Abstract
Aging is a genetically regulated process that happens in all organisms. Lifespan extension can be achieved through several mechanisms, including regulatory signaling from germline stem cells, which will be the focus of this chapter. The free-living nematode Caenorhabditis elegans has become the standard workhorse for aging studies due to its fast life cycle, short lifespan, and powerful functional genomics. In this chapter, we will first introduce germline organization and germline stem cell maintenance in C. elegans. Next, we will review the knowledge achieved by C. elegans research on how gonadal signaling pathways regulate organism longevity. Lastly, the current model of lipid metabolic reprogramming as the link between germline and longevity will be discussed as well.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Aging is a fundamental biological process occurring in the whole animal kingdom. It refers to the gradual loss of normal functions in organs, physiological systems, and ultimately the organism as a whole. Although this phenomenon is a familiar experience for all, it is only recently that people has started to realize aging is not just a simple process of wear and tear but rather a series of complex regulatory processes under tight control. Over the last two decades, groundbreaking work in model organisms has identified many genes and signaling pathways that regulate organism lifespan, including insulin/IGF-1 receptor, forkhead transcription factor, TOR kinase, AMP kinase, and Sirtuin (Blagosklonny et al. 2010; Kenyon 2010b). Strikingly, most of those molecular mechanisms were first linked to lifespan regulation by using a tiny free-living nematode, Caenorhabditis elegans, as a genetic model organism and lately shown to be well conserved across other species.
C. elegans is small (~1 mm in length) and transparent and has a fast life cycle, passing through four larval stages to adulthood in approximately 3 days. It also has short lifespan (~2–3 weeks), which is remarkably shorter than 2–3 months in Drosophila or 2–3 years in mice. The entire cell lineage of C. elegans has been completely mapped and found to be invariant among generations and populations (Sulston et al. 1983). Therefore, the developmental fate of each single cell in those tiny worms is clear throughout their lives. Furthermore, with fully sequenced genome (~97 megabases) (Consortium 1998), and the ease of RNA interference (RNAi) and transgenic techniques, C. elegans is highly amenable for genetic manipulation. Together, those features have made C. elegans a powerful animal model in studies of development, reproduction, metabolism, evolution, and aging. C. elegans is a multicellular organism like human beings, in which the functions of different cells and organs integrate and cooperate to support various biological processes, including aging. C. elegans lifespan can be modulated via a cell nonautonomous manner with the nervous system, intestine (the worm fat storage tissue), and germline stem cells (GSCs) serving as a major endocrine signaling network (Russell and Kahn 2007). In this chapter, we will focus on the endocrine function of GSCs and review the knowledge known to date on how those stem cells systemically modulate the aging process of the whole organism in C. elegans.
2 GSC Maintenance in C. elegans
Before we start to discuss the endocrine functions of GSCs in the regulation of organism aging, it is necessary to first overview their organization in the germline, and the signaling events that determine their stem cell fate and functions.
2.1 Germline Development in C. elegans
The reproductive system in adult C. elegans is consisting of two symmetrical U-shaped arms, connected by a common uterus. In each germline arm of adult worms, all stages of germ cells, from GSCs to differentiated gametes, are present at one time but finely regulated to specialize their positions. Germline specification occurs during embryogenesis and early L1 larval stage (Strome 2005). This process relies on the distribution of P granule at early embryogenesis (Updike and Strome 2010). P granule contains maternal RNAs and is asymmetrically segregated into P blastomeres, which eventually grow into Z2 and Z3 primordial germ cells (Fig. 3.1a). In early L1 worms, the Z2 and Z3 primordial germ cells (PGCs), together with Z1 and Z4 somatic gonad precursors (SGPs), are surrounded within a basement membrane (Fig. 3.1b). These four cells will give rise to the complete reproductive system during the postembryonic development.
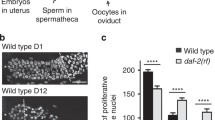
Germline organization in C. elegans. (a) Germline specification relies on maternally inherited germ granules, called P granules. P granules contain RNA and proteins required for germline determination and are progressively segregated into a series of P blastomeres (highlighted as turquoise cells) during embryonic development. P4 will eventually divide into two PGCs (Z2 and Z3, turquoise) at the ~100-cell stage. (b) In postembryonic development, the Z2 and Z3 PGCs will continue to proliferate and comprise the future germ cell population after hatch, which are supported by the DTCs (red) and sheath cells developed from Z1 and Z4 SGPs (white). The DTCs locate at the distal ends of gonad arms, acting as the niche for GSC maintenance and proliferation. Germ cells proliferate mitotically (turquoise) in the distal region; when progressing into the meiotic region, germ cells become meiotic (light blue) and undergo sperm (purple dot) differentiation at the L4 stage and oogenesis (yellow) later during adulthood. The matured oocyte goes through spermatheca and becomes self-fertilized egg (orange) in the uterus. (c) The DTC projects processes and surrounds the distal end of the gonad arm. Those germ cells adjacent to the DTC remain mitotic; otherwise, they turn into meiotic differentiation program. This is controlled via LAG-2/GLP-1 Notch-like signaling pathway. In brief, the DTC-expressed LAG-2 triggers the activation of its receptor GLP-1 and downstream FEB-1/2 signaling in GSCs. FEB-1/2 maintain mitotic proliferation fate by promoting mitosis-specific mRNA translation via inhibiting GLD-1/NOS-3 and by suppressing meiosis-specific mRNA translation via inhibiting GLD-2/3. When germ cells progress toward the proximal end, they lose the DTC signaling; thus, GLD-1/NOS-3 and GLD-2/3 take the lead to switch on meiotic program. Abbreviation: PGC primordial germ cell, SGP somatic gonad precursors, DTC distal tip cell, GSC germline stem cell
Germline proliferation, survival, and differentiation continue through L1 to L3 larval stages, followed by gametogenesis at L4 and early adulthood (Fig. 3.1b) (Hubbard and Greenstein 2005). When nutritional condition is favorable, PGCs undergo MES-mediated chromatin modification to activate proliferation genes, divide symmetrically to expand syncytial GSC population, and ultimately give rise to germline tissues. Meanwhile, each SGP differentiates asymmetrically into five proximal cells and one distal tip cell (DTC). The proximal cells eventually turn into sheath cells, forming single layer intimately surrounding the germ cells. The DTC, at the distal end of each germline arm, forms processes that stretch out proximally along germline surface and functions as a niche to maintain the adjacent GSCs (Fig. 3.1c). During the L4 larval stage, the polarity of the germline forms. As new cells continue proliferating at the distal end, the other germ cells are pushed toward the proximal side and farther away from the DTC. Only GSCs and early germ cells at the distal end remain mitotic. In contrast, those germ cells progressing to the proximal end switch fate from mitotic proliferation to meiotic differentiation and undergo spermatogenesis during the L4 stage, followed by oogenesis starting at adulthood (Fig. 3.1b). In adulthood, oogenesis can continue throughout the whole reproductive period until sperm exhaustion.
