Abstract
Background
The narrow genetic diversity of chickpea is a serious impediment to modern cultivar creation. Seed storage proteins (SSPs) are stable and have minimal or no degradation when subjected to isolation and SDS-PAGE.
Methods and results
We have characterized SSPs of 436 chickpea genotypes, belonging to nine annual Cicer species, originated from 47 countries by SDS-PAGE and determined the extent of genetic diversity in chickpea through clustering. Based on scoring, a total of 44 bands (10 to 170 kDa) were identified, which were all polymorphic. The least appeared protein bands were 11, 160 and 170 kDa where band of 11 and 160 kDa was present exclusively in wild type. Five bands were present in < 10% of genotypes. Bands appeared in 200–300 genotypes were suggested less polymorphic, on contrary bands present in 10–150 genotypes were suggested more polymorphic. Polymorphism of protein bands in context to their potential functions reported in literature were explored and suggested that the glubulins were most and glutelins were least abundant, whereas albumins with their known role in stress tolerance can be used as marker in chickpea breeding. Cluster analysis produced 14 clusters, interestingly three clusters contained only Pakistani genotypes and thus Pakistani genotypes appeared as a separate entity from the rest of the genotypes.
Conclusion
Our results indicate that SDS-PAGE of SSPs is a powerful technique in determining the genetic diversity plus it is easily adaptable, due to its cost effectiveness in comparison to other genomics tools.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Legumes chosen for seeds as a portion of food are known as pulses. These are cool seasons, annual species. Pulses are an alternate dominating dietary source of protein other than meat [1]. Among legumes, chickpea (Cicer arietinum L.) occupies the central stage for world production, imports, and exports, because of its extraordinary nutritional values. Globally, chickpea is the third most important food legume after dry bean (Phaseolus vulgaris L.) and field pea (Pisum sativum L.) [2]. It belongs to the family Fabaceae [3] and the tribe Cicereae.
Chickpea seeds are rich in protein, have low fat & sodium, and zero cholesterol. They are also a great source of soluble and insoluble fibers, carbohydrates, vitamins, folate, and minerals [4, 5]. Because of its highly balanced composition of amino acid and protein bioavailability, it is also considered a worthy source of dietary protein [6]. In addition to its cheap availability because of its low price, its consumption is also related to the avoidance of cardiovascular disorders, handling type-2 diabetes, and reducing the levels of LDL cholesterol. The dietary insoluble fibers are related to the prevention of colon cancer as well as soluble fibers help in the maintenance of healthy body weight [4]. In addition, vitamins [tocopherol (both γ and α)], folic acid, riboflavin (B2), pantothenic acid (B5), pyridoxine (B6), and carotenoids such as β-carotene, lutein, cryptoxanthin, and zeaxanthin are also in ample amount in chickpea [7].
The origin and cultivation of chickpea started in the Middle East era ~ 7 millennia ago [1]. Since then chickpea has found its place in fields of temperate and semi-arid states of Asia, Europe, Australia, and North America. Currently, chickpea is grown in nearly 52 countries [8]. Despite that, approximately three-quarters of the global chickpea producers are coming from South Asian countries [9] where India is by far the largest producer providing 75% of the world’s chickpea yield [10], followed by Turkey, Pakistan, and Iran with approximately 7%, 6%, and 4% world supply, respectively.
Two main types of cultivated chickpea are “Desi” and “Kabuli” where “Kabuli” genotypes are characterized by a larger, rounder, and cream-colored seeds (mainly grown in North Africa, West Asia, North America, and Europe) and “desi” genotypes have smaller, angular-shaped and dark colored seeds with wrinkled seed coat, mostly grown in Asia and Africa [11]. The flower color of the “desi” type is pink with a semi-erect or semi-spreading growth type and is grown mainly in Ethiopia and the Indian subcontinent. The flower color of the “Kabuli” type is a white semi spreading growth habit and cultivated in temperate regions of the world [4].
The genus Cicer contains (nine annual and 34 perennial) species. According to morphological characteristics, life span, and geographical allocation, Cicer species are organized into four Sect. [12]. Section Cicer (= Monocicer M.G. Popov) is characterized by annual, small-flowered species with firm, erect to inclined or prostrate stems and imparipinnate leaves, or the rachis ending in a tendril (carries eight annual species); Section Chamaecicer M.G. Popov contains shrubby species with thin, creeping branches and 3–7 leaflets per leaf (carries two perennial species viz. C. chorassanicum, C. incisum); Section Polycicer M.G. Popov contains the perennial, rather large-flowered species with imparipinnate leaves or the rachis ending in a tendril (contains 23 perennial species); and Section Acanthocicer M. G. Popov includes perennial species with a large flower, spiny leaf rachis and spiny calyx teeth (carries seven woody perennial species). Among these four sections, “Cicer” (containing C. reticulatum, C. echinospermum, C. pinnatifidum, C. arietinum, C. bijugum, C. judaicum, C. yamashitae, and C. cuneatum) has attracted considerable attention of breeders [12].
