Abstract
Sex chromosomes in males of most eutherian mammals share only a small homologous segment, the pseudoautosomal region (PAR), in which the formation of double-strand breaks (DSBs), pairing and crossing over must occur for correct meiotic segregation1,2. How cells ensure that recombination occurs in the PAR is unknown. Here we present a dynamic ultrastructure of the PAR and identify controlling cis- and trans-acting factors that make the PAR the hottest segment for DSB formation in the male mouse genome. Before break formation, multiple DSB-promoting factors hyperaccumulate in the PAR, its chromosome axes elongate and the sister chromatids separate. These processes are linked to heterochromatic mo-2 minisatellite arrays, and require MEI4 and ANKRD31 proteins but not the axis components REC8 or HORMAD1. We propose that the repetitive DNA sequence of the PAR confers unique chromatin and higher-order structures that are crucial for recombination. Chromosome synapsis triggers collapse of the elongated PAR structure and, notably, oocytes can be reprogrammed to exhibit spermatocyte-like levels of DSBs in the PAR simply by delaying or preventing synapsis. Thus, the sexually dimorphic behaviour of the PAR is in part a result of kinetic differences between the sexes in a race between the maturation of the PAR structure, formation of DSBs and completion of pairing and synapsis. Our findings establish a mechanistic paradigm for the recombination of sex chromosomes during meiosis.
Similar content being viewed by others
Main
During meiotic recombination in mice, DSBs must occur within the tiny (around 700 kb3,4) PAR2,3,4,5,6. Because on average one DSB forms per ten megabases, the PAR would be at risk of frequent recombination failure if it behaved similarly to a typical autosomal segment2. Consequently, DSBs and recombination are disproportionately frequent in the PAR2,6,7,8 (Supplementary Discussion). The mechanisms that promote these frequent DSBs are—in any species—not known.
DSBs arise concomitantly with linear axial structures that anchor the chromatin loops in which DSBs occur9,10. Axes begin to form during replication and become assembly sites for proteins that promote SPO11-mediated DSBs11,12,13. PAR chromatin in spermatocytes forms relatively short loops on a long axis2. However, only a low-resolution view of the structure of the PAR is presently available, and the controlling cis- and trans-acting factors are unknown. Moreover, it was unclear how spermatocytes—but not oocytes—make the PAR prone to such a high rate of recombination.
A distinctive PAR ultrastructure
The X and Y chromosomes usually pair late, with PARs paired in less than 20% of spermatocytes at late zygonema, when most autosomes are paired2,14. At this stage, unsynapsed PAR axes (defined by SYCP2 and SYCP3 immunostaining) appeared thickened relative to other unsynapsed axes and showed bright staining for HORMAD1 and HORMAD215 (Fig. 1a, Extended Data Fig. 1a, b). Moreover, the PAR was highly enriched for REC114, MEI4, MEI1 and IHO1—which are required for genome-wide DSB formation16,17,18,19—as well as ANKRD31, a partner of REC114 that is essential for DSBs in the PAR20,21.
a, Axis thickening (SYCP2 and SYCP3) and ANKRD31 accumulation on X and Y chromosome PARs (arrowheads) in late zygonema. The asterisk shows an autosomal ANKRD31 blob. Scale bars, 2 μm. b, Ultrastructure of the PAR before and after synapsis (montage of representative SIM images). Dashed lines indicate where chromosomes are cropped. c, Enrichment of ANKRD31 protein along split PAR axes in late zygonema (SIM image). d, Schematic showing the dynamic remodelling of the PAR loop–axis ensemble during prophase I. See measurements in Extended Data Fig. 3b. Scale bars, 1 μm (b–d).
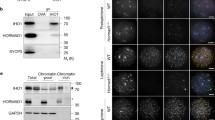
All five proteins (hereafter referred to as RMMAI) colocalized in several bright ‘blobs’ for most of prophase I (Fig. 1a, Extended Data Fig. 1c). Two blobs were on the PARs of X and Y chromosomes and others highlighted the ends of specific autosomes (Fig. 1a, Extended Data Fig. 1d). Similar blobs in previously published micrographs were not characterized16,17,19,22. The proteins also colocalized in smaller foci along unsynapsed axes16,17,19,20,21,22 (Extended Data Fig. 1c). Enrichment of RMMAI proteins at the PAR was already detectable at the pre-leptotene stage17,22 (Extended Data Fig. 1e), but not in spermatogonia (Extended Data Fig. 1f). Mass spectrometry of immunoprecipitates from testes also identified ZMYM3 and PTIP as ANKRD31-interacting proteins that were enriched at the PAR (Extended Data Fig. 1g–i, Supplementary Discussion).
Structured illumination microscopy (SIM) resolved the thickened PAR as two axial cores (Fig. 1b, Extended Data Fig. 2a, b) decorated with RMMAI proteins (Fig. 1c). The PAR axes were extended and separated in late zygonema before the synapsis of X and Y chromosomes, then collapsed during X–Y chromosome synapsis in early pachynema (Fig. 1b). Each axial core is a sister chromatid, with a ‘bubble’ from near the PAR boundary almost to the telomere (Extended Data Fig. 2c–h). This PAR structure is distinct from that seen at all other chromosome ends later in prophase I (Supplementary Discussion). Axis splitting and REC114 enrichment occurred independently of the formation of DSBs (Extended Data Fig. 2i).
Dynamic remodelling of the PAR structure
We investigated temporal patterns of axis differentiation, RMMAI composition and chromatin-loop configuration at the PAR using SIM and conventional microscopy (Fig. 1d, Extended Data Fig. 3a, b). The SYCP3-defined axis was long as soon as it became detectable in leptonema, and the fluorescence in situ hybridization (FISH) signal for the PAR boundary (PARb) was compact and remained so while the axis lengthened further through late zygonema, when the sister axes separated. Throughout, abundant ANKRD31 and REC114 signals stretched along the PAR axes, decorating the compact chromatin (Extended Data Fig. 3a, b). After synapsis, the axes shortened and the chromatin loops decompacted, with a concomitant dissociation of RMMAI proteins. A focus of the meiotic cohesin subunit REC8 was juxtaposed with ANKRD31 blobs at the pre-leptotene stage; REC8 was mostly restricted to the borders of the PAR as its axes elongated and split, and remained highly enriched on the short axis after RMMAI proteins disappeared (Extended Data Fig. 3a, b). Collapse of the loop–axis structure and dissociation of REC114 also occurred when the PAR underwent non-homologous synapsis in a Spo11−/− mutant (Extended Data Fig. 3c), showing that synapsis without recombination is sufficient for the reconfiguration of the PAR. DSB formation without synapsis may also be sufficient (Supplementary Discussion). These findings reveal a large-scale reconfiguration of the loop–axis structure and establish spatial and temporal correlations between the accumulation of RMMAI proteins at the PAR and the formation of a long axis associated with compact chromatin.
Heterochromatic mo-2 minisatellites
We deduced that specific DNA sequences might recruit RMMAI proteins because autosomal blobs also hybridized to the PARb probe (Extended Data Fig. 1d). This repetitive probe includes a tandem array of around 20 kb of a minisatellite (mo-2), with a 31-bp repeat23,24 (Fig. 2a). Clusters of mo-2 are also present at the non-centromeric ends of chromosomes 4, 9 and 1323,24 (Fig. 2a, b, Extended Data Fig. 4a, b). FISH with an mo-2 oligonucleotide probe showed that RMMAI blobs colocalize completely with mo-2 arrays (Fig. 2b, Extended Data Fig. 4c, d). Mo-2 arrays become enriched at the onset of meiosis for heterochromatic histone modifications (histone H3 trimethylated at K9 (H3K9me3) and H4K20me3) and proteins (heterochromatin protein 1β (HP1β), HP1γ and others), independent of DSB formation (Extended Data Fig. 5).
a, Left, self-alignment of the PARb FISH probe. The circled block is a 20-kb mo-2 cluster. Right, schematic showing the non-centromeric chromosome ends identified using a Basic Local Alignment Search Tool (BLAST) search with the mo-2 consensus sequence. b, Colocalization of REC114 blobs with the mo-2 oligonucleotide FISH signal (zygotene spermatocyte). Scale bar, 2 μm. c, Enrichment of ANKRD31 and RPA2 at the PAR correlates with mo-2 copy number. Top, late-zygotene spermatocyte from an F1 hybrid from crosses of B6 × MSM. Scale bar, 1 μm. Bottom, PAR-associated signals (AU, arbitrary units) on B6-derived (XB6) and MSM-derived (YMSM) chromosomes from the indicated number of spermatocytes (n). Red lines indicate mean ± s.d. Differences between X and Y chromosome PAR intensities are significant both for proteins and for mo-2 FISH (P < 10−6, paired t-test; for exact P values, see the associated Source Data).
To test whether mo-2 arrays are cis-acting determinants of RMMAI recruitment, we exploited the fact that Mus musculus molossinus has an mo-2 copy number substantially lower than that of other subspecies of Mus musculus24. Mice of the MSM/MsJ (MSM) strain showed a lower hybridization signal with the mo-2 FISH probe than did mice of the C57BL/6J (B6) strain, as well as a lower REC114 intensity in blobs (Extended Data Fig. 4e).
To avoid confounding strain effects, we examined spermatocytes of F1 hybrid mice (Fig. 2c, Extended Data Fig. 4f, g). Less ANKRD31 accumulated on the MSM PARs: in offspring from B6 mothers and MSM fathers, the YMSM PAR had eightfold less ANKRD31 than the XB6 PAR (Fig. 2c, Extended Data Fig. 4g), and in the reciprocal cross the XMSM PAR had 6.5-fold less ANKRD31 than the YB6 PAR (Extended Data Fig. 4f, g). The relative levels of ANKRD31 matched those that were observed in mo-2 FISH. Nevertheless, MSM PARs supported a similar efficiency and timing of sex-chromosome pairing to the B6 PARs (Extended Data Fig. 4h)—which is not surprising, as MSM mice are fertile. Notably, the single-strand DNA (ssDNA)-binding protein RPA2 was present at lower intensity on MSM PARs (Fig. 2c, Extended Data Fig. 4f).
Determinants that act in trans
To identify factors that are important for PAR behaviour, we eliminated different RMMAI proteins (MEI4 and ANKRD31) and axis proteins (HORMAD1 and REC8)16,20,25,26. Requirements for RMMAI blobs overlap with but are distinct from those for smaller RMMAI foci—for which HORMAD1 is key and MEI4 even more so, but ANKRD31 contributes only partially17,20,22 (Fig. 3a). HORMAD1 and REC8 were dispensable for RMMAI assembly in mo-2 regions, and for the elongation of the PAR axis, splitting of sister axes and formation of short loops (that is, compact mo-2 and REC114 signals) (Fig. 3a–c, Extended Data Fig. 6a, b). The distal PAR axes were separated in Rec8−/− mutants (Fig. 3c, Extended Data Fig. 6c), demonstrating that REC8 is essential for cohesion at the PAR end.
a, Top, quantification of REC114, ANKRD31, MEI4 and IHO1 foci along unsynapsed axes in wild-type and mutant leptotene or early-zygotene spermatocytes. Error bars are mean ± s.d. Comparisons to the wild type are indicated (*P < 0.02, **P ≤ 10−7, NS, not significant (P > 0.05), two-sided Student’s t-test; for exact P values, see the associated Source Data). Bottom, representative micrographs of REC114 staining; other proteins are shown in Extended Data Fig. 6a. Arrowheads indicate mo-2 associated blobs. Presence (+) or absence (–) of blobs is indicated below. Scale bars, 2 μm. b, Genetic requirements for PAR loop–axis organization (length of REC114 and mo-2 FISH signals along the PAR axis and axis-orthogonal extension of mo-2). Error bars are mean ± s.d. c, Representative SIM images of the loop–axis structure of the Y chromosome PAR in each mutant at late zygonema. Scale bar, 1 μm.
