Abstract
Historically, research has focused on the detrimental effects of ageing on fertility, but studies in the last two decades have shown that reproductive status profoundly impacts the length and quality of life. The nematode C. elegans has been at the forefront of these discoveries that have led to a fundamental transformation in our understanding of the relationship between procreation and lifespan in metazoans. In C. elegans, removing a population of proliferative germline-stem cells (GSCs) confers long life and enhances stress resilience. Germline loss is a major physiological challenge that compels the animal to arrest reproduction and reorganize its metabolic profile, so the phenomenon also provides a unique platform to understand how complex metazoans cope with changes in fertility and age. Recent studies have shown that GSC depletion triggers the activation of a group of conserved transcription factors in somatic cells. The transcriptional changes orchestrated by these proteins alter lipid metabolism, proteasomal function, autophagy and stress resistance, events that likely facilitate the adaptation to germline loss and lead to improved health and longevity. Here, we review the current literature on this longevity paradigm and the contributions made by C. elegans to understanding the molecular basis of the reproductive control of ageing.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
6.1 Introduction
As animals grow older, their somatic tissues undergo deterioration that is manifested as symptoms of ageing . However, the germline is passed on to the next generation in an immaculate condition for the maintenance of the species. This dichotomy between the preservation of the two tissues underscores the complex association between reproduction and ageing . This relationship is fascinating because both are such central, intimately linked aspects of an animal’s life history. Traditionally, research has focused on the lamentable, but well-documented, reduction in fertility associated with increasing maternal age. But, in recent years, there has been emerging evidence that the germline and soma exchange signals that help coordinate the health of both tissues. In this article, we provide an overview of the existing knowledge on how signals emanating from the proliferating germline influence the rate of somatic ageing in C. elegans . Studies on ageing of the reproductive system are addressed in Chap. 7.
The relationship between reproduction and ageing has proven to be intractable partly because it is rife with paradoxical observations. The mating process is known to have a negative impact on the lifespan and healthspan of females, not only due to the physical damage caused by copulation but also by the presence of mortality-inducing chemicals in the males’ seminal fluid. In Drosophila, males transfer peptides along with sperm that reduce female lifespan [1], whereas, in many Caenorhabditis species, mating causes hermaphrodites and females to undergo shrinkage [2] and reduces stress resistance and lifespan [3]. Similar observations have been made in other organisms [4, 5]. However, in numerous other species, especially arthropods, males are known to donate edible or glandular products during copulation that provide direct benefits to the female. These ‘nuptial gifts’ can range from simple food and energy supplies to seminal secretions that are immunoprotective to the recipient and its progeny, and even extend the females’ lifespan [6, 7]. Even in C. elegans , the deleterious impact of male pheromones on lifespan is accompanied with enhanced thermotolerance in the hermaphrodite [8]. Such paradoxes illustrate the complexity of the mutual interactions between germline and somatic health and have made it difficult to arrive at simplistic principles about the relationship between reproduction and ageing . This is particularly applicable to the effect that sterility has on the lifespan and healthspan of animals.
Correlative studies in many animal and some plant species have found sterility to be associated with increased lifespan [9–11], and these have informed the ‘Somatic Maintenance’ or ‘Disposable Soma’ theory of ageing [12]. It is founded on the consideration that reproduction is an energy intensive process that consumes cellular resources that could otherwise be devoted towards somatic repair and maintenance. This ‘trade off’ is postulated to be fundamental for the survival of the species but detrimental to the individual by increasing post-reproductive mortality. However, recent data negate this simplistic interpretation. Field studies on thirty mammalian and bird species showed either no correlation or even a positive correlation between fertility and longevity [13]. While some evaluations of human genealogical data have supported a ‘trade off’ phenomenon [14], there have been many more that have either found no association or detected positive correlations between fertility and lifespan [15]. In fact, both contemporary and historical data from European populations have revealed significant positive correlation in this relationship in both men and women [16–18]. Overall, observational human and animal studies provide a growing body of evidence that fails to support the theoretical notion of a cost of reproduction in fertile animals. Instead, recent experimental approaches in laboratory animals have suggested a complex, nuanced relationship between reproduction and ageing [9, 19]. In particular, studies in C. elegans have revealed a wealth of knowledge on the molecular underpinnings of this association, identifying numerous genes , cellular processes and metabolic pathways that are altered in response to germline removal and that consequently impact the rate of ageing.
6.2 The Influence of Germline Stem Cells on Ageing of Somatic Tissues
One of the first studies demonstrating a direct molecular-genetic link between germline status and longevity came from the observation that removing the germline increased C. elegans lifespan by ~60 % [20]. The worm was an excellent platform to explore the details of this phenomenon because of its invariant lineage and transparent body. Using temperature sensitive mutants of a gene, glp-1 , (essential for germline proliferation) [21] that lose select germ-cell types when exposed to non-permissive temperatures at different developmental stages, the Kenyon lab elegantly demonstrated that simply making the worms sterile did not confer longevity. Instead, removal of the germline , in particular a population of proliferating, totipotent germline stem cells (GSCs), while retaining the somatic structures required for reproduction (such as the uterus and spermatheca, collectively called the ‘somatic gonad’), was the key for extending the animal’s lifespan [22]. glp-1 mutants lived longer and were active for a greater fraction of life, and this enhancement was completely dependent on the presence of known lifespan-regulatory genes [22]. These observations undermined the ‘trade off’ hypothesis and opened up the possibility that signals from the germline may act on specific somatic tissues to influence overall organismal ageing. Shortly thereafter, eliminating GSCs in Drosophila melanogaster was found to cause a similar life lengthening, dependent on homologues of worm proteins needed for GSC-less longevity [23]. Subsequent studies have shown that transplanting ovaries of young mice into older ones increases their lifespan and reduces susceptibility to cardiovascular disease (CVD) observed in post-menopausal female mammals [24, 25]. Together, these studies provide compelling support to the worm data and suggest that the reproductive control of ageing is not unique to nematodes or invertebrates but a conserved aspect of the relationship between germline and somatic tissues of metazoans .
6.3 A Network of Transcription Factors Mediates Germline-Less Longevity
6.3.1 DAF-16/FOXO3A and TCER-1/TCERG1
The discovery, that reproductive signals reflective of the procreative status of the animal modulate lifespan, led to the inevitable search for genes involved in this soma-germline dialogue. Early observations suggested that GSC removal produces widespread transcriptional changes in the animal because two of the genes essential for longevity, daf-16 and daf-12, encode transcription factors [20]. DAF-16 is the worm homologue of FOXO3A and is the main pro-longevity factor repressed by insulin/IGF1 signalling (IIS), the conserved and most well known lifespan-regulatory pathway [26]. A detailed discussion of the role of IIS in ageing can be found in Chap. 4. DAF-16/FOXO3A involvement initially indicated that reproductive signals modulate IIS to alter ageing. But, DAF-16/FOXO3A relocates to the nuclei of intestinal cells in GSC-less young adults, whereas, upon reduced IIS, nuclear localization occurs in many tissues [27]. Similarly, GSC-less worms expressing DAF-16/FOXO3A only in the intestine undergo lifespan extension to the same extent as worms that have DAF-16/FOXO3A in all tissues, but the intestinal protein is not sufficient to increase lifespan in IIS mutants [28]. Other such observations suggested that reproductive cues and IIS are independent physiological stimuli that share DAF-16/FOXO3A as a downstream effector.
The discovery of two genes, kri-1 and tcer-1, that selectively enhance the longevity of glp-1 mutants was especially instrumental in consolidating this premise. kri-1, encoding an Ankyrin-repeat containing protein homologous to the human disease gene KRIT1/CCM1, is expressed only in gut cells and stimulates the nuclear localization of DAF-16/FOXO3A [29]. It also mediates the transcriptional upregulation of TCER-1/TCERG1 [30] (Fig. 6.1). We identified tcer-1, encoding the worm homologue of a human transcription elongation and splicing factor , TCERG1, in a screen designed to isolate genes essential for lifespan extension following GSC loss [30]. In exploring the role of TCER-1/TCERG1, it became apparent that DAF-16/FOXO3A regulated overlapping but distinct targets in GSC-less animals and IIS mutants, and that TCER-1/TCERG1 increased lifespan specifically following germline loss by facilitating a distinct pattern of DAF-16-dependent gene expression. TCER-1/TCERG1 is widely expressed in nuclei of somatic tissues and its expression is elevated following germline loss in intestinal cells and (unlike DAF-16/FOXO3A) in neurons . Elevating TCER-1/TCERG1 in the soma of fertile animals augments their lifespan without loss of fertility and is accompanied with increased expression of DAF-16/FOXO3A target genes [30]. This is an important discovery because it implies that TCER-1/TCERG1 serves as a switch that connects germline signals to the activity of a broadly deployed transcription factor such as DAF-16/FOXO3A, and because it opens up the possibility that health benefits accrued by GSC removal can be obtained by activating this pathway without loss of fertility. Recent experiments, including ours, with other pro-longevity genes acting in this pathway (discussed below) have further substantiated this beguiling possibility .
