Abstract
Biofuels are increasingly becoming popular due to the fact that they are renewable natural resources with low net emission of greenhouse gases. It is necessary to produce large enough biomass feedstock to get sustainable levels of biofuel production. Also, it is important to generate the biomass from non-food sources, and preferably using marginal lands. Therefore, it is important to plan a strategy to enhance the biomass production from non-food plants. Besides sugarcane, two of the plant species used to produce cellulosic ethanol are switchgrass and Miscanthus. To enhance the production of biomass in a unit area one can utilize breeding and biotechnological approaches. These would include genetic improvement of the species to withstand adverse conditions such as cold, drought, high salinity and attack by pathogens such as microbes. Also, it will be useful to modify plant architecture such as height and branching traits which are regulated by phytohormones. We discuss some of the strategies to genetically improve biofuel plant species in order to produce more biomass for future lignocellulosic ethanol production.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
8.1 Introduction
Biofuels are renewable and sustainable sources of energy that can be in the solid, liquid or gas forms. A major source of biofuels is the biomass of plants rendered as bioethanol, biodiesel and biogas. Biofuels are the natural alternative sources to fossil fuels and are environmentally friendly. The concept of biofuels is not new, with firewood as the most primitive form of solid biofuel used ever since the discovery of fire. In fact, wood is still being used for cooking food and to generate heat during winter in many parts of the world. The liquid form of biofuels is either vegetable oils or ethanol derived by fermentation of plant materials. The biogas produced by anaerobic digestion of animal manure and organic household wastes into gas (methane) used for cooking is also a biofuel. Biodiesel is obtained from the vegetable oils produced from several plant species including, oil palm, canola, soybean that are also used as food oils, and more recently, from non-food sources such as Jatropha seed oil. The liquid forms of biofuels are preferred over other forms due to the ease of storage and transportation; and in many cases these can directly replace petroleum fuels. Thus, the so-called “flex fuel vehicles” on the road today can use gasoline blended with 15–85% of bioethanol.
The world bioethanol production in 2010 was about 86 billion liters (Renewable Fuel Association: http://www.ethanolrfa.org/news/entry/global-ethanol-production-to-reach-85.9-billion-litres-22.7-billion-ga/). Bioethanol is currently produced mainly from corn starch in the USA and from sugarcane in Brazil. The use of food crops for fuel production affects the food chain and has the potential to lead to serious socioeconomic issues as reflected in escalating food price. Therefore, cellulosic ethanol is becoming a viable alternative for corn starch and sugarcane as the feedstock. Because cellulosic ethanol is produced from plant biomass such as crop residues (straw), forestry and wood waste it does not disturb the food chain. The use of bioethanol can greatly reduce the greenhouse gas (GHG) emission, which can reach up to 94% lower than gasoline GHG emission [1, 2]. Therefore, it is hoped that the use of more bioethanol in the coming decades can help to achieve the significant displacement of petroleum use mandated by the advanced energy initiative (AEI) in the USA [3, 4]. The AEI requires 30% reduction from the levels of 2005 petroleum use in the transportation sector to be replaced by domestically produced renewable bioethanol. Accordingly, numerous cellulosic ethanol production facilities are being opened or the existing facilities are expanding their capacities in the USA (Renewable Fuel Association).
Biomasses such as corn stover (stalk + leaves), rice straw and wheat straw are produced in large-scale as the by-products of food production and a large portion of it is going waste by getting burnt in the field and leading to more GHG emission. In 2009–2010, the world production of corn was about 890 million tons (mt) and at the proportion of 1:1 the corn stover produced will also be about 890 mt [5]. Similarly, around 730 mt of rice straw was reportedly produced in Africa, Asia, Europe and America, out which around 678 mt comes from Asia [6]. Also, the current global production of wheat is about 675 mt and the wheat grain to straw yield ratio is estimated at around 1:1.6 [7]. The yield of ethanol from corn grain is in the range of 400–500 liters/ton, and the yield of cellulosic ethanol from digestion of dried cellulosic biomass is (380 liters/ton) in the same range. Therefore, by not using the plant biomass from the major grain crops we are discarding an excellent renewable source of fuel. Nevertheless, it should be noted that even if the entire global non-grain biomass from the three main cereal crops (corn, wheat, rice) is used for ethanol fermentation, it can only yield about 25% of the annual use of petroleum in the world. Hence, we need to develop additional sources of lignocellulosic feedstock to generate higher amounts of bioethanol.
