Abstract
The pathological detergent-insoluble prion protein (PrPSc) is derived from its normal detergent-soluble cellular form (PrPC) through a structural transition from α-helixes into β-sheets, which is associated with a group of transmissible neurodegenerative diseases or prion diseases. According to the prevailing seeding model, PrPSc formation requires a precursor of PrPSc or an intermediate form between PrPC and PrPSc. However, the precursor or intermediate form in the brain remains to be determined. In 2006, we identified in uninfected human and animal brains a novel PrP conformer termed insoluble PrPC (iPrPC) that possesses PrPSc-like properties such as detergent-insolubility, resistance to protease, and tendency to form aggregates. Notably, other common neurodegenerative diseases such as Alzheimer’s disease (AD) and Parkinson’s disease (PD) have recently been proposed to share a prion-like seeding mechanism by which the detergent-soluble brain monomeric cellular proteins form the detergent-insoluble misfolded protein aggregates that transmit from cells to cells. This chapter reviews the physiochemical properties of iPrPC and discusses its formation and pathophysiology. It also highlights the findings and implications of other misfolded proteins such as amyloid-β, tau, and α-synuclein associated with AD and PD in the brain of asymptomatic individuals.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Prion protein
- Prion disease
- Insoluble prion protein
- α-Synuclein
- Amyloid-β
- Tau
- Parkinson’s disease
- Alzheimer’s disease
- Variably protease-sensitive prionopathy
- Dementia
- Memory
1 Introduction
The cellular prion protein (PrPC) is a universally expressed membrane protein present predominantly in the central nervous system (CNS). Deposition in the CNS of its pathologic isoform (PrPSc) derived from PrPC via a conformational transition is a molecular hallmark of prion diseases (PrDs), a group of fatal transmissible spongiform encephalopathies, neurodegenerative disorders, or prion diseases in humans and animals. Numbers of physiological and pathophysiological functions of PrPC have been reported, involved in copper transportation (Brown et al. 1997), anti-oxidative stress (Brown et al. 2001), neurotransmission (Ford et al. 2002), cell-cell adhesion (Málaga-Trillo et al. 2009), cell-cell junctions, signalling (Petit et al. 2013), Amyloid-β (Aβ) receptor in Alzheimer disease (AD) (Laurén et al. 2009; Chap. 22), and cancer biology (Liang et al. 2006; Meslin et al. 2007; Antonacopoulous et al.; Li et al. 2009; Chap. 23). It has been proposed that PrPC has beneficial and deleterious effects on cognition (Collinge et al. 1994; Laurén et al. 2009; Linden et al. 2008; Westaway et al. 2011; Das and Zou 2016). Moreover, it has been well demonstrated that the coexistence of PrPC and PrPSc is the prerequisite for the emergence of PrDs. The two PrP conformers mainly studied so far are believed to be implicated in these diseases. PrPC and PrPSc share the same primary sequence but have distinct secondary structures (Meyer et al. 1986; Caughey et al. 1991; Pan et al. 1993). PrPC is monomeric, rich in α-helical structure, sensitive to proteinase K (PK) digestion, soluble in non-denaturing detergents, non-infectious, and present in both uninfected and scrapie-infected brains. In contrast, PrPSc is oligomeric or aggregate, rich in β-sheet structure, partially resistant to PK digestion, insoluble in detergents, infectious, and present only in infected brains. Interestingly, we have previously demonstrated that PrPSc but not PrPC can be specifically captured by anti-DNA antibodies or DNA-binding proteins, suggesting that the PrPSc aggregates may bind to DNA or acquire a DNA-like structure (Zou et al. 2004). Soluble PrPC is the only conformer that has been detected in the uninfected mammalian brain. In contrast, insoluble PrPSc exhibits chameleon-like conformations, which may underlie the distinct prion strains and phenotypes of PrDs identified in animals and humans (Bessen and Marsh 1992; Parchi et al. 1996; Caughey et al. 1998; Safar et al. 1998; Zou and Gambetti 2007; Collinge and Clarke 2007). Our identification of insoluble cellular PrP (iPrPC) in the uninfected human and animal brain may raise two possibilities: that the PrPC molecule in the brain also exhibits chameleon-like conformations that are implicated in their beneficial or deleterious effects, and that these species may play a role in the pathogenesis of PrDs and other neurodegenerative disorders (Yuan et al. 2006; Zou 2010; Zou et al. 2011b).
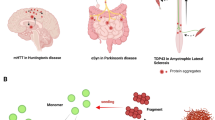
Notably, prion diseases have become a prototype of neurodegenerative diseases including but not limited to Alzheimer’s disease (AD) and Parkinson’s disease (PD) in terms of pathogenesis as well as related concepts and techniques used for investigating prions and prion diseases. For instance, the misfolded proteins including amyloid-β (Aβ) (Meyer-Luehmann et al. 2006; Stöhr et al. 2012), tau (Clavaguera et al. 2009; Iba et al. 2013; Lasagna-Reeves et al. 2012), α-synuclein (Luk et al. 2012a, b; Masuda-Suzukake et al. 2013), huntingtin with polyQ repeats (Ren et al. 2009), superoxide dismutase 1 (SOD1) (Münch et al. 2011), and TDP-43 (Chen et al. 2010; Nonaka et al. 2013) are also transmissible in vitro and/or in vivo. It has been proposed that neurodegenerative diseases share a prion-like self-propagating mechanism by which the misfolded proteins propagate and spread through cell-cell transmission as do prions (Prusiner 2013; Guo and Lee 2013; Goedert 2015). Like prions, they are derived from their normal cellular counterparts; moreover, insoluble Aβ, tau, and α-synuclein can be observed in the brain of asymptomatic, even very young individuals (Braak and Braak 1991; Savva et al. 2009; Braak and Del Tredici 2011; Braak et al. 2011; Jansen et al. 2015; Crary et al. 2014; Josephs et al. 2017; Braak and Braak 1995; Dickson 1998; Del Tredici et al. 2002; Braak et al. 2003).
2 Prion Protein Is Characterized by the Presence of an Intrinsically Chameleon-Like Conformation
Studies using recombinant PrP (rPrP) in vitro have indicated that PrP possesses a highly variable conformation. In aqueous solutions, rPrP could be folded into pH-dependent α-helical conformations, a thermodynamically more stable β-sheet, and various stable or transient intermediates (Zhang et al. 1997). A stopped-flow kinetic study demonstrated that PrP folded by a three-state mechanism involving a monomeric intermediate (Apetri and Surewicz 2002). It was found that the population of this partially structured PrP intermediate increased in the presence of relatively low concentrations of urea and was more stable at acidic pH 4.8, compared to neutral pH 7.0. Moreover, this approach revealed that PrP mutations, linked with naturally occurring familial prion diseases, showed a pronounced stabilization of the folding intermediate (Apetri et al. 2004). These findings suggest that the intermediates play a crucial role in PrP conversion and serve as direct precursors of the pathologic PrPSc isoform. The existence of a PrP folding intermediate was also indicated by hydrogen exchange experiments (Nicholson et al. 2002), and by studies using high-pressure NMR and fluorescence spectroscopy (Kuwata et al. 2002; Martins et al. 2003). In addition to a β-oligomer and an amyloid fibril (Baskakov et al. 2001; Morillas et al. 2001; Lu and Chang 2002; Sokolowski et al. 2003; Baskakov et al. 2004), two additional polymeric transient intermediates were also identified during fibrillogenesis of rPrP in vitro (Baskakov et al. 2002).
The cellular PrPC molecule is anchored to the cell membrane through a glycosylphosphatidylinositol (GPI) anchor. Several experiments have indicated that the PrP conformation is affected by its local conditions. For example, the interaction of the anchorless recombinant PrP with lipids in a membrane-like environment resulted in a conformational transition (Wang et al. 2007; Re et al. 2008). Increasing the local concentration of membrane-anchored PrPC seems to induce a conformational transition accompanied by oligomerization of PrPC (Elfrink et al. 2008). Recently, Faris et al. identified mitochondria PrPC in healthy mice, which is a transmembrane isoform with the C-terminus facing the mitochondrial matrix and the N-terminus facing the intermembrane space, which is PK-resistant (Faris et al. 2017). Therefore, the tendency of PrP to form multiple nonnative β-sheet-rich isoforms in vitro, as demonstrated in biophysical studies on rPrP, may represent a unique intrinsic feature of this protein.
Most of the N-terminal region of recombinant human and murine PrP has been observed to be disordered by NMR study (Riek et al. 1997; Zahn et al. 2000). The nucleic acid-binding intrinsically disordered proteins (IDPs) have recently been reported to be involved in diseases by driving liquid–liquid phase separation (LLPS) (Elbaum-Garfinkle 2019). Moreover, it is believed that the formation of membrane-less organelles in vivo follows the generation of protein-rich condensates or granules by LLPS (Brangwynne et al. 2009; Boeynaems et al. 2018). PrP is able to form liquid-like condensates (Kostylev et al. 2018). PrP interaction with nucleic acids (NAs) undergoes LLPS, modulates phase separation, and promotes PrP fibrillation in a NA structure and concentration-dependent manner (Matos et al. 2020). Interestingly, DNA/RNA-PrP is involved in the formation of dynamic compartments, which may be associated with various functions of PrPC and its misfolding; the condensates have been proposed to be part of the PrPSc pathway and therefore represent novel targetable structures for therapeutics (do Amaral and Cordeiro 2021).
