Abstract
The L-type amino acid transporter 1 (LAT1; also known as SLC7A5) catalyses the cross-membrane flux of large neutral amino acids in a sodium- and pH-independent manner1,2,3. LAT1, an antiporter of the amino acid–polyamine–organocation superfamily, also catalyses the permeation of thyroid hormones, pharmaceutical drugs, and hormone precursors such as l-3,4-dihydroxyphenylalanine across membranes2,3,4,5,6. Overexpression of LAT1 has been observed in a wide range of tumour cells, and it is thus a potential target for anti-cancer drugs7,8,9,10,11. LAT1 forms a heteromeric amino acid transporter complex with 4F2 cell-surface antigen heavy chain (4F2hc; also known as SLC3A2)—a type II membrane glycoprotein that is essential for the stability of LAT1 and for its localization to the plasma membrane8,9. Despite extensive cell-based characterization of the LAT1–4F2hc complex and structural determination of its homologues in bacteria, the interactions between LAT1 and 4F2hc and the working mechanism of the complex remain largely unknown12,13,14,15,16,17,18,19. Here we report the cryo-electron microscopy structures of human LAT1–4F2hc alone and in complex with the inhibitor 2-amino-2-norbornanecarboxylic acid at resolutions of 3.3 Å and 3.5 Å, respectively. LAT1 exhibits an inward open conformation. Besides a disulfide bond association, LAT1 also interacts extensively with 4F2hc on the extracellular side, within the membrane, and on the intracellular side. Biochemical analysis reveals that 4F2hc is essential for the transport activity of the complex. Together, our characterizations shed light on the architecture of the LAT1–4F2hc complex, and provide insights into its function and the mechanisms through which it might be associated with disease.
Similar content being viewed by others
Main
We isolated the human LAT1–4F2hc complex from HEK293F cells by recombinant expression. After purification, LAT1 and 4F2hc were examined by Coomassie blue staining of sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) gels (Fig. 1a). We reconstituted the complex into liposomes and performed a counterflow assay in which the uptake of the substrate [3H]leucine into proteoliposomes was monitored. Time-course analysis showed a typical profile for counterflow assays20 (Fig. 1b). The velocity of the enzyme-catalysed reaction at infinite concentration of substrate (Vmax) and the Michaelis constant (Km) for the wild-type LAT1–4F2hc complex were approximately 77.3 ± 3.9 nmol mg−1 min−1 and 111.6 ± 14.0 µM, respectively (mean ± s.d.) (Fig. 1c). The Km value was similar to that of T24 cells (~100 µM) and approximately six times larger than that of Xenopus oocytes (~20 µΜ)1,7,8.
a, Representative SEC purification of the LAT1(A36E)–4F2hc complex. The protein complex was purified in the presence of digitonin. Inset, SDS–PAGE under reducing (+dithiothreitol (DTT)) or oxidizing (−DTT) conditions, visualized by Coomassie blue staining. The experiment was repeated three times independently with similar results. M, molecular mass marker. b, Liposome-based counterflow assay for the purified wild-type complex. A typical time course of [3H]leucine uptake into liposomes is shown. c, Km and Vmax values for the wild-type complex. Inset, the [3H]leucine uptake in the first 60 s is almost linear, which enables the calculation of the initial rates within the first 30 s. d, Competition assay for Leu transport by the wild-type complex. See Extended Data Fig. 1a, b for the IC50 values of these inhibitors. e, 4F2hc is essential for the transport activity of LAT1. Data in b–e are mean ± s.d. of three independent experiments. Each dot represents a technical replicate.
Results of cold-competition experiments showed that the transport of [3H]leucine was notably inhibited by the large neutral amino acids Phe, Trp, Leu, Tyr, l-3,4-dihydroxyphenylalanine (l-DOPA), His, Met, 2-amino-2-norbornanecarboxylic acid (BCH), and Ile (Fig. 1d). We measured the half-maximal inhibitory concentration (IC50) of these compounds for the inhibition of [3H]leucine transport; all displayed an IC50 value at the micromolar level (Extended Data Fig. 1a, b).
