Abstract
SBL/RC-RNase was originally isolated from frog (Rana catesbeiana) oocytes and purified as a novel sialic acid-binding lectin (SBL) that displayed strong anti-cancer activity. SBL was later shown to be identical to a ribonuclease (RC-RNase) from oocytes of the same species. The administration of SBL/RC-RNase induced apoptosis (with nuclear condensation and DNA fragmentation) in mouse leukemia P388 cells but did not kill umbilical vein endothelial or fibroblast cells derived from normal tissues. The cytotoxic activity of SBL/RC-RNase was inhibited by desialylation of P388 cells and/or the co-presence of free bovine submaxillary mucin. FACS analysis showed that SBL/RC-RNase was incorporated into cells after attachment to cholesterol-rich microdomains. Addition of the cholesterol remover methyl-β-cyclodextrin reduced SBL/RC-RNase-induced apoptosis. Apoptosis occurred through the caspase-3 pathway following activation of caspase-8 by SBL/RC-RNase. A heat shock cognate protein (Hsc70) and a heat shock protein (Hsp70) (each 70 kDa) on the cell membrane were shown to bind to SBL/RC-RNase by mass spectrometric and flow cytometric analyses. Quercetin, an inhibitor of Hsc70 and Hsp70, significantly reduced SBL/RC-RNase-induced apoptosis. Taken together, our findings suggest that sialyl-glycoconjugates present in cholesterol-rich microdomains form complexes with Hsc70 or Hsp70 that act as triggers for SBL/RC-RNase to induce apoptosis through a pathway involving the activation of caspase-3 and caspase-8.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ribonucleases (RNases) from oocytes of the amphibian (frog) genus Rana have unique cytotoxic activities and are being investigated as innovative drugs in ongoing clinical trials. Studies along this line were initiated by the discovery in 1987 of a sialic acid-binding lectin (SBL) purified from Rana catesbeiana (American bullfrog) oocytes that displayed strong anti-proliferative activity against cancer cells [1]. SBL is a basic protein that consists of 111 amino acids (levels of Arg and Lys are high) with a pyroglutamyl residue at the N-terminus [2]. The primary structure of SBL did not show strong homology to those of 3,450 known proteins in a database comparison. SBL agglutinated a variety of tumor cells (but not normal lymphocytes, fibroblasts, or erythrocytes), and such agglutination was inhibited by the co-presence of highly sialylated acidic glycoproteins [1].
An RNase with a primary structure similar to that of SBL was subsequently isolated from R. catesbeiana liver [3]. SBL was shown to be a member of a RNase superfamily and to possess both enzymatic and sialic acid-binding activities [4–7]. An RNase isolated from oocytes of the Northern leopard frog (Rana pipiens), termed “onconase” or “ranpirnase” [8], was found to display apoptotic activity and 50 % primary structural homology with SBL [9]. An RNase isolated from R. catesbeiana oocytes (termed “RC-RNase”) showed cytotoxic properties similar to those of onconase [10–12]. The primary structures of SBL and RC-RNase were found to be identical, and the term “SBL/RC-RNase” is used to distinguish this structure from those isolated from R. catesbeiana liver.
The administration of onconase/ranpirnase was recently shown to down-regulate NFκB level and matrix metalloprotease-9 activity in malignant cells [13, 14]. SBL/RC-RNase was found to activate caspase-3 and down-regulate the levels of Bcl-2 and estrogen receptor genes in breast tumor cells [15, 16]. Three-dimensional structural analyses of SBL/RC-RNase and onconase helped explain the differences between their anticancer properties and that of pancreatic RNase A [17–20]. In addition to ongoing studies of the regulatory activities of amphibian oocyte RNases in cytosol, it is important to elucidate the target molecules that interact with RNases on the cell surface and the mechanisms whereby RNases are taken up by the cytosol of malignant cells. Such information will clarify the ability of these RNases to specifically kill cancer cells without harming normal cells.
Research to date on SBL/RC-RNase has been focused on specific aggregation of cancer cells and the inhibitory effects of the co-presence of sialomucins. Many types of signal transduction molecules within cells are enriched in glycoconjugates such as the gangliosides found in cholesterol-rich microdomains at the cell surface. In this study, we evaluated the importance of sialic acids and used specific inhibition assays to survey the macromolecules present in cholesterol-rich microdomain fractions that interact with SBL/RC-RNase. Mass spectrometric analyses indicated that heat shock cognate protein 70, heat shock protein 70, and ganglio-series gangliosides are potential trigger molecules for the uptake of SBL/RC-RNase and consequent induction of apoptosis.
Materials and methods
Cell lines and materials
Murine leukemia P388 cells were obtained from the Cell Resource Center of the Biomedical Research Institute of Development, Aging and Cancer, Tohoku University, Sendai, Japan. Human erythroleukemia K562 cells and promyelocytic leukemia HL-60 cells were from the Japanese Cancer Research Resources Bank, Tokyo, Japan. R. catesbeiana specimens were from Nippon Bio-Supply Center Co., Tokyo, Japan. Phenylmethylsulfonyl fluoride (PMSF) was from Wako Pure Chemical Co., Tokyo, Japan. Heparin-agarose gel was from Cosmo Bio Co., Tokyo, Japan. Sephadex G-75 was from GE Healthcare Japan, Tokyo, Japan. A standard protein marker mixture for sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE), 3,3’,5,5’-tetramethylbenzidine, and polyvinylidene difluoride (PVDF) membrane were from ATTO Co., Tokyo, Japan. Cell Counting Kit-8 including 2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium, monosodium salt (WST-8), and 3-[(3-cholamidopropyl)dimethylammonio]propanesulfonate (CHAPS) were from Dojindo Co., Kumamoto, Japan. RPMI 1640 medium was from Nissui Pharmaceutical Co., Tokyo, Japan. Fetal calf serum was from Life Technologies Co., Carlsbad, CA, USA. Penicillin-streptomycin was from Roche Diagnostics K. K., Tokyo, Japan. Trypan blue solution was from Nacalai Tesque, Inc., Kyoto, Japan. GloMax Multi Detection System was from Promega, Madison, WI, USA. UV absorbance plate reader, ImmunoMini NJ-2300, was from Biotec Co., Tokyo, Japan. Alamar Blue was from Kyowa Hakko Kogyo Co. Tokyo, Japan. Aphidicolin was from Wako Pure Chemical Industries, Osaka, Japan. Anti-HSP70 and anti-HSC70 antibodies were from Stressgen, Victoria, B.C., Canada. HiTrap protein G column was from GE Healthcare Japan. FACSCalibur was from Nippon Becton Dickinson Co., Tokyo, Japan.