The smaller MEI4 and REC114 foci still formed in Ankrd31−/− mutants, but were fewer and weaker20 (Fig. 3a, Extended Data Fig. 6a, d, e). In mo-2 regions, by contrast, RMMAI proteins did not accumulate detectably in Mei4−/− and Ankrd31−/− mutants (Fig. 3a, Extended Data Fig. 6a, b). ANKRD31 was dispensable for enrichment of heterochromatin factors (Extended Data Fig. 6f). REC114—though not IHO1—is also essential for RMMAI blobs21. The normal ultrastructure of the PAR was absent in Mei4−/− and Ankrd31−/− mutants: axes were short with no sign of splitting and mo-2 was decompacted (Fig. 3b, c, Extended Data Fig. 6b). We conclude that RMMAI blobs at the PAR share genetic requirements with autosomal mo-2 blobs, and that the presence of blobs correlates with normal structural differentiation of the PAR.
Axis remodelling and mo-2
If mo-2 arrays are cis-acting determinants of a high level of RMMAI recruitment, which in turn governs the structural dynamics of the PAR, then autosomal mo-2 regions should also form PAR-like structures. Consistent with this, the distal end of chromosome 9 underwent splitting in spermatocytes in which this region was late to synapse (Fig. 4a) and showed a PAR-like pattern of extended axes and compact chromatin dependent on Ankrd31 (Extended Data Fig. 7a). Thus, mo-2 (and/or linked elements) may be sufficient for both RMMAI recruitment and axis remodelling. We observed less axis remodelling for MSM PARs than B6 PARs (Extended Data Fig. 7b), which reinforces the correlation between mo-2 copy number, RMMAI levels and PAR ultrastructure.
a, The mo-2 region of chromosome 9 undergoes axis elongation and splitting similar to PARs (SIM image of a wild-type zygotene spermatocyte). Scale bar, 1 μm. b, ANKRD31 is required for high levels of DSB formation in mo-2 regions and for X–Y chromosome pairing. Immunolabelling with FISH (immuno-FISH) for RPA2 and mo-2 was used to detect DSBs. Immuno-FISH images for other mutants are shown in Extended Data Fig. 7c. Scale bar, 1 μm. c, PAR-like DSB formation near autosomal mo-2 regions. SSDS coverage7,20 is shown for the Y chromosome PAR (left) and the mo-2-adjacent region of chromosome 9 (right). The positions of mo-2 repeats are shown below. Examples for other chromosomes are shown in Extended Data Fig. 8a. d, Early pachytene XY oocyte showing bright RPA2 foci in the PAR (arrowheads). Scale bars, 2 μm (main image); 1 μm (inset).
DSB formation in spermatocytes
We hypothesized that RMMAI recruitment and axis remodelling create an environment that is conducive to a high level of DSB formation. This idea predicts that mutations should affect all of these processes (RMMAI recruitment, axis remodelling and DSB formation) coordinately and that PAR-like DSB formation should occur in autosomal mo-2 regions. We counted axial RPA2 foci as a proxy for the global number of DSBs and assessed the overlap of mo-2 with RPA2 (Fig. 4b, Extended Data Fig. 7c–f).
In wild-type zygotene spermatocytes, RPA2 foci overlapped with 35% of the mo-2 regions of each cell on average, increasing to 70% at pachynema (Extended Data Fig. 7e). Similar to the PAR2, autosomal mo-2 regions often acquired DSBs late (Extended Data Fig. 7g). By contrast, Ankrd31−/− mutants showed a substantial reduction in the overlap of RPA2 foci with mo-2, and X and Y chromosomes were paired in only 6% of mid-pachytene spermatocytes (Fig. 4b, Extended Data Fig. 7e, h). This is distinct from autosomes: global RPA2 foci were only modestly reduced in Ankrd31−/− cells (Extended Data Fig. 7d), and in most cells all autosomes were paired and synapsed20,21. (Ankrd31−/− mutants form fewer RPA2 foci at leptonema and early zygonema, but normal numbers thereafter20,21.)
Rec8 deficiency did not reduce the formation of RPA2 foci at mo-2 regions or more globally relative to a synapsis-deficient control (Syce1−/−) (Extended Data Fig. 7c–e). However, X–Y chromosome pairing was reduced (Extended Data Fig. 7h), presumably because REC8 promotes inter-homologue recombination27. Hormad1−/− spermatocytes had comparable or higher frequencies of mo-2-overlapping RPA2 foci and X–Y chromosome pairing to the Syce1−/− control (Extended Data Fig. 7e, h). The high frequency of RPA2 foci at mo-2 regions was notable, given the global reduction in the number of RPA2 foci (Extended Data Fig. 7d, f) and DSBs28, but is consistent with the dispensability of HORMAD1 both for the recruitment of RMMAI proteins to mo-2 and for the ultrastructure of the PAR (Fig. 3a–c).
These findings show that RMMAI recruitment and axis remodelling are tightly correlated with a high frequency of DSB formation. Further strengthening this correlation, we noted above that MSM PARs exhibit a lower RPA2 intensity than B6 PARs (Fig. 2c)—perhaps reflecting a lesser tendency to make multiple DSBs. Indeed, multiple PAR RPA2 foci were resolved more frequently in B6 than in MSM mice (Extended Data Fig. 7i, j).
We used maps of ssDNA bound by the strand-exchange protein DMC1 (ssDNA sequencing; SSDS)7,29,30 to test more directly whether PAR-like DSB formation occurs in autosomal mo-2 regions (that is, dependent on ANKRD31 but largely independent of the histone methyltransferase PRDM9)7,20,21 (Fig. 4c, Extended Data Fig. 8a). We found that the region that encompasses the chromosome 9 mo-2 cluster showed an accumulation of SSDS reads that was substantially reduced in Ankrd31−/− but not in Prdm9−/− mutants. A modest ANKRD31-dependent, PRDM9-independent peak was also observed near the mo-2 cluster on chromosome 13 (Extended Data Fig. 8a). Thus, autosomal mo-2 regions not only accumulate PAR-like levels of RMMAI proteins and undergo PAR-like axis remodelling in spermatocytes, but also frequently form DSBs in a PAR-like manner.
Mo-2 regions in oocytes
In females, recombination between the two X chromosomes is not restricted to the PAR, so oocytes do not require DSBs in the PAR as with spermatocytes31. We therefore asked whether the PAR undergoes spermatocyte-like structural changes in oocytes. RMMAI proteins robustly accumulated at the PAR and in autosomal mo-2 regions from leptonema to pachynema (Extended Data Fig. 9a), consistent with studies of MEI4 and ANKRD3116,21. Oocytes also had an extended PAR axis and a compact PARb FISH signal from leptonema to zygonema and transitioned to a shorter axis and a more extended PARb signal in pachynema, with a loss of REC114 signal upon synapsis (Extended Data Fig. 9b). Heterochromatin factors were also enriched (Extended Data Fig. 9c). However, in contrast to spermatocytes, we did not detect a thickening or splitting of the PAR axis, or the accumulation of REC8 (Extended Data Fig. 9d)—even in the absence of synapsis in Syce1−/− mutants (Extended Data Fig. 9e). Moreover, similar to the PAR31, autosomal mo-2 regions showed little enrichment for SSDS signal in wild-type ovaries (Extended Data Fig. 8b, c).
A low SSDS signal despite the enrichment of RMMAI proteins and the presence of long axes could indicate that oocytes lack a critical factor (or more than one factor) that promotes DSBs in the PAR of spermatocytes. Alternatively, oocyte PARs may not realize their full DSB potential because of negative feedback tied to homologue engagement32,33; perhaps synapsis that is initiated elsewhere on the X chromosome often spreads into the PAR and disrupts its ultrastructure before DSBs can form. To investigate this idea, we tested the effects of delaying or blocking PAR synapsis using sex-reversed XY females34 and Syce1−/− mutants.
XY oocytes pair and synapse their PARs relatively late—only 28% of late-zygotene cells had X and Y chromosomes that were paired and/or synapsed (25 of 90 cells from two mice), increasing to 66% at pachynema (115 of 174 cells). This late pairing and synapsis is reminiscent of spermatocytes, but appears to be less efficient. Most pachytene XY oocytes that synapsed their PARs had a PAR-associated RPA2 focus, which occurred at twice the frequency and with higher immunofluorescence intensity than in XX oocytes (Fig. 4d, Extended Data Fig. 9f). RPA2 foci were also seen on most PARs that failed to synapse (Extended Data Fig. 9g). By contrast, chromosomes 9 and 13 showed a lower frequency and intensity of RPA2 foci, comparable to those in XX PARs and not differing between XY and XX oocytes (Extended Data Fig. 9f).
These findings suggest that delayed PAR synapsis allows oocytes to form DSBs more efficiently. In support of this conclusion, the absence of synapsis in Syce1−/− oocytes was accompanied by an increase in both the frequency and the intensity of RPA2 foci at PARs and autosomal mo-2 regions alike (Extended Data Fig. 9h). Our results do not exclude the possibility of differences between the trans-acting factors of spermatocytes and oocytes, but we infer that the ability to manifest a high level of DSB formation depends substantially on the result of a race between the formation of DSBs and the completion of synapsis (Supplementary Discussion).
Discussion
We have shown here that the PAR in male mice undergoes a notable rearrangement of loop–axis structure before DSBs form, and that this rearrangement involves the recruitment of RMMAI proteins, dynamic axis elongation and splitting of sister chromatid axes (Extended Data Fig. 10). Most of these behaviours also occur in oocytes, which can also support a high rate of DSB formation in the PAR if synapsis is delayed. The mo-2 array may be a key cis-acting determinant, and RMMAI proteins are crucial trans-acting determinants. Although the function of sister axis splitting is unclear (Supplementary Discussion), the full suite of PAR behaviours appears essential for the pairing, recombination and segregation of heteromorphic sex chromosomes.
Budding yeast also uses the robust recruitment of Rec114 and Mer2 (the IHO1 orthologue) to ensure that its smallest chromosomes incur DSBs35. Thus, such preferential recruitment is an evolutionarily recurrent strategy for mitigating the risk of recombination failure when the length of chromosomal homology is limited.
The hyperaccumulation of RMMAI proteins may reflect the binding of one or more of these proteins to an mo-2-associated chromatin structure and/or direct binding to mo-2 repeats or another tightly linked DNA element. We note that the repetitive mo-2 array imposes a risk of unequal exchange23,36. Thus, paradoxically, the DNA structure of the PAR stabilizes the genome by supporting the segregation of sex chromosomes, but also promotes the rapid evolution of mammalian PARs4.
Methods
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Mice
Mice were maintained and killed under USA regulatory standards and experiments were approved by the Memorial Sloan Kettering Cancer Center (MSKCC) Institutional Animal Care and Use Committee (IACUC, protocol number 01-03-007). Mice were fed regular rodent chow with ad libitum access to food and water. The Ankrd31 knockout allele (Ankrd31em1Sky) is a single-base-insertion mutation (+A) in exon 3; its generation and phenotypic characterization have previously been described20. Mice with the Mei4 knockout allele16 were provided by B. de Massy. All other mouse strains were purchased from the Jackson Laboratory: C57BL/6J (stock no. 00664), MSM/MsJ (stock no. 003719), B6N(Cg)-Syce1tm1b(KOMP)Wtsi/2J (stock no. 026719), B6;129S7-Hormad1tm1Rajk/Mmjax (stock no. 41469-JAX), B6;129S4-Rec8mei8/JcsMmjax (stock no. 34762-JAX) and B6.Cg-Tg(Sry)2Ei Srydl1Rlb/ArnoJ (stock no. 010905). Mice were genotyped using Direct Tail lysis buffer (Viagen) following the manufacturer’s instructions.
