Abstract
Physical inactivity and disuse lead to a decrease in the functionality of skeletal muscles (oxidative capacity, insulin sensitivity, and performance), which is associated with a change in mitochondrial density. In contrast, aerobic exercise training is effective for maintaining/increasing skeletal muscle mitochondrial density and functionality. The review considers the effect of increasing and decreasing physical activity on the mitochondrial density of human skeletal muscles, as well as the main mechanisms responsible for these changes. It is discussed that the content of mitochondrial proteins can be regulated by changing the content of their mRNAs, changes in the rate of synthesis specific for mitochondrial proteins, as well as changes in the rate of degradation, transport, import, and stability of mitochondrial proteins. It has been shown that the mechanisms of regulation of the mitochondrial proteins content under various interventions are significantly different. At the same time, their contribution to the change in the content of mitochondrial proteins is characterized clearly insufficiently, which emphasizes the relevance of further research in this area.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Skeletal muscles make up more than a third of the body mass and, with a normal level of physical activity, are characterized by a high level of metabolic activity, which positively affects the functioning of other tissues, organs, and the organism. Physical inactivity and a sharp decrease in physical activity while on bed rest (disuse) have a complex effect on various systems and organs of the organism, including skeletal muscles. This is expressed in a decrease in the functional capacity of skeletal muscles (the maximum rate of fat and carbohydrate oxidation, insulin sensitivity, aerobic performance, and muscle strength), which is largely due to a decrease in mitochondrial density and muscle mass. These changes may be one of the causes of chronic diseases, such as obesity and type 2 diabetes, cardiovascular diseases, depression and chronic fatigue syndrome, as well as a decrease in lifespan [1–4]. In contrast, regular exercise, particularly aerobic exercise, is effective in maintaining/increasing skeletal muscle mitochondrial density, oxidative capacity, insulin sensitivity, and aerobic performance. The review briefly characterizes the most striking changes in the functional capacity and phenotype of human skeletal muscles that occur with regular aerobic training and with a decrease in the level of physical activity. The effect of increase and decrease in physical activity on mitochondrial density, which changes markedly and in a multidirectional manner under these interventions, as well as the main mechanisms responsible for these changes, is considered in detail.
Influence of Aerobic Exercise Training on the Functions and Phenotype of Skeletal Muscles
To study the effects of regular aerobic physical exercise or increasing the level of physical activity, bicycle exercise and treadmill training, as well as long-distance walking, is most often used [5].
The use of physical exercise with different intensity and duration allows to recruit different muscle fibers. As an example, low- and moderate-intensity aerobic exercise predominantly involves type I fibers that are resistant to fatigue and have high oxidative capacity. An increase in exercise intensity to submaximal and maximum aerobic power leads to additional involvement in the work of type II muscle fibers, which have low oxidative and high glycolytic potentials in an untrained muscle.
Regular aerobic exercise markedly increases the density and function of mitochondria in skeletal muscle, as well as the ability of muscles to oxidize fats and carbohydrates. This relates with an increase in the concentration of numerous proteins involved in the transport and β-oxidation of lipids, the tricarboxylic acid cycle, and oxidative phosphorylation, but there is no pronounced effect on the content of glycolytic enzymes [1, 6, 7]. In addition, regular exercise leads to an increase in capillary density [8], which plays an important role in increasing the capacity of skeletal muscles for oxygen diffusion. In addition, the skeletal muscles adapted to regular aerobic exercise show an increase in the glycogen and triglyceride stores. The latter are actively oxidized by trained skeletal muscles at rest and during physical activity of low and moderate intensity, while carbohydrates are the dominant substrate during high-intensity aerobic exercise [9].
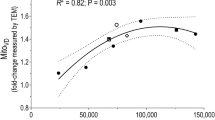
During the first weeks of regular aerobic exercise, mitochondrial density (as well as respiratory enzyme activity and mitochondrial respiratory rate) rapidly increases. Thus, the volume density of mitochondria increases by 40% as soon as after 6 weeks of regular aerobic training [10]. It is important to note that regular aerobic exercise, unlike strength exercise, does not lead to a pronounced change in the size of muscle fibers and trained skeletal muscles, as well as maximum voluntary contraction [11].
The Effect of Disuse on the Function and Phenotype of Skeletal Muscles
To study the effects of a chronic decrease in physical activity (disuse), immobilization of one limb, bed rest, and “dry” immersion are most often used. These models make it possible to significantly reduce the contractile activity and mechanical load on the studied skeletal muscles, as well as to significantly reduce the support afferentation in the case of “dry” immersion [12, 13].
Numerous studies have convincingly shown that a decrease in physical activity in human muscles is accompanied by a decrease in the density of mitochondria and the maximum rate of their respiration, oxidation of fats and carbohydrates, as well as a decrease in aerobic performance [14, 15]. As an example, after 5 to 6 weeks of disuse, the activity of various mitochondrial enzymes, a mitochondrial density marker, decreases by 20–40% [16]. Interestingly, a similar decrease in the concentration of several enzyme of the Krebs cycle and oxidative phosphorylation was observed in the vastus lateralis muscle after 8 and 35 days of bed rest [17]. Apparently, with a decrease in physical activity, the oxidative capacity of skeletal muscles decreases rapidly during the first week and then changes relatively slowly.
The density of muscle capillaries is one of the key indicators affecting the diffusion of oxygen to muscle mitochondria. A number of studies have not revealed a decrease in the number of capillaries per fiber [18–20] and capillary density [18–21] in the vastus lateralis and/or soleus muscles in humans after several weeks of disuse, while several others showed a decrease in these indices [22, 23]. Apparently, the absence of a decrease in capillarization is associated with a decrease in the size of muscle fibers. Indeed, it has been repeatedly shown that bed rest and immobilization of one limb for several days/weeks cause a decrease in muscle volume (at a rate of ~0.4% per day) and strength [14, 24]. In human skeletal muscle with a mixed fiber type composition (for example, in the vastus lateralis), the reduction in cross-sectional area is comparable to type I and type II muscle fibers, while a decrease in the size of these fibers is mainly observed in the soleus muscle with a predominant content of slow-twitch type I muscle fibers [14, 25–28]; it has also been shown that the decrease in the mass of the leg extensor muscles is more pronounced than that of the flexor muscles [13]. These data suggest that the mechanisms regulating the decline in skeletal muscle phenotype in different muscles may slightly differ.