Purification of SBL/RC-RNase and generation of polyclonal antibody
R. catesbeiana oocytes were collected by laparotomy. Lipids in the yolk were removed by filtration of a homogenate with cold acetone, and the precipitate was powdered by evaporation. Acetone-dried egg powder (50 g) was homogenized with 1 L of 150 mM sodium chloride and centrifuged at 9,000 × g for 30 min. The crude supernatant was saturated with ammonium sulfate to 40 % (w/v) concentration, centrifuged as above, dialyzed against distilled water, and lyophilized into powder. The powder (2 g) was dissolved with 10 mL of 1 mM phosphate-buffered saline (PBS) (pH 6.0), and the supernatant was collected by centrifugation at 27,500 × g for 1 h at 4 °C. SBL/RC-RNase was purified by successive isolations from the crude supernatant by application of DEAE-sepharose, heparin-sepharose, and hydroxyl apatite [1].
For the generation of polyclonal antibody against SBL/RC-RNase, 1 mg purified SBL/RC-RNase (0.5 mL) in PBS was mixed with 1 mg complete Freund’s adjuvant, and the emulsion was injected subdermally in the abdomen of Japanese white rabbits 3 times at monthly intervals. Blood containing polyclonal antibody was collected from the ear vein and applied to a HiTrap protein G column to isolate the IgG fraction.
Cytotoxicity and cell viability assays
P388 cells were maintained in RPMI 1640 medium supplemented with heat-inactivated fetal calf serum (10 %, v/v), penicillin (100 IU/mL), and streptomycin (100 μg/mL) at 37 °C in an atmosphere of 95 % air/5 % CO2. Cells (2 × 104, in 90 μL solution) were seeded into a 96-well flat-bottom plate and treated with various concentrations (0–50 μg/mL) of SBL/RC-RNase (10 μL) for 4 to 24 h. Cytotoxic activity and cell viability/growth were evaluated by trypan blue (0.5 % (w/v)) exclusion [21, 22] and WST-8 (10 μL) assays, respectively. Reductions in the proportion of living cells were assayed by measurement of absorbance at wavelength 450 nm (reference, 600 nm) using the GloMax Multi Detection System.
Fluorescence-activated cell sorting (FACS) analysis
Detection of SBL/RC-RNase on the surface or in the cytosol of P388 cells: Cells (2 × 105) were cultured in 6-well dishes with SBL/RC-RNase (1 μΜ) for 1 h at 4 °C, washed with PBS, and incubated for 30 min at 4 °C or 37 °C for detection of SBL/RC-RNase at the cell surface or cytosol, respectively. Cells were then treated with anti-SBL/RC-RNase polyclonal antibody (100 μL, 0.2 mg/mL in PBS) for 30 min at 4 °C, washed twice with PBS, and incubated with fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit secondary antibody at a dilution of 1:500 in PBS (100 μL) at 4 °C for 30 min. SBL/RC-RNase localization on the cell surface was detected by flow cytometry (FACSCalibur) with excitation wavelength 488 nm and emission wavelength 530 nm [23–25].
Detection of heat shock protein 70 and heat shock cognate protein 70 on the cell surface: Cells (2 × 105) were treated with anti-mouse heat shock protein 70 kDa (Hsp70) and anti-mouse heat shock cognate protein 70 kDa (Hsc70) mAbs at a dilution of 1:1000 in PBS (100 μL, 1 μg/mL) for 30 min at 4 °C, washed twice with PBS, and incubated with FITC-conjugated goat anti-rabbit IgG and goat anti-rat IgG antibodies (100 μL, 0.5 μg/mL in PBS) for 30 min at 4 °C. Cell surface expression of Hsp70 and Hsc70 was detected by flow cytometry (FACSCalibur).
Caspase activation and inhibition assays
Caspase activation assay: Cells (5 × 105) were cultured with SBL/RC-RNase (3 μM) for 24 h at 37 °C. The activation of caspase-3 and caspase-8 was evaluated using hydrolyzed artificial substrate CPP32 and FLICE colorimetric protease assay kits, respectively. Cells were extracted with Triton X-100 lysis buffer (25 mM Tris, pH 7.5, 150 mM NaCl, 5 mM EDTA, 1 % Triton X-100, 0.5 % sodium deoxycholate, 0.1 % SDS, 10 mM NaF), incubated for 10 min at 4 °C, and centrifuged at 10,000 × g for 1 min. The lysate was mixed with substrate buffer containing 10 mM dithiothreitol and 200 μM p-nitroanilide (pNA) for 2 h at 37 °C according to the manufacturer’s protocol. Caspase activation induced by SBL/RC-RNase was determined by measurement of absorbance at 405 nm.
Caspase inhibition assay: To confirm that caspase-3 and -8 were activated by administration of SBL/RC-RNase, their respective inhibitors conjugated with pNA (DEVD-pNA and IETD-pNA substrate; final concentration 200 μM) were co-incubated for 1 h at 37 °C, and the pNA amount was quantified by measurement of absorbance with an ImmunoMini NJ-2300 plate reader. For the inhibition assay, cells were preincubated with two other inhibitors of caspase-3 and -8, Z-Asp-Glu-Val-Asp-fluoromethylketone (Z-DEVD-fmk) and Z- Ile-Glu-Thr-Asp-fluoromethylketone (Z-IETD-fmk), respectively, for 1 h at 37 °C, followed by incubation with SBL/RC-RNase (3 μM) for 24 h. The activities of caspase-3 and -8 were quantified as described above.
Depletion of cholesterol-rich microdomains by methyl-β-D-cyclodextrin (MβCD) treatment
Cells (2 × 105) were grown in a 6-well plate for 24 h at 37 °C, and the medium was replaced by fresh medium with or without MβCD (3.1 or 12.5 mM). Cell surface cholesterol was completely depleted by treatment with MβCD for 1 h [26]. To determine whether cholesterol-rich microdomains were essential for induction of apoptosis, SBL/RC-RNase (3 μM) was incubated with MβCD-treated or -untreated cells for 24 h at 37 °C. Cell viability and caspase-3 activity were measured as described above.
DNA fragmentation
Cells (2 × 105) were cultured with SBL/RC-RNase (2–20 μM) for 24 h at 37 °C, harvested, and lysed by lysis buffer (20 μL Tris–HCl (50 mM, pH 7.8), 0.5 % (w/v) sodium-N-lauroylsarcosinate, 10 mM EDTA). RNase A and proteinase K (each 1 μL, 1 mg/mL) were added to the crude extract and incubated for 30 min at 50 °C. DNA was precipitated by treatment with isopropyl alcohol and ethanol, loaded onto a 1.8 % agarose gel, and subjected to electrophoresis. Fragmented DNA was visualized by staining with 0.002 % (w/v) ethidium bromide under UV illumination at 330 nm.
