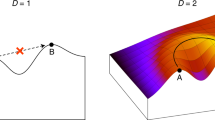
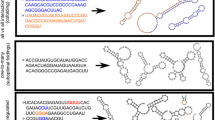
In this chapter, RNA secondary structures are used as an appropriate toy model to illustrate an application of the landscape concept to understand the molecular basis of structure formation, optimization, adaptation, and evolution in simple systems. Two classes of landscapes are considered (1) conformational landscapes mapping RNA conformations into free energies of formation and (2) sequence-structure mappings assigning minimum free energy structures to sequences. Even without referring to suboptimal conformations, optimization of RNA structures by mutation and selection reveals interesting features on the population level that can be interpreted by means of sequence-structure maps. The full power of the RNA model unfolds when sequence-structure maps and conformational landscapes are merged into a more advanced mapping that assigns a whole spectrum of conformations to the individual sequence. The scenario is complicated further – but at the same time made more realistic – by considering kinetic effects that allow for the assignment of two or more long-lived conformations, together with their suboptimal folds, to a single sequence. In this case, molecules can be designed, which fulfil multiple functions by switching back and forth from one stable conformation to the other or by changing conformation through allosteric binding of effectors. The evolution of noncoding RNAs is presented as an example for the application of landscape-based concepts.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
D. Thirumalai, Proc. Natl. Acad. Sci. 95, 11506 (1998)
D. Thirumalai, N. Lee, S.A. Woodson, D.K. Klimov, Annu. Rev. Phys. Chem. 52, 751 (2001)
D.E. Draper, RNA 10, 335 (2004)
M. Wu, I. Tinoco, Jr., Proc. Natl. Acad. Sci. USA 95, 11555 (1998)
S.R. Holbrook, Curr. Opt. Struct. Biol. 15, 302 (2005)
G. Varani, I. Tinoco, Jr., Q. Rev. Biophys. 24, 479 (1991)
S. Louise-May, P. Auffinger, E. Westhof, Curr. Opin. Struct. Biol. 6, 289 (1996)
P.F. Stadler, J. Math. Chem. 20, 1 (1996)
P. Schuster, P.F. Stadler, in Discrete Models of Biopolymers. ed. by M.J.C. Crabbe, M. Drew, A. Konopka. Handbook of Computational Chemistry (Marcel Dekker, New York, 2004) pp. 187-222
C.M. Reidys, P.F. Stadler, SIAM Rev. 44, 3 (2002)
E. Rivas, S.R. Eddy, J. Mol. Biol. 285, 2053 (1999)
I.L. Hofacker, P. Schuster, P.F. Stadler, Discr. Appl. Math. 89, 177 (1998)
J.S. McCaskill, Biopolymers 29, 1105 (1990)
M.S. Waterman, Introduction to Computational Biology: Maps Sequences and Genomes (Chapman and Hall/CRC, London/Boca Raton, 2000)
M.S. Waterman, T.F. Smith, Math. Biosci. 42, 257 (1978)
M. Tacker, P.F. Stadler, E.G. Bornberg-Bauer, I.L. Hofacker, P. Schuster, Eur. Biophys. J. 25, 115 (1996)
M. Zuker, D. Sankoff, Bull. Math. Biol. 46, 591 (1984)
J. Rogers, G. Joyce, Nature 402, 323 (1999)
