Abstract
Pediatric drug development faces many difficulties. Traditionally, pediatric drug doses are simply calculated linearly based on the body weight, age, and body surface area of adults. Due to the ontogeny of children, this simple linear scaling may lead to drug overdose in pediatric patients. The physiologically based pharmacokinetic (PBPK) model, as a mathematical model, contributes to the research and development of pediatric drugs. An example of a PBPK model guiding drug dose selection in pediatrics has emerged and has been approved by the relevant regulatory agencies. In this review, we discuss the principle of the PBPK model, emphasize the necessity of establishing a pediatric PBPK model, introduce the absorption, distribution, metabolism, and excretion of the pediatric PBPK model, and understand the various applications and related prospects of the pediatric PBPK model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Pediatric drug development is faced with many difficulties, such as ethical problems, low consent rates from neonate parents, and limited blood volume availability (1). In addition, the same drug may need to be studied separately for neonates, infants, children, and adolescents. For safety reasons, each subgroup may have different dose selections, different adverse reactions, and different dosage forms. The high cost and complexity of the research both lead to a lack of pediatric drug research, forcing pediatricians to use drugs with little evidence-based data to support drug and dose selections and none of the expected information about efficacy and side effects (2). Although legislative actions such as the Food and Drug Administration Safety and Innovation Act (FDASIA) and the Best Pharmaceuticals for Children Act (BPCA) have clearly defined and provided incentives for appropriate research on children, pediatric drug trial issues are still not properly solved (3, 4).
Physiologically based pharmacokinetic (PBPK) models, as mathematical models, can provide a quantitative mechanical framework for predicting the pharmacokinetics of exogenous substances in humans or animals. The physiological state and the process of absorption, distribution, metabolism, and excretion (ADME) can be reflected (5). Most organs related to ADME, such as the heart, lung, brain, stomach, spleen, pancreas, intestine, liver, kidney, adipose tissue, muscle, bone, and skin, can be integrated by the PBPK model. These tissues are connected by arteries and veins. Each of which is characterized by a relevant blood flow rate, volume, tissue partition coefficient, and permeability (6).
The PBPK model can serve to integrate multiple levels of information (i.e., in vitro, preclinical, or clinical) to elucidate PK changes among children, so it has advantages in predicting the pharmacokinetics of adults and extrapolating the data to children (7). Some details of the PBPK model, such as age-related PK differences in mABs (8), ontogenic classification (9), and strategies to handle PK parameters across different age groups (10), would add to the information regarding rational dose and trial designs. The PBPK model has been widely applied as a tool for decision making, study optimization, and data analysis by academia, the pharmaceutical industry, and the regulatory agencies of pediatric drug development and therapy (11,12,13). The PBPK model has a variety of applications, such as guiding first dosing (14,15,16), predicting tissue drug concentration (17), estimating potential drug-drug interactions (DDIs) in pediatric patients, and describing the effects of organ impairment on pediatric pharmacokinetics (18).
PHYSIOLOGICALLY BASED PHARMACOKINETIC MODEL
Development of PKPB Model
To develop the PKPB model, we need to collect some parameters. The parameters of the PBPK model are divided into three categories: (1) physiological parameters; (2) drug parameters; and (3) research design parameters (19). These parameters can be used to develop PBPK models through software such as SimCyp, GastroPlus, and PK-Sim, each of which has its own advantages (20) in simulating and predicting various hypothetical scenarios. The parameters in the model can either be obtained from the literature or calculated from verified formulas. One way to quantify the contribution of different organs to total clearance is to use in vitro and in vivo extrapolation (IVIVE) (21). When predicting the impact of internal and external factors on drug PK, the PBPK-IVIVE model shows its value in guiding decision making (22).
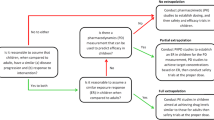
Whether the PBPK model can be widely used in pediatric drug development and personalized administration of drugs depends on the quality of the model and the accuracy of predicting exposure. However, considering the limitation of our knowledge, prediction may not always accurately match the clinical data. For example, some metabolic enzymes in children cannot be quantified (23). The PBPK model should reasonably describe all the parameters (12), but uncertainty analyses focus on the spread or distribution of simulated outputs due to the ambiguity surrounding input parameters and model structure. For sensitivity analyses, the emphasis is shifted toward quantitatively describing the influence of model parameters on simulated outputs. Uncertainty analysis can be conducted in an iterative manner by incorporating the potential range of ambiguous parameter values into simulations (24). Quantitative measures of sensitivity can be attained by varying individual model parameters by 1% and evaluating the impact on simulated outputs (7, 25). Model parameters with certainty can be determined in advance so that the computational cost can be greatly reduced (26, 27). Where some uncertainty about the similarity of disease and/or response to intervention remains, this is referred to as partial extrapolation. In partial extrapolation, “confirmation of efficacy” is required, generally either through a single, controlled or uncontrolled efficacy, and safety trial, as well as evidence demonstrating a similar exposure-response relationship (28).
The Necessity of Pediatric PBPK Model
Traditionally, drug doses for children are calculated based on weight (e.g., Clark’s rule), age (e.g., Young’s rule), and body surface area (e.g., mg/m2) (20). However, when the absorption and disposition of drugs are complicated, these rules cannot accurately predict the drug exposure of pediatric patients. For neonates, a huge physiological change will occur during the course of treatment. When adjusting the dose according to weight, pediatric patients may be premature babies weighing 400 g or morbidly obese adolescents weighing up to 250 kg. The dosage of the drug in both cases is adjusted according to the weight of adults (2). When adjusting the dose according to age, for children under five years old, the younger the age is, the greater the dose deviation (29, 30). Rapid changes in organ maturity, blood flow, body composition, drug elimination, and transport mechanisms in developing children of all age groups, especially in neonates, are not considered. During the neonatal period, some metabolic enzymes that do not exist at birth rise rapidly after a few days, while some other metabolic enzymes disappear quickly a few days after birth. Dose adjustment according to body surface area can lead to overdose in neonates and infants (31). The PBPK model can predict children’s clearance rate and guide children on medication by adjusting physiological parameters according to changes in children’s physiological processes (32, 33). To develop a PBPK model, all the factors that affect the ADME of the drug must be addressed, as shown in Fig. 1.
ADME OF PEDIATRIC PBPK MODEL
A common way to develop a pediatric PBPK model is to modify the adult PBPK model, which has been verified by adult PK data. An adult PBPK model enables researchers to understand the adult ADME of the drug before extrapolating the data to different age groups of children. However, a poor prediction by the adult PBPK model will also be reflected in the pediatric PBPK model, an issue of concern to researchers developing pediatric research (7). Therefore, the physiological differences between adults and children must be understood.