B6.Cg-Tg(Sry)2Ei Srydl1Rlb/ArnoJ males have a Y chromosome with a deletion of the sex-determining Sry gene and also have an Sry transgene integrated on an autosome. When these males are crossed with C57BL/6J females, those XY and XX mice that do not inherit the Sry transgene develop as females.
Generation of REC8 and REC114 antibodies
To produce antibodies against REC8, a fragment of the mouse Rec8 gene encoding amino acids 36 to 253 (NCBI reference sequence: NP_001347318.1) was cloned into a pGEX-4T-2 vector. The resulting fusion of the REC8 fragment fused to glutathione S-transferase (GST) was expressed in Escherichia coli, affinity-purified on glutathione sepharose 4B and cleaved with Precision protease. Antibodies were raised in rabbits by Covance against the purified recombinant REC8 fragment, and antibodies were affinity-purified using GST-REC836-253 that had been immobilized on glutathione sepharose by cross-linking with dimethyl pimelimidate; bound antibodies were eluted with 0.1 M glycine, pH 2.5. Purified antibodies were tested in western blots of testis extracts and specificity was validated by immunostaining of spread meiotic chromosomes from wild-type and Rec8−/− mice.
To produce antibodies against REC114, a fragment of the mouse Rec114 gene encoding a truncated polypeptide lacking the N-terminal 110 amino acids (NCBI reference sequence: NP_082874.1) was cloned into a pET-19b expression vector. The resulting His6-tagged REC114(111–259) fragment was insoluble when expressed in E. coli, so the recombinant protein was solubilized and affinity-purified on Ni-NTA resin in the presence of 8 M urea. Eluted protein was dialysed against 100 mM NaH2PO4, 10 mM Tris-HCl, 6 M urea, pH 7.3 and used to immunize rabbits (Covance). Antibodies were affinity-purified against purified recombinant His6–REC114(111–259) protein immobilized on cyanogen-bromide-activated sepharose and eluted in 0.2 M glycine, pH 2.5. The affinity-purified antibodies were used in a previous study17 that reported detection of a band of appropriate molecular weight in western blots of testis extracts. Subsequent analysis showed that this band is also present in extracts of Rec114−/− testes, and thus is non-specific (C. Brun and B. de Massy, personal communication). However, the initial study17 also reported detection of immunostaining foci on spread meiotic chromosomes similar to findings reported here and in another previous study20. This immunostaining signal is absent from chromosome spreads prepared from Rec114−/− mutant mice (C. Brun and B. de Massy, personal communication). Moreover, this immunostaining signal is indistinguishable from that reported using independently generated and validated anti-REC114 antibodies19. We conclude that our anti-REC114 antibodies are highly specific for the cognate antigen when used for immunostaining of meiotic chromosome spreads.
Chromosome spreads
Testes were dissected and deposited after removal of the tunica albuginea in 1 × PBS, pH 7.4. Seminiferous tubules were minced using forceps to form a cell suspension. The cell suspension was filtered through a 70-μm cell strainer into a 15-ml Falcon tube pre-coated with 3% (w/v) BSA, and was centrifuged at 200g for 5 min. The cell pellet was resuspended in 12 ml of 1 × PBS for an additional centrifugation step at 200g for 5 min and the pellet was resuspended in 1 ml hypotonic buffer containing 17 mM sodium citrate, 50 mM sucrose, 30 mM Tris-HCl, pH 8, 5 mM EDTA, pH 8, 0.5 mM dithiothreitol (DTT) and 10 μl of 100 × Halt protease inhibitor cocktail (Thermo Fisher Scientific), and incubated for 8 min. Next, 9 ml of 1 × PBS was added and the cell suspension was centrifuged at 200g for 5 min. The cell pellet was resuspended in 100 mM sucrose pH 8 to obtain a slightly turbid cell suspension, and incubated for 10 min. Superfrost glass slides were divided into two squares using an ImmEdge hydrophobic pen (Vector Labs), then 110 μl of 1% paraformaldehyde (PFA) (freshly dissolved in presence of NaOH at 65 °C, 0.15% Triton, pH 9.3, cleared through a 0.22-μm filter) and 30 μl of cell suspension were added per square and swirled three times for homogenization, and the slides were placed horizontally in a closed humid chamber for 2 h. The humid chamber was opened for 1 h to allow almost complete drying of the cell suspension. Slides were washed in a Coplin jar 2 × 5 min in 1 × PBS on a shaker, and 2 min with 0.4% Photo-Flo 200 solution (Kodak), air-dried and stored in aluminium foil at −80 °C.
Ovaries were extracted from mice 14.5–18.5 days post-coitum (dpc), and collected in 1 × PBS, pH 7.4. After a 15-min incubation in hypotonic buffer, the ovaries were placed on a slide containing 30 μl of 100 mM sucrose, pH 8, and dissected with forceps to form a cell suspension. The remaining tissues were removed, 110 μl of 1% PFA–0.15% Triton was added and the slides were gently swirled for homogenization, before incubation in a humid chamber as described above for spermatocyte chromosome spreads.
Immunostaining
Slides of meiotic chromosome spreads were blocked for 30 min at room temperature horizontally in a humid chamber with an excess of blocking buffer containing 1 × PBS, pH 7.4 with 0.05% Tween-20, 7.5% (v/v) donkey serum, 0.5 mM EDTA, pH 8.0 and 0.05% (w/v) sodium azide, and cleared by centrifugation at 13,000 rpm for 15 min in an Eppendorf microcentrifuge (with rotor FA-45-24-11). Slides were incubated with primary antibody overnight in a humid chamber at 4 °C, or for at least 3 h at room temperature. Slides were washed 3 × 5 min in 1 × PBS, 0.05% Tween-20, then blocked for 10 min and incubated with secondary antibody for 1–2 h at 37 °C in a humid chamber. Slides were washed 3 × 5 min in the dark on a shaker with 1 × PBS and 0.05% Tween-20, then rinsed in H2O, and mounted before air-drying with Vectashield (Vector Labs). Antibody dilutions were centrifuged at 13,000 rpm for at least 5 min before use. The primary antibodies used were rabbit and guinea pig anti-ANKRD3120 (1:200 dilution), rabbit anti-HORMAD2 (Santa Cruz, sc-82192, 1:50), guinea pig anti-HORMAD2 (1:200) and guinea pig anti-IHO1 (1:200) (gifts from A. Tóth), goat anti-MEI1 (Santa Cruz, sc-86732, 1:50), rabbit anti-MEI4 (gift from B. de Massy, 1:200), rabbit anti-REC8 (this study, 1:100), rabbit anti-REC114 (this study, 1:200), rabbit anti-RPA2 (Santa Cruz, sc-28709, 1:50), goat anti-SYCP1 (Santa Cruz, sc-20837, 1:50), rabbit anti-SYCP2 (Atlas Antibodies, HPA062401, 1:100), mouse anti-SYCP3 (Santa Cruz, sc-74569, 1:100), goat anti-SYCP3 (Santa Cruz, sc-20845, 1:50), rabbit anti-TRF1 (Alpha Diagnostic, TRF12-S, 1:100), rabbit anti-H4K20me3 (Abcam, ab9053, 1:200), rabbit anti-H3K9me3 (Abcam, ab8898, 1:200), mouse anti-macroH2A1.2 (Active Motif, 61428, 1:100), mouse anti-HP1γ (Millipore, MAB3450, 1:100), mouse anti-HP1β (Millipore, MAB3448, 1:100), rabbit anti-HP1β (Genetex, GTX106418, 1:100), rabbit anti-Mi2 (recognizes CHD3 and CHD4; Santa Cruz, sc-11378, 1:50), rabbit anti-ATRX (Santa Cruz, sc-15408, 1:50), mouse anti-DMRT1 (Santa Cruz, sc-377167, 1:50), rabbit anti-ZMYM3 (Abcam, ab19165, 1:300), rabbit anti-PAXIP1 (EMD Millipore, ABE1877, 1:300). The secondary antibodies used were CF405S anti-guinea pig (Biotium, 20356), CF405S anti-rabbit (Biotium, 20420), CF405S anti-mouse (Biotium, 20080), Alexa Fluor 488 donkey anti-mouse (Life Technologies, A21202), Alexa Fluor 488 donkey anti-rabbit (Life Technologies, A21206), Alexa Fluor 488 donkey anti-goat (Life Technologies, A11055), Alexa Fluor 488 donkey anti-guinea pig (Life Technologies, A11073), Alexa Fluor 568 donkey anti-mouse (Life Technologies, A10037), Alexa Fluor 568 donkey anti-rabbit (Life Technologies, A10042), Alexa Fluor 568 goat anti-guinea pig (Life Technologies, A11075), Alexa Fluor 594 donkey anti-mouse (Life Technologies, A21203), Alexa Fluor 594 donkey anti-rabbit (Life Technologies, A21207), Alexa Fluor 594 donkey anti-goat (Life Technologies, A11058), Alexa Fluor 647 donkey anti-rabbit (Abcam, ab150067), Alexa Fluor 647 donkey anti-goat (Abcam, ab150131), all at 1:250 dilution.
Immuno-FISH and DNA probe preparation
All steps were performed in the dark to prevent loss of fluorescence from prior immunostaining. After the last washing step in the immunostaining protocol, slides were placed horizontally in a humid chamber and the chromosome spreads were re-fixed with an excess of 2% (w/v) PFA in 1 × PBS (pH 9.3) for 10 min at room temperature. Slides were rinsed once in H2O, washed for 4 min in 1 × PBS, sequentially dehydrated with 70% (v/v) ethanol for 4 min, 90% ethanol for 4 min and 100% ethanol for 5 min, and air-dried vertically for 5–10 min. Next, 15 μl of hybridization mix was applied containing the DNA probe(s) in 70% (v/v) deionized formamide (Amresco), 10% (w/v) dextran sulfate, 2 × saline sodium citrate (SSC) buffer, 1 × Denhardt’s buffer, 10 mM EDTA, pH 8 and 10 mM Tris-HCl, pH 7.4. Cover glasses (22 × 22 mm) were applied and sealed with rubber cement (Weldwood contact cement), then the slides were denatured on a heat block for 7 min at 80 °C, followed by overnight incubation (for at least 14 h) at 37 °C. Cover glasses were carefully removed using a razor blade, slides were rinsed in 0.1 × SSC buffer, washed in 0.4 × SSC, 0.3% NP-40 for 5 min, washed in PBS–0.05% Tween-20 for 3 min, rinsed in H2O and mounted with Vectashield before air-drying.
To generate FISH probes, we used the nick translation kit from Abbott Molecular following the manufacturer’s instructions and using CF-dye-conjugated dUTP (Biotium), on bacterial artificial chromosome (BAC) DNA from the clones RP24-500I4 (maps to the region of the PAR boundary; PARb probe) CH25-592M6 (maps to the distal PAR, PARd probe), RP23-139J18, RP24-136G21 and CH36-200G6 (centromere-distal ends of chromosomes 4, 9 and 13, respectively). BAC clones were obtained from the BACPAC Resource Center (CHORI). Labelled DNA (500 ng) was precipitated during a 30-min incubation at −20 °C after adding 5 μl mouse Cot-1 DNA (Invitrogen), 0.5 volume of 7.5 M ammonium acetate and 2.5 volumes of cold 100% ethanol. After washing with 70% ethanol and air-drying in the dark, the pellet was dissolved in 15 μl hybridization buffer.