J.S. Reader, G.F. Joyce, Nature 420, 841 (2002)
C. Flamm, W. Fontana, I.L. Hofacker, P. Schuster, RNA 6, 325 (1999)
W. Zhang, S.J. Chen, J. Chem. Phys. 118, 3413 (2003)
W. Zhang, S.J. Chen, J. Chem. Phys. 119, 8716 (2003)
M. Zuker, P. Stiegler, Nucleic Acids Res. 9, 133 (1981)
D.H. Turner, N. Sugimoto, Annu. Rev. Biophys. Chem. 17, 167 (1988)
A.E. Walter, D.H. Turner, J. Kim, M.H. Lyttle, P. Müller, D.H. Mathews, M. Zuker, Proc. Natl. Acad. Sci. USA 91, 9218 (1994)
D.H. Mathews, J. Sabina, M. Zuker, D.H. Turner, J. Mol. Biol. 288, 911 (1999)
D.H. Mathews, M.D. Disney, J.L. Childs, S.J. Schroeder, M. Zuker, D.H. Turner, Proc. Natl. Acad. Sci. USA 101, 7287 (2004)
I.L. Hofacker, W. Fontana, P.F. Stadler, L.S. Bonhoeffer, M. Tacker, P. Schuster, Mh. Chemie 125, 167 (1994)
I.L. Hofacker, Nucleic Acids Res. 31, 3429 (2003)
B. Bollobás, Random Graphs (Academic, London, 1985)
C. Reidys, P.F. Stadler, P. Schuster, Bull. Math. Biol. 59, 339 (1997)
W. Grüner, R. Giegerich, D. Strothmann, C. Reidys, J. Weber, I.L. Hofacker, P. Schuster, Mh. Chemie 127, 355 (1996)
W. Grüner, R. Giegerich, D. Strothmann, C. Reidys, J. Weber, I.L. Hofacker, P. Schuster, Mh. Chemie 127, 375 (1996)
P. Schuster, J. Biotechnol. 41, 239 (1995)
E. Schultes, D. Bartel, Science 289, 448 (2000)
D.M. Held, S.T. Greathouse, A. Agrawal, D.H. Burke, J. Mol. Evol. 57, 299 (2003)
Z. Huang, J.W. Szostak, RNA 9, 1456 (2003)
W. Fontana, D.A.M. Konings, P.F. Stadler, P. Schuster, Biopolymers 33, 1389 (1993)
P.G. Higgs, J. Phys. I (France) 3, 43 (1993)
J. Gevertz, H.H. Gan, T. Schlick, RNA 11, 853 (2005)
P. Clote, F. Ferré, E. Kranakis, D. Krizanc, RNA 11, 578 (2005)
M. Zuker, Science 244, 48 (1989)
S. Wuchty, W. Fontana, I.L. Hofacker, P. Schuster, Biopolymers 49, 145 (1999)
M.S. Waterman, T.H. Byers, Math. Biosci. 77, 179 (1985)
B.A. Shapiro, K. Zhang, Comput. Appl. Biosci. 6, 309 (1990)
C. Reidys, P.F. Stadler, Comput. Chem. 20, 85 (1996)
V. Moulton, M. Zuker, M. Steel, R. Pointon, D. Penny, J. Comput. Biol. 7, 277(2000)
M. Höchsmann, T. Töller, R. Giegerich, S. Kurtz, Proceedings of the Computational Systems Bioinformatics Conference, vol. 159 (Stanford, CA, CSB 2003)
M. Andronescu, A.P. Fejes, F. Hutter, H.H. Hoos, A. Condon, J. Mol. Biol. 336, 607 (2004)
J.R. Fresco, A. Adains, R. Ascione, D. Henley, T. Lindahl, Cold Spring Harb. Symp. Quant. Biol. 31, 527 (1966)
E.R. Hawkins, S.H. Chang, W.L. Mattice, Biopolymers 16, 1557 (1977)
V.L. Emerick, S.A. Woodson, Biochemistry 32, 14062 (1993)
R. Micura, C. Höbartner, Chembiochem 4, 984 (2003)
T. Baumstark, A.R. Schroder, D. Riesner, EMBO J. 16, 599 (1997)
A.T. Perrotta, M.D. Been, J. Mol. Biol. 279, 361 (1998)
C.K. Biebricher, S. Diekmann, R. Luce, J. Mol. Biol. 154, 629 (1982)
C.K. Biebricher, R. Luce, EMBO J. 11, 5129 (1992)
H. Zamora, R. Luce, C.K. Biebricher, Biochemistry 34, 1261 (1995)