Agricultural productivity is faced with dual challenges in the form of an increasing human population and irregular climate changes [13]. Ecosystems and environments have been altered because of changes in rainfall patterns and seasons. This situation dictates that joint efforts may be made to enhance the productivity of breeding programs toward accelerated climate smart varietal development [10, 14]. The situation becomes even more challenging in the case of chickpea because of its narrow genetic diversity that impedes the improvement and characterization of molecular markers for any trait [15]. To add to it, genetic bottlenecks are created [16] because of the use of few successful varieties for future breeding. Hence, to hasten the development of new varieties with increased annual yield, selection and use of diverse genotypes in a breeding program are required to create genetic variability [17].
The adaptive behavior of organisms is determined by the genetic diversity that it contains which is developed over a vast period of evolution [18]. The raw material in the form of diversity is the basis of introducing desired alleles in modern cultivars. This diversity can be expanded by using wild and cultivated relatives of a given species. The first step towards diversity enhancement is its assessment. Diversity can be gauged by taking into account the pedigree data [19] passport data, morphological data [20], biochemical data [21], storage proteins [22], and DNA-based marker data [23]. Among them, the use of storage protein profiling has been widely adopted to understand the latent evolutionary aspects of different crops [24].
One of the most economical and widely adopted biochemical methods to analyze the genetic relationships in germplasm is Sodium Dodecyl Sulphate Polyacrylamide Gel Electrophoresis (SDS-PAGE) [25]. Seed storage proteins have been identified as potent markers for; (a) discerning the diversity between and within species; (b) domestication for genetic resources and conservation and breeding, (c) to ascertaining the genetic relationships, and (d) the improvement of important agronomic and economic traits in crop plants [24]. The electrophoretic separation of seed storage protein has also been used to investigate the variations in the genome and to identify plant varieties [26]. Seed proteins are not susceptible to ecological fluctuation; the banding pattern of seed protein is highly stable which encouraged the identification of cultivars. Protein banding patterns also serve as an additional tool for cultivar identification or when the cultivars are to be patented [27].
Owing to the importance of chickpea and the selection and use of diverse genotypes in a breeding program with the larger aim to create genetic variability, we initiated this study to dissect the genomic diversity of 436 accessions of nine annual Cicer species by seed storage profiling using SDS-PAGE technique. The general objectives were to find out the polymorphism in seed storage proteins of the germplasm and to identify markers that could successfully differentiate various genotypes.
Materials and methods
Plant materials
This research was conducted in Plant Breeding and Genetics Division, Nuclear Institute for Agriculture and Biology (NIAB) Faisalabad. This study was accomplished on a set of 436 accessions of 9 annual Cicer species. The germplasm was acquired from Australian Grain Genebank is originated from 47 different countries. It comprises 305 cultivated and 131 wild type chickpea genotypes. Passport information of germplasm used in this study is provided in Table S1. The Pakistani accessions were available at NIAB.
Methods
Sample preparation
One seed from each accession was ground to a fine powder in pastel & mortar. The measured quantity of powder from each ground sample (0.05 g) was solubilized in 1ml Tris Urea Extraction Buffer [0.5 M Tri buffer (pH8), Urea (30.3 g/dry), SDS (10%), and 1ml Beta-merceptoethanol in total volume of 100ml] employing modified standard protocol for SDS-PAGE [28] in eppendorf tubes and centrifuged for 10 min at 14,000 rpm in a micro-centrifuge machine. The supernatant was collected and stored at −20 °C for subsequent analysis.
Protein estimation and SDS-PAGE
A dye-binding assay by [29] was carried out to estimate the protein concentrations in the supernatant. A volume of 2 µl of supernatant and 8 µl of sample preparation buffer i.e. extraction buffer mixed with tracking dye Bromophenol Blue (BPB) were loaded into wells and 5 µl of pre-stained broad range protein marker (Bio-Rad Catt log#161–0317) was loaded in a separate well. Electrophoresis was performed using a Mini-PROTEAN BIO-RAD electrophoresis system at a constant voltage (35 V) for 3–4 h in 1X running buffer (For 5X: 15 g of Tris-base, 72 g Glycine, and 5 g SDS in 1 L of distilled water).
After completion of SDS-PAGE, the gel was stained for 5–6 h on the shaker in a staining solution containing H2O, ethanol, and acetic acid with a ratio of 5:4:1, respectively, and 1 g of Coomassie brilliant blue R250 dye. Later on, the gel was destained in a destaining solution overnight. Destaining solution was the same as the staining solution without Comassie Brilliant Blue R250 dye. The gel picture was captured and stored for further data analysis.
Band scoring and data analyses
Each lane exhibited protein bands from each sample representing one genotype. Prestained marker bands were set as a landmark for the estimation of the molecular weight of protein bands on the gel. All the sample bands were compared to landmarks using the software PyElph 1.4. The analysis of images through software allowed easy and rapid determination of numbers and molecular masses of protein bands resolved on an SDS-PAGE gel. Furthermore, scoring was performed and electropherograms were produced by recording the presence (1) or absence (0) of each band.