Detection of binding triggers of SBL/RC-RNase in low-density, detergent-insoluble membrane fractions
Cells (2 × 107) were harvested and centrifuged. Pelleted cells were suspended in 1 mL of 1 % Triton X-100 containing TNE buffer (10 mM Tris–HCl (pH 7.5), 150 mM NaCl, 5 mM EDTA) with 75 U aprotinin and 2 mM PMSF and lysed by standing for 30 min on ice. The lysed cells were Dounce homogenized with a tight-fitting piston for 10 strokes and centrifuged for 5 min at 3,000 × g to remove large cellular debris. The supernatant was subjected to sucrose density-gradient centrifugation [27], and 1 mL supernatant was mixed with 1 mL of 85 % sucrose (w/v) in TNE buffer. The resulting diluent (2 mL) was placed at the bottom of a 12-mL centrifuge tube and overlaid successively with 5.5 mL of 35 % sucrose in TNE and 5.5 mL of 5 % sucrose in TNE. The tubes were centrifuged using a Beckman SW40Ti rotor at 250,000 × g for 17 h at 4 °C. A series of 1-mL aliquots from top to bottom of the tube were collected.
A 500-μL sample from each tube, associated with low-density, detergent-insoluble membrane, was precipitated with 500 μL trichloroacetic acid, centrifuged at 16,000 × g for 30 min, and washed 3 times with 1 mL cold ethyl ether to eliminate trichloroacetic acid. The precipitate was dissolved in 10 μL sample buffer (62.5 mM Tris–HCl (pH 6.8) containing 10 % glycerol, 2 % SDS, 5 % mercaptoethanol, and 0.0025 % bromophenol blue). Proteins were separated by SDS-PAGE [28] and electrotransferred to a PVDF membrane [29]. The membrane was soaked in 2 % Triton X-100 and incubated with SBL/RC-RNase (40 μg/mL) for 1 h. Targets on the membrane were detected by incubation with anti-SBL/RC-RNase polyclonal antibody followed by HRP-conjugated goat anti-rat IgG (Santa Cruz Biotechnology, Santa Cruz, CA, USA). The trigger for SBL/RC-RNase was detected by exposure to X-ray film (Fuji Film Co., Tokyo, Japan) using an enhanced chemical luminescence western blotting detection kit (GE Healthcare Japan) [30].
The protein in the fifth tube, containing the low-density, detergent-insoluble membrane fraction, was subjected to SDS-PAGE. The gel at the position of the SBL/RC-RNase trigger as detected by the above assay was cut out and transferred into a plastic tube. The gel was dehydrated in 100 μL CH3CN for 10 min at 37 °C and dried by vacuum centrifugation for 5 min. The dried residue was successively extracted with (i) 50 % CH3CN and 0.1 % trifluoroacetic acid (with centrifugation); (ii) 15 % isopropyl alcohol/20 % formic acid/25 % CH3CN/40 % H2O; (iii) 80 % CH3CN. Each of these extracts was dried successively in a tube, and the residue was dissolved in 6 μL ultrapure water. Aliquots were used for protein identification by an API QSTAR pulsar hybrid mass spectrometer system connected to a micro-liquid chromatograph (Magic 2002, Michrom BioResource, Auburn, CA, USA) [31]. The micro-LC conditions were as follows: Magic C18 column (0.2 mm, inner diameter × 50 mm); elution with 0.1 % formic acid (solvent A) and 0.1 % formic acid in 90 % CH3CN (solvent B) using a program of 3 % solvent B for 2 min, gradient at 2.1 %/min for 45 min, 100 % solvent B for 5 min, flow rate 2.5 mL/min. The mass spectrometer system consisted of a nanoelectrospray ionization source and quadrupole time-of-flight MS. The mass accuracy was 0.1 mass unit. The MS conditions were as follows: ion spray voltage 3.0–3.8 kV, electron multiplier voltage 2,200 V for MS and MS/MS analyses; nitrogen 10 collision gas and collision energy 20–55 eV for MS/MS analysis. The micro-LC conditions were as follows: Magic C18 column (0.2 mm, inner diameter × 50 mm); elution with solvent A and solvent B using a program of 3 % solvent B for 2 min, gradient at 2.1 %/min for 45 min, 100 % solvent B for 5 min, flow rate 2.5 mL/min. The trigger protein for SBL/RC-RNase was identified by micro-LC/MS (QSTAR) using the PROWL (ProFound) search engine 2 and the NCBI database [32]. The major ion peaks of the total ion chromatogram were further analyzed to obtain the amino acid sequences of the tryptic peptides by LC/MS/MS using the Mascot search engine under the same conditions (QSTAR with Magic micro-liquid chromatograph) and the same database.
Statistical analysis
The results of experiments are presented as mean ± standard error (SE). Differences in means were evaluated by two-tailed Student’s t-test, with P values < 0.05 considered to be statistically significant.
Results
Antiproliferative effect of SBL/RC-RNase on leukemic cell lines
SBL/RC-RNase effectively reduced the viability of leukemic cell lines such as P388, K562, and HL60, but had no effect on the viability of human breast cancer MCF7 or Burkitt’s lymphoma Daudi or Raji. SBL/RC-RNase treatment did not kill normal human cells derived from epidermal fibroblasts (NHDF), epidermal melanocytes (NHEM), or keratinocytes (NHEK) (Fig. 1a). FACS analysis using anti-SBL/RC-RNase antibody showed that administered SBL/RC-RNase attached to the membranes of P388 and K562 cells but not to those of Raji or NHDF cells (Fig. 1b). SBL/RC-RNase was incorporated into the cytoplasm through binding to the cell surface (Fig. 1b, panel INC P388). These findings were consistent with those shown in Fig. 1a; i.e., SBL/RC-RNase bound to P388 and K562 cells and reduced cell viability, whereas SBL/RC-RNase did not bind to lymphoma Raji or normal tissue-derived NHDF and had no effect on the viability of these cells.