Regulation of Mitochondrial Biogenesis in Skeletal Muscles
Regular aerobic training and disuse produce a pronounced and multidirectional effect on the content of mitochondrial proteins in skeletal muscles. Theoretically, these changes can be regulated by changing the content of mRNA encoding mitochondrial proteins, changing the rate of mitochondrial protein synthesis (translation), and also by changing the rate of degradation of mitochondrial proteins (proteolysis).
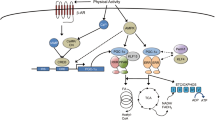
mRNA content. Currently, the activation of mitochondrial biogenesis after aerobic exercise is largely associated with regulation at the transcriptional level [29]. During aerobic exercise, significant metabolic changes occur in the muscle cell: in particular, the AMP/ATP ratio and the content of Ca2+ and reactive oxygen species increase. It was shown in various models (myoblast culture, rodent and human skeletal muscles) that this leads to activation of various signaling cascades, which, in turn, changes the activity of transcriptional regulators (coactivators, corepressors, and transcription factors) that control angiogenesis, carbohydrate and fat metabolism, and mitochondrial biogenesis [30–32]. Among the best studied transcriptional regulators of mitochondrial biogenesis are peroxisome proliferator-activated receptor γ coactivator 1α (PGC-1α) [33], estrogen-related receptor γ (ESRRG) [34], mitochondrial transcription factor A (TFAM) [35], nuclear receptor 4A3 (NR4A3) (nuclear receptor subfamily 4, group A, member 3) [36], cAMP-responsive element-binding protein (CREB)-regulated transcriptional coactivator 2 (CRTC2) [37], and nuclear receptor corepressor 1 (NCOR1) [38]. These transcriptional regulators can control mitochondrial biogenesis by increasing the expression of genes encoding other regulators of mitochondrial biogenesis and mitochondrial proteins.
In confirmation of this concept, a simultaneous increase in the content of mRNA encoding some mitochondrial enzymes (cytochrome b and c, cytochrome oxidase) and mitochondrial density markers (mitochondrial DNA content, activity of citrate synthase, cytochrome oxidase, etc.) has already been shown in early studies with chronic (1–3 weeks) low-frequency stimulation of animal muscles or with running training [39–42]. However, in other studies with electrical stimulation of the muscle, it was not always possible to detect a simultaneous increase in the content of cytochrome oxidase and cytochrome c mRNA with an increase in the activity and content of these proteins [43, 44].
It was noted previously that even several weeks of aerobic training led to an increase in mitochondrial density and the content of mitochondrial proteins in human skeletal muscles. A study evaluating the expression of a hundred genes in the vastus lateralis muscle after 12 weeks of aerobic bicycle exercise showed an increase in the content of about 40 mRNA encoding enzymes of the tricarboxylic acid cycle and the electron transport chain [45]. It is important to note that mitochondria are composed of more than 1000 proteins; moreover, about 150 of them belong to enzymes of the Krebs cycle and oxidative phosphorylation. Therefore, of great interest are studies that correlate changes in the transcriptome (in particular, the mRNA content of mitochondrial proteins) induced by regular aerobic training with changes in the content of many individual mitochondrial proteins. In three such studies, the effects of various 6–12-week aerobic training on the transcriptome and proteome of the vastus lateralis were investigated in young and old humans [7, 46, 47]. Mass spectrometry-based proteomics made it possible to detect about two hundred mitochondrial proteins (mainly mitochondrial enzyme proteins) most of which increased in content. At the same time, an increase in the expression of several hundred genes was found, but only about 20 of them encoded mitochondrial proteins. This means that in the basal state (i.e., 2 or 3 days after the last exercise), an increase in the content of mitochondrial proteins in human skeletal muscle induced by regular aerobic training does not correlate with an increase in the content of corresponding mRNAs. The absence of a massive increase in the mRNA content of mitochondrial proteins in human skeletal muscle after several weeks of regular aerobic training is confirmed by most (but not all [48–50]) transcriptomic studies, which showed, using functional enrichment analysis, that gene sets with increased expression were not enriched in mitochondrial protein genes [46, 47, 51–57]. This conclusion is confirmed by the results of a meta-analysis, which analyzed and summarized the original transcriptomic data of these studies [58]. Are there other examples of activation of mitochondrial biogenesis (an increase in the content of mitochondrial proteins) that occurs without an increase in the mRNA content of mitochondrial proteins? Indeed, using transcriptomic and proteomic analysis, it was shown that an increase in the content of many mitochondrial proteins is observed in human CD34 hematopoietic stem cells during their differentiation into proerythroblasts [59] and in HeLa cells under endoplasmic reticulum stress [60] and occurs without a massive increase in the expression of genes encoding these proteins.
It may be suggested that the lack of a massive increase in the mRNA content of mitochondrial proteins after a period of regular aerobic training can be explained by a short-term increase in the expression of these genes after each exercise. A number of studies have investigated transcriptome changes after a single aerobic exercise. In most studies, it was shown that a single aerobic exercise does not induce in vastus lateralis of healthy people a massive (several tens or more) increase in the expression of genes encoding mitochondrial proteins: immediately after [61, 62], after 1 h [56, 63], 2.5 h [64], 3 h [65 , 66], 4 h [56, 63, 67], 5 h [64], 8 h [63], 48 h [65, 66] and 96 h [66]. This is confirmed by a meta-analysis that studied transcriptome changes immediately after and 1, and 4 hours after an aerobic exercise [58]. It cannot be ruled out that a massive increase in mRNA expression of mitochondrial proteins may occur at other times after aerobic exercise, for example, after 10–20 hours. This supported by the fact that during the first hours of recovery, changes in the transcriptome after an aerobic exercise are mainly associated with an increase in the expression of several dozen genes that regulate transcription [56, 58], including a number of well-studied regulators of mitochondrial biogenesis, angiogenesis, carbohydrate and fat metabolism (PPARGC1A, ESRRG, TFAM, NR4A3, etc.).
For normal mitochondria function, it is necessary to maintain the stoichiometry of the proteins included in the respiratory complexes [68, 69]. Changes in the content of individual proteins in respiratory complexes can lead to an increase in the production of reactive oxygen species [70, 71]. If the increase in the content of mitochondrial proteins during regular aerobic training is regulated by an increase in the content of the corresponding mRNAs, then this should lead to an increase in the content of all mRNAs encoding proteins of the respiratory complexes. However, it was previously shown that acute aerobic exercise increases the content of only some mRNAs encoding mitochondrial enzymes. Few studies directly comparing changes in the content of many mitochondrial proteins and their mRNAs in human skeletal muscle [7, 46, 47] suggest that an increase in the content of mitochondrial proteins during aerobic training is regulated not only (and not so much) at the level of transcription, but by other mechanisms.