3 Insoluble Cellular Prion Protein Aggregates Are Present in Mammalian Brains Without Prion-Infection
If the tendency of PrP to form multiple conformations in vitro represents a unique intrinsic feature of this protein, it is conceivable that other PrP conformers would be present in the normal brain in addition to the well-characterized PrPC. To test this, we examined uninfected human and animal brains using a combination of biophysical and biochemical approaches to confirm the presence of additional PrP conformers (Yuan et al. 2006). Indeed, we identified a novel conformer that forms insoluble cellular PrP aggregates and protease-resistant PrP species in uninfected human brains (Yuan et al. 2006). Using gel filtration, we revealed that PrP in uninfected human brains is present not only in monomers with molecular weight less than 66 kDa, but also in oligomers between 66 kDa and 200 kDa, and large aggregates greater than 669 kDa, even 2000 kDa (Yuan et al. 2006) (Fig. 4.1). The new PrP conformer, termed insoluble cellular PrP (iPrPC), accounts for approximately 5–25% of total PrP including full-length and N-terminally truncated forms, and a portion of iPrPC is resistant to PK digestion even at 50 μg/mL (Yuan et al. 2006). Notably, the PK-resistant iPrPC has immunoreactive behaviour different from that of classic PrPSc detected in prion-infected brains; its affinity is much lower for 3F4 while higher for 1E4, compared to the affinity of those antibodies for classic PrPSc (Yuan et al. 2006, 2008; Zou et al. 2010a, 2011a) (Fig. 4.2). In contrast to the gel mobilities of the deglycosylated PrPSc type 1 and type 2 that are 21 kDa and 19 kDa, respectively, the 1E4-detected PK-resistant deglycosylated PrP has gel mobility at ~20 kDa (Fig. 4.2). The epitopes of the two antibodies 3F4 and 1E4 are adjacent and the C-terminus of the 1E4 epitope between PrP97–105 is connected to the N terminus of the 3F4 epitope between PrP 106–112 (Yuan et al. 2008; Zou et al. 2010a). 3F4 is the most widely used antibody in the detection of human PrPC and PrPSc, including PrPSc types 1 and 2 seen in sCJD and inherited CJD, and the internal PrPSc fragment PrP7–8 seen in GSS. Besides the 1E4-detected 20 kDa band, a PK-resistant PrP band migrating at ~18 kDa is also detectable with an antibody against the C-terminal PrP domain from residues 220–231 (anti-C antibody) (Fig. 4.2) (Yuan et al. 2006). In addition, the new conformer reveals a high affinity for the gene 5 protein (g5p, a single-stranded DNA-binding protein) and sodium phosphotungstate (NaPTA), both of which specifically bind to PrPSc but not to soluble PrPC (Zou et al. 2004; Yuan et al. 2006; Safar et al. 1998; Wadsworth et al. 2001). By using the g5p enrichment from 500 μL of normal human brain homogenate, two more PK-resistant PrP bands migrating at ~18–19 kDa and ~ 7–8 kDa are detected by 1E4 in the uninfected human brain (Yuan et al. 2006). To rule out the possibility that PrP aggregates detected in the uninfected human brain result from post-mortem autolysis of autopsy tissues or from other neurodegenerative disorders, we also examined frozen uninfected human biopsy brain tissues or normal animal brain tissues from hamsters and cows. We observed that the insoluble PrPC was also detectable in these tissues, a finding which confirmed that iPrPC is a de novo generated PrP conformer (Yuan et al. 2006). Using gel filtration, we recently further demonstrated that not only soluble PrPC monomers, but also soluble PrPC oligomers are present in the uninfected human brain (Xiao et al. 2012).
Western blotting of gel filtration fractions of PrP from uninfected human brains. Gel filtration fractions of uninfected brain homogenates were subjected to SDS-PAGE and Western blotting with 3F4. Molecular mass (kDa) of various PrP species recovered in different fractions is indicated by an arrow and molecular mass markers used include dextran blue (2000 kDa), thyroglobulin (669 kDa), apoferritin (443 kDa), β-amylase (200 kDa), and albumin (66 kDa). PrP was detected not only in fractions with molecular mass less than 66 kDa after fraction 59 but also in fractions with molecular mass greater than 66 kDa before fraction 59 including fraction 33 containing large PrP aggregates (2000 kDa)
Western blotting of PK-resistance of PrP in uninfected human brains. Brain homogenates from two uninfected human brains received at autopsy were treated with PK at 0, 5, 10, 25, 50, and 100 μg/mL (upper two panels a and b) or PK plus PNGase F (lower three panels c, d, and e). The samples were subjected to SDS-PAGE and Western blotting with 3F4, 1E4, and Anti-C antibodies. No PK-resistant PrP was detectable with 3F4 antibody. In contrast, PK-resistant PrP was detected with 1E4 and Anti-C up to 100 μg/mL. With PK alone, three PrP bands migrating at 30–29 kDa, 27–26 kDa, and 21–20 kDa were detected, in which the upper band (~30–29 kDa, blue asterisk) was predominant while the intensity of the middle band was lowest, which is apparently different from those of PrPSc type 1 (T1) and type 2 (T2). After PNGase F treatment, only one band was detected with 1E4 and Anti-C migrating at ~20 kDa and ~ 18 kDa, respectively (PrP*20 and PrP*18, red asterisk). Interestingly, a band migrating at ~12–13 kDa was also detected with anti-C at low PK concentration (5–10 μg/mL, yellow asterisk)
The presence of additional PrP oligomeric conformers besides the typical soluble PrPC monomers in uninfected brains was also implied in the observations reported by other groups. Consistent with our findings, small amounts of PrP (less than 5% of total PrPC) were also reported to be precipitated by NaPTA from uninfected human brains (Wadsworth et al. 2001). Moreover, by a differential SDS solubility assay, PrPC species with either lower or higher solubility were differentiated in brain homogenates of noninfected humans, sheep, and cattle (Kuczius et al. 2009, 2011). Based on the detergent-solubility, the PrPC phenotypes in cattle were similar to those in humans but not in sheep (Kuczius and Groschup 2013). Notably, a purified hamster brain PrPC displayed an unexpectedly high β-sheet component under native conditions (Pergami et al. 1999). This finding provided evidence that the full-length native PrPC isolated from animal brains exhibited intrinsic conformational plasticity. Moreover, mammalian brain PrPC from six species was observed to be initially degraded to an intermediate fragment prior to complete proteolysis, suggesting an intrinsic partial PK-resistance (Buschmann et al. 1998). Ward et al. have recently observed that in response to the inoculation of normal brain homogenates, the host brain PrPC exhibited increased insolubility and protease resistance at 72 h post-inoculation, similar to that of PrPSc (Ward et al. 2019). The authors proposed that the occurrence of PrP aggregation and protease-resistance results from brain injury due to the inoculation of normal brain homogenates. They believe that these changes were comparable to that observed in the examination of post-mortem human brain tissue (Esiri et al. 2000), in hypoxic human brain tissue from cases of cerebral ischemia (McLennan et al. 2004) and stroke (Mitsios et al. 2007), as well as in brain tissue of sheep with various neurological diseases (Jeffrey et al. 2012). Moreover, the same group has also previously identified a PK-resistant PrP species that is derived from the mitochondria of healthy mouse brain tissues (Faris et al. 2017). Interestingly, PrP aggregates have also been reported in pancreatic beta-cells of uninfected rats in response to hyperglycemia (Strom et al. 2007). In sum, the cumulative evidence shows that insoluble and PK-resistant PrPC aggregates are present in tissues and organs of uninfected animals and humans.
4 Spontaneous Formation of the Insoluble Cellular Prion Protein Has Been Modelled with Cultured Cells and May Result from PrP Cytosolic Accumulation
Lehmann and Harris (1996) modelled the spontaneous formation of PrPSc-like insoluble PrP in cultured Chinese hamster ovary (CHO) cells expressing wild-type or mutant mouse PrP. Significant amounts of mutant PrP with a point mutation at residue 199 (E199K) (~60%) or six octapeptide repeat insertion mutations between residues 51 and 90 (~90%) linked to inherited human prion disease were detergent-insoluble; notably, approximately 15% wild-type PrPC was also detergent-insoluble (Lehmann and Harris 1996). While approximately 5% of mutant PrP was resistant to the digestion by PK at 3.3 μg/ml for 20 min, wild-type PrP was completely degraded. Because the two mutant PrP molecules but not wild-type PrP were tightly associated with the plasma membrane, it was hypothesized that the acquisition of PrPSc-like properties results from an alternation in membrane topology or affinity (Lehmann and Harris 1996). Using the same models, they further identified a three-step endocytic pathway by which mutant PrP forms a PrPSc-like conformer: initially hydrophobic, then detergent-insoluble, and finally partially PK resistant (Daude et al. 1997). Using human neuroblastoma cells, Singh et al. also revealed that PrP with Q217R mutation linked to GSS formed a PrPSc-like form (Singh et al. 1997).
In addition to the above PrP mutations, the two N-linked glycosylation sites located at residue 181, Asn-Ile-Thr residues 181–183, and at residue 197, Asn-Phe-Thr residues 197–199 (Puckett et al. 1991) are believed to play a crucial role in the stabilization of prion protein conformation. The naturally occurring mutations at residue 183, Thr to Ala (PrPT183A), or at residue 198, Phe to Ser (PrPF198S), falling in the two consensus sites, are linked to two distinct familial prion diseases (Nitrini et al. 1997; Tagliavini et al. 1991). Elimination of either site or both by mutagenesis of hamster PrP in CV1 cells, induced intracellular accumulation of mutant proteins (Rogers et al. 1990). Lehmann and Harris observed that mouse PrP mutated at T182 alone, or at both T182 and T198 in CHO cells, failed to reach the cell surface but the PrP with T198 mutation did. Moreover, all three mutant PrP molecules acquired PrPSc-like physicochemical properties reminiscent of PrPSc; PrPWt did so only when synthesized in the presence of N-linked glycosylation inhibitor tunicamycin (Lehmann and Harris 1997). Using human neuroblastoma M17 cells expressing human PrPN181G or PrPT183A, Capellari et al. observed that PrPN181G, but not PrPT183A, reached the cell surface even though both mutations eliminated glycosylation at the first site (Capellari et al. 2000). This observation indicates that the Thr to Ala mutation itself, rather than the elimination of the first glycosylation site, altered the physical properties of the mutant protein (Capellari et al. 2000). Although the F198S mutation falls within the second glycosylation site, Asn-Phe-Thr residues 197–199, PrPF198S slightly increased the efficiency of glycosylation at the first glycosylation site (N181) and greatly increased it at the second site (N197) in cultured cells (Zaidi et al. 2005).