Cluster analyses
Cluster analysis was performed in R Studio (version 2021.09.0 Build 351) using the packages “cluster” and “factoextra” which are equipped with different methods of clustering. K-means clustering was performed using the function “fviz_cluster (k2, data = X)” where k2 is defined by the user and symbolizes the presence of two sub-groups in the data “X”. The elbow method was implemented using the function, “fviz_nbclust(X, kmeans, method= “wss”)” where “wss” was defined as within the cluster sum of squares. The silhouette method was implemented using the function “fviz_nbclust(X, kmeans, method= “silhouette”)” where “silhouette” determines the quality ,of a clustering i-e, how well a value fits within its group and a high mean silhouette width is proportional to a good clustering. The “Gap statistic” method was computed using the “clusGap” function which results in providing a gap statistic and the standard error of the output. The following function was used: “gap_stat<-clusGap(X, FUN = kmeans, nstart = 25, K.max = 50, B = 50)” where “K.max” is defined as the maximum number of sub-groups in the dataset. The results were visualized using the function “fviz_cluster(gap_stat)”. After defining the optimal number of clusters (which in our case was 14), k-means clustering was performed by defining K = 14 using the function “optimal<-kmeans(X, 14, nstart = 25)” and visualized using the function “fviz_cluster(optimal, data = X)”. Further sub-groups in each cluster were identified using the “FactorMineR” package which computes the hierarchical clustering using the principal components of the individual dataset.
Results
Scoring and abundance of SSPs bands across genotypes
Based on scoring, a total of 44 bands were identified which were all polymorphic, their molecular weights were in the range of 10 kDa (kilodalton) to 170 kDa. The least appeared protein bands were found in some genotypes, for instance, a band of 170 kDa was detected in two wild types species only and was absent in all cultivated and wild type chickpea. Similarly, another band of 160 kDa was present in 23 wild type and two cultivated accessions only. However, five bands of sizes 140, 150, 120, 16, and 12 kDa were present in < 10% of the accessions. Another band of 11 kDa was present in 33 wild types but was absent in all cultivated types. The band size of 126, and 145 kDa were present in 46 accessions. Moreover, the band of 130 kDa was present in 58 accessions. Two bands sizes of 10 and 15 kDa were present in 56 accessions. Likewise, bands of sizes 14 and 17 kDa were present in 70 accessions each.
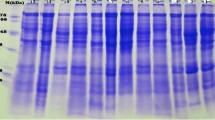
The 11 protein bands of sizes 115, 87, 75, 62, 60, 45, 48, 28, 27, 24, and 20 kDa were detected in up to 200 accessions. Likewise, peptides of sizes 113, 100, 98, 85, 70, 58, 55, 50, 30, 25, 21, and 13 kDa were detected in up to 300 genotypes of cultivated as well as wild type. Finally, the most abundant peptides were of the sizes 35 and 37 kDa detected in 398 and 395 accessions respectively. Additionally, there were five peptides of sizes 66, 42, 37, 35, and 33 kDa were detected in more than 300 genotypes. The results are described in detail for each band and accession in Table 1, and represented graphically in Fig. 1. A representation of the electrophoregram and names of the accessions is provided in Fig. 2.
A representation of electrophoregram; a serial number of accessions from Table S1, b protein banding patterns of the accessions on SDS-PAGE and Protein ladder (Broad range, Bio-Rad) with molecular weight (Dalton), c technical names of the accessions, d clusters of the respective accessions and e scoring of the bands according to which clusters were determined. Unique bands (present in only one accession) are highlighted in deep pink
Stratification and cluster analysis
We employed the three most commonly used methods viz. “elbow method” [30], “silhouette method” [31], and “gap statistic” [32] to determine the appropriate number of clusters in our germplasm. Using the “elbow method”, no optimal clusters could be determined in our dataset (Fig S1). The silhouette method indicated the presence of three possible clusters in our dataset (Fig S2). However, when the “silhouette method” model to cluster construction was fitted to three clusters, many accessions in clusters 1 and 3 were found concentrated together. As a result, we failed to differentiate among those accessions and hence, those accessions remained unrecognizable (Fig S3). Employing the “gap statistic” method [32], based on the presence or absence of protein bands, all genotypes could be divided into 14 clusters (Fig. 3) where the number of genotypes in a single cluster varied from eight (cluster 7) to 58 (cluster 12) (Fig. 4). However, an adequate amount of overlapping was also present in different clusters (Fig S4).