What happens to the mRNA content of mitochondrial proteins in human skeletal muscle during disuse? Regardless of the experimental model chosen, all studies found that the sets of genes that decreased expression after inactivity and disuse lasting from 2 to 90 days are enriched in several tens or even hundreds of genes encoding mitochondrial proteins (mainly respiratory complex proteins) [72–81]. The same result was shown in meta-analyses that studied the effects of several weeks of disuse [50, 58]. This allows us to conclude that the decrease in the content of mitochondrial proteins in human skeletal muscle induced by inactivity and disuse is associated with a massive decrease in the content of mRNA encoding mitochondrial proteins (a hundred or more genes, which is more than 10% of the total number of mitochondrial proteins). Interestingly, a similar result was obtained in vastus lateralis of old individuals with sarcopenia (characterized by reduced mitochondrial density) compared with age-related controls [82]. This suggests that the decrease in the concentration of some mitochondrial proteins (in particular, some enzymes of the Krebs cycle and oxidative phosphorylation) is regulated at the level of mRNA content, while the content of other mitochondrial proteins is regulated by other mechanisms.
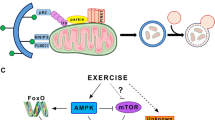
Translation of mitochondrial proteins. During the first hours after a single aerobic exercise, an increase in the rate of synthesis of mitochondrial proteins in vastus lateralis is observed in untrained and trained individuals; at the same time, the rate of synthesis of myofibrillar proteins does not change or increases less [83, 84]. Moreover, 12-week high-intensity interval training increases the rate of mitochondrial protein synthesis in vastus lateralis in the basal state (72 h after the last exercise) in both young and elderly volunteers [46]. It is important to note that there is no massive increase in the mRNA content of mitochondrial proteins in vastus lateralis at these time points. Based on this, it may be suggested that, after aerobic exercise, the regulation of translation plays an important role in increasing the content of mitochondrial proteins. If the rate of myofibrillar proteins synthesis does not change after an aerobic exercise, but mitochondrial protein synthesis increases, then the question arises how the translation of mRNAs encoding mitochondrial proteins can be regulated.
The mTORC1 complex is the main translation regulator that controls protein synthesis in the cell by regulating the binding of translation initiation factors to the 40S ribosomal subunit and to the cap at the 5' mRNA end (the canonical cap-dependent regulation) [85]. It was previously noted that during the differentiation of hematopoietic stem cells, an increase in the content of some (including mitochondrial) proteins occurs without an increase in the content of the corresponding mRNAs. It turned out that a distinctive feature of these mRNAs is the presence of motifs with a 5' terminal oligopyrimidine (TOP- and TOP-like motifs) in the 5' untranslated region (UTR) [59]. In another cell study, it was shown that the translation of most mRNAs is suppressed under stress induced by a decrease in glucose content, while the translation of mRNAs encodind proteins that regulate mitochondrial energy metabolism does not stop [86]. It appeared that these mRNAs contain translation initiator motifs of short 5' UTR (TISU- and TISU-like motifs). Translation of mRNAs containing TOP or TISU motifs is cap-dependent [86–88] but is regulated in a non-canonical way: these mRNAs have a short 5' UTR (an average of 12 nucleotides), which allows the cap-associated 43S complex to directly interact with the AUG start codon.
Under conditions of energy deficiency, the cell reduces energy consumption by suppressing the energy-intensive process of protein synthesis. The main role of the mechanism controlling mRNA specific regulation of translation (including the mRNAs encoding mitochondrial proteins) is to maintain the synthesis of proteins necessary for cell survival under changed environmental conditions. As an example, mRNAs with TISU motifs were found in only 4% of protein-coding genes, in particular, in genes that regulate protein synthesis, degradation and folding, mRNA metabolism, and mitochondrial biogenesis [89]. It may be suggested that the mechanisms described above may play a role in increasing the rate of mitochondrial protein synthesis in human skeletal muscle after an aerobic exercise. In addition, it should be noted that cap-independent translation initiation mechanisms exist, which allow maintaining the rate of translation of some mRNAs encoding proteins important for cell survival. These mechanisms associated with the structures that form internal ribosome entry sites (IRES) in the 5' UTR mRNA and N6-methyladenosine modifications (m6A) in the 5' and 3' UTR mRNA [85]. We have not found evidence that translation of mitochondrial proteins can be regulated in this way, but the potential role of these mechanisms in the regulation of mitochondrial biogenesis cannot be ruled out.
Disuse leads to a decrease in the content of mitochondrial proteins, which occurs along with muscle mass decrease. In human skeletal muscle with a mixed fiber type compositions (for example, in vastus lateralis), muscle mass decrease is closely associated with a decrease in the rate of protein synthesis [90, 91]. It may be suggested that during disuse, the decrease in the content of mitochondrial proteins is at least partially due to the suppression of translation, which occurs simultaneously with a decrease in the content of mRNA encoding some mitochondrial and ribosomal proteins [80]. At the same time, it remains unclear whether the translation of mRNA encoding mitochondrial proteins is regulated by the noncanonical cap-dependent and cap-independent mechanisms.
Degradation of mitochondrial proteins. The ubiquitin-proteasome system plays an important role in the regulation of the mitochondrial biogenesis and their functions by ubiquitinating maturing mitochondrial proteins in the cytoplasm, as well as proteins in various mitochondrial compartments [92, 93]. The autophagal–lysosomal system is involved in the degradation of large protein complexes, including damaged mitochondria [94]. Actively functioning mitochondria can accumulate damaged proteins; therefore, it is logical to assume that this should cause the activation of proteolytic systems that eliminate such proteins thereby maintaining proteostasis.
Indeed, an intensive aerobic exercise causes short-term dephosphorylation of transcription factor FOXO1Ser256 (an activation marker) and an increase in the expression of its target genes, muscle-specific E3 ubiquitin ligases TRIM63 (MURF1) and FBXO32 (MAFbx) in human skeletal muscle [67, 95–98], as well as an increase in phosphorylation of the ULK1Ser555 kinase (an activation marker) without altering ULK1Ser757 (an inhibition marker), which indicates the activation of autophagy [99]. Studies on rodents have shown that regular aerobic training increases the content of autophagy-regulating proteins (LC3-II, Beclin-1, ATG7, LAMP2a) and the activity of the 26S proteasome [100, 101]. On the other hand, after low-intensity aerobic exercise, there is no activation of markers of the ubiquitin-proteasome system and autophagy [67, 102, 103]. Apparently, the activation of proteolytic systems occurs mainly after intensive and/or prolonged physical activity, which causes the accumulation of damaged proteins in the muscle. At the same time, it has been repeatedly shown that the regular exercise training is a powerful stimulus for the activation of mitochondrial biogenesis and an increase in mitochondrial density. This suggests that short-term activation of proteolytic systems after each exercise does not limit the increase in the content of mitochondrial proteins during exercise training. On the other hand, mitochondrial proteins, compared to other muscle proteins, contain fewer degradation-regulating motifs, which correlates with their longer half-life [7]. This indirectly indicates that degradation does not play a key role in the regulation of the content of mitochondrial proteins, in particular, during regular aerobic training.