Absorption
Oral administration is the most commonly used method in pediatrics. However, absorption of oral administration is affected by a variety of physiological factors, such as gastric pH and emptying time, intestinal transit time, and intestinal volume, which vary with sex, race, food effects, and diseases (34). For example, changes in pH in different parts of the gastrointestinal tract directly affect the relative absorption of the drug by affecting the stability and ionization degree of the drug. For neonates, the pH in the stomach is relatively high (pH> 4) due to the decrease in the production of basal acids and the total amount of gastric secretions. Age-dependent changes in bile function also affect its ability to dissolve and absorb lipophilic drugs. For neonates, both the binding and transport of bile salts are immature, leading to a low level of absorption in the duodenum (35).
Some PBPK models simulate the oral absorption of biopharmaceutical classification system (BCS) class I compounds (such as theophylline and paracetamol) and BCS class II compounds (such as ketoconazole) (36, 37). The absorption of BCS class I compounds in neonates was predicted to be slower than that in the older age group, while the fraction absorption was the same. The Tmax of ketoconazole was predicted to be 1 h in both neonates and adults, while the FA of ketoconazole was higher in neonates (36). If given a high dose of BCS class II compounds, pediatric patients will have reduced drug absorption due to their reduced gastrointestinal size and transport time (37).
Moreover, the high frequency of neonatal feeding has a huge effect on oral drug absorption. Feeding can cause physiological changes, including gastrointestinal pH, delayed gastric emptying, bile secretion, and liver blood flow. It is very important to understand these changes for the development of the PBPK model (38).
Distribution
After the drug is absorbed, it will be distributed to tissues and organs. The PBPK model can describe this process by the tissue-plasma partition coefficient (Kp) of fat, bone, brain, intestine, heart, kidney, liver, lung, muscle, pancreas, skin, spleen, and thymus. The value of Kp depends on the fractional volume of water and lipids in tissue and the level of plasma binding protein (39, 40).
Age-dependent changes in body composition alter the physiological space of drug distribution. Compared with adults, the total-body water space in neonates and infants is relatively larger, and their ratio of water to lipids is also higher (35).
The level of plasma binding protein in neonates is lower than that in adults and increases with age. For neonates, the concentration of human serum albumin (HSA) is approximately 75–80% that of adults, while the concentration of alpha 1-glycoprotein (AAG) is initially half that of adults. Compared with drugs that bind to AAG, the distribution of drugs in neonates that bind to HSA is more similar to the distribution in adults. Bilirubin and free fatty acids competitively bind to plasma proteins, resulting in an increase in the free portion of the drug, and may influence the extent of binding of some drugs in neonates (41).
Metabolism
Drug-metabolizing enzymes can be divided into phase I enzymes (mainly cytochrome P450, CYP) and phase II enzymes (e.g., UDP-glucuronosyltransferase, UGT) (42). One of the main advantages of the pediatric PBPK model is the inclusion of enzyme ontogeny. CYP enzymes are a major contributor to the clearance of approximately 2/3 of prescription drugs (43).
The ontogeny of CYP is has been reported to be the main cause of age-dependent clearance (44). Drug metabolism occurs mainly in the liver, and the liver and kidney clearance systems of infants or neonates are more immature than those of adults, especially in the first few weeks of life. The delayed maturation of drug-metabolizing enzymes may be the reason for the significant drug toxicity in infants (45). The expression of phase I enzymes changes significantly during ontogeny. CYP3A7 is the main CYP3A subtype expressed in the fetal liver, the expression of which reaches a peak shortly after birth and then drops rapidly. CYP3A4 and CYP2C both appear in the first week of life, while the relative expression of most CYP enzymes is relatively low at birth and increases with age, reaching a peak at approximately five years old. The relative expression of 6 types of CYP is shown in Fig. 2 (35, 44, 46,47,48).
Metabolism mediated by the phase II enzyme UGT is an important binding reaction. UGT-mediated drug clearance was evaluated by ultrahigh-performance liquid chromatography-tandem mass spectrometry (UHPLC-MS/MS) (49). Studies have shown significant differences in the expression and activity of UGT among different subtypes in different age groups and age-related changes in the activity of different UGT types (50). The relative expression of 7 human phase II drug-metabolizing enzymes at different ages is shown in Fig. 3 (51,52,53).
At present, some studies have suggested that age-related changes in the clearance of drugs by major CYP enzymes in the liver, such as theophylline (CYP1A2), metronidazole (CYP2A6), isavelam (CYP2B6), desloratadine, montelukast (CYP2C8), diclofenac, (S)-warfarin (CYP2C9), esomeprazole, lansoprazole (CYP2C19), tramadol, tolterodine (CYP2D6), itraconazole, ondansetron, and sufentanil (CYP3A4), were accurately predicted (44, 54). Some studies have suggested hepatic intrinsic clearance and fraction metabolism (Fig. 4) (55, 56).
Hepatic clearance depends on several factors, including blood flow, hepatic enzyme activities, transport systems, and plasma protein binding. Blood flow and drug-metabolizing enzymes are low in children, while the former reaches adult levels by approximately one year of age (57). In the PBPK model of some drugs, liver clearance is often calculated according to metabolic enzyme activities and the proportion of the drug cleared by various metabolic enzymes (58). Drugs eliminated via the same enzymes require similar dose adjustments only in specific cases. We could establish a priori whether two drugs metabolized by the same isoenzyme will require the same or different dose adjustments in pediatric patients (59).
For pediatric patients, the elimination mechanism of the same drug may be different from that of adults, so these drugs require some adjustment based on the pharmacokinetic data. Esomeprazole is both a substrate and inhibitor of CYP2C19. The CYP2C19 autoinhibition model was appropriate for esomeprazole in adults and older children but could not be directly extended to infants. The prediction was markedly improved by assuming no autoinhibition of esomeprazole in infants in the PBPK model. This result may be due to a compensatory pathway mediated by other enzymes (60). This study reminds us that it is necessary to thoroughly understand the complex interactions among enzyme maturation, inhibition, and compensatory mechanisms for PBPK modeling in infants.
Excretion
The kidney is one of the main excretory organs, and most drugs are eliminated from the body through the kidney. Renal excretion of the drug depends on glomerular filtration rate, tubular secretion, and tubular reabsorption. The glomerular filtration rate of premature infants is approximately half of that of neonates. The permeability of the glomerulus increases rapidly in the first 2 weeks of life and then increases steadily until 8 to 12 months of age. Similarly, tubules are immature at birth and reach adult capacity in the first year (35). Tubular secretion can be extended to the PBPK model to make the prediction more accurate (61). For intrarenal elimination drugs in neonates and infants, clearance often depends on growth, age, and renal function (62).
The ontogeny of some renal membrane transporters shows an age-related model, indicating that clearance is consequently age-related (63). When the drug is eliminated only through the kidney, the model based on body weight and postmenstrual age can initially predict the drug clearance rate of neonates and infants with normal renal function and provide drug administration strategies for neonates and infants (64). Renal clearance models of linezolid and toratabine based on body surface area were established, and because their elimination pathway is relatively simple, the adult model can be directly and accurately extended to the pediatric population (65).