In addition to the agricultural by-products, fast growing grasses such as switchgrass (Panicum virgatum L.), Miscanthus X giganteus, reed canary and trees such as willows and hybrid poplar have been identified as dedicated biofuel crops. Of these, switchgrass and Miscanthus are the most favored candidates due to their low input needs and high yield that can be harvested with existing agricultural methods [8, 9]. There are varieties suitable for different ecosystems [10] with estimated net energy yield of over 60 GJ/hectare/year [1]. Similarly, Miscanthus has been shown to yield harvestable biomass between 30 and 60 t/hectare/year [4]. At the 30 t/hectare yield, it was estimated that 12 million hectares of US cropland can yield adequate volumes of ethanol (133 × 109 l) corresponding to about 20% of the annual gasoline used in the USA, and in comparison, corn starch grown in a similar land area would yield only about 49 × 109 liters of ethanol with much higher fertilizer needs and other inputs accounting for significantly higher GHG emission [4]. Hence, it is clear that the net GHG release will be highly reduced by using switchgrass and Miscanthus as feedstock for bioethanol.
To get sustainable amount of biomass for the future biofuel production needs it is important to enhance the biomass yield of these dedicated biofuel crops. In this chapter we will discuss some of the possible molecular and genetic strategies to enhance plant biomass.
8.2 Strategies for Enhancement of Biomass
The second-generation bioethanol production facilities depend on lignocellulosic biomass, unlike the first-generation bioethanol plants that use corn starch or sugar. Demands on agricultural land for food production are expected to increase significantly in the coming decades and hence use of marginal land to grow and harvest the highest possible levels of biomass using plants such as switchgrass and Miscanthus will contribute significantly to ensure sustainable production of renewable fuel in the future. In order to enhance their productivity, these grasses have to be targeted for intensive research aimed at improving the biomass yield and other attempts to change the characteristics of the chemical contents (e.g., lignin, hemicellulose, cellulose).
Expanding the industry to use biomass feedstock from the agricultural and forestry waste materials and enhanced plant biomass from biofuel crops from marginal lands might be the best ways to get more bioethanol and reduce net emission of GHG. Hence, it is important to develop strategies to increase the yield of plant biomass in a unit area of marginal land, and save the arable land for food production. In this context, the following strategies can be employed to enhance biomass production and ensure a sustainable and constant supply of lignocellulosic biomass for bioethanol production.
8.2.1 Genetic Basis of Plant Architecture
Plant architecture is one of the important points to be considered for biomass enhancement. It is clear that different plant species grow to different heights, sizes and shapes. The final size and shape are determined by genetic and environmental factors. Thus, it would be appropriate to conclude that plant architecture is determined and influenced by the genetic information and the environmental factors, respectively. The final shape of a mature plant is established by post-embryonic growth of the shoot apical meristem (SAM) and root apical meristem (RAM). SAM activity involves development of lateral organs such as leaves, flowers and branches as well as maintenance of the meristem identity in a pool of stem cells within the meristem. Recent data show that SAM is controlled by several genes such as SHOOTMERISTEMLESS, CLAVATA and WUSCHEL in dicotyledonous plants (e.g., Arabidopsis) and OSH1 and MOC1 in monocotyledonous plants (e.g., rice) (see [11] for detailed review). The involvement of various phytohormones such as cytokinin, gibberellin, auxin and abscisic acid in regulating shoot development has been well recognized by plant physiologists and developmental biologists. Therefore, it is interesting to note that besides the genes listed above, several key regulatory genes that influence shoot development have been identified, among which are phytohormone signaling intermediates such as ARR5, ARR6 and ARR7 [11].
A number of other genes are known to be involved in regulating branching. Table 8.1 lists some of the known mutants with increased or decreased branching (for review see [12, 13]). The process of branching could be viewed as a multipronged developmental event, because it will involve establishment of axillary meristem, development of axillary bud, promotion of the outgrowth of the branch by overcoming the apical dominance [13]. Therefore, one can expect to find genes regulating the various steps in this developmental program, and they can be the targets of genetic modification of branching.
Manipulation of selected genes that are involved in plant growth and development may lead to the increase in the biomass. For example, mutation in a cytochrome P450 gene called SUPERSHOOT resulted in significantly increased axillary bud growth and led to profuse branching and significant increase in biomass [14].