Heat shock proteins (chaperones) play an important role in the intracellular transport of proteins, in the formation and maintenance of the structure of proteins, and in the restoration of the structure of damaged proteins, thereby reducing the likelihood of their degradation [104, 105]. It turned out that overexpression of the 72 kDa heat shock protein (Hsp72) (a protein of the chaperone 70 family with mitochondrial localization) in murine skeletal muscles increases mitochondrial content, the time and maximum running speed of mice. In addition, Hsp72 overexpression increases the mitochondrial content in the heart of rodents [106, 107]. The influence of chaperones on the regulation of mitochondrial functions in mouse skeletal muscles is also confirmed by studies with modulation of the expression of the transcriptional factor Hsf1, which regulates the expression of various chaperones [108]. Interestingly, in human skeletal muscle, regular aerobic training leads to a simultaneous increase in the content of many mitochondrial proteins and two tens of chaperones and chaperone-associated proteins [6, 7], which is consistent with the results of the studies described previously. However, the presented studies did not give an unambiguous answer about the mechanisms of chaperone-dependent regulation of mitochondrial biogenesis. These mechanisms can be related with an increase in the transport and import of mitochondrial proteins from the cytoplasm (more than 99% of mitochondrial proteins are encoded by genomic DNA and synthesized in the cytoplasm) [106, 107, 109], as well as by an increase in the stability of mitochondrial proteins [7].
During disuse, a decrease in the content of contractile proteins and muscle mass in the mixed vastus lateralis is predominantly associated with a decrease in the rate of protein synthesis rather than with an increase in the rate of their degradation [90, 91]. However, a number of studies on models [110] and human muscle showed the activation of some elements of the ubiquitin–proteasome and autophage–lysosomal systems [72, 75, 77, 79, 80, 111–114]. This, as well as the chaperone-dependent decrease in the import of mitochondrial proteins from the cytoplasm [115], may be responsible for the decrease in the skeletal muscle content of mitochondrial proteins induced by disuse.
CONCLUSIONS
The use of high-throughput methods allowed to characterize changes in the content of almost all mRNAs encoding mitochondrial proteins and several hundred mitochondrial proteins induced by increase and decrease in physical activity in human skeletal muscle. It has been shown that the mechanisms of regulation of the content of mitochondrial proteins under such exposures are significantly different.
The results of most studies have shown that an increase in the content of various mitochondrial proteins induced by regular aerobic training is not associated with a massive increase in the content of the corresponding mRNAs either in the basal state (2 to 3 days after the last exercise) or during the first hours of recovery after a single aerobic exercise. It is important to note that in this case, there is a predominant increase in the rate of synthesis of mitochondrial rather than myofibrillar proteins. One of the most probable mechanism responsible for such changes is noncanonical cap-dependent and independent regulation of mRNA translation of mitochondrial proteins. Another potential mechanism for increasing the content of mitochondrial proteins during regular exercise may be a chaperone-dependent increase in their transport, import, and stability. Disuse-induced decrease in the content of mitochondrial proteins, at least in part, is associated with a decrease in the content of the corresponding mRNAs. In parallel with this, there is a decrease in the overall rate of muscle protein synthesis and import of mitochondrial proteins, as well as activation of some elements of various proteolytic systems. However, the contribution of these processes to the decrease in the content of mitochondrial proteins has not been fully characterized. In this context, it seems promising to directly assess the effect of increased and decreased levels of physical activity on the rate of translation and degradation of individual proteins using mass spectrometric analysis of proteins labeled with deuterium in vivo [116], as well as ribosomal profiling.
REFERENCES
Lanza, I.R., Short, D.K., Short, K.R., et al., Endurance exercise as a countermeasure for aging, Diabetes, 2008, vol. 57, no. 11, p. 2933.
Pedersen, B.K. and Febbraio, M.A., Muscles, exercise and obesity: skeletal muscle as a secretory organ, Nat. Rev. Endocrinol., 2012, vol. 8, no. 8, p. 457.
Demontis, F., Piccirillo, R., Goldberg, A.L., and Perrimon, N., The influence of skeletal muscle on systemic aging and lifespan, Aging Cell, 2013, vol. 12, no. 6, p. 943.
Agudelo, L.Z., Femenía, T., Orhan, F., et al., Skeletal muscle PGC-1α1 modulates kynurenine metabolism and mediates resilience to stress-induced depression, Cell, 2014, vol. 159, no. 1, p. 33.
Waller, L., Krüger, K., Conrad, K., et al., Effects of different types of exercise training on pulmonary arterial hypertension: a systematic review, J. Clin. Med., 2020, vol. 9, no. 6, p. 1689.
Schild, M., Ruhs, A., Beiter, T., et al., Basal and exercise induced label-free quantitative protein profiling of m. vastus lateralis in trained and untrained individuals, J. Proteomics, 2015, vol. 122, p. 119.
Makhnovskii, P.A., Zgoda, V.G., Bokov, R.O., et al., Regulation of proteins in human skeletal muscle: the role of transcription, Sci. Rep., 2020, vol. 10, no. 1, p. 3514.
Saltin, B., Henriksson, J., Nygaard, E., et al., Fiber types and metabolic potentials of skeletal muscles in sedentary man and endurance runners, Ann. N.Y. Acad. Sci., 1977, vol. 301, p. 3.
San-Millán, I. and Brooks, G.A., Assessment of metabolic flexibility by means of measuring blood lactate, fat, and carbohydrate oxidation responses to exercise in professional endurance athletes and less-fit individuals, Sports Med., 2018, vol. 48, no. 2, p. 467.
Montero, D., Cathomen, A., Jacobs, R.A., et al., Haematological rather than skeletal muscle adaptations contribute to the increase in peak oxygen uptake induced by moderate endurance training, J. Physiol., 2015, vol. 593, no. 20, p. 4677.
Hawley, J.A., Hargreaves, M., Joyner, M.J., and Zierath, J.R., Integrative biology of exercise, Cell, 2014, vol. 159, no. 4, p. 738.