For drugs eliminated by glomerular filtration (GF), the clearance is determined by the glomerular filtration rate (GFR) and the unbound portion of the drug. The clearance was measured by GFR function, and this method can minimize the effect of mature plasma protein concentration on GF (66, 67).
Transporter
Membrane transporters control the transport of drugs and affect the absorption, distribution, metabolism, and excretion of drugs (68). Cheung et al. introduced the ontogeny of intestinal, hepatic, and renal transporters and the recent development direction. The PBPK model of tazobactam was also established to illustrate the integrated renal transporter ontogeny function in simulating exposure (69). The PBPK model incorporated the ontogeny and pharmacogenetic effect of OCT1 and adequately predicted the pharmacokinetics of morphine (65). Integrating ontogeny data of these transporters in the PBPK model will be a key step in predicting drug disposition in pediatric patients, since the expression of P-gp, OCT1, and OATP1B3 was significantly lower in neonates or infants than in adolescents and adults (66). Protein expression of major hepatic uptake and efflux drug transporters in human pediatric and adult livers was quantified by liquid chromatography-tandem mass spectrometry. Transporter protein expression of OCT1, P-gp, MRP2, etc. was age-dependent (Fig. 5 (70,71,72,73)). The main reason for this difference between different age groups is the expression of mRNA (69).
Extrapolation of Pediatric PBPK Model
Understanding the pharmacokinetics of drugs in the body is important in the development of new drugs. We could combine the PBPK model with IVIVE to predict the pharmacokinetics of drugs (74). IVIVE could be used to determine the activity of metabolic enzymes and transporters. With the data-based development of PBPK models, in vitro metabolism data have been accepted as PBPK model parameters by the pharmaceutical industry. By scaling the enzyme content in vitro to the metabolic parameters in vivo, the metabolic constants measured in vitro can be “proportionally magnified” to the corresponding metabolic parameters in vivo in the PBPK model. The PBPK model could be developed by intrinsic clearance values collected from subcellular components, primary hepatocytes, and expressing enzymes (75,76,77). A PBPK model was established to predict the effect of rifampicin on CYP3A4 induction in vivo based on the in vitro data from primary human hepatocyte culture supernatants from 14 volunteers (78). However, to improve the accuracy of the model, it is recommended that the cell-based assay incubated in 100% human serum be used to estimate the relevant parameters or to correct the experimental data at the same time (79). Ex vivo cotyledon perfusion experiments are currently considered the gold standard for studying maternal-fetal drug transfer. The ex vivo cotyledon perfusion experiment could provide the key parameters governing placental transfer (the placental transfer constant and the partition coefficient) (80).
For animal-human extrapolation, at present, it is believed that data from young animals can better predict the pharmacokinetics of pediatric patients than that from adult animals. The deviation in the extrapolation process from young rats to neonates needs to be corrected by maximum lifespan potential (MLP), which could substantially improve the prediction of antimalarial drug clearance in children (81). The PBPK model, based on data extrapolated by MLP from juvenile mice to neonates, provided a consistent prediction of the pharmacokinetics of fluconazole in neonates and proved the feasibility of this method to verify the first dose of fluconazole in neonates (82). The results of the PBPK model of midazolam and clindamycin supported a more accurate prediction using MLP than brain weight (83). In these studies, MLP has been used as a correction factor of interspecies scaling from animals to humans.
The PBPK model allows reasonable scaling between adults and children according to related physiological differences. Scaling adult PBPK to pediatric PBPK for different age groups requires a sufficient understanding of disease pathogenesis, the mechanism of action of the drug, and its pharmacological behavior. We should assess the evidence supporting the similarity of the disease course between the reference and pediatric populations, whether the evidence supports a similar exposure-response between the reference and intended populations, and uncertainties or limitations of the existing data (e.g., clinical or historical data and published literature) (84). Some adult PBPK models have been successfully extended to children to predict plasma concentration-time profiles in pediatric patients. A PBPK model of lisinopril has been developed to predict pediatric doses in neonates to infants, infants to toddlers, children of preschool age, children of school age and adolescents (85, 86). PBPK models of ganciclovir and its prodrug valganciclovir were developed, and the clinical models were gradually verified using the pharmacokinetic data of adults, children, and neonates (87). PBPK models of buprenorphine and norbuprenorphine in adults have been developed and extrapolated to children and preterm neonates. PBPK models could be helpful to further investigate the pharmacokinetics of buprenorphine in pediatric patients (88).
APPLICATION OF PBPK IN PEDIATRIC DRUG
Dose Prediction
Compared with dose predictions calculated based on weight, age, and body surface area, the PBPK model is more accurate because related physiological changes are considered (86). In the absence of sufficient clinical data, the pediatric PBPK model can be chosen to use existing knowledge to indirectly infer pediatric dose.
The PBPK model of lisinopril, which was suitable for oral administration (feeding and fasting) and intravenous injection in healthy adults, was scaled to a virtual pediatric population. By developing a virtual PBPK model of neonates for 0–28 days, the pharmacokinetics of dolutegravir and several multidose regimens of dolutegravir were successfully simulated and predicted. We could choose a theoretical optimal dosing regimen (89). The PBPK model can be developed according to the pathophysiological changes of patients. For example, a drug-disease PBPK model of ibuprofen was developed by incorporating pathophysiological changes occurring with cystic fibrosis to present along with therapy optimization suggestions (90).
The PBPK model can also guide experimental designs through dose prediction (91). The risk involved in clinical trials of some drugs in neonates can be reduced through the PBPK model. The dose of trazodone predicted through PBPK modeling can be used to guide administration in initial clinical trials in pediatric patients. The conduct of a clinical trial in pediatric patients was based on previous dose predictions endorsed by the European Medicines Agency. Ethical and regulatory approvals for the clinical trial were also based on the doses predicted in this analysis (16). The pediatric PBPK model in combination with the population pharmacokinetics (PopPK) model could be further used to guide dose selections for treatment (92).
Dose prediction by the PBPK model can be rapid and effective. A previously developed PBPK model for chloroquine (CHQ) was used to simulate exposure in adults and children and was verified against published pharmacokinetic data. The results of the study guided dose selection for children of different ages (93). Although the efficacy of CHQ is still in dispute, it is undeniable that the PBPK model has rapid and economic advantages in dose selection. PBPK modeling of remdesivir and its metabolites also rapidly supports dose selection for the treatment of pediatric patients with COVID-19 (94).
Drug-Drug Interactions
Due to ethical and other reasons, pediatric DDI studies are rarely carried out, and most DDI data come from adult clinical studies and case reports. The absence of clinical information is the major sticking point to extrapolate adult DDI data to the pediatric population, and extrapolation from adult data may not be applicable across all pediatric age groups (50). Using the PBPK model can integrate our understanding in this field and may help to predict pediatric DDIs.