2.2 Molecular Mechanism Controlling GSC Maintenance and Activity
In C. elegans, GSCs are located at the most distal end of the germline. The somatic DTC extends thin cytoplasmic processes that encircle the most distal end germ cells and serves as a niche for adjacent GSCs (Fig. 3.1c) (Li and Xie 2005; Xie 2008). When DTC is killed by laser ablation, the germline mitotic region diminishes (Kimble and White 1981). Conversely, DTC relocation results in the development of ectopic proliferating germ cells at the corresponding position, and DTC duplication leads to doubling of germ cell pools (Kidd et al. 2005; Kipreos et al. 2000; Lam et al. 2006). The DTC promotes mitotic proliferation of GSCs via GLP-1/Notch signaling. LAG-2, a notch-like signaling ligand, is expressed in the DTC and activates its receptor GLP-1, located on the surface of GSCs (Fig. 3.1c) (Crittenden et al. 1994; Fitzgerald and Greenwald 1995; Henderson et al. 1994). The activation of GLP-1/Notch signaling is necessary and sufficient for GSC maintenance and proliferation. In the loss-of-function mutants of glp-1, all of the germ cells progress into meiotic differentiation, leading to a complete loss of the GSC population (Fitzgerald and Greenwald 1995). Conversely, gain-of-function mutations of glp-1 promote GSC overproliferation and the formation of germline tumor (Berry et al. 1997).
In response to the activation of GLP-1/Notch signaling, a battery of downstream RNA regulators cooperate to promote GSC self-renewal and prevent cell differentiation (Fig. 3.1c) (Byrd and Kimble 2009; Kimble and Crittenden 2005). In brief, GLP-1 can transcriptionally activate fbf-2 (along with other unidentified targets) (Lamont et al. 2004). FBF-2 is a pumilio-like RNA-binding translational repressor that can work together with another pumilio-like protein FBF-1 to repress the expression of gld-1 and gld-3 via binding to their 3′-untranslated regions (UTRs) (Crittenden et al. 2002; Eckmann et al. 2004). GLD-1 together with NOS-3 and GLD-3 together with GLD-2 form two parallel regulatory branches to control meiotic entry in the germline. In one branch, GLD-1 functions as a translational repressor that acts through regulatory elements present in the 3′-UTR of mitosis-promoting genes and represses their expression posttranscriptionally. NOS-3, a Nanos family of RNA-binding proteins, physically interacts with FBF-1 and promotes GLD-1 accumulation (Hansen et al. 2004). In the other branch, GLD-2, a cytoplasmic poly(A) polymerase and translational activator, activates meiosis-promoting genes while GLD-3, a member of the bicaudal-C family of RNA-binding protein, enhances the poly(A) polymerase activity of GLD-2 and antagonizes the FBF-mediated repression (Eckmann et al. 2002, 2004; Wang et al. 2002). Together, GLP-1/Notch signaling and FBF-GLD proteins constitute the control hub for GSC maintenance in adults (Fig. 3.1c).
3 GSC Arrest Promotes Longevity in C. elegans: Trade-Off Between Reproduction and Longevity?
As mentioned in the introduction, we now know that the aging process is under the control of various regulatory pathways (Blagosklonny et al. 2010; Kenyon 2010b). Altered activity of those pathways by genetic or pharmacological methods can effectively influence organism lifespan. The first of such pathways was characterized in C. elegans and named the insulin/IGF-1 signaling (IIS) pathway. Mutations of age-1 or daf-2, the worm homolog of phosphoinositide 3-kinase (PI3K) or insulin/IGF-1 receptor, respectively, were shown to increase C. elegans adult lifespan by two-folds (Friedman and Johnson 1988; Kimura et al. 1997; Samuelson et al. 2007). IIS activation is transduced through PDK-1 (PIP3-dependent kinase 1) and AKT-1/2 (AKT/protein kinase B) and inhibits the transcriptional activity of the FOXO forkhead transcription factor, DAF-16. Loss-of-function mutations of pdk-1 or akt-1/2 increase C. elegans lifespan (Paradis et al. 1999; Paradis and Ruvkun 1998) while the increased lifespan associated with daf-2 or age-1 mutants is suppressed by mutations in daf-16 (Ogg et al. 1997). Following C. elegans studies, the IIS pathway has been shown to regulate lifespan also in yeast, fruit flies, and mice (Longo et al. 2005). In human, several insulin receptor and IGF-1 receptor variants are linked to extremely long life in Ashkenazi Jewish centenarian and Japanese centenarian, respectively (Kojima et al. 2004; Suh et al. 2008). Furthermore, a number of AKT and FOXO3 variants have been implicated to be responsible for human exceptional longevity, as shown in multiple independent genetic association studies (Anselmi et al. 2009; Flachsbart et al. 2009; Pawlikowska et al. 2009; Willcox et al. 2008).
To date, the IIS pathway has been well recognized as a remarkable conserved mechanism to modulate the aging process across species. However, when first identified, its life-extending effect was suspected to be a result of fertility trade-off. Could it be possible that the daf-2 and age-1 mutants live longer simply because of their comprised reproductive abilities and consequent allocation of energy resources from reproduction to somatic maintenance? Several lines of evidences do not agree with this assumption. First of all, sterility alone, either by physical removal of whole gonad or using non-reproducing mutants, is not sufficient to ensure longevity (Kenyon et al. 1993). Secondly, fer-15 rescue can restore normal fertility in the age-1 mutants but has no effect on the lifespan extension (Johnson et al. 1993). In addition, the daf-2 mutants remain long lived at 15 °C, where the animals have similar reproductive ability as wild type (Tissenbaum and Ruvkun 1998). Last but not least, knockdown of daf-2 by RNAi only during adulthood extends lifespan without affecting reproduction while daf-2 inactivation during development reduces fertility with no lifespan extension (Dillin et al. 2002). Therefore, decreased fertility in the daf-2 mutants is likely due to its requirement during development, but not related to its longevity effects at adulthood.
So, what is the relationship between reproduction and longevity, if there is no simple trade-off between them? The merging view is that the reproductive system can produce regulatory signals that actively coordinate the metabolic state of the organism toward reproduction or toward survival. In the following sections, we will focus on the mechanistic details of how reproductive signaling might regulate lifespan in C. elegans.
4 Germline, GSCs, and Longevity
The first line of evidence that the reproductive system produces signals to regulate lifespan came from the classic experiments by the Kenyon laboratory (Hsin and Kenyon 1999). They showed that removal of the germline precursors Z2/3 PGCs by laser microsurgery results in germline-less animals that live 60 % longer than the mock control, while additional removal of the somatic precursors Z1/4 SGPs, which eliminates somatic gonad surrounding and supporting the germ cells, abrogates this lifespan extension (Hsin and Kenyon 1999). These suggest that the longevity effect conferred by germline ablation is not a simple consequence of sterility, but rather arises from certain regulatory signals. Removal of somatic gonad may antagonize those longevity-promoting signals (Hsin and Kenyon 1999). In addition, the longevity phenotype in the germline-less animals resulting from physical ablation can also be recapitulated using genetic mutants. Loss-of-function mutations in mes-1 that have no germ cells can live twofold longer, and the glp-1 mutants that arrest germ cell proliferation can live 1.5-fold longer (Arantes-Oliveira et al. 2002). In both studies of mes-1 and glp-1, the presence of somatic gonad is required for the longevity effects.
