Abstract
This paper presents the results of a study on genetic diversity from seed storage proteins of six species of plants from two families using sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE). The two families were; Melastomataceae (Senduduk); Melastoma malabathricum, Clidemia hirta and Lijndenia laurina and Fagaceae (Berangan, chesnut or timber); Lithocarpus wallichianus, Castanopsis lucida and Castanopsis schefferiana. The total numbers of polypeptide bands of Melastomataceae which resolved in 14% gel SDS were 33 bands, where three bands (9%) were monomorphic, while 30 bands (91%) were polymorphic with the size ranging from 12.89 to 95.24 kDa. In Fagaceae a total of 55 polypeptides bands were produced. Out of these one band (2%) was monomorphic among all three species and 54 (98%) were polymorphic. Cluster analysis for seed storage protein clearly distinguished between six species. Seed storage protein profiling of Melastomatecea and Fagaceae from this study highlighted a high degree of genetic diversity within the families. The high polymorphism shown by each family revealed that all the species are genetically variable. Therefore, it is recommended that the species should be conserved in order to establish in their natural habitat .
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Genetic diversity is the heritable variation within and between population of organisms and the conservation of plant genetic diversity is essential for present and future human wellbeing (Rao and Hodgkin 2002). The loss of genetic diversity of plants can effect in the loss of valuable and desirable traits and reduce options to use unexplored resources for food production , industry and medicine (Falk et al. 2001). A better understanding of genetic diversity of plant and its distribution is essential for its conservation and use. At the same time will improve our understanding of the taxonomy and origin and evolution of plant species of interest, therefore, some population need a sufficient genetic diversity to survive to a new condition for adaptation (Rao and Hodgkin 2002; Falk et al. 2001).
Morphological trait can be used for assessing genetic diversity in plants but it is often influenced by the environmental factors (Siddique and Naz 2009). Biochemical markers such as proteins can served as genetic markers as they are direct products of active genes and are quite polymorphic, heritable and highly independent of environmental fluctuations (Chittora and Purohit 2012; Gepts 1990). In order to study the genetic diversity of plant, sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) is one of the technique can be used to analyse the variation of polypeptide band of seeds storage protein (Ungureanu et al. 2007). In this study, two plant families have been chosen in order to study their genetic diversity; (a) Melastomataceae and (b) Fagaceae . Melastomataceae plants originate in the tropic and subtropic regions, with a total of more than 4000 species in the world and 22 species found in Southeast Asian region including Malaysia (Joffry et al. 2011). It is ubiquitous, species-rich, and dominant in the forest area (Silveira et al. 2013) either as herbs or shrubs or tress and the leaf lamina is dorsiventral or centric (Watson and Dallwitz 1992). Three species of Melastomataceae that have been chosen in this research were Melastoma malabathricum (Mm, Senduduk Ungu), Clidemia hirta (Ch, Senduduk Bulu) and Lijndenia laurina (Ll, Nipis Kulit). M. malabathricum has ethno-pharmacological value where certain parts of the plant can be used in medicinal treatment such as the leaves, shoots, barks, seeds and roots (Joffry et al. 2011). M. malabathricum (Fig. 1a) has gained herbal status and have been used to treat diarrhoea , dysentery, haemorrhoids , cuts and wounds, toothache, and stomach -ache (Joffry et al. 2011) while C. hirta (Fig. 1b) also has been used as medicinal treatment for traditional folk medicine to treat some bacterial infection (Dianita et al. 2011), treat for skin infection (Franca et al. 1996), and treat for venom fever (Latiff and Mat-Salleh 2002). Whereas, there is no known study on the medicinal status of L. laurina (Fig. 1c).
Another plant family have been studied was Fagaceae or chestnut family which have four genera and 64 species were found in the Malay Peninsula (Ng 1991) either as trees or shrubs and the leaf lamina is dorsiventral to bifacial (Watson and Dallwitz 1992). Many species of Fagaceae have important economic uses such as oak, chestnut, and beech are commonly used as timber for floors, furniture, cabinets, and wine barrels and it is widespread in the Northern hemisphere, with a centre of diversity found in tropical Southeast Asia (Nixon 1993; Manos et al. 2001). To understand more about the genetic diversity of Fagaceae, there were three species of Fagaceae have been studied in this research such as Castanopsis lucida (Cl) (Fig. 2a), Castanopsis schefferiana (Cs) (Fig. 2b) and Lithocarpus wallichianus (Lw) (Fig. 2c) were used in this study.
Castanopsis lucida, Cl (a), Castanopsis schefferiana, Cs (b) and Lithocarpus wallichianus, Lw (c). (From Wan Bayani et al. 2013)
Information regarding to the application of molecular technique in genetic study of Melastomatacea and Fagacea is very little. Therefore, this study was conducted to assess the genetic diversity of this two plants by using sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) with two different protein buffers; (a) Phosphate buffer and (b) Tris-HCL buffer (Chittora and Purohit 2012). The polypeptide bands were analyzed by using Dendrogram Unweighted Pair Group Method with Arithmetic (D-UPGMA) software.