In addition to the main 14 clusters, the genotypes within each cluster were further sub-divided into several sub-clusters (Table S2-S15, Fig S5-S18). The number of sub-clusters within each cluster varied from three (clusters 2, 4, 8, 10, and 12) to six (cluster 3). Five subgroups were detected in cluster 9 whereas the rest of the clusters (1, 5, 6, 7, 11, 13, and 14) contained four sub-clusters (Fig S2-S15). Cluster 1 contained 48 genotypes originated from 16 different countries where the major contribution is from India (12 genotypes) and Turkey (13 genotypes) (Table S2). In addition, a total of 15 were advanced cultivars, three were breeder’s line, 21 were traditional cultivars and four were wild types. Except for the wild types (Cicer reticulatum), all were C. arietinum. The second, fourth, and seventh clusters contained 10, nine, and eight genotypes, correspondingly, all of which were elite cultivars originating from Pakistan (Tables S3, S5, and S8). The third cluster carried 31 genotypes originating from 10 different countries. Nine of them were advanced and three were traditional cultivars. The rest of the 19 genotypes were wild types. The wild types belonged to C. echinospermum (one genotype), C. reticulatum (8 genotypes), and C. judaicum (10 genotypes) (Table S4). Cluster 5 contained 34 wild types that originated pre-dominantly from Turkey (Table S5). There were 40 genotypes grouped in cluster 6 originating from Turkey (27 genotypes), Russia (eight genotypes), India (one genotype), Spain (one genotype), and Syria (one genotype) whereas the origin of two genotypes remained unknown (Table S7). In this cluster, the Syrian and other 26 Turkish genotypes were wild types while the Russian, Indian, Spanish and one Turkish genotype were cultivated types. The eighth cluster was comprised of nine cultivated types originating from different countries (Table S9). There were 37 genotypes in cluster 9 of cultivated type (advanced or traditional cultivars or elite lines) originating from many different countries including 14 Syrian genotypes (Table S10). The 10th cluster carried 51 genotypes originating from 14 different countries where all except one were cultivated types (Table S11). Cluster 11 contained 34 genotypes, among those all were cultivated species and most of them (19) have Indian origin, and three genotypes were from Turkey. (Table S12). In cluster 12, 50% of the genotypes originated from India (15 genotypes) and Turkey (14 genotypes) (Table S13). In the 13th cluster, there were 35 genotypes where 26 were cultivated types and 9 were wild type belonging to 15 different countries (Table S14). Among them, the wild types originated from Israel (two genotypes), Morocco, Russia, Syria, and Turkey (four genotypes). The 14th cluster carried 32 genotypes originating from Turkey (except one genotype that originated from Israel), all of which were wild types (Table S15).
Discussion
Among old techniques, one of the reliable and economic techniques to study population genetics is the SDS-PAGE of seed storage proteins because of little or no influence of environmental fluctuations on them [33]. Earlier, the characterization of seeds storage proteins are part of the basic and applied studies of any crop plant research [34]. Plant populations of chickpea [25, 35] and cowpea [34] have successfully been differentiated based on seeds storage proteins. This differentiation is necessary to select superior but diverse parents for future breeding [23].
Polymorphism of protein bands in context to their potential functions reported in previous studies
Seed storage proteins are proteins that accumulate significantly in the developing seed. The majority of seeds contain one to many groups of proteins, which serves as a source of amino acids, metal ions, nitrogen, carbon, and sulfur. These proteins are rapidly mobilized during seed germination [36]. In chickpea, globulins are the dominant storage proteins and account for 50–90% of seed protein followed by glutelins (18.10%), albumins (12.0%), prolamins (2.8%), and others [37]. These groups of seed storage proteins have been characterized and assigned a specific range of molecular weights which helps to locate the presence and absence of these proteins on a Coomassie stained SDS-PAGE gel. For example, according to previous reports the protein bands of 10, 11, and 12 kDa correspond to albumin (2 S) proteins [38]. Likewise, bands of 15, 18, 33, 35, 37, and 50 kDa correspond to vicilin (7 S) proteins [39]. Similarly, peptides of sizes 24, 25, and 42 belong to legumin (11 S) proteins whereas those of sizes 55 and 58 kDa belong to glutelin proteins [40, 41]. Besides, we have identified 44 protein bands across all 436 chickpea genotypes which could belong to albumin, vicilin, legumin, and glutelin proteins.
Globulins
Globulins are the most abundant form of seed storage proteins and were monomorphic as these were found in all genotypes. Vicilins together with legumins form globulins fraction of the legume seeds representing 70% of SSPs [42]. Legumins constitute 97% of globulins and are considered a major impetus during germination and seedling growth in chickpea [43]. Our results validate these findings as globulins were present in all accessions and were most abundant. All accessions carried at least one of the six vicilin (7 S) proteins (Fig. 5a). A total of 17 accessions exhibited one vicilin protein. Likewise, another 17 accessions showed all the six vicillin protein bands. Forty-three accessions had at least two vicilin proteins whereas 139 and 150 accessions carried three and four vicillin proteins, respectively. Among all 66 accessions carried five vicilin proteins. Legumin (11 S) proteins were present in 399 of the total 436 accessions whereas 57 accessions carried legumins of all sizes (Fig. 5b). In addition, 112 and 230 accessions correspondingly carried two legumins.