To further investigate the formation of iPrPC and the effect of mutations on the formation of iPrPC, we examined iPrPC in cultured M17 cells expressing human wild-type (PrPWt) and mutant PrP (Yuan et al. 2008; Zou et al. 2011a). We confirmed that the de novo generated iPrP was detectable not only in cells expressing mutant PrP (PrPT183A or PrPF198S) linked to naturally occurring genetic Creutzfeldt–Jakob disease and Gerstmann–Sträussler–Scheinker disease, respectively, but also in cells expressing wild-type PrP. Compared to cells expressing wild-type PrP, cells expressing mutant PrP exhibit significantly increased amounts of iPrP forming PrP aggregates and PK-resistant PrP. Most of PrPT183A was composed of oligomers and large aggregates; virtually no monomeric form was present. In PrPF198S, however, monomeric species were still dominant despite an increase in the amounts of aggregates. The enhanced tendency of PrPT183A to form aggregates may result from the intracellular accumulation of the mutant protein. The F198S mutation did not significantly diminish the ability of PrPF198S to reach the cell surface (Zaidi et al. 2005), although the mutation may change the structure around the V14 epitope previously found to be localized between human PrP168-181 (Zou et al. 2011a; Moudjou et al. 2004; Rezaei et al. 2005; Zhang et al. 2021). Therefore, the majority of the iPrPC associated with the T183A mutation may result from PrP intracellular accumulation, raising the possibility that iPrPC is derived predominantly from intracellular PrP species. Immunofluorescence microscopy of tagged PrP also indicated that PrPT183A accumulates within the cell, whereas PrPF198S was distributed both inside the cell and on the cell surface, consistent with previous observations (Zou et al. 2011a; Capellari et al. 2000; Zaidi et al. 2005).
In uninfected cultured cells, we also confirmed that the PK-resistant iPrPC exhibited higher affinity for 1E4 than for 3F4, which was initially observed in brain tissue samples (Zou et al. 2011a; Yuan et al. 2006, 2008). In Western blotting with cell lysates, 1E4 virtually detected no PrP before PK treatment, and it detected PrP only after PK treatment. However, PrP was stainable by 1E4 in fixed cultured cells treated with or without PK although the PrP signal was weaker in treated than in untreated cells (Zou et al. 2011a). It is worth noting that an antibody against human PrP95-110 (termed 8G8), that actually extends merely two more amino acids toward the N- and C-terminuses of the 1E4 epitope, respectively, stained PrP-expressing cells with a brilliant cytoplasmic fluorescence (Krasemann et al. 1999). However, the number of positive cells was smaller than that of cells stained with antibodies against other PrP regions. Moreover, despite sharing a similar amino acid sequence within the corresponding epitope region, only cattle, but not mouse and hamster PrP, was observed to react with 8G8 (Krasemann et al. 1999). In contrast to 3F4, 1E4 seems to detect intracellular PrP in cultured cells (Zou et al. 2011a). Therefore, like 8G8, 1E4 may recognize a PrP species with a unique conformation in its epitope region.
In the absence of scrapie infection, aggregation of the cellular wild-type PrP in cultured cells was also observed only when proteasome inhibitors were used (Yedidia et al. 2001). It was later reported that PrPWt accumulated in the cytoplasm of cultured cells under other conditions as well, such as in a reducing environment, or when expressing PrP without both N and C terminal signal peptides (Ma and Lindquist 2001, 2002; Drisaldi et al. 2003; Grenier et al. 2006). Cytosolic PrP forms aggregates that are insoluble in non-ionic detergents and partially resistant to PK (Ma and Lindquist 2001). Accumulated cytosolic PrP aggregates induced by ER stress and inhibition of proteasomal activity were recently observed to travel through the secretory pathway and reach the plasma membrane (Nunziante et al. 2011). Cytosolic PrP was observed not only in cultured cells but also in subpopulations of neurons in the hippocampus, neocortex, and thalamus in uninfected wild-type mice (Mironov Jr et al. 2003). In addition, soluble PrPC in human brain homogenate was observed to switch to insoluble PrPC by treatment with acidic buffers in vitro (Zou and Cashman 2002).
The above observations may suggest that the formation of iPrPC or the aggregation of PrPC is associated not only with mutations of the protein but also with altered cellular conditions that cause abnormal traffic and distribution of PrP in cells including reductive/oxidative stress and low pH.
5 Physiology and Pathophysiology of Insoluble PrPC Aggregates
5.1 Long-Term Memory Storage
The iPrPC with a conformation likely different from soluble PrPC may have a physiologic function. It has been hypothesized that prion-like conformational changes of related proteins are indispensable for the maintenance of structural synaptic changes required for long-term memory (Si et al. 2003, 2010; Papassotiropoulos et al. 2005; Shorter and Lindquist 2005). Interestingly, 24 h after a word-list learning task, carriers of either PrP polymorphism methionine/methionine (M/M) at residue 129 (129MM) or M/valine (V) (129 MV) genotype were observed to recall 17% more information than did 129VV carriers (Papassotiropoulos et al. 2005). Their further investigation of brain activity with event-related functional magnetic resonance imaging (fMRI) during a word recognition task suggested that the PrP-129 polymorphism affects neural plasticity following learning at a time scale of minutes to hours (Buchmann et al. 2008). The authors proposed that the PrP gene is genetically associated with human long-term memory performance. It is possible that the polymorphism at residue 129 of PrP participates in mediating human memory, in which the 129 M allele has a beneficial effect on long-term memory. Moreover, the impact of a putative PrP conformation rather than pathologic PrPSc on long-term memory in healthy humans was proposed to be related to physiologically occurring conformational changes (Tompa and Friedrich 1998; Papassotiropoulos et al. 2005).
It would be interesting to determine whether the conversion of soluble PrPC monomers into insoluble PrP oligomers or aggregates is directly associated with long-term memory storage in the normal human brain (Zou et al. 2011c). The possibility cannot be ruled out that iPrPC is involved in long-term memory since it is the specific isoform of PrPC that binds to nucleic acids, an important feature of proteins involved in long-term memory (Sudhakaran and Ramaswami 2017). For instance, the iPrPC molecule is able to gene five protein (g5p), the single-stranded DNA-binding protein (Yuan et al. 2006, 2008). The binding of recombinant PrP to different types of RNAs has been observed in vitro (Bera and Biring 2018) and the possible binding of iPrPC to mRNA in vivo cannot be ruled out. RNA has been found to modulate the aggregation of recombinant murine PrP by direct interaction in vitro (Kovachev et al. 2019).
5.2 Prion Disease
The in vivo pathway by which PrPC forms PrPSc remains poorly understood. Two non-exclusive conversion models were proposed: refolding (Griffith 1967; Prusiner 1991) and seeding (Jarrett and Lansbury Jr 1993). In the former, the exogenous PrPSc binds to the PrPC species that has been partially unfolded and the PrPSc-bound PrPC molecule undergoes a refolding process during which the nascent PrPSc is derived from this PrPC species via a conformational transition. The latter proposes that a small amount of abnormal PrPSc or PrPSc-like form (PrP*) is present in the normal brain and is in reversible equilibrium with PrPC. When several monomeric PrP* molecules form a highly ordered nucleus, PrPC is converted to PrPSc polymers. Obviously, two key elements are required by the seeding model. One is the presence in the uninfected brain of a small amount of endogenous PrPSc or PrP* and the second is the formation of PrPSc-derived oligomers. The seeding model, with the two elements, has been recapitulated in vitro using PrP from various fungal and mammalian sources (Ross et al. 2005; Castilla et al. 2005; Tanaka et al. 2005). Indeed, this model well explains the replication pattern of PrPSc in which a newly recruited polypeptide chain accurately replicates that of a PrPSc template.
Recent studies also observed that replication of PrPSc does not always follow the refolding and seeding models, especially in vitro propagation of PrPSc in the presence of recombinant PrP substrate by serial protein misfolding cyclic amplification (sPMCA). For instance, Baskakov and co-workers have recently proposed an alternative model of PrPSc replication designated as deformed templating (see Chap. 5; Makarava and Baskakov 2012; Requena 2020; Spagnolli et al. 2020). It appears to involve switching from one cross-β folding pattern present in a template to an altered folding pattern, which undergoes a deformed process.
Given that iPrPC aggregates possess PrPSc-like physicochemical properties, we propose that iPrPC could represent endogenous PrPSc (Yuan et al. 2006; Zou et al. 2011a; Das and Zou 2016), an intermediate form (PrP*) between PrPC and PrPSc, or a silent prion, required for seeding model of PrPSc formation (Jarrett and Lansbury Jr 1993; Hall and Edskes 2004; Weissmann 2004). Based on the observation that the brain of bigenic mice is capable of clearing prions, it has been proposed that the normal brain contains low levels of PrPSc (Safar et al. 2005). Under normal circumstances, despite the presence of a small amount of PrPSc, the brain may maintain an equilibrium between the formation and clearance of PrPSc. The amount of PrPSc is expected to be too small to induce a neurodegenerative disorder, which presumably, remains in a silent state. However, prion diseases may be triggered when the levels of the silent prions are significantly increased due to infection, PrP mutation, or unknown causes. Using PMCA, Barria and co-workers generated a new infectious prion without adding exogenous PrPSc seeds (Barria et al. 2009). This study raises two possibilities (1) PMCA replicates an intermediate PrPSc that is present in the brain homogenate; or (2) the silent prion is activated by the sonication–incubation cycles during PMCA. Further studies to address these questions will be critical to the understanding of initial molecular events in prion formation.