Cytotoxic activity and cell binding of SBL/RC-RNase. a Cell viability (%) (vertical axis) was quantified by WST-8 assay. Cancer cells: P388, K562, HL60 (leukemia), Raji, Daudi (Burkitt’s lymphoma), and MCF7 (breast cancer). Normal cells: epidermal fibroblasts (NHDF), epidermal melanocytes (NHEM), and keratinocytes (NHEK). SBL/RC-RNase (3 μM) was administered to each cell line. Error bars: SE calculated from three different cell preparations assayed individually. *, P < 0.01 in comparison with control (white bar). b Binding of SBL/RC-RNase to cells as in Panel a was analyzed by flow cytometry. Cells were treated with 3 μM SBL/RC-RNase and incubated with anti-SBL/RC-RNase polyclonal antibody and FITC-conjugated goat anti-rabbit IgG, and fluorescence was detected by flow cytometry (FACSCalibur). Panel INC P388: incorporation of SBL/RC-RNase into P388 cells (2 × 105) cultured with 3 μM SBL/RC-RNase for 1 h at 4 °C. The cells were washed with PBS, incubated for 1 h at 4 °C (dotted line) or 37 °C (solid line), treated with anti-SBL/RC-RNase antibody for 30 min at 4 °C, washed twice with PBS, and incubated with FITC-conjugated goat anti-rabbit secondary antibody at a dilution of 1:500 in PBS (100 μL) for 30 min at 4 °C. The peak positions of the dotted and solid lines indicate cells before and after treatment with SBL/RC-RNase (3 μM) for 30 min, respectively
Apoptotic effect of SBL/RC-RNase on P388 cells
Chromatin condensation of P388 cells was observed by staining with Hoechst 33342 after treatment with 3 μM SBL/RC-RNase for 24 h (Fig. 2a, panel SBL/RC-RNase). In addition to morphological changes of nuclei in P388 cells, DNA fragmentation as indicated by the appearance of ladders was induced in a time-dependent manner by SBL/RC-RNase administration (Fig. 2b, lanes 0 to 48). In P388 cells pretreated with sialidase, SBL/RC-RNase administration did not induce DNA fragmentation (Fig. 2b, lane Sia). SBL/RC-RNase bound to P388 and K562 cells and induced DNA fragmentation. In contrast, SBL/RC-RNase did not bind to Raji or NHDF cells and did not induce DNA degradation of these cells (Fig. 2c). Consideration of these findings together with those shown in Fig. 1a, and the fact that cell lines whose viability was reduced by SBL/RC-RNase were detected by WST-8 assay, indicate that both nuclear condensation and DNA fragmentation occurred. Reduction of cell viability is hereafter considered as equivalent to the degree of apoptosis induced by SBL/RC-RNase.
Induction of nuclear condensation and DNA fragmentation (ladder formation) by SBL/RC-RNase. a Nuclei of P388 cells treated (right panel) or not treated (control; left panel) with 3 μM SBL/RC-RNase. Cells were cultured for 24 h and observed by Hoechst33342 staining using fluorescence microscopy (excitation and emission wavelengths 350 and 461 nm). b Cells were treated with SBL/RC-RNase (3 μM) and cultured for 0–48 h (numbers at top). DNA ladders were detected by agarose-electrophoresis stained with ethidium bromide under UV light. Sia: sialidase-pretreated cells were incubated for 24 h with SBL/RC-RNase (3 μM). M: molecular marker for DNA fragment (100-bp DNA ladder). c DNA electrophoresis pattern extracted from SBL-RC-RNase-treated cells. Cancer cells (P388, K562, Raji) and normal cells (NHDF) were treated with SBL/RC-RNase (3 or 20 μM) for 24 h. C: control (not treated with SBL/RC-RNase)
Importance of sialyl-glycoconjugates for the cytotoxic effect of SBL/RC-RNase on P388 cells
The inhibitory effect of 3 μM SBL/RC-RNase on P388 cell viability (Fig. 3a, black bar) was enhanced ~40–50 % in the co-presence of bovine submaxillary mucin (BSM), heparin, or heparan sulfate (Fig. 3a, bars BSM, Hepa, and HepS), whereas asialo-BSM and sialic acid had no significant enhancing effect (Fig. 3a, bars asialo-BSM and Sia). These findings were consistent with those for inhibition of DNA fragmentation. BSM, heparin, and heparan sulfate inhibited DNA ladder formation induced by SBL/RC-RNase (Fig. 3b). SBL/RC-RNase administration reduced cell viability in a dose-dependent manner, whereas the viability of sialidase-treated cells (Fig. 3c, striped bars) was 15 % or 30 % higher than that of untreated cells (Fig. 3c, white bars). These findings suggest that SBL/RC-RNase bound to sialic acid-containing complex glycoconjugates on the cell membrane and that desialylation reduced the apoptotic effect of SBL/RC-RNase.
Inhibition of the cytotoxic effect of SBL/RC-RNase on P388 cells by co-treatment with free acidic glycoproteins or sialidase. a White bar: negative control (no SBL/RC-RNase treatment). Black bar: positive control (SBL/RC-RNase treatment without glycoproteins). Hepa: heparin; HepS: heparan sulfate; Sia: sialic acid. Saccharides and glycoproteins were co-incubated with 3 μM SBL/RC-RNase with 25 μg/mL glycoproteins (BSM, asialo-BSM) or 50 μg/mL glycans (Hep, HepS). Sialic acid was administered at a concentration of 50 μg/mL. Error bars: SE from three independent experiments. *, P < 0.01 from comparison with negative control. **, P < 0.01 from comparison with positive control. b Inhibition of SBL/RC-RNase-induced DNA fragmentation by the co-presence of sialyl-glycoproteins. Abbreviations as in panel a. c Recovery of cell viability against SBL/RC-RNase by pretreatment with sialidase. Shaded bars: pretreatment with 1 U sialidase prior to administration of SBL/RC-RNase (0–3 μM). White bars: no sialidase pretreatment. Error bars: SE from three different cell preparations assayed individually. *, P < 0.05
Cholesterol-rich microdomains contain apoptotic ligands of SBL/RC-RNase that activate caspase-3
The inhibitory effect of SBL/RC-RNase on viability of P388 cells was significantly reduced by treatment with the cholesterol inhibitor MβCD (40 % reduction for 3.1 μg/mL MβCD; 50 % reduction for 12.5 μg/mL MβCD) (Fig. 4a, shaded bars). MβCD disrupts the construction within the cell membrane of cholesterol-rich microdomains, which contain many types of glycosphingolipids (GSLs) and glycoproteins. The above findings suggest that desialylation of cells reduces the cytotoxicity of SBL/RC-RNase because SBL/RC-RNase binds to glycoconjugates located in cholesterol-rich microdomains that are involved in apoptotic cascade signaling within the cell. SBL/RC-RNase administration activated caspase-3, an effector of apoptosis (Fig. 4b, black bar), and such activation was inhibited in a dose-dependent manner by the co-presence of MβCD (Fig. 4b, gray bars). MβCD in the absence of SBL/RC-RNase had no effect on caspase-3 activity (Fig. 4b, white bars).
