Abstract
The objective of nuclear forensic science is to link an event that involves a radioactive or nuclear material with the personal and material means that have facilitated it. This implies the collection and analysis of any physical evidence of the scene, both radioactive and nuclear material for its characterization as well as classical evidence like DNA, hair, fingerprints or blood. Collecting evidence in these circumstances can be potentially dangerous for the respondent due to the risk of radiation or radioactive contamination, so studying the stability of forensic evidence in the presence of radiation will allow taking a reasonable decision whether the probative utility that the evidence may have exceeds the dangers involved in its collection. In this context, this work addresses the resistance of classical forensic evidences to radiation. Thus, gamma post-irradiation results of DNA profiling from relevant biological samples are presented and discussed providing threshold values of radiation that, depending on the matrix, degrade DNA evidence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Nuclear forensic science has evolved in response to the growing illicit incidents related with radioactive or nuclear materials outside of regulatory control and its possible use in a malicious act. The ultimate goal of nuclear forensic discipline in an intentional incident involving these kinds of materials is the attribution. Therefore, a forensic study should be undertaken in a way that could answer the two main questions that are posed: the origin of the material and the identification of the human contribution in the event and the processes involved. The origin of the material is determined through its physical (e.g., morphology) and chemical characterization (trace level composition, isotopic ratios…), although clarifying the authorship requires the use of traditional forensic science disciplines such as DNA and fingerprints analysis [1].
Since its appearance, nuclear forensic science is focused fundamentally on the studies and advances related to the characterization of materials, leaving as secondary until recent years the evaluation of classical forensic evidences. The International Atomic Energy Agency reports [2, 3] that physical evidence like DNA, hair, fingerprints, tool marks, urine or blood, and explosive residue can link specific people to possible locations or events where nuclear activity may have occurred or radioactive material was released, so a forensic specialist must gather, test, and preserve this evidence. In the context of a terrorist scenario, the nature of the event itself causes on the potential probative samples being exposed to rugged environment effects that can impact on its successful analysis. So far, there have been relatively few investigations into the impacts and effects of radioactivity (mostly electromagnetic radiation) on DNA samples and the consequences that exposure may have for the use of this evidence for identification purposes [4,5,6,7,8].
Based on these reviews [5, 8], an experiment comprising the exposure to gamma radiation of relevant biological matrices at accumulated doses in the range of 10–1000 kGy and subsequent analysis of DNA extracted from irradiated samples was carried out [9]. The range of absorbed dose considered is based on the scenarios related to CBRN attack with radionuclide sources where those are removed from the shield and manipulated (in a workshop and or storage in a place for malicious use). The aforementioned papers demonstrate that dose values lower than 10 kGy have not any effect on DNA samples. Moreover, it is important to remain that the investigation in case of CBRN attack takes a large time, due to the first step is minimize the effects, protect and save the population and contain the radionuclide released, reason for which when the first responders begin to take the evidences, the dose absorbed by material surrounding the CBRN device will be in the range dose selected in the paper. In this scenario, the purpose of this paper is to assess the impact of sample degradation on forensic genotyping and well as postulated dose threshold above which DNA degradation begins. Furthermore, in this paper, it presented the method used for identification, in which the DNA results are input in the database for searching the personal profile and when, e.g., the dose amount, it is impossible to coupling with the person [10].
2 Experimental procedure
2.1 Samples
The DNA studied came from blood, skin and saliva, the most frequent evidence in the scene of a crime. Once deposited on swabs (blood), keys (epithelial cells) and plastic bottle (saliva), the samples were, in this order, placed in a laminar flow hood until dry, packaged in zip lock plastic bags and stored at room temperature. All samples proceeded from a single individual and contained enough DNA to carry out unequivocal identification of the donor.
Prepared samples were irradiated at the CIEMAT 60Co facility (NAYADE), a pool-type radioactive unit composed by 72 spatially configurable sources (current activity 423 TBq) which allows controlling both the total absorbed dose and the homogeneity of dose rate applied. In summary, samples were irradiated at 0, 10, 50, 100, 500, 1000 kGy with a dose rate of 20.314 kGy h−1. Once irradiated, they were delivered to the crime laboratory of the Civil Guard for analysis.
