et al (2013a) Involvement of ER stress in apoptosis induced by sialic acid-binding lectin (leczyme) from bullfrog eggs. Int J Oncol 43:1799–1808. https://doi.org/10.3892/ijo.2013.2128
Tatsuta T, Hosono M, Sugawara S et al (2013b) Sialic acid-binding lectin (leczyme) induces caspase-dependent apoptosis-mediated mitochondrial perturbation in Jurkat cells. Int J Oncol 43:1402–1412. https://doi.org/10.3892/ijo.2013.2092
Tatsuta T, Hosono M, Ogawa Y et al (2014a) Downregulation of Hsp70 inhibits apoptosis induced by sialic acid-binding lectin (leczyme). Oncol Rep 31:13–18. https://doi.org/10.3892/or.2013.2814
Tatsuta T, Hosono M, Takahashi K et al (2014b) Sialic acid-binding lectin (leczyme) induces apoptosis to malignant mesothelioma and exerts synergistic antitumor effects with TRAIL. Int J Oncol 44:377–384. https://doi.org/10.3892/ijo.2013.2192
Tatsuta T, Sugawara S, Takahashi K et al (2014c) Cancer-selective induction of apoptosis by leczyme. Front Oncologia 4:1–6. https://doi.org/10.3389/fonc.2014.00139
Tatsuta T, Sato S, Sato T et al (2018a) Sialic acid-binding lectin from bullfrog eggs exhibits an anti-tumor effect against breast cancer cells including triple-negative phenotype cells. Molecules 23(10):2714. https://doi.org/10.3390/molecules23102714
Tatsuta T, Satoh T, Sugawara S et al (2018b) Sialic acid-binding lectin from bullfrog eggs inhibits human malignant mesothelioma cell growth in vitro and in vivo. PLoS One 13:1–15. https://doi.org/10.1371/journal.pone.0190653
Tatsuta T, Nakasato A, Sugawara S, Hosono M (2021) Transcriptomic alterations in malignant pleural mesothelioma cells in response to long-term treatment with bullfrog sialic acid-binding lectin. Mol Med Rep 23(6):467. https://doi.org/10.3892/mmr.2021.12106
Tikhonov A, Smoldovskaya O, Feyzkhanova G et al (2020) Glycan-specific antibodies as potential cancer biomarkers: a focus on microarray applications. Clin Chem Lab Med 58:1611–1622. https://doi.org/10.1515/cclm-2019-1161
Titani K, Takio K, Kuwada M et al (1987) Amino acid sequence of sialic acid binding lectin from frog (Rana catesbeiana) eggs. Biochemistry 26:2189–2194. https://doi.org/10.1021/bi00382a018
Tonkinson JL, Worzalla JF, Teng CH, Mendelsohn LG (1999) Cell cycle modulation by a multitargeted antifolate, LY231514, increases the cytotoxicity and antitumor activity of gemcitabine in HT29 colon carcinoma. Cancer Res 59:3671–3676
Tubert P, Rodríguez M, Ribó M et al (2011) The nuclear transport capacity of a human-pancreatic ribonuclease variant is critical for its cytotoxicity. Investig New Drugs 29:811–817. https://doi.org/10.1007/s10637-010-9426-2
Van Breedam W, Pöhlmann S, Favoreel HW et al (2014) Bitter-sweet symphony: glycan-lectin interactions in virus biology. FEMS Microbiol Rev 38:598–632. https://doi.org/10.1111/1574-6976.12052
Varki A (2017) Biological roles of glycans. Glycobiology 27:3–49. https://doi.org/10.1093/glycob/cww086
Vert A, Castro J, Ruiz-Martínez S et al (2012) Generation of new cytotoxic human ribonuclease variants directed to the nucleus. Mol Pharm 9:2894–2902. https://doi.org/10.1021/mp300217b
Vert A, Castro J, Ribó M et al (2016) A nuclear-directed human pancreatic ribonuclease (PE5) targets the metabolic phenotype of cancer cells. Oncotarget 7:18309–18324. https://doi.org/10.18632/oncotarget.7579
Vicentini AM, Kieffer B, Matthies R et al (1990) Protein chemical and kinetic characterization of recombinant porcine ribonuclease inhibitor expressed in Saccharomyces cerevisiae. Biochemistry 29:8827–8834. https://doi.org/10.1021/bi00489a046
Vogelzang NJ, Stadler WM (1999) Gemcitabine and other new chemotherapeutic agents for the treatment of metastatic bladder cancer. Urology 53:243–250. https://doi.org/10.1016/s0090-4295(98)00501-9
Wang F-L, Cui S-X, Sun L-P et al (2009) High expression of alpha 2, 3-linked sialic acid residues is associated with the metastatic potential of human gastric cancer. Cancer Detect Prev 32:437–443. https://doi.org/10.1016/j.cdp.2009.01.001
Wei CWW, Hu CCACA, Tang CHAHA et al (2002) Induction of differentiation rescues HL-60 cells from Rana catesbeiana ribonuclease-induced cell death. FEBS Lett 531:421–426. https://doi.org/10.1016/S0014-5793(02)03577-9
WHO (WHO) (2020) Global Health estimates 2020: deaths by cause, age, sex, by country and by region, 2000–2019. World Health Organization, Geneva
Wu Y, Mikulski SM, Ardelt W et al (1993) A cytotoxic ribonuclease: study of the mechanism of onconase cytotoxicity. J Biol Chem 268:10686–10693. https://doi.org/10.1016/s0021-9258(18)82252-9
Yamamoto S, Tomiyama M, Nemoto R et al (2013) Effects of food lectins on the transport system of human intestinal CaCO-2 cell monolayers. Biosci Biotechnol Biochem 77:1917–1924. https://doi.org/10.1271/bbb.130367
Yokota M, Sakakibara H, Kawauchi H (1975) Separation of agglutinin from eggs of Rana nigromaculata nigromaculata Hallowell and its interaction with ascites hepatoma (author’s transl). Yakugaku Zasshi 95:50–55. https://doi.org/10.1248/yakushi1947.95.1_50
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Tatsuta, T., Hosono, M. (2023). Effects of Bullfrog Sialic Acid–Binding Lectin in Cancer Cells. In: Furukawa, K., Fukuda, M. (eds) Glycosignals in Cancer. Springer, Singapore. https://doi.org/10.1007/978-981-19-7732-9_7
Download citation
DOI: https://doi.org/10.1007/978-981-19-7732-9_7
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-7731-2
Online ISBN: 978-981-19-7732-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)