Inhibition of the cytotoxic effect of SBL/RC-RNase on P388 cells by pretreatment with MβCD. a Cell viability (%) (vertical axis) was quantified by trypan blue exclusion assay. White bars: viability of cells nont treated with SBL/RC-RNase and treated or not with MβCD. Black bar: SBL/RC-RNase alone (control). Shaded bars: cells pretreated with MβCD (3.1 or 12.5 μg/mL) and then treated with SBL/RC-RNase. b Caspase-3 activity was measured by absorbance at 405 nm. Black bar: cells treated with SBL/RC-RNase (3 μM) but not MβCD (positive control). Gray bars: cells pretreated with MβCD (3.1 or 12.5 μg/mL) and then treated with SBL/RC-RNase (3 μM). White bars: caspase-3 activity of cells not treated with SBL/RC-RNase (negative controls). Error bars: SE from three different cell preparations assayed individually. *, P < 0.01 from comparison with control (black bar)
These findings indicate that apoptotic-trigger molecules for SBL/RC-RNase are located in cholesterol-rich microdomains on the P388 cell surface and are essential for induction of caspase-3 activity by SBL/RC-RNase.
Activation of caspase-8 by SBL/RC-RNase is suppressed by removal of sialic acids
Inhibition assays showed that caspase-8 as an initiator of caspase-3 was activated in a time-dependent manner by SBL/RC-RNase administration (Fig. 5a). Levels of activated caspase-8 increased during 3 h following SBL/RC-RNase administration, and the removal of sialic acids from the cell surface reduced the activation (Fig. 5c). The addition of the caspase-8 inhibitor Z-IETD-fmk increased cell viability following SBL/RC-RNase administration (Fig. 5b, gray bars) and reduced the activity of caspase-3 as an effector of apoptosis (Fig. 5b, bold line). Taken together, these findings suggest that SBL/RC-RNase causes apoptosis through a caspase-8 to caspase-3 pathway.
Activation of caspases was induced by SBL/RC-RNase administration. a Time course of caspase-8 activity induced by administration of 3 μM SBL/RC-RNase. b Co-inhibition of caspase-3 and SBL/RC-RNase-induced cytotoxicity by administration of caspase-8 inhibitor Z-IETD-fmk. Caspase-8 and -3 activities were measured as described in Materials and methods, “Caspase activation and inhibition assays” section. White bars: viability of cells not treated with Z-IETD-fmk or SBL/RC-RNase. Black bars: cells treated with SBL/RC-RNase only. Gray bars: cells co-treated with SBL/RC-RNase (3 μM) and Z-IETD-fmk (10–40 μM). *, P < 0.01 from comparison with control (black bar). c Inhibition of SBL/RC-RNase-induced caspase-8 activation by pretreatment with sialidase. Cells were pretreated or not with sialidase (1 U), then treated with SBL/RC-RNase, and caspase-8 activity was measured as described above. *, P < 0.01 from comparison with negative control. **, P < 0.01 from comparison with positive control. Error bars: SE from three different cell preparations assayed individually
Expression of heat shock proteins on the cell membrane
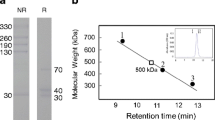
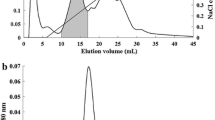
SBL/RC-RNase clearly bound to a 70-kDa band in fractions 4 to 7, which contained Triton X-100-insoluble cell membrane (Fig. 6a). Mascot analysis identified the protein as 70-kDa heat shock cognate protein (Hsc70). Minor components identified were 4F2 cell surface antigen heavy chain (CD98) (Mr 90,180) and guanine nucleotide-binding protein (Mr 41,010).
Expression of Hsc70 and Hsp70 on the P388 cell surface. a Results of western blotting. Following lysis of cells, detergent-insoluble fractions were obtained by sucrose density gradient centrifugation as described in Materials and methods, “Detection of binding triggers of SBL/RC-RNase in low-density, detergent-insoluble membrane fractions” section. Protein (10 μg) from each fraction was subjected to SDS-PAGE and transferred onto a PVDF membrane. Left: detection of SBL/RC-RNase receptor protein by treatment with SBL/RC-RNase, anti-SBL/RC-RNase antibody, and HRP-conjugated goat anti-rabbit IgG. Right: detection of HSC70 by treatment with its primary and secondary antibody as described in Materials and methods. b Expression of Hsp70 and Hsc70 was quantified by flow cytometry. Cells were treated with anti-Hsc70 mAb or anti-Hsp70 mAb and FITC-conjugated goat anti-mouse IgG, and fluorescence was detected by flow cytometry (FACSCalibur). c Specific binding of Hsp70 to SBL/RC-RNase. Various concentrations of Hsp70 (0.1–10 μg/mL) were placed on an ELISA plate, and SBL/RC-RNase (circles) or bovine pancreatic RNase A (squares) (each 50 μg/mL) was added. Bound RNases were detected by anti-SBL/RC-RNase or anti-bovine pancreatic RNase A polyclonal antibody as primary antibody and horseradish peroxidase-conjugated anti-rabbit IgG goat antibody as secondary antibody. Absorbances from reaction products were detected using an ImmunoMini NJ-2300 plate recorder. d Detection of SBL/RC-RNase and Hsc70 on the cell surface. Cells were treated with SBL/RC-RNase and anti-SBL/RC-RNase antibody, anti-Hsc70 antibody, and Alexa 488 goat anti-rabbit IgG antibody or Alexa 568-conjugated goat anti-mouse IgG antibody. Merge: partial co-localization of SBL/RC-RNase and Hsc70 on the cell surface
In addition to Hsc70, heat shock protein 70 (Hsp70) bound to SBL/RC-RNase and was present in the Triton X-100-insoluble fractions derived from P388 cells by sucrose density ultracentrifugation (data not shown). FACS analysis showed that Hsp70 was also localized on the cell membrane (Fig. 6b). Binding of SBL/RC-RNase to Hsp70 on the cell membrane was inhibited by the co-presence of free Hsp70 in an ELISA plate. The binding of frog RNase to Hsp70 (concentration 10 μg/mL) was significantly (~3-fold) stronger than that of bovine pancreatic RNase (Fig. 6c, circles vs. squares). The co-localization of SBL/RC-RNase and Hsc70 on the cell surface was demonstrated using fluorescence-conjugated antibodies against SBL/RC-RNase and Hsc70 (Fig. 6d).
The heat shock protein inhibitor quercetin suppresses apoptotic activity of SBL/RC-RNase
Pretreatment of P388 cells with 5 μM quercetin reduced the expression of Hsp70 and Hsc70 by 60 % and 20 %, respectively (Fig. 7a). SBL/RC-RNase-induced apoptosis was 20 % lower in cells pretreated with 5 μM quercetin than in non-pretreated cells (Fig. 7b, column 2 vs. 4). DNA ladder formation caused by SBL/RC-RNase in non-pretreated cells (Fig. 7c, lane 2) was reduced by pretreatment with 5 μM quercetin (Fig. 7c, lane 4). These findings suggest that SBL/RC-RNase bound to heat shock proteins and that this molecular interaction was not only a physicochemical phenomenon but also a physiologically-influenced cytotoxic effect acting as a trigger for SBL/RC-RNase.