As mentioned above, iPrPC possesses a unique immunoreactive behaviour of poor affinity for 3F4 and higher affinity for 1E4, compared to other types of human PrPSc identified so far (Yuan et al. 2006, 2008; Zou et al. 2011a). The two antibodies have adjacent epitopes on PrP (Yuan et al. 2008; Zou et al. 2010b). Thus, the possibility cannot be ruled out that iPrP is a distinct PrP species with an altered conformation and that it may be a conformer which, when it increases, induces an atypical form of prion disease. Some previous observations with experimental animals may favour this hypothesis. A novel neurologic syndrome was reported in Tg mice overexpressing wild-type PrP and these mice exhibited degeneration of skeletal muscle, peripheral nerves, and the central nervous system (Westaway et al. 1994). The increased amounts of wild-type PrPC might form aggregates that induce degeneration in those mice. Chiesa et al. observed that homozygous Tg mice overexpressing wild-type PrP at approximately ten-fold but not hemizygous mice overexpressing wild-type PrP at approximately five-fold developed a spontaneous neurodegenerative disorder manifesting tremor and paresis (Chiesa et al. 2008). Nevertheless, abnormal PrP deposits and enlarged synaptic terminals with a dramatic proliferation of membranous structures were found in both types of mice. It was also observed that the overexpressed PrP assembled into insoluble aggregates with mild PK resistance but acquired no infectivity (Chiesa et al. 2008). Misfolding and neurotoxicity of wild-type PrP in transgenic flies were observed to be sequence dependent: Hamster PrP formed large amounts of PrP aggregates with spongiform degeneration, whereas rabbit PrP formed only small amounts of PrP aggregates without spongiform degeneration (Fernandez-Funez et al. 2010). Moreover, the same study also found that although small amounts of PrP aggregates were similarly detected in young flies expressing hamster PrP (day 1), spongiform degeneration was not evident. Therefore, the small amounts of PrP aggregates were unable to induce spongiform degeneration. Interestingly, spongiform degeneration occurred in older flies only when the concentrations of PrP aggregates increased (day 30).
The same unique immunoreactivity behaviour with 1E4 has also been observed in an atypical PrPSc species we recently identified from variably protease-sensitive prionopathy (VPSPr), a novel human prion disease (Gambetti et al. 2008; Zou et al. 2010b, 2013; Chap. 20). VPSPr exhibits an abnormal PrP species with peculiar glycosylation, enzymatic proteolysis, in vitro seeding activity, and in vivo infectivity (Zou et al. 2010b, 2011c; Wang et al. 2019; Zhang et al. 2021; Notari et al. 2014; Diack et al. 2014; Nonno et al. 2019). The 1E4-detected pathogenetic PK-resistant PrPSc with a ladder-like electrophoretic profile in the brain is the molecular hallmark of VPSPr. PrPSc from VPSPr exhibits not only the peculiar immunoreactivity behaviour but also three PK-resistant core fragments, which is similar to iPrPC (Zou et al. 2010b, 2011c, 2013). These similarities may suggest that they share a common molecular metabolic pathway. Similar to sCJD, VPSPr affects patients regardless of their PrP genotypes defined by 129 MV polymorphism; however, the allelic prevalence is distinct in the two diseases (Zou et al. 2010b; Gambetti et al. 2011a; Notari et al. 2018). Notably, the amounts of PK-resistant PrPSc in VPSPr seem to be dependent on the polymorphism, a characteristic that has not been observed in sCJD. Recent studies revealed that the infectivity of PrPSc from VPSPr is incomplete or inefficient in humanized transgenic mice expressing human PrP while it is transmissible in bank voles with attack rates of 5–35% in the first passage and 100% in the second passage (Nonno et al. 2019). Therefore, it is possible that VPSPr characterized by the deposition in the brain of iPrPC-like PrPSc represents a prion disease, distinct from classical prion diseases and bearing more resemblance to other neurodegenerative diseases such as AD and tauopathies (Gambetti et al. 2011b; Zou et al. 2013; Chap. 20). Because of the similarities between iPrPC and PrPSc from VPSPr, the possibility that VPSPr results from an increase in the amount of iPrPC cannot be excluded (Zou et al. 2013; Chap. 20).
5.3 Alzheimer’s Disease
PrPC has been observed to be the receptor of amyloid-β (Aβ) in AD (Laurén et al. 2009; Balducci et al. 2010; Chap. 22). In 2011, we demonstrated for the first time that the insoluble PrPC is the main PrP species that interacts with Aβ in the brain of AD patients and transgenic mice expressing human amyloid precursor protein, carrying both the Swedish (K670N and M671L) and Indiana (V717F) mutations (Zou et al. 2011b). This study made the following seven novel findings. First, large PrP and Aβ aggregates are eluted in the same gel filtration fractions from the brains of AD patients and AD mouse models. Second, more than 95% of Aβ co-immunoprecipitated with PrP by 3F4 from these brains is insoluble, while less than 5% of Aβ is soluble. Third, Aβ is co-captured with iPrPC by gene 5 protein (g5p) from AD brains. Fourth, 6 Aβ42-specific binding regions on the human PrP molecule are identified with a peptide membrane array involving 13-mer human PrP peptides and two Aβ peptides (Aβ42 and Aβ40). Fifth, 4 of 6 Aβ42-specific binding areas are observed in the PrP octapeptide repeat domain of the unstructured N-terminal domain and only one is in the folded C-terminal region between residues 151 and 165. The other Aβ42-specific binding sites are located between the N- and C-terminal domains (residues 119–137). Sixth, compared with its nonspecific binding PrP sites (non-distinguishingly binding to both Aβ42 and Aβ40), the affinity of Aβ42 for its specific binding sites (binding to Aβ42 only) is significantly lower. Finally, the oligomeric state or conformation of Aβ42 and Aβ40 may determine the affinity of the two Aβ peptides for human PrP.
Our findings were largely confirmed by a subsequent study using both Aβ-PrP interaction and co-immunoprecipitation assays in a large AD patient cohort (Dohler et al. 2014). Specifically, they revealed that (1) significant binding of Aβ to PrPC only occurs in AD, (2) Aβ aggregates bind particularly to the N-terminus of PrPC, (3) optimal binding of PrPC to Aβ is observed in the insoluble fraction of AD brain homogenates, and (4) neither expression levels nor PrP-129 polymorphisms of PrPC influence their binding. The C-terminal PrPC also has been found to play a role in the interaction between the protein and Aβ. PrPC inhibits Aβ fibril growth via its C-terminal domain and the proposed binding of Aβ to the N-terminal domain of PrP may cause a conformational change in the C-terminal domain that unmasks additional Aβ-binding sites in that region (Bove-Fenderson et al. 2017). A new study with solid-state MAS NMR spectroscopy revealed that most of the C-terminal domain of PrP is part of the rigid complex with a loss in regular secondary structure in the two C-terminal α-helices (König et al. 2021), which could well explain why the complexes of PrPC and Aβ are mainly detected in the insoluble fractions (Zou et al. 2011b; Dohler et al. 2014). Notably, the PrPC-dependent, Aβ oligomers-induced Fyn activation was observed in detergent-insoluble subcellular fractions of cultured N2A neuroblastoma, suggesting that insoluble PrPC is involved in Aβ-induced PrPC-Fyn signalling pathway (Um et al. 2012). Larson et al. revealed that the anti-PrP antibody C20 was able to immunoprecipitate Aβ dimers and activate Fyn, triggering tau aberrant mis-sorting and hyperphosphorylation (Larson et al. 2012). However, they claimed that their results are in contrast with our findings because no Aβ monomers coimmunoprecipitating with PrPC from AD brains were detected using five anti-PrP antibodies (8B4, C20, 6D11, M20, and 7D11) and four anti-Aβ antibodies (6E10, 4G8, and 40/4-end specific Mab2.1.3 and Mab 13.1.1). The discrepancy between Larson et al. and Zou et al./Dohler et al. remains unknown. One of the possibilities could be due to different antibodies (3F4 antibody used in studies by Zou et al. and Dohler et al.) and lysis buffer. Larson et al. used the RIPA buffer that contains 3% SDS that may dissociate large PrPC-Aβ assemblies.
The findings that iPrPC mainly or optimally binds to Aβ aggregates observed by us and Dohler et al. are consistent with other previous observations. For instance, PrP deposits often histologically accompany Aβ-positive plaques in AD brains (Esiri et al. 2000; Ferrer et al. 2001; Kovacs et al. 2002). In addition, Freir et al. displayed that interaction between PrP and toxic Aβ assemblies can be therapeutically targeted at multiple sites (Freir et al. 2011), indicating that their binding sites are not limited only to the internal domain. Remarkably, Kudo et al. showed that not only anti-PrP antibodies but also PrPC peptides identified in our previous study (Zou et al. 2011b) rescued Aβ oligomer-induced neurotoxicity (Kudo et al. 2012).
Although the exact biological relevance and pathophysiology of the interaction between iPrPC and Aβ remain unclear, aggregation of one protein was observed to facilitate aggregation of the others (Morales et al. 2010). Moreover, synergistic interactions between other amyloidogenic proteins associated with neurodegeneration have also been reported to promote each other’s fibrillization, amyloid deposition, and formation of filamentous inclusions in transgenic mice (Schwarze-Eicker et al. 2005). An increase in the efficiency of Aβ42 aggregation in vitro was dependent on PrPSc dosage (Morales et al. 2010). Moreover, insoluble PrPSc aggregates also seemed to facilitate Aβ42 aggregation in vivo; AD mice developed a strikingly higher load of cerebral amyloid plaques that appeared much faster in prion-infected than in uninfected mice (Morales et al. 2010). Our finding that Aβ42 binds to iPrP may suggest that iPrP facilitates the fibrillization of Aβ42 in AD. Similarly, the possibility should be considered that a significant increase in the total number of Aβ plaques observed in bigenic mice overexpressing PrP (Schwarze-Eicker et al. 2005) might result from an increase in the formation of iPrP. Since the less toxic insoluble Aβ42 aggregates constitute the end products of highly toxic soluble Aβ42 oligomers, it is conceivable that the formation of the large aggregates facilitated by iPrPC may reduce the amount of Aβ42 oligomers. The decrease in the levels of toxic Aβ42 oligomers would then attenuate the cognitive impairment induced by Aβ42 oligomers in AD. If this is the case, iPrPC may play a protective role in AD. Given that iPrPC interacts with insoluble Aβ42, whereas soluble PrPC binds soluble Aβ42 in vivo (Zou et al. 2011b), it is possible that distinct PrP conformers binding to different Aβ42 species thereby function either as receptors for soluble Aβ42 oligomers or as modulators of insoluble Aβ42 deposition. It would be interesting to test this hypothesis by intracerebrally injecting anti-PrP antibodies against either soluble or insoluble PrP species in AD animal models. This experiment would establish that the multiple conformers of PrPC are coupled with their beneficial and deleterious effects.