In the adult germline, different stages of proliferative and differentiated germ cells, including GSCs, mitotic germ cells, meiotic germ cells, oocytes, and sperms, are present at one time (Fig. 3.1b). Which of those specific germ cell groups contribute to the lifespan regulation? Surprisingly, several different sperm-deficient (fem-1, fog-1, fog-2, or fog-3) or oocyte-deficient (daz-1) mutants all exhibit normal lifespan without extension (Arantes-Oliveira et al. 2002). It indicates the negligible functions of gametogenesis in the regulation of organism lifespan. This is particularly unexpected for the oocyte-deficient mutants, given that vast energy is invested during oocyte formation. These observations further support that the lifespan extension conferred by germline deficiency is not a simple passive result of energy reallocation. On the other hand, both GLP-1/Notch signaling and FBF-GLD proteins are required for GSC maintenance and proliferation. Interestingly, loss-of-function mutations of glp-1 that arrest GSCs promote longevity, while glp-1 gain-of-function mutations or gld-1 loss-of-function mutations that cause GSC overproliferation in contrast shorten lifespan (Arantes-Oliveira et al. 2002). These observations reveal the crucial function of GCSs in regulating longevity. Furthermore, in the temperature-sensitive glp-1 mutants, GSC arresting upon temperature shift can always lead to increased longevity, even at day-1 adulthood when the whole germline development is completely accomplished (Arantes-Oliveira et al. 2002). Thus, GSC arrest itself is sufficient to promote longevity, and the lifespan regulatory signals could directly associate with the presence of GSCs.
Importantly, this GSC longevity regulatory mechanism is evolutionarily conserved. In Drosophila, adult flies lacking GSCs live 50 % longer than the controls (Flatt et al. 2008). In mice, ovary transplantation from young females increases the lifespan of older female recipients (Cargill et al. 2003), suggesting that unknown lifespan-enhancing endocrine signals are produced from mammalian ovary tissues.
5 Signaling Pathways Regulating Longevity in Response to Gonadal Signals
5.1 Steroid Hormone Control from the Reproductive System
Reproductive tissues are crucial functional parts of the endocrine system. Steroid hormones synthesized and released from the reproductive system can systemically coordinate the whole body physiology. Consistently, steroidal signaling has been implicated as a key factor contributing to the regulation of GSC longevity (Fig. 3.2). This includes the bile acid-like steroids (dafachronic acids (DA)), its receptor DAF-12, and several enzymes involved in DA synthesis, such as DAF-36/Rieske-like oxygenase, DHS-16/3-hydroxysteroid dehydrogenase, and DAF-9/cytochrome P450 (Lee and Schroeder 2012). Loss-of-function mutations of daf-12, daf-36, dhs-16, or daf-9 completely abrogate the longevity in the germline-less animals conferred by the glp-1 mutations or Z2/3 laser ablation (Beckstead and Thummel 2006; Gerisch et al. 2001; Wollam et al. 2012). Moreover, DA supplementation restores longevity in the daf-9;glp-1, dhs-16;glp-1, and daf-36;glp-1 double mutants back to that of glp-1 but has no effect in extending the shortened lifespan of daf-12;glp-1 (Gerisch et al. 2007; Wollam et al. 2012). Thus, DA biosynthesis and its transcriptional activation of DAF-12 are both required in promoting longevity of the germline-less animals.
Longevity regulation in germline-deficient C. elegans. Germ cell loss works in concert with somatic gonad, intestine, and neuronal cells to regulate lifespan extension. In somatic gonad, steroid synthesis via DAF-36, DHS-16, and DAF-9 generates hormone ligands, like DA and PREG, for DAF-12 activation. DAF-12 promotes longevity via regulating downstream target gene expression and via assisting FOXO/DAF-16 nuclear localization in the intestine. DAF-16 activation also requires KRI-1 and neuronal mir-71 activity. Once DAF-16 enters nucleus, it activates assorted downstream targets through forming distinct complexes with different factors. When DAF-16 interacts with TCER-1, PHI-62, and FTT-2, it is known to regulate the gene dod-8. DAF-16 also plays a central role in regulating lipl-4 and other lipolysis genes. With the aid of lipl-4, FOXA/PHA-4 regulates autophagy gene expression, e.g., lgg-1, bec-1, and unc-51. On the other hand, NHR-80 and NHR-49 activate the expression of SCD, including fat-6, to alter lipid composition by increasing oleic acid synthesis. These subsets of regulatory modules do not work self-sufficiently; instead, they belong to a closely linked network and may have partial dependency on others. These findings also suggest there is cross-tissue endocrine signaling to communicate the whole organism in the germline-depleted scenario and to achieve lifespan extension. Abbreviation: DA dafachronic acid, PREG pregnenolone, SCD stearoyl-CoA-Δ9-desaturases
As mentioned above, an intact somatic gonad is required for the longevity of the germline-less animals, suggesting somatic gonad as a source of life-extending signals. Notably, DA supplementation can restore longevity of the whole gonad-deficient worms lacking both germline and somatic gonad, which requires the presence of daf-12 (Gerisch et al. 2007). In addition, DAF-12 transcriptional activities are stimulated by germline loss, but further removal of somatic gonad diminishes this induction, which can be rescued by supplying DA exogenously (Yamawaki et al. 2010). These indicate that DA biosynthesis and DAF-12 signaling are both involved in mediating life-extending signals from the somatic gonad. In wild-type animals, those signals may be hindered by the presence of germline. Besides DA, C. elegans also contain several other hormonal steroids that are present in humans, including pregnenolone (3β-hydroxy-pregn-5-en-20-one; PREG) and other pregnane and androstane derivatives (Broue et al. 2007). Some of those steroids are also involved in the regulation of longevity in the germline-less animals. For example, in the glp-1 mutants, PREG levels are elevated in a DAF-9-dependent fashion. Moreover, PREG supplementation restores longevity in the daf-9;glp-1 double mutants, but fails to do so in the daf-12;glp-1 mutants, suggesting the involvement of PREG in mediating gonadal longevity signals (Broue et al. 2007). However, neither DA nor PREG supplementation is sufficient to extend lifespan in wild type or further increase longevity of germline-deficient animals. Therefore, there must be also other signaling components mediating the longevity effects conferred by germline loss.
5.2 Regulation of Intestinal FOXO Activity Upon Germline Loss
The FOXO forkhead transcription factor DAF-16 serves as another central regulator of longevity by germline deficiency (Fig. 3.2). Null mutation of daf-16 completely abolishes the longevity of the germline-ablated animals (Hsin and Kenyon 1999) and the glp-1 germline-defective mutants (Arantes-Oliveira et al. 2002; Berman and Kenyon 2006). Despite ubiquitous distribution in all worm tissues, DAF-16 activities in the intestine may be particularly critical to mediate the longevity effect brought about by germline loss. Upon laser microsurgery of germline precursor cells or in the glp-1 germline-deficient mutants, DAF-16 is translocalized to the nucleus in intestinal cells during the first day of adulthood, but its subcellular localization remains unchanged in other tissues (Berman and Kenyon 2006; Lin et al. 2001). Similar to mammalian FOXO transcription factors, translocation of DAF-16 from the cytoplasm to the nucleus triggers its activation, and three conserved AKT phosphorylation sites are required for its cytoplasmic retention. Substitution of those three consensus sites with alanines generates a constitutively nuclear DAF-16 protein (referred to as DAF-16 AM) (Lin et al. 2001). Importantly, expression of this mutated form of DAF-16 AM within the intestine fully restores the longevity of the glp-1;daf-16 double mutants, showing the functional significance of DAF-16 nuclear localization in this tissue (Berman and Kenyon 2006). The worm intestine is not only a digestive organ, but also a key metabolic and endocrine organ to store lipid and release humoral signals to the rest of the organism. Thus, activation of DAF-16 in the intestine by germline signals may generate secondary effects and hence have global impacts on organism longevity.