SBL/RC-RNase-induced cytotoxicity was reduced by inhibition of Hsc70 and Hsp70 biosynthesis. a Reduced expression of Hsc70 and Hsp70 on the P388 cell surface by culturing with the heat shock protein inhibitor quercetin (Quer). Gene expression levels of Hsc70 and Hsp70 were reduced after quercetin treatment, as detected by quantitative RT-PCR. b The reduction in cell viability induced by 3 μM SBL/RC-RNase was recovered by co-culture with 5 μM quercetin. c Inhibition of SBL/RC-RNase-induced DNA fragmentation by the co-presence of quercetin. The numbers of samples loaded onto agarose gel for electrophoresis are described in panel b. Error bars: SE from three different cell preparations assayed individually. *, P < 0.01
Discussion
Despite earlier studies of the molecular basis of apoptosis induction by frog oocyte RNases [19], little is known regarding the receptors or trigger molecules of the RNases for induction of anticancer activity.
Starting with our original 1987 studies [1, 2], we have investigated the role of glycoconjugates that contain sialic acids as triggers for apoptosis by SBL/RC-RNase. We have observed that the amount of sialic acids on cancer cells varies depending on the cell line and that different cell lines show differing sensitivities to SBL/RC-RNase; e.g., P388, K562, and HL-60 cells undergo apoptosis following SBL/RC-RNase treatment, whereas SBL/RC-RNase does not cause apoptosis or any other effect in MCF7, Daudi, or Raji cells or in cell lines derived from normal tissues. These differing sensitivities are presumably due to differences in cell surface components.
Our present findings indicate that contact with molecules on the cell surface promotes incorporation of SBL/RC-RNase into the cytoplasm (Fig. 1b) prior to occurrence of apoptosis (with nuclear condensation) (Fig. 2a) or DNA fragmentation (ladder formation) (Fig. 2b, c). We are currently investigating the specific role in cytotoxicity of pyroglutamic acid (5-pyrolidone-2-carboxylic acid) located at the N-terminus of SBL/RC-RNase [2]. The properties arising from pyroglutamylation at the N-terminus of amphibian RNase differ from those for bovine pancreatic RNase A [33], and the role in cytotoxicity of post-translational modifications of this protein have not been well studied. These are essential points for elucidating the cytotoxic mechanisms of proteins such as amyloid-β, which is accumulated in the brains of patients with Alzheimer’s disease [34].
We found that the increased P388 cell viability (Fig. 3a) and decreased DNA fragmentation (Fig. 3b) caused by SBL/RC-RNase administration occurred through desialylation and the co-presence of sialomucin. These findings suggest that cell surface sialyl-glycoconjugates function as receptors for SBL/RC-RNase that allow cellular uptake (endocytosis) leading to apoptosis induction. A recent study by Chao et al. showed that specific amino acid residue-modified bovine RNase A induced cytotoxicity against Chinese Hamster Ovary (CHO) cells, and that cellular uptake increased through binding to heparan sulfate and chondroitin sulfate of proteoglycans on the cell surface [35]. We found that binding of SBL/RC-RNase to P388 cells was specifically inhibited by the co-presence of heparin, heparan sulfate, or sialyl-glycoprotein (BSM). The co-presence of a monosaccharide, sialic acid, did not inhibit SBL/RC-RNase cytotoxicity. Thus, some form of acidic saccharides, e.g., glycoconjugates, may be necessary for recognition of SBL/RC-RNase. This concept is supported by the observation that DNA ladder formation in P388 cells was inhibited by the co-presence of acidic glycans such as sialomucin, heparin, and heparan sulfate (Fig. 3b). Deconstruction of cholesterol-rich microdomains by pretreatment with the cholesterol inhibitor MβCD [26] inhibited the cytotoxic activity of SBL/RC-RNase, indicating that sialyl-glycoconjugates were a trigger for SBL/RC-RNase localization at these microdomains (Fig. 4a).
SBL/RC-RNase-induced apoptosis was shown to be triggered by caspase-3 activation (Fig. 4b, black bar). Reduction of caspase-3 activation by SBL/RC-RNase resulted from deconstruction of cholesterol-rich microdomains by MβCD treatment (Fig. 4b, gray bar). SBL/RC-RNase was activated through binding with sialyl-glycoconjugates in cholesterol-rich microdomains and was incorporated into cells to activate caspase-3, which induced apoptosis (Fig. 4b). Caspase-8 was activated by SBL/RC-RNase administration, and apoptosis was inhibited by pretreatment with the synthetic peptide Z-IETD-fmk (a specific inhibitor of caspase-8), indicating that SBL/RC-RNase-induced apoptosis occurred through a membrane-associated pathway involving both caspase-8 and -3 (Fig. 5b). Desialylation of cells also inhibited caspase-8 activation by SBL/RC-RNase (Fig. 5c).
Heat shock protein (an acute stress protein) and heat shock cognate protein were found in the Triton X-100-insoluble fractions of P388 cells; these fractions also contained GSL-enriched microdomains. SBL/RC-RNase bound to these heat shock proteins even though they are not sialic acid-containing glycoproteins (Fig. 6a). These findings indicate that the binding of SBL/RC-RNase to heat shock proteins (Hsc70 and Hsp70) is based on protein-protein interaction rather than protein-glycan interaction. FACS analysis showed that Hsc70 and Hsp70 were located on cell membranes (Fig. 6b). Binding of SBL/RC-RNase to Hsp70 was stronger than that of bovine RNase (Fig. 6c).
The 70-kDa heat shock protein family has two major members, the inducible Hsp70 and the constitutive Hsc70. Both Hsp70 and Hsc70 are intracellular proteins and function as molecular chaperones, playing important roles in folding and transporting newly synthesized peptides [36, 37]. Hsp70 expression is also an important mechanism whereby cells adapt to stress. Recombinant Hsc70 was recently reported to interact directly with TLR4 in macrophages. It appears that extracellular Hsc70 activates “inert” immunity to induce pro-inflammatory cytokine expression [38].
Previous studies indicated that heat shock proteins, in their role as molecular chaperones involved in protein folding, form complexes with gangliosides, and that the ganglioside/Hsp70 complex is formed through the N-terminal ATPase and the C-terminal peptide-binding domains [39, 40]. Hsp70 and its structural homologue Hsc70 are most likely present in cholesterol-rich microdomains of P388 cells in similar fashion. Our immunohistochemical experiments showed that heat shock proteins are localized on the cell membrane and may function as indirect receptors for SBL/RC-RNase (Fig. 6d). A complex of SBL/RC-RNase, sialyl-glycoconjugates, and heat shock proteins in the cholesterol-rich microdomain of P388 cells promoted apoptosis, and this promoting effect was blocked by pretreatment with quercetin, a specific inhibitor of heat shock protein synthesis (Fig. 7).