Likewise mutations in the MAX1 and MAX2 loci resulted in bushy shoots in Arabidopsis [15]. The presence of OsMAX gene family in rice suggests that similar functions may be conserved in monocotyledonous plants as well. Also, overexpression of a gene called OsSPL14 in rice increased shoot branching in the vegetative stage and panicle branching in the reproductive stage [16]. The feasibility of modifying plant architecture was demonstrated with the bahiagrass (Paspalum notatum), which is a low input requiring turf grass, but with the undesirable trait of tall seedheads. Application of plant growth retardants can lead to shorter stature, but long-term use of chemicals may lead to phytotoxicity and environmental pollution. Hence, in an attempt to modify the architecture to shorter tillers with shorter leaves, transgenic plants expressing ATHB16 gene were generated [17]. These transgenic plants expressing the repressor of cell expansion (ATHB16 gene) exhibited the more desirable shorter tiller phenotype, likely to be conferred by the transgene. The teosinte branched1 (tb1) gene in maize, and homologs in wheat, rice and Arabidopsis regulate tillering or branching [13]. The loss of function of the probable rice ortholog OsTB1 gene (fine culm 1) leads to increased tillering in rice, and its overexpression leads to decreased tillering [18]. Similarly, overexpression of the wild-type form of maize tb1 gene in wheat leads to decreased tillering, suggesting that this gene function is conserved among a variety of plant species [18]. The action of tb1 gene in sorghum (SbTB1) has been demonstrated to be under the control of phytochrome B, with suppression of the gene by the active Pfr form leading to promotion of tillering [19]. Conversely, when light conditions cause inactivation of phytochrome B, SbTB1 expression is increased and tillering is inhibited, which explains the light-mediated control of branching
This is supportive of the proposal of a combinatorial model of shoot development proposed according to which a series of independently regulated but overlapping programs modify a common set of processes leading to change from juvenile to mature phase [20]. This concept holds good and the identification of various regulatory genes and the complex genetic interactions among these genes as well as their interactions with biochemical (phytohormones) as well as environmental factors are beginning to emerge. Thus, a recent study showed that growing maize in clumps rather than equidistant planting under dryland conditions results in less tillering and biomass accumulation [21]. Due to the fact that plant architecture is significantly influenced by the phytohormones we will discuss how they may be used to enhance biomass in selected species.
8.2.2 Phytohormone-Related Genes and Developmental Regulation
Phytohormones control every aspect of plant growth and development, including seed germination, seedling growth, branching, plant height, flowering, seed development and senescence. A few major phytohormones and their roles in regulating plant growth and development are listed in Table 8.2. Also, phytohormones such as auxins, gibberellins, cytokinins and ethylene can modify fiber and wood formation during growth [22]. Auxin is required for cell division and axial plant growth and it helps to enforce apical dominance (where the shoot tip exerts inhibitory action on the axillary bud outgrowth). The primary site of biosynthesis of auxins is at the shoot tip. It is transported basipetally to other parts of the plant via an elaborate transport mechanism involving a number of members of the PIN family of proteins [23]. The involvement of auxins and cytokinins is proven in organ development and controlling organ size. Cytokinins help to break apical dominance and promote the outgrowth of lateral shoots. Hence, interactions of auxin and cytokinin control the shoot branching in plants [24, 25]. More recently, another phytohormone strigolactones has been shown to be necessary to inhibit shoot branching, and mutants in the biosynthesis pathway exhibit plants with more branches [26]. Increased gibberellin biosynthesis by ectopically expressing AtGA20ox promotes growth rate and biomass increase in hybrid aspen [27] and tobacco [28]. Furthermore, in a recent study silencing of AtGA2ox homolog in tobacco was demonstrated to enhance plant biomass [29].
The effect of phytohormones can be examined from the biosynthesis and their biological actions. Thus, plants exhibiting wide variations in structure have been observed when key genes involved in phytohormone biosynthesis and signaling have been mutated. The classic examples of gibberellin-deficient plants (e.g., Arabidopsis ga1-3 mutant; [30]) showing extreme dwarfism is a good illustration of the importance of this hormone in regulating plant architecture. This mutant arose from a deletion in the ent-kaurene synthase enzyme that catalyzes an early step in gibberellic acid biosynthesis. However, it retains the ability to respond to exogenously added gibberellins to grow to normal size. Mutants in other phytohormone biosynthetic pathways are also known to result in similarly striking changes in plant morphology.
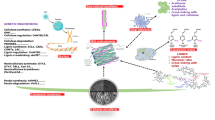
The discovery of specific receptors for the different phytohormones and their elaborate signaling pathways [31–33] is another area of interest for this discussion. The signaling cascade for cytokinins involves sequential phosphorylation and activation of intermediate proteins [34]. There are generally multiple receptors and intermediate proteins for the phytohormones. Thus, for cytokinin signaling, more than three receptors, five phosphotransfer proteins (cytoplasm to nucleus shunting) and over 20 response regulator proteins are known (Fig. 8.1). Similarly, auxin signaling cascade has multiple receptors and effector proteins [33]. Another major aspect of phytohormone signaling is the crosstalk between different phytohormones [35], which adds a new dimension of control of plant development by this group of rather simple chemical molecules. Mutants in various intermediates along the signaling pathway can lead to interesting agronomic traits such as altered organ size, altered branching and overall changes to plant architecture. Thus we observed that suppression of AtHOG1 expression, which is a putative cytokinin signaling intermediate, leads to enhanced branching in Arabidopsis and petunia [36]. It is not our intention to review phytohormone signaling in detail here, but this brief description is used to illustrate the genetic complexity of phytohormone signaling. Therefore, the various intermediates of the phytohormone signaling pathways may be explored as targets for genetic modification to achieve desired plant architecture.
