Abstract
There remains to this day a great gap in understanding as to the role of B cells and their products—antibodies and cytokines—in mediating the protective response to Francisella tularensis, a Gram-negative coccobacillus belonging to the group of facultative intracellular bacterial pathogens. We previously have demonstrated that Francisella interacts directly with peritoneal B-1a cells. Here, we demonstrate that, as early as 12 h postinfection, germ-free mice infected with Francisella tularensis produce infection-induced antibody clones reacting with Francisella tularensis proteins having orthologs or analogs in eukaryotic cells. Production of some individual clones was limited in time and was influenced by virulence of the Francisella strain used. The phylogenetically stabilized defense mechanism can utilize these early infection-induced antibodies both to recognize components of the invading pathogens and to eliminate molecular residues of infection-damaged self cells.
Similar content being viewed by others
Introduction
Francisella tularensis is a Gram-negative coccobacillus belonging to the group of facultative intracellular bacterial pathogens proliferating in phagocytic cells, including neutrophils. Several subspecies of F. tularensis cause serious tularemia infection in both animals and humans. A safe and effective vaccine continues to be sought to protect against respiratory infection with hypervirulent F. tularensis subsp. tularensis1. One of the reasons we do not yet have a suitable vaccine relates to the considerable gaps in our knowledge of the body’s defense mechanisms against this dangerous pathogen.
The immune mechanisms contributing to the expression of protective immune response against F. tularensis remain a matter of debate. The theories and views as to the roles of individual cell types and subtypes in generating protective immune response are as diverse as are the infection models used to reveal their function. Originally, the T cell-mediated immune processes based on IFN-γ-dependent activation of macrophages were regarded as constituting the decisive factor in host defense against tularemia2,3,4,5. Although we have intensively studied macrophages as the primary hosts for Francisella replication6,7,8,9,10, the evaluation of their role in protection remains to this day dependent on the experimental approach taken and virulence of the Francisella strains used11,12. The contribution of polymorphonuclear leukocytes to the limitation and/or dissemination of infection around the body is similarly ambiguous13,14,15. The currently prevailing opinion is that neutrophils create detrimental effects by participating in tissue destruction following F. tularensis infection and thus contribute to pathogenic processes16. The primary interaction of dendritic cells with F. tularensis, which, as in other phagocytic cells, escapes from the phagosome and proliferates within the dendritic cell cytosol, ends with cytokine-producing cells that have not been sufficiently stimulated and therefore have limited capacity to influence the protective immune response17,18. As in the case of macrophages, however, the signaling pathways of dendritic cells are differentially regulated by virulent and attenuated strains of Francisella19.
Since the 1970s, cellular immunity to tularemia has been regarded as an immunologically specific component of protective response20,21,22,23. Logically, the T lymphocytes have become the target of attention24,25,26. The alpha/beta CD4 + and CD8 + lymphocytes are equally effective in resolving infection by primary sublethal F. tularensis attenuated live vaccine strain (LVS) and establishing protective response to secondary lethal challenge27. Moreover, gamma/delta T cells28,29,30 and such other rare T cell populations as, for example, CD4−CD8−CD3+alpha/beta+ gamma/delta− NK1.1− T cells31 can participate in the control of intracellular bacterium proliferation.
There remains to this day, a great gap in our knowledge as to the role of B cells in mediating protective response to F. tularensis infection. The basic function of B lymphocytes within the humoral branch of the adaptive immune system is to generate a spectrum of specific antibodies. According to the prevailing notion, however, the specific antibodies are not a decisive component of protective immunity. Early studies on lymphocyte response to the vaccine strain of F. tularensis demonstrated response of both T and B cell populations32. Gradually, it was shown that strong early protection, detectable on the third day after infection, is highly dependent on B cells33. Limitation of bacterial growth was in this case dependent on IFN-γ and probably independent of the antibodies. Subsequently, a regulatory role of B cells in controlling the secondary immune response to F. tularensis LVS infection that is independent of antibody production was demonstrated. The influence of B cells in controlling the classical primary response to F. tularensis was marginal34. When F. novicida was used instead of F. tularensis LVS, however, both primary and secondary murine infections were critically dependent on the B cells35. Both in vitro and in vivo experiments later proved that murine and human B cell lines, as well as murine peritoneal B cell subsets, are targets of F. tularensis invasion36. The internalization process in B-1a cells was observed to be dependent on functional surface B cell receptors alone. In B-1b and B-2 cells, by contrast, these receptors needed co-signaling from the co-receptor CR1/2 to initiate F. tularensis engulfment37. Infected B cells were activated and produced a range of cytokines, including IFN-γ, IL-12, and TNF-α38.
An early attempt to resolve the controversial role of humoral immunity in protecting against F. tularensis infection utilized the passive transfer of immune sera collected from human recipients of the live tularemia vaccine to naïve mice. Pooled immune sera were shown to protect mice fully against a lethal challenge with the LVS strain39. Contributions of specific antibodies to protection against F. tularensis infection were then repeatedly demonstrated in several infection models40,41,42,43,44,45. The immune sera used in the aforementioned studies were isolated from infected individuals during either the acute or convalescent phase of infection. In the course of our studies on the mutual interaction of F. tularensis with B cells36,37,45,46, we inadvertently identified that the antibody clones reacted with the F. tularensis proteins ATP synthase α chain, elongation factor TU, universal stress protein, ABC transporter, putative outer membrane protein OmpH, peroxidase/catalase, and chaperone protein DnaK in murine sera isolated 12 h postinfection from specific-pathogen free (SPF) Balb/c mice. These specificities were not detected in control sera from naïve SPF mice. To determine whether these clones are cross-reacting antibodies in sera of SPF mice or if there are natural antibodies induced by infection, we used germ-free (GF) mice for further study. Germ-free mice, unlike SPF mice, are not attacked by internal or external bacteria, so the possibility for cross-reactivity of infection-induced antibody clones is limited47,48.
Here, we demonstrate that GF mice infected with F. tularensis produce natural antibody clones reacting with 87 F. tularensis proteins, of which 63 antibody specificities are targeted to proteins having orthologs or analogs in eukaryotic cells. Production of some individual clones was limited in time and was influenced by virulence of the Francisella strain used. This early infection-induced antibody production might be a phylogenetically stabilized defense mechanism both to recognize the components of the invading pathogens and to eliminate the molecular residues of infection-damaged self cells.
Results
F. tularensis elicits early robust total serum antibody response
Our initial finding of early antibody production determined by ELISA in SPF mice after F. tularensis infection allowed for two interpretations. This reflected either a reaction of cross-reacting pre-existing antibody clones or the rapid natural response of innate B cell subsets. For this reason, we used the GF mice for further experiments and we compared their antibody response with the response of SPF mice. GF animals have no pre-existing contacts with bacteria whereas SPF mice have a natural, pre-existing microbiota, including many non-Francisella species. To determine the kinetics of early antibody response to infection, SPF and GF Balb/c mice were infected with F. tularensis subsp. holarctica FSC200. To ensure direct interaction of antibody-producing cells with bacteria, we used intraperitoneal infection, where direct interaction of individual B cell subtypes is possible without prior processing of the immunogen by antigen-presenting cells. Moreover, the peritoneal cavity provides a suitable model for the study of B cells because of the presence there of a unique peritoneal cavity-resident B cell subset known as B1a cells, in addition to B1b cells and conventional B2 cells. After 12, 24, and 48 h of colonization, total serum antibody levels were measured in each group of mice (Fig. 1). The antibody isotypes composition of Balb/c SPF and GF murine sera obtained from naïve mice without any bacterial colonization were very low and therefore are not shown in this figure. The basal individual antibody isotypes levels for GF mice were significantly lower in comparison with the corresponding isotypes levels for SPF mice. A lower level of circulating IgG1 and IgG2b was observed in normal uninfected GF mice when compared with uninfected SPF mice (Supplementary Fig. S1). A higher level of light kappa and lambda chains in analyzed SPF sera, however, are probably associated with higher content of antibodies in the SPF sera in both normal and infected mice (Supplementary Fig. S2).