Tomilovskaya, E., Shigueva, T., Sayenko, D., et al., Dry immersion as a ground-based model of microgravity physiological effects, Front. Physiol., 2019, vol. 10, p. 284.
Sharlo, K., Tyganov, S.A., Tomilovskaya, E., et al., Effects of various muscle disuse states and countermeasures on muscle molecular signaling, Int. J. Mol. Sci., 2022, vol. 23, no. 1, p. 468.
Hackney, K.J. and Ploutz-Snyder, L.L., Unilateral lower limb suspension: integrative physiological knowledge from the past 20 years (1991–2011), Eur. J. Appl. Physiol., 2012, vol. 112, no. 1, p. 9.
Hyatt, H., Deminice, R., Yoshihara, T., and Powers, S.K., Mitochondrial dysfunction induces muscle atrophy during prolonged inactivity: a review of the causes and effects, Arch. Biochem. Biophys., 2019, vol. 662, p. 49.
Gram, M., Dahl, R., and Dela, F., Physical inactivity and muscle oxidative capacity in humans, Eur. J. Sport Sci., 2013, vol. 14, no. 4, p. 376.
Brocca, L., Cannavino, J., Coletto, L., et al., The time course of the adaptations of human muscle proteome to bed rest and the underlying mechanisms, J. Physiol., 2012, vol. 590, no. 20, p. 5211.
Hather, B.M., Adams, G.R., Tesch, P.A., and Dudley, G.A., Skeletal muscle responses to lower limb suspension in humans, J. Appl. Physiol., 1992, vol. 72, no. 4, p. 1493.
Berg, H.E., Dudley, G.A., Hather, B., and Tesch, P.A., Work capacity and metabolic and morphologic characteristics of the human quadriceps muscle in response to unloading, Clin. Physiol., 1993, vol. 13, no. 4, p. 337.
Salanova, M., Schiffl, G., Püttmann, B., et al., Molecular biomarkers monitoring human skeletal muscle fibers and microvasculature following long-term bed rest with and without countermeasures, J. Anat., 2008, vol. 212, no. 3, p. 306.
Vigelsø, A., Gram, M., Wiuff, C., et al., Six weeks’ aerobic retraining after two weeks’ immobilization restores leg lean mass and aerobic capacity but does not fully rehabilitate leg strength in young and older men, J. Rehabil. Med., 2015, vol. 47, no. 6, p. 552.
Rudnick, J., Püttmann, B., Tesch, P.A., et al., Differential expression of nitric oxide synthases (NOS 1–3) in human skeletal muscle following exercise countermeasure during 12 weeks of bed rest, FASEB J., 2004, vol. 18, no. 11, p. 1228.
Arentson-Lantz, E.J., English, K.L., Paddon-Jones, D., and Fry, C.S., Fourteen days of bed rest induces a decline in satellite cell content and robust atrophy of skeletal muscle fibers in middle-aged adults, J. Appl. Physiol., 2016, vol. 120, no. 8, p. 965.
Narici, M.V. and de Boer, M.D., Disuse of the musculo-skeletal system in space and on earth, Eur. J. Appl. Physiol., 2011, vol. 111, no. 3, p. 403.
Bamman, M.M., Clarke, M.S.F., Feeback, D.L., et al., Impact of resistance exercise during bed rest on skeletal muscle sarcopenia and myosin isoform distribution, J. Appl. Physiol., 1998, vol. 84, no. 1, p. 157.
Hortobágyi, T., Dempsey, L., Fraser, D., et al., Changes in muscle strength, muscle fibre size and myofibrillar gene expression after immobilization and retraining in humans, J. Physiol., 2000, vol. 524, p. 293.
Yasuda, N., Glover, E.I., Phillips, S.M., et al., Sex-based differences in skeletal muscle function and morphology with short-term limb immobilization, J. Appl. Physiol., 2005, vol. 99, no. 3, p. 1085.
Hvid, L., Aagaard, P., Justesen, L., et al., Effects of aging on muscle mechanical function and muscle fiber morphology during short-term immobilization and subsequent retraining, J. Appl. Physiol., 2010, vol. 109, no. 6, p. 1628.
Hood, D.A., Memme, J.M., Oliveira, A.N., and Triolo, M., Maintenance of skeletal muscle mitochondria in health, exercise, and aging, Annu. Rev. Physiol., 2019, vol. 81, p. 19.
Popov, D.V., Adaptation of skeletal muscles to contractile activity of varying duration and intensity: the role of PGC-1α, Biochemistry (Moscow), 2018, vol. 83, no. 6, p. 613.
Perry, C.G.R. and Hawley, J.A., Molecular basis of exercise-induced skeletal muscle mitochondrial biogenesis: historical advances, current knowledge, and future challenges, Cold Spring Harb. Perspect. Med., 2018, vol. 8, no. 9, p. 1.
Memme, J.M., Erlich, A.T., Phukan, G., and Hood, D.A., Exercise and mitochondrial health, J. Physiol., 2021, vol. 599, no. 3, p. 803.
Olesen, J., Kiilerich, K., and Pilegaard, H., PGC-1α-mediated adaptations in skeletal muscle, Pflugers Arch., 2010, vol. 460, no. 1, p. 153.
Narkar, V.A., Fan, W., Downes, M., et al., Exercise and PGC-1α-independent synchronization of type i muscle metabolism and vasculature by ERRγ, Cell Metab., 2011, vol. 13, no. 3, p. 283.
Scarpulla, R.C., Transcriptional paradigms in mammalian mitochondrial biogenesis and function, Physiol. Rev., 2008, vol. 88, no. 2, p. 611.
Pearen, M.A. and Muscat, G.E.O., The nuclear receptor Nor-1 is a pleiotropic regulator of exercise-induced adaptations, Exercise Sports Sci. Rev., 2018, vol. 46, no. 2, p. 97.
Wu, Z., Huang, X., Feng, Y., et al., Transducer of regulated CREB-binding proteins (TORCs) induce PGC-1α transcription and mitochondrial biogenesis in muscle cells, Proc. Natl. Acad. Sci. U.S.A., 2006, vol. 103, no. 39, p. 14379.
Pérez-Schindler, J., Summermatter, S., Salatino, S., et al., The corepressor NCoR1 antagonizes PGC-1α and estrogen-related receptor α in the regulation of skeletal muscle function and oxidative metabolism, Mol. Cell. Biol., 2012, vol. 32, no. 24, p. 4913.
Williams, R.S., Salmons, S., Newsholme, E.A., et al., Regulation of nuclear and mitochondrial gene expression by contractile activity in skeletal muscle, J. Biol. Chem., 1986, vol. 261, no. 1, p. 376.