Schematic representation of cytokinin signal transduction pathway (based on [34, 37]). This is an example of the signaling intermediates of one of the phytohormones. Similarly, the signaling pathways of other phytohormones have many intermediates, genes for which can be the targets of biotechnological improvements of biomass yield in selected plants. AHK2, 3, 4 Arabidopsis Histidine Kinase2, 3, 4 are cytokinin receptors on cell membranes. Dimers of the receptors bind cytokinins such as zeatin. AHP Arabidopsis Histidine Phosphotransfer proteins serve as phosphate shuttle from the cytoplasm to nucleus. ARR Arabidopsis Response Regulator proteins are the response regulators that affect the transcription of downstream target genes that are activated by cytokinins
8.2.3 Functional Genomics Approaches for Identification of Useful Genes
A widely used and accepted method of functional genomics is to disrupt the genes through mutations and study the effects in the following generations. There are several methods employed for this, e.g., the insertional mutagenesis such as T-DNA insertions in model plants such as Arabidopsis [38] and rice [39]. Also, transposon tagging is another method of choice for functional genomics in the model plants. The transposons or jumping genes were identified and isolated by Barbara McClintock from maize and it was cloned by [40]. The Ac/Ds system based on the transposons is being used as a tool for functional genomics in several plant systems [41, 42]. Using these methods one can generate and study a pool of insertion mutants in the biofuel crops and look for desirable phenotypes and genes associated with them. Such experiments will increase our understating of the genetics of biofuel crops and will open the doors for genetic modifications of such crops to enhance biomass and biofuel production. However, generating large number of mutants is not feasible in all cases, and to address that there are several alternate tools. For example, if the crop has synteny with model crops such as rice that can be tested in the biofuel species. Thus, an aluminum tolerance (Alt3) locus was mapped in rye using rice/rye synteny [43]. Another valuable reverse genetics technique called ‘Targeting Induced Local Lesions IN Genomes’ (TILLING), involves high throughput PCR screens of genomic DNA from M2 mutant populations induced by chemical mutagens [44, 45].
Genomic and functional genomics projects are being applied to one of the model grass species, Brachypodium, and its whole genome sequence has been released [46] similar to what has been achieved with the rice genome project. Also, functional genomics tools are being employed to the biomass crops such as switchgrass, Miscanthus and sorghum. These studies include genome-wide analysis of miRNA targets, developing low input switchgrass biomass using its bacterial endophytes and studying root physiology using root hair response to abiotic stresses. To study the gene functions there are other functional genomics approaches such as microarray studies which are useful to understand the global changes in gene expression. All these approaches will yield valuable information on the genetic nature of the biofuel crops, which have been ignored for a long time primarily due to the lack of investment in this area of research. Using these powerful genetic tools, one can generate and study a pool of insertion mutants in the biofuel crops and look for desirable phenotypes and genes associated with it. A better understating of genetic nature of biofuel crops will open the doors for genetic modifications to enhance biomass and biofuel production.
8.2.4 Plant Breeding
Plant breeding is the traditional way of improving plants by selecting for desirable phenotypes. In the simplest form, this can involve changing the ploidy of a plant to enhance the biomass production. Plant breeding is a laborious and time-consuming process that requires significant investment of resources as well, and perhaps it is for this reason there has been very little effort focused on plants used for biofuels compared to the crop species such as rice and wheat. Most of the biofuel crops are polyploid and display self-incompatibility [47]. It is a well-established fact that intensive plant breeding efforts during the early 1960s led to the production of high yielding dwarf and semi-dwarf hybrids of wheat, corn and rice, which formed the basis of the Green Revolution. Some of the key features modified were plant height, tillering habit and grain yield relative to straw yield. One of the more recent success stories of marker-assisted breeding is the submergence tolerant rice where the SUB1 locus was introgressed into several commercial cultivars of rice [48]. Hence, with the relatively high degree of synteny among grass species, opportunities exist for adaptation of observations from model species to the biofuel species by marker-assisted breeding. It is evident from these examples that grasses are amenable for considerable increases in yield and alterations to overall plant architecture. If concerted breeding efforts are applied for the biofuel crops, we can realize remarkable enhancements of these species as with the cereal crops during the Green Revolution.