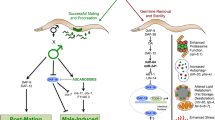
Transcription factors activated by GSC removal in C. elegans’ intestinal cells and cellular processes modulated by them. Proteins undergoing nuclear relocation (DAF-16/FOXO3A , SKN-1/NRF2 , HLH-30/TFEB and MML-1) are shown on membrane of, and within, the nucleus; upward arrow next to proteins indicates transcriptional upregulation upon GSC loss (TCER-1/TCERG1, PHA-4 /FOXA, NHR-80/HNF4 and NHR-49/PPARα). DAF-16/FOXO3A nuclear localization is governed by multiple inputs including the dafachronic-acid (DA) cascade (featuring DAF-12/VDR and its miRNA targets), a neuronal miRNA, mIR71, and KRI-1/KRIT1. KRI-1 also enhances TCER-1/TCERG1 transcription and SKN-1/NRF2 nuclear entry. NHR-80/HNF4 upregulation is controlled by DAF-12/VDR and, in part, by DAF-16/FOXO3A. NHR-49/PPARα upregulation is partially triggered by DAF-16/FOXO3A and TCER-1/TCERG1. NHR-49/PPARα participates in a positive feed-back loop, possibly in collaboration with NHR-71/HNF4, to potentiate DAF-16/TCER-1 activity by altering the subcellular localization of KRI-1/KRIT1. The main cellular processes modulated by these factors include lipid metabolism , autophagy and protein homeostasis . DAF-16/FOXO3A acts with TCER-1/TCERG1 to elevate both lipid-synthetic and lipid-degradative pathways. SKN-1/NRF2 shares the regulation of some of these processes. NHR-49/PPARα (likely in cooperation with MDT-15) stimulates β-oxidation and fatty-acid desaturation, whereas, NHR-80/HNF4 promotes fatty-acid desaturation alone. SKN-1/NRF2 and DAF-16/FOXO3A enhance proteasomal activity, while autophagy is augmented by PHA-4/FOXA, HLH-30/TFEB and the MML-1/MXL-2 complex. Improved heat- and oxidative stress resistance is mediated by HSF-1/HSF, SKN-1/NRF2 and, partly, DAF-16/FOXO3A
6.3.2 DAF-12/VDR, microRNAs and Steroid Signalling
DAF-16/FOXO3A intestinal nuclear localization in glp-1 mutants is also governed by DAF-12 [29], a nuclear hormone receptor (NHR) homologous to vertebrate vitamin D receptor (VDR) [31]. DAF-12/VDR, similar to other NHRs, responds to lipophilic hormones and steroids to modulate gene expression, including the expression of regulatory microRNAs (miRNAs). The Antebi lab has shown that DAF-12/VDR influences the choice between normal growth and diapause during larval development through binding and activation by 3-keto bile acid-like steroid ligands called Δ4 and Δ7 dafachronic acids (DAs) [32]. In GSC-less worms, DAs activate DAF-12/VDR which in turn promotes DAF-16/FOXO3A nuclear localization [33] by increasing the levels of (at least) two miRNAs, mIR-84 and mIR-241 [34]. Both were reported to be upregulated by DAF-12/VDR in intestinal and epidermal tissues upon GSC ablation and were redundantly required for DAF-16/FOXO3A nuclear localization and glp-1 mutants’ longevity . Genetic evidence suggests that mIR84 and mIR241 promote DAF-16/FOXO3A nuclear traffic by repressing the expression of (at least) two known anti-longevity proteins [34]. One of these, SGK-1, is a kinase that phosphorylates DAF-16/FOXO3A to inhibit nuclear entry [35]. The second, LIN-14, a developmental timing protein has also been previously shown to limit lifespan by repressing DAF-16/FOXO3A [36]. In contrast, DAF-12/VDR exerts no influence on mIR-71, another pro-longevity miRNA shown by the Horvitz lab to act in neuronal cells to augment DAF-16/FOXO3A’s intestinal nuclear passage [37]. It remains to be seen if additional miRNAs , regulated by DAF-12/VDR or not, have roles in this longevity pathway.
In a series of exhaustive studies, the Antebi lab also identified multiple genes of the biosynthetic pathway responsible for DA production, including daf-9 (encodes a cytochrome P450 enzyme) [38], daf-36 (encodes a Rieske-like oxygenase enzyme) [39] and dhs-16 (encodes a 3-hydroxysteroid dehydrogenase) [40] (Fig. 6.1). Expectedly, each of these genes is essential for lifespan extension upon GSC ablation, and supplementation of DA in GSC-less daf-9 and daf-36 (but not daf-12) mutants extends lifespan [33]. Interestingly, worms lacking both the germline and the somatic gonad, that would otherwise exhibit wild-type lifespan, live significantly longer upon DA supplementation [41]. This observation raises the possibility that the somatic gonad may be the site of DA production following GSC removal, and may partly explain the importance of the organ in GSC-less longevity. However, DA levels between wild-type worms and glp-1 mutants have been reported to be similar, although the study examined whole worms and would not have detected localized changes in levels [42]. daf-12 mutants expressing a nucleus-restricted version of DAF-16/FOXO3A selectively in intestinal cells do not undergo life lengthening suggesting that, besides regulating nuclear relocation, DAF-12/VDR is also required for DAF-16/FOXO3A’s transcriptional activity [29]. Unlike the well-characterized role of DAF-12/VDR in modulating DAF-16/FOXO3A sub-cellular traffic, how it influences the latter’s activity within the nucleus is unknown.
6.3.3 NHR-49/PPARα, NHR-80/HNF4 and NHR-71/HNF4
Besides DAF-12/VDR, the ‘NHR’ gene family in C. elegans includes ~284 members. The Aguilaniu lab showed that one of these, NHR-80, an ortholog of mammalian hepatocyte nuclear factor 4 (HNF4), is essential for glp-1 mutants’ longevity [43]. Similar to TCER-1/TCERG1, NHR-80/HNF4 specifically promotes GSC-less longevity, is transcriptionally upregulated upon GSC removal in a DAF-16/FOXO3A-independent manner, and on overexpression increases lifespan of fertile animals. However, unlike TCER-1/TCERG1, NHR-80/HNF4 expression is enhanced only in intestinal cells of glp-1 mutants (and not in the neurons) by DAF-12/VDR [30, 43].
Recently, we identified a group of 13 NHRs that are essential for GSC-less longevity [44]. NHR-80 was included in this group, but the most striking phenotypes were produced by NHR-49. NHR-49 is structurally similar to HNF4, but it performs the functions undertaken by Peroxisome Proliferator-Activated Receptor alpha (PPARα), a key regulator of energy metabolism in vertebrates [45–47]. In literature, NHR-49 is referred to as the functional homologue of PPARα [48–50], and we use the same nomenclature in this article for consistency, although the homology remains to be demonstrated conclusively. NHR-49/PPARα is critical for glp-1 mutants’ long life as well as the lifespan of fertile worms, but has no role in the lifespan extension of IIS mutants [44], similar to NHR-80/HNF4 and TCER-1/TCERG1. Both mRNA and protein levels of NHR-49/PPARα are up-regulated upon GSC loss, partially dependent upon DAF-16/FOXO3A and TCER-1/TCERG1 (Fig. 6.1). Overexpressing the protein in fertile animals lengthens their life significantly without concomitant loss of fertility [44]. NHR-49/PPARα also appears to operate in a positive feed-back loop to potentiate the activity of DAF-16/FOXO3A and TCER-1/TCERG1, likely by controlling their common upstream regulator KRI-1/KRIT1, because, in nhr-49; glp-1 mutants, KRI-1/KRIT1 undergoes relocation to intestinal membranes, especially the apical surface bordering the gut lumen [51]. In contrast, glp-1 mutants display largely diffused KRI-1/KRIT1 expression. NHR-49/PPARα may collaborate with NHR-71/HNF4, one of the pro-longevity NHRs we identified, in mediating this positive feedback because inactivating either factor prevents the upregulation of DAF-16/FOXO3A and TCER-1/TCERG1 targets [51].
6.3.4 PHA-4 /FOXA, HLH-30/TFEB, MML-1 and MXL-2
The forkhead box, or FOX, gene family includes transcription factors belonging to subfamilies ranging from FoxA to FoxP. While DAF-16 is a member of the FoxO sub-family, PHA-4 represents the FoxA branch and is orthologous to genes encoding mammalian FOXA1, FOXA2 and FOXA3 proteins. PHA-4/FOXA was initially reported to be critical only for worms that are long lived due to reduced food intake or dietary restriction (DR), a paradigm that also extends lifespan in a variety of metazoan species [52, 53]. However, this study did not examine GSC-less mutants and in a subsequent report from the Hansen lab, PHA-4 was shown to be essential for GSC-less longevity as well [54]. PHA-4 mRNA levels rise upon GSC-removal, independent of DAF-16/FOXO3A . Although the tissues where this upregulation is orchestrated are not known, it is brought about by repression of the nutrient sensing kinase, target of rapamycin (TOR ) , and results in increased expression of multiple autophagy genes [54].
The Hansen lab identified another autophagy-promoting transcription factor HLH-30, with homology to the mammalian transcription factor EB (TFEB), as being essential for this longevity paradigm [55]. TFEB is a key regulator of mammalian autophagy and translocates to the nucleus upon nutrient deprivation to increase the transcription of autophagy genes [56]. HLH-30/TFEB also exhibits nuclear translocation in glp-1 mutants. Similar to PHA-4/FOXA, HLH-30 up-regulates the expression of a host of autophagy genes, and it’s overexpression increases the lifespan of fertile animals too [55]. Unlike PHA-4/FOXA that is transcriptionally upregulated in glp-1 mutants, HLH-30/TFEB undergoes increased nuclear traffic, although both events are precipitated by TOR downregulation [54, 55]. While PHA-4/FOXA is critical for GSC-less and DR-mediated longevity, HLH-30 promotes lifespan extension in multiple longevity pathways, including IIS signalling [55]. HLH-30/TFEB and PHA-4/FOXA co-regulate a large number of the autophagy-related genes [54, 55], but genome-scale studies are needed to define the extent and nature of overlap between these proteins.
HLH-30/TFEB has been shown to share similarities with two members of the Mondo/Max-like transcriptional complex, MML-1 and MXL-2. MML-1 (Myc/Mondo-like) is homologous to vertebrate Mondo/ChREBP protein and functions in an activation complex with its partner MXL-2 (Max/Max-like), homologue of Max-like. These proteins were previously identified as promoters of longevity in IIS and DR mutants [57]. The Antebi lab described the necessity of both factors for glp-1 longevity [58]. They reported that MML-1, that is widely expressed in somatic cells and shows nuclear, mitochondrial and cytoplasmic localization, was elevated upon GSC removal, in nuclei and mitochondria. MXL-2, which shows basal cytoplasmic expression, did not show an increased protein expression. Inactivation of either protein prevented the nuclear localization of HLH-30/TFEB. Intriguingly, hlh-30 RNAi diminished the expression of MML-1 in germline-less animals suggesting that these proteins mutually regulate each other. MML-1 overexpression produced variable but significant extension of wild-type lifespan. Surprisingly, mml-1/mxl-2 did not impact PHA-4/FOXA (or DAF-16/FOXO3A ), although molecular genetic analyses provided strong evidence that the complex promotes autophagy upon GSC loss by repressing TOR and facilitating HLH-30/TFEB nuclear relocation. Amongst the target genes of MML-1/MXL-2, another member Myc/Mondo complex, MDL-1, was also required for glp-1 longevity. MDL-1 overexpression augmented lifespan in fertile adults suggesting that multiple members of this family may play roles in germline-less longevity [58].