Williams, R.S., Garcia-Moll, M., Mellor, J., et al., Adaptation of skeletal muscle to increased contractile activity. Expression nuclear genes encoding mitochondrial proteins, J. Biol. Chem., 1987, vol. 262, no. 6, p. 2764.
Morrison, P.R., Biggs, R.B., and Booth, F.W., Daily running for 2 wk and mRNAs for cytochrome c and α-actin in rat skeletal muscle, Am. J. Physiol., 1989, vol. 257, no. 5, p. C936.
Freyssenet, D., Connor, M.K., Takahashi, M., and Hood, D.A., Cytochrome c transcriptional activation and mRNA stability during contractile activity in skeletal muscle, Am. J. Physiol., 1999, vol. 277, no. 1, p. E26.
Hood, D.A., Zak, R., and Pette, D., Chronic stimulation of rat skeletal muscle induces coordinate increases in mitochondrial and nuclear mRNAs of cytochrome-c-oxidase subunits, Eur. J. Biochem., 1989, vol. 179, no. 2, p. 275.
Freyssenet D, Connor MK, Takahashi M, et al. Cytochrome c transcriptional activation and mRNA stability during contractile activity in skeletal muscle. Am. J. Physiol. 1999. V. 277. № 6. P E26.
Nishida, Y., Tanaka, H., Tobina, T., et al., Regulation of muscle genes by moderate exercise, Int. J. Sports Med., 2010, vol. 31, no. 9, p. 656.
Robinson, M.M., Dasari, S., Konopka, A.R., et al., Enhanced protein translation underlies improved metabolic and physical adaptations to different exercise training modes in young and old humans, Cell Metab., 2017, vol. 25, no. 3, p. 581.
Granata, C., Caruana, N.J., Botella, J., et al., Training-induced bioenergetic improvement in human skeletal muscle is associated with non-stoichiometric changes in the mitochondrial proteome without reorganization of respiratory chain content, Nat. Commun., 2021, vol. 12, no. 1, p. 7056.
Radom-Aizik, S., Hayek, S., Shahar, I., et al., Effects of aerobic training on gene expression in skeletal muscle of elderly men, Med. Sci. Sports Exercise, 2005, vol. 37, no. 10, p. 1680.
Chapman, M.A., Arif, M., Emanuelsson, E.B., et al., Skeletal muscle transcriptomic comparison between long-term trained and untrained men and women, Cell Rep., 2020, vol. 31, no. 12, p. 107808.
Pillon, N.J., Gabriel, B.M., Dollet, L., et al., Transcriptomic profiling of skeletal muscle adaptations to exercise and inactivity, Nat. Commun., 2020, vol. 11, no. 1, p. 470.
Keller, P., Vollaard, N.B.J., Gustafsson, T., et al., A transcriptional map of the impact of endurance exercise training on skeletal muscle phenotype, J. Appl. Physiol., 2011, vol. 110, no. 1, p. 46.
Turan, N., Kalko, S., Stincone, A., et al., A systems biology approach identifies molecular networks defining skeletal muscle abnormalities in chronic obstructive pulmonary disease, PLoS Comput. Biol., 2011, vol. 7, no. 9, p. e1002129.
Lindholm, M.E., Marabita, F., Gomez-Cabrero, D., et al., An integrative analysis reveals coordinated reprogramming of the epigenome and the transcriptome in human skeletal muscle after training, Epigenetics, 2014, vol. 9, no. 12, p. 1557.
Lindholm, M.E., Giacomello, S., Werne Solnestam, B., et al., The impact of endurance training on human skeletal muscle memory, global isoform expression and novel transcripts, PLoS Genet., 2016, vol. 12, no. 9, p. e1006294.
Clarke, K., Ricciardi, S., Pearson, T., et al., The role of Eif6 in skeletal muscle homeostasis revealed by endurance training co-expression networks, Cell Rep., 2017, vol. 21, no. 6, p. 1507.
Popov, D.V., Makhnovskii, P.A., Shagimardanova, E.I., et al., Contractile activity-specific transcriptome response to acute endurance exercise and training in human skeletal muscle, Am. J. Physiol.: Endocrinol. Metab., 2019, vol. 316, no. 4, p. E605.
Knuiman, P., Hangelbroek, R., Boekschoten, M., et al., Impact of protein supplementation during endurance training on changes in skeletal muscle transcriptome, BMC Genomics, 2020, vol. 21, no. 1, p. 397.
Makhnovskii, P.A., Bokov, R.O., Kolpakov, F.A., and Popov, D.V., Transcriptomic signatures and upstream regulation in human skeletal muscle adapted to disuse and aerobic exercise, Int. J. Mol. Sci., 2021, vol. 22, no. 3, p. 1208.
Liu, X., Zhang, Y., Ni, M., et al., Regulation of mitochondrial biogenesis in erythropoiesis by mTORC1-mediated protein translation, Nat. Cell Biol., 2017, vol. 19, no. 6, p. 626.
Cheng, Z., Teo, G., Krueger, S., et al., Differential dynamics of the mammalian mRNA and protein expression response to misfolding stress, Mol. Syst. Biol., 2016, vol. 12, no. 1, p. 855.
Catoire, M., Mensink, M., Boekschoten, M.V., et al., Pronounced effects of acute endurance exercise on gene expression in resting and exercising human skeletal muscle, PLoS One, 2012, vol. 7, no. 11, p. e51066.
McLean, C.S., Mielke, C., Cordova, J.M., et al., Gene and microRNA expression responses to exercise; relationship with insulin sensitivity, PLoS One, 2015, vol. 10, no. 5, p. e0127089.
Dickinson, J.M., D’Lugos, A.C., Naymik, M.A., et al., Transcriptome response of human skeletal muscle to divergent exercise stimuli, J. Appl. Physiol., 2018, vol. 124, no. 6, p. 1529.
Vissing, K. and Schjerling, P., Simplified data access on human skeletal muscle transcriptome responses to differentiated exercise, Sci. Data, 2014, vol. 1, p. 140041.
Rowlands, D.S., Thomson, J.S., Timmons, B.W., et al., Transcriptome and translational signaling following endurance exercise in trained skeletal muscle: impact of dietary protein, Physiol. Genomics, 2011, vol. 43, no. 17, p. 1004.
Neubauer, O., Sabapathy, S., Ashton, K.J., et al., Time course-dependent changes in the transcriptome of human skeletal muscle during recovery from endurance exercise: from inflammation to adaptive remodeling, J. Appl. Physiol., 2014, vol. 116, no. 3, p. 274.