Mo-2 oligonucleotide probes were synthesized by Integrated DNA Technologies, with 6-FAM or TYE 665 fluorophores added to both 5′ and 3′ ends of the oligonucleotide. The DNA sequence was designed on the basis of the previously defined consensus sequence24, and the probe was used at a final concentration of 10 pmol μl−1 in hybridization buffer without Cot-1 DNA. The Y chromosome paint probe was purchased from ID Labs and used at 1:30 dilution in hybridization buffer without Cot-1 DNA.
EdU incorporation
Seminiferous tubules were incubated in Dulbecco’s modified Eagle’s medium (DMEM) with 10% FCS and 10 μM 5-ethynyl-3′-deoxyuridine (EdU) at 37 °C for 1 h for in vitro labelling. EdU incorporation was detected using the Click-iT EdU Alexa Fluor 647 imaging kit (Invitrogen) according to the manufacturer’s instructions.
Image acquisition
Images of spread spermatocytes were acquired on a Zeiss Axio Observer Z1 Marianas Workstation, equipped with an ORCA-Flash 4.0 camera and DAPI, CFP, FITC, TEXAS red and Cy5 filter sets, illuminated by an X-Cite 120 PC-Q light source, with either a 63×/1.4 NA oil immersion objective or a 100×/1.4 NA oil-immersion objective. Marianas Slidebook 5.0 (Intelligent Imaging Innovations) software was used for acquisition.
Structured illumination microscopy (3D-SIM) was performed at the Bio-Imaging Resource Center in Rockefeller University using an OMX Blaze 3D-SIM super-resolution microscope (Applied Precision), equipped with 405-nm, 488-nm and 568- nm lasers, and a 100×/1.40 NA UPLSAPO oil objective (Olympus). Image stacks of several-μm thickness were taken with 0.125-μm z-steps, and were reconstructed in Deltavision softWoRx 6.1.1 software with a Wiener filter of 0.002 using wavelength-specific experimentally determined optical transfer functions. Slides were prepared and stained as described above, except that chromosomes were spread only on the central portion of the slides, and the slides were mounted using 18 × 18-mm coverslips (Zeiss).
Image analysis
The 3D-SIM images are shown either as a z-stack using the sum slices function in Fiji, or as a unique slice. The X and/or Y chromosomes were cropped, rotated and further cropped for best display. For montage display, the X and Y chromosome images were positioned on a black background using Adobe Illustrator 2020 (v.24.1). In the instances in which the axes of the X and Y chromosomes were cropped, the area of cropping was labelled with a light grey dotted line. Loop and axis measurements, foci counts and fluorescence intensity quantification were only performed on images from conventional microscopy using the original, unmodified data.
To measure the colocalization between RMMAI proteins, we costained for SYCP3 and ANKRD31 along with MEI4, REC114 or IHO1, and manually counted the number of ANKRD31 foci that overlapped with SYCP3 and colocalized (or not) with MEI4, REC114 or IHO1. These counts were performed in 16 spermatocytes from leptonema to early or mid-zygonema.
To quantify the total number of RPA2, MEI4, REC114, ANKRD31 and IHO1 foci, single cells were manually cropped and analysed with semi-automated scripts in Fiji37 (v.2.0.0-rc-69/1.52p) as previously described in detail20. In brief, images were auto-thresholded on SYCP3 staining, which was used as a mask to use ‘Find Maxima’ to determine the number of foci. Images were manually inspected to determine that there were no obvious defects in determining SYCP3 axes, that no axes from neighbouring cells were counted, that no artefacts were present, and that no foci were missed by the script.
To test for colocalization between RPA2 and mo-2 FISH signals, we manually scored the percentage of mo-2 FISH signals colocalizing at least partly with RPA2. Depending on the progression of synapsis during prophase I, between eight and four discrete mo-2 FISH signals could be detected, corresponding to (with increasing signal intensity) chromosome 4, chromosome 13, chromosome 9 and the PAR (two signals for each when unpaired, or a single signal for each after homologous pairing and synapsis). Notably, the RPA2 focus was most often found in a slightly more centromere-proximal position compared to the bulk of mo-2 FISH signals, and therefore colocalized partly with mo-2 FISH signals. In the case of the PAR, this position corresponds closely to the region of the PAR boundary (PARb probe). A similar trend was observed on autosomal mo-2 clusters.
For estimates of chromatin extension, we measured the maximal axis-orthogonal distance between the FISH signal and the centre of the PAR axis, or the centromere-distal axis for chromosome 9 stained by SYCP3. In mutant mice defective for RMMAI protein recruitment in the mo-2 regions, the PAR axis was defined as the nearest SYCP3 segment adjacent to the telomeric SYCP3 signal.
For quantification of RPA2, ANKRD31, REC8 and mo-2 signal intensity in B6 × MSM and MSM × B6 F1 hybrids, late-zygotene spermatocytes with at least one RPA2 focus on the X or Y PAR were analysed. We used the elliptic selection tool in Fiji to define a region of interest around the largest signal in the PAR, and the same selection tool was then positioned on the other PAR axis for comparison. The fluorescence intensity was measured as the integrated density with background subtraction.
Prophase I sub-staging and identification of the PAR
Nuclei were staged according to the dynamic behaviour of the autosome and sex-chromosome axes during prophase I, using SYCP3 staining. Leptonema was defined as having short stretches of SYCP3 but no evidence of synapsis; early or mid-zygonema as having longer stretches of SYCP3 staining and some synapsis; and late zygonema as having fully assembled chromosome axes and substantial (more than 70%) synapsis. The X and Y chromosomes generally can be identified at this stage, and the PAR axis is distinguishable because it appears thicker than the centromeric end—particularly near the end of zygonema, when autosomes are almost fully synapsed. Early pachynema was defined as complete autosomal synapsis, whereas the X and Y chromosomes could display various configuration: i) unsynapsed, with thickened PAR axes; ii) engaged in PAR synapsis; and iii) synapsed in the PAR and non-homologously synapsed along the full (or nearly full) Y chromosome axis. Mid-pachynema was defined as showing bright signal from autosome axes, with desynapsing X and Y axes remaining synapsed only in the PAR and a short PAR axis. During this stage, the autosomes and the non-PAR X and Y chromosome axes are initially short and thick, and progressively become longer and thinner. Late pachynema was defined as brighter autosome axes with a characteristic thickening of all autosome ends. The X and Y chromosome non-PAR axes are then long and thin and show excrescence of axial elements. Diplonema was defined as brighter axes and desynapsing autosome, associated with prominent thickening of the autosome ends, particularly the centromeric ends. In early diplonema, the non-PAR axes of X and Y chromosomes are still long and thin and progressively condense to form bright axes, associated with bulges. Most experiments were conducted using SYCP3 in combination with a RMMAI protein, which allows easier distinction between synapsing and desynapsing X and Y chromosomes.
By using only SYCP3 staining, the PARs can only be identified unambiguously from the late zygonema-to-early pachynema transition through to diplonema. From pre-leptonema to mid-or late zygonema, the PARs were identified as the two brightest RMMAI signals, the two brightest mo-2 FISH signals, the two brightest PARb FISH signals or the two FISH signals from the PARd probe. The Y chromosome PAR could be distinguished from the X chromosome PAR using the PARb probe, as this probe also weakly stains the chromatin of the non-PAR portion of the Y chromosome.
Measurements of the PAR chromatin loops and axis length in oocytes were performed on two 14.5–15.5 dpc female fetuses (enriched for leptotene and zygotene oocytes) and two 18.5 dpc female fetuses (enriched for pachytene oocytes).
We found notable variability in the length of the X or Y chromosome PAR axis between different mice in our mouse colony maintained in a C57BL/6J congenic background, and even between different C57BL/6J males obtained directly from the Jackson Laboratory. This is in agreement with previous reports about the hypervariable nature of the mo-2 minisatellite and its involvement in unequal crossing over in the mouse6,24,36,38,39 (mo-2 was also named DXYmov15 or Mov15 flanking sequences). However, the intensity and elongation of the RMMAI signal and the length of the PAR axis were always correlated with mo-2 FISH signal intensity. Despite this variability, mo-2 and RMMAI proteins were enriched in the PAR and autosome ends of all mice analysed.
Analysis of SSDS data
SSDS sequencing data were from previously described studies7,20,31 and are all available at the Gene Expression Omnibus (GEO) under the accession numbers GSE35498, GSE99921 and GSE118913. To define the enrichment values presented in Extended Data Fig. 8b, the SSDS coverage was summed across the indicated coordinates adjacent to the mo-2 repeats. A chromosomal mean and standard deviation for chromosome 9 was estimated by dividing the chromosome into 4-kb bins, summing the SSDS coverage in each bin and calculating the mean and standard deviation after excluding those bins that overlapped a DSB hotspot. The enrichment score was then defined as the difference between the coverage in the mo-2-adjacent region and the mean coverage for chromosome 9, divided by the chromosome 9 standard deviation.
Immunoprecipitation and mass spectrometry
Immunoprecipitations were carried out on samples from wild-type and Ankrd31−/− mice using two separate polyclonal anti-ANKRD31 antibodies raised in rabbit and guinea pig20 (four samples total). Two additional immunoprecipitations were performed using an anti-cyclin B3 antibody on either wild-type or Ccnb3-knockout testes40,41; these samples serve as additional negative controls for the ANKRD31 interaction screen. For each sample, protein extracts were prepared from testes of three 12-day-old mice in 1 ml RIPA buffer (50 mM Tris-HCl, 150 mM NaCl, 0.1% SDS, 0.5% sodium deoxycholate, 1% NP-40, 10 mM MgCl2 and 100 units of benzonase for 1h at 4 °C. After centrifugation at 13,000 rpm for 20 min at 4 °C, the lysate was pre-cleared using 30 μl of a slurry of protein A/G Dynabeads for 1 h at 4 °C. Next, 50 μl of protein A/G beads coupled for 30 min with 10 μg of anti-ANKRD31 or anti-cyclin B3 antibody (monoclonal antibody 5 from ref. 40) were added and the solution was incubated overnight at 4 °C on a rotating rack. Beads were washed 3 times in 1 ml RIPA buffer and once with 1 ml of 50 mM ammonium bicarbonate. Samples were then digested overnight with 2 μg trypsin in 80 μl of 50 mM ammonium bicarbonate at 37 °C on a thermo mixer (850 rpm). Peptides were desalted using C18 zip tips, and then dried by vacuum centrifugation. Each sample was reconstituted in 10 μl 0.1% (v/v) formic acid and 4 μl was analysed by microcapillary liquid chromatography with tandem mass spectrometry (MS/MS) using the NanoAcquity (Waters) with an ACQUITY UPLC BEH C18 Column (Waters) configured with an ACQUITY UPLC M-Class Symmetry C18 trap column (Waters) coupled to a QExactive Plus mass spectrometer (Thermo Fisher Scientific). Peptides were eluted with a linear gradient of 0–35% acetonitrile (0.1% formic acid) in water (0.1% formic acid) over 150 min with a flow rate of 300 nl per min. The QE Plus was operated in automatic, data-dependent MS/MS acquisition mode with one MS full scan (380–1,800 m/z) at 70,000 mass resolution and up to ten concurrent MS/MS scans for the ten most intense peaks that were selected from each survey scan. Survey scans were acquired in profile mode and MS/MS scans were acquired in centroid mode at 17,500 resolution with an isolation window of 1.5 amu and normalized collision energy of 27. Automatic gain control (AGC) was set to 1 × 10 for MS1 and 5 × 10 and 100 ms IT for MS2. Charge state exclusion rejected ions having unassigned charge states or having a charge state above 6, with a dynamic exclusion of 15 s. All MS/MS samples were analysed using MaxQuant v.1.5.3.3 (Max Planck Institute of Biochemistry) at default settings with a few modifications.