6 Insoluble Aβ, Tau, and α-Synuclein Aggregates in the Brain of Asymptomatic Individuals
Accumulation and deposition of insoluble Aβ, tau, and α-synuclein aggregates in the brain are the molecular hallmark of AD and PD. These insoluble aggregates are also derived from their soluble monomeric counterparts through a structural transition, a mechanism similar to the conversion of PrPC into PrPSc. They form extracellular Aβ plaques and intracellular phosphorylated neurofibrillary tangles in AD brains and α-synuclein-containing Lewy bodies in PD brains. Interestingly, examination of brains obtained at autopsy from nondemented and demented cadavers has demonstrated that accumulation of these AD- and PD-pathologies commences before the appearance of clinical symptoms (preclinical phase) (Braak and Braak 1991; Braak and Del Tredici 2011; Braak et al. 2011; Forno 1969; Bloch et al. 2006; Mikolaenko et al. 2005; Del Tredici and Braak 2008). Moreover, in living patients, positron emission tomography (PET) of Aβ revealed that the accumulation of Aβ can be detected approximately 20 years before dementia onset in AD (Gordon et al. 2018; Villemagne et al. 2013; Hansson 2021). Moreover, several lines of evidence also have revealed that they are detectable in the brain of aged individuals who never developed AD and PD clinical symptoms and signs. Positive cortical Aβ-PET was observed in ~10–15% of individuals with normal cognition at age 60 and in ~40% at age 90 (Hansson 2021; Savva et al. 2009; Jansen et al. 2015). Neuropathological examinations have demonstrated that stages A and B of Aβ pathology can be found before clinical dementia (Braak and Braak 1991; Braak et al. 2011). Tau pathology has been observed in primary age-related tauopathy (PART) (Crary et al. 2014; Hansson 2021). It is often confined to the medial temporal lobe area; moreover, PART displays minimal or no Aβ pathology, and seldom has dementia if it has no other primary co-pathology (Crary et al. 2014; Josephs et al. 2017). Lewy bodies and Lewy neurites were also observed in the brain of older individuals without clinical histories of PD (Forno 1969; Bloch et al. 2006; Mikolaenko et al. 2005; Del Tredici and Braak 2008).
7 Conclusions
Chameleon-like conformations of PrPSc are believed to link to transmissible and non-transmissible prion diseases with highly heterogeneous phenotypes (Collinge and Clarke 2007; Zou 2007; Zou and Gambetti 2007). Identification of iPrPC suggests that the normal protein also has chameleon-like conformations. It has been proposed that the variable conformations of PrPC are linked to its beneficial and deleterious effects (Zou et al. 2011c). Demonstration of the presence of insoluble PrP in normal mammalian brains and its potential association with AD and atypical prion disease may open a new avenue in the exploration of prion formation and in the physiology and pathophysiology of the prion protein. Similarly, findings of insoluble misfolded Aβ, tau, and α-synuclein proteins in individuals without AD or PD clinical manifestations such as dementia or motor and non-motor symptoms and signs are also significant because they imply that other co-factors are required for these insoluble misfolded proteins to cause neuronal death. Further investigation on the differences in neurotoxic and non-toxic insoluble misfolded proteins would be critical to our understanding of the pathogenesis of these diseases and developing effective therapeutic compounds for these disorders.
References
Apetri AC, Surewicz WK. Kinetic intermediate in the folding of human prion protein. J Biol Chem. 2002;277:44589–92.
Apetri AC, Surewicz K, Surewicz WK. The effect of disease-associated mutations on the folding pathway of human prion protein. J Biol Chem. 2004;279:18008–14.
Balducci C, Beeg M, Stravalaci M, Bastone A, Sclip A, Biasini E, et al. Synthetic amyloid-beta oligomers impair long-term memory independently of cellular prion protein. Proc Natl Acad Sci U S A. 2010;107:2295–300.
Barria MA, Mukherjee A, Gonzalez-Romero D, Morales R, Soto C. De novo generation of infectious prions in vitro produces a new disease phenotype. PLoS Pathog. 2009;5:e1000421.
Baskakov IV, Legname G, Prusiner SB, Cohen FE. Folding of prion protein to its native alpha-helical conformation is under kinetic control. J Biol Chem. 2001;276:19687–90.
Baskakov IV, Legname G, Baldwin MA, Prusiner SB, Cohen FE. Pathway complexity of prion protein assembly into amyloid. J Biol Chem. 2002;277:21140–8.
Baskakov IV, Legname G, Gryczynski Z, Prusiner SB. The peculiar nature of unfolding of the human prion protein. Protein Sci. 2004;13:586–95.
Bera A, Biring S. A quantitative characterization of interaction between prion protein with nucleic acids. Biochem Biophys Rep. 2018;14:114–24.
Bessen RA, Marsh RF. Identification of two biologically distinct strains of transmissible mink encephalopathy in hamsters. J Gen Virol. 1992;73:329–34.
Bloch A, Probst A, Bissig H, Adams H, Tolnay M. Alpha-synuclein pathology of the spinal and peripheral autonomic nervous system in neurologically unimpaired elderly subjects. Neuropathol Appl Neurobiol. 2006;32:284–95.
Boeynaems S, Alberti S, Fawzi NL, Mittag T, Polymenidou M, Rousseau F, et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 2018;28:420–35.
Bove-Fenderson E, Urano R, Straub JE, Harris DA. Cellular prion protein targets amyloid-β fibril ends via its C-terminal domain to prevent elongation. J Biol Chem. 2017;292:16858–71.
Braak H, Braak E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 1991;82:239–59.
Braak H, Braak E. Staging of Alzheimer’s disease-related neurofibrillary changes. Neurobiol Aging. 1995;16:271–84.
Braak H, Del Tredici K. The pathological process underlying Alzheimer’s disease in individuals under thirty. Acta Neuropathol. 2011;121:171–81.
Braak H, Del Tredici K, Rüb U, de Vos RA, Jansen Steur EN, Braak E. Staging of brain pathology related to sporadic Parkinson’s disease. Neurobiol Aging. 2003;24:197–211.
Braak H, Thal DR, Ghebremedhin E, Del Tredici K. Stages of the pathologic process in Alzheimer disease: age categories from 1 to 100 years. J Neuropathol Exp Neurol. 2011;70:960–9.
Brangwynne CP, Eckmann CR, Courson DS, Rybarska A, Hoege C, Gharakhani J, et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science. 2009;324:1729–32.
Brown DR, Qin K, Herms JW, Madlung A, Manson J, Strome R, et al. The cellular prion protein binds copper in vivo. Nature. 1997;390:684–7.
Brown DR, Clive C, Haswell SJ. Antioxidant activity related to copper binding of native prion protein. J Neurochem. 2001;76:69–76.
Buchmann A, Mondadori CR, Hänggi J, Aerni A, Vrticka P, Luechinger R, Boesiger P, Hock C, Nitsch RM, de Quervain DJ, Papassotiropoulos A, Henke K. Prion protein M129V polymorphism affects retrieval-related brain activity. Neuropsychologia. 2008;46:2389–402.
Buschmann A, Kuczius T, Bodemer W, Groschup MH. Cellular prion proteins of mammalian species display an intrinsic partial proteinase K resistance. Biochem Biophys Res Commun. 1998;253:693–702.
Capellari S, Zaidi SI, Long AC, Kwon EE, Petersen RB. The Thr183Ala mutation, not the loss of the first glycosylation site, alters the physical properties of the prion protein. J Alzheimers Dis. 2000;2:27–35.
Castilla J, Saá P, Hetz C, Soto C. In vitro generation of infectious scrapie prions. Cell. 2005;121:195–206.
Caughey BW, Dong A, Bhat KS, Ernst D, Hayes SF, Caughey WS. Secondary structure analysis of the scrapie-associated protein PrP 27-30 in water by infrared spectroscopy. Biochemistry. 1991;30:7672–80.
Caughey B, Raymond GJ, Bessen RA. Strain-dependent differences in beta-sheet conformations of abnormal prion protein. J Biol Chem. 1998;273:32230–5.
Chen AK, Lin RY, Hsieh EZ, Tu PH, Chen RP, Liao TY, et al. Induction of amyloid fibrils by the C-terminal fragments of TDP-43 in amyotrophic lateral sclerosis. J Am Chem Soc. 2010;132:1186–7.
Chiesa R, Piccardo P, Biasini E, Ghetti B, Harris DA. Aggregated, wild-type prion protein causes neurological dysfunction and synaptic abnormalities. J Neurosci. 2008;28:13258–67.
Clavaguera F, Bolmont T, Crowther RA, et al. Transmission and spreading of tauopathy in transgenic mouse brain. Nat Cell Biol. 2009;11:909–13.
Collinge J, Clarke AR. A general model of prion strains and their pathogenicity. Science. 2007;318:930–6.
Collinge J, Whittington MA, Sidle KC, Smith CJ, Palmer MS, Clarke AR, et al. Prion protein is necessary for normal synaptic function. Nature. 1994;370:295–7.
Crary JF, Trojanowski JQ, Schneider JA, Abisambra JF, Abner EL, Alafuzoff I, et al. Primary age-related tauopathy (PART): a common pathology associated with human aging. Acta Neuropathol. 2014;128:755–66.
Das AS, Zou WQ. Prions: beyond a single protein. Cli Microbio Rev. 2016;29:633–58.
Daude N, Lehmann S, Harris DA. Identification of intermediate steps in the conversion of a mutant prion protein to a scrapie-like form in cultured cells. J Biol Chem. 1997;272(17):11604–12. Erratum in J Biol Chem 2000;275:1520
Del Tredici K, Braak H. Neurofibrillary changes of the Alzheimer type in very elderly individuals: neither inevitable nor benign: commentary on “No disease in the brain of a 115-year-old woman”. Neurobiol Aging. 2008;29:1133–6.
Del Tredici K, Rüb U, De Vos RA, Bohl JR, BraakH. Where does Parkinson disease pathology begin in the brain? J Neuropathol Exp Neurol. 2002;61:413–26.
Diack AB, Ritchie DL, Peden AH, Brown D, Boyle A, Morabito L, Maclennan D, Burgoyne P, Jansen C, Knight RS, Piccardo P, Ironside JW, Manson JC. Variably protease-sensitive prionopathy, a unique prion variant with inefficient transmission properties. Emerg Infect Dis. 2014;20:1969–79.