Glutelins
Albumins
Sulfur amino acids constitute a large proportion of Albumin proteins. Our results described that two, out of 3 peptides of Albumin (2 S) proteins were present in 80 accessions out of which 61 were wild and 19 were cultivated species (Fig. 5d), however, 10 wild type accessions from Turkish origin carried all three peptides. All 19 cultivated types carried either 10 (9 accessions from Pakistan) or 12 kDa (nine Russian and one Tajik accession) peptides of Albumin. Albumins are known to influence certain physiological functions including activities of lipoxygenases, glycosidases, or proteases involved in the degradation of storage proteins. Lipoxygenases influenced longevity in wheat [46]. Other Albumins like lectins are reported to play roles in plant’s defense against various stresses [39, 47]. Furthermore, our results need to be evaluated for their potential role in stress tolerance and use Albumin protein bands as a protein marker for germplasm screening against plant stresses.
Cluster analysis
Sometimes, agronomic characteristics and ecotypes are not sufficient to differentiate between various accessions of the same plant species, nevertheless, cluster analysis based on the SDS-PAGE scoring of seed storage proteins (SSPs) has differentiated among Vigna radiata and Vigna mungo accessions [24]. Clustering finds subgroups of observations within a dataset based on similarity, were observations in the same group tend to be similar and observations in different groups tend to be dissimilar. K-means, clustering method splits the data into k groups through the k-mean algorithm which is a local search procedure but suffers from serious drawbacks [48]. In the case of chickpea, cluster analysis has been proved as a powerful tool to distinguish between a set of germplasm [35, 49]. The reported studies used only a few accessions, eight by [25]), 34 by [35], and 118 by [49] whereas we have used 436 accessions in this investigation. No studies are available with respect to seed storage protein markers based clustering for this size of the dataset. We used three clustering methods i.e. elbow method, silhouette method, and gap statistics to understand the latent structure of the genotypes. Our results established that the gap statistics method is the most appropriate to determine the number of clusters in a large dataset as described earlier in the text.
Previous studies showed that 20 clusters were identified in 118 genotypes of exotic and local germplasm, where the maximum number of genotypes in one single cluster was 47, mainly from Syria and the USA [49]. Three clusters out of eight Kabuli chickpea mutants/hybrids and four clusters out of a set of 34 genotypes were detected in previous studies by [25, 35] respectively. Moreover, no sub-clusters were reported in the aforementioned studies.
We have identified 14 main clusters as well as each cluster harboring three to six sub-clusters in this study. Our results demonstrated that 27 Pakistani genotypes clustered separately in cluster numbers 2, 4, and 7 and further into 3, 3, and 4 subclusters respectively (Fig S7, S9, and S12). Nevertheless, all the clusters (2, 4 and 7) containing Pakistani germplasm did not contain any exotic or wild genotypes implying the possible use of few progenitors during their domestication and inbreeding.
Based on our findings it is assumed that one method to increase the diversity in our germplasm which is a prerequisite for getting wheat varieties with desirable traits; is to include wild as well as exotic genotypes in future breeding [23]. For instance, in the case of wheat, the reason behind that the existence of a relatively small gene pool of wheat cultivars in Pakistan [50] is the use of recurrent selection which further created the omnipresent problem of rust epidemics. The same could happen in case of chickpea.
Hence, we propose a three-way cross of distinctly related genotypes, for example by including genotypes from cluster 2 (for example genotype number 188 viz. CM2000) and 7 (for example genotype number 197 viz. PakK01219) and any of the wild or exotic accessions from (for example genotype number 36 WALSGOTT-3 from Egypt) to increase diversity as well as selection gain in Pakistani chickpea breeding program.
It has been demonstrated before [25] that protein markers of certain sizes (KSSP-100, KSSP-93, and KSSP-64) can be used for the identification of CM-98/99, ILC-195 and CM-2000 genotypes, respectively. We propose here that polymorphic protein bands ranging in size from 10, 11, 12, 14, and 15 kDa and 120,126, 140, 145, 150, 160, and 170 kDa can be used to detect the introgression of blood from wild species into the hybrids produced by crossing wild species harboring desired traits like stress tolerance with cultivated chickpea genotype. We have observed that proteins markers of 10, 126,140, and 160 kDa in size were mostly present in wild type as compared to cultivated types. Specifically, we found that two Pakistani-approved cultivars showed a protein marker of 160 kDa named CH-2016 and Pb-2008. As the name indicated, CH-2016 is a recent variety from NIAB, Pakistan with excellent yield potential and high demand among chickpea growers. Pb-2008 was also a successful variety in 2008 and yet is being cultivated in some areas where it is specifically adapted. We propose that the yield advantage of these two genotypes and their wider adaptability could be due to the incorporation of genetic background from wild type in their ancestry and can be used as a marker in chickpea breeding. This need to be further investigated which is not within the scope of this study.