Yeast two-hybrid assay
Mouse testis cDNAs for Ptip (also known as Paxip1), Zmym3 and Ankrd31 were amplified and cloned in vectors to generate fusion proteins with the Gal4 DNA-binding domain (Gal4BD) or activation domain (Gal4AD). Assays were conducted according to the manufacturer’s instructions (Clontech).In brief, Y2HGold and Y187 (Clontech) yeast haploid strains were transformed with constructs encoding Gal4BD and Gal4AD fusion proteins. After mating on YPD plates, diploid cells expressing Gal4BD and Gal4AD fusion proteins were selected on double dropout medium lacking leucine and tryptophan. Protein interactions were assayed by spotting diploid-cell suspensions on selective medium lacking leucine, tryptophan, histidine and adenine (quadruple dropout), and quadruple dropout medium containing X-α-gal (5-bromo-4-chloro-3-indolyl α-d-galactopyranoside) and aureobasidin A, and growing for 3 days at 30 °C.
Statistical analysis
All statistical tests were performed in R (v.3.4.4)42 and RStudio (v.1.1.442). Negative binomial regression was calculated using the glm.nb function from the MASS package (v.7.3-49)43.
Statistics and reproducibility
The pictures shown in this article are representative images that aim to illustrate the findings in the clearest manner. Any conclusion or statement regarding the results that is not associated with explicit quantification is based on the imaging and analysis of at least 20 cells, sometimes hundreds, usually from multiple mice. Details for the main figures are as follows.
Fig. 1a: The thickening of the PAR axis (using SYCP3 staining) and the elongation of the RMMAI signal along the PAR axis were observed in more than three different mice in hundreds of late-zygotene spermatocytes, mostly using our homemade antibodies against REC114 and ANKRD31. Other antibodies such as anti-SYCP2 and anti-HORMAD2 were used to confirm the PAR axis thickening, and anti-MEI1, anti-MEI4 and anti-IHO1 were used to confirm the elongation of the REC114 and ANKRD31 signal along the PAR axis, in more than 20 spermatocytes for each antibody. Fig. 1b: The PAR axis splitting, the extension of the RMMAI signal and the collapse of the PAR structure during X–Y chromosome synapsis were observed by SIM in more than 60 spermatocytes in more than 3 different mice. Fig. 2b: The colocalization between REC114 blobs (or RMMAI blobs in general) and mo-2 FISH signals was observed in all spermatocytes analysed (more than 200), from leptotene to early pachytene in more than 3 different mice. Fig. 3c: Axis splitting on the Y chromosome PAR was observed by SIM in more than 100 late-zygotene spermatocytes and in more than 20 zygotene-like spermatocytes from Hormad1−/− mice. The fork-shaped PAR structure in Rec8−/− mice was observed in more than 20 spermatocytes. The absence of PAR differentiation and decompaction of mo-2-containing chromatin was observed in more than 30 Ankrd31−/− spermatocytes and 20 Mei4−/− spermatocytes. This specific pattern was confirmed in at least three different mice of each genotype using conventional microscopy. The differentiation of the PAR axis becomes hardly detectable in Hormad1−/− mutants at a later stage in some pachytene-like spermatocytes as cells enter apoptosis, similar to Spo11−/− mutants. Fig. 4a: The differentiation of the non-centromeric end of chromosome 9 was observed in 6 spermatocytes by SIM and in more than 20 late-zygotene spermatocytes by conventional microscopy in 3 different mice.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
SSDS data are publicly available at the GEO under the accession numbers GSE35498, GSE99921 and GSE118913. Source Data for all graphs in the figures are provided with the paper (Figs. 2, 3, Extended Data Figs. 1, 3, 4, 6, 7, 9). The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository44 with the dataset identifier PXD017191. Processed mass spectrometry data are provided in the Source Data for Extended Data Fig. 1.
Code availability
Image analysis scripts are available on GitHub (https://github.com/Boekhout/ImageJScripts).
References
Raudsepp, T. & Chowdhary, B. P. The eutherian pseudoautosomal region. Cytogenet. Genome Res. 147, 81–94 (2015).
Kauppi, L. et al. Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331, 916–920 (2011).
Perry, J., Palmer, S., Gabriel, A. & Ashworth, A. A short pseudoautosomal region in laboratory mice. Genome Res. 11, 1826–1832 (2001).
Raudsepp, T., Das, P. J., Avila, F. & Chowdhary, B. P. The pseudoautosomal region and sex chromosome aneuploidies in domestic species. Sex Dev. 6, 72–83 (2012).
Palmer, S., Perry, J., Kipling, D. & Ashworth, A. A gene spans the pseudoautosomal boundary in mice. Proc. Natl Acad. Sci. USA 94, 12030–12035 (1997).
Soriano, P. et al. High rate of recombination and double crossovers in the mouse pseudoautosomal region during male meiosis. Proc. Natl Acad. Sci. USA 84, 7218–7220 (1987).
Brick, K., Smagulova, F., Khil, P., Camerini-Otero, R. D. & Petukhova, G. V. Genetic recombination is directed away from functional genomic elements in mice. Nature 485, 642–645 (2012).
Lange, J. et al. The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167, 695–708 (2016).
Kleckner, N. Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma 115, 175–194 (2006).
Zickler, D. & Kleckner, N. Meiotic chromosomes: integrating structure and function. Annu. Rev. Genet. 33, 603–754 (1999).
Panizza, S. et al. Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146, 372–383 (2011).
Lam, I. & Keeney, S. Mechanism and regulation of meiotic recombination initiation. Cold Spring Harb. Perspect. Biol. 7, a016634 (2014).
de Massy, B. Initiation of meiotic recombination: how and where? Conservation and specificities among eukaryotes. Annu. Rev. Genet. 47, 563–599 (2013).
Kauppi, L., Jasin, M. & Keeney, S. The tricky path to recombining X and Y chromosomes in meiosis. Ann. NY Acad. Sci. 1267, 18–23 (2012).
Page, J. et al. Inactivation or non-reactivation: what accounts better for the silence of sex chromosomes during mammalian male meiosis? Chromosoma 121, 307–326 (2012).
Kumar, R., Bourbon, H. M. & de Massy, B. Functional conservation of Mei4 for meiotic DNA double-strand break formation from yeasts to mice. Genes Dev. 24, 1266–1280 (2010).
Stanzione, M. et al. Meiotic DNA break formation requires the unsynapsed chromosome axis-binding protein IHO1 (CCDC36) in mice. Nat. Cell Biol. 18, 1208–1220 (2016).
Reinholdt, L. G. & Schimenti, J. C. Mei1 is epistatic to Dmc1 during mouse meiosis. Chromosoma 114, 127–134 (2005).
Kumar, R. et al. Mouse REC114 is essential for meiotic DNA double-strand break formation and forms a complex with MEI4. Life Sci. Alliance 1, e201800259 (2018).
Boekhout, M. et al. REC114 partner ANKRD31 controls number, timing, and location of meiotic DNA breaks. Mol. Cell 74, 1053–1068 (2019).
Papanikos, F. et al. Mouse ANKRD31 regulates spatiotemporal patterning of meiotic recombination initiation and ensures recombination between X and Y sex chromosomes. Mol. Cell 74, 1069–1085 (2019).
Kumar, R. et al. MEI4 – a central player in the regulation of meiotic DNA double-strand break formation in the mouse. J. Cell Sci. 128, 1800–1811 (2015).
Harbers, K., Francke, U., Soriano, P., Jaenisch, R. & Müller, U. Structure and chromosomal mapping of a highly polymorphic repetitive DNA sequence from the pseudoautosomal region of the mouse sex chromosomes. Cytogenet. Cell Genet. 53, 129–133 (1990).
Takahashi, Y. et al. Methylation imprinting was observed of mouse mo-2 macrosatellite on the pseudoautosomal region but not on chromosome 9. Chromosoma 103, 450–458 (1994).
Shin, Y. H. et al. Hormad1 mutation disrupts synaptonemal complex formation, recombination, and chromosome segregation in mammalian meiosis. PLoS Genet. 6, e1001190 (2010).
Bannister, L. A., Reinholdt, L. G., Munroe, R. J. & Schimenti, J. C. Positional cloning and characterization of mouse mei8, a disrupted allelle of the meiotic cohesin Rec8. Genesis 40, 184–194 (2004).
Kim, K. P. et al. Sister cohesion and structural axis components mediate homolog bias of meiotic recombination. Cell 143, 924–937 (2010).
Daniel, K. et al. Meiotic homologue alignment and its quality surveillance are controlled by mouse HORMAD1. Nat. Cell Biol. 13, 599–610 (2011).
Brick, K., Pratto, F., Sun, C. Y., Camerini-Otero, R. D. & Petukhova, G. Analysis of meiotic double-strand break initiation in mammals. Methods Enzymol. 601, 391–418 (2018).
Khil, P. P., Smagulova, F., Brick, K. M., Camerini-Otero, R. D. & Petukhova, G. V. Sensitive mapping of recombination hotspots using sequencing-based detection of ssDNA. Genome Res. 22, 957–965 (2012).
Brick, K. et al. Extensive sex differences at the initiation of genetic recombination. Nature 561, 338–342 (2018).
Thacker, D., Mohibullah, N., Zhu, X. & Keeney, S. Homologue engagement controls meiotic DNA break number and distribution. Nature 510, 241–246 (2014).
Kauppi, L. et al. Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev. 27, 873–886 (2013).
Arnold, A. P. Mouse models for evaluating sex chromosome effects that cause sex differences in non-gonadal tissues. J. Neuroendocrinol. 21, 377–386 (2009).
Murakami, H., Lam, I., Song, J., Huang, P.-C., Song, J., van Overbeek, M. & Keeney, S. Multilayered mechanisms ensure that short chromosomes recombine in meiosis. Nature https://doi.org/10.1038/s41586-020-2248-2 (2020).
Harbers, K., Soriano, P., Müller, U. & Jaenisch, R. High frequency of unequal recombination in pseudoautosomal region shown by proviral insertion in transgenic mouse. Nature 324, 682–685 (1986).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Kipling, D. et al. Structural variation of the pseudoautosomal region between and within inbred mouse strains. Proc. Natl Acad. Sci. USA 93, 171–175 (1996).
Kipling, D., Salido, E. C., Shapiro, L. J. & Cooke, H. J. High frequency de novo alterations in the long-range genomic structure of the mouse pseudoautosomal region. Nat. Genet. 13, 78–82 (1996).
Karasu, M. E. & Keeney, S. Cyclin B3 is dispensable for mouse spermatogenesis. Chromosoma 128, 473–487 (2019).
Karasu, M. E., Bouftas, N., Keeney, S. & Wassmann, K. Cyclin B3 promotes anaphase I onset in oocyte meiosis. J. Cell Biol. 218, 1265–1281 (2019).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2018).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S 4th edn, (Springer, 2002).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Wojtasz, L. et al. Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet. 5, e1000702 (2009).
Gaysinskaya, V. & Bortvin, A. Flow cytometry of murine spermatocytes. Curr. Protoc. Cytom. 72, 7 44.1–7.44.24 (2015).
Chong, L. et al. A human telomeric protein. Science 270, 1663–1667 (1995).
Tres, L. L. Extensive pairing of the XY bivalent in mouse spermatocytes as visualized by whole-mount electron microscopy. J. Cell Sci. 25, 1–15 (1977).
Bergs, J. W. et al. Differential expression and sex chromosome association of CHD3/4 and CHD5 during spermatogenesis. PLoS ONE 9, e98203 (2014).