Polymorphism of Polypeptide Bands
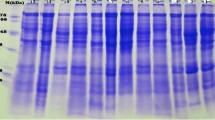
In this study, the protein samples were extracted by using two different buffers which were phosphate buffer and Tris-HCl extraction buffer. The range of the protein concentration from 0.1 to 2.0 mg/ml. Based on Jaccard similarity matrix for binary data , the relationships among species have been determined. Figure 3 shows the protein profile of Melastomatacea. The total protein band of seed protein extracts of three species of Melastomataceae from both buffers was 33 protein bands (range from 12.9 to 95.2 kDa), 30 were polymorphic (91%) and only three bands were monomorphic (9%) or common (32.12 kDa, 17.09 kDa and 15.34 kDa) among three species of Melastomataceae. However, three bands have been shared between Mm and Ch (51.86 kDa, 45.49 kDa and 28.17 kDa), two shared bands between Mm and Ll (63.33 kDa and 29.48 kDa), whereas between Ch and Ll only shared one band (64.42 kDa). The cluster analysis was done by using Unweighted Pair-Group Method of Arithmetic (UPGMA) to produce dendrogram (Fig. 4). When the lower similarity, it is means that higher genetic distance among species. Mm and Ch were clustered in the same group which the genetic distance was 0.76 whereas the genetic distance between Mm and Ch with Ll was 0.83 (the highest genetic distance). The similarity between Mm and Ch was more than between Mm and Ll or the most closely related species were between Mm and Ch, may be from their morphology, Mm and Ch look similar to each other.
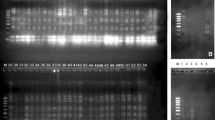
In Fagacea a total of 55 polypeptide bands were recorded from three species by using both extraction buffers (range from 151 to 380 kDa). Figure 5 shows the protein profile of Fagacea. Fifty-four bands were polymorphic (98%) and only one band (190 kDa) was monomorphic (2%) or common among three species. Five bands have been shared between Cl and Cs (330 kDa, 303 kDa, 180 kDa, 175 kDa and 166 kDa), four shared bands between Cl and Lw (196 kDa, 194 kDa, 193 kDa and 186 kDa), whereas between Cs and Lw shared two band (172 kDa and 182 kDa). The genetic distance between Cl and Cs was 0.77. Meanwhile the genetic distance between Cl and Cs to Lw was 0.88 (Fig. 6). The similarity between Cl and Cs was more than between Cl and Lw or the most closely related species were between Cl and Cs may be from the appearance of their seed, Cl and Cs look similar to each other.
Dendrogram of three species of Fagaceae. (From Wan Bayani et al. 2013)
As the comparison the genetic diversity by using SDS-PAGE in some plants indicated the following results; 5–11% polymorphism in three genotypes of Abrus precatorius (Chittora and Purohit 2012), 5–84% polymorphism in ten genotypes of wheat (Siddique and Naz 2009), 0–60% polymorphism in twenty genotypes of walnut (Khan et al. 2010), 0–80% polymorphism in eleven genotypes of Oryza sativa (Inamullah et al. 2010) and 0–100% polymorphism in nineteen genotypes of Capsicum (Akbar et al. 2010).
Conclusion
In conclusion, the genetic diversity of six species of two plants family were successfully determined from seed storage proteins by using SDS-PAGE. The seeds have an important genetic information of the plant and it must be conserved to avoid genetic loss by environmental factors or some diseases . This information will be useful as the basic genetic information for the species. The Melastomatacea is very useful as ethnomedicine since many races using this as their medicine and for Fagaceae , this family has important economic uses especially in timber industry.
References
Akbar, A., Ahmed, H., Ghafoor, S., & Khan, A. I. (2010). Phylogeny and genetic diversity studies in capsicum using seed storage proteins. Current Research Journal Biological Science, 72, 250–252.
Chittora, M., & Purohit, S. D. (2012). Optimization and analysis of seed protein diversity in Abrus precatorius genotypes with different seed coat colour. International Journal of Life Sciences Biotechnology and Pharma Research, 1, 268–277.
Dianita, R., Ramasamy, K., & Ab Rahman, N. (2011). Antibacterial activity of different extracts of Clidemia hirta (L.) D. Don leaves. Planta Medica, Journal of Medical Plant and Natural Product Research, 77, https://doi.org/10.1055/s-0031-1282769.
Falk, D. A., Knapp, E., & Guerrant, E. O. 2001. An introduction to restoration genetics. Society for Ecological Restoration. Retrieved from http://www.gov/pubs/rsetgene/restgenepdf. [12 February 2014].