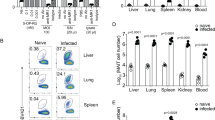
Isotypes of early infection-induced antibodies and their kinetics in Balb/c GF mice sera and Balb/c SPF mice sera obtained 12, 24, and 48 h after F. tularensis FSC200 intraperitoneal infection, dose of 102 CFU in total volume of 0.2 mL of saline. The immunoglobulin subclasses of serum samples were determined using a Quantibody mouse immunoglobulin isotype array while following the manufacturer’s instructions. The antibody isotypes compositions of Balb/c SPF and GF murine sera obtained from naïve mice without any bacterial colonization were very low and are not shown in these graphs. The basal individual antibody isotypes levels for GF mice were significantly lower in comparison with the corresponding isotypes levels for SPF mice (data not shown). The Quantibody mouse immunoglobulin isotype arrays were performed in biological triplicates (individual sera) for all time intervals and were independently repeated at least three times. Reference isotypes are presented. Values are expressed as mean ± standard deviation (SD) and analyzed for significance using Student’s two-tailed t-test. Differences were considered statistically significant at **p < 0.01 and *p < 0.05 and are denoted by asterisks.
The infection of GF as well as SPF mice with F. tularensis confirmed that infection induced the early antibody response as soon as 12 h post infection in both mice strains (Fig. 1). The infection with F. tularensis significantly increased the levels of all antibody isotypes in GF sera and in sera of SPF mice with the exceptions of IgD and IgE. The IgG1 and IgG2b isotypes dominated in sera of SPF mice, while IgM, IgA, and IgG3 predominated in GF mice. The kinetics of individual antibody isotypes 48 h post infection suggest the infection induces an immediate humoral stress response that is dominantly produced by early-induced natural antibodies.
Early robust IgM response occurred only after the infection of GF mice with F. tularensis subsp. holarctica FSC200 at all monitored time points. The IgM antibodies constitute the major component of the natural antibodies and should also be the first class of antibodies produced during early antibody response to infection. Our results demonstrate that this corresponds only to the response of GF mice, as the dominant isotype of early infection-induced antibodies of SPF mice are of IgG2b isotype. Assessed globally, the serum IgM and IgG antibodies were barely detectable in both GF and SPF mice before Francisella infection. The early humoral response of GF mice to infection is more intense, however, than is that of SPF mice 12 h post infection (Fig. 2).
Western blot analysis to evaluate F. tularensis FSC200 whole-cell lysate protein targets recognized by sera from F. tularensis FSC200 infected Balb/c mice and detected by chemiluminescence. The 2-D reference immunoblots demonstrated the reaction of antibody clones in murine sera obtained from Balb/c GF (upper) and SPF (bottom) mice at 12 h post infection. A total 200 μg of protein was separated by immobilized pH gradient (IPG) strips (3–10) and 12% (w/v) SDS-PAGE gels. Polyclonal peroxidase-conjugated goat anti-mouse IgG antibody was used for secondary antibody detection. All experiments were performed in biological triplicates (individual sera) for all time intervals and were independently repeated at least three times.
The increased level of infection-induced IgM antibody isotype in GF mice at 12 h postinfection and other time intervals in comparison with the levels of IgM isotype in sera of infected SPF mice may be caused by there being a different spectrum of IgM antibody clones (specificities) that reacted with different Francisella proteins (Supplementary Fig. S3).
Molecular targets of the early produced antibodies
Francisella tularensis subs. holarctica FSC200 whole-cell lysate was separated by 2-DE using broad range IPG strips of pH 3–10. To validate reproducibility of the separation technique, two independent whole-cell lysates were prepared. Reference silver-stained gel was prepared. Other gels were electroblotted onto polyvinylidene difluoride membrane. Immunoblotting was subsequently performed with mice tularemic sera from SPF and GF mice. Biological triplicates of SPF and GF sera were prepared at selected time points and immunoblotting experiments were performed twice with each serum. Normal murine SPF and GF serum was prepared from blood of Balb/c mice without exposure to any F. tularensis strain. Coomassie-stained 2-DE gels were then prepared to excise the selected spots that were digested and the reactivities on bacterial extracts were determined. The mass spectrometry approach successfully assessed in total 87 different F. tularensis proteins that reacted with the sera obtained from mice at 12, 24, and 48 h post infection (Fig. 3). A similarity analysis demonstrated that the proteins reacted according to the model (GF versus SPF) that was used and the time intervals when the sera were obtained after infection. The dominant immunoreactive protein spots of F. tularensis FSC200 detected on at least three immunoblots are shown on the reference 2-D protein map (Fig. 4) and the proteins are summarized in Supplementary Table S1. A total 50 immunoreactive Francisella proteins were unique targets of GF murine sera with no appearance of interaction with SPF sera isolated during the intervals of innate immune response to F. tularensis infection (see Fig. 3 and Supplementary Tab. S1).
The 2-D reference gels of F. tularensis FSC200 whole-cell lysate protein targets, which are recognized by sera from F. tularensis FSC200 infected Balb/c GF mice. (a) The targets of Balb/c GF mice antibody response at 12 h post infection, (b) the depicted unique targets of Balb/c GF mice antibody response at 24 and 48 h post infection, (c) the depicted unique antibody response of Balb/c SPF mice at 24 and 48 h post infection. A total 150 μg of protein was separated by immobilized pH gradient (IPG) strips (3–10) and 12% (w/v) SDS-PAGE gels. Identified immunogenic protein spots were numbered in preparative 2-D SDS-PAGE gels, excised for MS/MS analysis, corresponding to the proteins in Table S1 (designated letters and numbers of protein spots were used only for internal purposes and do not necessarily imply the same identity of the labeled proteins). Polyclonal peroxidase-conjugated goat anti-mouse IgG antibody was used for secondary antibody detection. All experiments were performed in biological triplicates (individual sera) for all time intervals and were independently repeated at least three times.
A number of immunoreactive proteins identified in this study already have been described using different infection models, including human tularemia at the stage of adaptive immune response (see references for Supplementary Table S1). Some studies were conducted to identify and characterize the virulence factors that F. tularensis uses to infect a wide variety of hosts, induce disease, and evade immune defense mechanisms. Some of them are listed as immunoreactive proteins. Sera isolated from GF as well as SPF mice during very early time points (i.e., 12, 24, and 48 h) postinfection reacted also with several virulence factors already identified. In this respect, the robust reactions to Francisella virulence factors were mainly observed using GF murine serum obtained 12 h after intraperitoneal inoculation by bacteria.
In this study, 19 F. tularensis proteins that were recognized by the early infection-induced antibody clones had already been recognized as potential virulence factors with specified function and importance. However, if F. tularensis survives in arthropods, fresh water amoeba, and mammals, where it has both intracellular and extracellular phases, it could be reasonable to expect changes in expression of (virulence) factors enabling Francisella to survive in these various environments. For these reasons, denomination of Francisella virulence factors is debatable49,50,51,52,53.
Namely, these are fructose-1,6-bisphosphatase (FTS_1656), chaperone ClpB (FTS_0084), intracellular growth locus A protein (FTS_0097/1125), intracellular growth locus C protein (FTS_0099/1127), superoxide dismutase (FTS_1747), peroxidase/catalase (FTS_1471), host factor I for bacteriophage Q beta replication (FTS_0889), heat shock protein GrpE (FTS_1166), chaperone protein DnaK (FTS_1167), co-chaperonin GroES (FTS_1671), OmpA family protein (FTS_1295), fumarate hydratase (FTS_0527), succinyl-CoA synthetase subunit beta (FTS_1517), succinyl-CoA synthetase subunit alpha (FTS_1518), DNA-directed RNA polymerase subunit alpha (FTS_0258), transcription elongation factor GreA (FTS_1440), glycerophosphoryl diester phosphodiesterase (FTS_1476), 30S ribosomal protein S1 (FTS_1862), bacterioferritin (FTS_0617), and trigger factor (FTS_0882). Moreover, 2 proteins, namely OmpA family protein (FTS_1295) and glycerophosphoryl diester phosphodiesterase (FTS_1476), were annotated as outer membrane associated proteins that might play an important role in immunity during host–pathogen interaction.