The PBPK model has been used to evaluate DDI liability at the drug discovery or development stage. Caffeine and ciprofloxacin were used as tool compounds to illustrate the application of the PBPK method in predicting human pharmacokinetics and DDI. Overall, the increase in Cmax of caffeine by ciprofloxacin was not significant. However, an increase in AUC was observed and was proportional to the administered dose of ciprofloxacin (95). When the clinical DDI data are limited, PBPK modeling can potentially predict pediatric DDI.
PBPK models, including the activity and ontogeny of CYP3A and CYP2C9, were established and verified by PK data from premature infants who received sildenafil with or without fluconazole. The PBPK model described age-related differences in CYP3A metabolism. This sildenafil PBPK model can be used to develop PBPK models for other CYP3A inhibitors or inducers. PBPK models of drugs that have DDI with sildenafil, such as erythromycin and protease inhibitors, can guide dose selection (96). The PBPK model of efavirenz was established by incorporating in vitro and clinical PK data. The DDI study was utilized to evaluate the CYP3A4 induction component in the liver and gut. This validated mechanistic model can now be applied in clinical pharmacology studies to prospectively assess the potential DDI of efavirenz as the victim and perpetrator (27).
Physiological Changes
The PBPK model can help researchers better understand the pharmacokinetic changes caused by these physiological differences. Voriconazole is an effective antifungal drug. After the first-pass intestinal metabolism was incorporated into the pediatric PBPK model of voriconazole, the prediction accuracy was significantly improved. Voriconazole was affected by first-pass intestinal metabolism in children but not in adults (97). The PBPK model was used to predict the PK profile of methadone in the treatment of neonatal abstinence syndrome (NAS). The analysis suggested that the activity of the P450 enzyme impacted the clearance of methadone in adults and neonates, and enzymatic polymorphisms (e.g., CYP2B6 and CYP3A4) may contribute to the interpatient variability in PK (98).
Prediction of Tissue Concentration
For some tissues from which drug concentrations are difficult to measure directly, it is ideal to use PBPK modeling to predict drug concentrations (90). Under conditions other than cardiac surgery, it seems impossible to analyze and evaluate drug concentrations in human heart tissue. Especially for children, the risk of surgery is high. Therefore, a PBPK model predicting drug concentration in the human heart can be developed to assess model-based cardiac safety (24). Drug disposition in cerebrospinal fluid (CSF) is also affected by age-related variations as well as by brain diseases that affect the integrity of the blood-brain barrier, such as meningitis. A generic pediatric brain PBPK model was developed to predict the CSF concentrations of drugs that undergo passive transfer, which was verified by multiple drugs (99).
Understanding the concentration-time profile of targets in the central nervous system (CNS) is very important for the development of targeted drugs in the CNS. A PBPK model was developed using the physiological characteristics of children that predicted the plasma morphine concentration in the CNS of individual patients. This model can study the potential mechanism occurring in the CNS and explain the PK differences related to pathophysiological changes (17).
Monoclonal Antibodies in Pediatric Populations
In the past two decades, there has been a surge in the number of macromolecular drugs registered in children, most of which are monoclonal antibodies (mAbs), which can also be modeled by PBPK. Unlike smaller molecular drugs, mAbs are easily hydrolyzed by digestive enzymes in the gastrointestinal tract and are usually injected intravenously. MAbs have high molecular weights and strong hydrophilicity, hindering their ability to freely spread into the tissue. Therefore, mAbs are generally limited to the vascular and tissue interstitial space. MAbs are eliminated mainly by proteolytic enzyme catabolism, and the FcRn (neonatal Fc receptor) has a protective effect on the elimination of drugs. After binding, mAbs can avoid being degraded by intracellular lysosomes, and the cycle of mAbs in vivo can be strengthened (100, 101). With the relatively high concentrations of endogenous IgG after birth, the lower FcRn concentration would lead to a higher body weight-normalized clearance of therapeutic proteins for very young children (102). This leads to a slightly higher body weight-normalized dose in younger children than in adults to obtain equivalent exposure (103).
Infliximab is a chimeric mAb directed against tumor necrosis factor alpha (TNF) that is used to treat inflammation associated with many autoimmune conditions. Antidrug antibody molecules (IgM, IgG) bind to infliximab to form a compound, which causes infliximab to lose its therapeutic effect and eventually be degraded. The PBPK model was successfully developed by optimizing two parameters: the rate of uptake into endosomal space and the zero-order molar synthesis rate of antidrug antibody molecules against infliximab. The findings can be generalized to pediatric PBPK models of other monoclonal antibody drugs with similar FcRn affinity and a low target burden to guide drug development and selection of first-in-pediatric doses (104). For the PBPK model of palivizumab, after introducing ontogenies of the capillary density, lymph flow, and leukocyte concentration, the predictions were in good agreement with the observed data (105).
Limit and Expectation
However, inevitably, the PBPK model also has some limitations in its theoretical development and practical application. Published data are usually presented only graphically. Researchers usually use software to extract data from the graphs, which produces errors in the process of digitization and interferes with the process of model development and evaluation. Therefore, the original data from each study should be provided in publications (106). The credibility of the parameters often represents the credibility of the model. For some uncertain parameters, although sensitivity analysis can be used to check the impact of each parameter, there is no general guide to select an appropriate range of variability for the model parameters (20). Therefore, we need greater understanding of the physiological profile of children and the process of drug metabolism in children. By incorporating more accurate model parameters, PBPK models can be continuously improved through the learning and confirmation cycle (107).
One promising field in the future is the application of PBPK models in personalized medicine, which requires more detailed characteristic physiological information. Developing a “virtual twin” that takes patient-specific characteristics into account in the PBPK model will help to better predict and personalize doses in specific age groups. Combining real patients with “virtual twins” will allow pediatric PBPK models to be better applied in clinical practice (108). In summary, using the PBPK model requires a detailed understanding of the drugs to be modeled. For pediatric applications, it includes the in vitro data on age-related physiological and biochemical changes regarding the most likely ontogeny scenario. Overall, the application of PBPK to guide pediatric clinical trials is a personalized medical application (109). In all stages of pediatric drug development, PBPK models should be conducted routinely to help pediatric drug research and development (110).
CONCLUSION
In this review, the development and necessity of the pediatric PBPK model have been introduced. The ADME of and extrapolation of the pediatric PBPK model have been summarized, especially the ontogeny of metabolic enzymes and the application, limits, and expectation of PBPK in pediatric drugs, such as the pediatric PBPK model of monoclonal antibody drugs. With further understanding of physiological mechanisms and clinical patient data, the PBPK model could provide greater accuracy at the individual patient level to support earlier decisions for new drug development for children.
References
Laughon MM, Avant D, Tripathi N, Hornik CP, Cohen-Wolkowiez M, Clark RH, et al. Drug labeling and exposure in neonates. JAMA Pediatr. 2014;168(2):130–6.
van den Anker JN, Coppes MJ, Koren G. Neonatal and pediatric clinical pharmacology. Pediatr Clin N Am. 2012;59(5):xv–xviii.