Conclusion
At large, cluster analysis of 436 chickpea cultivated and wild type genotypes on the basis of SSPs profiling could be differentiated into 14 main clusters with three to six sub-clusters in each main cluster. There were three main clusters of 27 Pakistani genotypes only. Pakistani germplasm showed rare protein bands which were not otherwise present in any other genotype including both cultivated and wild types. Further, genotypes from the three different main clusters could be crossed with other distinct genotypes in the study to develop hybrids to hasten the breeding high yielding cultivars with wider adaptability. In addition, the very high and very low sized protein bands can be used as possible markers of exotic genetic background in the cultivated germplasm, identification of hybrids and to stress tolerant genotypes, for instance albumins with their known role in stress tolerance can be used as marker in chickpea breeding. In the end, it is recommended that SDS-PAGE of SSPs is an important and cost effective technique and can be used successfully to identify hybrids among distantly related parents, measure genetic diversity across genotypes grown under a range of global environments with authenticity and it is worth to be established in laboratories where cost matters, due to its cost effectiveness.
Data availability
All data generated or analysed during this study are being included in the manuscript and its supplementary information files.
References
Maiti R, Ebeling WP (2001) Advances in chickpea science. Science Publishers, New York
Siddique K, Bultynck L (2004) Chicpea/Agronomy. In: Wrigley C et al (eds) Encyclopedia of grain science. Marcel Dekker, New York, pp 287–294
Nwokolo E, Smartt JJ (1996) Food and feed from legumes and oilseeds. Chapman & Hall, London
Kaushik I, Singh R, Bhisnoi JP (2017) Effect of barley malt, chickpea and peanut on quality of barley based beverage. J Appl Nat Sci 9:1182–1186
Roy F, Boye J, Simpson B (2010) Bioactive proteins and peptides in pulse crops: pea, chickpea and lentil. Food Res Int 43:432–442
Kaur R, Prasad K (2021) Technological, processing and nutritional aspects of chickpea (Cicer arietinum)-A review. Trends Food Sci Technol 109:448–463
Jukanti AK, Gaur PM, Gowda C, Chibbar RN (2012) Nutritional quality and health benefits of chickpea (Cicer arietinum L.): a review. Br J Nutr 108:S11–S26
Heidarvand L, Maali-Amiri R (2013) Physio-biochemical and proteome analysis of chickpea in early phases of cold stress. J Plant Physiol 170:459–469
Joshi PK, Rao PP (2017) Global pulses scenario: status and outlook. Ann N Y Acad Sci 1392(1):6–17
Gaur PM, Samineni S, Thudi M, Tripathi S, Sajja SB, Jayalakshmi V, Mannur DM, Vijayakumar AG, Ganga Rao NV, Ojiewo C (2019) Integrated breeding approaches for improving drought and heat adaptation in chickpea (Cicer arietinum L.). Plant Breed 138:389–400
Rachwa-Rosiak D, Nebesny E, Budryn G (2015) Chickpeas—Composition, nutritional value, health benefits, application to bread and snacks: a review. Crit Rev Food Sci Nutr 55:1137–1145
Van der Maesen L (1987) Origin, history and taxonomy of chickpea. In: Saxena MC, Singh KB (eds) The chickpea. CAB Int Publ, UK, pp 11–34
Arif MAR, Waheed MQ, Lohwasser U, Shokat S, Alqudah AM, Volkmar C, Börner A (2022) Genetic insight into the insect resistance in bread wheat exploiting the untapped natural diversity. Front Genet 13:828905
Arif A, Parveen N, Waheed MQ, Atif RM, Waqar I, Shah TM (2021) A comparative study for assessing the drought-tolerance of chickpea under varying natural growth environments. Front Plant Sci 11:607869
Li H, Rodda M, Gnanasambandam A, Aftab M, Redden R, Hobson K, Rosewarne G, Materne M, Kaur S, Slater AT (2015) Breeding for biotic stress resistance in chickpea: progress and prospects. Euphytica 204:257–288
Lopes M, Dreisigacker S, Peña R, Sukumaran S, Reynolds MP (2015) Genetic characterization of the wheat association mapping initiative (WAMI) panel for dissection of complex traits in spring wheat. Theor Appl Genet 128:453–464
Kumar P, Yadava RK, Kumar S, Kumar P (2016) Molecular diversity analysis in wheat genotypes using SSR markers. Electron J Plant Breed 7(2):464–468
Ayala FJ, Kiger JA (1984) Modern genetics. The Benjamins/Cummings Publishing Company, Menlo Park, pp 123–134
Körber N, Bus A, Li J, Parkin IA, Wittkop B, Snowdon RJ, Stich B (2016) Agronomic and seed quality traits dissected by genome-wide association mapping in Brassica napus. Front Plant Sci 7:386
Osawaru ME, Ogwu MC, Aiwansoba RO (2015) Hierarchical approaches to the analysis of genetic diversity in plants: a systematic overview. Univ Mauritius Res J 21
Prasad DR, Minocha T, Maurya B, Yousuf PY (2022) Advancement in molecular tools of plant population genetics. Plant ecogenomics. Apple Academic Press, New Jersey, pp 19–45
Rasheed A, Xia X, Yan Y, Appels R, Mahmood T, He Z (2014) Wheat seed storage proteins: advances in molecular genetics, diversity and breeding applications. J Cereal Sci 60(1):11–24