8.2.5 Biotechnological Approaches to Further Improve Biofuel Crops
Biotechnological approaches are well known rapid ways of enhancing the plant traits. Genetic transformation of useful genes into the biofuel crops is demonstrated to be feasible. Thus, there are several successful reports on genetic transformation of switchgrass [49–52], Miscanthus [53] and sugarcane [54, 55]. Selected genes (or their homologs) that cause biomass enhancement in a given (crop) plant species can be the candidate genes for genetic transformation of biofuel crops. For example root-specific expression of cell wall invertase gene CIN1 from Chenopodium rubrum displayed enhanced shoot and root biomass in Arabidopsis [56]. Hence, similar genetic modification using either this gene or its homolog from the biofuel species may increase the shoot and root biomass. In another study, the overexpression of sugar metabolism enzymes such as UDP-glucose pyrophosphorylase, sucrose synthase and sucrose phosphate synthase was shown to result in increased plant biomass [57]. Suppression of Arabidopsis GA2ox homolog in tobacco enhanced fiber, wood formations and overall biomass yields [29]. Mutation in one of the WRKY transcription factors induces secondary wall formation in pith cells and leads to increased stem biomass in Medicago [58]. Also, delayed flowering will increase the biomass due to the availability of more time for vegetative growth. Thus, overexpression of floral repressor FLC in tobacco caused delayed flowering and as a result, the plants had accumulated significantly more biomass [59]. Similarly, another flowering time regulator mutant in maize called indeterminate1 (id1) showed delayed flowering and increase in biomass [60], suggesting that various candidate genes are already available for genetic modification of the plants used for cellulosic bioethanol production.
Although biofuel crops can grow in marginal land and yield significant amounts of biomass there are several potential problems with them. These include susceptibility to abiotic, biotic stresses and difficulties associated with conversion of cellulose into simple sugars during downstream processing. Prolonged cold and drought stresses may lead to significant yield loss, and to overcome this biofuel crops can be genetically engineered using proven cold- and drought-tolerant genes. Another major problem identified is biotic stresses such as insect and pest attack (e.g., plant-parasitic root nematodes) associated with decline in biomass production [61]. Generation of plants resistant to nematode and insect attacks might be the solution for this problem, which is possible to be achieved by genetic modification and biotechnological approaches.
Other approaches for genetic improvement of plants used for cellulosic ethanol production are to modify the chemical composition of the cell wall, specifically, to alter the lignocellulosic content or to incorporate genes for stable/inducible forms of enzymes such as cellulase into the plants so that downstream processing will be facilitated. Recently, genetic modification involving RNAi suppression of caffeic acid 3-O-methyltransferase (COMT) gene in switchgrass has been demonstrated to reduce lignin content and increase ethanol yield by up to 30% [51]. To convert the cellulose (which is a polymer of glucose units) to simple sugars either acid hydrolysis at high temperatures (with high energy input) or treatment with fungal cellulase enzyme is used. This is a rate limiting and costly step and to avoid this, temporal expression of cellulase gene in biofuel crops using specific promoters has been suggested. Efforts are underway in various laboratories to achieve this.
8.3 Conclusions and Future Perspectives
Bioethanol appears to have been firmly established as an important form of alternate fuel. With the second and later generation of bioethanol production focusing on the use of cellulosic biomass, the need for improvement of biomass plants is evident from the above discussion. Despite the occasional controversies raised, bioethanol is an environmentally friendly renewable energy source, and its large-scale use will lead to significant reduction in net emission of GHG. Alternate forms of biofuels such as oils to be used as biodiesel either from plants or from algae are also being explored. The emerging field of synthetic biology strives to convert microalgae into an efficient fuel oil production system. Although it is in its infancy, based on the underlying biological facts, synthetic biology for biofuel production by microalgae is expected to be successful in the coming decades.
It is important to phase out the use of food grains for fuel production in the coming decades. Because of the significant increase in demand for food grain expected, the conflicting demands on agricultural land will lead to serious social conflicts. Therefore, improving the efficiency and scaling up production of cellulosic ethanol is imperative. In order to achieve this, it is important to generate sufficient amounts of cellulosic biomass. Well over a trillion liters of ethanol (theoretical yield per year) can be obtained if all the available corn stover, rice straw and wheat straw (estimated 3 billion tons per year, [6]) are utilized for biofuel production. This represents one year’s oil demand of USA or approximately 25% of the annual world usage of petroleum. Currently, a significant amount of straw is either burnt and disposed off or used for animal feed. Therefore, use of non-food crop biomass plants becomes essential to broaden the availability of raw material for bioethanol production. Unlike with food crops, objections will be minimal if genetic modification strategies are applied to the biofuel plants to enhance yield, be tolerant to stresses and adverse growth conditions.
We have identified manipulation of the intermediates of phytohormone signaling pathways as an important strategy for enhancing plant biomass. The key developmental processes affecting biomass, which include reduced apical dominance and increased branching, plant height, leaf area and root to shoot ratio etc., are strongly influenced by phytohormones. The fact that phytohormones have pleiotropic effects on growth and development combined with the recent findings of the multiple signaling intermediates presents tremendous untapped opportunities for modifying specific traits listed above for improvement of the biofuel plants. The various signaling intermediates and downstream target genes can serve as candidates for biotechnological improvement or future marker-assisted breeding efforts.
The foregoing discussion has highlighted the need for and feasibility of using genetic and biotechnological approaches to enhance biomass production from a unit land area. Knowledge gained from model plants can be adapted to the biofuel crops in order to achieve this and to ensure sustainable biofuel production as a valuable alternative fuel in the decades to come.