We also assessed the inhibition of the LAT1–4F2hc complex by the LAT1-specific inhibitor, JPH2033. Consistent with previous reports, JPH203 had a more potent inhibitory effect than the large neutral amino acids that we tested. However, owing to the low solubility of JPH203, its IC50 value could not be reliably derived (Extended Data Fig. 1a, b).
To investigate whether 4F2hc contributes to the transport activity of the complex, we purified LAT1 alone. The elution volumes of the size exclusion chromatography (SEC) that corresponded to LAT1 alone were reinjected to remove endogenous 4F2hc (Extended Data Fig. 1c). No transport activity was detected for LAT1 alone (Fig. 1e), indicating that 4F2hc is essential for the transport activity of the complex.
The cryo-electron microscopy (cryo-EM) structures of the LAT1(A36E)–4F2hc complex incubated with BCH or JPH203 were determined at 3.5 Å or 3.3 Å resolution, respectively (Extended Data Figs. 2–4). The A36E mutation was an unexpected variant that might have been derived from the cDNA library used for subcloning, or as a result of polymerase chain reaction (PCR) error. The transport activity of this variant was similar to the wild-type complex (Extended Data Fig. 1d). Nevertheless, we also corrected this mutation and determined the structure of the apo wild-type complex at 4.0 Å resolution (Extended Data Fig. 2h–k). These three samples will hereafter be referred to as heteromeric amino acid transporter (HAT) + BCH, HAT + JPH203, and apo-HAT, respectively. The three overall structures are nearly identical (Extended Data Fig. 2l). A density with a shape consistent with that of a BCH molecule was present in the cryo-EM map of HAT + BCH, but was absent in apo-HAT, supporting the assignment of BCH (Fig. 2a, Extended Data Fig. 4c). Despite the effective inhibition, no density corresponding to JPH203 was observed in the cryo-EM map for HAT + JPH203. As JPH203 is strongly hydrophobic, it is possible that JPH203 may favour binding to the detergent micelles (by contrast, the counterflow assay was performed in proteoliposomes). In the following sections, we focus mainly on the structure of HAT + BCH (Fig. 2).
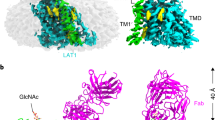
a, Cryo-EM map of the human LAT1–4F2hc complex in the presence of the inhibitor BCH. 4F2hc and LAT1 are coloured wheat and cyan, respectively. Inset, the density that corresponds to BCH is coloured yellow. b, Topology of the complex. The transmembrane segments are numbered from 1 to 12. c, Overall structure of the complex. The glycosylation moieties are shown as sticks. For the complex bound to BCH, LAT1 carries an A36E mutation. ECD, extracellular domain; H, helix; TM, transmembrane domain.
The N-terminal residues 1–162 for 4F2hc and 1–50 for LAT1 were not visible in the cryo-EM map. The extracellular loop 5–6 (EL5–6) of LAT1—which connects the transmembrane segments TM5 and TM6—was poorly resolved because of its flexibility, and the side chains in this loop were not assigned. With the exception of these areas, the sequences of both LAT1 and 4F2hc were clearly resolved.
The extracellular domain (residues 213–631) of 4F2hc contains four glycosylation sites and is connected to the transmembrane helix with a short linker (Fig. 2b, c). The transmembrane helices of 4F2hc and LAT1 traverse the membrane at a highly tilted angle (Fig. 2c). The 12 transmembrane segments of LAT1 are arranged in a canonical LeuT fold. TM1 and TM6 of LAT1 are disrupted by a short loop, and the half helices are named TM1a/1b and TM6a/6b (Fig. 2b, c).
There are several helical and loop segments in the solvent-exposed region of the complex. The intracellular loop 2–3 (IL2–3) between TM2 and TM3 of LAT1 contains a short helix, H1, and the extracellular loop EL7–8 between TM7 and TM8 contains two short helices, H2 and H3. In the C-terminal loop that follows TM12 of LAT1 is an amphipathic helix, H4. The C-terminal end of H4 approaches the transmembrane segment of 4F2hc and is located above the short helix H1′ in the N-terminal loop that precedes the transmembrane segment of 4F2hc (Fig. 2b, c).