Multiple factors genetically act upstream of DAF-16 in response to germline loss. First of all, DA/DAF-12 signaling assists DAF-16 nuclear localization in the germline-less animals. Loss-of-function mutations of either daf-9, daf-36, dhs-16, or daf-12 substantially compromise the nuclear translocation of DAF-16 in the intestine, and DA supplementation restores DAF-16 nuclear localization in the glp-1 double mutants with those DA biosynthesis genes (Gerisch et al. 2007; Libina et al. 2003; Wollam et al. 2012). Noteworthily, expression of constitutively nuclear localized DAF-16 AM fails to restore longevity in glp-1; daf-16; daf-12 mutants (Berman and Kenyon 2006). It suggests that DAF-12 may not only facilitate DAF-16 nuclear translocalization, but also aid its transcriptional activation or have other unidentified downstream pathways. Secondly, scientists also apply unbiased genetic screens to search for other components involved in the germline longevity signaling. kri-1 (Berman and Kenyon 2006) and tcer-1 (Ghazi et al. 2009) are two genes identified in large-scale RNAi screens that remarkably suppress the longevity of the glp-1 mutants but have no effect on wild-type lifespan. KRI-1, a worm homolog of the human KRIT/CCM1 protein, is expressed in the intestine and is required for the intestinal nuclear localization of DAF-16 in the glp-1 mutants (Berman and Kenyon 2006). In the presence of DAF-16 AM, kri-1 loss-of-function mutations cannot abrogate the longevity of the glp-1 mutants (Berman and Kenyon 2006). Thus, KRI-1 is likely a downstream intestinal factor of the germline longevity signaling pathway, and its primary function is to promote DAF-16 nuclear localization in this tissue. As a transcription elongation factor, TCER-1 works together with DAF-16 in regulating the expression of specific target genes in response to germline depletion. tcer-1 is specifically upregulated in the nuclei of intestine and neurons in the germline-less animals, and its intestinal upregulation is dependent on kri-1, but not daf-12 (Ghazi et al. 2009). Although loss-of-function mutations of tcer-1 do not affect DAF-16 nuclear localization in the germline-less animals, they reduce the induction of several DAF-16 target genes and abrogate the longevity (Ghazi et al. 2009). Conversely, TCER-1 overexpression significantly triggers specific DAF-16 target gene expressions, along with lifespan extension by 15 %, which is dependent on DAF-16 activity (Ghazi et al. 2009). The mammalian homolog of TCER-1, TCERG-1, associates with RNA polymerase II and regulates transcription elongation and pre-mRNA splicing (Carty et al. 2000; Goldstrohm et al. 2001; Lin et al. 2004; Pearson et al. 2008; Smith et al. 2004; Sune and Garcia-Blanco 1999). A plausible model is that germline removal triggers the formation of specialized transcriptional complexes including TCER-1, DAF-16, and other unidentified factors, which modulate the expression of a specific subset of genes that are crucial for organism longevity. This is supported by the findings that TCER-1 and DAF-16 physically interact with FTT-2 (14-3-3 protein) and PHI-62 (RNA-binding protein) to regulate dod-8 (putative steroid dehydrogenase) expression, and both ftt-2 and phi-62 are required for the longevity in germline-less animals (Li et al. 2007; McCormick et al. 2012). However, the expression of sod-3 (Mn++ superoxide dismutase), another daf-16 downstream target, does not require ftt-2 nor phi-62 (McCormick et al. 2012). This reveals that DAF-16 may form a variety of complexes with different factors to exert diverse regulatory functions on multiple subsets of downstream genes and together create a network that promotes organism longevity (Fig. 3.2).
Interestingly, recent studies have also implicated neuronal microRNA mir-71 in the regulation of DAF-16 nuclear localization and longevity in the germline-less animals (Boulias and Horvitz 2012). In a large screen of microRNA loss-of-function mutants, mir-71 loss-of-function was identified to shorten wild-type lifespan by 40 %. More importantly, loss of mir-71 fully abrogates the longevity in the glp-1 mutants or the germline-ablated animals and also blocks intestinal nuclear localization of DAF-16 and its target gene expressions in those germline-less animals (Boulias and Horvitz 2012). Conversely, mir-71 overexpression modestly increases lifespan in wild type and further enhances the glp-1 longevity. This increased longevity fully requires DAF-16 and TCER-1 activities within the intestine but is not affected by DAF-12. Although broadly expressed in multiple tissues, mir-71 activity within neurons is particularly important to mediate the longevity effects by germline loss. Thus, mir-71 acts in a cell nonautonomous manner to regulate DAF-16 activity in the intestine. A complex crosstalk among the reproductive system, neurons, and intestine systemically regulates the organism aging process (Fig. 3.2).
5.3 Interaction with the IIS Pathway in the Regulation of Longevity
As mentioned above, the IIS pathway is the first identified, the best characterized, and the most conserved longevity mechanism to date. Whether and how does the germline longevity signaling interact with the IIS pathway? So far, the evidences suggest that these two mechanisms largely work independently (Kenyon 2010a; Panowski and Dillin 2009). First, germline deficiency and reduced daf-2 activity have synergistic effects genetically in promoting longevity. Ablation of germ cells in the daf-2 loss-of-function mutants further enhances already increased lifespan in the mutants (Hsin and Kenyon 1999). Second, DAF-16 responds to germline loss signals differently from reduced IIS. Removal of germline causes DAF-16 localization in intestinal nuclei at adulthood, while daf-2 loss-of-function mutants display DAF-16 nuclear accumulation in all somatic cells throughout life (Henderson and Johnson 2001; Lee et al. 2001; Lin et al. 2001). Third, as discussed above, multiple factors are required to promote longevity in the germline-less animals, including daf-12, kri-1, tcer-1, and mir-71. However, none of those components are necessary for the longevity conferred by reduced daf-2 activity (Berman and Kenyon 2006; Gems et al. 1998; Ghazi et al. 2009; Hsin and Kenyon 1999).
Nevertheless, the IIS pathway is likely involved in mediating the lifespan extending signals from the somatic gonad. In wild-type animals, removal of somatic gonad precursors abrogates the longevity brought by ablating germline precursors. However, in the strong daf-2 loss-of-function mutants, longevity remains even with whole gonad depletion (Hsin and Kenyon 1999), indicating that the somatic gonad promotes longevity via reducing IIS. Furthermore, a subset of DAF-16 target genes is induced by both daf-2 reduction and germline removal, such as sod-3, gpd-2 (glyceraldehyde 3-phosphate dehydrogenase), and dod-8 (Murphy et al. 2003). Interestingly, the inductions of sod-3 and gpd-2 both require the presence of somatic gonad (Yamawaki et al. 2008). Thus, the somatic gonad may exert influence through a regulatory network that at least partially overlaps with the IIS pathway, while germ cells act in parallel with the somatic reproductive tissue to regulate lifespan in a counterbalancing manner.