6.3.5 SKN-1/NRF2 , HSF-1/HSF
SKN-1 , homologue of mammalian NRF2 (nuclear factor-erythroid related factor 2), is another longevity determinant shared between multiple longevity pathways [53, 59]. The Nrf family regulates cellular detoxification, proteasomal degradation and metabolic pathways in vertebrates and invertebrates. skn-1, first identified for its role in embryonic development [60], has been shown to extensively influence adult health and longevity by regulating multiple stress-response pathways, in normal worms as well as in diverse long-lived mutants [53, 59]. SKN-1/NRF2 overexpression extends lifespan modestly as well [59]. The protein translocates to the nuclei of intestinal cells in IIS mutants, wherein, it promotes longevity and stress resistance in a genetically parallel pathway from DAF-16/FOXO3A [59]. However, in mutants representing DR longevity, its function in a pair of sensory neurons is sufficient for lifespan to be augmented [53]. skn-1 inactivation was previously reported to shorten glp-1 mutants’ longevity [61], and subsequently, the Blackwell and Kenyon laboratories confirmed this observation [62, 63]. Both groups showed that SKN-1/NRF2 nuclear localization occurs in intestinal cells of glp-1 mutants. The Blackwell lab reported that the nuclear entry was partially dependent upon tcer-1 and kri-1, and completely under control of pmk-1 that encodes a p38 MAP Kinase known to phosphorylate SKN-1/NRF2 in other contexts [62]. However, by testing the induction of a SKN-1/NRF2-target gene, gst-4, the Kenyon lab found that SKN-1/NRF2 is activated only marginally by the p38 MAP Kinase pathway, and strongly by the transsulphuration pathway [63]. The transsulphuration pathway leads to the production of sulphur-containing metabolites, including hydrogen sulphide (H2S) and previous reports have shown that H2S extends worm lifespan via SKN-1/NRF2 activity [64]. Interestingly, KRI-1/KRIT1 was partially responsible for H2S production and gst-4 induction, whereas, both events were DAF-16/FOXO3A independent, suggesting that KRI-1/KRIT1 has independent effects on DAF-16/FOXO3A and SKN-1/NRF2 . Using both site-specific overexpression and loss-of-action studies, the authors also showed that SKN-1/NRF2 acts in the adult intestine to extend lifespan [63].
The worm homologue of the human heat-shock factor (HSF), HSF-1, an essential component of the heat-shock response (HSR) and proteostasis is also required for glp-1 mutants’ longevity [65]. Similar to SKN-1/NRF2, HSF-1/HSF impacts multiple longevity paradigms as well as normal lifespan [66]. Other transcription regulators such as MDT-15 (component of the mediator complex that putatively works as a co-activator of NHR-49/PPARα) [67] and SBP-1 (homologue of human SREBP1 transcription factor) [68] have also been implicated in glp-1 longevity [69] and stress resistance [62], respectively, but details of their regulation and molecular function are as yet unaddressed.
6.4 Cellular Processes Mobilized in Response to Germline Depletion
As evinced from the paragraphs above, eliminating the worm germline triggers the activation of a host of transcription regulators largely in intestinal cells (Fig. 6.1) suggesting that GSC removal is accompanied by a major gene-expression shift. It raises questions about the nature of these transcriptional changes, their physiological outcomes and the relationships between the regulatory factors involved. Contemporary studies have provided some insights into these queries, although there are many more questions than there are answers in the field. In a microarray-based study, about 3440 genes were reported to be upregulated and 150 downregulated (of the approximately ~18,000 genes in the worm genome) in GSC-ablated worms as compared to whole-gonad ablated sterile worms [70]. Similar transcriptional changes accompany GSC removal in Pristionchus pacificus, a related nematode that exhibits GSC-less life extension too, with a significant overlap between the transcriptomes of GSC-less worms of the two species [70]. In contrast, an RNA-Sequencing (RNA-Seq)-based study identified a smaller group upregulated in glp-1 mutants’- 1306 and 615 genes, by more than fourfold and fivefold, respectively [62]. The differences in numbers between these studies notwithstanding, they support broad transcriptional remodelling upon GSC removal. Experiments aimed at identifying the targets of some of the transcription factors discussed above have suggested the involvement of specific biochemical pathways. McCormick et al. reported the identification of 230 and 130 genes whose expression was altered in glp-1 mutants dependent upon DAF-16/FOXO3A and DAF-12/VDR, respectively [69], whereas, Steinbaugh et al. identified 529 SKN-1/NRF2 targets in glp-1 mutants [62]. In a recent study, we discovered 835 and 801 downstream genes whose expression is governed by TCER-1/TCERG1 and DAF-16/FOXO3A, respectively, in glp-1 adults. About one-third of the targets are shared between the two factors [71]. A similar comparison of the downstream targets of HLH-30/TFEB and the MML-1/MXL-2 complex identified a substantial number of co-regulated genes; MML-1 and MXL-2 shared 827 targets, whereas, 202 were common between all three factors [58]. In all these studies, the gene lists were strongly enriched for lipid-metabolic functions. In addition, proteasomal degradation, autophagy and stress resistance are also consistently represented. In the following sections, we focus on each of these cellular processes and discuss their impacts on the reproductive control of ageing .
6.4.1 Lipid Metabolism and the Reproductive Control of Longevity
Germline removal in C. elegans not only increases lifespan and stress resistance it also causes elevated fat accumulation, observed using lipid-labelling dyes as well as biochemical approaches [72]. At first glance this is astonishing as obesity is associated with increased mortality, not better health and long life. However, gonadectomy precipitates enhanced fat accumulation in many organisms besides worms, in both invertebrates (e.g., fruit flies, blow flies, locusts, grasshoppers) [73–77] and vertebrates (e.g., mice, rats, cats and monkeys) [78–81]. In humans, deficient gonadal hormone production results in obesity and metabolic disorders [82]. But, all fat is not equal and all fat accumulation is not detrimental to the organism. Long-lived IIS worm mutants manifest greater adiposity but are healthier and longer-lived than their leaner, wild-type counterparts [72, 83]. In Drosophila, interventions that extend lifespan, such as reduced IIS and TOR inhibition, elevate fat [84, 85]. Obese mice that exhibit healthy metabolic profiles have been described too [86, 87]. Similarly, a small but striking group of ‘metabolically healthy obese’ individuals are notable because they retain excessive weight without developing clinical pathologies [88]. Altogether, there is increasing awareness of the nuanced and multi-layered relationship between adiposity, reproduction and lifespan (for reviews see [89, 90]). Studies in GSC-less worms have revealed interesting answers to questions regarding this trifecta .
6.4.2 Coordinate Induction of Lipogenesis and Lipolysis Following GSC Loss
In our efforts to map the transcriptomes dictated by TCER-1/TCERG1 and DAF-16/FOXO3A following GSC loss, we made the intriguing discovery that the targets of these factors were enriched in both lipid production and degradation functions [71]. Overall, we found that the lipogenic classes included genes involved in de novo fatty-acid synthesis, fat desaturation and elongation and conversion of diglycerides (DAGs) to the storage form, triglycerides (TAGs). Lipolytic processes were equally represented with the inclusion of multiple peroxisomal and mitochondrial β-oxidation genes as well as numerous lipases. Functional studies showed that both anabolic and catabolic genes contributed to longevity, but the mechanism and overall significance of this apparent widespread lipid turnover is unknown. It likely reflects an adaptive mechanism that allows the animal to preserve metabolic homeostasis when faced with fertility loss, consistent with observations in other species. For instance, DR flies [91] and mice [92] display enhanced fatty-acid turnover. In humans, inefficient lipid turnover is associated with metabolic diseases [93]. Alternatively, endurance-trained athletes who have increased insulin sensitivity display high lipid content in muscles , a phenomenon termed ‘athlete’s paradox’ [94]. Such observations suggest that concordant modulation of fat buildup and breakdown may be a conserved mechanism utilized by organisms facing diverse physiological challenges. We summarize below data on the different lipid-metabolic pathways that have been studied in GSC-less worms. A discussion of the broader role of lipids in ageing can be found in Chap. 14.
6.4.2.1 de novo Fatty-Acid Synthesis and TAG Production
In our RNA-Seq study [71], key conserved genes encoding enzymes responsible for initiating de novo fatty acid synthesis were identified as being upregulated by DAF-16/FOXO3A and/or TCER-1/TCERG1 upon GSC removal. These included pod-2 (encodes acetyl CoA carboxylase, ACC), fasn-1 (encodes fatty-acid synthase, FAS) and mlcd-1 (malonyl CoA decarboxylase 1, MLCD). Accordingly, lipid-labelling studies demonstrated elevated de novo fatty-acid synthesis following GSC removal. This is a functionally relevant metabolic shift because inactivation of each of these genes suppresses glp-1 mutants’ longevity. Similarly, five of the six genes encoding diacyl glycerol acyl transferase (DGAT) enzymes that catalyse the final step in TAG production (dgat-2, acs-22, mboa-2, Y53G8B.2 and K07B1.4) were upregulated by DAF-16/FOXO3A and/or TCER-1/TCERG1. Expectedly, lipid staining and GC/MS analyses showed increased TAG levels in glp-1 adults. Functional studies indicate that this is achieved, at least partly, through DAF-16/FOXO3A and TCER-1/TCERG1-mediated increase in expression of ‘dgat’ genes, each making modest and redundant contributions to GSC-less longevity [71].
6.4.2.2 Fatty-Acid Desaturation
In glp-1 mutants’, both NHR-49/PPARα and NHR-80/HNF4 enhance the expression of desaturase enzymes that catalyse the conversion of saturated fatty acids (SFAs) into unsaturated fatty acids (UFAs), including mono- and poly- unsaturated fatty acids (MUFAs and PUFAs, respectively). Accordingly, glp-1 mutants exhibit increased MUFA and reduced SFA levels [71]. The expressions of fat-6 and fat-7, genes encoding stearoyl-CoA 9-desaturase (SCD) enzymes that redundantly transform stearic acid (SA, a SFA) to oleic acid (OA, a MUFA) are strongly altered. Genetic evidence suggests that DAF-16/FOXO3A and TCER-1/TCERG1 may exert indirect control over these genes, through upregulation of NHR-49/PPARα (but not NHR-80/HNF4) [44, 71]. GSC-less animals exhibit enhanced OA levels as compared to fertile worms but the precise function performed by OA is unknown [43]. In the absence of NHR-49/PPARα or NHR-80/HNF4, OA supplementation does not rescue the long life of GSC-less worms. So, it may function as a lipid ligand for these factors. Indeed, OA supplementation also enhances SKN-1/NRF2 nuclear localization suggesting that this protein may also be activated by lipid ligands [62]. Interestingly, a recent study found that an OA metabolite, oleoylethanol amide (OEA), binds NHR-80/HNF4 and promotes the expression of downstream targets of both NHR-80/HNF4 and NHR-49/PPARα in fertile animals [48]. Levels of other fatty acids besides OA are modulated upon GSC loss as well, and their function remains unknown [71]. For instance, SKN-1/NRF2 nuclear localization can also be triggered by treatment with coconut oil, which includes OA and many other UFAs and SFAs [62]. glp-1 mutants exhibit higher levels of other UFAs, including ones with ≥18 carbon chains, and one or more of these may be important as well [71].
