Abstract
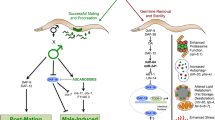
Classical evolutionary theories propose tradeoffs among reproduction, damage repair and lifespan. However, the specific role of the germline in shaping vertebrate aging remains largely unknown. In this study, we used the turquoise killifish (Nothobranchius furzeri) to genetically arrest germline development at discrete stages and examine how different modes of infertility impact life history. We first constructed a comprehensive single-cell gonadal atlas, providing cell-type-specific markers for downstream phenotypic analysis. We show here that germline depletion—but not arresting germline differentiation—enhances damage repair in female killifish. Conversely, germline-depleted males instead showed an extension in lifespan and rejuvenated metabolic functions. Through further transcriptomic analysis, we highlight enrichment of pro-longevity pathways and genes in germline-depleted male killifish and demonstrate functional conservation of how these factors may regulate longevity in germline-depleted Caenorhabditis elegans. Our results, therefore, demonstrate that different germline manipulation paradigms can yield pronounced sexually dimorphic phenotypes, implying alternative responses to classical evolutionary tradeoffs.








Similar content being viewed by others
Data availability
All raw RNA sequencing data (bulk and single-cell) and Hi-C data, as well as processed datasets, can be found in the Gene Expression Omnibus database under accession numbers GSE248741 and GSE218971, respectively. Metabolomics data are available in Supplementary Table 9. All other data are available from the corresponding author upon reasonable request. Individual supplementary tables are available in Mendeley Data (https://data.mendeley.com/datasets/ggys689v6x).
Code availability
The code supporting the current study is available in the following GitHub repository: https://github.com/Harel-lab/germline-regulation-of-the-vertebrate-lifespan. The Hi-C code is available at https://gitlab.com/mcfrith/last/-/blob/main/doc/last-papers.rst.
References
Ricklefs, R. E. Life-history connections to rates of aging in terrestrial vertebrates. Proc. Natl Acad. Sci. USA 107, 10314–10319 (2010).
de Magalhães, J. P., Costa, J. & Church, G. M. An analysis of the relationship between metabolism, developmental schedules, and longevity using phylogenetic independent contrasts. J. Gerontol. A Biol. Sci. Med. Sci. 62, 149–160 (2007).
Austad, S. N. & Hoffman, J. M. Is antagonistic pleiotropy ubiquitous in aging biology? Evol. Med. Public Health 2018, 287–294 (2018).
Kirkwood, T. B. L. Evolution of ageing. Nature 270, 301–304 (1977).
Kirkwood, T. B. L. & Austad, S. N. Why do we age? Nature 408, 233–238 (2000).
Schumacher, B., Garinis, G. A. & Hoeijmakers, J. H. J. Age to survive: DNA damage and aging. Trends Genet. 24, 77–85 (2008).
Chen, H. Y., Jolly, C., Bublys, K., Marcu, D. & Immler, S. Trade-off between somatic and germline repair in a vertebrate supports the expensive germ line hypothesis. Proc. Natl Acad. Sci. USA 117, 8973–8979 (2020).
Hsin, H. & Kenyon, C. Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399, 362–366 (1999).
Khodakarami, A., Saez, I., Mels, J. & Vilchez, D. Mediation of organismal aging and somatic proteostasis by the germline. Front. Mol. Biosci. 2, 3 (2015).
Hamilton, J. B. & Mestler, G. E. Mortality and survival: comparison of eunuchs with intact men and women in a mentally retarded population. J. Gerontol. 24, 395–411 (1969).
Min, K. J., Lee, C. K. & Park, H. N. The lifespan of Korean eunuchs. Curr. Biol. 22, R792–R793 (2012).
Benedusi, V. et al. Ovariectomy shortens the life span of female mice. Oncotarget 6, 10801 (2015).
Asdell, S. A., Doornenbal, H., Joshi, S. R. & Sperling, G. A. The effects of sex steroid hormones upon longevity in rats. Reproduction 14, 113–120 (1967).
Cargill, S. L., Carey, J. R., Müller, H. G. & Anderson, G. Age of ovary determines remaining life expectancy in old ovariectomized mice. Aging Cell 2, 185–190 (2003).
Hoffman, J. M., Creevy, K. E. & Promislow, D. E. L. Reproductive capability is associated with lifespan and cause of death in companion dogs. PLoS ONE 8, e61082 (2013).
Hamilton, J. B. Relationship of castration, spaying, and sex to survival and duration of life in domestic cats. J. Gerontol. 20, 96–104 (1965).
Torre, S., Della, Benedusi, V., Fontana, R. & Maggi, A. Energy metabolism and fertility—a balance preserved for female health. Nat. Rev. Endocrinol. 10, 13–23 (2014).