C. Flamm, I.L. Hofacker, S. Maurer-Stroh, P.F. Stadler, M. Zehl, RNA 7, 254 (2000)
I. Abfalter, C. Flamm, P.F. Stadler, in Design of Multistable Nucleic Acid Sequences. ed. by H.W. Mewes, V. Heun, D. Frishman, S. Kramer. Proceedings of the German Conference on Bioinformatics (GCB 2003), vol. 1 (Belleville Verlag Michael Farin, München, 2003) pp.1-7
P. Clote, L. Gasieniec, R. Kolpakov, E. Kranakis, D. Krizanc, J. Theor. Biol. 236, 216 (2005)
E. Merino, C. Yanofsky, in Regulation by Termination-Antitermination: A Genomic Approach. ed. by A.L. Sonenshein, J.A. Hoch, R. Losick. Bacillus subtilis and its Closest Relatives: From Genes to Cells (ASM, Washington, DC, 2002) pp. 323-336
T.M. Henkin, C. Yanofsky, Bioessays 24, 700 (2002)
A.G. Vitreschak, D.A. Rodionov, A.A. Mironov, M.S. Gelfand, Trends Genet. 20, 44 (2004)
W.C. Winkler, R.R. Breaker, Chembiochem 4, 1024 (2003)
S. Brantl, Trends Microbiol. 12, 473 (2004)
E. Nudler, A.S. Mironov, Trends Biochem. Sci. 29, 11 (2004)
J.E. Barrick, K.A. Corbino, W.C. Winkler, A. Nahvi, M. Mandal, J. Collins, M. Lee, A. Roth, N. Sudarsan, I. Jona, J.K. Wickiser, R.R. Breaker, Proc. Natl. Acad. Sci. USA 101, 6421 (2004)
E.A. Lesnik, G.B. Fogel, D. Weekes, T.J. Henderson, H.B. Levene, R. Sampath, D.J. Ecker, Biosystems 80, 145 (2005)
R.R. Breaker, Curr. Opin. Biotechnol. 13, 31 (2002)
S.K. Silverman, RNA 9, 377 (2003)
M.T. Wolfinger, W.A. Svrcek-Seiler, C. Flamm, I.L. Hofacker, P.F. Stadler, J. Phys. A: Math. Gen. 37, 4731 (2004)
M. Eigen, Naturwissenschaften 58, 465 (1971)
M. Eigen, P. Schuster, Naturwissenschaften 64, 541 (1977)
M. Eigen, P. Schuster, Naturwissenschaften 65, 7 (1978)
J. Swetina, P. Schuster, Biophys. Chem. 16, 329 (1982)
M. Eigen, J. McCaskill, P. Schuster, Adv. Chem. Phys. 75, 149 (1989)
P. Jagers, Branching Processes with Biological Applications (Wiley, London, 1975)
P.A.P. Moran, The Statistical Processes of Evolutionary Theory (Clarendon, Oxford, UK, 1962)
L. Demetrius, P. Schuster, K. Sigmund, Bull. Math. Biol. 47, 239 (1985)
W. Fontana, P. Schuster, Biophys. Chem. 26, 123 (1987)
W. Fontana, W. Schnabl, P. Schuster, Phys. Rev. A 40, 3301 (1989)
W. Fontana, P. Schuster, Science 280, 1451 (1998)
D.T. Gillespie, J. Comput. Phys. 22, 403 (1976)
D.T. Gillespie, J. Phys. Chem. 81, 2340 (1977)
P. Schuster, in Molecular Insight into the Evolution of Phenotypes. ed. by J.P. Crutchfield, P. Schuster. Evolutionary Dynamics: Exploring the Interplay of Accident, Selection, Neutrality, and Function (Oxford University Press, New York, 2003) pp. 163-215
B.R.M. Stadler, P.F. Stadler, G.P. Wagner, W. Fontana, J. Theor. Biol. 213, 241(2001)
M. Kimura, The Neutral Theory of Molecular Evolution(Cambridge University Press, Cambridge, UK, 1983)
M.A. Huynen, P.F. Stadler, W. Fontana, Proc. Natl. Acad. Sci. USA 93, 397 (1996)