6 Metabolic Reprogramming in Mediating GSC Longevity
Following the removal of germline and activation of a series of downstream signaling pathways, what might be the mechanisms that confer longevity? Although not conclusive, recent studies reveal that the germline removal actively modulates the organism metabolic state, especially the lipid metabolism. This metabolic reprogramming may play a crucial role in promoting longevity.
6.1 LIPL-4 Lysosomal Lipid Signaling
The germline-less glp-1 mutants have substantial fat accumulation in the intestine and also in the hypodermis, another major fat storage tissue in C. elegans (Ashrafi 2007; Yokota et al. 2002). On the other hand, they have reduced levels of Nile red staining signals, which labels worm lysosomal-like organelles involved in lipid metabolism (Greenspan and Fowler 1985; Wang et al. 2008). To understand the mechanisms by which germline signals regulate lipid metabolism, scientists screened through an RNAi library targeting various key metabolic genes and identified a group of genes whose inactivation specifically increase Nile red staining levels in the glp-1 mutants (Wang et al. 2008). Interestingly, most of those genes are involved in catalyzing lipid hydrolysis, fatty acid transport and β-oxidation, and citric acid cycle, revealing an altered lipid metabolic state in the germline-less animals (Wang et al. 2008). Importantly, further characterization of one candidate, lipl-4, showed that its knockdown completely abrogates longevity of glp-1 and its overexpression robustly extends lifespan in gonad intact wild-type animals (Wang et al. 2008).
lipl-4 expression is upregulated in the germline-less animals in a daf-16- and kri-1-dependent manner (Wang et al. 2008) (Fig. 3.2). LIPL-4 is a lysosomal acid lipase, and its overexpression alone sufficiently promotes longevity through triggering the nuclear localization of LBP-8, a lysosomal lipid chaperone. This allows LBP-8 to deliver a lipid messenger oleoylethanolamide (OEA) to the nucleus and to consequently activate nuclear hormone receptors NHR-80 and NHR-49 (Folick et al. 2015). Activation of this lysosome-to-nucleus lipid-signaling pathway promotes a metabolic shift toward lipid catabolism (Folick et al. 2015). Further studies to fully understand the signaling role of lysosomes will be crucial to unveil the metabolic link between GSCs and longevity.
6.2 NHR-80/NHR-49 Mediated Fatty Acid Metabolic Changes
Not only total lipid content increases in the germline-less animals, but also lipid composition is altered in those animals. The monounsaturated oleic acid (OA, C18:1(n-9)) levels and the ratio of OA/stearic acid (SA, C18:0) are specifically increased in the germline-less animals (Goudeau et al. 2011). In C. elegans, OA is synthesized from SA by stearoyl-CoA-Δ9-desaturases (SCDs), fat-6, and faf-7 and is the key precursor for a variety of polyunsaturated fatty acids (PUFAs) (Watts and Browse 2002). Loss-of-function mutations of both fat-6 and fat-7 abrogate longevity of glp-1, which can be perfectly rescued if OA is supplied exogenously (Goudeau et al. 2011). This suggests that elevation of OA synthesis is necessary for the longevity in the germline-less animals. However, OA dietary supplementation is not sufficient to extend lifespan in gonad intact wild-type animals or further enhance longevity in the germline-less animals (Goudeau et al. 2011). The induction of OA synthesis is not mediated by daf-16 or daf-12 but requires other nuclear hormone receptors, including nhr-80 and nhr-49 (Brock et al. 2006; Goudeau et al. 2011) (Fig. 3.2).
Knockdown of nhr-80 by RNAi or mutation completely blocks glp-1 longevity, but has no effect on the lifespan of wild type or other long-lived conditions, such as daf-2 mutants, dietary restricted worms, or worms with reduced mitochondrial functions (Goudeau et al. 2011). NHR-80 is expressed in both neurons and intestine, and only its intestinal levels are dramatically increased in germline-less animals, suggesting the intestine as its functional site in mediating the germline signals (Goudeau et al. 2011). Interestingly, nhr-80 overexpression leads to further lifespan extension in the glp-1 mutants, but is not sufficient to prolong wild-type lifespan (Goudeau et al. 2011). Thus, activating ligands of NHR-80 are likely generated in the glp-1 mutants, which are absent in wild type.
On the other hand, nhr-49 expression is also upregulated in the glp-1 mutant and is required for its longevity (Ratnappan et al. 2014). NHR-49 and NHR-80 form a nuclear complex and regulate fatty acid desaturation coordinately (Pathare et al. 2012; Ratnappan et al. 2014). nhr-49 also controls the induction of a variety of genes involved in fatty acid beta-oxidation in the glp-1 mutant, including acs-2, acs-22, acdh-11, and hacd-1 (Ratnappan et al. 2014). Those fatty acid beta-oxidation genes are required for the longevity of the glp-1 mutant, suggesting an increased lipid catabolism in germline-less animals despite their increased fat content levels.
6.3 Autophagy
Autophagy is a cellular process where cells deliver unnecessary or dysfunctional cellular organelles and protein aggregates to the lysosomes for degradation and recycling. This process is crucial for a healthy turnover of cellular organelles and proteins and also produces energy from unwanted waste under nutrient starvation. Interestingly, autophagy has been revealed as a crucial link between lipid metabolism and germline-loss-associated longevity. Autophagy is induced in the germline-less animals, as seen by the accumulation of autophagic particles in the intestine and hypodermis (Lapierre et al. 2011). Consistently, several autophagy genes, like unc-51 (ULK1 homolog), bec-1 (Beclin1 homolog), and lgg-1 (LC3 homolog), are upregulated in the glp-1 mutant (Lapierre et al. 2011). Importantly, inactivation of those autophagy genes by RNAi specifically abrogates longevity of the glp-1 mutant, but does not affect wild-type lifespan (Lapierre et al. 2011). The induction of autophagy in the germline-less animals requires the activity of the FOXA forkhead transcription factor, PHA-4 (Lapierre et al. 2011). PHA-4 has been previously identified as required for dietary restriction-mediated longevity (Panowski et al. 2007). Adult-specific knockdown of pha-4 abrogates glp-1 longevity with minor shortening effects on wild-type lifespan (Goudeau et al. 2011). Autophagy has also been linked to lipid catabolism where it stimulates lipolysis in lysosomal compartments (lipophagy) (Singh et al. 2009). Interestingly, overexpression of the LIPL-4 lipase alone leads to increased autophagic particles in the hypodermal seam cells. Knockdown of pha-4 or of autophagy genes blocks the lifespan extension conferred by lipl-4 overexpression (Goudeau et al. 2011). These findings suggest a possible induction of lipophagy mechanisms following germline removal and its crucial roles in regulating longevity (Fig. 3.2).