Gems, D., Kern, C. C., Nour, J. & Ezcurra, M. Reproductive suicide: similar mechanisms of aging in C. elegans and Pacific salmon. Front. Cell Dev. Biol. 9, 688788 (2021).
Antebi, A. Regulation of longevity by the reproductive system. Exp. Gerontol. 48, 596–602 (2013).
Uno, M. & Nishida, E. Lifespan-regulating genes in C. elegans. NPJ Aging Mech. Dis. 2, 16010 (2016).
Ermolaeva, M. A. et al. DNA damage in germ cells induces an innate immune response that triggers systemic stress resistance. Nature 501, 416–420 (2013).
Calculli, G. et al. Systemic regulation of mitochondria by germline proteostasis prevents protein aggregation in the soma of C. elegans. Sci. Adv. 7, eabg3012 (2021).
Austad, S. N. Sex differences in longevity and aging. In Handbook of the Biology of Aging 7th edn (eds Masoro, E. J. & Austad, S. N.) 479–495 (2011).
Tacutu, R. et al. Human Ageing Genomic Resources: new and updated databases. Nucleic Acids Res. 46, D1083–D1090 (2018).
Schartl, M. Beyond the zebrafish: diverse fish species for modeling human disease. Dis. Model. Mech. 7, 181–192 (2014).
Nielsen, J. et al. Eye lens radiocarbon reveals centuries of longevity in the Greenland shark (Somniosus microcephalus). Science 353, 702–704 (2016).
Kolora, S. R. R. et al. Origins and evolution of extreme life span in Pacific Ocean rockfishes. Science 374, 842–847 (2021).
Vrtílek, M., Žák, J., Pšenička, M. & Reichard, M. Extremely rapid maturation of a wild African annual fish. Curr. Biol. 28, R822–R824 (2018).
Harel, I. The turquoise killifish. Nat. Methods 19, 1150–1151 (2022).
Harel, I. et al. A platform for rapid exploration of aging and diseases in a naturally short-lived vertebrate. Cell 160, 1013–1026 (2015).
Astre, G., Moses, E. & Harel, I. The African turquoise killifish (Nothobranchius furzeri): biology and research applications. In Laboratory Fish in Biomedical Research (eds DʼAngelo, L. & de Girolamo, P.) 245–287 (Academic Press, 2021).
Astre, G. et al. Genetic perturbation of AMP biosynthesis extends lifespan and restores metabolic health in a naturally short-lived vertebrate. Dev. Cell 58, 1350–1364 (2023).
Harel, I., Valenzano, D. R. & Brunet, A. Efficient genome engineering approaches for the short-lived African turquoise killifish. Nat. Protoc. 11, 2010–2028 (2016).
Reichwald, K. et al. Insights into sex chromosome evolution and aging from the genome of a short-lived fish. Cell 163, 1527–1538 (2015).
Valenzano, D. R. et al. The African turquoise killifish genome provides insights into evolution and genetic architecture of lifespan. Cell 163, 1539–1554 (2015).
Rozenberg, I., Moses, E. & Harel, I. CRISPR–Cas9 genome editing in Nothobranchius furzeri for gene knockout and knock-in. Cold Spring Harb. Protoc. 2023, 90–99 (2023).
Wang, W. et al. Changes in regeneration-responsive enhancers shape regenerative capacities in vertebrates. Science 369, eaaz3090 (2020).
Moses, E., Franek, R. & Harel, I. A scalable and tunable platform for functional interrogation of peptide hormones in fish. eLife 12, e85960 (2023).
Slanchev, K., Stebler, J., de la Cueva-Mendez, G. & Raz, E. Development without germ cells: the role of the germ line in zebrafish sex differentiation. Proc. Natl Acad. Sci. USA 102, 4074–4079 (2005).
Kurokawa, H. et al. Germ cells are essential for sexual dimorphism in the medaka gonad. Proc. Natl Acad. Sci. USA 104, 16958–16963 (2007).
Kossack, M. E. & Draper, B. W. Genetic regulation of sex determination and maintenance in zebrafish (Danio rerio). In Current Topics in Developmental Biology (ed Capel, B.) 134, 119–149 (Elsevier, 2019).
Liu, Y. et al. Single-cell transcriptome reveals insights into the development and function of the zebrafish ovary. eLife 11, e76014 (2022).
Qian, P., Kang, J., Liu, D. & Xie, G. Single cell transcriptome sequencing of zebrafish testis revealed novel spermatogenesis marker genes and stronger Leydig-germ cell paracrine interactions. Front. Genet. 13, 851719 (2022).
Sun, X. et al. CloneSeq—single-cell clonal 3D culture and analysis protocol. STAR Protoc. 2, 100794 (2021).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Chan, J. T. H., Kadri, S., Köllner, B., Rebl, A. & Korytář, T. RNA-seq of single fish cells—seeking out the leukocytes mediating immunity in teleost fishes. Front. Immunol. 13, 798712 (2022).