6.4.2.3 Mitochondrial β-Oxidation
While NHR-80/HNF4 influences desaturation alone, NHR-49/PPARα also upregulates the expression of genes involved in β-oxidation, a multi-step pathway that results in breakdown of fatty acids into Acetyl CoA moieties. Genes encoding enzymes operating at each of these steps are elevated in glp-1 mutants in an NHR-49/PPARα-dependent manner, suggesting that fatty-acid oxidation may be elevated upon GSC removal [44]. This enhancement is likely to be physiologically important because inactivation of many of these genes reduces glp-1 mutants’ longevity. The DAF-16/FOXO3A and TCER-1/TCERG1 transcriptomes are also enriched for β-oxidation genes, and genetic evidence suggests that these proteins modulate the process indirectly through their effect on NHR-49/PPARα [44, 71]. Interestingly, the SKN-1/NRF2 target list also includes many β-oxidation genes [62]. The adiposity of GSC-less mutants is further enhanced in a skn-1 mutant background, implying that it acts to limit fat accrual before it reaches toxic levels. Intriguingly, nhr-49;glp-1 mutants do not show further increase in stored lipids. Instead, they undergo a striking depletion of fat with age [44]. In literature, similar phenotypes have been observed when cells and animal tissues exposed to unesterified ‘free’ fatty acids (FFA), undergo ‘lipotoxicity’ [95, 96]. It is plausible that in the absence of NHR-49/PPARα, the simultaneous inhibition of β-oxidation and desaturation causes accumulation of FFAs and consequent lipotoxicity.
6.4.2.4 Lipolysis
One of the first fat-metabolic genes found to be essential for glp-1 mutants’ longevity by the Ruvkun lab encoded a lipase, LIPL-4, orthologous to a human lipase LIPA. lipl-4 expression is intestine restricted and upregulated by DAF-16/FOXO3A in glp-1 mutants; it’s overexpression lengthens fertile animals’ lifespan [97]. Besides LIPL-4, at least six other lipases and lipase-like proteins encoded in the C. elegans genome are upregulated (or predicted to be so) following GSC loss, dependent upon DAF-16/FOXO3A, TCER-1/TCERG1 or SKN-1/NRF2 [62, 69, 71]. RNAi of many of these reduces glp-1 longevity [71]. The involvement of multiple lipases in this lifespan paradigm is inexplicable, especially in the light of the marked adiposity manifested by GSC-less animals. LIPL-4/LIPA has been shown to link lipid metabolism to autophagic flux upon GSC removal (discussed in the next section) [54] and the SKN-1/NRF2 target LIPL-3 is postulated to prevent excess fat accumulation [62]. In fertile worms, OEA production is LIPL-4 dependent [48, 97] and in human cardiac cells, the cytosolic lipase ATGL-1 is essential for PPARα ligand synthesis [98]. Hence, it is plausible that one or more of these lipases may also facilitate the production of signalling molecules or ligands for factors such as NHR-49/PPARα or NHR-80/HNF4. But, overall little is understood about the function of the lipolytic genes and is likely to be a major focus of future studies.
The current data on lipid metabolism and reproductive control of ageing raises many important questions. Why does GSC-ablation increase fat content and what is the nature of these adipose depots? Why are lipid catabolism and anabolism simultaneously augmented and how? Lipid content is increased largely in the intestine , and to some extent, in the epidermis. However, the worm intestine is not simply a part of the alimentary canal and the major fat-storage depot, it also subsumes functions of the liver and pancreas, is the main site for induction of immunogenic responses and the sole centre for yolk synthesis. Yolk, made up of lipids and proteins, is generated in gut cells, secreted into the body cavity and transported to the gonad to be deposited into oocytes for nourishment of the embryo. It is a logical supposition that the adiposity of GSC-less animals is derived from yolk lipids that can no longer be deposited into eggs, and much of it is likely to be so. Indeed, Steinbaugh et al. showed that GSC-less mutants continue to produce a yolk lipoprotein, vitellogenin 2 (VIT-2) [62]. But in this study, VIT-2 localization was predominantly in the body cavity, not the intestine. Instead, there was increased level of GFP-labelled DHS-3 (DHS-3::GFP), a protein that almost exclusively labels TAGs, in gut cells of GSC-less mutants [62]. Additionally, the gene-expression data and biochemical evidences showing elevated de novo fatty-acid synthesis [71], TAG production [44, 71, 72] and lipid desaturation [43, 44, 71] suggest that the excess fat in GSC-less worms is derived at least in part from bona fide increase in lipid production. But why is lipogenesis triggered when fertility is thwarted? While DGATs can help immure fats (normally designated for oocytes in fertile animals) into lipid droplets, what purpose is served by elevating de novo fatty-acid synthesis? The reason for this is unknown, but vertebrate and worm evidences suggest that it may serve signalling functions. Fatty acids have been known as lipid ligands for long. But, recent reports have begun to emphasize the importance of the ‘source’ of lipid signals . For instance, mice incapable of synthesizing ‘new fat’ due to FAS/FASN-1 deletion in the liver or hypothalamus cannot activate PPARα [99]. So, augmenting de novo lipid synthesis may help synthesize ligands for factors activated upon germline depletion. As mentioned above, PPARα ligand production in cardiac cells is dependent upon the lipase ATGL-1 [98], so it is equally possible that one or more of the lipases upregulated upon GSC removal also contributes to the synthesis of such ligands.
Why is lipid desaturation increased upon GSC loss? The elevated MUFA:SFA ratio observed in glp-1 mutants suggests that the metabolic shift involves not only quantitative but also a qualitative remodelling of the lipid profile. SFAs are critical for reproductive health as they make up >70 % of human oocyte lipids, whereas, MUFAs make up <15 % [100]. But, SFAs are poor substrates for incorporation into TAGs and major causes of lipotoxicity [95, 96]. Alternatively, lipids with higher UFAs are generally associated with improved cellular maintenance [101], and with enhanced lifespan in human centenarian studies [102]. The transformation of a SFA-rich, reproduction -oriented lipid profile of a fertile adult into one that is enriched in UFAs by proteins such as NHR-49/PPARα and NHR-80/HNF4 may mitigate the deleterious effects of GSC loss and organize a lipid profile conducive for somatic maintenance and health. Whether this transformation is simply an adaptation to sterility or the bedrock for longevity remains to be discovered, and the two possibilities are not mutually exclusive either. Another important question emerging from these studies is how these transcription factors simultaneously elevate ostensibly antagonistic lipid-metabolic steps in the same animal. Future experiments will address if, and how, this coordination is managed, and if the strategy is widely used in the animal kingdom for retaining lipid homeostasis in the face of metabolic challenges.
6.4.3 Role of Autophagy in Linking Germline Status to Lifespan
Autophagy is a conserved cellular recycling process in which cytosolic components, including damaged organelles, misfolded proteins and sometimes pathogens are sequestered in vesicles called autophagosomes [103] (see also Chap. 15). The contents are degraded when autophagosomes fuse with lysosomes gaining access to the lysosomal degradative enzymes and recycling machinery. Autophagy has emerged as a shared mechanism for various longevity paradigms in multiple species [104]. In C. elegans , mutants representing the IIS and DR longevity pathways exhibit increased autophagy and require autophagy genes for their long lives [55, 105]. The Hansen lab showed that autophagic events are enhanced in glp-1 mutants, in the two main fat depots, intestine and hypodermis [54]. This enhancement is likely under transcriptional control because the expression of multiple autophagic genes, including bec-1/BECN-1, lgg-1/LC3 and unc-51/ULK1, is increased in glp-1 mutants and these genes promote longevity . PHA-4 /FOXA was found to be responsible for mediating this transcriptional alteration- a surprising finding since autophagy was traditionally thought to be under post-transcriptional regulation. This study helped establish a link between autophagy and lipid metabolism in the long life of GSC-less animals. The increased autophagy of GSC-less mutants was found to be dependent on the lipase LIPL-4/LIPA, and conversely, autophagy genes were essential for elevated lipase activity. The lifespan extension obtained by LIPL-4/LIPA overexpression [97] is accompanied with increased autophagic events, and is abolished when pha-4 or any of the autophagy genes are inactivated [54]. The downregulation of TOR is a common upstream event that triggers both enhanced lipase activity and autophagy. The Hansen lab provided further proof for the instrumental role of autophagy in this longevity paradigm through identification of HLH-30/TFEB [55]. As described above, it is a conserved transcription factor that upregulates autophagic and lysosomal genes, via a TOR -dependent pathway. In glp-1 mutants, HLH-30/TFEB mediates enhanced expression of numerous predicted orthologues of TFEB targets involved in autophagosome formation and autophagic flux (e.g., lgg-1/LC3 and sqst-1/SQSTM1/p62) as well as genes with lysosomal functions (e.g., vacuolar ATPase subunits, cathepsin peptidases and sulphatases). Conversely, HLH-30/TFEB impairment prevents autophagy augmentation. The Antebi lab’s studies added MML-1/MXL-2 to this select group of factors augmenting autophagy upon germline loss [58]. Several autophagy genes were identified amongst MML-1/MXL-2 downstream targets. The MML-1/MXL-2 complex was shown to transcriptionally regulate the TOR pathway by repressing lars-1, a gene encoding a leucine t-RNA synthetase that stimulates TOR. TOR repression triggers HLH-30/TFEB nuclear localization, in part, initiating the autophagy cascade. Accordingly, mml-1/mxl-2 knockdown causes diminished appearance of autophagy markers such as LGG-1 puncta in worm tissues. However, neither HLH-30/TFEB overexpression nor TOR downregulation restored longevity in glp1mml1 or glp-1mxl-2 mutants. Although the three factors regulate autophagy, they control distinct autophagic gene sets suggesting the existence of overlapping genetic networks rather than a linear pathway [58].
In fasted worms, HLH-30/TFEB elevates the expression of multiple lipases in addition to up-regulating autophagy genes, but in this activity it is antagonized by MXL-3, another Mondo family member [106]. Interestingly, the increased lipolysis observed during starvation also leads to enrichment of PUFAs, such as arachidonic acid (AA) and di-homo-γ-linoleic acid (DGLA). Genetic evidence implies that these PUFAs, in turn, activate autophagy to confer resistance against nutritional deprivation [107]. HLH-30/TFEB and MML-1/MXL-2 may perform similar functions in response to GSC removal. Indeed, many lipid-metabolic genes were included in the genomic targets of these proteins [58]. Alternatively, autophagy may be required for lipid hydrolysis, or ‘lipophagy’. Recent evidence suggests that lipophagy serves multiple functions, depending on physiological context, including the large-scale hydrolysis of neutral lipids by lysosomal lipases, and production of signalling lipids, as seen in pancreatic β cells [108]. In GSC-less worms, the process may be utilized to catalyse the production of signalling lipids that activate one of the many ligand-activated transcription factors involved in this pathway. It may also promote lifespan by simply ensuring advantageous yolk repartitioning or by preventing the accumulation of toxic lipid intermediates, including FFAs. Interestingly, autophagy genes have been reported to be important for maintaining adequate TAG levels in fertile animals, although reducing their functions does not shorten wild-type lifespan [54, 109]. This further substantiates that this catabolic process has broad and possibly interlinked roles in influencing lipid homeostasis and lifespan, and the relationship may be physiological-context dependent. Much remains obscure about the intriguing links between autophagy, lifespan and lipid metabolism and is likely to be the focus of current and future efforts in the field.