Dickson DW. Pick’s disease: a modern approach. Brain Pathol. 1998;8:339–54.
do Amaral MJ, Cordeiro Y. Intrinsic disorder and phase transitions: pieces in the puzzling role of the prion protein in health and disease. Prog Mol Biol Transl Sci. 2021;183:1–43.
Dohler F, Sepulveda-Falla D, Krasemann S, Altmeppen H, Schlüter H, Hildebrand D, Zerr I, Matschke J, Glatzel M. High molecular mass assemblies of amyloid-beta oligomers bind prion protein in patients with Alzheimer’s disease. Brain. 2014;137:873–86.
Drisaldi B, Stewart RS, Adles C, Stewart LR, Quaglio E, Biasini E, Fioriti L, Chiesa R, Harris DA. Mutant PrP is delayed in its exit from the endoplasmic reticulum, but neither wild-type nor mutant PrP undergoes retrotranslocation prior to proteasomal degradation. J Biol Chem. 2003;278:21732–43.
Elbaum-Garfinkle S. Matter over mind: liquid phase separation and neurodegeneration. J Biol Chem. 2019;294:7160–8.
Elfrink K, Ollesch J, Stöhr J, Willbold D, Riesner D, Gerwert K. Structural changes of membrane-anchored native PrP(C). Proc Natl Acad Sci U S A. 2008;105:10815–9.
Esiri MM, Carter J, Ironside JW. Prion protein immunoreactivity in brain samples from an unselected autopsy population: findings in 200 consecutive cases. Neuropathol Appl Neurobiol. 2000;26:273–84.
Faris R, Moore RA, Ward A, Race B, Dorward DW, Hollister JR, Fischer ER, Priola SA. Cellular prion protein is present in mitochondria of healthy mice. Sci Rep. 2017;7:41556.
Fernandez-Funez P, Zhang Y, Casas-Tinto S, Xiao X, Zou WQ, Rincon-Limas DE. Sequence-dependent prion protein misfolding and neurotoxicity. J Biol Chem. 2010;285:36897–908.
Ferrer I, Blanco R, Carmona M, Puig B, Ribera R, Rey MJ, et al. Prion protein expression in senile plaques in Alzheimer’s disease. Acta Neuropathol. 2001;101:49–56.
Forno LS. Concentric hyalin intraneuronal inclusions of Lewy type in the brains of elderly persons (50 incidental cases): relationship to parkinsonism. J Am Geriatr Soc. 1969;17:557–75.
Ford MJ, Burton LJ, Li H, Graham CH, Frobert Y, Grassi J, et al. A marked disparity between the expression of prion protein and its message by neurons of the CNS. Neuroscience. 2002;111:533–51.
Freir DB, Nicoll AJ, Klyubin I, Panico S, Mc Donald JM, Risse E, Asante EA, Farrow MA, Sessions RB, Saibil HR, Clarke AR, Rowan MJ, Walsh DM, Collinge J. Interaction between prion protein and toxic amyloid β assemblies can be therapeutically targeted at multiple sites. Nat Commun. 2011;2:336–40.
Gambetti P, Dong Z, Yuan J, Xiao X, Zheng M, Alshekhlee A, et al. A novel human disease with abnormal prion protein sensitive to protease. Ann Neurol. 2008;63:697–708.
Gambetti P, Puoti G, Zou WQ. Variably protease-sensitive prionopathy: a novel disease of the prion protein. J Mol Neurosci. 2011a;45:422–4.
Gambetti P, Zou WQ, Torres JM, Soto C, Notari S, Espinosa JC, et al. Variably protease-sensitive prionopathy: transmissibility and PMCA studies. Prion. 2011b;5:14.
Goedert M. NEURODEGENERATION. Alzheimer’s and Parkinson’s diseases: the prion concept in relation to assembled Aβ, tau, and α-synuclein. Science. 2015;349:1255555.
Gordon BA, Blazey TM, Su Y, Hari-Raj A, Dincer A, Benzinger T, et al. Spatial patterns of neuroimaging biomarker change in individuals from families with autosomal dominant Alzheimer’s disease: a longitudinal study. Lancet Neurol. 2018;17:241–50.
Grenier C, Bissonnette C, Volkov L, Roucou X. Molecular morphology and toxicity of cytoplasmic prion protein aggregates in neuronal and non-neuronal cells. J Neurochem. 2006;97:1456–66.
Griffith JS. Self-replication and scrapie. Nature. 1967;215:1043–4.
Guo JL, Lee VM. Neurofibrillary tangle-like tau pathology induced by synthetic tau fibrils in primary neurons over-expressing mutant tau. FEBS Lett. 2013;587:717–23.
Hall D, Edskes H. Silent prions lying in wait: a two-hit model of prion/amyloid formation and infection. J Mol Biol. 2004;336:775–86.
Hansson O. Biomarkers for neurodegenerative diseases. Nat Med. 2021;27:954–63.
Iba M, Guo JL, McBride JD, Zhang B, Trojanowski JQ, Lee VM. Synthetic tau fibrils mediate transmission of neurofibrillary tangles in a transgenic mouse model of Alzheimer’s-like tauopathy. J Neurosci. 2013;33:1024–37.
Jansen WJ, Ossenkoppele R, Knol DL, Tijms BM, Scheltens P, Verhey FR, et al. Prevalence of cerebral amyloid pathology in persons without dementia: a meta-analysis. JAMA. 2015;313(19):1924–38.
Jarrett JT, Lansbury PT Jr. Seeding “one-dimensional crystallization” of amyloid: a pathogenic mechanism in Alzheimer’s disease and scrapie? Cell. 1993;73:1055–8.
Jeffrey M, Scholes SF, Martin S, McGovern G, Sisó S, González L. Increased immunohistochemical labelling for prion protein occurs in diverse neurological disorders of sheep: relevance for normal cellular PrP function. J Comp Pathol. 2012;147:46–54.
Josephs KA, Murray ME, Tosakulwong N, Whitwell JL, Knopman DS, Machulda MM, et al. Tau aggregation influences cognition and hippocampal atrophy in the absence of beta-amyloid: a clinico-imaging-pathological study of primary age-related tauopathy (PART). Acta Neuropathol. 2017;133:705–15.
König AS, Rösener NS, Gremer L, Tusche M, Flender D, Reinartz E, Hoyer W, Neudecker P, Willbold D, Heise H. Structural details of amyloid beta oligomers in complex with human prion protein as revealed by solid-state MAS NMR spectroscopy. J Biol Chem. 2021;296:100499.
Kostylev MA, Tuttle MD, Lee S, Klein LE, Takahashi H, Cox TO, et al. Liquid and hydrogel phases of PrPC linked to conformation shifts and triggered by Alzheimer’s amyloid-β oligomers. Mol Cell. 2018;72:426–443.e12.
Kovachev PS, Gomes MPB, Cordeiro Y, Ferreira NC, Valadão LPF, Ascari LM, Rangel LP, Silva JL, Sanyal S. RNA modulates aggregation of the recombinant mammalian prion protein by direct interaction. Sci Rep. 2019;9:12406.
Kovacs GG, Zerbi P, Voigtländer T, Strohschneider M, Trabattoni G, Hainfellner JA, et al. The prion protein in human neurodegenerative disorders. Neurosci Lett. 2002;329:269–72.
Krasemann S, Jürgens T, Bodemer W. Generation of monoclonal antibodies against prion proteins with an unconventional nucleic acid-based immunization strategy. J Biotechnol. 1999;73:119–29.
Kuczius T, Groschup MH. Cellular prion proteins in humans and cattle but not sheep are characterized by a low-solubility phenotype. Comp Immunol Microbiol Infect Dis. 2013;36:599–605.
Kuczius T, Karch H, Groschup MH. Differential solubility of prions is associated in manifold phenotypes. Mol Cell Neurosci. 2009;42:226–33.
Kuczius T, Wohlers J, Karch H, Groschup MH. Subtyping of human cellular prion proteins and their differential solubility. Exp Neurol. 2011;227:188–94.
Kudo W, Lee HP, Zou WQ, Wang X, Perry G, Zhu X, Smith MA, Petersen RB, Lee HG. Cellular prion protein is essential oligomeric amyloid-β induced neuronal cell death. Hum Mol Genet. 2012;21(5):1138–44.
Kuwata K, Li H, Yamada H, Legname G, Prusiner SB, Akasaka K, et al. Locally disordered conformer of the hamster prion protein: a crucial intermediate to PrPSc? Biochemistry. 2002;41:12277–83.
Larson M, Sherman MA, Amar F, Nuvolone M, Schneider JA, Bennett DA, Aguzzi A, Lesné SE. The complex PrP(c)-Fyn couples human oligomeric Aβ with pathological tau changes in Alzheimer’s disease. J Neurosci. 2012;32:16857–71.
Lasagna-Reeves CA, Castillo-Carranza DL, Sengupta U, Guerrero-Munoz MJ, Kiritoshi T, Neugebauer V, Jackson GR, Kayed R. Alzheimer brain-derived tau oligomers propagate pathology from endogenous tau. Sci Rep. 2012;2:700.
Laurén J, Gimbel DA, Nygaard HB, Gilbert JW, Strittmatter SM. Cellular prion protein mediates impairment of synaptic plasticity by amyloid-beta oligomers. Nature. 2009;457:1128–32.
Lehmann S, Harris DA. Two mutant prion proteins expressed in cultured cells acquire biochemical properties reminiscent of the scrapie isoform. Proc Natl Acad Sci U S A. 1996;93:5610–4.
Lehmann S, Harris DA. Blockade of glycosylation promotes acquisition of scrapie-like properties by the prion protein in cultured cells. J Biol Chem. 1997;272:21479–87.
Li C, Yu S, Nakamura F, Yin S, Xu J, Petrolla AA, et al. Binding of pro-prion to filamin A disrupts cytoskeleton and correlates with poor prognosis in pancreatic cancer. J Clin Invest. 2009;119:2725–36.
Liang J, Wang JB, Pan YL, Wang J, Liu LL, Guo XY, et al. High frequency occurrence of 1-OPRD variant of PRNP gene in gastric cancer cell lines and Chinese population with gastric cancer. Cell Biol Int. 2006;30:920–3.