References
Schmer MR, Vogel KP, Mitchell RB, Perrin RK (2008) Net energy of cellulosic ethanol from switchgrass. Proc Natl Acad Sci USA 105:464–469. doi:10.1073/pnas0704767105
Demirbas A (2009) Political, economic and environmental impacts of biofuels: a review. Appl Energ 86:S108–S117. doi:10.1016/j.apenergy.2009.04.036
Milliken J, Joseck F, Wang M, Yuzugullu E (2007) The advanced energy initiative. J Power Sources 172:121–131. doi:10.1016/j.jpowsour.2007.05.030
Heaton EA, Long SP, Dohleman FG (2008) Meeting US biofuel goals with less land: the potential of Miscanthus. Glob Change Biol 14:2000–2014. doi:10.1111/j.1365-2486.2008.01662.x
Kim S, Dale BE (2004) Global potential bioethanol production from wasted crops and crop residues. Biomass Bioenerg 26:361–375. doi:10.1016/j.biombioe.2003.08.002
Binod P, Sindhu R, Singhania RR, Vikram S, Devi L, Nagalakshmi S, Kurien N, Sukumaran RK, Pandey A (2010) Bioethanol production from rice straw: an overview. Bioresour Technol 101:4767–4774. doi:10.1016/j.biortech.2009.10.079
Engel R, Long D, Carlson G, Wallander R (2005) Estimating straw production of spring and winter wheat. Fertilizer Facts Montana State University Extention Bulletin No.33
Vogel KP, Brejda JJ, Walters DT, Buxton DR (2002) Switchgrass biomass production in the Midwest USA: harvest and nitrogen management. Agron J 94:413–420
McLaughlin SB, Kiniry JR, Taliaferro CM, Ugarte DD (2006) Projecting yield and utilization potential of switchgrass as an energy crop. Adv Agron 90:267–297. doi:10.1016/S0065-2113(06)90007-8
Parrish DJ, Fike JH (2009) Selecting, establishing, and managing switchgrass (Panicum virgatum L.) for biofuels. Methods Mol Biol 581:27–40. doi:10.1007/978-3-642-28418-2_2
Wang Y, Li J (2008) Molecular basis of plant architecture. Annu Rev Plant Biol 59:253–279. doi:10.1146/annurev.arplant.59.032607.092902
Bennett T, Leyser O (2006) Something on the side: axillary meristems and plant development. Plant Mol Biol 60:843–854. doi:10.1007/s11103-005-2763-4
Lewis JM, Mackintosh CA, Shin S, Gilding E, Kravchenko S, Baldridge G, Zeyen R, Muehlbauer GJ (2008) Overexpression of the maize Teosinte Branched1 gene in wheat suppresses tiller development. Plant Cell Rep 27:1217–1225. doi:10.1007/s00299-008-0543-8
Tantikanjana T, Yong JW, Letham DS, Griffith M, Hussain M, Ljung K, Sandberg G, Sundaresan V (2001) Control of axillary bud initiation and shoot architecture in Arabidopsis through the SUPERSHOOT gene. Genes Dev 15:1577–1588. doi:10.1101/gad.887301
Stirnberg P, van De Sande K, Leyser HM (2002) MAX1 and MAX2 control shoot lateral branching in Arabidopsis. Development 129:1131–1141
Miura K, Ikeda M, Matsubara A, Song X-J, Ito M, Asano K, Matsuoka M, Kitano H, Ashikari M (2010) OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat Genet 42:545–549. doi:10.1038/ng.592
Zhang HN, Altpeter F, Lomba P (2007) Improved turf quality of transgenic bahiagrass (Paspalum notatum Flugge) constitutively expressing the ATHB16 gene, a repressor of cell expansion. Mol Breeding 20:415–423. doi:10.1007/s11032-007-9101-2
Takeda T, Suwa Y, Suzuki M, Kitano H, Ueguchi-Tanaka M, Ashikari M, Matsuoka M, Ueguchi C (2003) The OsTB1 gene negatively regulates lateral branching in rice. Plant J 33:513–520
Kebrom TH, Burson BL, Finlayson SA (2006) Phytochrome B represses Teosinte Branched1 expression and induces sorghum axillary bud outgrowth in response to light signals. Plant Physiol 140:1109–1117. doi:10.1104/pp.105.074856
Poethig RS (1990) Phase-change and the regulation of shoot morphogenesis in plants. Science 250:923–930
Kapanigowda M, Stewart BA, Howell TA, Kadasrivenkata H, Baumhardt RL (2010) Growing maize in clumps as a strategy for marginal climatic conditions. Field Crop Res 118:115–125. doi:10.1016/j.fcr.2010.04.012
Aloni R (2001) Foliar and axial aspects of vascular differentiation: hypotheses and evidence. J Plant Growth Regul 20:22–34