.FDA. Food and Drug Administration Safety and Innovation Act (FDASIA). Silver spring: FDA; 2012.https://www.fda.gov.Accessed 9 JULY, 2012.
Avant D, Baer G, Moore J, Zheng P, Sorbello A, Ariagno R, et al. Neonatal safety information reported to the FDA during drug development studies. Ther Innov Regul Sci. 2017;2017:1–9.
Lucas AJ, Sproston JL, Barton P, Riley RJ. Estimating human ADME properties, pharmacokinetic parameters and likely clinical dose in drug discovery. Expert Opin Drug Discovery. 2019;14(12):1313–27.
Kuepfer L, Niederalt C, Wendl T, Schlender JF, Willmann S, Lippert J, et al. Applied concepts in PBPK modeling: how to build a PBPK/PD model. CPT Pharmacometrics Syst Pharmacol. 2016;5(10):516–31.
Maharaj AR, Edginton AN. Physiologically based pharmacokinetic modeling and simulation in pediatric drug development. CPT Pharmacometrics Syst Pharmacol. 2014;3(11):e150.
Wong H, Chow TW. Physiologically based pharmacokinetic modeling of therapeutic proteins. J Pharm Sci. 2017;106(9):2270–5.
Ince I, Solodenko J, Frechen S, et al. Predictive pediatric modeling and simulation using ontogeny information. J Clin Pharmacol. 2019;59(Suppl 1):S95–S103.
Stass H, Lettieri J, Vanevski KM, Willmann S, James LP, Sullivan JE, et al. Pharmacokinetics, safety, and tolerability of single-dose intravenous moxifloxacin in pediatric patients: dose optimization in a phase 1 study. J Clin Pharmacol. 2019;59(5):654–67.
Jamei M. Recent advances in development and application of physiologically-based pharmacokinetic (PBPK) models: a transition from academic curiosity to regulatory acceptance. Curr Pharmacol Rep. 2016;2(3):161–9.
Wagner C, Zhao P, Pan Y, et al. Application of physiologically based pharmacokinetic (PBPK) modeling to support dose selection: report of an FDA public workshop on PBPK. CPT Pharmacometrics Syst Pharmacol. 2015;4(4):226–30.
Zhao P. Report from the EMA workshop on qualification and reporting of physiologically based pharmacokinetic (PBPK) modeling and simulation. CPT Pharmacometrics Syst Pharmacol. 2017;6(2):71–2.
Adiwidjaja J, Boddy AV, McLachlan AJ. Implementation of a physiologically based pharmacokinetic modeling approach to guide optimal dosing regimens for imatinib and potential drug interactions in paediatrics. Front Pharmacol. 2020;10:1672.
Martins FS, Zhu P, Heinrichs MT, Sy SKB. Physiologically based pharmacokinetic-pharmacodynamic evaluation of meropenem plus fosfomycin in paediatrics. Br J Clin Pharmacol. 2020;87:1012–23. https://doi.org/10.1111/bcp.14456.
Oggianu L, Ke AB, Chetty M, Picollo R, Petrucci V, Calisti F, et al. Estimation of an appropriate dose of trazodone for pediatric insomnia and the potential for a trazodone-atomoxetine interaction. CPT Pharmacometrics Syst Pharmacol. 2020;9(2):77–86.
Ketharanathan N, Yamamoto Y, Rohlwink UK, Wildschut ED, Mathôt RAA, de Lange ECM, et al. Combining brain microdialysis and translational pharmacokinetic modeling to predict drug concentrations in pediatric severe traumatic brain injury: the next step toward evidence-based pharmacotherapy? J Neurotrauma. 2019;36(1):111–7.
Ye L, Ke M, You X, Huang P, Lin C. A physiologically based pharmacokinetic model of ertapenem in pediatric patients with renal impairment. J Pharm Sci. 2020;109(9):2909–18.
Jamei M, Dickinson GL, Rostami-Hodjegan A. A framework for assessing inter-individual variability in pharmacokinetics using virtual human populations and integrating general knowledge of physical chemistry, biology, anatomy, physiology and genetics: a tale of ‘bottom-up’ vs ‘top-down’ recognition of covariates. Drug Metab Pharmacokinet. 2009;24(1):53–75.
Yellepeddi V, Rower J, Liu X, Kumar S, Rashid J, Sherwin CMT. State-of-the-art review on physiologically based pharmacokinetic modeling in pediatric drug development. Clin Pharmacokinet. 2019;58(1):1–13.
Proctor NJ, Tucker GT, Rostami-Hodjegan A. Predicting drug clearance from recombinantly expressed CYPs: intersystem extrapolation factors. Xenobiotica. 2004;34(2):151–78.
Rostami-Hodjegan A. Physiologically based pharmacokinetics joined with in vitro-in vivo extrapolation of ADME: a marriage under the arch of systems pharmacology. Clin Pharmacol Ther. 2012;92(1):50–61.
Johnson TN, Cleary Y, Parrott N, Reigner B, Smith JR, Toovey S. Development of a physiologically based pharmacokinetic model for mefloquine and its application alongside a clinical effectiveness model to select an optimal dose for prevention of malaria in young Caucasian children. Br J Clin Pharmacol. 2019;85(1):100–13.
Tylutki Z, Mendyk A, Polak S. Mechanistic physiologically based pharmacokinetic (PBPK) model of the heart accounting for inter-individual variability: development and performance verification. J Pharm Sci. 2018;107(4):1167–77.
Ke A, Barter Z, Rowland-Yeo K, Almond L. Towards a best practice approach in PBPK modeling: case example of developing a unified efavirenz model accounting for induction of CYPs 3A4 and 2B6. CPT Pharmacometrics Syst Pharmacol. 2016;5(7):367–76.
van Groen BD, Pilla Reddy V, Badée J, et al. Pediatric pharmacokinetics and dose predictions: a report of a satellite meeting to the 10th Juvenile Toxicity Symposium. Clin Transl Sci. 2020;14:29–35. https://doi.org/10.1111/cts.12843.
Calvier EAM, Nguyen TT, Johnson TN, Rostami-Hodjegan A, Tibboel D, Krekels EHJ, et al. Can population modelling principles be used to identify key PBPK parameters for paediatric clearance predictions? An Innovative Application of Optimal Design Theory. Pharm Res. 2018;35(11):209.
Conklin LS, Hoffman EP, Van den Anker J. Developmental pharmacodynamics and modeling in pediatric drug development. J Clin Pharmacol. 2019;(Suppl 1):S87–94. https://doi.org/10.1002/jcph.1482.
Calvier EA, Krekels EH, Välitalo PA, et al. Allometric scaling of clearance in paediatric patients: when does the magic of 0.75 fade? Clin Pharmacokinet. 2017;56(3):273–85.
Mahmood I, Staschen CM, Goteti K. Prediction of drug clearance in children: an evaluation of the predictive performance of several models. AAPS J. 2014;16(6):1334–43.