The early Francisella-induced antibodies reacted also with SodB (FTS_1747), KatG (FTS_1471), DnaK (FTS_1167), and GroES (FTS_1671), which are proteins that already have been reported to be released or secreted by F. tularensis but the mechanisms for their secretion remain unclear. In addition, we identified 6 immunoreactive proteins annotated with signal peptide according to SignalP 5.0 software, namely KatG (FTS_1471), OmpA (FTS_1295), GTP-dependent nucleic acid-binding protein EngD (FTS_0935), Omp protein of unknown function (FTS_0008), glycerophosphoryl diester phosphodiesterase (FTS_1476), and hypothetical protein (FTS_0815). All the mentioned proteins are listed along with their characteristics in Supplementary Table S1.
All identified proteins reacting with the sera from Francisella-infected GF and SPF mice were sorted into Clusters of Orthologous Groups (COG) functional categories (Fig. 5). Two dominant COGs relate to biogenesis at the stage of protein post-translational modification and chaperon-accompanied protein translocation. Sixty-three of the 87 identified antibody clones reacted with bacterial proteins that have orthologs or analogs in eukaryotic cells. The remaining 24 prokaryotic targets are characteristic for F. tularensis, of which 7 are hypothetical proteins. The identities of those proteins were determined according to the UniProtKB database. Among the eukaryotic analogs or orthologs, mitochondrial and cytosolic proteins were found to be the dominant targets of early Francisella-induced antibodies (Fig. 6). The reactivities of all SPF and GF murine sera obtained at the selected time points are described in Supplementary Table S1. Of 50 unique GF antibody specificities, the serum of GF mice contains 12 with no eukaryotic orthologs (Table S2). These antibody specificities have protein targets that, according to the Clusters of Orthologous Groups categorization, are involved on the one hand in amino acid metabolism and transport, protein intracellular trafficking and secretion, and membrane biogenesis, and on the other hand in cell cycle control and replication and repair (see Fig. 6).
Categorization of infection induced antibody response protein targets according to Clusters of Orthologous Groups (COGs) characterizing the functional as well as topographical characteristics of identified protein groups as predicted by EggNOG v. 5.0.0. All identified protein targets by immune sera of both mouse models and all time intervals tested are summarized.
Venn diagram representing the eukaryotic cell compartments within which are summarized orthologs or analogs of all F. tularensis proteins reactive with early infection-induced antibody clones. The numbers within sectors represent a set of proteins located at a given cellular compartment. Data on cell localization were taken from the UniProtKB database.
Francisella-induced early unspecific antibody response
The sera from Balb/c mice infected with F. tularensis subs. holarctica FSC200 obtained in early 12 h interval after infection reacted with different sets of F. tularensis subsp. holarctica FSC200 and F. tularensis LVS proteins used in the form of whole-cell lysates for immunoblotting. The intensity of reaction corresponded to virulence of the bacterial strain used for infection. The early humoral response of mice to F. tularensis subs. holarctica FSC200 is more intense in terms of both the quantity of antibodies in serum and the spectrum of recognized targets when compared with LVS counterparts. In parallel with immunoblotting of sera obtained from GF and SPF mice infected by F. tularensis against F. tularensis whole-cell lysate proteins we used as targets also the whole-cell lysates of Klebsiella pneumonia (clinical isolate) and Pseudomonas aeruginosa CCM 3955. Both bacteria are conditionally pathogenic and can incidentally colonize the airways. The reaction of early Francisella-induced antibodies on these targets was weaker than was the response against F. tularensis whole-cell lysate, but it was still very significant (Supplementary Fig. S4). The reactions of protein targets of both alternative microbes with the sera obtained from F. tularensis-infected GF mice were more intense than was the response of sera obtained from SPF mice. The GF control sera failed to react with Klebsiella pneumoniae and Pseudomonas aeruginosa proteins. The control sera obtained from SPF mice reacted, but only slightly, with the proteins of both bacteria. The target proteins for the sera from Francisella-infected mice were not identified because we did not search for them.
If the sera from both GF and SPF Francisella-infected mice reacted with proteins of unrelated bacteria we tested also the response of these sera with the whole-cell lysate of Balb/c peritoneal cells. In this case, too, we have demonstrated the interaction of early infection-induced antibody clones obtained from GF and SPF mice with protein targets of whole-cell lysate of peritoneal cells. The responses of both GF and SPF mice appear to be similar in intensity, but they respond differently in terms of their protein targets (Supplementary Fig. S5).
Discussion
The immune response to natural infection with F. tularensis as well as the response to immunization of experimental animals or vaccination of humans with attenuated strains induced strong cell-mediated immunity. Originally, the delayed-type skin test or the in vitro lymphocyte stimulation test demonstrated the expression of cell-mediated immune response at the end of the first week after natural infection or vaccination20,54. Similarly, the utilization of whole-blood culture DNA synthesis response and IL-2 and IFN-gamma secretion authenticated the second week as marking the time interval needed for the Th1 type of T-cell-mediated immunity expression after vaccination of human volunteers with a live vaccine strain55. In parallel, it was demonstrated that a substantial antibody response of human patients occurred within the second week of infection54. Studies on the host genetic background of resistance to experimental Francisella infections56,57,58 and the participation of neutrophils, NK cells, or B cells in the resistance to primary tularemia infection13,33,59,60 drew attention to early innate immunity. This innate (natural, genetic, or inherent) immunity (or resistance) has been defined as “the ability of an organism to resist the effect of an ecological or physiological agent in the absence of any acquired (reactive) immunity”61. In other words, innate immune responses occur because cells are able rapidly to react to invading microorganisms without the requirement for an antigen-specific adaptation. Innate immune responses in the context of Francisella models operate until the fifth day post infection6. During this time period, the unspecific early protective immunity can be demonstrated in murine models between the second and third days post infection with vaccine strain of F. tularensis33,62; human volunteers respond to LVS vaccination by activation of γδ T cells and NK cells 24 h post vaccination63. Transcriptome analysis has demonstrated the induction of proinflammatory and innate immunity-related genes at early post-vaccination time points, meaning ≤ 48 h64.
Although we have been investigating immunity against F. tularensis infection for many decades our knowledge as to the role of natural humoral mechanisms remains relatively rudimentary. We have identified, rather serendipitously, several antibody clones isolated 12 h post infection from sera of SPF mice immunized with LVS that reacted with F. tularensis proteins and were absent in the sera of control mice. Such an early antibody response could be either a response of memory B cells resulting from previous interactions with microorganisms or an induction of T-cell-independent early natural antibody production induced by F. tularensis strain FSC200 infection. We decided to resolve this dilemma using GF mice that should not have had any interaction with bacteria during their ontogeny, and not even with bacteria that constitute the gut microbiome of SPF mice.
According to earlier studies, the antibody production of GF mice in response to corpuscular antigens, such as from sheep red blood cells65,66 or hamster erythrocytes67, was comparable to that seen in conventional mice. Similarly, only limited differences were observed between conventional and GF mice in response to hydrophilic polysaccharide dinitrophenyl-lysine-Ficoll or dinitrophenylated bovine serum albumin66. In all these early studies, the humoral immune response or competence was monitored in the intervals corresponding to induction of acquired immune responses. Antibody production during early time intervals corresponding to induction of innate immune responses was not in focus, despite the fact that the natural antibodies have been shown—beyond immediate protection against infection—to play a substantial regulatory role in the actions of the defense system and homeostasis68.