Throughout the process, a chain of custody similar to that implemented in the case of real samples is followed, documenting the tasks and people involved in each stage.
2.2 DNA analysis
The analyses were performed according to the DNA standard procedure [11] of crime lab (Civil Guard) that includes the following steps:
-
Extraction and purification from the biological matrices using the Automate Express™ platform (Thermo Fisher Scientific), a solid-phase extraction system based on the use of paramagnetic silica beads that simultaneously extract and purify the DNA in a single stage.
-
DNA quantification and amplification by means of real-time PCR (fluorescence reporter probe method) through the 7500 Real-Time PCR System and QuantifilerTM kits, both of Applied Biosystems.
-
Separation and detection of amplified DNA fragments with capillary electrophoresis (CE). The CE separates DNA fragments by size (molecular weight) detecting them by excitation of the fluorescent labels incorporated in the amplification step.
-
Profile interpretation of the 24 short tandems repeats (STRs) or markers (loci) for every sample. These included those collected in the European Standard Set (ESS) and in the Combined DNA Index System (CODIS) from FBI adding another five markers and takes place in three stages: raw data collection, length approximation, and allelic assignment.
The last step was conducted using GlobalFilerTM PCR Amplification Kit and GeneMapper ID-XTM software (Applied Biosystems).
3 Results and discussion
3.1 DNA degradation with radiation
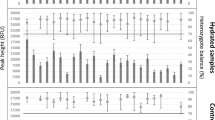
Results obtained for each sample at every absorbed dose were compared to the DNA reference profiling of the donor, that is the profiles obtained from unirradiated blood, skin and saliva samples. From the data obtained in the analysis of the reference profile, the points that will be used to determine alterations due to radiation are established. Thus, Fig. 1 the DNA profile of an unirradiated sample (reference profile) appears on the top and that of the same type of sample after irradiation (500 kGy) below.
In each electropherogram, DNA fragments or markers are represented along the x-axis of each graph in order from smallest to largest.
The pattern of DNA profiling obtained for blood, epithelial cells and saliva in function of radiation dose showed a progressive decrease in the intensity of the signals of the different markers as the radiation dose increased (from 0 to 1000 kGy). This effect is more pronounced for the higher molecular weight markers which at high doses even disappear. This behaviour seems to indicate that the degradation of DNA by the effect of irradiation is mainly due to fragmentation of the DNA chain which will be more likely the longer the length of the fragment.
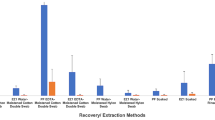
Normalising the intensity of peak measured of each STR to its reference value (non-irradiated sample) as a function of the absorbed dose and for three biological matrices studied (Figs. 2, 3, 4). These plots are presented the data obtained as a function of the dose absorbed (10 kGy, 50 kGy, 100 kGy, 500 kGy and 1 MGy, plotted as square, sphere, up-triangle, diamond and down triangle, respectively) and the lines represents the tendency of decrease the intensity for each dose absorbed as a function of the STR base pair.
Data obtained at least of 3 replicates for matrix are summarised in Figs. 2, 3, 4. These plots show that due to radiolysis effect a decreasing in the STR intensity value as a function of the dose absorbed is detected. This effect is better observed in saliva (Fig. 5) and epithelial cells (Fig. 4). In this last serial of samples, an effect of the amount of sample is observed, i.e., in case of samples taken from caps the intensity measured is higher than from keys due to the amount of DNA in each surface. Although the absorbed those effect on the intensity is exactly the same. Furthermore, from these plots it can also be concluded that the loss of these peaks occurs at lower doses in DNA extracted from epithelial cells (Fig. 4) and saliva (Fig. 5) than for DNA from blood (Fig. 6) as can be seen where the evolution of 5 DNA markers for the three matrices is shown.
Although according to these findings (Figs. 2, 3, 4), DNA of biological matrices is relatively resistant to gamma irradiation until 50 kGy, at higher doses significant differences appear that allows us to establish a threshold for each one. Thus:
-
At 100 kGy, DNA of saliva and blood samples preserves almost all markers, while DNA of epithelial samples has suffered further degradation (Figs. 2, 3, 4).