Studies since 2009 on specific RNA-binding protein families have shown that Ser- and Arg-rich mRNA splicing factors have carbohydrate-binding activity, whereas small nuclear ribonucleoproteins interact with carbohydrate-binding proteins such as galectin-1 and -3 [41–43]. This concept is supported by our observation of specific sialyl-glycoconjugate-binding activity by SBL/RC-RNase. We are currently attempting to characterize the glycoproteins that act as receptors of SBL/RC-RNase. Further elucidation of the relationship between quantitative expression of sialyl-glycoconjugates and heat shock proteins on the cell surface and SBL/RC-RNase-induced apoptosis will increase our understanding of the cell surface receptors of cytotoxic proteins and promote new concepts of synergistic mechanisms for apoptosis induction.
Abbreviations
- FACS:
-
Fluorescence-activated cell sorting
- GSL:
-
Glycosphingolipid
- HSC70:
-
70-kDa heat shock cognate protein
- HSP70:
-
70-kDa heat shock protein
- MβCD:
-
Methyl β-D-cyclodextrin
- NHDF:
-
Normal human epidermal fibroblast
- NHEM:
-
Normal human epidermal melanocyte
- NHEK:
-
Normal human keratinocyte cell
- RC-RNase:
-
Rana catesbeiana ribonuclease
- SBL:
-
Sialic acid-binding lectin
References
Nitta, K., Takayanagi, G., Kawauchi, H., Hakomori, S.: Isolation and characterization of Rana catesbeiana lectin and demonstration of the lectin-binding glycoprotein of rodent and human tumor cell membranes. Cancer Res. 47, 4877–4883 (1987)
Titani, K., Takio, K., Kuwada, M., Nitta, K., Sakakibara, F., Kawauchi, H., Takayanagi, G., Hakomori, S.: Amino acid sequence of sialic acid binding lectin from frog (Rana catesbeiana) eggs. Biochemistry 26, 2189–2194 (1987)
Nitta, R., Katayama, N., Okabe, Y., Iwama, M., Watanabe, H., Abe, Y., Okazaki, T., Ohgi, K., Irie, M.: Primary structure of a ribonuclease from bullfrog (Rana catesbeiana) liver. J. Biochem. 106, 729–735 (1989)
Nitta, K., Oyama, F., Oyama, R., Sekiguchi, K., Kawauchi, H., Takayanagi, Y., Hakomori, S., Titani, K.: Ribonuclease activity of sialic acid-binding lectin from Rana catesbeiana eggs. Glycobiology 3, 37–45 (1993)
Nitta, K., Ozaki, K., Tsukamoto, Y., Furusawa, S., Ohkubo, Y., Takimoto, H., Murata, R., Hosono, M., Hikichi, N., Sasaki, K., Kawauchi, H., Takayanagi, Y., Tsuiki, S., Hakomori, S.: Characterization of a Rana catesbeiana lectin-resistant mutant of leukemia P388 cells. Cancer Res. 54, 928–934 (1994)
Nitta, K., Ozaki, K., Tsukamoto, Y., Hosono, M., Ogawa-Konno, Y., Kawauchi, H., Takayanagi, Y., Tsuiki, S., Hakomori, S.: Catalytic lectin (leczyme) from bullfrog (Rana catesbeiana) eggs. Int. J. Oncol. 9, 19–23 (1996)
Nitta, K.: Leczyme. Methods Enzymol. 341, 368–374 (2001)
Wu, Y.N., Mikulski, S.M., Adelt, W., Rybak, S.M., Youle, R.J.: A cytotoxic ribonuclease. Study of the mechanism of onconase cytotoxicity. J. Biol. Chem. 268, 10668–10693 (1993)
Ardelt, W., Mikulski, S.M., Shogen, K.: Amino acid sequence of an anti-tumor protein from Rana pipiens oocytes and early embryos. Homology to pancreatic ribonucleases. J. Biol. Chem. 266, 245–251 (1991)
Liao, Y.D.: A pyrimidine-guanine sequence-specific ribonuclease from Rana catesbeiana (bullfrog) oocytes. Nucleic Acids Res. 20, 1371–1377 (1992)
Liao, Y.D., Wang, J.J.: Yolk granules are the major compartment for bullfrog (Rana catesbeiana) oocyte-specific ribonuclease. Eur. J. Biochem. 222, 215–220 (1994)
Liao, Y.D., Huang, H.C., Chan, H.J., Kuo, S.J.: Large-scale preparation of a ribonuclease from Rana catesbeiana (bullfrog) oocytes and characterization of its specific cytotoxic activity against tumor cells. Protein Expr. Purif. 7, 194–202 (1996)
Goparaju, C.M., Blasberg, J.D., Volinia, S., Palatini, J., Ivanov, S., Donington, J.S., Croce, C., Yang, H., Pass, H.I.: Onconase mediated NFKβ downregulation in malignant pleural mesothelioma. Oncogene 30, 2767–2777 (2011)
Nasu, M., Carbone, M., Gaudino, G., Ly, B.H., Bertino, P., Shimizu, D., Morris, P., Pass, H.I., Yang, H.: Ranpirnase interferes with NF-κB pathway pathway and MMP9 activity, inhibiting malignant mesothelioma cell invasiveness and xenograft growth. Genes Cancer 2, 576–584 (2011)
Hu, C.C., Tang, C.H., Wang, J.J.: Caspase activation in response to cytotoxic Rana catesbeiana ribonuclease in MCR-7 cells. FEBS Lett. 503, 65–68 (2001)
Tseng, H.H., Yu, Y.L., Chen, Y.L.S., Chen, J.H., Chou, C.L., Kou, T.Y., Wang, J.J., Lee, M.C., Huang, T.H., Chen, M.H.C., Yiang, G.T.: RC-RNase-induced cell death in estrogen receptor positive breast tumor through down-regulation of Bcl-2 and estrogen receptor. Oncol. Rep. 25, 849–853 (2011)
Chang, C.F., Chen, C., Chen, Y.C., Hom, K., Huang, R.F., Huang, T.H.: The solution structure of a cytotoxic ribonuclease from the oocytes of Rana catesbeiana (bullfrog). J. Mol. Biol. 283, 231–244 (1998)
Liu, J.H., Liao, Y.D., Sun, Y.J.: Crystallization and preliminary X-ray diffraction analysis of cytotoxic ribonucleases from bullfrog Rana catesbeiana. Acta Crystallogr. D Biol. Crystallogr. 57, 1697–1699 (2001)
Turcotte, R.F., Lavis, L.D., Raines, R.T.: Onconase cytotoxicity relies on the distribution of its positive charge. FEBS J. 276, 3846–3857 (2009)
Sundless, N.K., Raines, R.T.: Arginine residues are more effective than lysine residues in eliciting the cellular uptake of onconase. Biochemistry 50, 10293–10299 (2011)
Tennant, J.R.: Evaluation of the trypan blue technique for determination of cell viability. Transplantation 2, 685–694 (1964)