Surface analysis showed that LAT1 exhibits an inward open conformation, with BCH bound in close proximity to the side chain of Phe252 (Fig. 3a, b) and in the centre of the putative transport path—a location similar to the substrate-binding site in the amino acid transporters GkApcT and AdiC15,17 (Extended Data Fig. 5). The hydrophobic moiety of BCH participates in hydrophobic interactions with the benzene ring of Phe252, whereas the carboxyl group and amino group form hydrogen bonds with the main-chain atoms of TM1 and TM6 of LAT1, respectively (Fig. 3b). Notably, Phe252 corresponds to Phe231 in GkApcT, Phe253 in LeuT and Trp202 in AdiC, all of which have previously been shown to be critical for substrate binding15,17,21.
a, LAT1 adopts an inward open conformation with BCH (yellow sticks) bound in the centre of the putative transport path. b, The BCH-binding site. The BCH density, contoured at 6σ, is shown as blue mesh. c, The intracellular vestibule of the transport path. Some of the key residues that are involved in transport are shown as green sticks. d, Mutations of the transport-path residues lead to a marked reduction in transport activity. WT, wild type. Data are mean ± s.d. of three independent experiments. Each dot represents a technical replicate.
On the basis of structural analysis and sequence alignment with other amino acid antiporters (Extended Data Fig. 6), we characterized six potential gating residues on LAT1—Tyr117, Phe252, Trp257, Asn258, Tyr259, and Arg348—all of which are located along the putative transport path (Fig. 3c). Mutation of each residue to Ala led to a marked decrease in transport activity compared with the wild-type complex (Fig. 3d).
Tyr117 is located on the H1 helix of LAT1, which forms hydrogen bonds with Glu356 and His358 at the end of TM8 (Fig. 3c). The Y117A mutant retained less than 10% of its transport activity compared with the wild type, suggesting a key role of Tyr117. Consistent with a previous study22, LAT1(F252A) displayed almost no transport activity. The three consecutive residues Trp257, Asn258, and Tyr259 are located on the unwound region of TM6. When these residues were mutated to Ala, the W257A variant almost completely lost transport activity, and the N258A and Y259A variants retained approximately 25% and 20% transport activities, respectively (Fig. 3d).
Asn258 of LAT1 corresponds to Glu208 of AdiC; this is an essential residue that is thought to be located in the distal gate16. To further investigate the functional effect of Asn258, we designed and purified three different variants: N258A, N258D, and N258Q. The transport activities of the N258A and N258Q mutants both decreased to approximately 30% compared with the wild type, and that of N258D was reduced to less than 10% (Fig. 3d).
4F2hc interacts extensively with LAT1 via several interfaces (Fig. 4a). Confirming previous characterizations, a disulfide bond is present between Cys211 of 4F2hc and Cys164 of LAT15 (Fig. 4b). On the extracellular side, 4F2hc interacts with LAT1 mainly via polar interactions mediated by Lys533 and Arg535 of 4F2hc, and Thr163, Glu303, and Gln304 of LAT1 (Fig. 4b). Notably, Lys533 and Arg535 are conserved in 4F2hc homologues among different species (Extended Data Fig. 7a).
a, 4F2hc interacts with LAT1 via several interfaces. Left, the coloured boxes correspond to those analysed in detail in b, d, and e. Right, the interaction of 4F2hc and LAT1 is restricted to one side of LAT1, away from the transport path. b, The extracellular interface. Polar interactions are shown as red dashed lines. EL, extracellular loop. c, The extracellular interactions between 4F2hc and LAT1 are important for transport activity. Charge-swapping mutations (4F2hc(K533E) and LAT1(E303K)) rescue the transport activity. d, The hydrophobic interactions between the transmembrane segment of 4F2hc and TM4 of LAT1. e, The intracellular interface. Bound lipids are shown as black sticks. IL, intracellular loop. f, Arg183 of 4F2hc is involved in lipid coordination and is critical for transport activity. LAT1(ΔC), LAT1 with residues 483–507 (including the C-terminal helix H4) deleted. g, Model for the putative working mechanism of the complex. 4F2hc may facilitate transport by stabilizing the scaffolding domain of LAT1 during an alternating-access cycle. Data in c and f are presented as mean ± s.d. of three independent experiments. Each dot represents a technical replicate.