Hassan R, Waheed M, Shokat S, Arif MAR, Tariq R, Arif M, Arif A (2020) Estimation of genomic diversity using sequence related amplified polymorphism (SRAP) markers in a mini core collection of wheat germplasm from Pakistan. Cereal Res Commun 48:33–44
Sayed MR, Alshallash KS, Safhi FA, Alatawi A, Al-shamrani SM, Dessoky ES, Althobaiti AT, Althaqafi MM, Gharib HS, Shafie WW, Awad-Allah MM (2022) Genetic diversity, analysis of some agro-morphological and quality traits and utilization of plant resources of alfalfa. Genes 13(9):1521
Hameed A, Shah TM, Atta BM, Iqbal N, Haq MA, Ali H (2009) Comparative seed storage protein profiling of Kabuli chickpea genotypes. Pak J Bot 41:703–710
Sharma M, Chaudhary M, Raina SN, Sahoo D, Bhavesh NS, Thakur RK, Rajpal VR, Raturi D, Singh A (2023) Comparative analysis of the efficiency of seed protein profiles in assessing genetic variation and population structure among indigenous Manipur black rice cultivars. Mol Biol Rep. https://doi.org/10.1007/s11033-022-08228-w
Tanksley S, Jones R (1981) Application of alcohol dehydrogenase allozymes in testing the genetic purity of F1 hybrids of tomato. Hort Sci 16:179–181
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Annal Biochem 72:248–254
Kodinariya TM, Makwana PR (2013) Review on determining number of Cluster in K-Means clustering. Int J Adv Res Comput Sci Manage Stud 1(6):90–95
Saputra DM, Saputra D, Oswari LD (2020) Effect of distance metrics in determining k-value in k-means clustering using elbow and silhouette method. SICONIAN.2019. Atlantis Press, Paris, pp 341–346
El-Mandouh AM, Abd-Elmegid LA, Mahmoud HA, Haggag MH (2019) Optimized K-means clustering model based on gap statistic. Int J Adv Comput Sci Appl. https://doi.org/10.14569/IJACSA.2019.0100124
Pal T, Ghosh S, Mondal A, De KK (2016) Evaluation of genetic diversity in some promising varieties of lentil using karyological characters and protein profiling. JGEB 14(1):39–48
Alghamdi SS, Khan MA, Migdadi HM, El-Harty EH, Afzal M, Farooq M (2019) Biochemical and molecular characterization of cowpea landraces using seed storage proteins and SRAP marker patterns. Saudi J Biol Sci 26:74–82
Hameed A, Saddiqa A, Nadeem S, Iqbal N, Atta BM, Shah TM (2012) Genotypic variability and mutant identification in Cicer arietinum L. by seed storage protein profiling. Pak J Bot 44:1303–1310
Wakasa Y, Takaiwa F (2013) The use of rice seeds to produce human pharmaceuticals for oral therapy. Biotechnol J 8:1133–1143
Salunkhe DK, Kadam SS (1989) CRC handbook of world food legumes: nutritional chemistry, processing technology, and utilization. CRC Press, Boca Raton
Vioque J, Clemente A, Sánchez-Vioque R, Pedroche J, Bautista J, Millán F (1998) Comparative study of chickpea and pea PA2 albumins. J Agric Food Chem 46(9):3609–3613
Gueguen J (1991) Pea and fababean proteins. In: Hudson BJF (ed) Development in food proteins. Elsevier, London, pp 35–78
Agrawal L, Narula K, Basu S, Shekhar S, Ghosh S, Datta A, Chakraborty N, Chakraborty S (2013) Comparative proteomics reveals a role for seed storage protein AmA1 in cellular growth, development, and nutrient accumulation. J Proteome Res 12(11):4904–4930
Grasso N, Lynch NL, Arendt EK, O’Mahony JA (2022) Chickpea protein ingredients: a review of composition, functionality, and applications. CRFSFS 21(1):435–452
Gravel A, Doyen A (2023) Pulse globulins 11S and 7S: origins, purification methods, and techno-functional properties. J Agric Food Chem. https://doi.org/10.1021/acs.jafc.2c07507
Plietz B, Damaschum G, Schwenke KD (1980) Quaternary structure of lIS globulin from different plant seeds. Stud Biophys 79:145–146
Singh U, Jambunatha R (1982) Distribution of seed protein fractions and amino acids in different anatomical parts of chickpea (Cicer arietinum L.) and pigeon pea (Cajanus cajan L.). Plant Foods for Hum Nutr 31(10):347–354
Chang YW, Alli I, Molina AT, Konishi Y, Boye JI (2012) Isolation and characterization of chickpea (Cicer arietinum L.) seed protein fractions. Food Bioprocess Technol 5(2):pp618–625
Arif MAR, Bux H, Kazi AG, Rasheed A, Napar AA, Riaz A, Mujeeb-Kazi A (2012) Stripe rust analysis of D-genome synthetic wheats (2 n = 6 x = 42, AABBDD) and their molecular diversity. Arch Phytopathol Pfl 45:1479–1487
Rubio LA, Grant G, Cavalle C, Martinez-Aragon A, Pusztai A (1994) High in vivo (rat) digestibility of faba bean (Vicia faba), lupin (Lupinus angustifolius) and soya bean (Glycine max) soluble globulins. J Sci Food Agric 66:289–292
Pena JM, Lozano JA, Larranaga P (1999) An empirical comparison of four initialization methods for the k-means algorithm. Pattern Recognit Lett 20:1027–1040
Nisar M, Ghafoor A, Khan MR, Ahmad H, Qureshi AS, Ali H (2007) Genetic diversity and geographic relationship among local and exotic chickpea germplasm. Pak J Bot 39:1575–1581
Arif MAR, Nagel M, Neumann K, Kobiljski B, Lohwasser U, Börner A (2012) Genetic studies of seed longevity in wheat using segregation and association mapping approaches. Euphytica 186:1–13
Acknowledgements
The authors are thankful to the Australian Grain Genebank, Victoria, Australia and chickpea group at NIAB, Pakistan for providing the seeds of exotic and Pakistani chickpea cultivars respectively.