-
At 500 kGy, DNA of blood preserves almost all markers (Figs. 2, 3, 4).
As consequence, the dose threshold that commits the validity of the samples as forensic evidence depends on the type of matrix.
3.2 Validity for identification
Considering the quality of the genetic profiles obtained, they have been classified into three groups:
-
Complete Genetic Profile: when results are obtained for all markers and they are correct.
-
Partial Genetic Profile: when no results are obtained for all the studied markers or an incorrect marker appears, provided that the profile has a minimum of 10 correct markers (9 markers + the marker for determining sex, amelogenin).
-
No Profile: when no results are obtained or less than 10 correct markers are obtained.
In Figs. 5, 6 and 7 are represented as color map the results obtained in the irradiated samples. As can be observed in case of irradiated DNA samples from epithelial cells and saliva (Figs. 6, 7), it is possible to recognize the individual (following the standard method Fig. 8) until absorbed dose level of 100 kGy, whereas in case of those samples taken from blood the threshold value is 500 kGy. Those samples that were irradiated until absorbed dose higher than these values (aforementioned) cannot give any information which can help to recognize any person.
4 Conclusions
The resistance to radiation of DNA evidences has a strength dependence of the initial concentration in the samples taken. The radiolysis effect shows a homogenous behaviour with independence of the alleles considered.
-
DNA degradation induced by irradiation did not lead to the appearance of additional peaks in the profiles.
As consequence, the dose threshold that commits the validity of the samples as forensic evidence depends on the type of matrix.
-
According to the obtained results, these differences can be significant and allows to establish a threshold for each one. Thus, at:
-
50 kGy DNA of epithelial cells, saliva and blood samples preserve almost all markers.
-
100 kGy, DNA of saliva and blood samples preserves almost all markers. Epithelial samples taken from keys remain enough STR for identification.
-
500 kGy DNA of blood samples preserves almost all markers.
-
The results obtained will be applied in a CBRN scenario wherein will be necessary to recollect evidences in order to open a criminal persecution both ALARA point of view and accuracy in the data.
References
L. Wilson-Wilde, The international development of forensic science standards—a review. Forensic Sci. Int. (2018). https://doi.org/10.1016/j.forsciint.2018.04.009
IAEA, Radiological Crime Scene Management, 2014.
IAEA, Risk informed approach for nuclear security measures for nuclear and other radioactive material out of regulatory control. Nucl. Secur. Ser. 2015.
M. Colella, S. Thomson, S. Macintosh, M. Logan, An introduction to radiological terrorism. Aust. J. Emerg. Manag. 20(2), 9–17 (2005)
S. F. Abbondante, The effect of radioactive materials on forensic DNA evidence : procedures and interpretation. University of Camberra, 2009.
C. S. Goodwin, In vitro repair of gamma-irradiated DNA for forensic analysis. University of Camberra, 2013.
R. Hoile, C. Banos, M. Colella, S.J. Walsh, C. Roux, Gamma irradiation as a biological decontaminant and its effect on common fingermark detection techniques and DNA profiling. J. Forensic Sci. (2010). https://doi.org/10.1111/j.1556-4029.2009.01233.x
K.L. Monson et al., Potential effects of ionizing radiation on the evidentiary value of DNA, latent fingerprints, hair, and fibers: a comprehensive review and new results. Forensic Sci. Int. (2018). https://doi.org/10.1016/j.forsciint.2018.01.012
J. Serrano Checa, J. Quiñones, Desarrollo de las técnicas forenses nucleares: valoración de riesgos. Universidad Complutense de Madrid, 2019.
J.S. Buckleton, J.A. Bright, D. Taylor, Forensic DNA Evidence Interpretation (CRC Press, Boca Raton, 2018).
J.M.J.M. Butler, Forensic DNA Typing: Biology, Technology, and Genetics of STR Markers (Elsevier, Amsterdam, 2005).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Quinones Diez, J., Fernández, M., Mingorance, E. et al. DNA resistance to radiation field: forensic genotyping in a radiological incident scenario. Eur. Phys. J. Plus 136, 413 (2021). https://doi.org/10.1140/epjp/s13360-021-01407-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjp/s13360-021-01407-x