Linden R, Martins VR, Prado MA, Cammarota M, Izquierdo I, Brentani RR. Physiology of the prion protein. Physiol Rev. 2008;88:673–728.
Lu BY, Chang JY. Isolation and characterization of a polymerized prion protein. Biochem J. 2002;364:81–7.
Luk KC, Kehm V, Carroll J, Zhang B, O’Brien P, Trojanowski JQ, Lee VM. Pathological α-synuclein transmission initiates Parkinson-like neurodegeneration in nontransgenic mice. Science. 2012a;338:949–53.
Luk KC, Kehm VM, Zhang B, O’Brien P, Trojanowski JQ, Lee VM. Intracerebral inoculation of pathological α-synuclein initiates a rapidly progressive neurodegenerative α-synucleinopathy in mice. J Experi Med. 2012b;209:975–86.
Ma J, Lindquist S. Wild-type PrP and a mutant associated with prion disease are subject to retrograde transport and proteasome degradation. Proc Natl Acad Sci U S A. 2001;98:14955–60.
Ma J, Lindquist S. Conversion of PrP to a self-perpetuating PrPSc-like conformation in the cytosol. Science. 2002;298:1785–8.
Makarava N, Baskakov IV. Purification and fibrillation of full-length recombinant PrP. Methods Mol Biol. 2012;849:33–52.
Málaga-Trillo E, Solis GP, Schrock Y, Geiss C, Luncz L, Thomanetz V, et al. Regulation of embryonic cell adhesion by the prion protein. PLoS Biol. 2009;7:e55.
Martins SM, Chapeaurouge A, Ferreira ST. Folding intermediates of the prion protein stabilized by hydrostatic pressure and low temperature. J Biol Chem. 2003;278:50449–55.
Masuda-Suzukake M, Nonaka T, Hosokawa M, Oikawa T, Arai T, Akiyama H, Mann DM, Hasegawa M. Prion-like spreading of pathological α-synuclein in brain. Brain. 2013;136:1128–38.
Matos CO, Passos YM, do Amaral MJ, Macedo B, Tempone MH, Bezerra O, et al. Liquid-liquid phase separation and fibrillation of the prion protein modulated by a high-affinity DNA aptamer. FASEB J. 2020;34:365–85.
McLennan NF, Brennan PM, McNeill A, Davies I, Fotheringham A, Rennison KA, et al. Prion protein accumulation and neuroprotection in hypoxic brain damage. Am J Pathol. 2004;165:227–35.
Meslin F, Hamaï A, Gao P, Jalil A, Cahuzac N, Chouaib S, et al. Silencing of prion protein sensitizes breast adriamycin-resistant carcinoma cells to TRAIL-mediated cell death. Cancer Res. 2007;67:10910–9.
Meyer RK, McKinley MP, Bowman KA, Braunfeld MB, Barry RA, Prusiner SB. Separation and properties of cellular and scrapie prion proteins. Proc Natl Acad Sci U S A. 1986;83:2310–4.
Meyer-Luehmann M, Coomaraswamy J, Bolmont T, Kaeser S, Schaefer C, Kilger E, et al. Exogenous induction of cerebral beta-amyloidogenesis is governed by agent and host. Science. 2006;313:1781–4.
Mikolaenko I, Pletnikova O, Kawas CH, O’Brien R, Resnick SM, Crain B, Troncoso JC. Alpha-synuclein lesions in normal aging, Parkinson disease, and Alzheimer disease: evidence from the Baltimore longitudinal study of aging (BLSA). J Neuropathol Exp Neurol. 2005;64:156–62.
Mironov A Jr, Latawiec D, Wille H, Bouzamondo-Bernstein E, Legname G, Williamson RA, Burton D, DeArmond SJ, Prusiner SB, Peters PJ. Cytosolic prion protein in neurons. J Neurosci. 2003;23:7183–93.
Mitsios N, Saka M, Krupinski J, Pennucci R, Sanfeliu C, Miguel Turu M, et al. Cellular prion protein is increased in the plasma and periinfarcted brain tissue after acute stroke. J Neurosci Res. 2007;85:602–11.
Morales R, Estrada LD, Diaz-Espinoza R, Morales-Scheihing D, Jara MC, Castilla J, et al. Molecular cross talk between misfolded proteins in animal models of Alzheimer’s and prion diseases. J Neurosci. 2010;30:4528–35.
Morillas M, Vanik DL, Surewicz WK. On the mechanism of alpha-helix to beta-sheet transition in the recombinant prion protein. Biochemistry. 2001;40:6982–7.
Moudjou M, Treguer E, Rezaei H, Sabuncu E, Neuendorf E, Groschup MH, Grosclaude J, Laude H. Glycan-controlled epitopes of prion protein include a major determinant of susceptibility to sheep scrapie. J Virol. 2004;78:11449.
Münch C, O’Brien J, Bertolotti A. Prion-like propagation of mutant superoxide dismutase-1 misfolding in neuronal cells. Proc Natl Acad Sci U S A. 2011;108:3548–53.
Nicholson EM, Mo H, Prusiner SB, Cohen FE, Marqusee S. Differences between the prion protein and its homolog Doppel: a partially structured state with implications for scrapie formation. J Mol Biol. 2002;316:807–15.
Nitrini R, Rosemberg S, Passos-Bueno MR, da Silva LS, Iughetti P, Papadopoulos M, Carrilho PM, Caramelli P, Albrecht S, Zatz M, LeBlanc A. Familial spongiform encephalopathy associated with a novel prion protein gene mutation. Ann Neurol. 1997;42:138–46.
Nonaka T, Masuda-Suzukake M, Arai T, Hasegawa Y, Akatsu H, Obi T, Yoshida M, Murayama S, Mann DM, Akiyama H, Hasegawa M. Prion-like properties of pathological TDP-43 aggregates from diseased brains. Cell Rep. 2013;4:124–34.
Nonno R, Notari S, Di Bari MA, Cali I, Pirisinu L, d’Agostino C, Cracco L, Kofskey D, Vanni I, Lavrich J, Parchi P, Agrimi U, Gambetti P. Variable protease-sensitive Prionopathy transmission to Bank voles. Emerg Infect Dis. 2019;25:73–81.
Notari S, Xiao X, Espinosa JC, Cohen Y, Qing L, Aguilar-Calvo P, Kofskey D, Cali I, Cracco L, Kong Q, Torres JM, Zou W, Gambetti P. Transmission characteristics of variably protease-sensitive prionopathy. Emerg Infect Dis. 2014;20:2006–14.
Notari S, Appleby BS, Gambetti P. Variably protease-sensitive prionopathy. Handb Clin Neurol. 2018;153:175–90.
Nunziante M, Ackermann K, Dietrich K, Wolf H, Gädtke L, Gilch S, Vorberg I, Groschup M, Schätzl HM. Proteasomal dysfunction and endoplasmic reticulum stress enhance trafficking of prion protein aggregates through the secretory pathway and increase accumulation of pathologic prion protein. J Biol Chem. 2011;286:33942–53.
Pan KM, Baldwin M, Nguyen J, Gasset M, Serban A, Groth D, et al. Conversion of alpha-helices into beta-sheets features in the formation of the scrapie prion proteins. Proc Natl Acad Sci U S A. 1993;90:10962–6.
Papassotiropoulos A, Wollmer MA, Aguzzi A, Hock C, Nitsch RM, de Quervain DJ. The prion gene is associated with human long-term memory. Hum Mol Genet. 2005;14:2241–6.
Parchi P, Castellani R, Capellari S, Ghetti B, Young K, Chen SG, et al. Molecular basis of phenotypic variability in sporadic Creutzfeldt–Jakob disease. Ann Neurol. 1996;39:767–78.
Pergami P, Poloni TE, Corato M, Camisa B, Ceroni M. Prions and prion diseases. Funct Neurol. 1999;14:241–52.
Petit CS, Besnier L, Morel E, Rousset M, Thenet S. Roles of the cellular prion protein in the regulation of cell-cell junctions and barrier function. Tissue Barriers. 2013;1:e24377.
Prusiner SB. Molecular biology of prion diseases. Science. 1991;252:1515–22.
Prusiner SB. Biology and genetics of prions causing neurodegeneration. Annu Rev Genet. 2013;47:601–23.
Puckett C, Concannon P, Casey C, Hood L. Genomic structure of the human prion protein gene. Am J Hum Genet. 1991;49:320–9.
Re F, Sesana S, Barbiroli A, Bonomi F, Cazzaniga E, Lonati E, Bulbarelli A, Masserini M. Prion protein structure is affected by pH-dependent interaction with membranes: a study in a model system. FEBS Lett. 2008;582:215–20.
Ren PH, Lauckner JE, Kachirskaia I, Heuser JE, Melki R, Kopito RR. Cytoplasmic penetration and persistent infection of mammalian cells by polyglutamine aggregates. Nat Cell Biol. 2009;11:219–25.
Requena JR. The protean prion protein. PLoS Biol. 2020;18:e3000754.
Rezaei H, Eghiaian F, Perez J, Doublet B, Choiset Y, Haertle T, Grosclaude J. Sequential generation of two structurally distinct ovine prion protein soluble oligomers displaying different biochemical reactivities. J Mol Biol. 2005;347:665–79.
Riek R, Hornemann S, Wider G, Glockshuber R, Wüthrich K. NMR characterization of the full-length recombinant murine prion protein, mPrP(23-231). FEBS Lett. 1997;413:282–8.
Rogers M, Taraboulos A, Scott M, Groth D, Prusiner SB. Intracellular accumulation of the cellular prion protein after mutagenesis of its Asn-linked glycosylation sites. Glycobiology. 1990;1:101–9.
Ross ED, Minton A, Wickner RB. Prion domains: sequences, structures and interactions. Nat Cell Biol. 2005;7:1039–44.
Safar J, Wille H, Itri V, Groth D, Serban H, Torchia M, et al. Eight prion strains have PrP(Sc) molecules with different conformations. Nat Med. 1998;4:1157–65.
Safar JG, DeArmond SJ, Kociuba K, Deering C, Didorenko S, Bouzamondo-Bernstein E, et al. Prion clearance in bigenic mice. J Gen Virol. 2005;86:2913–23.