Krecek P, Skupa P, Libus J, Naramoto S, Tejos R, Friml J, Zazimalova E (2009) The PIN-FORMED (PIN) protein family of auxin transporters. Genome Biol 10:249. doi:10.1186/gb-2009-10-12-249
Ongaro V, Leyser O (2007) Hormonal control of shoot branching. J Exp Bot 59:67–74. doi:10.1093/jxb/erm134
Shimizu-Sato S, Tanaka M, Mori H (2008) Auxin–cytokinin interactions in the control of shoot branching. Plant Mol Biol 69:429–435. doi:10.1007/s11103-008-9416-3
Umehara M, Hanada A, Yoshida S, Akiyama K, Arite T, Takeda-Kamiya N, Magome H, Kamiya Y, Shirasu K, Yoneyama K, Kyozuka J, Yamaguchi S (2008) Inhibition of shoot branching by new terpenoid plant hormones. Nature 455:195–200. doi:10.1038/nature07272
Eriksson ME, Israelsson M, Olsson O, Moritz T (2000) Increased gibberellin biosynthesis in transgenic trees promotes growth, biomass production and xylem fiber length. Nat Biotechnol 18:784–788
Biemelt S (2004) Impact of altered gibberellin metabolism on biomass accumulation, lignin biosynthesis, and photosynthesis in transgenic tobacco plants. Plant Physiol 135:254–265. doi:10.1104/pp103.036988
Dayan J, Schwarzkopf M, Avni A, Aloni R (2010) Enhancing plant growth and fiber production by silencing GA2-oxidase. Plant Biotechnol J 8:425–435. doi:10.1111/j.1467-7652.2009.00480.x
Michaels SD, Amasino RM (1999) The gibberellic acid biosynthesis mutant ga1-3 of Arabidopsis thaliana is responsive to vernalization. Dev Genet 25:194–198. doi:10.1002/(SICI)1520-6408(1999)
Bonetta D, McCourt P (2005) Plant biology: a receptor for gibberellin. Nature 437:627–628. doi:10.1038/437627a
Riefler M, Novak O, Strnad M, Schmulling T (2006) Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 18:40–54. doi:10.1105/tpc.105.037796
Mockaitis K, Estelle M (2008) Auxin receptors and plant development: a new signaling paradigm. Annu Rev Cell Dev Biol 24:55–80. doi:10.1146/annurev.cellbio.23.090506.123214
Argueso CT, Raines T, Kieber JJ (2010) Cytokinin signaling and transcriptional networks. Curr Opin Plant Biol 13:533–539. doi:10.1016/j.pbi.2010.08.006/s1369-5266(10)00108-1
Stamm P, Ramamoorthy R, Kumar PP (2011) Feeding the extra billions: strategies to improve crops and enhance future food security. Plant Biotechnol Rep 5:107–120. doi:10.1007/s11816-011-0169-0
Godge MR, Kumar D, Kumar PP (2008) Arabidopsis HOG1 gene and its petunia homolog PETCBP act as key regulators of yield parameters. Plant Cell Rep 27:1497–1507. doi:10.1007/s00299-008-0576-z
Wulfetange K, Lomin SN, Romanov GA, Stolz A, Heyl A, Schmulling T (2011) The cytokinin receptors of Arabidopsis are located mainly to the endoplasmic reticulum. Plant Physiol 156:1808–1818. doi:10.1104/pp111.180539
Krysan PJ, Young JC, Sussman MR (1999) T-DNA as an insertional mutagen in Arabidopsis. Plant Cell 11:2283–2290
Jeon JS, Lee S, Jung KH, Jun SH, Jeong DH, Lee J, Kim C, Jang S, Yang K, Nam J, An K, Han MJ, Sung RJ, Choi HS, Yu JH, Choi JH, Cho SY, Cha SS, Kim SI, An G (2000) T-DNA insertional mutagenesis for functional genomics in rice. Plant J 22:561–570
Fedoroff N, Wessler S, Shure M (1983) Isolation of the transposable maize controlling elements Ac and Ds. Cell 35:235–242. doi:10.1016/0092-8674(83)90226-X
Tissier AF, Marillonnet S, Klimyuk V, Patel K, Torres MA, Murphy G, Jones JD (1999) Multiple independent defective suppressor-mutator transposon insertions in Arabidopsis: a tool for functional genomics. Plant Cell 11:1841–1852
Ramachandran S, Sundaresan V (2001) Transposons as tools for functional genomics. Plant Physiol Bioch 39:243–252
Miftahudin T, Chikmawati K, Ross GJ Scoles, Gustafson JP (2005) Targeting the aluminum tolerance gene Alt3 region in rye, using rice/rye micro-colinearity. Theor Appl Genet 110:906–913. doi:10.1007/s00122-004-1909-0
McCallum CM, Comai L, Greene EA, Henikoff S (2000) Targeting induced local lesions IN genomes (TILLING) for plant functional genomics. Plant Physiol 123:439–442
Tadele Z, Esfeld K, Plaza S (2010) Applications of high-throughput techniques to the understudied crops of Africa. Aspects Appl Biol 96:223–240