Stévant, I. et al. Dissecting cell lineage specification and sex fate determination in gonadal somatic cells using single-cell transcriptomics. Cell Rep. 26, 3272–3283 (2019).
Li, L. et al. Single-cell RNA-seq analysis maps development of human germline cells and gonadal niche interactions. Cell Stem Cell 20, 858–873 (2017).
Estermann, M. A., Major, A. T. & Smith, C. A. Gonadal sex differentiation: supporting versus steroidogenic cell lineage specification in mammals and birds. Front. Cell Dev. Biol. 8, 1711 (2020).
Karlsson, M. et al. A single–cell type transcriptomics map of human tissues. Sci. Adv. 7, eabh2169 (2021).
Niu, S.-F. et al. Characterization of a novel piscidin-like antimicrobial peptide from Pseudosciaena crocea and its immune response to Cryptocaryon irritans. Fish Shellfish Immunol. 35, 513–524 (2013).
Yan, T. et al. Estradiol upregulates the expression of the TGF-β receptors ALK5 and BMPR2 during the gonadal development of Schizothorax prenanti. Animals 11, 1365 (2021).
Fan, X. et al. Single-cell reconstruction of follicular remodeling in the human adult ovary. Nat. Commun. 10, 3164 (2019).
Rotgers, E., Jørgensen, A. & Yao, H. H.-C. At the crossroads of fate—somatic cell lineage specification in the fetal gonad. Endocr. Rev. 39, 739–759 (2018).
Yoon, C., Kawakami, K. & Hopkins, N. Zebrafish vasa homologue RNA is localized to the cleavage planes of 2-and 4-cell-stage embryos and is expressed in the primordial germ cells. Development 124, 3157–3165 (1997).
Nishimura, T. & Tanaka, M. Gonadal development in fish. Sex. Dev. 8, 252–261 (2014).
Schulz, R. W. et al. Spermatogenesis in fish. Gen. Comp. Endocrinol. 165, 390–411 (2010).
Lubzens, E., Young, G., Bobe, J. & Cerdà, J. Oogenesis in teleosts: how fish eggs are formed. Gen. Comp. Endocrinol. 165, 367–389 (2010).
Li, J. & Cheng, C. H. K. Evolution of gonadotropin signaling on gonad development: insights from gene knockout studies in zebrafish. Biol. Reprod. 99, 686–694 (2018).
Liu, Y. & Lin, H. Genetic analysis of the reproductive axis in fish using genome-editing nucleases. Sci. Bull. (Beijing) 62, 302–308 (2017).
McBride, R. S. et al. Energy acquisition and allocation to egg production in relation to fish reproductive strategies. Fish Fisheries 16, 23–57 (2015).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Weidinger, G. et al. dead end, a novel vertebrate germ plasm component, is required for zebrafish primordial germ cell migration and survival. Curr. Biol. 13, 1429–1434 (2003).
Žák, J., Dyková, I. & Reichard, M. Good performance of turquoise killifish (Nothobranchius furzeri) on pelleted diet as a step towards husbandry standardization. Sci. Rep. 10, 8986 (2020).
Dyková, I. et al. Histology of major organ systems of Nothobranchius fishes: short-lived model species. J. Vertebr. Biol. 71, 21071–21074 (2022).
Minchin, J. E. N. & Rawls, J. F. A classification system for zebrafish adipose tissues. Dis. Model. Mech. 10, 797–809 (2017).
Liu, C., Peng, J., Matzuk, M. M. & Yao, H. H.-C. Lineage specification of ovarian theca cells requires multicellular interactions via oocyte and granulosa cells. Nat. Commun. 6, 6934 (2015).
Koley, S., Rozenbaum, M., Fainzilber, M. & Terenzio, M. Translating regeneration: local protein synthesis in the neuronal injury response. Neurosci. Res. 139, 26–36 (2019).
Blanco, S. et al. Stem cell function and stress response are controlled by protein synthesis. Nature 534, 335–340 (2016).
Riehle, K. J., Dan, Y. Y., Campbell, J. S. & Fausto, N. New concepts in liver regeneration. J. Gastroenterol. Hepatol. 26, 203–212 (2011).
Wang, S., Meyer, D. H. & Schumacher, B. H3K4me2 regulates the recovery of protein biosynthesis and homeostasis following DNA damage. Nat. Struct. Mol. Biol. 27, 1165–1177 (2020).
Lopez-Otin, C. et al. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Tyshkovskiy, A. et al. Identification and application of gene expression signatures associated with lifespan extension. Cell Metab 30, 573–593 (2019).
Shi, C. & Murphy, C. T. Sex and death. In Current Topics in Developmental Biology (eds Jarriault, S. & Podbilewicz, B.) 144, 353–375 (Elsevier, 2021).