Kourmouli, N. et al. Heterochromatin and tri-methylated lysine 20 of histone H4 in animals. J. Cell Sci. 117, 2491–2501 (2004).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Turner, J. M., Burgoyne, P. S. & Singh, P. B. M31 and macroH2A1.2 colocalise at the pseudoautosomal region during mouse meiosis. J. Cell Sci. 114, 3367–3375 (2001).
Costa, Y. et al. Two novel proteins recruited by synaptonemal complex protein 1 (SYCP1) are at the centre of meiosis. J. Cell Sci. 118, 2755–2762 (2005).
Acknowledgements
We thank A. Tóth and B. de Massy for antibodies, mice, discussions and sharing of unpublished information; A. North and the Bio-Imaging Resource Center at Rockefeller University for assistance with SIM (supported by award number S10 RR031855 from the National Center For Research Resources); and R. Hendrickson, R. Soni and Z. Li (MSKCC Proteomics Core) for assistance with mass spectrometry. This work used the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov). MSKCC core facilities are supported by a Cancer Center Support Grant (P30 CA008748). L.A. was supported in part by a fellowship from the Lalor Foundation; M.B. was supported in part by a Rubicon fellowship from the Netherlands Organization for Scientific Research; and M.v.O. was supported in part by an NIH fellowship (F32 GM096692). This work was supported by NIGMS grants R35 GM118092 (S.K.) and R35 GM118175 (M.J.).
Author information
Authors and Affiliations
Contributions
L.A. designed and conducted all of the cytogenetic experiments presented and analysed the data; M.E.K. generated Ankrd31 mutant mice and anti-ANKRD31 antibodies; M.B. and M.E.K. provided Ankrd31 mutant mice and unpublished data; M.E.K. performed the immunoprecipitation for mass spectrometry and T.L. validated the ANKRD31-interacting proteins; K.B. and F.P. performed SSDS and analysed the data under the supervision of R.D.C.-O., with input from L.A. and S.K.; M.v.O. generated REC8 and REC114 antibodies; L.K. performed initial characterization and provided unpublished data on the PAR ultrastructure and cohesin enrichment; M.J. and S.K. designed and supervised the research, analysed the data and secured funding; L.A. and S.K. wrote the manuscript with input from M.J. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Francesca Cole Bergemann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 PAR axis thickening and accumulation of RMMAI proteins.
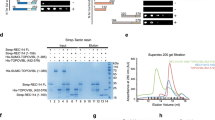
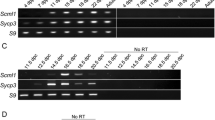
a, Axis thickening (SYCP3 and HORMAD2 staining) at the PAR (arrowheads) in a late-zygotene spermatocyte. HORMAD2 staining in the PAR at late zygonema mimics SYCP3 staining in all late-zygonema spermatocytes analysed (n > 20) in three mice. Scale bars, 2 μm. b, Image (adapted under a Creative Commons CC-BY licence from a previous study45) showing enrichment of HORMAD1 on the thick PAR axis of the Y chromosome. c, Colocalization of ANKRD31 and MEI4, REC114, IHO1 and MEI1. Representative zygotene spermatocytes are shown. Arrowheads indicate densely staining blobs. Areas indicated by dashed boxes are shown at higher magnification. The graphs show the total number of foci colocalized in leptotene or zygotene spermatocytes (mean ± s.d.). N.D., not determined: the low immunofluorescence signal for MEI1 did not allow us to quantify the colocalization with ANKRD31, although MEI1 showed clear colocalization with ANKRD31 in the blobs and at least some autosomal foci (insets). Scale bars, 2 μm. d, PARb FISH probe colocalizes with REC114 blobs. Two blobs are on the PAR, as judged by chromosome morphology and bright FISH with a PARb probe, and others highlight specific autosome ends. The colocalization between REC114 blobs and PARb FISH signals was observed in all spermatocytes analysed (n > 60), from pre-leptonema to early pachynema, in more than three mice. Scale bar, 2 μm. e, ANKRD31, REC114 and MEI1 immunostaining starts to appear in pre-leptonema. Seminiferous tubules were cultured with EdU to label replicating cells, and then chromosome spreads were stained for SYCP3 and either MEI1 with REC114 or ANKRD31 with PARb FISH. Colocalized foci appear in pre-leptonema (EdU-positive cells that are weakly SYCP3 positive), as previously shown for MEI4 and IHO117,22. Because we can already detect ANKRD31 accumulation at sites of PARb hybridization, we infer that the stronger sites of accumulation of MEI1 and REC114 also include PARs. PARb colocalized with ANKRD31 blobs (top) and MEI1 with REC114 (bottom) in all pre-leptotene spermatocytes analysed (n > 20) in one mouse. Scale bars, 2 μm. f, REC114 is not detected in the mo-2 regions in spermatogonia. Seminiferous tubules were cultured with EdU, and chromosome spreads were stained for DMRT1 (a marker of spermatogonia46) and REC114 plus mo-2 FISH. REC114 blobs colocalized with mo-2 FISH signals in the pre-leptotene spermatocyte (bottom) but were not apparent in the DMRT1-positive spermatogonium (top). Both cells shown were captured in a single microscopic field. Mo-2 FISH signals do not colocalize with the REC114 signal in all of the spermatogonia analysed (n > 20) in one mouse. Scale bar, 2 μm. g, Candidate ANKRD31-interacting proteins. To identify other PAR-associated proteins, ANKRD31 was immunoprecipitated from extracts made from whole testes of 12-day-old mice using two different polyclonal antibodies. This table shows a subset of proteins that were identified by mass spectrometry in immunoprecipitates from the testes of wild-type but not Ankrd31−/− mice, and not in immunoprecipitates using an irrelevant antibody (anti-cyclin B3). LFQ, label-free quantification. h, Enrichment of ZMYM3 (top) and PTIP (bottom) at the PAR. Sex chromosomes of representative early-pachytene spermatocytes are shown. ZMYM3 and PTIP were enriched in the PAR in all spermatocytes analysed (n > 20) in three mice. Scale bars, 2 μm. i, Yeast two-hybrid assays testing the interaction of full-length (FL) ANKRD31 fused to Gal4AD with either full-length PTIP or the C-terminal 191 amino acids of ZMYM3 fused to Gal4BD (full-length ZMYM3 autoactivates in this assay). Double dropout (DDO) medium selects for the presence of both the Gal4AD and the Gal4BD vector (positive control for growth); quadruple dropout (QDO) and QXA (QDO plus X-α-gal and aureobasidin A) media select for a productive two-hybrid interaction at lower and higher stringency, respectively. The image is representative of two experiments using the same yeast strains.
Extended Data Fig. 2 PAR ultrastructure.
a, Comparison of conventional microscopy and SIM, showing that the thickened PAR axis in conventional microscopy is resolved as separated axial cores (arrowheads). The thickening of the PAR axis in conventional microscopy and the splitting of the PAR axis in SIM was observed in more than 60 spermatocytes at late zygonema in at least three mice. Scale bars, 2 μm. b, Ultrastructure of the axis proteins SYCP2, SYCP3 and HORMAD2 in the PAR. SYCP2 (left) and HORMAD2 (right) staining mimic SYCP3 staining in late zygonema in all cells analysed (n > 30 in at least three mice by conventional microscopy; n = 5 in one mouse by SIM) (except that HORMAD2 appears rather depleted at the telomeres compared to SYCP3 and SYCP2). Scale bars, 1 μm. c–e, Distinguishing between splitting apart of sister chromatid axes and a crozier configuration in which a single conjoined axis for both sister chromatids is folded back on itself. A crozier (cartooned in c) was ruled out because the telomere-binding protein TRF147 decorates the tip of the PAR bubble (d) and the FISH signal for the PARb probe is arrayed relatively symmetrically on both axial cores (e), consistent with separated sister chromatid axes. Scale bars, 1 μm. The presence of TRF1 at the distal tip of the PAR was observed in all spermatocytes analysed (n > 20 by conventional microscopy; n = 3 by SIM) in one mouse. PARb FISH signals were relatively symmetrically arranged along the split PAR axes (n > 100 by conventional microscopy; n = 9 by SIM) in three mice. f, Schematic of the PAR ultrastructure and distribution of axis and RMMAI proteins at late zygonema. Tel., telomere. g, h, Paired PARs with elongated and split axes occur in late zygonema to early pachynema. Electron micrographs (adapted with permission from a previous study48) are shown in comparison with SIM immunofluorescence images of spermatocytes at early pachynema (g) or late zygonema (h; cyan arrowheads indicate examples of incomplete autosomal synapsis). Elongation and splitting of PAR axes occurs earlier than originally thought48 (Supplementary Discussion). Scale bars in SIM images, 1 μm (g); 2 μm (h). Extended and split PAR axes were observed by SIM (n > 30 spermatocytes) around the zygonema–pachynema transition in more than three mice. i, REC114 enrichment and axis splitting occurs in the absence of SPO11, thus neither is provoked by DSB formation. PAR axis splitting and extension of the RMMAI signal were observed by SIM in Spo11−/− mice in more than 20 late-zygotene-like spermatocytes in more than three mice. The differentiation of the PAR axis became hardly detectable at later stages in some pachytene-like spermatocytes as cells entered apoptosis. Scale bar, 1 μm.
Extended Data Fig. 3 Time course of the spatial organization of the PAR loop–axis ensemble.
a, Time course of REC8 and ANKRD31 immunostaining along the PAR axis from pre-leptonema (preL, left) to mid-pachynema (right). A montage of representative SIM images is shown. Chromosomes a–e are presumptive X or Y, but could be the distal end of chromosome 9. Chromosomes at later stages were unambiguously identified by morphology. Chromosomes i–k show examples in which the initial pairing (probably synaptic) contact between X and Y chromosomes is centromere-proximal (that is, closer to the PAR boundary) (i), distal (closer to the telomere) (k), or interstitial (j). The preferential enrichment of REC8 at the border of the PAR split axes was observed in more than 30 zygotene spermatocytes by SIM in more than three mice. Scale bar, 1 μm. b, We collected three measurements of conventional immuno-FISH images from leptonema through to mid-pachynema: length of the REC114 signal along the PAR axis; maximum distance from the PARb FISH signal to the distal end of the SYCP3-defined axis; and axis-orthogonal extension of the FISH signal for the PARb probe (a proxy for loop sizes). Data were collected from three males. Insets show examples of each type of measurement at each stage. Horizontal black lines indicate means. Means of each measurement for each mouse at each stage are given below, along with the means across all three mice. Means are rounded to two significant figures; the grand means were calculated using unrounded values from individual mice. The number of cells of each stage from each mouse is given. Modest variability in the apparent dimensions of the Y chromosome PAR between different mice may be attributable to variation in the copy number of mo-2 and other repeats because of unequal exchange during meiosis. Nonetheless, highly similar changes in spatial organization over time in prophase were observed in all mice examined—namely, progressive elongation then shortening of axes and concomitant lengthening of loops. Scale bar, 1 μm. In brief, a and b show the following. At pre-leptonema, ANKRD31 blobs had a closely juxtaposed focus of the meiotic cohesin subunit REC8 (chromosome a). In leptonema and early zygonema, ANKRD31 and REC114 signals stretched along the presumptive PAR axes, with REC8 restricted to the borders (a, chromosomes b–e). The SYCP3-defined axis was already long as soon as it was detectable (0.73 μm) and the PARb FISH signal was compact (0.52 μm) (b, i). At late zygonema, the PAR axis had lengthened still further (1.0 μm), whereas the PARb signal remained compact (b, ii). The PAR split into separate axes during this stage, each with abundant RMMAI (a, chromosomes f–h). The split was a REC8-poor zone bounded by REC8 foci (a, chromosomes f–h; Extended Data Fig. 2f). After synapsis, the axes shortened and chromatin loops decompacted, with concomitant dissociation of RMMAI proteins. As cells transitioned into early pachynema and the X and Y chromosome PARs synapsed (a, chromosomes i–m), the PAR axes began to shorten slightly (0.85 μm) and the PARb signal expanded (0.85 μm) (b, iii). Meanwhile, the elongated ANKRD31 signals progressively decreased in intensity, collapsed along with the shortening axes and separated from the axis while remaining nearby (a, chromosomes l–m). By mid-pachynema, the PAR axes collapsed still further, to about half their zygotene length (0.50 μm) and the PARb chromatin expanded to more than twice the zygotene measurement (1.3 μm). ANKRD31 and REC114 enrichment largely disappeared, leaving behind a bright bolus of REC8 on the short remaining axis (a, chromosomes n–o; b, iv). c, Non-homologous synapsis appears sufficient to trigger collapse of the PAR loop–axis structure. We measured the length of the REC114 signal along the PAR axis and the extension of mo-2 chromatin orthogonal to the axis in Spo11−/− spermatocytes in which the X chromosome PAR had non-homologously synapsed with an autosome while the Y chromosome PAR remained unsynapsed. Within any given cell, the unsynapsed Y chromosome PAR maintained the characteristic late-zygotene configuration (long axis, short loops) whereas the synapsed X chromosome PAR adopted the configuration characteristic of pachynema (short axis, long loops). Error bars are mean ± s.d. Scale bar, 2 μm.