How can lipid metabolism influence lifespan? Current hypotheses include lipophilic signaling, alteration of lipid composition or distribution, and lipotoxicity (Ackerman and Gems 2012). A study found that lipolytic products and intermediates participate in specific cellular signaling and can systemically influence the whole organism activity (Haemmerle et al. 2011), which unveils a previously underappreciated aspect of lipolysis. Interestingly, a lipolytic product OEA has been identified to activate NHR-49/NHR-80 nuclear hormone receptor complex and consequently promote longevity (Folick et al. 2015). Lipid composition of saturated versus unsaturated fat may decide oxidative stress tolerance capacity; the ratio of short- versus long-chain length, or the lipid turnover rate, may all affect fat profile contributed by healthy or unhealthy lipid (Shmookler Reis et al. 2011). Yet, more supporting biochemical evidence is required to back up this model. On the other hand, in the mammalian system, it has long been supposed that good fat or bad fat could also be defined by where it locates. Indeed, visceral fat or fat in non-adipocyte is considered deleterious and may cause metabolic syndromes while subcutaneous fat or adipocyte fat is recognized fine (Huffman and Barzilai 2009; Schaffer 2003). In C. elegans, yolk lipoprotein ectopic accumulation has been observed post-reproductively and may lead to age-related tissue dysfunction. Long-lived daf-2 mutants display lower yolk levels after reproduction; and RNAi knockdown of yolk genes extends worm lifespan (Murphy et al. 2003). Hence, ectopic accumulation of fat may lead to lipotoxicity and thus trigger aging. Up to now, it is still under debate which of these hypotheses is correct, notwithstanding that they might not be mutually exclusive with each other.
7 Perspectives
During evolution, the maximum reproductive success is the ultimate purpose of all organisms. In the wild, the animals that lack the germline will not continue living, so why does this germline-mediated regulatory mechanism exist? Interestingly, germline proliferation is under tight control of nutrient availability. Several nutrient-sensing pathways regulate germline proliferation and maintenance, such as IIS, TOR, and dietary restriction (Narbonne and Roy 2006). Under unfavorable environmental conditions, germline proliferation is expected to stall. This is carried out by several developmental and reproductive arrest checkpoints in C. elegans. In particular, C. elegans can enter into an adult reproductive diapause in response to starvation, during which only a few GSCs are maintained in the germline. Animals can survive in this diapause state for extremely long time until nutrient becomes available and will reconstitute the germline and resume full reproductive capacity. We could expect that the germline-mediated longevity mechanism might be responsible for the long-term survival under diapause. In the wild, these mechanisms could assist animals survive through harsh environment and ensure the maximum reproductive success.
In the past, lifespan extension sounded like an unrealistic dream. Through decades of efforts, scientists have demonstrated that lifespan can be prolonged via various genetic mechanisms. Nevertheless, there are still many questions in the field of germline-regulated longevity. Loss of germ cells drives signaling responses in neurons, intestine, and somatic gonads that lead to enhanced somatic maintenance and healthy aging. How do different tissues crosstalk with each other? The lipid metabolism in intestine is altered in germline-less animals. What are the intestinal receptors interacting with the germline-less signals? What are the downstream mechanisms of lipid metabolism that directly affect lifespan? Somatic gonad regulates lifespan via daf-12 lipophilic signaling. How does daf-2 IIS signaling pathway intervene in daf-12 signaling? All these interesting questions (and many more to come) remain to be answered. There is always more knowledge of aging waiting for us to be explored and amazed.
References
Ackerman D, Gems D (2012) The mystery of C. elegans aging: an emerging role for fat. Distant parallels between C. elegans aging and metabolic syndrome? Bioessays 34:466–471
Anselmi CV, Malovini A, Roncarati R, Novelli V, Villa F, Condorelli G, Bellazzi R, Puca AA (2009) Association of the FOXO3A locus with extreme longevity in a southern Italian centenarian study. Rejuvenation Res 12:95–104
Arantes-Oliveira N, Apfeld J, Dillin A, Kenyon C (2002) Regulation of life-span by germ-line stem cells in Caenorhabditis elegans. Science 295:502–505
Ashrafi K (2007) Obesity and the regulation of fat metabolism. WormBook, ed. The C. elegans Research Community, WormBook 1–20, doi:10.1895/wormbook.1.130.1, http://www.wormbook.org
Beckstead RB, Thummel CS (2006) Indicted: worms caught using steroids. Cell 124:1137–1140
Berman JR, Kenyon C (2006) Germ-cell loss extends C. elegans life span through regulation of DAF-16 by kri-1 and lipophilic-hormone signaling. Cell 124:1055–1068
Berry LW, Westlund B, Schedl T (1997) Germ-line tumor formation caused by activation of glp-1, a Caenorhabditis elegans member of the Notch family of receptors. Development 124:925–936
Blagosklonny MV, Campisi J, Sinclair DA, Bartke A, Blasco MA, Bonner WM, Bohr VA, Brosh RM Jr, Brunet A, Depinho RA et al (2010) Impact papers on aging in 2009. Aging (Albany NY) 2:111–121
Boulias K, Horvitz HR (2012) The C. elegans microRNA mir-71 acts in neurons to promote germline-mediated longevity through regulation of DAF-16/FOXO. Cell Metab 15:439–450
Brock TJ, Browse J, Watts JL (2006) Genetic regulation of unsaturated fatty acid composition in C. elegans. PLoS Genet 2, e108
Broue F, Liere P, Kenyon C, Baulieu EE (2007) A steroid hormone that extends the lifespan of Caenorhabditis elegans. Aging Cell 6:87–94
Byrd DT, Kimble J (2009) Scratching the niche that controls Caenorhabditis elegans germline stem cells. Semin Cell Dev Biol 20:1107–1113
C. elegans Sequencing Consortium (1998) Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282:2012–2018
Cargill SL, Carey JR, Muller HG, Anderson G (2003) Age of ovary determines remaining life expectancy in old ovariectomized mice. Aging Cell 2:185–190
Carty SM, Goldstrohm AC, Sune C, Garcia-Blanco MA, Greenleaf AL (2000) Protein-interaction modules that organize nuclear function: FF domains of CA150 bind the phosphoCTD of RNA polymerase II. Proc Natl Acad Sci U S A 97:9015–9020
Crittenden SL, Troemel ER, Evans TC, Kimble J (1994) GLP-1 is localized to the mitotic region of the C. elegans germ line. Development 120:2901–2911
Crittenden SL, Bernstein DS, Bachorik JL, Thompson BE, Gallegos M, Petcherski AG, Moulder G, Barstead R, Wickens M, Kimble J (2002) A conserved RNA-binding protein controls germline stem cells in Caenorhabditis elegans. Nature 417:660–663