6.4.4 Proteasome Activity in Germline-Less Animals
One of the major quality control mechanisms that influence cellular homeostasis is the ability to degrade proteins. Autophagy and the ubiquitin proteasome system (UPS) are the two main proteolytic systems, the UPS being the primary pathway for protein degradation in eukaryotic cells. The most well-known function of UPS is the spatially and temporally controlled destruction of regulatory proteins that inform various cellular processes. Regulatory proteasomal activity has been implicated in the lifespan extension of C. elegans mutants, including GSC-less worms [110–112] (see also Chap. 12). In addition, the ‘housekeeping’ function of UPS in destroying damaged, misfolded, old and aggregation-prone proteins is critical for maintenance of the proteostasis network . Loss of proteostasis is one of the hallmarks of ageing to which numerous age-related pathologies are attributed [113]. In C. elegans, the ability to retain proteostasis falls dramatically once peak reproductive age is reached, and declines over time [114]. Conversely, germline-deficient animals exhibit enhanced resistance to conditions that tax the proteostasis machinery such as high temperatures or toxic protein aggregates [115]. This striking resilience is dependent on many of the transcription factors discussed here, including DAF-16/FOXO3A , NHR-80/HNF4, PHA-4 /FOXA, DAF-12/VDR and SKN-1/NRF2 [62, 115]. Expectedly, proteostasis and UPS are highly represented gene-categories in glp-1 genomic studies. In particular, SKN-1/NRF2 targets include a large number of proteasomal subunits [62]. The Dillin lab first showed that glp-1 mutants exhibit elevated proteasomal activity in their somatic tissues as compared to fertile worms [61]. This enhancement could be attributed to elevated expression of the gene rpn-6.1. rpn-6.1 encodes the worm ortholog of PSMD11, a protein that stabilizes the interactions of the 20S proteasomal core with the 19S regulatory cap, a critical step for proteasomal activity [61, 116]. In this report, rpn-6.1 expression was found to be elevated in glp-1 mutants in a DAF-16/FOXO3A-dependent manner and, surprisingly, independent of SKN-1/NRF2 and HSF-1/HSF. RPN-6.1/PSMD11 overexpression augments overall UPS activity, increases lifespan and confers protection against toxic aggregates in Huntington’s Disease (HD) models [61]. Strikingly, immortal human embryonic stem cells (hESCs) also manifest high PSMD11 levels and elevated UPS activity, dependent upon FOXO4 , one of DAF-16 orthologs [116]. Data from the Blackwell lab confirmed the elevated proteasomal activity of glp-1 mutants, but found it to be strongly dependent upon SKN-1/NRF2. Indeed, rpn-6.1 appears to be a direct target of SKN-1/NRF2 [62]. The discrepancies notwithstanding, these studies have provided strong evidence that improved protein homeostasis is one of the consequences of GSC removal and very possibly a major contributing factor to the ensuing enhancement in lifespan and health.
6.4.5 Stress-Response Mechanisms and GSC-Less Longevity
Many interventions that prolong life also confer tolerance to environmental stressors, in worms and in many other species, though exceptions exist [117]. glp-1 mutants also exhibit this positive correlation between longevity and stress resistance. In C. elegans , the most commonly studied stress paradigms include oxidative stress, heat shock, protein misfolding in the endoplasmic reticulum (ER) and mitochondria that evokes an unfolded protein response (UPR) in these organelles (UPRERand UPRmt, respectively) and immuno-competence, or ability to combat pathogen attack [118]. Arantes-Oliveira et al. first reported the enhanced oxidative-stress resistance of glp-1 mutants [22]. Steinbaugh et al. substantiated this data and characterized the vital role for SKN-1/NRF2 in mediating this resilience [62]. Wei and Kenyon linked GSC removal to altered redox signalling and showed that GSC removal causes elevation in the levels of reactive oxygen species (ROS) and H2S cell non-autonomously [63]. These events are important because quenching ROS with anti-oxidants or inhibiting the transsulphuration pathway responsible for H2S production curtailed longevity. Interestingly, ROS and H2S appear to activate different stress-response paradigms during adulthood; ROS leads to UPRmt, whereas, H2S causes SKN-1/NRF2 activation. Using two chemical redox sensors, it was found that ROS production was induced in two waves. The first one, a mitochondrial signal, was detected late in larval life, just before the animal reaches adulthood. The second cytoplasmic ROS signal appeared during early adulthood and activated UPRmt (as evidenced by the induction of the UPRmt reporter, hsp-6) through upregulation of the transcription factors DVE-1 that mediates UPRmt, and its co-activator UBL-5 [63]. Interestingly, atfs-1, that encodes the key mediator of UPRmt, is included in the list of genes upregulated by DAF-16/FOXO3A and its RNAi knockdown suppresses glp-1 longevity [71], but hsp-6 induction was found to be DAF-16/FOXO3A independent [63]. The finding that SKN-1/NRF2 -dependent detoxification systems and DVE-1-dependent UPRmt are induced by different redox systems is highly intriguing, and may reflect the response of somatic tissues to loss of individual aspects of germ-cell physiology. How the other transcription factors of the network play into these stress-response initiatives remains to be described.
Studies focusing on the interaction between reproductive fitness and somatic endurance have revealed considerable information on the ability of GSC-less mutants to mount a chaperone-driven heat-shock response (HSR) following exposure to high temperatures. In C. elegans, thermo-resistance declines with the onset of reproduction , at least in part due to diminished expression of a histone H3 trimethyllysine-27 (H3K27me3) demethylase, JMJD-3.1, that antagonizes transcription-repressive chromatin marks [119]. glp-1 mutants are exceptionally thermotolerant, dependent upon many of the genes discussed here, including daf-16, tcer-1, kri-1, hsf-1 and jmjd-3.1 [115, 119]. However, other mutants that exhibit sterility due to gonadogenesis defects (not GSC loss) and have normal lifespan (e.g. glp-4, gon-2) are also thermotolerant, although genetic evidence hints that GSC removal (not just arresting reproduction) may trigger their HSR.
As with ageing , immuno-competence also manifests an inverse correlation with reproduction in many organisms, including C. elegans. Expectedly, glp-1 mutants exhibit superior resistance against gram-negative pathogens such as Salmonella enterica [120], Pseudomonas aeruginosa [121, 122] and Serratia marcescens [122], the gram-positive pathogen Enterococcus faecalis [120], and the fungal pathogen Cryptococcus neoformans [120]. Of these, the response to P. aeruginosa has been well studied. Alper et al. reported that DAF-16/FOXO3A’s requirement for glp-1 mutants’ immunoresistance was influenced by worm-culture conditions, indicating that nutrient status may have an impact on the reproduction-immunity relationship as well [122]. Interestingly, the DAF-16/FOXO3A-driven immuno-resistance appears to be associated with sterility and not especially GSC status, as similar resistance is exhibited by other gonadal mutants that are sterile but not long lived (e.g., the feminized mutant, fog-2 and the somatic-gonad defective mutant, glp-4) [122, 123]. DAF-16/FOXO3A also undergoes nuclear relocation in many gonadal mutants not just those lacking GSCs. However, GSC removal seems essential to confer broader and stronger immunity, because both glp-1 and glp-4 mutants are resilient against P. aeruginosa, whereas, only glp-1 mutants are resistant to S. enterica infection [120]. Altogether, these findings reiterate that GSCs may signal to inhibit immunocompetence, and upon their removal innate immunity is enhanced. In few cases, the genetic basis of this improved immunity has been dissected and appears to be partially DAF-16/FOXO3A dependent. But, it remains to be seen if the other transcription factors that are important for longevity have similar roles , and if so, whether they are stress specific or shared.
6.4.6 Pathways Repressed Upon GSC Loss
Fertile, young adults are highly invested in macromolecular synthesis to support their reproductive physiology. Loss of the germline induces a fundamental change in this metabolic state, and an inability to suppress the growth programmes already in place can be detrimental to the animal. Thus, genes that are downregulated following GSC ablation are likely as crucial as those that are upregulated. However, these factors have received little attention so far. In examining the targets repressed by DAF-16/FOXO3A and TCER-1/TCERG1, we noted high enrichment of molecular pathways associated with active procreation such as protein translation and reproduction , respectively. Indeed, DAF-16/FOXO3A appears to repress the transcription of at least 18 genes involved in translation including those encoding ribosomal subunits and translation initiation factors [71]. Since protein synthesis is a key requirement for a proliferating germline, it is plausible that GSC removal triggers DAF-16/FOXO3A -dependent translation repression. On the other hand, the list of genes repressed specifically by TCER-1/TCERG1 is enriched for factors with predicted roles in reproduction [71], suggesting that that upon GSC loss, TCER-1/TCERG1 actively represses the somatic programme of reproduction. Thus, DAF-16/FOXO3A and TCER-1/TCERG1 may together facilitate the adaptation to GSC depletion by terminating the somatic gene-expression programmes that support reproductive physiology. The repressive functions undertaken by other proteins in this longevity network remain to be described.
6.5 Contributions from C. elegans to the Reproductive Control of Ageing
The initial observations of Hsin and Kenyon that demonstrated the control exerted by the germline on the lifespan of the animal [20], and challenged the simplistic ‘trade off’ interpretation, laid the groundwork for a field that has burgeoned into great significance and mainstream science interest. The early worm studies led to a renewed examination of the germline-soma dialogue in other model organisms and species. The remarkable ease of molecular-genetic analysis and large-scale RNAi screening in worms allowed the identification of innumerable genetic players with roles in this dialogue, many of them with conserved functions in lifespan regulation. These studies have not only revealed knowledge about ageing but have also led to important discoveries in the fields of metabolism, autophagy and proteostasis . The mechanisms found to be operating in GSC-less worms, at least with respect to proteostasis and lipid metabolism , appear to be recapitulated in higher organisms in other physiological contexts [116, 124]. Recent human studies have begun to closely examine the effect of reproductive status on health and longevity. Age at menarche and timing of menopause in women have both been found to impact susceptibility to age-linked diseases such as cancer, cardiovascular disease (CVD) and osteoporosis [125]. Early gonadal failure is associated with reduced life span and increased morbidity from CVD in both men and women [126, 127]. Thus, worm studies have provided greater urgency and impetus to emerging evidence that the influence of germline on overall health has significant biomedical relevance. Indeed, it is difficult to overstate the contributions of C. elegans research to the understanding of the relationship between procreation and lifespan .