Johnson TN. The problems in scaling adult drug doses to children. Arch Dis Child. 2008;93(3):207–11.
Jadhav PR, Kern SE. The need for modeling and simulation to design clinical investigations in children. J Clin Pharmacol. 2010;50(9 Suppl):121S–9S.
Bellanti F, Della Pasqua O. Modelling and simulation as research tools in paediatric drug development. Eur J Clin Pharmacol. 2011;1(Suppl 1):75–86.
Jamei M, Turner D, Yang J, Neuhoff S, Polak S, Rostami-Hodjegan A, et al. Population-based mechanistic prediction of oral drug absorption. AAPS J. 2009;11(2):225–37.
Kearns GL, Abdel-Rahman SM, Alander SW, Blowey DL, Leeder JS, Kauffman RE. Developmental pharmacology--drug disposition, action, and therapy in infants and children. N Engl J Med. 2003;349(12):1157–67.
Johnson TN, Bonner JJ, Tucker GT, Turner DB, Jamei M. Development and applications of a physiologically-based model of paediatric oral drug absorption. Eur J Pharm Sci. 2018;115:57–67.
Kohlmann P, Stillhart C, Kuentz M, Parrott N. Investigating oral absorption of carbamazepine in pediatric populations. AAPS J. 2017;19(6):1864–77.
Cheng L, Wong H. Food effects on oral drug absorption: application of physiologically-based pharmacokinetic modeling as a predictive tool. Pharmaceutics. 2020;12(7):672.
Rodgers T, Leahy D, Rowland M. Physiologically based pharmacokinetic modeling 1: predicting the tissue distribution of moderate-to-strong bases. J Pharm Sci. 2005;94(6):1259–76.
Rodgers T, Rowland M. Physiologically based pharmacokinetic modelling 2: predicting the tissue distribution of acids, very weak bases, neutrals and zwitterions. J Pharm Sci. 2006;95(6):1238–57.
McNamara PJ, Alcorn J. Protein binding predictions in infants. AAPS PharmSci. 2002;4(1):E4.
Mahmood I, Ahmad T, Mansoor N, Sharib SM. Prediction of clearance in neonates and infants (≤ 3 months of age) for drugs that are glucuronidated: a comparative study between allometric scaling and physiologically based pharmacokinetic modeling. J Clin Pharmacol. 2017;57(4):476–83.
Williams JA, Hyland R, Jones BC, Smith DA, Hurst S, Goosen TC, et al. Drug-drug interactions for UDP-glucuronosyltransferase substrates: a pharmacokinetic explanation for typically observed low exposure (AUCi/AUC) ratios. Drug Metab Dispos. 2004;32(11):1201–8.
Upreti VV, Wahlstrom JL. Meta-analysis of hepatic cytochrome P450 ontogeny to underwrite the prediction of pediatric pharmacokinetics using physiologically based pharmacokinetic modeling. J Clin Pharmacol. 2016;56(3):266–83.
Johnson TN, Rostami-Hodjegan A. Resurgence in the use of physiologically based pharmacokinetic models in pediatric clinical pharmacology: parallel shift in incorporating the knowledge of biological elements and increased applicability to drug development and clinical practice. Paediatr Anaesth. 2011;21(3):291–301.
Song G, Sun X, Hines RN, McCarver DG, Lake BG, Osimitz TG, et al. Determination of human hepatic CYP2C8 and CYP1A2 age-dependent expression to support human health risk assessment for early ages. Drug Metab Dispos. 2017;45(5):468–75.
Stevens JC, Hines RN, Gu C, Koukouritaki SB, Manro JR, Tandler PJ, et al. Developmental expression of the major human hepatic CYP3A enzymes. J Pharmacol Exp Ther. 2003;307(2):573–82.
Stevens JC, Marsh SA, Zaya MJ, Regina KJ, Divakaran K, le M, et al. Developmental changes in human liver CYP2D6 expression. Drug Metab Dispos. 2008;36(8):1587–93.
Badée J, Qiu N, Parrott N, Collier AC, Schmidt S, Fowler S. Optimization of experimental conditions of automated glucuronidation assays in human liver microsomes using a cocktail approach and ultra-high performance liquid chromatography-tandem mass spectrometry. Drug Metab Dispos. 2019;47(2):124–34.
Badée J, Fowler S, de Wildt SN, Collier AC, Schmidt S, Parrott N. The ontogeny of UDP-glucuronosyltransferase enzymes, recommendations for future profiling studies and application through physiologically based pharmacokinetic modelling. Clin Pharmacokinet. 2019;58(2):189–211.
Abduljalil K, Jamei M, Rostami-Hodjegan A, Johnson TN. Changes in individual drug-independent system parameters during virtual paediatric pharmacokinetic trials: introducing time-varying physiology into a paediatric PBPK model. AAPS J. 2014;16(3):568–76.
Bhatt DK, Basit A, Zhang H, Gaedigk A, Lee SB, Claw KG, et al. Hepatic abundance and activity of androgen- and drug-metabolizing enzyme UGT2B17 are associated with genotype, age, and sex. Drug Metab Dispos. 2018;46(6):888–96.
van Groen BD, Nicolaï J, Kuik AC, van Cruchten S, van Peer E, Smits A, et al. Ontogeny of hepatic transporters and drug-metabolizing enzymes in humans and in nonclinical species. Pharmacol Rev. 2021;73(2):597–678.
Zhou W, Johnson TN, Bui KH, Cheung SYA, Li J, Xu H, et al. Predictive performance of physiologically based pharmacokinetic (PBPK) modeling of drugs extensively metabolized by major cytochrome P450s in children. Clin Pharmacol Ther. 2018;104(1):188–200.
Lang J, Vincent L, Chenel M, Ogungbenro K, Galetin A. Impact of hepatic CYP3A4 ontogeny functions on drug-drug interaction risk in pediatric physiologically-based pharmacokinetic/pharmacodynamic modeling: critical literature review and ivabradine case study. Clin Pharmacol Ther. 2020;109:1618–30. https://doi.org/10.1002/cpt.2134.
T’jollyn H, Vermeulen A, Van Bocxlaer J. PBPK and its virtual populations: the impact of physiology on pediatric pharmacokinetic predictions of tramadol. AAPS J. 2018;21(1):8.
Fernandez E, Perez R, Hernandez A, Tejada P, Arteta M, Ramos J. Factors and mechanisms for pharmacokinetic differences between pediatric population and adults. Pharmaceutics. 2011;3(1):53–72.
Thompson EJ, Wu H, Maharaj A, Edginton AN, Balevic SJ, Cobbaert M, et al. Physiologically based pharmacokinetic modeling for trimethoprim and sulfamethoxazole in children. Clin Pharmacokinet. 2019;58(7):887–98.