Wang, C., Li, Q., Redden, D. T., Weindruch, R. & Allison, D. B. Statistical methods for testing effects on ‘maximum lifespan’. Mech. Ageing Dev. 125, 629–632 (2004).
Han, S. K. et al. OASIS 2: online application for survival analysis 2 with features for the analysis of maximal lifespan and healthspan in aging research. Oncotarget 7, 56147 (2016).
Garratt, M. et al. Male lifespan extension with 17‐α estradiol is linked to a sex‐specific metabolomic response modulated by gonadal hormones in mice. Aging Cell 17, e12786 (2018).
Stout, M. B. et al. 17α-Estradiol alleviates age-related metabolic and inflammatory dysfunction in male mice without inducing feminization. J. Gerontol. Ser. A Biomed. Sci. Med. Sci. 72, 3–15 (2017).
Blencowe, M. et al. Relative contributions of sex hormones, sex chromosomes, and gonads to sex differences in tissue gene regulation. Genome Res. 32, 807–824 (2022).
Nikkanen, J. et al. An evolutionary trade-off between host immunity and metabolism drives fatty liver in male mice. Science 378, 290–295 (2022).
Mann, S. N. et al. Health benefits attributed to 17α-estradiol, a lifespan-extending compound, are mediated through estrogen receptor α. eLife 9, e59616 (2020).
Garratt, M., Bower, B., Garcia, G. G. & Miller, R. A. Sex differences in lifespan extension with acarbose and 17‐α estradiol: gonadal hormones underlie male‐specific improvements in glucose tolerance and mTORC 2 signaling. Aging Cell 16, 1256–1266 (2017).
Arantes-Oliveira, N., Apfeld, J., Dillin, A. & Kenyon, C. Regulation of life-span by germ-line stem cells in Caenorhabditis elegans. Science 295, 502–505 (2002).
Ripa, R. et al. Refeeding-associated AMPKγ1 complex activity is a hallmark of health and longevity. Nat. Aging 3, 1544–1560 (2023).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes—a 2019 update. Nucleic Acids Res. 48, D445–D453 (2020).
Pang, Z. et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 49, W388–W396 (2021).
Lee, M. H., Malloy, C. R., Corbin, I. R., Li, J. & Jin, E. S. Assessing the pentose phosphate pathway using [2, 3‐13C2]glucose. NMR Biomed. 32, e4096 (2019).
Bennett, C. F. et al. Transaldolase inhibition impairs mitochondrial respiration and induces a starvation-like longevity response in Caenorhabditis elegans. PLoS Genet. 13, e1006695 (2017).
Harshman, L. G. & Zera, A. J. The cost of reproduction: the devil in the details. Trends Ecol. Evol. 22, 80–86 (2007).
Maklakov, A. A. & Chapman, T. Evolution of ageing as a tangle of trade-offs: energy versus function. Proc. R. Soc. B 286, 20191604 (2019).
Leroi, A. M. Molecular signals versus the Loi de Balancement. Trends Ecol. Evol. 16, 24–29 (2001).
Valenzano, D. R., Terzibasi, E., Cattaneo, A., Domenici, L. & Cellerino, A. Temperature affects longevity and age-related locomotor and cognitive decay in the short-lived fish: Nothobranchius furzeri. Aging Cell 5, 275–278 (2006).
Reichard, M. & POLAČIK, M. Reproductive isolating barriers between colour-differentiated populations of an African annual killifish, Nothobranchius korthausae (Cyprinodontiformes). Biol. J. Linn. Soc. 100, 62–72 (2010).
List, E. O. et al. Endocrine parameters and phenotypes of the growth hormone receptor gene disrupted (GHR−/−) mouse. Endocr. Rev. 32, 356–386 (2011).
Aleksic, S. et al. Integrity of hypothalamic–pituitary–testicular axis in exceptional longevity. Aging Cell 21, e13656 (2022).
Harrison, D. E. et al. Acarbose, 17‐α‐estradiol, and nordihydroguaiaretic acid extend mouse lifespan preferentially in males. Aging Cell 13, 273–282 (2014).
Williams, T. D. Mechanisms underlying the costs of egg production. Bioscience 55, 39–48 (2005).
Kleppe, L. et al. Sex steroid production associated with puberty is absent in germ cell-free salmon. Sci. Rep. 7, 12584 (2017).
Siegfried, K. R. & Nüsslein-Volhard, C. Germ line control of female sex determination in zebrafish. Dev. Biol. 324, 277–287 (2008).
Habermehl, T. L. et al. Extension of longevity and reduction of inflammation is ovarian-dependent, but germ cell-independent in post-reproductive female mice. Geroscience 41, 25–38 (2019).