Extended Data Fig. 4 RMMAI enrichment at mo-2 minisatellite arrays in the PAR and on specific autosomes.
a, Top, self-alignment of the PARb FISH probe (reproduced from Fig. 2a). The circled block is a 20-kb mo-2 cluster. Bottom, schematic depicting the last 1.4 Mb of the non-centromeric ends of the indicated chromosomes, showing the positions of mo-2 repeats (green) adjacent to assembly gaps (mm10); mo-2 repeats were identified by BLAST search using the mo-2 consensus sequence. Mo-2 repeats also appear at the distal end of chromosome 4 in the Celera assembly (Mm_Celera, 2009/03/04). PARb and PARd BAC clones are indicated. b, Confirmation that autosomal mo-2 FISH signals match the chromosomal locations indicated by mm10 or Celera genome assemblies. FISH was performed using an oligonucleotide probe containing the mo-2 consensus sequence in combination with BAC probes for adjacent segments of chromosomes 13, 9 and 4, as indicated. Magenta arrows point to concordant FISH signals. The chromosome 9 BAC probe also hybridizes to the PAR. The colocalization of mo-2 and the three autosomal FISH signals was observed in two mice (n > 20 spermatocytes). Scale bars, 2 μm. c, Comparison of mo-2 FISH with REC114 localization relative to the PAR boundary (PARb FISH probe) and the distal PAR (PARd probe). In mid-zygonema, the mo-2 FISH signal colocalizes well with REC114 staining in between the PARb and PARd FISH signals. In late zygonema, mo-2 and REC114 are similar to one another and are elongated along the thickened SYCP3 staining of the PAR axis. From early to mid-pachynema, REC114 progressively disappears, whereas the mo-2 FISH signal becomes largely extended away from the PAR axes. The relative positions of the PARb and PARd probes reinforce the conclusion that the PAR does not adopt a crozier configuration. The different positioning of PARb and PARd FISH signals compared to mo-2 or REC114 signals was observed in more than 30 spermatocytes in at least three mice. Scale bar, 1 μm. d, Illustration of the compact organization of the PAR chromatin (mo-2 FISH signal) compared to a whole Y chromosome paint probe. The costaining of mo-2 and the full chromosome Y probe was evaluated in one mouse (n > 20 spermatocytes). Scale bar, 2 μm. e, Lower mo-2 copy number in the M. musculus molossinus subspecies correlates with lower REC114 staining in mo-2 regions. The left panels compare MSM and B6 mice for colocalization between REC114 immunostaining and mo-2 FISH in leptotene spermatocytes. The REC114 and SYCP3 channels are shown at equivalent exposure for the two strains, whereas a longer exposure is shown for the mo-2 FISH signal in the MSM spermatocyte. The mo-2-associated REC114 blobs are much brighter relative to the smaller dispersed REC114 foci in the B6 spermatocyte than in the MSM spermatocyte. The right panel shows representative pachytene spermatocytes to confirm the locations of mo-2 clusters at autosome ends and at the PAR in the MSM background. The lower intensity of REC114 blobs in MSM compared to B6 spermatocytes was observed in more than 30 spermatocytes in three different pairs of mice. Scale bars, 2 μm. f, PAR enrichment for ANKRD31 and RPA2 correlates with mo-2 copy number. Top, late-zygotene spermatocytes from MSM × B6 F1 hybrids. Scale bar, 1 μm. Bottom, PAR-associated signals on B6-derived (YB6) and MSM-derived (XMSM) chromosomes from the indicated number of spermatocytes. Red lines indicate mean ± s.d. Differences between the PAR intensities of X and Y chromosomes are significant both for proteins and for mo-2 FISH in both F1 hybrids (P < 10−13, paired t-test; for exact P values, see the associated Source Data). g, Representative micrographs of late-zygotene spermatocytes from reciprocal F1 hybrid males from crosses of B6 (high mo-2 copy number) and MSM (low mo-2 copy number) parents. Scale bars, 1 μm. h, Frequency of paired X and Y chromosomes at late zygonema and mid-pachynema analysed in three MSM and three B6 males. Differences between strains were not statistically significant at either stage (P = 0.241 for late zygonema and P = 0.136 for mid-pachynema; two-sided Student’s t-test). X and Y are late-pairing chromosomes in the MSM background—as in the B6 background. The similar pairing kinetics indicates that the lower intensity of RMMAI staining on the MSM PAR is not attributable to earlier PAR pairing and synapsis in this strain. The number of spermatocytes analysed is indicated.
Extended Data Fig. 5 Mo-2 regions accumulate heterochromatin factors.
a, Costaining of ANKRD31 or mo-2 with the indicated proteins and histone marks known to localize at the pericentromeric heterochromatin (mouse major satellite), in zygotene spermatocytes (left) and pre-leptotene spermatocytes (right). Each of the heterochromatin factors shows a locally enriched signal coincident with mo-2 regions (arrowheads), in addition to broader staining of other sub-nuclear regions. The CHD3/4 antibody recognizes both proteins49. The colocalization of ANKRD31 blobs with heterochromatin blobs was observed in all zygotene spermatocytes analysed (n > 20) in at least three mice for each antibody (left) and in one mouse for pre-leptotene spermatocytes (n > 10) for each antibody (right). Scale bars, 2 μm. b, CHD3/4, ATRX, HP1β, H4K20me3, H3K9me3 and macroH2A1.2 are not detectably enriched at mo-2 regions in spermatogonia (small, DMRT1-positive cells). These factors may be present at mo-2 regions in these cells, but do not appear to accumulate to elevated levels. The absence of colocalization between mo-2 FISH signals and heterochromatin factors was noted in all spermatogonia analysed (n > 30) from one mouse. Scale bars, 2 μm. c, Heterochromatin factors can be detected in the PAR up to late pachynema. Each of the assayed proteins and histone marks showed staining on the autosomal and X-specific pericentromeric heterochromatin, the sex body and euchromatin—albeit with variations between sites in the timing and level of accumulation. Notably, however, they also showed enriched staining at all mo-2 regions up to early or mid-pachynema, as shown for H4K20me3 (top). By mid-to-late pachynema, as shown here for H3K9me3, the signal persisted in the PAR but was usually barely detectable at mo-2 regions of chromosome 9 or chromosome 13. This observation indicates that—at least for the PAR—the heterochromatin factors can continue to be enriched on mo-2 chromatin after RMMAI proteins have dissociated. These results substantially extend previous observations about CHD3/4 colocalizing with PAR FISH signals; H4K20me3 being localized in the PAR and at the ends of other chromosomes; and detection of H3K9me3, HP1β and macroH2A1.2 in the PAR in late pachynema49,50,51,52. The colocalization between major satellite (maj sat) and H4K20me3 and H3K9me3 was observed in all spermatocytes analysed (n > 20) in one mouse. The colocalization between H4K20me3 and mo-2 FISH signals was observed in all spermatocytes analysed (n > 60) from the pre-leptotene to mid-pachytene stage in more than three mice. Scale bars, 2 μm. d, Enrichment of the heterochromatin factors is independent of SPO11. Representative images of Y chromosomes from a Spo11–/– mouse are shown. The colocalization between PAR mo-2 FISH signals and heterochromatin factors was observed in all Spo11−/− spermatocytes analysed (n > 30) in more than three mice for CHD3/4 and at least one mouse each for ATRX, HP1β, HP1γ, macroH2A1.2, H3K9me3 and H4K20me3. Scale bar, 1 μm.
Extended Data Fig. 6 Genetic requirements for RMMAI assembly on chromosomes and for PAR loop–axis organization.
a, Representative micrographs of ANKRD31, MEI4, IHO1 and MEI1 staining in wild-type spermatocytes and the indicated mutants (quantification is in Fig. 3a). Scale bars, 2 μm. b, Measurements of PAR loop–axis organization, as in Fig. 3b, in two additional males. The data from mouse 1 are reproduced from Fig. 3b to facilitate comparison. Means of each measurement for each mouse at each stage are given below, along with the means across all three mice. Means are rounded to two significant figures; the grand means were calculated using unrounded values from individual mice. The number of cells of each stage from each mouse is given. c, REC8 is dispensable for the splitting apart of PAR sister chromatid axes, but is required to maintain the connection between sisters at the distal tip of the chromosome. A representative SIM image is shown of a Y chromosome from a late-zygotene Rec8−/− spermatocyte. The SYCP3-labelled axes adopt an open-fork configuration. The distal FISH probe (PARd) shows that there are clearly disjoined sisters, whereas the PAR boundary (PARb) shows only a single compact signal comparable to the wild type. The disposition of the probes and SYCP3 further rules out the crozier configuration as an explanation for split PAR axes. The structure of the Y or X chromosome PAR was resolved by SIM as ‘fork-shaped’ in all spermatocytes analysed (n > 20) from three mice. Scale bar, 1 μm. d, Quantification of REC114 and MEI4 foci in two additional pairs of wild-type and Ankrd31−/− mice. Horizontal lines indicate means. Fewer foci were observed in the Ankrd31−/− mutant (two-sided Student’s t-tests for each comparison of mutant to wild type: P = 5.6 × 10−6 (second set, REC114); P = 1.1 × 10−5 (second set, MEI4); P = 2.1 × 10−6 (third set, REC114); P = 0.017 (third set, MEI4)). e, Reduced REC114 staining intensity of axis-associated foci in Ankrd31−/− mutants. To rigorously control for slide-to-slide and within-slide variation in immunostaining, we mixed together wild-type and Ankrd31−/− testis cell suspensions before preparing chromosome spreads. A representative image is shown of a region from a single microscopic field containing two wild-type zygotene spermatocytes (left) and two Ankrd31−/− spermatocytes of the equivalent stage (right). Note the diminished intensity of REC114 foci in the Ankrd31−/− spermatocytes. Scale bar, 2 μm. REC114 (non-blob) foci showed a lower fluorescence intensity in Ankrd31–/– mutant compared to wild-type spermatocytes in all pairs of spermatocytes captured in the same imaging field (n = 8 pairs), from one pair of mice. f, PAR enrichment of heterochromatin-associated factors is independent of ANKRD31. Representative images of the Y chromosome at late zygonema or early pachynema, showing colocalization between the decompacted mo-2 chromatin and the indicated proteins. Both the FISH and the immunofluorescence signals are localized mostly off the axis (compare with the same signals in the absence of SPO11; Extended Data Fig. 5d). The mo-2 FISH signal colocalized off the axis with the heterochromatin factors in Ankrd31−/− mice in all spermatocytes analysed (n > 30) in more than three mice for CHD3/4 and at least one mouse for ATRX, HP1β, HP1γ, macroH2A1.2, H3K9me3 and H4K20me3. Scale bar, 1 μm.