Lys533 of 4F2hc appears to interact with Glu303 of LAT1. Single-point mutations of these residues to 4F2hc(K533E) and LAT1(E303K) reduced transport activities to approximately 35% and 60%, respectively, compared with the wild type. The charge-swapped mutations 4F2hc(K533E) and LAT1(E303K) completely restored transport activity. Truncation of the 4F2hc extracellular domain residues 235–631 (4F2hc(ΔC)) resulted in a loss of activity that was greater than 90% (Fig. 4c). Together, these results suggest that the extracellular interface is important for the transport activity of the complex.
In the membrane region, the transmembrane segment of 4F2hc interacts with TM4 of LAT1 via extensive hydrophobic interactions (Fig. 4d) that are mediated by highly conserved residues (Extended Data Figs. 6, 7b).
Two lipid-like densities were observed on the intracellular side of the membrane. One aliphatic tail was better resolved than the other for both molecules (Extended Data Fig. 8a–c). The discernible densities enabled the modelling of two phosphatidic acids, with their polar moieties each inserting into a positively charged pocket (Fig. 4e, Extended Data Fig. 8d). The first of these two pockets is lined with the transmembrane segment of 4F2hc, together with the LAT1 components TM4, TM9, the C-terminal end of the H4 helix, and the ensuing C-terminal loop. The second pocket consists of TM8, IL8–9, TM12, and the H4 N-terminal end of LAT1, together with the N-terminal loop of 4F2hc (Fig. 4e). Within the first pocket, Arg183 of 4F2hc appears to interact with the polar head of the first lipid molecule (Fig. 4e, Extended Data Fig. 8d left panel). To investigate the role of this pocket, we generated five mutants (R183A, R183L, R183E, R183K, and LAT1(ΔC)), all of which displayed a pronounced reduction in transport activity (Fig. 4f).
There is also an elongated density in a cleft enclosed by the LAT1 transmembrane helices TM3, TM9, TM10, and TM12 (Extended Data Fig. 8e). Although this might correspond to a digitonin, its structure suggests that a physiologically relevant cholesterol molecule could be accommodated23. Supporting this notion, transport activity was enhanced by more than twofold when 10% cholesterol was added into the proteoliposomes (Extended Data Fig. 8f).
The structures we have described here provide a molecular framework that can be used to investigate disease-related mutations. The A246V and P375L mutations in LAT1 are related to autism spectrum disorders6. Both of these mutants, in complex with 4F2hc, showed similar behaviour in solution to the wild type (Extended Data Fig. 9a). However, the transport activity for each of these mutants was less than 5% that of the wild-type complex (Extended Data Fig. 9b). As Ala246 is located on TM6a, the substitution of this residue with Val might affect the helix packing between TM1 and TM6 of LAT1 (Extended Data Fig. 9c). Pro375—which is on the intracellular tip of TM9—appears to be a coordinating centre for the surrounding hydrophobic residues on IL8–9 and H4, as well as the aliphatic tails of the two bound lipids (Extended Data Fig. 9d). Substitution of Pro with Leu—which is a bulkier residue—might alter the local conformation and affect binding with the nearby 4F2hc, hence reducing the transport activity.
4F2hc has a single transmembrane segment that is similar to many other auxiliary subunits—for example, the β subunits in eukaryotic sodium channels, and neuroplastin in human plasma membrane Ca2+ ATPases24. Previous studies have shown that 4F2hc is required for the cellular localization and stability of LAT18,9. Here, using purified proteins, we have shown that 4F2hc is also essential for the transport activity of LAT1.