Our study revealed the early humoral innate immune response of GF as well as SPF mice to the intracellular bacterium F. tularensis at least from the 12th hour after intraperitoneal inoculation with the bacteria. The majority of antibody clones reacted with bacterial proteins that have orthologs or analogs in eukaryotic cells. The dominant targets were cytosolic and mitochondrial proteins. Some of the clones reacted also directly with eukaryotic proteins separated by SDS PAGE of naïve murine peritoneal cells lysate. Considering that natural antibodies (autoantibodies) have been shown to recognize evolutionarily fixed epitopes shared between foreign and self-antigens69, the results of our study correspond to the generally accepted target profile of natural antibodies. The isotypes composition and their relative proportions in the serum of GF mice also correspond to the general characteristics of natural antibodies. The kinetics of antibody production suggest a temporary nature of humoral innate response, at least in the sense that some antibody clones (not the B cell clones) disappear within the next 12 h. Such kinetics suggest that different (sub)types of B cells and plasma cells might participate in the early natural humoral immune response to infection. Part of the clones persists, however, and these specificities can be identified even in the acquired immune response phase. Antibody specificities that were detected in both the innate immunity phase and the acquired immunity phase may result from the B cell responses regulation, selection of the B cell repertoires, and/or the regulation of B cell development, all of which are processes modulated by early produced natural antibodies70. An anamnestic response of memory B cells derived from an innate B cell pool by infection may ensure selected specific and long-term acquired humoral responses.
This comparison of the GF and SPF mice antibody response intensities leads to the conclusion that previous interaction with bacteria instructed the immune system of SPF mice and that the uniform humoral response of the naïve innate immune system of GF mice is more limited, and especially at IgM isotype level. Moreover, data in the literature proves that the exogenous antigenic stimulation has changed the relative proportion of cells secreting IgM isotype antibody on the one hand and cells secreting IgG and IgA antibody isotypes on the other. This means that exogenous antigenic stimulation has a great effect on the background IgM-, IgG-, and IgA-secreting cells, and, most likely, it also changes the repertoire of antigenic specificities of the antibodies produced71.
It has been demonstrated that antibodies—including natural antibodies—can influence the immune processes through diverse mechanisms, among these being opsonization of microbes, complement activation, formation of circulating antigen–antibody complexes and their trapping in lymphoid organs, antibody-dependent cell-mediated bactericidal activity, signaling through Fc receptors, and modulation of the inflammatory response. Moreover, natural antibodies may prevent pathogen dissemination to target organs and influence the immunogenicity through enhanced antigen-trapping in secondary lymphoid organs72,73. All the activities of natural antibodies mentioned above can modulate the induction or regulation of acquired immune responses73.
Among the early induced natural antibody clones were clones reacting with ClpB, SodB, KatG, Hfq, IglA, IglB, and IglC proteins, which already had been demonstrated to be Francisella virulence factors responsible for intracellular survival and stress response74. In addition to the KatG and SodB proteins that are secreted, antibody clones reacting with the other secreted proteins DnaK and GroES also have been identified. It is noteworthy that antibody clones reacting with these virulence factors encompassing both unique bacterial proteins and proteins having eukaryotic orthologs or analogs, with the exception of SodB, were identified only at time point 12 h postinfection. All these antibody specificities may potentially modulate the early stages of interaction between Francisella and the host cell. In this context, we would mention that KL Elkins et al. have disclosed in several studies an early T-cell-independent and B-cell-dependent specific or eventually nonspecific protective response that can be demonstrated between the second and third day after primary sublethal infection of mice33,62,75,76. Due to the fact that, according to a notion prevailing since the late 1990s, the antibodies are produced up to about the fifth or seventh day after vaccination or infection, early resistance to Francisella and/or Listeria has not been attributed to the involvement of antibodies. According to the results presented here, however, the early production of infection-induced natural antibody clones reacting with F. tularensis proteins may be the critical partner of the early cellular responses controlling the presence of bacteria in the body. A search for direct proof of antibody involvement in the induction of this early anti-infective resistance should be a matter for further experiments.
Conclusion and perspectives
We conclude that both GF and SF mice respond to F. tularensis strain FSC200 intraperitoneal infection by early IgM, IgG3, and IgA antibody production that can be detected in serum as early as 12 h after infection. Some antibody specificities production has a temporary character. The spectrum of 87 antibody specificities, consisting of a group of bacterial proteins, of which 63 have orthologs or analogs in eukaryotic cell proteins, along with the kinetics of their production, lead to the conclusion that this is a production of natural antibodies, probably by B-1a cells induced by F. tularensis intraperitoneal infection. Preliminary testing of cell subtypes infection in murine peritoneum proved B-1a cells to be the dominant target of Francisella invasion (data not shown). It had already been demonstrated that body cavity B-1 cells are activated by infection and can be considered—unlike lymph tissues B-1 cells—as responder cells70,77,78. Our results, however, do not correspond to those obtained in a model of influenza infection where no specific IgM antibodies in serum were detected at early intervals after intranasal infection77,78. The reason for this difference can be seen in the different cell types, and including B cell subtypes, that encounter infectious agents as first responders on the nasal mucosa and in the lung in comparison with those which are first responders in peritoneum. Differences in pathogen type (i.e., virus versus bacterium) also can play a substantial role.
Such an early production of antibodies raised several questions. First, where is the origin of this antibody production? Is it directly in the peritoneum or does it occur after translocation of B cells to lymph organs? Further, which B cell subset(s) really produce(s) these antibodies and which receptor(s) and signaling pathways are involved in this phenomenon? What is the primary target of the antibodies thus produced? Are they primarily autoantibodies or are they already targeted, due to the phylogenetic relationship between eukaryotes and prokaryotes, as well as phylogenetic memory, to components of an invading microorganism? Finally, how do these early produced antibodies participate in the Francisella–host cell interactions and to what extent can they influence the induction of protective immune response? The positive effects of B-1a cells and of antibodies produced early by B-1a cells already have been reported79,80,81.
The phylogenetically stabilized defense mechanism can utilize these early infection-induced antibodies both to recognize the components of the invading pathogens and to eliminate the pathogen-damaged own cells. Such early defense steps can thus be characterized as a booster of opsonophagocytosis (or neutralization) of pathogens that is one of the key events in the induction of acquired immunity mechanisms that already are targeted upon eliminating the pathogen82 (Table 1). Nevertheless, there remain substantial gaps in our understanding of cooperation between the early humoral and cellular branches of the innate and acquired immune systems in expression of protective responses to F. tularensis infection that can lead us to the design of an effective and safe vaccine.
Material and methods
Bacteria
Francisella tularensis subsp. holarctica strain FSC200 (F. tularensis FSC200) was kindly provided by Åke Forsberg (Swedish Defence Research Agency, Umeå, Sweden). Klebsiella pneumonia was a clinical isolate, and Pseudomonas aeruginosa CCM 3955 was obtained from Czech Collection of Microorganisms (Masaryk University, Brno, Czech Republic). Stocks of F. tularensis were precultivated on McLeod agar supplemented with bovine hemoglobin and IsoVitalex (Becton Dickinson, New Jersey, USA) at 37 °C for 24 h. Chamberlain medium83 was prepared according to the manufacturer’s instructions, pH was adjusted with HCl to 6.8 (unless otherwise stated), and the medium was sterile-filtered rather than autoclaved. Following incubation, bacterial colonies were lifted from the plates and resuspended in saline to reach optical density 1.00 (corresponding to bacterial 5 × 109 CFU/ml). The actual number of bacteria in the suspension utilized for the experiments was determined by serial dilutions and the number of colony-forming units (CFU) was calculated. Klebsiella pneumonia and Pseudomonas aeruginosa CCM 3955 were cultivated under standard cultivation procedures using blood agar plates.
Animals
Female BALB/c-SPF mice (SPF) were purchased from Velaz (Unetice, Czech Republic). Female germ-free (GF) BALB/c (hereafter GF mice) mice were kindly provided by the Institute of Microbiology of the Czech Academy of Sciences (Department of Gnotobiology, Novy Hradek, Czech Republic). The GF mice were housed under sterile conditions in Trexler-type plastic isolators with free access to autoclaved tap water and 50 kGy-irradiated sterile pellet diet Altromin 1414 (Altromin, Lage, Germany). Before experiments GF mice were transferred into sterile indiviually ventilated cages (IVC, Tecniplast, Italy). The SPF mice were housed under specific-pathogen-free conditions and GF mice were housed in micro-isolator cages under germ-free conditions. The mice were placed into sterile boxes with air-conditioning and stabilized temperature of 22 ± 2 °C. Light mode was 12 h of light and 12 h darkness45. All experiments on mice were conducted under supervision of the institution’s Animal Unit and were approved by the Animal Care and Use Committee of the Faculty of Military Health Sciences, University of Defence, Hradec Kralove, Czech Republic under project number 35/17. All experiments were carried out in compliance with the ARRIVE guidelines and regulations (http://www.nc3rs.org.uk/page.asp?id=1357).