Calvier EAM, Krekels EHJ, Yu H, Välitalo PAJ, Johnson TN, Rostami-Hodjegan A, et al. Drugs being eliminated via the same pathway will not always require similar pediatric dose adjustments. CPT Pharmacometrics Syst Pharmacol. 2018;7(3):175–85.
Duan P, Wu F, Moore JN, Fisher J, Crentsil V, Gonzalez D, et al. Assessing CYP2C19 ontogeny in neonates and infants using physiologically based pharmacokinetic models: impact of enzyme maturation versus inhibition. CPT Pharmacometrics Syst Pharmacol. 2019;8(3):158–66.
Edginton AN, Schmitt W, Voith B, Willmann S. A mechanistic approach for the scaling of clearance in children. Clin Pharmacokinet. 2006;45(7):683–704.
Wilbaux M, Fuchs A, Samardzic J, Rodieux F, Csajka C, Allegaert K, et al. Pharmacometric approaches to personalize use of primarily renally eliminated antibiotics in preterm and term neonates. J Clin Pharmacol. 2016;56(8):909–35.
Cheung KWK, van Groen BD, Spaans E, et al. A comprehensive analysis of ontogeny of renal drug transporters: mRNA analyses, quantitative proteomics, and localization. Clin Pharmacol Ther. 2019;106(5):1083–92.
Wang J, Kumar SS, Sherwin CM, Ward R, Baer G, Burckart GJ, et al. Renal clearance in newborns and infants: predictive performance of population-based modeling for drug development. Clin Pharmacol Ther. 2019;105(6):1462–70.
Duan P, Fisher JW, Yoshida K, Zhang L, Burckart GJ, Wang J. Physiologically based pharmacokinetic prediction of linezolid and emtricitabine in neonates and infants. Clin Pharmacokinet. 2017;56(4):383–94.
Salem F, Johnson TN, Abduljalil K, Tucker GT, Rostami-Hodjegan A. A re-evaluation and validation of ontogeny functions for cytochrome P450 1A2 and 3A4 based on in vivo data. Clin Pharmacokinet. 2014;53(7):625–36.
Cristea S, Krekels EHJ, Allegaert K, Knibbe CAJ. The Predictive value of glomerular filtration rate-based scaling of pediatric clearance and doses for drugs eliminated by glomerular filtration with varying protein-binding properties. Clin Pharmacokinet. 2020;59(10):1291–301.
Zhang L, Huang SM, Reynolds K, Madabushi R, Zineh I. Transporters in drug development: scientific and regulatory considerations. Clin Pharmacol Ther. 2018;104(5):793–6.
Cheung KWK, van Groen BD, Burckart GJ, et al. Incorporating ontogeny in physiologically based pharmacokinetic modeling to improve pediatric drug development: what we know about developmental changes in membrane transporters. J Clin Pharmacol. 2019;1(Suppl 1):S56–69.
Hahn D, Emoto C, Euteneuer JC, Mizuno T, Vinks AA, Fukuda T. Influence of OCT1 ontogeny and genetic variation on morphine disposition in critically ill neonates: lessons from PBPK modeling and clinical study. Clin Pharmacol Ther. 2019;105(3):761–8.
Prasad B, Gaedigk A, Vrana M, Gaedigk R, Leeder JS, Salphati L, et al. Ontogeny of hepatic drug transporters as quantified by LC-MS/MS proteomics. Clin Pharmacol Ther. 2016;100(4):362–70.
Bhatt DK, Prasad B. Critical issues and optimized practices in quantification of protein abundance level to determine interindividual variability in DMET proteins by LC-MS/MS proteomics. Clin Pharmacol Ther. 2018;103(4):619–30.
Mooij MG, van de Steeg E, van Rosmalen J, Windster JD, de Koning BAE, Vaes WHJ, et al. Proteomic analysis of the developmental trajectory of human hepatic membrane transporter proteins in the first three months of life. Drug Metab Dispos. 2016;44(7):1005–13.
Jaroch K, Jaroch A, Bojko B. Cell cultures in drug discovery and development: the need of reliable in vitro-in vivo extrapolation for pharmacodynamics and pharmacokinetics assessment. J Pharm Biomed Anal. 2018;147:297–312.
Bosgra S, van de Steeg E, Vlaming ML, Verhoeckx KC, Huisman MT, Verwei M, et al. Predicting carrier-mediated hepatic disposition of rosuvastatin in man by scaling from individual transfected cell-lines in vitro using absolute transporter protein quantification and PBPK modeling. Eur J Pharm Sci. 2014;65:156–66.
Schneckener S, Krauss M, Ghallab A, et al. A systematic evaluation of the use of physiologically based pharmacokinetic modeling for cross-species extrapolation. J Pharm Sci. 2015;104(1):191–206.
Jamei M, Marciniak S, Edwards D, Wragg K, Feng K, Barnett A, et al. The simcyp population based simulator: architecture, implementation, and quality assurance. In Silico Pharmacol. 2013;1:9.
Baneyx G, Parrott N, Meille C, Iliadis A, Lavé T. Physiologically based pharmacokinetic modeling of CYP3A4 induction by rifampicin in human: influence of time between substrate and inducer administration. Eur J Pharm Sci. 2014;56:1–15.
Poulin P, Haddad S. Albumin and uptake of drugs in cells: additional validation exercises of a recently published equation that quantifies the albumin-facilitated uptake mechanism(s) in physiologically based pharmacokinetic and pharmacodynamic modeling research. J Pharm Sci. 2015;104(12):4448–58.
Dallmann A, Liu XI, Burckart GJ, van den Anker J. Drug transporters expressed in the human placenta and models for studying maternal-fetal drug transfer. J Clin Pharmacol. 2019;1(Suppl 1):S70–81. https://doi.org/10.1002/jcph.1491.
Mahmood I. Interspecies scaling for the prediction of drug clearance in children: application of maximum lifespan potential and an empirical correction factor. Clin Pharmacokinet. 2010;49(7):479–92.
Zhao W, Le Guellec C, Benjamin DK Jr, et al. First dose in neonates: are juvenile mice, adults and in vitro-in silico data predictive of neonatal pharmacokinetics of fluconazole. Clin Pharmacokinet. 2014;53(11):1005–18.
Ye PP, Zheng Y, Du B, et al. First dose in neonates: pharmacokinetic bridging study from juvenile mice to neonates for drugs metabolized by CYP3A. Xenobiotica. 2020;50(11):1275–84.
EMA. ICH E11(R1) guideline on clinical investigation of medicinal products in the pediatric population. 2016. https://www.ema.europa.eu/
Yu Y, DuBois SG, Wetmore C, et al. Physiologically based pharmacokinetic modeling and simulation of sunitinib in pediatrics. AAPS J. 2020;22(2):31.
Rashid M, Sarfraz M, Arafat M, Hussain A, Abbas N, Sadiq MW, et al. Prediction of lisinopril pediatric dose from the reference adult dose by employing a physiologically based pharmacokinetic model. BMC Pharmacol Toxicol. 2020;21(1):56.