Notably, the contact between 4F2hc and LAT1 is restricted to one side of LAT1 (Fig. 4a, right panel), and TM1 and TM6 of LAT1—which are conformational switches that are critical for the alternating-access transport mechanisms of LeuT-fold transporters—are positioned away from the coordination of 4F2hc. 4F2hc therefore appears to stabilize the scaffolding domain of LAT1 in the membrane, which may facilitate the local conformational shifts of gating elements such as TM1, TM2, TM6, and TM10 during an alternating-access cycle (Fig. 4g).
Methods
Protein expression and purification
The cDNAs of full-length LAT1 (NCBI reference sequence NM_003486.7) and 4F2hc (isoform b, NCBI reference sequence NM_001012662.2) were subcloned separately into pCAG. An N-terminal Flag tag was fused to LAT1, and an N-terminal 8× His tag was fused to 4F2hc. The mutation was introduced using a standard two-step PCR.
HEK293F cells (Invitrogen) were cultured in SMM 293T-I medium (Sino Biological) at 37 °C under 5% CO2 in a Multitron-Pro shaker (Infors, 130 r.p.m.). Cells were not tested for mycoplasma contamination. When the cell density reached 2.0 × 106 cells per ml, the cells were transiently transfected with the plasmids and polyethylenimines (Polysciences). Approximately 0.75 mg of the LAT1 plasmid and 0.75 mg of the 4F2hc plasmid were premixed with 3 mg of polyethylenimines in 50 ml of fresh medium for 15–30 min before transfection. For the expression of LAT1 alone, 1.5 mg of the LAT1 plasmid was used in the premixture before transfection.
For transfection, 50 ml of the mixture was added to one litre of cell culture, and incubated with the cells for 15–30 min. The transfected cells were cultured for 48 h before collection.
For purification of the LAT1–4F2hc complex, the cells were collected and resuspended in a buffer containing 25 mM Tris (pH 8.0), 150 mM NaCl,and a protease inhibitor mixture that included aprotinin (0.8 µM, AMRESCO), pepstatin (2 µM, AMRESCO), and leupeptin (2.5 µM, AMRESCO).
For incubation of the complex sample with BCH, the membrane fraction was solubilized at 4 °C for 2 h with 1% (w/v) digitonin (Sigma). After ultracentrifugation at 100,000g for 1 h, the supernatant was collected and loaded onto anti-Flag M2 affinity resin (Sigma). The resin was rinsed with the wash buffer containing 25 mM Tris (pH 8.0), 150 mM NaCl, and 0.08% digitonin (w/v). The protein was eluted with wash buffer supplemented with 0.2 mg ml−1 Flag peptide. The eluent was then applied to nickel resin (Ni-NTA, Qiagen). The resin was rinsed with wash buffer supplemented with 20 mM imidazole. The protein complex was eluted from the nickel resin with wash buffer supplemented with 300 mM imidazole. The eluent was then concentrated and subjected to SEC (Superose 6 Increase 10/300 GL, GE Healthcare) in buffer containing 25 mM Tris (pH 8.0), 150 mM NaCl, and 0.08% digitonin. The peak fractions were pooled and concentrated for electron microscopy analysis. The purification procedure of wild-type LAT1–4F2hc protein for electron microscopy analysis was the same.
For incubation of the complex sample with JPH203, the membrane fraction was solubilized at 4 °C for 2 h with 1% (w/v) lauryl maltose neopentyl glycol (LMNG) (Anatrace) supplemented with 0.1% (w/v) cholesteryl hemisuccinate Tris salt (Anatrace). The buffer used for affinity chromatography and gel filtration was the same as that described above except for the detergent, which was changed from 0.08% digitonin to 0.05% GDN (Anatrace).