Mason, J. B., Cargill, S. L., Anderson, G. B. & Carey, J. R. Transplantation of young ovaries to old mice increased life span in transplant recipients. J. Gerontol. A Biol. Sci. Med. Sci. 64, 1207–1211 (2009).
Toran-Allerand, C. D., Tinnikov, A. A., Singh, R. J. & Nethrapalli, I. S. 17α-Estradiol: a brain-active estrogen? Endocrinology 146, 3843–3850 (2005).
Lind, M. I. et al. Sex-specific growth and lifespan effects of germline removal in the dioecious nematode Caenorhabditis remanei. Preprint at bioRxiv https://doi.org/10.1101/2023.12.07.570570 (2023).
Barnes, A. I., Boone, J. M., Jacobson, J., Partridge, L. & Chapman, T. No extension of lifespan by ablation of germ line in Drosophila. Proc. R. Soc. B Biol. Sci. 273, 939–947 (2006).
Flatt, T. et al. Drosophila germ-line modulation of insulin signaling and lifespan. Proc. Natl Acad. Sci. USA 105, 6368–6373 (2008).
Zhao, Y. et al. Two forms of death in ageing Caenorhabditis elegans. Nat. Commun. 8, 15458 (2017).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Jao, L. E., Wente, S. R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl Acad. Sci. USA 110, 13904–13909 (2013).
Astre, G., Moses, E. & Harel, I. Laboratory fish in biomedical research. In Laboratory Fish in Biomedical Research (eds D’Angelo, L. & de Girolamo, P.) (Academic Press, 2021).
Mazutis, L. et al. Single-cell analysis and sorting using droplet-based microfluidics. Nat. Protoc. 8, 870–891 (2013).
Bavli, D. et al. CloneSeq: a highly sensitive analysis platform for the characterization of 3D-cultured single-cell-derived clones. Dev. Cell 56, 1804–1817.e7 (2021).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Carlson, M., Falcon, S., Pages, H. & Li, N. org.Hs.eg.db: genome wide annotation for human. R package version 3.8.2. https://doi.org/10.18129/B9.bioc.org.Hs.eg.db (2019).
Pagès, H., Carlson, M., Falcon, S. & Li, N. AnnotationDbi: manipulation of SQLite-based annotations in Bioconductor. R package version1. 54.1. https://bioconductor.org/packages/devel/bioc/manuals/AnnotationDbi/man/AnnotationDbi.pdf (2021).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Krueger, F. Trim Galore. https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Leek, J. T. Surrogate variable analysis (PhD thesis, Univ. of Washington, 2007).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Carlson, M., Falcon, S., Pages, H. & Li, N. org.Mm.eg.db: genome wide annotation for mouse. R package version 3.8.2. https://doi.org/10.18129/B9.bioc.org.Mm.eg.db (2019).
Dodzian, J., Kean, S., Seidel, J. & Valenzano, D. R. A protocol for laboratory housing of turquoise killifish (Nothobranchius furzeri). J. Vis. Exp. 57073 (2018).
Boocholez, H. et al. Neuropeptide signaling and SKN-1 orchestrate differential responses of the proteostasis network to dissimilar proteotoxic insults. Cell Rep. 38, 110350 (2022).
Rual, J.-F. et al. Toward improving Caenorhabditis elegans phenome mapping with an ORFeome-based RNAi library. Genome Res. 14, 2162–2168 (2004).
Malitsky, S. et al. Viral infection of the marine alga Emiliania huxleyi triggers lipidome remodeling and induces the production of highly saturated triacylglycerol. New Phytol. 210, 88–96 (2016).
Salem, M., Bernach, M., Bajdzienko, K. & Giavalisco, P. A simple fractionated extraction method for the comprehensive analysis of metabolites, lipids, and proteins from a single sample. J. Vis. Exp. 55802 (2017).
Zheng, L. et al. Fumarate induces redox-dependent senescence by modifying glutathione metabolism. Nat. Commun. 6, 6001 (2015).
Harel, I. & Brunet, A. The African turquoise killifish: a model for exploring vertebrate aging and diseases in the fast lane. Cold Spring Harb. Symp. Quant. Biol. 80, 275–279 (2015).
Van Keymeulen, A. et al. Epidermal progenitors give rise to Merkel cells during embryonic development and adult homeostasis. J. Cell Biol. 187, 91–100 (2009).
Harel, I. et al. Identification of protein aggregates in the aging vertebrate brain with prion-like and phase separation properties. Preprint at bioRxiv https://doi.org/10.1101/2022.02.26.482115 (2022).
Gruenbaum-Cohen, Y. et al. The actin regulator N-WASp is required for muscle-cell fusion in mice. Proc. Natl Acad. Sci. USA 109, 11211–11216 (2012).
Theis, S. et al. The occipital lateral plate mesoderm is a novel source for vertebrate neck musculature. Development 137, 2961–2971 (2010).