Popov, D.V., Makhnovskii, P.A., Kurochkina, N.S., et al., Intensity-dependent gene expression after aerobic exercise in endurance-trained skeletal muscle, Biol. Sports, 2018, vol. 35, no. 3, p. 277.
Rudler, D.L., Hughes, L.A., Perks, K.L., et al., Fidelity of translation initiation is required for coordinated respiratory complex assembly, Sci. Adv., 2019, vol. 5, no. 12, p. eaay2118.
Rudler, D.L., Hughes, L.A., Viola, H.M., et al., Fidelity and coordination of mitochondrial protein synthesis in health and disease, J. Physiol., 2021, vol. 599, no. 14, p. 3449.
Khalimonchuk, O., Bird, A., and Winge, D.R., Evidence for a pro-oxidant intermediate in the assembly of cytochrome oxidase, J. Biol. Chem., 2007, vol. 282, no. 24, p. 17442.
Hu, D., Liu, Z., Qi, X., and Perez, M.J., Mitochondrial quality control strategies: potential therapeutic targets for neurodegenerative diseases? Front. Neurosci., 2021, vol. 15, p. 746873.
Chopard, A., Lecunff, M., Danger, R., et al., Large-scale mRNA analysis of female skeletal muscles during 60 days of bed rest with and without exercise or dietary protein supplementation as countermeasures, Physiol. Genomics, 2009, vol. 38, no. 3, p. 291.
Abadi, A., Glover, E.I., Isfort, R.J., et al., Limb immobilization induces a coordinate down-regulation of mitochondrial and other metabolic pathways in men and women, PLoS One, 2009, vol. 4, no. 8, p. e6518.
Reich, K.A., Chen, Y.W., Thompson, P.D., et al., Forty-eight hours of unloading and 24 h of reloading lead to changes in global gene expression patterns related to ubiquitination and oxidative stress in humans, J. Appl. Physiol., 2010, vol. 109, no. 5, p. 1404.
Alibegovic, A.C., Sonne, M.P., Højbjerre, L., et al., Insulin resistance induced by physical inactivity is associated with multiple transcriptional changes in skeletal muscle in young men, Am. J. Physiol.: Endocrinol. Metab., 2010, vol. 299, no. 5, p. 752.
Lammers, G., Poelkens, F., Duijnhoven, N.T.L., et al., Expression of genes involved in fatty acid transport and insulin signaling is altered by physical inactivity and exercise training in human skeletal muscle, Am. J. Physiol.: Endocrinol. Metab., 2012, vol. 303, no. 10, p. e1245.
Rullman, E., Fernandez-Gonzalo, R., Mekjavić, I.B., et al., MEF2 as upstream regulator of the transcriptome signature in human skeletal muscle during unloading, Am. J. Physiol.: Regul. Integr. Comp. Physiol., 2018, vol. 315, no. 4, p. R799.
Trevino, M.B., Zhang, X., Standley, R.A., et al., Loss of mitochondrial energetics is associated with poor recovery of muscle function but not mass following disuse atrophy, Am. J. Physiol.: Endocrinol. Metab., 2019, vol. 317, no. 5, p. E899.
Mahmassani, Z.S., Reidy, P.T., McKenzie, A.I., et al., Age-dependent skeletal muscle transcriptome response to bed rest-induced atrophy, J. Appl. Physiol., 2019, vol. 126, no. 4, p. 894.
Standley, R.A., Distefano, G., Trevino, M.B., et al., Skeletal muscle energetics and mitochondrial function are impaired following 10 days of bed rest in older adults, J. Gerontol. A, 2020, vol. 75, no. 9, p. 1744.
Fernandez-Gonzalo, R., Tesch, P.A., Lundberg, T.R., et al., Three months of bed rest induce a residual transcriptomic signature resilient to resistance exercise countermeasures, FASEB J., 2020, vol. 34, no. 6, p. 7958.
Migliavacca, E., Tay, S.K.H., Patel, H.P., et al., Mitochondrial oxidative capacity and NAD+ biosynthesis are reduced in human sarcopenia across ethnicities, Nat. Commun., 2019, vol. 10, no. 1, p. 5808.
Wilkinson, S.B., Phillips, S.M., Atherton, P.J., et al., Differential effects of resistance and endurance exercise in the fed state on signalling molecule phosphorylation and protein synthesis in human muscle, J. Physiol., 2008, vol. 586, no. 15, p. 3701.
Donges, C.E., Burd, N.A., Duffield, R., et al., Concurrent resistance and aerobic exercise stimulates both myofibrillar and mitochondrial protein synthesis in sedentary middle-aged men, J. Appl. Physiol., 2012, vol. 112, no. 12, p. 1992.
Leppek, K., Das, R., and Barna, M., Functional 5′ UTR mRNA structures in eukaryotic translation regulation and how to find them, Nat. Rev. Mol. Cell Biol., 2018, vol. 19, no. 3, p. 158.
Sinvani, H., Haimov, O., Svitkin, Y., et al., Translational tolerance of mitochondrial genes to metabolic energy stress involves TISU and eIF1-eIF4GI cooperation in start codon selection, Cell Metab., 2015, vol. 21, no. 3, p. 479.
Morita, M., Gravel, S.P., Chénard, V., et al., mTORC1 controls mitochondrial activity and biogenesis through 4E-BP-dependent translational regulation, Cell Metab., 2013, vol. 18, no. 5, p. 698.
Haimov, O., Sinvani, H., and Dikstein, R., Cap-dependent, scanning-free translation initiation mechanisms, Biochim. Biophys. Acta, Gene Regul. Mech., 2015, vol. 1849, no. 11, p. 1313.
Elfakess, R. and Dikstein, R., A translation initiation element specific to mRNAs with very short 5′UTR that also regulates transcription, PLoS One, 2008, vol. 3, no. 8, p. e3094.
Rudrappa, S.S., Wilkinson, D.J., Greenhaff, P.L., et al., Human skeletal muscle disuse atrophy: effects on muscle protein synthesis, breakdown, and insulin resistance—A qualitative review, Front. Physiol., 2016, vol. 7, p. 361.
Crossland, H., Skirrow, S., Puthucheary, Z.A., et al., The impact of immobilization and inflammation on the regulation of muscle mass and insulin resistance: different routes to similar end-points, J. Physiol., 2019, vol. 597, no. 5, p. 1259.
Bragoszewski, P., Turek, M., and Chacinska, A., Control of mitochondrial biogenesis and function by the ubiquitin-proteasome system, Open Biol., 2017, vol. 7, no. 4, art. ID 170007.