Franca, F., Lago, E. L., & Marsden, P. D. (1996). Plants used in the treatment of leishmanial ulcers due to Leishmania (Vannia) brazilliensis in an endemic area of Bahia, Brazil. Revista da Sociedade Brasileira de Medicina Tropical, 9(3), 229–232.
Gepts, P. (1990). Genetic diversity of seed storage proteins in plants. In A. H. D. Brown, M. T. Clegg, A. L. Kahler, & B. S. Weir (Eds.), Plant population genetic, breeding and genet resource (pp. 64–82). Sunderland: Sinauer Associates.
Inamullah, K. I. A., Ahmed, H., Ghafoor, S., Gul, S., Muhammad, I., & Iiyas, M. (2010). Seed storage protein profile of rice varieties commonly grown in Pakistan. Asian Journal of Agricultural Science, 4, 120–123.
Joffry, S. M., Yob, N., Rofiee, M., Affandi, M., Suhaili, Z., Othman, F., Akim, A. M., Desa, M. N. M, & Zakaria, Z. (2011). Melastoma malabathricum (L.) Smith ethnomedicinal uses, chemical constituents, and pharmacological properties: A review. Evidence-based Complementary and Alternative Medicine, 2012, https://doi.org/10.1155/2012/258434.
Khan, M. W., Khan, I. A., Ahmed, H., Ali, H., Ghafoor, S., Afzal, M., Khan, F. A., Shah, M., & Afridi, S. G. (2010). Estimation genetic diversity in walnut. Pakistan Journal of Botany, 42, 1791–1796.
Latiff, A., & Mat-Salleh, K. (2002). Medicinal plants of Malaysia. Bangi: Universiti Kebangsaan Malaysia.
Manos, P. S., Zhou, Z. K., & Cannon, C. H. (2001). Systematics of Fagaceae: Phylogenetic tests of reproductive trait evolution. International Journal of Plant Sciences, 162(6), 1361–1379.
Ng, F. S. P. (1991). Variety of forest fruits, seeds & seedling. Malayan Forest Record, 1(34), 85–87.
Nixon, K. C. (1993). Infrageneric classification of Quercus (Fagaceae) and typification of sectional names. Annals of Forest Science, 50, 25–34.
Rao, V. R., & Hodgkin, T. (2002). Genetic diversity and conservation and utilization of plant genetic resources. Plant Cell, Tissue and Organ Culture, 68(1), 1–19.
Siddique, M. F., & Naz, N. (2009). Protein landmarks for diversity assessment in wheat genotypes. African Journal of Biotechnology, 8(9), 1855–1859.
Silveira, F. A. O., Fernandes, G. W., & Lemos-Filho, J. P. (2013). Seed and seedling ecophysiology of neotropical Melastomataceae: Implications for conservation and restoration of savannas and rainforests. Annals of the Missouri Botanical Garden, 99(1), 82–99.
Ungureanu, E., Artenie, V., & Ungureanu, A. (2007). SDS-PAGE analysis of protein from the seed of honeylocust (Gleditsia Triacanthos) during germination. Analele Stiintifice ale Universitatii “Alexandru Ioan Cuza” din Iasi Sec. II a. Genetica si Biologie Moleculara, 8, 31–35.
Wan Bayani, W. O., Hazlina A. Z., & Azhar, A. M. (2013). Optimization of SDS-PAGE conditions and analysis of seed protein diversity in Fagaceae (Chestnut) based on seed storage protein pattern from Taman Tropika Kenyir. The 13th Symposium of the Malaysian Society of Applied Biology. Cherating, Pahang, 8–10 June 2013. Malaysian Society of Applied Biology, Kuala Lumpur.
Watson, L., & Dallwitz, M. J. (1992). The families of the flowering plants: Descriptions, illustrations, identification and information. Retrieved from http://www.delta-intkey.com/angio.html.
Acknowledgements
The author wish to acknowledge University Malaysia Terengganu (UMT) for providing the opportunity to pursue this study. I would also like thanks MARDI at Tasik Kenyir for providing the samples and also wish thank you to Mr. Muhamad Razali Bin Salam for identify the samples.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Wan Omar, W.B., Zakeri, H.A., Salim, J.M., Fisall, U.F.M., Amran, M.A. (2019). Genetic Diversity of Seed Storage Protein in Selected Melastomataceae and Fagaceae from Tasik Kenyir. In: Abdullah, M., Mohammad, A., Nor Zalipah, M., Safiih Lola, M. (eds) Greater Kenyir Landscapes. Springer, Cham. https://doi.org/10.1007/978-3-319-92264-5_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-92264-5_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-92263-8
Online ISBN: 978-3-319-92264-5
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)