The International Brachypodium Initiative (2010) Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 463:763–768. doi:10.1038/nature08747
Martinez-Reyna JM, Vogel KP (2002) Incompatibility systems in switchgrass. Crop Sci 42:1800–1805
Septiningsih EM, Pamplona AM, Sanchez DL, Neeraja CN, Vergara GV, Heuer S, Ismail AM, Mackill DJ (2009) Development of submergence-tolerant rice cultivars: the Sub1 locus and beyond. Ann Bot 103:151–160. doi:10.1093/aob/mcn206
Somleva MN, Tomaszewski Z, Conger BV (2002) Agrobacterium-mediated genetic transformation of switchgrass. Crop Sci 42:2080–2087
Xi Y, Fu C, Ge Y, Nandakumar R, Hisano H, Bouton J, Wang Z-Y (2009) Agrobacterium-mediated transformation of switchgrass and inheritance of the transgenes. BioEnerg Res 2:275–283. doi:10.1007/s12155-009-9049-7
Fu C, Mielenz JR, Xiao X, Ge Y, Hamilton CY, Rodriguez M, Chen F, Foston M, Ragauskas A, Bouton J, Dixon RA, Wang ZY (2011) Genetic manipulation of lignin reduces recalcitrance and improves ethanol production from switchgrass. Proc Natl Acad Sci USA 108:3803–3808. doi:10.1073/pnas.1100310108
Li R, Qu R (2011) High throughput Agrobacterium-mediated switchgrass transformation. Biomass Bioenerg 35:1046–1054. doi:10.1016/j.biombioe.2010.11.025
Zili Y, Zhou P, Chu C, Li X, Li X, Tian W, Wang L, Cao S, Tang Z (2004) Establishment of genetic transformation system for Miscanthus sacchariflorus and obtaining of its transgenic plants. High Tech Lett 10:27–31
Arencibia AD, Carmona ER, Tellez P, Chan MT, Yu SM, Trujillo LE, Oramas P (1998) An efficient protocol for sugarcane (Saccharum spp. L.) transformation mediated by Agrobacterium tumefaciens. Transgenic Res 7:213–222
Santosa DA, Hendroko R, Farouk A, Greiner R (2004) A rapid and highly efficient method for transformation of sugarcane callus. Mol Biotechnol 28:113–119. doi:10.1385/MB:28:2:113
Von Schweinichen C, Buttner M (2005) Expression of a plant cell wall invertase in roots of Arabidopsis leads to early flowering and an increase in whole plant biomass. Plant Biol (Stuttg) 7:469–475. doi:10.1055/s-2005-865894
Coleman HD, Beamish L, Reid A, Park JY, Mansfield SD (2010) Altered sucrose metabolism impacts plant biomass production and flower development. Transgenic Res 19:269–283. doi:10.1007/s11248-009-9309-5
Wang H, Avci U, Nakashima J, Hahn MG, Chen F, Dixon RA (2010) Mutation of WRKY transcription factors initiates pith secondary wall formation and increases stem biomass in dicotyledonous plants. Proc Natl Acad Sci USA 107:22338–22343. doi:10.1073/pnas.1016436107
Salehi H (2005) Delay in flowering and increase in biomass of transgenic tobacco expressing the floral repressor gene. J Plant Physiol 162:711–717. doi:10.1016/j.jplph.2004.12.002
Colasanti J, Yuan Z, Sundaresan V (1998) The indeterminate gene encodes a zinc finger protein and regulates a leaf-generated signal required for the transition to flowering in maize. Cell 93:593–603. doi:10.1016/S0092-8674(00)81188-5
Cassida KA, Kirkpatrick TL, Robbins RT, Muir JP, Venuto BC, Hussey MA (2005) Plant-parasitic nematodes associated with switchgrass (Panicum virgatum L.) grown for biofuel in the South Central United States. Nematropica 35:1–10
Acknowledgments
We thank Ms. Petra Stamm for helping to prepare Fig. 8.1. Research in the author’s laboratory is funded by the Science and Engineering Research Council (SERC Grant No.: 0921390036) of the Agency for Science Technology and Research, Singapore; and the National University of Singapore.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Ramamoorthy, R., Kumar, P.P. (2012). Molecular Genetic Strategies for Enhancing Plant Biomass for Cellulosic Ethanol Production. In: Baskar, C., Baskar, S., Dhillon, R. (eds) Biomass Conversion. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-28418-2_8
Download citation
DOI: https://doi.org/10.1007/978-3-642-28418-2_8
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-28417-5
Online ISBN: 978-3-642-28418-2
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)