Vyas, H.K., Pal, R., Vishwakarma, R., Lohiya, N.K., Talwar, G.P.: Selective killing of leukemia and lymphoma cells ectopically expressing hCGβ by a conjugate of curcumin with an antibody against hCGβ subunit. Oncology 76, 101–111 (2009)
Pepper, C., Thomas, A., Tucker, H., Hoy, T., Bentley, P.: Flow cytometric assessment of three different methods for the measurement of in vivo apoptosis. Leuk. Res. 22, 439–444 (1998)
Sugawara, S., Hosono, M., Ogawa, Y., Takayanagi, M., Nitta, K.: Catfish egg lectin causes rapid activation of multidrug resistance 1 P-glycoprotein as a lipid translocase. Biol. Pharm. Bull. 28, 434–441 (2005)
Sugawara, S., Kawano, T., Omoto, T., Hosono, M., Tatsuta, T., Nitta, K.: Binding of Silurus asotus lectin to Gb3 on Raji cells causes disappearance of membrane-bound form of HSP70. Biochim. Biophys. Acta 1790, 101–109 (2009)
Gniadecki, R., Christoffersen, N., Wulf, H.C.: Cholesterol-rich plasma membrane domains (lipid rafts) in keratinocytes: importance in the baseline and UVA-induced generation of reactive oxygen species. J. Investig. Dermatol. 118, 582–588 (2001)
Hakomori, S., Handa, K.: Interaction of glycosphingolipids with signal transducers and membrane proteins in glycosphingolipid-enriched microdomains. Methods Enzymol. 363, 191–207 (2003)
Laemmli, U.K.: Cleavage of structural proteins during assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970)
Matsudaira, P.: Sequence from picomole quantities of proteins electroblotted onto polyvinylidene difluoride membranes. J. Biol. Chem. 262, 10035–10038 (1987)
Krajewski, S., Zapata, J.M., Reed, J.C.: Detection of multiple antigens on western blots. Anal. Biochem. 236, 221–228 (1996)
Hamaguchi, A., Suzuki, E., Murayama, K., Fujimura, T., Hikita, T., Iwabuchi, K., Handa, K., Withers, D.A., Casters, S.C., Fu, H., Hakomori, S.: Sphingosine-denependent protein kinase-1, directed to 14-3-3, is identified as the kinase domain of protein kinase Cd. J. Biol. Chem. 278, 41557–41565 (2003)
Perkins, D.N., Pappin, D.J.C., Creasy, D.M., Cottrell, J.S.: Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999)
Kamiya, Y., Oyama, F., Oyama, R., Sakakibara, F., Nitta, K., Kawauchi, H., Takayanagi, Y., Titani, K.: Amino acid sequence of a lectin from Japanese frog (Rana japonica) eggs. J. Biochem. 108, 139–143 (1990)
Morawski, M., Hartlage-Rubsamen, M., Jager, C., Waniek, A., Schilling, S., Schwab, C., McGeer, P.L., Arendt, T., Demuth, H.U., Rossner, S.: Distinct glutaminyl cyclase expression in Edinger-Westphal nucleus, locus coeruleus and nucleus basalis Meynert contributes to pGlu-Aβ pathology in Alzheimer’s disease. Acta Neuropathol. 120, 195–207 (2010)
Chao, T.Y., Lavis, L.D., Raines, R.T.: Cellular uptake of ribonuclease A relies an anionic glycans. Biochemistry 49, 10666–10673 (2010)
Su, X., Sykes, J.B., Ao, L., Raeburn, C.D., Fullerton, D.A., Meng, X.: Extracellular heat shock cognate protein 70 induces cardiac functional tolerance to endotoxin: differential effect on TNF-α and ICAM-1 levels on heart tissue. Cytokine 51, 60–66 (2010)
Young, J.C., Barral, J.M., Hartl, F.U.: More than folding: localized functions of cytosolic chaperones. Trends Biochem. Sci. 28, 541–547 (2003)
Zou, N., Ao, L., Cleveland Jr., J.C., Yang, X., Su, X., Cai, G.Y., Banerjee, A., Fullerton, D.A., Meng, X.: Critical role of extracellular heat shock cognate protein 70 in the myocardial inflammatory response and cardiac dysfunction after global ischemia-reperfusion. Am. J. Physiol. Heart Circ. Physiol. 294, H2805–H2813 (2008)
Harada, Y., Sato, C., Kitajima, K.: Complex formation of 70 kDa heat shock protein with acidic glycolipids and phospholipids. Biochem. Biophys. Res. Commun. 353, 655–660 (2007)
Maehashi, E., Sato, C., Ohta, K., Harada, Y., Matsuda, T., Hirohashi, N., Lennarz, W.J., Kitajima, K.: Identification of the sea urchin 350-kDa sperm-binding protein as a new sialic acid-binding lectin that belongs to the heat shock protein 110 family: implication of its binding to gangliosides in sperm lipid rafts in fertilization. J. Biol. Chem. 278, 42050–42057 (2003)
Hatakeyama, S., Sugihara, K., Nakayama, J., Akama, T.O., Wong, S.M., Kawashima, H., Zhang, J., Smith, D.F., Ohyama, C., Fukuda, M., Fukuda, M.N.: Identification of mRNA splicing factors as the endothelial receptor for carbohydrate-dependent lung colonization of cancer cells. Proc. Natl. Acad. Sci. U. S. A. 106, 3095–3100 (2009)
Haudek, K.C., Patterson, R.J., Wang, J.L.: SP proteins and galectins: what’s in a name? Glycobiology 20, 1199–1207 (2010)
Haudek, K.C., Spronk, K.J., Voss, P.G., Patterson, R.J., Wang, J.L., Arnoys, E.J.: Dynamics of galectin-3 in the nucleus and cytoplasm. Biochim. Biophys. Acta 1800, 181–189 (2010)
Acknowledgments
This study was supported in part by Grants-In-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT) Japan, the Japan Society for the Promotion of Science (JSPS) and Nagasaki International University. The authors are grateful to Dr. Stephen Anderson for English editing of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Y. Ogawa and S. Sugawara made equal contributions to the study and are both considered as first authors.
Rights and permissions
About this article
Cite this article
Ogawa, Y., Sugawara, S., Tatsuta, T. et al. Sialyl-glycoconjugates in cholesterol-rich microdomains of P388 cells are the triggers for apoptosis induced by Rana catesbeiana oocyte ribonuclease. Glycoconj J 31, 171–184 (2014). https://doi.org/10.1007/s10719-013-9513-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-013-9513-7