Extended Data Fig. 7 PAR-associated RPA2 foci.
a, Loop–axis organization of the mo-2 region of chromosome 9 in late zygonema. Compare with the PAR (Fig. 3b). Error bars are mean ± s.d. Scale bars, 1 μm. b, Low mo-2 copy number correlates with less loop–axis reorganization (SIM images of late-zygotene F1 hybrid spermatocytes). The differentiation of the B6 PAR was observed in both hybrids (B6 × MSM and MSM × B6) in three and four spermatocytes, respectively, by SIM (one mouse for each), and in more than 20 spermatocytes by conventional microscopy (two mice of each genotype). Scale bars, 1 μm. c–e, Immuno-FISH for RPA2 and mo-2 was used to detect DSBs cytologically in wild-type mice and the indicated mutants. To analyse Rec8 and Hormad1 mutations, we compared to mutants lacking SYCE1 (a synaptonemal complex central element component53) because Syce1–/– mutants show similar meiotic progression defects without defective RMMAI recruitment. c, Representative images. Scale bars, 2 μm (main image); 1 μm (inset). d, Global counts of RPA2 foci for zygotene (zyg) or zygotene-like cells and for pachytene (pach) or pachytene-like cells. e, Fraction of mo-2 regions, for each cell, that had a colocalized RPA2 focus. Red lines indicate mean ± s.d. Statistical significance is indicated in d, e for comparisons (two-sided Student’s t-tests) of wild type to Ankrd31−/− or of Syce1−/− to either Rec8−/− or Hormad1−/− for matched stages; for exact P values, see the associated Source Data. The number of discretely scorable mo-2 regions in e varied from cell to cell depending on pairing status. f, Frequent formation of DSBs at mo-2 regions in the PAR and on autosomes does not require HORMAD1. Left, micrograph showing two adjacent spermatocytes (boundary indicated by dashed line). Scale bar, 2 μm. Right, insets show higher-magnification views of the numbered mo-2 regions, all of which are associated with RPA2 immunostaining of varying intensity. This image illustrates the preferential formation of RPA2 foci in mo-2 regions in a Hormad1−/− mouse; quantification is in e. g, Autosomal mo-2 regions often form DSBs late. Immuno-FISH for RPA2, mo-2 and PARb was used to detect DSBs cytologically in wild-type mice from leptonema to mid-pachynema, and to distinguish the X or Y chromosome PAR from chromosomes 9 and 13. Chromosome 4 was not assayed because the mo-2 FISH signal was often barely detectable. Top, global number of RPA2 foci per cell. Black lines indicate mean ± s.d. Bottom, percentage of spermatocytes with an RPA2 focus overlapping the PAR (X, Y or both) or overlapping chromosome 9 or 13. A representative image of an early-pachytene spermatocyte is shown. As previously shown for the PAR2, autosomal mo-2 regions continue to accumulate RPA2 foci beyond the time point at which global RPA2 foci have largely or completely ceased accumulating. Scale bar, 2 μm. h, X–Y chromosome pairing status, quantified by immuno-FISH for SYCP3 and the PARd probe. i, Montage of SIM images from a B6 male, showing that multiple, distinct RPA2 foci can be detected from late zygonema to mid-pachynema. This suggests that multiple PAR DSBs can be formed during one meiosis2. The presence of multiple RPA2 foci in the PAR was observed by SIM in more than 20 spermatocytes from late zygonema to mid-pachynema in one mouse. Scale bar, 1 μm. j, Percentage of spermatocytes at the zygotene–pachytene transition with no (0), one, two or three distinguishable RPA2 foci on the unsynapsed Y chromosome PAR of MSM and B6 mice. The difference between the strains is statistically significant (negative binomial regression, P = 7.2 × 10−5). The number of spermatocytes analysed is indicated. A representative image is shown for each genotype, with one RPA2 focus on the MSM PAR and two apparent sites of RPA2 accumulation on the B6 PAR. The detection of multiple foci is consistent with reported double crossovers6. Scale bar, 1 μm.
Extended Data Fig. 8 DSB maps on the PAR and autosomal mo-2 regions.
a, SSDS sequence coverage (data from previous reports7,20) is shown for the X chromosome PAR (shown in a different form previously20), the Y chromosome PAR and the mo-2-adjacent regions of chromosomes 9 and 13. The dashed segments indicate gaps in the mm10 genome assembly. We did not assess chromosome 4 because available assemblies are too incomplete. b, Regions adjacent to the mo-2 region on chromosome 9 show an SSDS signal that is reproducibly increased relative to the chromosome 9 average in wild-type testis samples but not in maps from Ankrd31−/− testes or wild-type ovaries. Two of the SSDS browser tracks are reproduced from a. The bar chart shows enrichment values from individual SSDS maps (T1–T9 are maps from wild-type testes; O1 and O2 are from wild-type ovaries31). Enrichment values are defined as coverage across the indicated coordinates relative to the mean coverage for chromosome 9 (see Methods for details). Ovary sample O1 and the Ankrd31−/− adult sample are known to have poorer signal-to-noise ratios than the other samples20,31. For all SSDS coverage tracks, reads that map to multiple locations are included after random assignment to one of their mapped positions. However, the same conclusions are reached about ANKRD31 dependence and PRDM9 independence of the signal on chromosomes 9 and 13 if only uniquely mapped reads are used. c, Oocytes incur substantially fewer DSBs than spermatocytes near the mo-2 region on chromosome 9. The SSDS signal is from a previous study31 (samples T1 and O2). The X chromosome PAR is shown for comparison (previously shown to be essentially devoid of DSBs in ovary samples31). See b for quantification.
Extended Data Fig. 9 RMMAI accumulation and low frequency of DSB formation at mo-2 regions in oocytes.
a, Examples of zygotene oocytes showing the colocalization between blobs of IHO1 and REC114, MEI4 and MEI1, or ANKRD31 and mo-2 FISH signal (arrowheads). RMMAI blobs colocalized with mo-2 FISH signals in all zygotene oocytes analysed (n > 30) from at least three mice. Scale bars, 2 μm. b, PAR ultrastructure in oocytes, quantified as in Extended Data Fig. 3b. Late-zygotene cells with PAR synapsis are compiled separately from other zygotene cells. Error bars are mean ± s.d. Scale bar, 1 μm. c, Examples of zygotene oocytes showing colocalization of ANKRD31 blobs with enrichment for heterochromatin factors. ANKRD31 colocalized with blobs of heterochromatin factors in all zygotene oocytes analysed (n > 20) from one mouse. Scale bars, 2 μm. d, Representative SIM image of a wild-type late-zygotene oocyte showing neither detectable splitting of the PAR axis nor REC8 enrichment. The absence of spermatocyte-like differentiation of the PAR axis was observed (n > 30 zygotene oocytes) in more than three mice. A modest degree of differentiation was observed in a minority of oocytes (5/45) analysed by SIM, but this did not resemble the typical PAR axis splitting found in spermatocytes. Scale bar, 2 μm. e, Prolonged asynapsis does not allow axis splitting to occur in oocytes. Because synapsis appears to be sufficient to trigger the collapse of the PAR ultrastructure in spermatocytes (Extended Data Fig. 3b), we asked whether preventing synapsis (that is, in a Syce1−/− mutant) could reveal a tendency towards axis splitting in oocytes. Whereas axis splitting was clearly observed by SIM in Syce1−/− mutant spermatocytes, PAR axes were not detectably split in oocytes. Axis splitting of chromosome 9 was observed by SIM in multiple (n > 20) Syce1−/− spermatocytes from three different mice. The centromere-distal axes were also occasionally seen to be split in chromosomes 13 or 4, but we did not quantify this for these chromosomes. In males, the differentiation of the axes of the PAR or chromosome 9 becomes hardly detectable at later stages in some pachytene-like spermatocytes as cells enter apoptosis, similar to Spo11−/− or Hormad1−/− mice. However, in Syce1−/− oocytes, no obvious axis differentiation or splitting was observed by conventional microscopy or by SIM in multiple oocytes (n > 30) from three different mice—similar to what was observed in wild-type oocytes. Scale bars, 2 μm (main image); 1 μm (insets). f, h, Delaying synapsis promotes the formation of DSBs in the PAR in oocytes. Top, representative micrographs of pachytene XY (f) and Syce1−/− XX (h) oocytes. Middle, RPA2 fluorescence intensity at the border of mo-2 FISH signals from the PAR, chromosome 9 and chromosome 13. Bottom, percentage of oocytes with an RPA2 focus colocalizing with mo-2 regions on the PAR, chromosome 9 and chromosome 13. Graphs show data only for pachytene oocytes in which PARs are synapsed (two mice of each genotype). Red lines indicate mean ± s.d. Scale bars, 2 μm. g, Percentage of pachytene oocytes with one or more RPA2 foci colocalizing with the mo-2 FISH signal from the PAR, chromosome 9 and chromosome 13 in XY pachytene oocytes that had unsynapsed X and Y chromosomes. Scale bars, 2 μm (main image); 1 μm (inset).
Extended Data Fig. 10 Summary of PAR ultrastructure and molecular determinants of axis remodelling and DSB formation.
Schematic of the meiotic Y chromosome loop–axis structure before the pairing or synapsis of X and Y chromosomes at the transition between zygonema and pachynema. The chromosome axis comprises the meiosis-specific axial proteins SYCP2, SYCP3, HORMAD1 and HORMAD2; cohesin subunits (only REC8 is represented); and RMMAI proteins (REC114, MEI4, MEI1, ANKRD31 and IHO1). On the non-PAR portion of the Y chromosome axis (left), RMMAI protein loading and DSB formation are partly dependent on HORMAD1 and ANKRD31, and strictly dependent on MEI4, REC11419, IHO121 and, presumably, MEI118. The DNA is organized into large loops, with a low number of axis-associated RMMAI foci. By contrast, in the PAR (right), the hyperaccumulation of RMMAI proteins at mo-2 minisatellites (possibly spreading into the adjacent chromatin) promotes the elongation and subsequent splitting of the PAR sister chromatid axes. Short mo-2-containing chromatin loops stretch along this extended PAR axis, which increases the physical distance between the PAR boundary and the distal PAR sequences, including the telomere. The degree of RMMAI protein loading, PAR axis differentiation and DSB formation are proportional to the mo-2 FISH signal (which we interpret as reflecting mo-2 copy number), and depend on MEI4, ANKRD31 and, presumably, REC114.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion and Supplementary References.
Rights and permissions
About this article
Cite this article
Acquaviva, L., Boekhout, M., Karasu, M.E. et al. Ensuring meiotic DNA break formation in the mouse pseudoautosomal region. Nature 582, 426–431 (2020). https://doi.org/10.1038/s41586-020-2327-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2327-4
- Springer Nature Limited
This article is cited by
-
Divergence and conservation of the meiotic recombination machinery
Nature Reviews Genetics (2024)
-
Seeding the meiotic DNA break machinery and initiating recombination on chromosome axes
Nature Communications (2024)
-
Recent advances in mechanisms ensuring the pairing, synapsis and segregation of XY chromosomes in mice and humans
Cellular and Molecular Life Sciences (2024)
-
Meiosis through three centuries
Chromosoma (2024)
-
Bi-allelic MEI1 variants cause meiosis arrest and non-obstructive azoospermia
Journal of Human Genetics (2023)