Benayoun, B. A. et al. Remodeling of epigenome and transcriptome landscapes with aging in mice reveals widespread induction of inflammatory responses. Genome Res. 29, 697–709 (2019).
Harel, I. et al. Pharyngeal mesoderm regulatory network controls cardiac and head muscle morphogenesis. Proc. Natl Acad. Sci. USA 109, 18839–18844 (2012).
Longenecker, K. & Langston, R. The Jungle Histology Atlas of Gonad Stages in Coral-Reef Fishes (Bishop Museum, 2016).
Imrie, D. & Sadler, K. C. White adipose tissue development in zebrafish is regulated by both developmental time and fish size. Dev. Dyn. 239, 3013–3023 (2010).
Abràmoff, M. D., Magalhães, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Intern. 11, 36–42 (2004).
Hollander-Cohen, L., Golan, M., Aizen, J., Shpilman, M. & Levavi-Sivan, B. Characterization of carp gonadotropins: structure, annual profile, and carp and zebrafish pituitary topographic organization. Gen. Comp. Endocrinol. 264, 28–38 (2018).
Liu, Z. et al. Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci. Rep. 7, 2193 (2017).
Wendler, S., Hartmann, N., Hoppe, B. & Englert, C. Age-dependent decline in fin regenerative capacity in the short-lived fish Nothobranchius furzeri. Aging Cell 14, 857–866 (2015).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Acknowledgements
We thank A. Zaslaver, E. Meshorer, A. Brunet, Y. Tzur, N. E. Stroustrup, M. Nitzan and the Harel laboratory for stimulating discussion and feedback on the manuscript. We thank A. Velan, E. Yanay, A. Abu-tair, Y. Birenbaum, F. Idrees and R. Barakat for help with killifish maintenance and N. Melamed-Book from the imaging facility (HUJI) and S. Malitsky and M. Itkin from the Life Sciences Core Facilities, Metabolic Profiling Unit (Weizmann Institute of Science). This study was supported by ERC StG no. 101078188 (I.H.), the Zuckerman Program (I.H.), the Abisch-Frenkel Foundation 19/HU04 (I.H.), ISF 2178/19 (I.H.), Israeli Ministry of Science 3-17631 (I.H.) and 3-16872 (I.H.), the Moore Foundation GBMF9341 (I.H.), BSF-NSF 2020611 (I.H.), the Israeli Ministry of Agriculture 12-16-0010 (I.H.), the Levi Eshkol Scholarship of the Israeli Ministry of Science (E.M.), the Pamela and Paul Austin Research Center on Aging fellowship (T.A.), the Czech Science Foundation (no. 22-01781O), the Ministry of Education, Youth and Sports of the Czech Republic (no. CZ.02.1.01/0.0/0.0/16_025/0007370) (R.F.), Chan Zuckerberg Initiative 2017-174468 and 2018-182817 (W.J.G.) and National Institutes of Health grants P50HG007735, UM1HG009442 and 1UM1HG009436 (W.J.G.).
Author information
Authors and Affiliations
Contributions
E.M., T.A. and I.H. designed the study. E.M. and R.F. performed experiments. E.M. generated the HPG and dnd1 mutant killifish lines. X.S. and E.M. prepared the single-cell RNA libraries, under the supervision of O.R. and I.H., respectively. T.A. designed and performed the analysis of RNA sequencing, under the supervision of I.H. A.S. performed worm lifespan experiments, under the supervision of E.C. T.A. and E.M. performed statistical analyses, with help from S.S. and D.M.Z. A.O.-G., G.K.M. and W.J.G. performed and analyzed the Hi-C data. T.A., E.M. and I.H. wrote the manuscript. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks David Vilchez and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Filtering germ cell clusters.
(a) Gene expression UMAP plot of piwil1, a germ cell marker gene. (b) Sub-clustering of the germ cells, performed according to Fig. 1c, color-coded by sex (left). Within this cluster, a marker of erythroid cells (top right) and previtellogenic oocytes (bottom right) is also indicated. A full list of erythroid marker genes can be found in Supplementary Table S3.
Extended Data Fig. 2 Pseudotime of germ cell clusters.
(a) Sub-clustering of the male germ cells excluding the female germ cells and erythroid cells. Cells are color-coded and labeled pseudo-time.
Extended Data Fig. 3 Perturbation and rescue of the reproductive axis in killifish.