Dillin A, Crawford DK, Kenyon C (2002) Timing requirements for insulin/IGF-1 signaling in C. elegans. Science 298:830–834
Eckmann CR, Kraemer B, Wickens M, Kimble J (2002) GLD-3, a bicaudal-C homolog that inhibits FBF to control germline sex determination in C. elegans. Dev Cell 3:697–710
Eckmann CR, Crittenden SL, Suh N, Kimble J (2004) GLD-3 and control of the mitosis/meiosis decision in the germline of Caenorhabditis elegans. Genetics 168:147–160
Fitzgerald K, Greenwald I (1995) Interchangeability of Caenorhabditis elegans DSL proteins and intrinsic signalling activity of their extracellular domains in vivo. Development 121:4275–4282
Flachsbart F, Caliebe A, Kleindorp R, Blanche H, von Eller-Eberstein H, Nikolaus S, Schreiber S, Nebel A (2009) Association of FOXO3A variation with human longevity confirmed in German centenarians. Proc Natl Acad Sci U S A 106:2700–2705
Flatt T, Min KJ, D’Alterio C, Villa-Cuesta E, Cumbers J, Lehmann R, Jones DL, Tatar M (2008) Drosophila germ-line modulation of insulin signaling and lifespan. Proc Natl Acad Sci U S A 105:6368–6373
Folick A, Oakley HD, Yu Y, Armstrong EH, Kumari M, Sanor L, Moore DD, Ortlund EA, Zechner R, Wang MC (2015) Aging. Lysosomal signaling molecules regulate longevity in Caenorhabditis elegans. Science 347:83–86
Friedman DB, Johnson TE (1988) A mutation in the age-1 gene in Caenorhabditis elegans lengthens life and reduces hermaphrodite fertility. Genetics 118:75–86
Gems D, Sutton AJ, Sundermeyer ML, Albert PS, King KV, Edgley ML, Larsen PL, Riddle DL (1998) Two pleiotropic classes of daf-2 mutation affect larval arrest, adult behavior, reproduction and longevity in Caenorhabditis elegans. Genetics 150:129–155
Gerisch B, Weitzel C, Kober-Eisermann C, Rottiers V, Antebi A (2001) A hormonal signaling pathway influencing C. elegans metabolism, reproductive development, and life span. Dev Cell 1:841–851
Gerisch B, Rottiers V, Li D, Motola DL, Cummins CL, Lehrach H, Mangelsdorf DJ, Antebi A (2007) A bile acid-like steroid modulates Caenorhabditis elegans lifespan through nuclear receptor signaling. Proc Natl Acad Sci U S A 104:5014–5019
Ghazi A, Henis-Korenblit S, Kenyon C (2009) A transcription elongation factor that links signals from the reproductive system to lifespan extension in Caenorhabditis elegans. PLoS Genet 5, e1000639
Goldstrohm AC, Albrecht TR, Sune C, Bedford MT, Garcia-Blanco MA (2001) The transcription elongation factor CA150 interacts with RNA polymerase II and the pre-mRNA splicing factor SF1. Mol Cell Biol 21:7617–7628
Goudeau J, Bellemin S, Toselli-Mollereau E, Shamalnasab M, Chen Y, Aguilaniu H (2011) Fatty acid desaturation links germ cell loss to longevity through NHR-80/HNF4 in C. elegans. PLoS Biol 9:e1000599
Greenspan P, Fowler SD (1985) Spectrofluorometric studies of the lipid probe, nile red. J Lipid Res 26:781–789
Haemmerle G, Moustafa T, Woelkart G, Buttner S, Schmidt A, van de Weijer T, Hesselink M, Jaeger D, Kienesberger PC, Zierler K et al (2011) ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-alpha and PGC-1. Nat Med 17:1076–1085
Hansen D, Wilson-Berry L, Dang T, Schedl T (2004) Control of the proliferation versus meiotic development decision in the C. elegans germline through regulation of GLD-1 protein accumulation. Development 131:93–104
Henderson ST, Johnson TE (2001) daf-16 integrates developmental and environmental inputs to mediate aging in the nematode Caenorhabditis elegans. Curr Biol 11:1975–1980
Henderson ST, Gao D, Lambie EJ, Kimble J (1994) lag-2 may encode a signaling ligand for the GLP-1 and LIN-12 receptors of C. elegans. Development 120:2913–2924
Hsin H, Kenyon C (1999) Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399:362–366
Hubbard EJA, Greenstein D (2005) Introduction to the germ line. WormBook, ed. The C. elegans Research Community, WormBook 1–4, doi:10.1895/wormbook.1.18.1, http://www.wormbook.org
Huffman DM, Barzilai N (2009) Role of visceral adipose tissue in aging. Biochim Biophys Acta 1790:1117–1123
Johnson TE, Tedesco PM, Lithgow GJ (1993) Comparing mutants, selective breeding, and transgenics in the dissection of aging processes of Caenorhabditis elegans. Genetica 91:65–77
Kenyon C (2010a) A pathway that links reproductive status to lifespan in Caenorhabditis elegans. Ann N Y Acad Sci 1204:156–162
Kenyon CJ (2010b) The genetics of ageing. Nature 464:504–512
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366:461–464
Kidd AR 3rd, Miskowski JA, Siegfried KR, Sawa H, Kimble J (2005) A beta-catenin identified by functional rather than sequence criteria and its role in Wnt/MAPK signaling. Cell 121:761–772
Kimble J, Crittenden SL (2005) Germline proliferation and its control. WormBook, ed. The C. elegans Research Community, WormBook 1–14, doi:10.1895/wormbook.1.13.1, http://www.wormbook.org
Kimble JE, White JG (1981) On the control of germ cell development in Caenorhabditis elegans. Dev Biol 81:208–219
Kimura KD, Tissenbaum HA, Liu Y, Ruvkun G (1997) daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science 277:942–946
Kipreos ET, Gohel SP, Hedgecock EM (2000) The C. elegans F-box/WD-repeat protein LIN-23 functions to limit cell division during development. Development 127:5071–5082
Kojima T, Kamei H, Aizu T, Arai Y, Takayama M, Nakazawa S, Ebihara Y, Inagaki H, Masui Y, Gondo Y et al (2004) Association analysis between longevity in the Japanese population and polymorphic variants of genes involved in insulin and insulin-like growth factor 1 signaling pathways. Exp Gerontol 39:1595–1598
Lam N, Chesney MA, Kimble J (2006) Wnt signaling and CEH-22/tinman/Nkx2.5 specify a stem cell niche in C. elegans. Curr Biol 16:287–295
Lamont LB, Crittenden SL, Bernstein D, Wickens M, Kimble J (2004) FBF-1 and FBF-2 regulate the size of the mitotic region in the C. elegans germline. Dev Cell 7:697–707
Lapierre LR, Gelino S, Melendez A, Hansen M (2011) Autophagy and lipid metabolism coordinately modulate life span in germline-less C. elegans. Curr Biol 21:1507–1514
Lee SS, Schroeder FC (2012) Steroids as central regulators of organismal development and lifespan. PLoS Biol 10, e1001307
Lee RY, Hench J, Ruvkun G (2001) Regulation of C. elegans DAF-16 and its human ortholog FKHRL1 by the daf-2 insulin-like signaling pathway. Curr Biol 11:1950–1957
Li L, Xie T (2005) Stem cell niche: structure and function. Annu Rev Cell Dev Biol 21:605–631
Li J, Tewari M, Vidal M, Lee SS (2007) The 14-3-3 protein FTT-2 regulates DAF-16 in Caenorhabditis elegans. Dev Biol 301:82–91
Libina N, Berman JR, Kenyon C (2003) Tissue-specific activities of C. elegans DAF-16 in the regulation of lifespan. Cell 115:489–502
Lin K, Hsin H, Libina N, Kenyon C (2001) Regulation of the Caenorhabditis elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat Genet 28:139–145
Lin KT, Lu RM, Tarn WY (2004) The WW domain-containing proteins interact with the early spliceosome and participate in pre-mRNA splicing in vivo. Mol Cell Biol 24:9176–9185