References
Chapman T, Liddle LF, Kalb JM, Wolfner MF, Partridge L (1995) Cost of mating in Drosophila melanogaster females is mediated by male accessory gland products. Nature 373(6511):241–244. doi:10.1038/373241a0
Shi C, Murphy CT (2014) Mating induces shrinking and death in Caenorhabditis mothers. Science 343(6170):536–540. doi:10.1126/science.1242958
Maures TJ, Booth LN, Benayoun BA, Izrayelit Y, Schroeder FC, Brunet A (2014) Males shorten the life span of C. elegans hermaphrodites via secreted compounds. Science 343(6170):541–544. doi:10.1126/science.1244160
Crudgington HS, Siva-Jothy MT (2000) Genital damage, kicking and early death. Nature 407(6806):855–856. doi:10.1038/35038154
Blanckenhorn WU, Hosken DJ, Martin OY, Reim C, Teuschl Y, Ward PI (2002) The costs of copulating in the dung fly Sepsis cynipsea. Behav Ecol 13(3):353–358. doi:10.1093/beheco/13.3.353
Gwynne DT (2008) Sexual conflict over nuptial gifts in insects. Annu Rev Entomol 53:83–101. doi:10.1146/annurev.ento.53.103106.093423
Boggs CL, Gilbert LE (1979) Male contribution to egg production in butterflies: evidence for transfer of nutrients at mating. Science 206(4414):83–84. doi:10.1126/science.206.4414.83
Aprison EZ, Ruvinsky I (2015) Sex pheromones of C. elegans males prime the female reproductive system and ameliorate the effects of heat stress. PLoS Genet 11(12):e1005729. doi:10.1371/journal.pgen.1005729
Partridge L, Gems D, Withers DJ (2005) Sex and death: what is the connection? Cell 120(4):461–472. doi:10.1016/j.cell.2005.01.026
Law R (1979) The cost of reproduction in annual meadow grass. Am Nat 113(1):3–16
Butlin RK, Day TH (1985) Adult size, longevity and fecundity in the seaweed fly, Coelopa-Frigida. Heredity 54:107–110. doi:10.1038/hdy.1985.14
Kirkwood TB (1977) Evolution of ageing. Nature 270(5635):301–304
Ricklefs RE, Cadena CD (2007) Lifespan is unrelated to investment in reproduction in populations of mammals and birds in captivity. Ecol Lett 10(10):867–872. doi:10.1111/j.1461-0248.2007.01085.x
Westendorp RG, Kirkwood TB (1998) Human longevity at the cost of reproductive success. Nature 396(6713):743–746. doi:10.1038/25519
Gavrilova NS, Gavrilov LA (2005) Human longevity and reproduction: an evolutionary perspective. In: Voland E, Chasiotis A, Schiefenhoevel W (eds) Grandmotherhood – the evolutionary significance of the second half of female life. Rutgers University Press, Piscataway, pp 59–80
Korpelainen H (2000) Fitness, reproduction and longevity among European aristocratic and rural Finnish families in the 1700s and 1800s. Proc Biol Sci 267(1454):1765–1770. doi:10.1098/rspb.2000.1208
Le Bourg É (2001) A mini-review of the evolutionary theories of aging: is it the time to accept them? Demogr Res 4(1):1–28. doi:10.4054/DemRes.2001.4.1
Mitteldorf J (2010) Female fertility and longevity. Age (Dordr) 32(1):79–84. doi:10.1007/s11357-009-9116-1
Keith SA, Ghazi A (2015) Recent discoveries in the reproductive control of aging. Curr Genet Med Rep 3(1):26–34
Hsin H, Kenyon C (1999) Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399(6734):362–366. doi:10.1038/20694
Austin J, Kimble J (1987) glp-1 is required in the germ line for regulation of the decision between mitosis and meiosis in C. elegans. Cell 51(4):589–599
Arantes-Oliveira N, Apfeld J, Dillin A, Kenyon C (2002) Regulation of life-span by germ-line stem cells in C. elegans. Science 295(5554):502–505. doi:10.1126/science.1065768
Flatt T, Min KJ, D’Alterio C, Villa-Cuesta E, Cumbers J, Lehmann R, Jones DL, Tatar M (2008) Drosophila germ-line modulation of insulin signaling and lifespan. Proc Natl Acad Sci U S A 105(17):6368–6373. doi:10.1073/pnas.0709128105
Cargill SL, Carey JR, Muller HG, Anderson G (2003) Age of ovary determines remaining life expectancy in old ovariectomized mice. Aging Cell 2(3):185–190
Mason JB, Cargill SL, Anderson GB, Carey JR (2009) Transplantation of young ovaries to old mice increased life span in transplant recipients. J Gerontol Ser A Biol Sci Med Sci 64(12):1207–1211. doi:10.1093/gerona/glp134
Kenyon CJ (2010) The genetics of ageing. Nature 464(7288):504–512. doi:10.1038/nature08980
Lin K, Hsin H, Libina N, Kenyon C (2001) Regulation of the C. elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat Genet 28(2):139–145. doi:10.1038/88850
Libina N, Berman JR, Kenyon C (2003) Tissue-specific activities of C. elegans DAF-16 in the regulation of lifespan. Cell 115(4):489–502
Berman JR, Kenyon C (2006) Germ-cell loss extends C. elegans life span through regulation of DAF-16 by kri-1 and lipophilic-hormone signaling. Cell 124(5):1055–1068. doi:10.1016/j.cell.2006.01.039
Ghazi A, Henis-Korenblit S, Kenyon C (2009) A transcription elongation factor that links signals from the reproductive system to lifespan extension in C. elegans. PLoS Genet 5(9):e1000639. doi:10.1371/journal.pgen.1000639
Antebi A, Yeh WH, Tait D, Hedgecock EM, Riddle DL (2000) daf-12 encodes a nuclear receptor that regulates the dauer diapause and developmental age in C. elegans. Genes Dev 14(12):1512–1527
Motola DL, Cummins CL, Rottiers V, Sharma KK, Li T, Li Y, Suino-Powell K, Xu HE, Auchus RJ, Antebi A, Mangelsdorf DJ (2006) Identification of ligands for DAF-12 that govern dauer formation and reproduction in C. elegans. Cell 124(6):1209–1223. doi:10.1016/j.cell.2006.01.037
Gerisch B, Rottiers V, Li D, Motola DL, Cummins CL, Lehrach H, Mangelsdorf DJ, Antebi A (2007) A bile acid-like steroid modulates C. elegans lifespan through nuclear receptor signaling. Proc Natl Acad Sci U S A 104(12):5014–5019. doi:10.1073/pnas.0700847104
Shen Y, Wollam J, Magner D, Karalay O, Antebi A (2012) A steroid receptor-microRNA switch regulates life span in response to signals from the gonad. Science 338(6113):1472–1476. doi:10.1126/science.1228967
Hertweck M, Gobel C, Baumeister R (2004) C. elegans SGK-1 is the critical component in the Akt/PKB kinase complex to control stress response and life span. Dev Cell 6(4):577–588
Boehm M, Slack F (2005) A developmental timing microRNA and its target regulate life span in C. elegans. Science 310(5756):1954–1957. doi:10.1126/science.1115596
Boulias K, Horvitz HR (2012) The C. elegans microRNA mir-71 acts in neurons to promote germline-mediated longevity through regulation of DAF-16/FOXO. Cell Metab 15(4):439–450. doi:10.1016/j.cmet.2012.02.014
Gerisch B, Weitzel C, Kober-Eisermann C, Rottiers V, Antebi A (2001) A hormonal signaling pathway influencing C. elegans metabolism, reproductive development, and life span. Dev Cell 1(6):841–851
Rottiers V, Motola DL, Gerisch B, Cummins CL, Nishiwaki K, Mangelsdorf DJ, Antebi A (2006) Hormonal control of C. elegans dauer formation and life span by a Rieske-like oxygenase. Dev Cell 10(4):473–482. doi:10.1016/j.devcel.2006.02.008
Wollam J, Magner DB, Magomedova L, Rass E, Shen Y, Rottiers V, Habermann B, Cummins CL, Antebi A (2012) A novel 3-hydroxysteroid dehydrogenase that regulates reproductive development and longevity. PLoS Biol 10(4):e1001305. doi:10.1371/journal.pbio.1001305
Yamawaki TM, Berman JR, Suchanek-Kavipurapu M, McCormick M, Gaglia MM, Lee SJ, Kenyon C (2010) The somatic reproductive tissues of C. elegans promote longevity through steroid hormone signaling. PLoS Biol 8(8):e1000468. doi:10.1371/journal.pbio.1000468
Li TM, Liu W, Lu S, Zhang YP, Jia LM, Chen J, Li X, Lei X, Dong MQ (2015) No significant increase in the delta4- and delta7-dafachronic acid concentration in the long-lived glp-1 mutant, nor in the mutants defective in dauer formation. G3 (Bethesda) 5(7):1473–1479. doi:10.1534/g3.115.018812
Goudeau J, Bellemin S, Toselli-Mollereau E, Shamalnasab M, Chen Y, Aguilaniu H (2011) Fatty acid desaturation links germ cell loss to longevity through NHR-80/HNF4 in C. elegans. PLoS Biol 9(3):e1000599. doi:10.1371/journal.pbio.1000599
Ratnappan R, Amrit FR, Chen SW, Gill H, Holden K, Ward J, Yamamoto KR, Olsen CP, Ghazi A (2014) Germline signals deploy NHR-49 to modulate fatty-acid beta-oxidation and desaturation in somatic tissues of C. elegans. PLoS Genet 10(12):e1004829. doi:10.1371/journal.pgen.1004829
Evans RM, Mangelsdorf DJ (2014) Nuclear receptors, RXR, and the big bang. Cell 157(1):255–266. doi:10.1016/j.cell.2014.03.012
Van Gilst MR, Hadjivassiliou H, Jolly A, Yamamoto KR (2005) Nuclear hormone receptor NHR-49 controls fat consumption and fatty acid composition in C. elegans. PLoS Biol 3(2):e53. doi:10.1371/journal.pbio.0030053
Van Gilst MR, Hadjivassiliou H, Yamamoto KR (2005) A C. elegans nutrient response system partially dependent on nuclear receptor NHR-49. Proc Natl Acad Sci U S A 102(38):13496–13501. doi:10.1073/pnas.0506234102