(a) Histological sections from one-month-old males. n ≥ 4 individuals from each genotype (except for lhb, which was sex-linked, n = 1). Scale bar: 50 µm. Sperm developmental stages according to102: SG: spermatogonia; SC: spermatocytes; ST: spermatids; SZ: spermatozoa. (b) Quantification of sperm maturation, examples in (a). Data presented as proportion of each developmental stage. n ≥ 4 individuals for each experimental group (except for lhb). Significance was measured by a two-sided χ2 test with the WT proportion as the expected model. (c) Distribution of genotype progeny from heterozygous pairs (stratified by sex). n = 30-130 individuals, per genotype/sex. Significance was measured by a two-sided χ2 test with Mendelian proportions (25:50:25) as the expected model and FDR correction. (d) Quantification of somatic growth by calculating the standard length of one-month-old males (left) and females (right) of the indicated genotypes. n = 8-12 individuals for all experimental groups. Error bars indicate mean ± SEM. Significance was calculated using one-way ANOVA. (e) Quantification of male fertility. Each dot represents the number of eggs per breeding pair, per week of egg collection (except for lhb, which was sex-linked). n = 4-6 pairs for each group, over 4 collections. Error bars indicate mean ± SEM with individual points. Significance was calculated using one-way ANOVA. (f) Schematic illustration (left). Representative of n = 3 females. Quantification of fertility output (right) in lhbΔ8/Δ8 mutant females following plasmid electroporation. Each dot represents the number of eggs per breeding pair per week. n = 3-6 pairs over 4 collections. Error bars show mean ± SEM. Significance was calculated using a two-sided Student’s t-test. (g) smFISH in the ovaries of the indicated genetic models. Marker for immature germ-cells (ddx4) in red, and for fshr and lhr in green. Representative of n ≥ 6 individuals. Scale bar: 50 µm. (h) Oxford Grid plots (left) showing correspondence between the Hi-C linkage groups (Y-axis) and previously predicted chromosomes (X-axis). Positive strand in blue, and negative strand matches in red. Cumulative fraction of genes (right) located on Hi-C pseudochromosomes and on contigs in the original assembly (gray). In the Hi-C scaffolded assembly (black), most small contigs containing genes are placed into the 19 main pseudochromosomes.
Extended Data Fig. 4 Physiological characterization germline-depleted fish.
(a) Quantification of somatic growth (standard length) of WT or dnd1Δ4/Δ4 one-month-old mutants: males (top) and females (bottom), n = 8-12 individuals for all experimental groups. Error bars show mean ± SEM. Significance was measured using a two-sided Student’s t-test. (b) Representative images of adult (2.5-month-old) males (top) and females (bottom) WT or dnd1Δ4/Δ4 mutant fish. Black arrowheads highlight the presence of eggs in the female representative of n ≥ 4 individuals. Scale bar: 3.5 mm. (c) Distribution of genotype progeny from dnd1Δ4/+ heterozygous pairs (stratified by sex). Significance was measured by two-sided χ2 test with Mendelian proportions (25:50:25) as the expected model and FDR correction. n ≥ 500 fish for each group and p-values are indicated. (d) Tail-fin regeneration following whole-body irradiation in males. Genotypes and treatments are indicated (n = 4 individuals for non-irradiated WT and 6 individuals for WT and dnd1). The length of the outgrowth was calculated as the percentage of the original fin size (before the amputation). Error bars indicate mean ± SEM. Significance was calculated using repeated-measures two-way ANOVA with a Dunnet post-hoc compared to the irradiated WT, and p-values are indicated. (e) Tail-fin regeneration during aging, in males (left) and females (right). Fish sex, genotypes, and age are indicated (n = 5-7 individuals for all experimental groups). Error bars indicate mean ± SEM. Significance was calculated using repeated measures two-way ANOVA with a Tukey post-hoc, and p-values of the comparison between young and old fish of each genotype are indicated by color-coding.
Extended Data Fig. 5 Calibration of DNA damage detection.
(a) Representative images of apoptosis detected by the TUNEL assay (green and green arrows) and DNA (PI, in red). Assay was performed in gut sections of non-irradiated (left) or irradiated fish (right). n = 6 individuals in each experimental group. Scale bar: 50 µm.
Extended Data Fig. 6 Characterization of cell type markers and differential gene expression in dnd1Δ4/Δ4 mutant gonads.
(a-b) Dot-plot (top left, a, left, b) and UMAPs presenting the expression of select marker genes for males (a) and females (b). Dot-plot clusters are color-coded according to Fig. 6a, b (for males and females separately). Note that most of the markers overlap with Fig. 1c. A full list of markers can be found in Supplementary Tables S5, S6. (c) smFISH in ovaries for selected markers in dnd1Δ4/Δ4 mutants. Amh (red), a marker for Pre- and mature Granulosa cells, and cyp19a1 (green), a specific marker for mature Granulosa. Note that cyp19a1 is expressed in the adjacent adipose tissue. Representative of n ≥ 4 individuals. Scale bar: 20 µm. (d) Log2 fold-change heatmap represents the gene expression ratio between WT and dnd1Δ4/Δ4 fish of the indicated cell-types. The genes were selected as they were similarly altered in several cell types. (e) smFISH in WT fish and dnd1Δ4/Δ4 ovaries. Eef1a1, a marker for translation initiation (green), and ptgds, a marker for ovarian epithelium (in red). Representative of n ≥ 4 individuals. Scale bar: 20 µm.