Savva GM, Wharton SB, Ince PG, Forster G, Matthews FE, Brayne C, Medical Research Council Cognitive Function and Ageing Study. Age, neuropathology, and dementia. N Engl J Med. 2009;360:2302–9.
Schwarze-Eicker K, Keyvani K, Görtz N, Westaway D, Sachser N, Paulus W. Prion protein (PrPc) promotes beta-amyloid plaque formation. Neurobiol Aging. 2005;26:1177–82.
Shorter J, Lindquist S. Prions as adaptive conduits of memory and inheritance. Nat Rev Genet. 2005;6:435–50.
Si K, Lindquist S, Kandel ER. A neuronal isoform of the aplysia CPEB has prion-like properties. Cell. 2003;115:879–91.
Si K, Choi YB, White-Grindley E, Majumdar A, Kandel ER. Aplysia CPEB can form prion-like multimers in sensory neurons that contribute to long-term facilitation. Cell. 2010;140:421–35.
Singh N, Zanusso G, Chen SG, Fujioka H, Richardson S, Gambetti P, Petersen RB. Prion protein aggregation reverted by low temperature in transfected cells carrying a prion protein gene mutation. J Biol Chem. 1997;272:28461–70.
Sokolowski F, Modler AJ, Masuch R, Zirwer D, Baier M, Lutsch G, et al. Formation of critical oligomers is a key event during conformational transition of recombinant Syrian hamster prion protein. J Biol Chem. 2003;278:40481–92.
Spagnolli G, Rigoli M, Novi Inverardi G, Codeseira YB, Biasini E, Requena JR. Modeling PrPSc Generation Through Deformed Templating. Front Bioeng Biotechnol. 2020;8:590501.
Stöhr J, Watts JC, Mensinger ZL, et al. Purified and synthetic Alzheimer’s amyloid beta (Aβ) prions. Proc Natl Acad Sci U S A. 2012;109(27):11025–30.
Strom A, Wang GS, Reimer R, Finegood DT, Scott FW. Pronounced cytosolic aggregation of cellular prion protein in pancreatic beta-cells in response to hyperglycemia. Lab Investig. 2007;87:139–49.
Sudhakaran IP, Ramaswami M. Long-term memory consolidation: the role of RNA-binding proteins with prion-like domains. RNA Biol. 2017;14:568–86.
Tagliavini F, Prelli F, Ghiso J, Bugiani O, Serban D, Prusiner SB, Farlow MR, Ghetti B, Frangione B. Amyloid protein of Gerstmann-Sträussler-Scheinker disease (Indiana kindred) is an 11 kd fragment of prion protein with an N-terminal glycine at codon 58. EMBO J. 1991;10:513–9.
Tanaka M, Chien P, Yonekura K, Weissman JS. Mechanism of cross-species prion transmission: an infectious conformation compatible with two highly divergent yeast prion proteins. Cell. 2005;121:49–62.
Tompa P, Friedrich P. Prion proteins as memory molecules: an hypothesis. Neuroscience. 1998;86:1037–43.
Um JW, Nygaard HB, Heiss JK, Kostylev MA, Stagi M, Vortmeyer A, Wisniewski T, Gunther EC, Strittmatter SM. Alzheimer amyloid-beta oligomer bound to postsynaptic prion protein activates Fyn to impair neurons. Nat Neurosci. 2012;15:1227–35.
Villemagne VL, Burnham S, Bourgeat P, Brown B, Ellis KA, Salvado O, et al. Amyloid β deposition, neurodegeneration, and cognitive decline in sporadic Alzheimer’s disease: a prospective cohort study. Lancet Neurol. 2013;12(4):357–67.
Wadsworth JD, Joiner S, Hill AF, Campbell TA, Desbruslais M, Luthert PJ, Collinge J. Tissue distribution of protease resistant prion protein in variant Creutzfeldt–Jakob disease using a highly sensitive immunoblotting assay. Lancet. 2001;358:171–80.
Wang F, Yang F, Hu Y, Wang X, Wang X, Jin C, Ma J. Lipid interaction converts prion protein to a PrPSc-like proteinase K-resistant conformation under physiological conditions. Biochemistry. 2007;46:7045–53.
Wang Z, Yuan J, Shen P, Abskharon R, Lang Y, Dang J, Adornato A, Xu L, Chen J, Feng J, Moudjou M, Kitamoto T, Lee HG, Kim YS, Langeveld J, Appleby B, Ma J, Kong Q, Petersen RB, Zou WQ, Cui L. In vitro seeding activity of Glycoform-deficient prions from variably protease-sensitive Prionopathy and familial CJD associated with PrP(V180I) mutation. Mol Neurobiol. 2019;56:5456–69.
Ward A, Hollister JR, Choi YP, Race B, Williams K, Shoup DW, Moore RA, Priola SA. Altered distribution, aggregation, and protease resistance of cellular prion protein following intracranial inoculation. PLoS One. 2019;14:e0219457.
Weissmann C. The state of the prion. Nat Rev Microbiol. 2004;2:861–71.
Westaway D, DeArmond SJ, Cayetano-Canlas J, Groth D, Foster D, Yang SL, et al. Degeneration of skeletal muscle, peripheral nerves, and the central nervous system in transgenic mice overexpressing wild-type prion proteins. Cell. 1994;76:117–29.
Westaway D, Alier K, Vergote D, MacTavish D, Mercer R, Fu W, et al. Prion proteins and the Alzheimer disease Aβ amyloid cascade. Prion. 2011;5:1–2.
Xiao X, Yuan J, Zou WQ. Isolation of soluble and insoluble PrP oligomers in the normal human brain. J Vis Exp. 2012;68:e3788. https://doi.org/10.3791/3788.
Yedidia Y, Horonchik L, Tzaban S, Yanai A, Taraboulos A. Proteasomes and ubiquitin are involved in the turnover of the wild-type prion protein. EMBO J. 2001;20:5383–91.
Yuan J, Xiao X, McGeehan J, Dong Z, Cali I, Fujioka H, et al. Insoluble aggregates and protease-resistant conformers of prion protein in uninfected human brains. J Biol Chem. 2006;281:34848–58.
Yuan J, Dong Z, Guo JP, McGeehan J, Xiao X, Wang J, et al. Accessibility of a critical prion protein region involved in strain recognition and its implications for the early detection of prions. Cell Mol Life Sci. 2008;65:631–43.
Zahn R, Liu A, Lührs T, Riek R, von Schroetter C, López García F, et al. NMR solution structure of the human prion protein. Proc Natl Acad Sci U S A. 2000;97:145–50.
Zaidi SI, Richardson SL, Capellari S, Song L, Smith MA, Ghetti B, Sy MS, Gambetti P, Petersen RB. Characterization of the F198S prion protein mutation: enhanced glycosylation and defective refolding. J Alzheimers Dis. 2005;7:159–71. discussion 173–180
Zhang H, Stockel J, Mehlhorn I, Groth D, Baldwin MA, Prusiner SB, et al. Physical studies of conformational plasticity in a recombinant prion protein. Biochemistry. 1997;36:3543–53.
Zhang W, Xiao X, Ding M, Yuan J, Foutz A, Moudjou M, Kitamoto T, Langeveld JPM, Cui L, Zou WQ. Further characterization of Glycoform-selective prions of variably protease-sensitive Prionopathy. Pathogens. 2021;10:513.
Zou WQ. Transmissible spongiform encephalopathy and beyond (E-letter). Science; 2007. https://www.science.org/doi/10.1126/science.1114168. Accessed 12 Mar 2022
Zou WQ. Chameleon-like prion protein and human cognition. Curr Top Biochem Res. 2010;12:1–8.
Zou WQ, Cashman NR. Acidic pH and detergents enhance in vitro conversion of human brain PrPC to a PrPSc-like form. J Biol Chem. 2002;277:43942–7.
Zou WQ, Gambetti P. Prion: the chameleon protein. Cell Mol Life Sci. 2007;64:3266–70.
Zou WQ, Zheng J, Gray DM, Gambetti P, Chen SG. Antibody to DNA detects scrapie but not normal prion protein. Proc Natl Acad Sci U S A. 2004;101:1380–5.
Zou WQ, Langeveld J, Xiao X, Chen S, McGeer PL, Yuan J, et al. PrP conformational transitions alter species preference of a PrP-specific antibody. J Biol Chem. 2010a;285:13874–84.
Zou WQ, Puoti G, Xiao X, Yuan J, Qing L, Cali I, et al. Variably protease-sensitive prionopathy: a new sporadic disease of the prion protein. Ann Neurol. 2010b;68:162–72.
Zou RS, Fujioka H, Guo JP, Xiao X, Shimoji M, Kong C, Chen C, Tasnadi M, Voma C, Yuan J, Moudjou M, Laude H, Petersen RB, Zou WQ. Characterization of spontaneously generated prion-like conformers in cultured cells. Aging. 2011a;3:968–84.
Zou WQ, Xiao X, Yuan J, Puoti G, Fujioka H, Wang X, et al. Amyloid-{beta}42 interacts mainly with insoluble prion protein in the Alzheimer brain. J Biol Chem. 2011b;286:15095–105.
Zou WQ, Zhou X, Yuan J, Xiao X. Insoluble cellular prion protein and its association with prion and Alzheimer diseases. Prion. 2011c;5:172–8.
Zou WQ, Gambetti P, Xiao X, Yuan J, Langeveld J, Pirisinu L. Prions in variably protease-sensitive prionopathy: an update. Pathogens. 2013;2:457–71.
Acknowledgements
This work was supported by the National Institutes of Health (NIH) grant NS112010, NIH NS109532, the BAND grant jointly funded by the Alzheimer’s Association, Alzheimer’s Research, UK, Michael J. Fox Foundation for Parkinson’s Research (MJFF), and Weston Brain Institute, α-synuclein seed amplification grant supported by MJFF, and the CJD Foundation grant.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Zou, WQ. (2023). Insoluble Cellular Prion Protein and Other Neurodegeneration-Related Protein Aggregates in the Brain of Asymptomatic Individuals. In: Zou, WQ., Gambetti, P. (eds) Prions and Diseases. Springer, Cham. https://doi.org/10.1007/978-3-031-20565-1_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-20565-1_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-20564-4
Online ISBN: 978-3-031-20565-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)