For the transport-activity assay, the membrane fraction was solubilized at 4 °C for 2 h with 1% (w/v) LMNG (Anatrace) supplemented with 0.1% (w/v) cholesteryl hemisuccinate Tris salt (Anatrace). The buffer used for affinity chromatography and gel filtration was the same as that used for cryo-sample preparation except for the detergent, which was changed from 0.08% digitonin to 0.01% LMNG supplemented with 0.001% cholesteryl hemisuccinate Tris salt. For purification of LAT1 alone, the affinity purification step using nickel resin was omitted and one more step of SEC was added. After the first step of SEC, the fractions of LAT1 alone were concentrated and SEC was applied again to further separate the LAT1–4F2hc complex formed by overexpressed LAT1 and endogenous 4F2hc from LAT1 alone.
Preparation of liposomes and proteoliposomes
Escherichia coli polar lipids (Avanti) supplemented with 10% (wt%) cholesterol (Sigma) were dissolved at a concentration of 20 mg ml−1 in chloroform:methanol (3:1, v/v). The lipid mixture was dried under nitrogen in clean glass bottles. After solvent evaporation, thin lipid films were further dried under vacuum for 2 h. The lipids were resuspended at a concentration of 20 mg ml−1 with buffer containing 20 mM potassium phosphate (pH 6.5), 150 mM KCl, and 5 mM l-leucine (Sigma). After 10 rounds of freezing and thawing by liquid nitrogen, the liposomes were extruded through 0.4-µm polycarbonate membranes (Avanti) up to 21 times.
The liposomes were pre-incubated with 1% n-octyl-β-d-glucoside (β-OG) for 30 min at 4 °C before protein incorporation. The protein was incubated with liposomes for 60 min at a concentration of 10 µg per mg of lipid. Then, β-OG was removed by incubation overnight with 260 mg ml−1 Bio-Beads SM2 (Bio-Rad), and the resulting solution was further incubated with 90 mg ml−1 Bio-Beads for an additional hour. The proteoliposomes were collected by ultracentrifugation at 4 °C for 1 h at 100,000g and rinsed twice with resuspension buffer (20 mM potassium phosphate (pH 6.5) and 150 mM KCl). The proteoliposomes were resuspended in resuspension buffer to a final lipid concentration of 50 mg ml−1.
In vitro transport-activity assay
All transport-activity assays were performed at room temperature. The reaction was initiated by adding 100 µl of reaction buffer containing 20 mM potassium phosphate (pH 6.5), 150 mM KCl, and 0.08 µM (1 µCi) l-[3H]leucine (PerkinElmer Life Sciences) to 4 µl of proteoliposome. [3H]leucine uptake was stopped after 2 min by rapidly filtering the reaction solution through a 0.22-µm GSTF filter (Millipore) and washing with 2 ml of ice-cold wash buffer (20 mM potassium phosphate (pH 6.5) and 150 mM KCl). The filter was then used for liquid scintillation counting.
The substrate competition assay for the LAT1–4F2hc complex was performed in the presence of 1 mM unlabelled substrate outside the proteoliposome. The measurements of Km and Vmax were performed in the presence of unlabelled leucine at the indicated concentrations in reaction buffer outside the proteoliposome, and the uptake of [3H]leucine was stopped after 30 s.
IC50 values were measured by adding unlabelled substrates at the indicated concentrations in reaction buffer outside the proteoliposome. For measurement of the IC50 value of JPH203, 4 µl proteoliposome was pre-incubated with the indicated concentration of JHP203 in a final volume of 5 µl for 30 min before the addition of reaction buffer. The uptake of [3H]leucine was stopped after 2 min. Liposome containing no protein was introduced as an empty control for each measurement. The data for liposome-based transport assays were analysed by GraphPad Prism 7.00. For all of the counterflow activities included in the manuscript, the no-protein control reading—which was 5–10% of the reading of wild type LAT1–4F2hc in the raw data—has been subtracted.