Lukacova V, Goelzer P, Reddy M, Greig G, Reigner B, Parrott N. A physiologically based pharmacokinetic model for ganciclovir and its prodrug valganciclovir in adults and children. AAPS J. 2016;18(6):1453–63.
Kovar L, Schräpel C, Selzer D, Kohl Y, Bals R, Schwab M, et al. Physiologically-based pharmacokinetic (PBPK) modeling of buprenorphine in adults, children and preterm neonates. Pharmaceutics. 2020;12(6):578.
Bunglawala F, Rajoli RKR, Mirochnick M, Owen A, Siccardi M. Prediction of dolutegravir pharmacokinetics and dose optimization in neonates via physiologically based pharmacokinetic (PBPK) modelling. J Antimicrob Chemother. 2020;75(3):640–7.
Cicali B, Long T, Kim S, Cristofoletti R. Assessing the impact of cystic fibrosis on the antipyretic response of ibuprofen in children: Physiologically-based modeling as a candle in the dark. Br J Clin Pharmacol. 2020;86(11):2247–55.
Yellepeddi VK, Baker OJ. Predictive modeling of aspirin-triggered resolvin D1 pharmacokinetics for the study of Sjögren’s syndrome. Clin Exp Dent Res. 2020;6(2):225–35.
Willmann S, Thelen K, Kubitza D, Lensing AWA, Frede M, Coboeken K, et al. Pharmacokinetics of rivaroxaban in children using physiologically based and population pharmacokinetic modelling: an EINSTEIN-Jr phase I study. Thromb J. 2018;16:32.
Verscheijden LFM, van der Zanden TM, van Bussel LPM, et al. Chloroquine dosing recommendations for pediatric COVID-19 supported by modeling and simulation. Clin Pharmacol Ther. 2020;108(2):248–52.
Lutz JD, Mathias A, German P, Pikora C, Reddy S, Kirby BJ. Physiologically-based pharmacokinetic modeling of remdesivir and its metabolites to support dose selection for the treatment of pediatric patients with COVID-19. Clin Pharmacol Ther. 2021;109:1116–24. https://doi.org/10.1002/cpt.2176.
Park MH, Shin SH, Byeon JJ, Lee GH, Yu BY, Shin YG. Prediction of pharmacokinetics and drug-drug interaction potential using physiologically based pharmacokinetic (PBPK) modeling approach: a case study of caffeine and ciprofloxacin. Korean J Physiol Pharmacol. 2017;21(1):107–15.
Salerno SN, Edginton A, Gerhart JG, Laughon MM, Ambalavanan N, Sokol GM, et al. Physiologically-based pharmacokinetic modeling characterizes the CYP3A-mediated drug-drug interaction between fluconazole and sildenafil in infants. Clin Pharmacol Ther. 2020;109:253–62. https://doi.org/10.1002/cpt.1990.
Zane NR, Thakker DR. A physiologically based pharmacokinetic model for voriconazole disposition predicts intestinal first-pass metabolism in children. Clin Pharmacokinet. 2014;53(12):1171–82.
McPhail BT, Emoto C, Fukuda T, et al. Utilizing pediatric physiologically based pharmacokinetic models to examine factors that contribute to methadone pharmacokinetic variability in neonatal abstinence syndrome patients. J Clin Pharmacol. 2020;60(4):453–65.
Verscheijden LFM, Koenderink JB, de Wildt SN, Russel FGM. Development of a physiologically-based pharmacokinetic pediatric brain model for prediction of cerebrospinal fluid drug concentrations and the influence of meningitis. PLoS Comput Biol. 2019;15(6):e1007117.
Sun X, Ponte JF, Yoder NC, Laleau R, Coccia J, Lanieri L, et al. Effects of drug-antibody ratio on pharmacokinetics, biodistribution, efficacy, and tolerability of antibody-maytansinoid conjugates. Bioconjug Chem. 2017;28(5):1371–81.
Castelli MS, McGonigle P, Hornby PJ. The pharmacology and therapeutic applications of monoclonal antibodies. Pharmacol Res Perspect. 2019;7(6):e00535.
Malik P, Edginton A. Pediatric physiology in relation to the pharmacokinetics of monoclonal antibodies. Expert Opin Drug Metab Toxicol. 2018;14(6):585–99.
Hanke N, Kunz C, Thiemann M, Fricke H, Lehr T. Translational PBPK modeling of the protein therapeutic and CD95L inhibitor asunercept to develop dose recommendations for its first use in pediatric glioblastoma patients. pharmaceutics. 2019;11(4):152.
Malik PRV, Edginton AN. Physiologically-based pharmacokinetic modeling vs. allometric scaling for the prediction of infliximab pharmacokinetics in pediatric patients. CPT Pharmacometrics Syst Pharmacol. 2019;8(11):835–44.
Malik PRV, Edginton AN. Integration of ontogeny into a physiologically based pharmacokinetic model for monoclonal antibodies in premature infants. J Clin Pharmacol. 2020;60(4):466–76.
Wojtyniak JG, Britz H, Selzer D, Schwab M, Lehr T. Data digitizing: accurate and precise data extraction for quantitative systems pharmacology and physiologically-based pharmacokinetic modeling. CPT Pharmacometrics Syst Pharmacol. 2020;9(6):322–31.
Templeton IE, Jones NS, Musib L. Pediatric dose selection and utility of PBPK in determining dose. AAPS J. 2018;20(2):31.
Tucker GT. Personalized drug dosage - closing the loop. Pharm Res. 2017;34(8):1539–43.
Johnson TN, Abduljalil K, Nicolas JM, Muglia P, Chanteux H, Nicolai J, et al. Use of a physiologically based pharmacokinetic-pharmacodynamic model for initial dose prediction and escalation during a paediatric clinical trial. Br J Clin Pharmacol. 2020;87:1378–89. https://doi.org/10.1111/bcp.14528.
Bi Y, Liu J, Li L, Yu J, Bhattaram A, Bewernitz M, et al. Role of Model-informed drug development in pediatric drug development, regulatory evaluation, and labeling. J Clin Pharmacol. 2019;59(Suppl 1):S104–11. https://doi.org/10.1002/jcph.1478.
Acknowledgments
This work was supported by the Shenzhen Fundamental Research Program (JCYJ20180301170422030) from the Shenzhen Innovation of Science and Technology Committee and Science and Technology Key Project (2021CZ16) from the Guangdong Administration for Market Regulation. The funding agencies did not have any role in collection, analysis, interpretation of data, writing of the report, or the decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
K. W and J. K contribute to the conception of the work and draft the work. X. W and Y. L analyze the data for the work. T. W and Y. S approve the version to be published.
Corresponding authors
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, K., Jiang, K., Wei, X. et al. Physiologically Based Pharmacokinetic Models Are Effective Support for Pediatric Drug Development. AAPS PharmSciTech 22, 208 (2021). https://doi.org/10.1208/s12249-021-02076-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12249-021-02076-w