Folick A, Oakley HD, Yu Y, Armstrong EH, Kumari M, Sanor L, Moore DD, Ortlund EA, Zechner R, Wang MC (2015) Aging. Lysosomal signaling molecules regulate longevity in C. elegans. Science 347(6217):83–86. doi:10.1126/science.1258857
Antebi A (2015) Nuclear receptor signal transduction in C. elegans. In: WormBook. The C. elegans research community, WormBook
Burkewitz K, Morantte I, Weir HJ, Yeo R, Zhang Y, Huynh FK, Ilkayeva OR, Hirschey MD, Grant AR, Mair WB (2015) Neuronal CRTC-1 governs systemic mitochondrial metabolism and lifespan via a catecholamine signal. Cell 160(5):842–855. doi:10.1016/j.cell.2015.02.004
Ratnappan R, Ward JD, Yamamoto KR, Ghazi A (2016) Nuclear hormone receptors as mediators of metabolic adaptability following reproductive perturbations. Worm 5:e1151609. doi:10.1080/21624054.2016.1151609
Panowski SH, Wolff S, Aguilaniu H, Durieux J, Dillin A (2007) PHA-4/Foxa mediates diet-restriction-induced longevity of C. elegans. Nature 447(7144):550–555. doi:10.1038/nature05837
Bishop NA, Guarente L (2007) Two neurons mediate diet-restriction-induced longevity in C. elegans. Nature 447(7144):545–549. doi:10.1038/nature05904
Lapierre LR, Gelino S, Melendez A, Hansen M (2011) Autophagy and lipid metabolism coordinately modulate life span in germline-less C. elegans. Curr Biol CB 21(18):1507–1514. doi:10.1016/j.cub.2011.07.042
Lapierre LR, De Magalhaes Filho CD, McQuary PR, Chu CC, Visvikis O, Chang JT, Gelino S, Ong B, Davis AE, Irazoqui JE, Dillin A, Hansen M (2013) The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in C. elegans. Nat Commun 4:2267. doi:10.1038/ncomms3267
Settembre C, Di Malta C, Polito VA, Garcia Arencibia M, Vetrini F, Erdin S, Erdin SU, Huynh T, Medina D, Colella P, Sardiello M, Rubinsztein DC, Ballabio A (2011) TFEB links autophagy to lysosomal biogenesis. Science 332(6036):1429–1433. doi:10.1126/science.1204592
Johnson DW, Llop JR, Farrell SF, Yuan J, Stolzenburg LR, Samuelson AV (2014) The C. elegans Myc-Mondo/Mad complexes integrate diverse longevity signals. PLoS Genet 10(4):e1004278. doi:10.1371/journal.pgen.1004278
Nakamura S, Karalay O, Jager PS, Horikawa M, Klein C, Nakamura K, Latza C, Templer SE, Dieterich C, Antebi A (2016) Mondo complexes regulate TFEB via TOR inhibition to promote longevity in response to gonadal signals. Nat Commun 7:10944. doi:10.1038/ncomms10944
Tullet JM, Hertweck M, An JH, Baker J, Hwang JY, Liu S, Oliveira RP, Baumeister R, Blackwell TK (2008) Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132(6):1025–1038. doi:10.1016/j.cell.2008.01.030
Bowerman B, Draper BW, Mello CC, Priess JR (1993) The maternal gene skn-1 encodes a protein that is distributed unequally in early C. elegans embryos. Cell 74(3):443–452
Vilchez D, Morantte I, Liu Z, Douglas PM, Merkwirth C, Rodrigues AP, Manning G, Dillin A (2012) RPN-6 determines C. elegans longevity under proteotoxic stress conditions. Nature 489(7415):263–268. doi:10.1038/nature11315
Steinbaugh MJ, Narasimhan SD, Robida-Stubbs S, Moronetti Mazzeo LE, Dreyfuss JM, Hourihan JM, Raghavan P, Operana TN, Esmaillie R, Blackwell TK (2015) Lipid-mediated regulation of SKN-1/Nrf in response to germ cell absence. Elife 4. doi:10.7554/eLife.07836
Wei Y, Kenyon C (2016) Roles for ROS and hydrogen sulfide in the longevity response to germline loss in C. elegans. Proc Natl Acad Sci U S A 113(20):E2832–E2841. doi:10.1073/pnas.1524727113
Miller DL, Budde MW, Roth MB (2011) HIF-1 and SKN-1 coordinate the transcriptional response to hydrogen sulfide in C. elegans. PLoS ONE 6(9):e25476. doi:10.1371/journal.pone.0025476
Hansen M, Hsu AL, Dillin A, Kenyon C (2005) New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a C. elegans genomic RNAi screen. PLoS Genet 1(1):119–128. doi:10.1371/journal.pgen.0010017
Hsu AL, Murphy CT, Kenyon C (2003) Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science 300(5622):1142–1145. doi:10.1126/science.1083701
Taubert S, Van Gilst MR, Hansen M, Yamamoto KR (2006) A mediator subunit, MDT-15, integrates regulation of fatty acid metabolism by NHR-49-dependent and -independent pathways in C. elegans. Genes Dev 20(9):1137–1149. doi:10.1101/gad.1395406
Yang F, Vought BW, Satterlee JS, Walker AK, Jim Sun ZY, Watts JL, DeBeaumont R, Saito RM, Hyberts SG, Yang S, Macol C, Iyer L, Tjian R, van den Heuvel S, Hart AC, Wagner G, Naar AM (2006) An ARC/mediator subunit required for SREBP control of cholesterol and lipid homeostasis. Nature 442(7103):700–704. doi:10.1038/nature04942
McCormick M, Chen K, Ramaswamy P, Kenyon C (2012) New genes that extend C. elegans lifespan in response to reproductive signals. Aging Cell 11(2):192–202. doi:10.1111/j.1474-9726.2011.00768.x
Sinha A, Rae R (2014) A functional genomic screen for evolutionarily conserved genes required for lifespan and immunity in germline-deficient C. elegans. PLoS ONE 9(8):e101970. doi:10.1371/journal.pone.0101970
Amrit FR, Steenkiste EM, Ratnappan R, Chen SW, McClendon TB, Kostka D, Yanowitz J, Olsen CP, Ghazi A (2016) DAF-16 and TCER-1 facilitate adaptation to germline loss by restoring lipid homeostasis and repressing reproductive physiology in C. elegans. PLoS Genet 12(2):e1005788. doi:10.1371/journal.pgen.1005788
O’Rourke EJ, Soukas AA, Carr CE, Ruvkun G (2009) C. elegans major fats are stored in vesicles distinct from lysosome-related organelles. Cell Metab 10(5):430–435. doi:10.1016/j.cmet.2009.10.002
Doane WW (1961) Developmental physiology of the mutant female sterile(2)adipose of Drosophila melanogaster. III. Corpus allatum-complex and ovarian transplantations. J Exp Zool 146:275–298
Butterworth FM, Bodenstein D (1968) Adipose tissue of Drosophila melanogaster. 3. The effect of the ovary on cell growth and the storage of lipid and glycogen in the adult tissue. J Exp Zool 167(2):207–217. doi:10.1002/jez.1401670209
Judd ET, Wessels FJ, Drewry MD, Grove M, Wright K, Hahn DA, Hatle JD (2011) Ovariectomy in grasshoppers increases somatic storage, but proportional allocation of ingested nutrients to somatic tissues is unchanged. Aging Cell 10(6):972–979. doi:10.1111/j.1474-9726.2011.00737.x
Strong L (1967) Feeding activity, sexual maturation, hormones, and water balance in the female African migratory locust. J Insect Physiol 13(4):495–507, doi:http://dx.doi.org/10.1016/0022-1910(67)90061-3
Thomsen E, Hamburger K (1955) Oxygen consumption of castrated females of the blow-fly, Calliphora Erythrocephala Meig. J Exp Biol 32(4):692–699
Pallier E, Aubert R, Lemonnier D (1980) Effect of diet and ovariectomy on adipose tissue cellularity in mice. Reprod Nutr Dev 20(3A):631–636
McElroy JF, Wade GN (1987) Short- and long-term effects of ovariectomy on food intake, body weight, carcass composition, and brown adipose tissue in rats. Physiol Behav 39(3):361–365
Fettman MJ, Stanton CA, Banks LL, Hamar DW, Johnson DE, Hegstad RL, Johnston S (1997) Effects of neutering on bodyweight, metabolic rate and glucose tolerance of domestic cats. Res Vet Sci 62(2):131–136
Crane SW (1991) Occurrence and management of obesity in companion animals. J Small Anim Pract 32(6):275–282. doi:10.1111/j.1748-5827.1991.tb00930.x
Corona G, Mannucci E, Forti G, Maggi M (2009) Hypogonadism, ED, metabolic syndrome and obesity: a pathological link supporting cardiovascular diseases. Int J Androl 32(6):587–598. doi:10.1111/j.1365-2605.2008.00951.x
Hahm JH, Kim S, DiLoreto R, Shi C, Lee SJ, Murphy CT, Nam HG (2015) C. elegans maximum velocity correlates with healthspan and is maintained in worms with an insulin receptor mutation. Nat Commun 6:8919. doi:10.1038/ncomms9919
Bohni R, Riesgo-Escovar J, Oldham S, Brogiolo W, Stocker H, Andruss BF, Beckingham K, Hafen E (1999) Autonomous control of cell and organ size by CHICO, a Drosophila homolog of vertebrate IRS1-4. Cell 97(7):865–875
Zhang H, Stallock JP, Ng JC, Reinhard C, Neufeld TP (2000) Regulation of cellular growth by the Drosophila target of rapamycin dTOR. Genes Dev 14(21):2712–2724
Liu X, Huh JY, Gong H, Chamberland JP, Brinkoetter MT, Hamnvik OP, Mantzoros CS (2015) Lack of mature lymphocytes results in obese but metabolically healthy mice when fed a high-fat diet. Int J Obes (Lond) 39(10):1548–1557. doi:10.1038/ijo.2015.93
Kim JY, van de Wall E, Laplante M, Azzara A, Trujillo ME, Hofmann SM, Schraw T, Durand JL, Li H, Li G, Jelicks LA, Mehler MF, Hui DY, Deshaies Y, Shulman GI, Schwartz GJ, Scherer PE (2007) Obesity-associated improvements in metabolic profile through expansion of adipose tissue. J Clin Invest 117(9):2621–2637. doi:10.1172/JCI31021
Naukkarinen J, Heinonen S, Hakkarainen A, Lundbom J, Vuolteenaho K, Saarinen L, Hautaniemi S, Rodriguez A, Fruhbeck G, Pajunen P, Hyotylainen T, Oresic M, Moilanen E, Suomalainen A, Lundbom N, Kaprio J, Rissanen A, Pietilainen KH (2014) Characterising metabolically healthy obesity in weight-discordant monozygotic twins. Diabetologia 57(1):167–176. doi:10.1007/s00125-013-3066-y