Preparation of bacterial whole-cell lysates
The suspensions of bacterial cells (F. tularensis, Klebsiella pneumonia, Pseudomonas aeruginosa) were disrupted using a French press twice at 16,000 psi and cell debris was removed by centrifugation at 12,600×g for 30 min at 4 °C. Benzonase nuclease (250 U/μL, Sigma, St. Louis, MO, USA) was added to the supernatant, resulting in a final concentration of 0.5 U/ml of lysate.
Preparation of eukaryotic whole-cell lysate
For peritoneal cavity lavage, the mice were anesthetized under complete anesthesia; the abdominal skin was cleaned thoroughly with ethanol pads. For each mouse, a sterile tweezer was used to pinch and lift the middle section of the abdominal skin approximately 2 cm to create a pocket of peritoneum void of internal organs. A syringe with needle was filled with 10 ml phosphate-buffered saline (PBS) and the needle was inserted vertically into the pocket within the peritoneal cavity while being cautious to avoid injuring the internal organs. The PBS solution was injected into the peritoneum. Recovery of the lavage fluid was in the range of about 80–90%, depending on the peritoneal fat content and lavage technique. The retrieved suspension was centrifuged at 400×g for 5 min to concentrate any cellular components. Peritoneal lavage cells were then lysed with RIPA buffer (Thermo Fisher Scientific, USA) to obtain extracted proteins. The extraction was done according to the manufacture’s procedure. The protein suspension was subsequently precipitated using trichloroacetic acid/sodium deoxycholate (TCA-DOC) with acetone wash. Briefly, the protein suspension was mixed with 1/10 of the volume of 0.15% sodium deoxycholate (DOC) and incubated on ice for 15 min. The sample was then mixed with 1/10 of the original volume of 72% trichloroacetic acid (TCA) and the suspension was again incubated on ice for 15 min. Subsequently, the mixture was microfuged at 12,000×g, at 4 °C for 15 min. Supernatant was discarded and 1 ml of acetone chilled to − 20 °C was added. All steps of TCA-DOC precipitation were repeated twice. The pellet was air-dried for 5 min, subsequently resuspended in a minimal volume with 2-D sample buffer, then subjected to 2-D gel analysis.
Infection of mice and collection of murine sera
The SPF and GF mice were intraperitoneally inoculated with an F. tularensis FSC200 dose of 102 CFU in 0.2 ml total volume of saline. At three time intervals post-challenge (12 h, 24 h, and 48 h), infected mice were killed by cervical dislocation, blood was collected, and serum was prepared according to a common procedure. Sterility of serum was determined by plating on McLeod agar plates for 72 h. Obtained serum was pooled and tested for specific antibody titers using enzyme-linked immunosorbent assay (ELISA). Serum aliquots were stored in volume 500 μl sera at − 80 °C. Normal murine serum was prepared from blood of Balb/c mice without exposure to any F. tularensis strain.
Serum analysis procedures
Determination of antibody isotypes
The immunoglobulin subclasses of serum samples were determined using a Quantibody mouse immunoglobulin isotype array (RayBiotech Life, Peachtree Corners, GA, USA) while following the manufacturer’s instructions. The mouse array is a glass chip-based multiplex assay that detects the 8 mouse immunoglobulin subclasses (IgG1, IgG2a, IgG2b, IgG3, IgA, IgD, IgE, and IgM) plus the 2 light chain types (kappa and lambda). Briefly, samples were mixed with the Cy3 equivalent dye-labeled detection antibody and applied to the array slide. After a wash step, the slides were dried and scanned with a laser scanner. Results were evaluated by densitometric data extraction (Quantibody Q Analyzer, RayBiotech Life, Peachtree Corners, GA, USA) (https://www.raybiotech.com/analysis-tools/).
ELISA-based method for titer determination
ELISA laboratory technique was selected to assess the magnitude of humoral immune response for evaluating the time course and immune status of an ongoing infection. Briefly, ELISA plates were coated with F. tularensis FSC200 autolysate Ag-2 in concentration corresponding to 1.1 × 109 bacteria/well and incubated overnight at 4 °C. The following day, plates were washed with PBS/Tween 20 (0.05% [v/v]) solution and blocked with 200 μl of 10% calf serum in PBS for 30 min at 37 °C. After washing, serum samples were serially diluted with PBS onto plates and incubated at 37 °C for 90 min. Subsequently, plates were washed and enzyme-labeled antibodies were added for another 90 min at 37 °C. Polyclonal HRP-conjugated goat anti-mouse Ig (Bio-Rad AbD Serotec Limited, Oxford, UK) was used for detection. Subsequently, plates were washed and TMB solution was used for color development. Final optical density values were measured at 450 nm using a plate reader MultiDetector Magic XBC Paradigm Detection Platform (Beckman Coulter, Pasadena, CA, USA). Uncoated plates were also assayed to ensure that antibodies in immune sera did not bind a plastic. Antibody titers were expressed as optical density values obtained after subtraction of the blank value, which was obtained from the duplicate wells containing wash buffer.
Immunoproteomic analysis of murine sera
Mini two-dimensional electrophoresis
Bacterial whole-cell lysates were solubilized in isoelectric focusing (IEF) rehydration buffer containing 6 M urea, 2 M thiourea, 4% CHAPS, 40 mM TRIS-base, 0.12% De Streak, and 0.5% bromophenol blue with 1% carrier ampholytes (pH 3–10) and 0.5% carrier ampholytes (pH 6–11). Protein concentration was defined by bicinchoninic acid assay. The IEF was performed using a Multiphor II electrophoresis system (GE Healthcare, Chicago, Illinois, USA) on 7 cm gradient pI 3–10 Immobiline DryStrip gels84. Briefly, proteins were loaded by in-gel rehydration onto polyacrylamide gel strips with a nonlinear immobilized 3–10 pH gradient (IPG) from GE Healthcare (Chicago, IL, USA) and separated according to their different pI values by IEF. Following IEF, the IPG strips were treated in equilibration buffer containing 2% (w/v) sodium dodecyl sulfate (SDS), 50 mM Tris/HCl (pH 8.8), 6 M urea, 30% (v/v) glycerol, and 1% (w/v) dithiothreitol (DTT). This was immediately followed by a second equilibration of strip in the same solution containing 4% (w/v) iodoacetamide (instead of DTT). In the second dimension, the IPG strips were embedded onto 12% homogeneous SDS polyacrylamide gels. Electrophoresis was performed on a Protean II Multi-Cell (Bio-Rad, Hercules, CA, USA) according to the standard running conditions84. Blue staining was then used to visualize proteins on a reference protein map while colloidal blue staining was performed for mass-spectrometric purposes. Analyses of reference maps were made by visual inspection and data were evaluated using Melanie 3 software (Bio-Rad Laboratories, Hercules, CA, USA)84.
Semi-dry western blotting
Selected 2-DE gels were electroblotted onto polyvinylidene difluoride membrane. All immunoreactive spots were detected using incubation with control or immune sera diluted 1:100 in the blocking buffer at 4 °C overnight. The next day, membranes were washed with 0.05% Tween 20 in Tris-buffered saline. Polyclonal peroxidase-conjugated goat anti-mouse IgG antibody (Dako, Copenhagen, Denmark) or monoclonal HRP-conjugated goat anti-mouse IgM antibody (Invitrogen, Carlsbad, Ca, USA) was diluted 1:100 in blocking buffer and used for secondary antibody detection. Membranes were then washed and prepared for visualizing using BM Chemiluminescence Blotting Substrate kit (Boehring, Mannheim, Germany) and CL-XPosure films (Pierce, Rockford, IL, USA).