Extended Data Fig. 7 Germline depletion extends male maximal lifespan.
(a) Quantile-Quantile (Q-Q) plots for the survival data of WT and dnd1Δ4/Δ4 fish (shown in Fig. 7b), assessed separately for males (left) and females (right). (b) Heatmap for normalized enrichment scores (NESs) in the male liver, comparing the response to gonadal and hormonal treatments in mice with germline-depleted young fish (left, selected pathways highlighted on the right). A full list of pathways can be found in Supplementary Table S8. Esr1: estrogen receptor KO; E2: estrogen treatment; T: testosterone treatment. (c) Lifespan of C. elegans from the glp-1(e2144) mutants, grown either at the permissive 15 °C (fertile, left) or restrictive 25 °C (germline depleted, right), and fed with either EV or eef1a1 RNAi. P-values were calculated according to log-rank. Worm numbers and raw data can be found Supplementary Table S4.
Extended Data Fig. 8 Germline depletion enhances regeneration under several stressors.
(a) PCA for hepatic transcript levels. Groups include male (blue) or female (red), WT (dark shades) or dnd1Δ4/Δ4 (light shades), either young (circles) or old (triangles). n = 3-4 samples per condition, and each symbol represents an individual fish. (b) Top, left: gene expression UMAP plot of dnd1 gene. Right and bottom: tail-fin regeneration of WT and dnd1morphant fish following whole-body irradiation. n = 4 individuals for WT and dnd1morphant males, 7 for WT and dnd1morphant females and 10 for non-irradiated males and females. Error bars indicate mean ± SEM. Significance was calculated using repeated-measures two-way ANOVA with a Dunnet post-hoc, compared to WT irradiated fish, and p-values are indicated. (c) Tail-fin regeneration of WT fish following treatment by chloroquine or paraquat. Treatments and sexes are indicated. n = 5 individuals for untreated females, 11 for chloroquine treated females, 12 for untreated and chloroquine treated males, and paraquat treated females and 15 for paraquat treated males. The same untreated fish were used as control for both treatments. Error bars indicate mean ± SEM. Significance was calculated using repeated-measures two-way ANOVA with a Sidak post-hoc, and p-values are indicated. (d) Tail-fin regeneration of WT and dnd1morphant fish following treatment by chloroquine or paraquat. Treatments and sexes are indicated. n = 5 individuals for chloroquine treated dnd1morphant males, 7 for paraquat treated dnd1morphant males and chloroquine treated dnd1morphant females, 8 for chloroquine treated WT and paraquat treated WT and dnd1morphant females, 9 for paraquat treated WT males and 10 for chloroquine treated WT males. Error bars indicate mean ± SEM. Significance was calculated using repeated-measures two-way ANOVA with a Sidak post-hoc, and p-values are indicated. (e) Right: Representative images of proliferation detected in the livers of male fish of the indicated experimental groups, following a 3-day treatment of EdU (green, see Methods) with DAPI (blue) for detecting nuclei. Representative of n = 5-7 individuals per group. Scale bar: 50 μm. Left: Quantification of the percentage of proliferating cells. Error bars indicate mean ± SEM. Significance was calculated using one-way ANOVA with a Tukey post-hoc, and p-values are indicated.
Supplementary information
Supplementary information
Supplementary Fig. 1 and Supplementary Methods
Source data
Source Data Fig. 1
Unprocessed images
Source Data Fig. 2
Unprocessed images
Source Data Fig. 3
Numerical source data
Source Data Fig. 3
Unprocessed images
Source Data Fig. 4
Numerical source data
Source Data Fig. 4
Unprocessed images
Source Data Fig. 5
Numericalsource data
Source Data Fig. 5
Unprocessed images
Source Data Fig. 6
Unprocessed images
Source Data Fig. 7
Numerical source data
Source Data Fig. 8
Numerical source data
Source Data Extended Data Fig. 3
Numerical source data
Source Data Extended Data Fig. 3
Unprocessed images
Source Data Extended Data Fig. 4
Numerical source data
Source Data Extended Data Fig. 4
Unprocessed images
Source Data Extended Data Fig. 5
Unprocessed images
Source Data Extended Data Fig. 6
Unprocessed images
Source Data Extended Data Fig. 7
Numerical source data
Source Data Extended Data Fig. 8
Numerical source data
Source Data Extended Data Fig. 8
Unprocessed images
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moses, E., Atlan, T., Sun, X. et al. The killifish germline regulates longevity and somatic repair in a sex-specific manner. Nat Aging 4, 791–813 (2024). https://doi.org/10.1038/s43587-024-00632-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-024-00632-0
- Springer Nature America, Inc.