Funding
No external funding was received for this work.
Author information
Authors and Affiliations
Contributions
AA conceived the idea; UK and MQW performed the original experiments and carried out the gel visualization and band scoring; MARA, AA, UK, MQW and NP performed the analysis; UK, MQW, AA & MARA wrote the manuscript; NP and AA reviewed the manuscript by providing inputs to improve the manuscript. All authors read and approve the final manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors have no relevant financial and non-financial interest to disclose.
Ethical approval
This article does not contain any studies with animals or humans performed by any of the authors.
Additional information
Uswah Khalid and Muhammad Qandeel Waheed shared first authors.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11033_2023_8358_MOESM5_ESM.png
Supplementary material 5 (PNG 67.7 kb) Sub clusters in cluster 1 where different shaded regions indicate different sub groups. For details, see Table S2
11033_2023_8358_MOESM6_ESM.png
Supplementary material 6 (PNG 30.3 kb) Sub clusters in cluster 2 where different colors indicate different sub groups. For details, see Table S3
11033_2023_8358_MOESM7_ESM.png
Supplementary material 7 (PNG 65.4 kb) Sub clusters in cluster 3 where different colors indicate different sub groups. For details, see Table S4
11033_2023_8358_MOESM8_ESM.png
Supplementary material 8 (PNG 28.0 kb) Sub clusters in cluster 4 where different colors indicate different sub groups. For details, see Table S5
11033_2023_8358_MOESM9_ESM.png
Supplementary material 9 (PNG 46.8 kb) Sub clusters in cluster 5 where different colors indicate different sub groups. For details, see Table S6
11033_2023_8358_MOESM10_ESM.png
Supplementary material 10 (PNG 45.6 kb) Sub clusters in cluster 6 where different colors indicate different sub groups. For details, see Table S7
11033_2023_8358_MOESM11_ESM.png
Supplementary material 11 (PNG 29.1 kb) Sub clusters in cluster 7 where different colors indicate different sub groups. For details, see Table S8
11033_2023_8358_MOESM12_ESM.png
Supplementary material 12 (PNG 30.7 kb) Sub clusters in cluster 8 where different colors indicate different sub groups. For details, see Table S9
11033_2023_8358_MOESM13_ESM.png
Supplementary material 13 (PNG 48.0 kb) Sub clusters in cluster 9 where different colors indicate different sub groups. For details, see Table S10
11033_2023_8358_MOESM14_ESM.png
Supplementary material 14 (PNG 54.0 kb) Sub clusters in cluster 10 where different colors indicate different sub groups. For details, see Table S11
11033_2023_8358_MOESM15_ESM.png
Supplementary material 15 (PNG 49.7 kb) Sub clusters in cluster 11 where different colors indicate different sub groups. For details, see Table S12
11033_2023_8358_MOESM16_ESM.png
Supplementary material 16 (PNG 58.4 kb) Sub clusters in cluster 12 where different colors indicate different sub groups. For details, see Table S13
11033_2023_8358_MOESM17_ESM.png
Supplementary material 17 (PNG 43.8 kb) Sub clusters in cluster 13 where different colors indicate different sub groups. For details, see Table S14
11033_2023_8358_MOESM18_ESM.png
Supplementary material 18 (PNG 44.1 kb) Sub clusters in cluster 14 where different colors indicate different sub groups. For details, see Table S15
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khalid, U., Waheed, M.Q., Parveen, N. et al. Estimation of genetic diversity using seed storage protein (SSP) profiling in wild and cultivated species of Cicer L. Mol Biol Rep 50, 4175–4185 (2023). https://doi.org/10.1007/s11033-023-08358-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08358-9