Mass spectrometry profiling
In-gel digestion
Selected protein spots were excised from representative colloidal blue-stained gels and subjected to in-gel digestion. Briefly, gel pieces were covered with 100 mM Tris/HCl (pH 8.5) in 50% acetonitrile (ACN) at 30 °C for 20 min. Next, 50 mM ammonium bicarbonate (pH 7.8) in 5% ACN was added to each gel piece. Subsequently, gel pieces were vacuum-dried, covered with 0.1 μg of trypsin in 50 mM ammonium bicarbonate (pH 7.8) in 5% ACN, then gently shaken at 37 °C overnight. Protein digests were then extracted using 2% trifluoroacetic acid and 98% ACN. The final extracts were reduced using a vacuum dryer to approximately 15 μl and frozen at − 80 °C. The resulting peptides were mixed with 5 mg/ml of α-cyano-4-hydroxycinnamic acid in 50% acetonitrile and 0.1% trifluoroacetic acid. Peptide samples with matrix solution were spotted onto a MALDI plate.
Protein identification and characterization
The protein identification and characterization analysis of the murine sera have been done according45,84. Briefly, the mass spectra were recorded in the positive reflectron mode on a 4800 MALDI-TOF/TOF mass spectrometer (AB Sciex, Foster City, CA, USA) and operated in its delayed extraction mode. Internal calibration of mass spectra was conducted utilizing the tryptic autolytic peptides. Acquired data were evaluated using GPS Explorer Software v.3.6 (AB Sciex, Australia) (https://sciex.com/products/software), which integrates the Mascot search algorithm against the Francisella tularensis subsp. holarctica OSU18 genome database. The following parameters were used for protein identification: 100 ppm error, optional oxidation of methionine, fixed carbamidomethylation of cysteine residues, and one possible missed cleavage site allowed. Proteins were considered to be identified with confidence when a minimum of two peptide sequences per protein were identified and the GPS protein score confidence interval was equal to 100%. Trypsin was selected as the proteolytic enzyme. The identities of all proteins were determined according to the UniProtKB database85 (https://www.uniprot.org).
Bioinformatic analysis
A number of online programs were used for bioinformatic analysis and characterization of identified proteins. EggNOG 5.0.0 algorithm was applied to sort the Francisella-identified proteins into functional categories86 (http://eggnog5.embl.de/#/app/results). PSORTb 3.0. was used to predict localization based on the ORF sequences of identified proteins87 (https://www.psort.org/psortb/). SignalP 5.0 was used for predicting signal peptide presence (type II secretion pathway)88 (http://www.cbs.dtu.dk/services/SignalP/). LipoP 1.0 was applied to predict lipoproteins and to discriminate between lipoprotein signal peptides, other signal peptides, and n-terminal membrane helices in Gram-negative bacteria89 (http://www.cbs.dtu.dk/services/LipoP/).
Statistical analysis
Each immunoblot experiment was conducted at least in triplicate while using three biological replicates. Pearson correlation coefficient was calculated using GraphPad Prism 5.01 software (GraphPad Software, San Diego, CA, USA) (https://www.graphpad.com/scientific-software/prism/). All statistical analyses were also performed using GraphPad Prism 5.01 software (GraphPad Software, San Diego, CA, USA). In case of Quantibody immunoglobulin isotype array, four technical replicates were measured (according to the manufacturer’s instructions). Reference antibody isotypes and light kappa and lambda chains are presented as well as reference immunoblots. Values are expressed as mean ± standard deviation (SD) and analyzed for significance using Student’s two-tailed t-test. Differences were considered statistically significant at **p < 0.01 and *p < 0.05, unless otherwise stated, and are denoted by asterisks.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Jia, Q. & Horwitz, M. A. Live attenuated tularemia vaccines for protection against respiratory challenge with virulent F. tularensis subsp. tularensis. Front. Cell. Infect. Microbiol. 8, 154. https://doi.org/10.3389/fcimb.2018.00154 (2018).
Anthony, L. S., Ghadirian, E., Nestel, F. P. & Kongshavn, P. A. The requirement for gamma interferon in resistance of mice to experimental tularemia. Microb. Pathog. 7, 421–428 (1989).
Fortier, A. H., Polsinelli, T., Green, S. J. & Nacy, C. A. Activation of macrophages for destruction of Francisella tularensis: Identification of cytokines, effector cells, and effector molecules. Infect. Immun. 60, 817–825 (1992).
Leiby, D. A., Fortier, A. H., Crawford, R. M., Schreiber, R. D. & Nacy, C. A. In vivo modulation of the murine immune response to Francisella tularensis LVS by administration of anticytokine antibodies. Infect. Immun. 60, 84–89 (1992).
Elkins, K. L., Rhinehart-Jones, T. R., Culkin, S. J., Yee, D. & Winegar, R. K. Minimal requirements for murine resistance to infection with Francisella tularensis LVS. Infect. Immun. 64, 3288–3293 (1996).
Macela, A. Interaction of Francisella tularensis wirh the cells of mononuclear phagocytic system in the course of early stages of infection. Thesis, (Faculty of Science, Charles University, 1980).
Kovarova, H., Marcela, A. & Stulik, J. Macrophage activating factors produced in the course of murine tularemia: Effect on multiplication of microbes. Arch. Immunol. Ther. Exp. (Warsz.) 40, 183–190 (1992).
Hernychova, L. et al. Early consequences of macrophage-Francisella tularensis interaction under the influence of different genetic background in mice. Immunol. Lett. 57, 75–81 (1997).
Hrstka, R., Stulík, J. & Vojtesek, B. The role of MAPK signal pathways during Francisella tularensis LVS infection-induced apoptosis in murine macrophages. Microbes Infect. Inst. Pasteur 7, 619–625 (2005).
Hrstka, R. et al. Francisella tularensis strain LVS resides in MHC II-positive autophagic vacuoles in macrophages. Folia Microbiol. (Praha) 52, 631–636 (2007).
Steiner, D. J., Furuya, Y., Jordan, M. B. & Metzger, D. W. Protective role for macrophages in respiratory Francisella tularensis infection. Infect. Immun. 85, 00064–00117. https://doi.org/10.1128/IAI.00064-17 (2017).
Steiner, D. J., Furuya, Y. & Metzger, D. W. Detrimental influence of alveolar macrophages on protective humoral immunity during Francisella tularensis SchuS4 pulmonary infection. Infect. Immun. 86, 00787–00817. https://doi.org/10.1128/IAI.00787-17 (2018).
Sjöstedt, A., Conlan, J. W. & North, R. J. Neutrophils are critical for host defense against primary infection with the facultative intracellular bacterium Francisella tularensis in mice and participate in defense against reinfection. Infect. Immun. 62, 2779–2783 (1994).
Bosio, C. M. & Elkins, K. L. Susceptibility to secondary Francisella tularensis live vaccine strain infection in B-cell-deficient mice is associated with neutrophilia but not with defects in specific T-cell-mediated immunity. Infect. Immun. 69, 194–203 (2001).
KuoLee, R., Harris, G., Conlan, J. W. & Chen, W. Role of neutrophils and NADPH phagocyte oxidase in host defense against respiratory infection with virulent Francisella tularensis in mice. Microbes Infect. 13, 447–456 (2011).
Malik, M. et al. Matrix metalloproteinase 9 activity enhances host susceptibility to pulmonary infection with type A and B strains of Francisella tularensis. J. Immunol. Baltim. Md 1950(178), 1013–1020 (2007).
Bosio, C. M. & Dow, S. W. Francisella tularensis induces aberrant activation of pulmonary dendritic cells. J. Immunol. Baltim. Md 1950(175), 6792–6801 (2005).
Fabrik, I., Härtlova, A., Rehulka, P. & Stulik, J. Serving the new masters—dendritic cells as hosts for stealth intracellular bacteria. Cell. Microbiol. 15, 1473–1483 (2013).
Fabrik, I. et al. The early dendritic cell signaling induced by virulent Francisella tularensis strain occurs in phases and involves the activation of extracellular signal-regulated kinases (ERKs) and p38 In the later stage. Mol. Cell. Proteom. MCP 17, 81–94 (2018).