Hansen M, Flatt T, Aguilaniu H (2013) Reproduction, fat metabolism, and life span: what is the connection? Cell Metab 17(1):10–19. doi:10.1016/j.cmet.2012.12.003
Ackerman D, Gems D (2012) The mystery of C. elegans aging: an emerging role for fat. Distant parallels between C. elegans aging and metabolic syndrome? Bioessays 34(6):466–471. doi:10.1002/bies.201100189
Katewa SD, Demontis F, Kolipinski M, Hubbard A, Gill MS, Perrimon N, Melov S, Kapahi P (2012) Intramyocellular fatty-acid metabolism plays a critical role in mediating responses to dietary restriction in Drosophila melanogaster. Cell Metab 16(1):97–103. doi:10.1016/j.cmet.2012.06.005
Bruss MD, Khambatta CF, Ruby MA, Aggarwal I, Hellerstein MK (2010) Calorie restriction increases fatty acid synthesis and whole body fat oxidation rates. Am J Physiol Endocrinol Metab 298(1):E108–E116. doi:10.1152/ajpendo.00524.2009
Arner P, Bernard S, Salehpour M, Possnert G, Liebl J, Steier P, Buchholz BA, Eriksson M, Arner E, Hauner H, Skurk T, Ryden M, Frayn KN, Spalding KL (2011) Dynamics of human adipose lipid turnover in health and metabolic disease. Nature 478(7367):110–113. doi:10.1038/nature10426
Goodpaster BH, He J, Watkins S, Kelley DE (2001) Skeletal muscle lipid content and insulin resistance: evidence for a paradox in endurance-trained athletes. J Clin Endocrinol Metab 86(12):5755–5761. doi:10.1210/jcem.86.12.8075
Savary S, Trompier D, Andreoletti P, Le Borgne F, Demarquoy J, Lizard G (2012) Fatty acids – induced lipotoxicity and inflammation. Curr Drug Metab 13(10):1358–1370
Zhou YP, Grill V (1995) Long term exposure to fatty acids and ketones inhibits B-cell functions in human pancreatic islets of Langerhans. J Clin Endocrinol Metab 80(5):1584–1590. doi:10.1210/jcem.80.5.7745004
Wang MC, O’Rourke EJ, Ruvkun G (2008) Fat metabolism links germline stem cells and longevity in C. elegans. Science 322(5903):957–960. doi:10.1126/science.1162011
Haemmerle G, Moustafa T, Woelkart G, Buttner S, Schmidt A, van de Weijer T, Hesselink M, Jaeger D, Kienesberger PC, Zierler K, Schreiber R, Eichmann T, Kolb D, Kotzbeck P, Schweiger M, Kumari M, Eder S, Schoiswohl G, Wongsiriroj N, Pollak NM, Radner FP, Preiss-Landl K, Kolbe T, Rulicke T, Pieske B, Trauner M, Lass A, Zimmermann R, Hoefler G, Cinti S, Kershaw EE, Schrauwen P, Madeo F, Mayer B, Zechner R (2011) ATGL-mediated fat catabolism regulates cardiac mitochondrial function via PPAR-alpha and PGC-1. Nat Med 17(9):1076–1085. doi:10.1038/nm.2439
Chakravarthy MV, Pan Z, Zhu Y, Tordjman K, Schneider JG, Coleman T, Turk J, Semenkovich CF (2005) “New” hepatic fat activates PPARalpha to maintain glucose, lipid, and cholesterol homeostasis. Cell Metab 1(5):309–322. doi:10.1016/j.cmet.2005.04.002
Ferreira CR, Saraiva SA, Catharino RR, Garcia JS, Gozzo FC, Sanvido GB, Santos LF, Lo Turco EG, Pontes JH, Basso AC, Bertolla RP, Sartori R, Guardieiro MM, Perecin F, Meirelles FV, Sangalli JR, Eberlin MN (2010) Single embryo and oocyte lipid fingerprinting by mass spectrometry. J Lipid Res 51(5):1218–1227. doi:10.1194/jlr.D001768
Morgan NG, Dhayal S, Diakogiannaki E, Welters HJ (2008) The cytoprotective actions of long-chain mono-unsaturated fatty acids in pancreatic beta-cells. Biochem Soc Trans 36(Pt 5):905–908. doi:10.1042/bst0360905
Gonzalez-Covarrubias V, Beekman M, Uh HW, Dane A, Troost J, Paliukhovich I, van der Kloet FM, Houwing-Duistermaat J, Vreeken RJ, Hankemeier T, Slagboom EP (2013) Lipidomics of familial longevity. Aging Cell 12(3):426–434. doi:10.1111/acel.12064
Wileman T (2013) Autophagy as a defence against intracellular pathogens. Essays Biochem 55:153–163. doi:10.1042/bse0550153
Madeo F, Zimmermann A, Maiuri MC, Kroemer G (2015) Essential role for autophagy in life span extension. J Clin Invest 125(1):85–93. doi:10.1172/JCI73946
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B (2003) Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301(5638):1387–1391. doi:10.1126/science.1087782
O’Rourke EJ, Ruvkun G (2013) MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability. Nat Cell Biol 15(6):668–676. doi:10.1038/ncb2741
O’Rourke EJ, Kuballa P, Xavier R, Ruvkun G (2013) Omega-6 polyunsaturated fatty acids extend life span through the activation of autophagy. Genes Dev 27(4):429–440. doi:10.1101/gad.205294.112
Liu K, Czaja MJ (2013) Regulation of lipid stores and metabolism by lipophagy. Cell Death Differ 20(1):3–11. doi:10.1038/cdd.2012.63
Lapierre LR, Silvestrini MJ, Nunez L, Ames K, Wong S, Le TT, Hansen M, Melendez A (2013) Autophagy genes are required for normal lipid levels in C. elegans. Autophagy 9(3):278–286. doi:10.4161/auto.22930
Ghazi A, Henis-Korenblit S, Kenyon C (2007) Regulation of C. elegans lifespan by a proteasomal E3 ligase complex. Proc Natl Acad Sci U S A 104(14):5947–5952. doi:10.1073/pnas.0700638104
Carrano AC, Liu Z, Dillin A, Hunter T (2009) A conserved ubiquitination pathway determines longevity in response to diet restriction. Nature 460(7253):396–399. doi:10.1038/nature08130
Li W, Gao B, Lee SM, Bennett K, Fang D (2007) RLE-1, an E3 ubiquitin ligase, regulates C. elegans aging by catalyzing DAF-16 polyubiquitination. Dev Cell 12(2):235–246. doi:10.1016/j.devcel.2006.12.002
Vilchez D, Saez I, Dillin A (2014) The role of protein clearance mechanisms in organismal ageing and age-related diseases. Nat Commun 5:5659. doi:10.1038/ncomms6659
Ben-Zvi A, Miller EA, Morimoto RI (2009) Collapse of proteostasis represents an early molecular event in C. elegans aging. Proc Natl Acad Sci U S A 106(35):14914–14919. doi:10.1073/pnas.0902882106
Shemesh N, Shai N, Ben-Zvi A (2013) Germline stem cell arrest inhibits the collapse of somatic proteostasis early in C. elegans adulthood. Aging Cell 12(5):814–822. doi:10.1111/acel.12110
Vilchez D, Boyer L, Morantte I, Lutz M, Merkwirth C, Joyce D, Spencer B, Page L, Masliah E, Berggren WT, Gage FH, Dillin A (2012) Increased proteasome activity in human embryonic stem cells is regulated by PSMD11. Nature 489(7415):304–308. doi:10.1038/nature11468
Zhou KI, Pincus Z, Slack FJ (2011) Longevity and stress in C. elegans. Aging (Albany NY) 3(8):733–753
Keith SA, Amrit FR, Ratnappan R, Ghazi A (2014) The C. elegans healthspan and stress-resistance assay toolkit. Methods (San Diego, Calif) 68(3):476–486. doi:10.1016/j.ymeth.2014.04.003
Labbadia J, Morimoto RI (2015) Repression of the heat shock response is a programmed event at the onset of reproduction. Mol Cell 59(4):639–650. doi:10.1016/j.molcel.2015.06.027
TeKippe M, Aballay A (2010) C. elegans germline-deficient mutants respond to pathogen infection using shared and distinct mechanisms. PLoS ONE 5(7):e11777. doi:10.1371/journal.pone.0011777
Evans EA, Kawli T, Tan MW (2008) Pseudomonas aeruginosa suppresses host immunity by activating the DAF-2 insulin-like signaling pathway in C. elegans. PLoS Pathog 4(10):e1000175. doi:10.1371/journal.ppat.1000175
Alper S, McElwee MK, Apfeld J, Lackford B, Freedman JH, Schwartz DA (2010) The C. elegans germ line regulates distinct signaling pathways to control lifespan and innate immunity. J Biol Chem 285(3):1822–1828. doi:10.1074/jbc.M109.057323
Miyata S, Begun J, Troemel ER, Ausubel FM (2008) DAF-16-dependent suppression of immunity during reproduction in C. elegans. Genetics 178(2):903–918. doi:10.1534/genetics.107.083923
Medina DL, Di Paola S, Peluso I, Armani A, De Stefani D, Venditti R, Montefusco S, Scotto-Rosato A, Prezioso C, Forrester A, Settembre C, Wang W, Gao Q, Xu H, Sandri M, Rizzuto R, De Matteis MA, Ballabio A (2015) Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat Cell Biol 17(3):288–299. doi:10.1038/ncb3114
Hartge P (2009) Genetics of reproductive lifespan. Nat Genet 41(6):637–638. doi:10.1038/ng0609-637
De Vos M, Devroey P, Fauser BC (2010) Primary ovarian insufficiency. Lancet 376(9744):911–921. doi:10.1016/S0140-6736(10)60355-8
Perheentupa A, Huhtaniemi I (2009) Aging of the human ovary and testis. Mol Cell Endocrinol 299(1):2–13. doi:10.1016/j.mce.2008.11.004
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Amrit, F.R.G., Ghazi, A. (2017). Influences of Germline Cells on Organismal Lifespan and Healthspan. In: Olsen, A., Gill, M. (eds) Ageing: Lessons from C. elegans. Healthy Ageing and Longevity. Springer, Cham. https://doi.org/10.1007/978-3-319-44703-2_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-44703-2_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-44701-8
Online ISBN: 978-3-319-44703-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)