Lavie, J., De Belvalet, H., Sonon, S., et al., Ubiquitin-dependent degradation of mitochondrial proteins regulates energy metabolism, Cell Rep., 2018, vol. 23, no. 10, p. 2852.
Johnson, M.L., Robinson, M.M., and Nair, S.K., Skeletal muscle aging and the mitochondrion, Trends Endocrinol. Metab., 2013, vol. 24, no. 5, p. 247.
Louis, E., Raue, U., Yang, Y., et al., Time course of proteolytic, cytokine, and myostatin gene expression after acute exercise in human skeletal muscle, J. Appl. Physiol., 2007, vol. 103, no. 5, p. 1744.
Harber, M.P., Crane, J.D., Dickinson, J.M., et al., Protein synthesis and the expression of growth-related genes are altered by running in human vastus lateralis and soleus muscles, Am. J. Physiol.: Regul. Integr. Comp. Physiol., 2009, vol. 296, no. 3, p. 708.
Pasiakos, S.M., McClung, H.L., McClung, J.P., et al., Molecular responses to moderate endurance exercise in skeletal muscle, Int. J. Sport Nutr. Exercise Metab., 2010, vol. 20, no. 4, p. 282.
Stefanetti, R.J., Lamon, S., Wallace, M., et al., Regulation of ubiquitin proteasome pathway molecular markers in response to endurance and resistance exercise and training, Pflugers Arch., 2015, vol. 467, no. 7, p. 1523.
Møller, A.B., Vendelbo, M.H., Christensen, B., et al., Physical exercise increases autophagic signaling through ULK1 in human skeletal muscle, J. Appl. Physiol., 2015, vol. 118, no. 8, p. 971.
Cunha, T.F., Moreira, J.B.N., Paixão, N.A., et al., Aerobic exercise training upregulates skeletal muscle calpain and ubiquitin-proteasome systems in healthy mice, J. Appl. Physiol., 2012, vol. 112, no. 11, p. 1839.
Kim, Y.A., Kim, Y.S., Oh, S.L., et al., Autophagic response to exercise training in skeletal muscle with age, J. Physiol. Biochem., 2013, vol. 69, no. 4, p. 697.
Kim, Y.A., Kim, Y.S., and Song, W., Autophagic response to a single bout of moderate exercise in murine skeletal muscle, J. Physiol. Biochem., 2012, vol. 68, no. 2, p. 229.
Popov, D.V., Lysenko, E.A., Miller, T.F., et al., The effect of single aerobic exercise on the regulation of mitochondrial biogenesis in skeletal muscles of trained men: a time-course study, Hum. Physiol., 2015, vol. 41, no. 3, p. 296.
Young, J.C., Hoogenraad, N.J., and Hartl, F.U., Molecular chaperones Hsp90 and Hsp70 deliver pre-proteins to the mitochondrial import receptor Tom70, Cell, 2003, vol. 112, no. 1, p. 41.
Evgen’ev, M.B., Garbuz, D.G., and Zatsepina, O.G., Heat shock proteins: functions and role in adaptation to hyperthermia, Russ. J. Dev. Biol., 2005, vol. 36, no. 4, p. 218.
Williamson, C.L., Dabkowski, E.R., Dillmann, W.H., and Hollander, J.M., Mitochondria protection from hypoxia/reoxygenation injury with mitochondria heat shock protein 70 overexpression, Am. J. Physiol.: Hear. Circ. Physiol., 2008, vol. 294, no. 1, p. 249.
Shepherd, D.L., Hathaway, Q.A., Nichols, C.E., et al., Mitochondrial proteome disruption in the diabetic heart through targeted epigenetic regulation at the mitochondrial heat shock protein 70 (mtHsp70) nuclear locus, J. Mol. Cell. Cardiol., 2018, vol. 119, p. 104.
Ma, X., Xu, L., Alberobello, A.T., et al., Celastrol protects against obesity and metabolic dysfunction through activation of a HSF1-PGC1α transcriptional axis, Cell Metab., 2015, vol. 22, no. 4, p. 695.
Takahashi, M., Chesley, A., Freyssenet, D., and Hood, D.A., Contractile activity-induced adaptations in the mitochondrial protein import system, Am. J. Physiol., 1998, vol. 274, no. 5, p. 1380.
Memme, J.M., Slavin, M., Moradi, N., and Hood, D.A., Mitochondrial bioenergetics and turnover during chronic muscle disuse, Int. J. Mol. Sci., 2021, vol. 22, no. 10, p. 5179.
Jones, S.W., Hill, R.J., Krasney, P.A., et al., Disuse atrophy and exercise rehabilitation in humans profoundly affects the expression of genes associated with the regulation of skeletal muscle mass, FASEB J., 2004, vol. 18, no. 9, p. 1025.
Gustafsson, T., Osterlund, T., Flanagan, J.N., et al., Effects of 3 days unloading on molecular regulators of muscle size in humans, J. Appl. Physiol., 2010, vol. 109, no. 3, p. 721.
Møller, A.B., Vendelbo, M.H., Schjerling, P., et al., Immobilization decreases foxo3a phosphorylation and increases autophagy-related gene and protein expression in human skeletal muscle, Front. Physiol., 2019, vol. 10, p. 736.
Leermakers, P.A., Kneppers, A.E.M., Schols, A.M.W.J., et al., Skeletal muscle unloading results in increased mitophagy and decreased mitochondrial biogenesis regulation, Muscle Nerve, 2019, vol. 60, no. 6, p. 769.
Singh, K. and Hood, D.A., Effect of denervation-induced muscle disuse on mitochondrial protein import, Am. J. Physiol.: Cell Physiol., 2011, vol. 300, no. 1, p. C138.
Camera, D.M., Burniston, J.G., Pogson, M.A., et al., Dynamic proteome profiling of individual proteins in human skeletal muscle after a high-fat diet and resistance exercise, FASEB J., 2017, vol. 31, no. 12, p. 5478.
Funding
This study was supported by the Russian Science Foundation, project no. 21-15-00405.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This work does not contain any studies involving animals or human subjects performed by any of the authors.
CONFLICT OF INTERESTS
The authors declare that they do not have a conflict of interests.
Additional information
Translated by E. Babchenko
Rights and permissions
About this article
Cite this article
Bokov, R.O., Popov, D.V. Regulation of Mitochondrial Biogenesis in Human Skeletal Muscles Induced by Aerobic Exercise and Disuse. Hum Physiol 48, 261–270 (2022). https://doi.org/10.1134/S0362119722030033
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0362119722030033