Claflin, J. L. & Larson, C. L. Infection-immunity in tularemia: Specificity of cellular immunity. Infect. Immun. 5, 311–318 (1972).
Kostiala, A. A., McGregor, D. D. & Logie, P. S. Tularaemia in the rat. I. The cellular basis on host resistance to infection. Immunology 28, 855–869 (1975).
Tärnvik, A. & Löfgren, S. Stimulation of human lymphocytes by a vaccine strain of Francisella tularensis. Infect. Immun. 12, 951–957 (1975).
Koskela, P. & Herva, E. Cell-mediated immunity against Francisella tularensis after natural infection. Scand. J. Infect. Dis. 12, 281–287 (1980).
Sjöstedt, A., Sandström, G., Tärnvik, A. & Jaurin, B. Molecular cloning and expression of a T-cell stimulating membrane protein of Francisella tularensis. Microb. Pathog. 6, 403–414 (1989).
Surcel, H. M., Ilonen, J., Poikonen, K. & Herva, E. Francisella tularensis-specific T-cell clones are human leukocyte antigen class II restricted, secrete interleukin-2 and gamma interferon, and induce immunoglobulin production. Infect. Immun. 57, 2906–2908 (1989).
Surcel, H. M., Tapaninaho, S. & Herva, E. Cytotoxic CD4+ T cells specific for Francisella tularensis. Clin. Exp. Immunol. 83, 112–115 (1991).
Yee, D., Rhinehart-Jones, T. R. & Elkins, K. L. Loss of either CD4+ or CD8+ T cells does not affect the magnitude of protective immunity to an intracellular pathogen, Francisella tularensis strain LVS. J. Immunol. Baltim. Md 1950(157), 5042–5048 (1996).
Poquet, Y. et al. Expansion of Vgamma9 Vdelta2 T cells is triggered by Francisella tularensis-derived phosphoantigens in tularemia but not after tularemia vaccination. Infect. Immun. 66, 2107–2114 (1998).
Henry, T. et al. Type I IFN signaling constrains IL-17A/F secretion by gammadelta T cells during bacterial infections. J. Immunol. Baltim. Md 1950(184), 3755–3767 (2010).
Crane, D. D., Scott, D. P. & Bosio, C. M. Generation of a convalescent model of virulent Francisella tularensis infection for assessment of host requirements for survival of tularemia. PLoS ONE 7, 33349. https://doi.org/10.1371/journal.pone.0033349 (2012).
Cowley, S. C. et al. CD4-CD8- T cells control intracellular bacterial infections both in vitro and in vivo. J. Exp. Med. 202, 309–319 (2005).
Tärnvik, A. & Holm, S. E. Stimulation of subpopulations of human lymphocytes by a vaccine strain of Francisella tularensis. Infect. Immun. 20, 698–704 (1978).
Culkin, S. J., Rhinehart-Jones, T. & Elkins, K. L. A novel role for B cells in early protective immunity to an intracellular pathogen, Francisella tularensis strain LVS. J. Immunol. Baltim. Md 1950(158), 3277–3284 (1997).
Elkins, K. L., Bosio, C. M. & Rhinehart-Jones, T. R. Importance of B cells, but not specific antibodies, in primary and secondary protective immunity to the intracellular bacterium Francisella tularensis live vaccine strain. Infect. Immun. 67, 6002–6007 (1999).
Chou, A. Y., Kennett, N. J., Melillo, A. A. & Elkins, K. L. Murine survival of infection with Francisella novicida and protection against secondary challenge is critically dependent on B lymphocytes. Microbes Infect. 19, 91–100 (2017).
Krocova, Z. et al. Interaction of B cells with intracellular pathogen Francisella tularensis. Microb. Pathog. 45, 79–85 (2008).
Plzakova, L., Krocova, Z., Kubelkova, K. & Macela, A. Entry of Francisella tularensis into murine B cells: The role of B cell receptors and complement receptors. PLoS ONE 10, 0132571. https://doi.org/10.1371/journal.pone.0132571 (2015).
Plzakova, L. et al. B cell subsets are activated and produce cytokines during early phases of Francisella tularensis LVS infection. Microb. Pathog. 75, 49–58 (2014).
Drabick, J. J., Narayanan, R. B., Williams, J. C., Leduc, J. W. & Nacy, C. A. Passive protection of mice against lethal Francisella tularensis (live tularemia vaccine strain) infection by the sera of human recipients of the live tularemia vaccine. Am. J. Med. Sci. 308, 83–87 (1994).
Stenmark, S., Lindgren, H., Tärnvik, A. & Sjöstedt, A. Specific antibodies contribute to the host protection against strains of Francisella tularensis subspecies holarctica. Microb. Pathog. 35, 73–80 (2003).
Kirimanjeswara, G. S., Golden, J. M., Bakshi, C. S. & Metzger, D. W. Prophylactic and therapeutic use of antibodies for protection against respiratory infection with Francisella tularensis. J. Immunol. Baltim. Md 1950(179), 532–539 (2007).
Klimpel, G. R. et al. Levofloxacin rescues mice from lethal intra-nasal infections with virulent Francisella tularensis and induces immunity and production of protective antibody. Vaccine 26, 6874–6882. https://doi.org/10.1016/j.vaccine.2008.09.077 (2008).
Sebastian, S. et al. Cellular and humoral immunity are synergistic in protection against types A and B Francisella tularensis. Vaccine 27, 597–605 (2009).
Mara-Koosham, G., Hutt, J. A., Lyons, C. R. & Wu, T. H. Antibodies contribute to effective vaccination against respiratory infection by type A Francisella tularensis strains. Infect. Immun. 79, 1770–1778 (2011).
Kubelkova, K. et al. Specific antibodies protect gamma-irradiated mice against Francisella tularensis infection. Microb. Pathog. 53, 259–268 (2012).
Krocova, Z., Plzakova, L., Benuchova, M., Macela, A. & Kubelkova, K. Early cellular responses of germ-free and specific-pathogen-free mice to Francisella tularensis infection. Microb. Pathog. 123, 314–322 (2018).
Lamousé-Smith, E., Tzeng, A. & Starnbach, M. The intestinal flora is required to support antibody responses to systemic immunization in infant and germ free mice. PLoS ONE 6, 27662. https://doi.org/10.1371/journal.pone.0027662 (2011).
Kubelkova, K. et al. Gnotobiotic mouse model’s contribution to understanding host-pathogen interactions. Cell. Mol. Life Sci. CMLS 73, 3961–3969 (2016).
Zarrella, T. M. et al. Host-adaptation of Francisella tularensis alters the bacterium’s surface-carbohydrates to hinder effectors of innate and adaptive immunity. PLoS ONE 6, 22. https://doi.org/10.1371/journal.pone.0022335 (2011).
Wallqvist, A. et al. Using host-pathogen protein interactions to identify and characterize Francisella tularensis virulence factors. BMC Genom. 16, 1106. https://doi.org/10.1186/s12864-015-2351-1 (2015).
Holland, K. et al. Differential growth of francisella tularensis, which alters expression of virulence factors, dominant antigens, and surface-carbohydrate synthases, governs the apparent virulence of Ft SchuS4 to immunized animals. Front. Microbiol. 8, 1158. https://doi.org/10.3389/fmicb.2017.01158 (2017).
Köppen, K. et al. Screen for fitness and virulence factors of Francisella sp. strain W12–1067 using amoebae. Int. J. Med. Microbiol. 309, 151341. https://doi.org/10.1016/j.ijmm.2019.151341 (2019).
Spidlova, P., Stojkova, P., Sjöstedt, A. & Stulik, J. Control of Francisella tularensis virulence at gene level: Network of transcription factors. Microorganisms 8, 1622. https://doi.org/10.3390/microorganisms8101622 (2020).
Syrjälä, H., Herva, E., Ilonen, J., Saukkonen, K. & Salminen, A. A whole-blood lymphocyte stimulation test for the diagnosis of human tularemia. J. Infect. Dis. 150, 912–915 (1984).