K. Grünberger, U. Langhammer, A. Wernitznig, P. Schuster, RNA evolution in Silico (Technical Report, Institut für Theoretische Chemie, Universität Wien, 2005)
D.T. Gillespie, J. Stat. Phys. 16, 311 (1977)
D.P. Bartel, C.Z. Chen, Nat. Genet. 5, 396 (2004)
O. Hobert, Trends Biochem. Sci. 29, 462 (2004)
J.S. Mattick, Bioessays 25, 930 (2003)
J.S. Mattick, Nat. Genet. 5, 316 (2004)
M. Szymański, M.Z. Barciszewska, M.Zywicki, J. Barciszewski, J. Appl. Genet. 44, 1 (2003)
S.R. Eddy, Nat. Genet. 2, 919 (2001)
M. Schöninger, A. von Haeseler, J. Mol. Evol. 49, 691 (1999)
B. Knudsen, J.J. Hein, Bioinformatics 15, 446 (1999)
N.J. Savill, D.C. Hoyle, P.G. Higgs, Genetics 157, 399 (2001)
J. Otsuka, N. Sugaya, J. Theor. Biol. 222, 447 (2003)
H. Jow, C. Hudelot, M. Rattray, P.G. Higgs, Mol. Biol. Evol. 19, 1591 (2002)
C. Hudelot, V. Gowri-Shankar, H. Jow, M. Rattray, P.G. Higgs, Mol. Phylogenet. Evol. 28, 241 (2003)
E. Rivas, R.J. Klein, T.A. Jones, S.R. Eddy, Curr. Biol. 11, 1369 (2001)
S. Washietl, I.L. Hofacker, P.F. Stadler, Proc. Natl. Acad. Sci. USA 102, 2454 (2005)
A.F. Bompfünewerer, C. Flamm, C. Fried, G. Fritzsch, I.L. Hofacker, J. Lehmann, K. Missal, A. Mosig, B. Müller, S.J. Prohaska, B.M.R. Stadler, P.F. Stadler, A. Tanzer, S. Washietl, C. Witwer, Theor. Biosci. 123, 301 (2005)
S. Carranza, J. Baguñà, M. Riutort, J. Mol. Evol. 49, 250 (1999)
A.P. Rooney, Mol. Biol. Evol. 21, 1704 (2004)
M.J. Telford, P.W.H. Holland, J. Mol. Evol. 44, 135 (1997)
F.E. Frenkel, M.B. Chaley, E.V. Korotkov, K.G. Skryabin, Gene 335, 57 (2004)
M. Eigen, B.F. Lindemann, M. Tietze, R. Winkler-Oswatitsch, A.W.M. Dress, A. von Haeseler, Science 244, 673 (1989)
M. Eigen, R. Winkler-Oswatitsch, Naturwissenschaften 68, 282 (1981)
S. Rodin, S. Ohno, A. Rodin, Proc. Natl. Acad. Sci. USA 90, 4723 (1993)
M. Di Giulio, J. Theor. Biol. 226, 89 (2004)
M.P. Terns, R.M. Terns, Gene Expr. 10, 17 (2002)
D. Lafontaine, D. Tollervey, Trends Biochem. Sci. 23, 383 (2002)
Y. Lee, K. Jeon, J.T. Lee, S. Kim, V.N. Kim, EMBO J. 21, 4663 (2002)
J. Hertel, M. Lindemeyer, K. Missal, C. Fried, A. Tanzer, C. Flamm, I.L. Hofacker, P.F Stadler, The Students of Bioinformatics Computer Labs 2004 and 2005. BMC Genomics 7, 25 (2006)
A. Tanzer, P.F. Stadler, J. Mol. Biol. 339, 327 (2004)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2007 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Schuster, P., Stadler, P.F. (2007). Modeling Conformational Flexibility and Evolution of Structure: RNA as an Example. In: Bastolla, U., Porto, M., Roman, H.E., Vendruscolo, M. (eds) Structural Approaches to Sequence Evolution. Biological and Medical Physics, Biomedical Engineering. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-35306-5_1
Download citation
DOI: https://doi.org/10.1007/978-3-540-35306-5_1
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-35305-8
Online ISBN: 978-3-540-35306-5
eBook Packages: Physics and AstronomyPhysics and Astronomy (R0)