Longo VD, Mitteldorf J, Skulachev VP (2005) Programmed and altruistic ageing. Nat Rev Genet 6:866–872
McCormick M, Chen K, Ramaswamy P, Kenyon C (2012) New genes that extend Caenorhabditis elegans’ lifespan in response to reproductive signals. Aging Cell 11:192–202
Murphy CT, McCarroll SA, Bargmann CI, Fraser A, Kamath RS, Ahringer J, Li H, Kenyon C (2003) Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424:277–283
Narbonne P, Roy R (2006) Regulation of germline stem cell proliferation downstream of nutrient sensing. Cell Div 1:29
Ogg S, Paradis S, Gottlieb S, Patterson GI, Lee L, Tissenbaum HA, Ruvkun G (1997) The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389:994–999
Panowski SH, Dillin A (2009) Signals of youth: endocrine regulation of aging in Caenorhabditis elegans. Trends Endocrinol Metab 20:259–264
Panowski SH, Wolff S, Aguilaniu H, Durieux J, Dillin A (2007) PHA-4/Foxa mediates diet-restriction-induced longevity of C. elegans. Nature 447:550–555
Paradis S, Ruvkun G (1998) Caenorhabditis elegans Akt/PKB transduces insulin receptor-like signals from AGE-1 PI3 kinase to the DAF-16 transcription factor. Genes Dev 12:2488–2498
Paradis S, Ailion M, Toker A, Thomas JH, Ruvkun G (1999) A PDK1 homolog is necessary and sufficient to transduce AGE-1 PI3 kinase signals that regulate diapause in Caenorhabditis elegans. Genes Dev 13:1438–1452
Pathare PP, Lin A, Bornfeldt KE, Taubert S, Van Gilst MR (2012) Coordinate regulation of lipid metabolism by novel nuclear receptor partnerships. PLoS Genet 8, e1002645
Pawlikowska L, Hu D, Huntsman S, Sung A, Chu C, Chen J, Joyner AH, Schork NJ, Hsueh WC, Reiner AP et al (2009) Association of common genetic variation in the insulin/IGF1 signaling pathway with human longevity. Aging Cell 8:460–472
Pearson JL, Robinson TJ, Munoz MJ, Kornblihtt AR, Garcia-Blanco MA (2008) Identification of the cellular targets of the transcription factor TCERG1 reveals a prevalent role in mRNA processing. J Biol Chem 283:7949–7961
Ratnappan R, Amrit FR, Chen SW, Gill H, Holden K, Ward J, Yamamoto KR, Olsen CP, Ghazi A (2014) Germline signals deploy NHR-49 to modulate fatty-acid beta-oxidation and desaturation in somatic tissues of C. elegans. PLoS Genet 10:e1004829
Russell SJ, Kahn CR (2007) Endocrine regulation of ageing. Nat Rev Mol Cell Biol 8:681–691
Samuelson AV, Klimczak RR, Thompson DB, Carr CE, Ruvkun G (2007) Identification of Caenorhabditis elegans genes regulating longevity using enhanced RNAi-sensitive strains. Cold Spring Harb Symp Quant Biol 72:489–497
Schaffer JE (2003) Lipotoxicity: when tissues overeat. Curr Opin Lipidol 14:281–287
Shmookler Reis RJ, Xu L, Lee H, Chae M, Thaden JJ, Bharill P, Tazearslan C, Siegel E, Alla R, Zimniak P et al (2011) Modulation of lipid biosynthesis contributes to stress resistance and longevity of C. elegans mutants. Aging (Albany NY) 3:125–147
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M, Tanaka K, Cuervo AM, Czaja MJ (2009) Autophagy regulates lipid metabolism. Nature 458:1131–1135
Smith MJ, Kulkarni S, Pawson T (2004) FF domains of CA150 bind transcription and splicing factors through multiple weak interactions. Mol Cell Biol 24:9274–9285
Strome S (2005) Specification of the germ line. WormBook, ed. The C. elegans Research Community, WormBook 1–10, doi:10.1895/wormbook.1.9.1, http://www.wormbook.org
Suh Y, Atzmon G, Cho MO, Hwang D, Liu B, Leahy DJ, Barzilai N, Cohen P (2008) Functionally significant insulin-like growth factor I receptor mutations in centenarians. Proc Natl Acad Sci U S A 105:3438–3442
Sulston JE, Schierenberg E, White JG, Thomson JN (1983) The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol 100:64–119
Sune C, Garcia-Blanco MA (1999) Transcriptional cofactor CA150 regulates RNA polymerase II elongation in a TATA-box-dependent manner. Mol Cell Biol 19:4719–4728
Tissenbaum HA, Ruvkun G (1998) An insulin-like signaling pathway affects both longevity and reproduction in Caenorhabditis elegans. Genetics 148:703–717
Updike D, Strome S (2010) P granule assembly and function in Caenorhabditis elegans germ cells. J Androl 31:53–60
Wang L, Eckmann CR, Kadyk LC, Wickens M, Kimble J (2002) A regulatory cytoplasmic poly(A) polymerase in Caenorhabditis elegans. Nature 419:312–316
Wang MC, O’Rourke EJ, Ruvkun G (2008) Fat metabolism links germline stem cells and longevity in C. elegans. Science 322:957–960
Watts JL, Browse J (2002) Genetic dissection of polyunsaturated fatty acid synthesis in Caenorhabditis elegans. Proc Natl Acad Sci U S A 99:5854–5859
Willcox BJ, Donlon TA, He Q, Chen R, Grove JS, Yano K, Masaki KH, Willcox DC, Rodriguez B, Curb JD (2008) FOXO3A genotype is strongly associated with human longevity. Proc Natl Acad Sci U S A 105:13987–13992
Wollam J, Magner DB, Magomedova L, Rass E, Shen Y, Rottiers V, Habermann B, Cummins CL, Antebi A (2012) A novel 3-hydroxysteroid dehydrogenase that regulates reproductive development and longevity. PLoS Biol 10, e1001305
Xie T (2008) Germline stem cell niches. StemBook, ed. The Stem Cell Research Community, StemBook, doi:10.3824/stembook.1.23.1, http://www.stembook.org
Yamawaki TM, Arantes-Oliveira N, Berman JR, Zhang P, Kenyon C (2008) Distinct activities of the germline and somatic reproductive tissues in the regulation of Caenorhabditis elegans’ longevity. Genetics 178:513–526
Yamawaki TM, Berman JR, Suchanek-Kavipurapu M, McCormick M, Gaglia MM, et al (2010) The Somatic Reproductive Tissues of C. elegans Promote Longevity through Steroid Hormone Signaling. PLoS Biol 8(8): e1000468. doi:10.1371/journal.pbio.1000468
Yokota S, Togo SH, Maebuchi M, Bun-Ya M, Haraguchi CM, Kamiryo T (2002) Peroxisomes of the nematode Caenorhabditis elegans: distribution and morphological characteristics. Histochem Cell Biol 118:329–336
Acknowledgment
We would like to acknowledge all the scientists in the C. elegans research community for their contributions to the field of reproductive system-regulated aging. We also thank Cheng-lin Li and Timothy R. Mahoney for helpful comments and valuable advice on the manuscript. The work in the authors’ laboratory is supported by the National Institutes of Health grant RO0 AG034988 and RO1 AG045183 and the Ellison Medical Foundation grant AG-NS-0756-11.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag Wien
About this chapter
Cite this chapter
Lin, Cc.J., Wang, M.C. (2015). Germline Stem Cells and Their Roles in the Regulation of Organism Longevity. In: Geiger, H., Jasper, H., Florian, M. (eds) Stem Cell Aging: Mechanisms, Consequences, Rejuvenation. Springer, Vienna. https://doi.org/10.1007/978-3-7091-1232-8_3
Download citation
DOI: https://doi.org/10.1007/978-3-7091-1232-8_3
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-1231-1
Online ISBN: 978-3-7091-1232-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)