Karttunen, R., Surcel, H. M., Andersson, G., Ekre, H. P. & Herva, E. Francisella tularensis-induced in vitro gamma interferon, tumor necrosis factor alpha, and interleukin 2 responses appear within 2 weeks of tularemia vaccination in human beings. J. Clin. Microbiol. 29, 753–756 (1991).
Anthony, L. S., Skamene, E. & Kongshavn, P. A. Influence of genetic background on host resistance to experimental murine tularemia. Infect. Immun. 56, 2089–2093 (1988).
Macela, A. et al. The immune response against Francisella tularensis live vaccine strain in Lps(n) and Lps(d) mice. FEMS Immunol. Med. Microbiol. 13, 235–238 (1996).
Kovárová, H., Hernychová, L., Hajdúch, M., Sírová, M. & Macela, A. Influence of the bcg locus on natural resistance to primary infection with the facultative intracellular bacterium Francisella tularensis in mice. Infect. Immun. 68, 1480–1484 (2000).
López, M. C., Duckett, N. S., Baron, S. D. & Metzger, D. W. Early activation of NK cells after lung infection with the intracellular bacterium Francisella tularensis LVS. Cell. Immunol. 232, 75–85 (2004).
Gosselin, E. J., Gosselin, D. R. & Lotz, S. A. Natural killer and CD8 T cells dominate the response by human peripheral blood mononuclear cells to inactivated Francisella tularensis live vaccine strain. Hum. Immunol. 66, 1039–1049 (2005).
Rumyantsev, S. Constitutional and non-specific immunity to infection. Revue scientifique et technique (International Office of Epizootics) 17, 26–42 (1998).
Elkins, K. L., MacIntyre, A. T. & Rhinehart-Jones, T. R. Nonspecific early protective immunity in Francisella and Listeria infections can be dependent on lymphocytes. Infect. Immun. 66, 3467–3469 (1998).
Fuller, C. L. et al. Dominance of human innate immune responses in primary Francisella tularensis live vaccine strain vaccination. J. Allergy Clin. Immunol. 117, 1186–1188 (2006).
Fuller, C. L. et al. Transcriptome analysis of human immune responses following live vaccine strain (LVS) Francisella tularensis vaccination. Mol. Immunol. 44, 3173–3184 (2007).
Seibert, K., Pollard, M. & Nordin, A. Some aspects of humoral immunity in germ-free and conventional SJL-J mice in relation to age and pathology. Cancer Res. 34, 1707–1719 (1974).
Ohwaki, M., Yasutake, N., Yasui, H. & Ogura, R. A comparative study on the humoral immune responses in germ-free and conventional mice. Immunology 32, 43–48 (1977).
Taniguchi, T. et al. Difference in antibody production to heterologius erythrocytes in conventional, specific-pathogen-free (SPF), Germfree and antigen-free mice. Microbiol. Immunol. 22, 793–802 (1978).
Maddur, M. S. et al. Natural antibodies: From first-line defense against pathogens to perpetual immune homeostasis. Clin. Rev. Allergy Immunol. 58, 213–228. https://doi.org/10.1007/s12016-019-08746-9 (2020).
Cohen, I. R. & Cooke, A. Natural autoantibodies might prevent autoimmune disease. Immunol. Today 7, 363–364 (1986).
Baumgarth, N., Waffarn, E. E. & Nguyen, T. T. T. Natural and induced B-1 cell immunity to infections raises questions of nature versus nurture. Ann. N. Y. Acad. Sci. 1362, 188–199 (2015).
Bos, N. A., Meeuwsen, C. G., Wostmann, B. S., Pleasants, J. R. & Benner, R. The influence of exogenous antigenic stimulation on the specificity repertoire of background immunoglobulin-secreting cells of different isotypes. Cell. Immunol. 112, 371–380 (1988).
Ochsenbein, A. F. et al. Control of early viral and bacterial distribution and disease by natural antibodies. Science 286, 2156–2159 (1999).
Ochsenbein, A. F. & Zinkernagel, R. M. Natural antibodies and complement link innate and acquired immunity. Immunol. Today 21, 624–630 (2000).
Su, J. et al. Genome-wide identification of Francisella tularensis virulence determinants. Infect. Immun. 75, 3089–3101 (2007).
Elkins, K. L., Leiby, D. A., Winegar, R. K., Nacy, C. A. & Fortier, A. H. Rapid generation of specific protective immunity to Francisella tularensis. Infect. Immun. 60, 4571–4577 (1992).
Elkins, K. L., Rhinehart-Jones, T., Nacy, C. A., Winegar, R. K. & Fortier, A. H. T-cell-independent resistance to infection and generation of immunity to Francisella tularensis. Infect. Immun. 61, 823–829 (1993).
Baumgarth, N. et al. Innate and acquired humoral immunities to influenza virus are mediated by distinct arms of the immune system. Proc. Natl. Acad. Sci. U.S.A. 96, 2250–2255 (1999).
Choi, Y. S. & Baumgarth, N. Dual role for B-1a cells in immunity to influenza virus infection. J. Exp. Med. 205, 3053–3064 (2008).
Cole, L. E. et al. Antigen-specific B-1a antibodies induced by Francisella tularensis LPS provide long-term protection against F. tularensis LVS challenge. Proc. Natl. Acad. Sci. U.S.A. 106, 4343–4348 (2009).
Cole, L. E. et al. Role of TLR signaling in Francisella tularensis-LPS-induced, antibody-mediated protection against Francisella tularensis challenge. J. Leukoc. Biol. 90, 787–797 (2011).
Yang, Y. et al. Antigen-specific antibody responses in B-1a and their relationship to natural immunity. Proc. Natl. Acad. Sci. U. S. A. 109, 5382–5387 (2012).
Krocova, Z., Macela, A. & Kubelkova, K. Innate immune recognition. Implications for the interaction of Francisella tularensis with the host immune system. Front. Cell Infect. Microbiol. https://doi.org/10.3389/fcimb.2017.00446 (2017).
Chamberlain, R. Evaluation of live tularemia vaccine prepared in a chemically defined medium. Appl. Microbiol. 13, 232–235 (1965).
Havlasova, J. et al. Proteomic analysis of anti-Francisella tularensis LVS antibody response in murine model of tularemia. Proteomics 5, 2090–2103 (2005).
The UniProt Consortium. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 47, 506–515 (2019).
Jaime Huerta-Cepas, J. et al. eggNOG 5.0: A hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucl Acids Res. 47, 309–314 (2019).
Yu, N. Y. et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics (Oxford, England) 26, 1608–1615 (2010).
Almagro Armenteros, J. J. et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37, 420–423 (2019).
Rahman, O. et al. Methods for the bioinformatic identification of bacterial lipoproteins encoded in the genomes of Gram-positive bacteria. World J. Microbiol. Biotechnol. 24, 2377–2382 (2008).
Acknowledgements
The authors wish to thank the Institute of Microbiology, Laboratory of Gnotobiology in Novy Hradek, Czech Republic for providing GF Balb/c mice for our experiments. The authors also thank Jitka Zakova and Miroslav Fidransky for their excellent technical support during our work, Pavla Stojkova for valuable bioinformatic hints and analysis, and Marek Sinkora for the critical review of the manuscript. This study was supported by the grant VJ01030003 from the Ministry of Interior of the Czech Republic, grant 19-08294S obtained from the Czech Science Foundation, and by the Ministry of Defence of the Czech Republic—long-term organization development plan Medical Aspects of Weapons of Mass Destruction of the Faculty of Military Health Sciences, University of Defence.
Author information
Authors and Affiliations
Contributions
K.K. participated in conceptualization, investigation, decisions on methodology, data curation, and writing of the original draft, H.T. participated in decisions on methodology K.H. participated in decisions on methodology, writing of the original draft, P.J. participated in decisions on methodology M.A. participated in conceptualization, investigation, decisions on methodology, and writing of the original draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kubelkova, K., Hudcovic, T., Kozakova, H. et al. Early infection-induced natural antibody response. Sci Rep 11, 1541 (2021). https://doi.org/10.1038/s41598-021-81083-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81083-0
- Springer Nature Limited