Cryo-EM sample preparation and data acquisition
The purified LAT1–4F2hc complex was concentrated to approximately 15 mg ml−1 and incubated with 10 mM BCH or 0.2 mM JPH203 for 2 h before being applied to the grids. For clarity, the samples are hereafter referred to as LAT1–4F2hc + BCH and LAT1–4F2hc + JPH203. Aliquots (4 µl) of the protein complex were placed on glow-discharged holey carbon grids (Quantifoil Cu R1.2/1.3, 300 mesh). Then, the grids were blotted for 3.5 s and flash-frozen in liquid ethane cooled by liquid nitrogen with Vitrobot Mark IV (Thermo Fisher Scientific). The prepared grids were transferred to a Titan Krios, operating at 300 kV, which was equipped with a Gatan K2 Summit detector and a GIF Quantum energy filter. Totals of 2,700, 5,908, and 5,207 movie stacks were automatically collected using AutoEMation25 for LAT1–4F2hc + BCH, LAT1–4F2hc + JPH203, and apo wild-type LAT1–4F2hc, respectively, with a slit width of 20 eV on the energy filter and a preset defocus range of −1.8 µm to −1.5 µm in super-resolution mode at a nominal magnification of 105,000×. Each stack was exposed for 5.6 s with an exposure time of 0.175 s per frame, resulting in a total of 32 frames per stack. The total dose rate was approximately 48 e− per Å2 for each stack. The stacks were motion corrected with MotionCor226 and binned twofold, resulting in a pixel size of 1.091 per pixel. Dose weighting27 was performed. The defocus values were estimated with Gctf28.
Data processing
Totals of 1,247,297, 3,775,582, and 2,507,734 particles were automatically picked from 2,458, 5,623, and 4,293 manually selected micrographs, using RELION-229,30,31,32, for LAT1–4F2hc + BCH, LAT1–4F2hc + JPH203, and apo wild-type LAT1–4F2hc, respectively. After 2D classification, totals of 1,128,961, 2,425,306, and 1,218,886 particles were selected for LAT1–4F2hc + BCH, LAT1–4F2hc + JPH203, and apo wild-type LAT1–4F2hc, respectively. The selected LAT1–4F2hc + BCH particles were subjected to several rounds of 3D classification with a Gaussian sphere as an initial model to obtain a suitable reference map. Then, a global angular searching 3D classification against this map with only one class was performed. For each of the last several iterations of the global angular searching 3D classification, a local angular searching 3D classification was performed, during which the particles were classified into four classes. In total, 1,120,028 non-redundant LAT1–4F2hc + BCH particles were selected from the local angular searching 3D classification. The selected LAT1–4F2hc + JPH203 and apo wild-type LAT1–4F2hc particles were processed with similar procedures, except that the template for global angular searching was obtained from the 3D map of LAT1–4F2hc + BCH. Totals of 1,707,839 and 714,680 non-redundant particles were selected from the local angular searching 3D classification for LAT1–4F2hc + JPH203 and apo wild-type LAT1–4F2hc, respectively. The selected particles were then subjected to several cycles of multi-reference 3D classification with local defocus values calculated with Gctf28 for each particle. The final resolution of the 3D auto-refinement after post-processing was 3.5 Å for LAT1–4F2hc + BCH, with a final particle number of 262,949; 3.3 Å for LAT1–4F2hc + JPH203, with a final particle number of 480,596; and 4.0 Å for apo wild-type LAT1–4F2hc, with a final particle number of 308,781. To improve the BCH density quality, the LAT1–4F2hc + BCH map was further focused by applying a spherical mask covering BCH during refinement; this resulted in a 3.7 Å resolution map with better density for BCH. The 2D classification, 3D classification, and auto-refinement were performed with RELION-2 or RELION-3. The resolution was estimated with the gold-standard Fourier shell correlation 0.143 criterion with high-resolution noise substitution.
Model building and structure refinement
Model building of the LAT1–4F2hc complex was based on the 3.5 Å cryo-EM map of LAT1–4F2hc + BCH. The structure of the extracellular domain of 4F2hc (Protein Data Bank (PDB) ID 2DH2) was fitted into the density map using Chimera33. The transmembrane helices were modelled with EMBuilder34. The subsequent modelling was performed in Coot35 with aromatic residues as land markers, as most of these residues were clearly visible in our cryo-EM map. Each residue was manually checked with the chemical properties considered during model building.
A total of 926 amino acid residues were constructed for the complex structure, with 909 side chains assigned. The N-terminal sequences of both 4F2hc and LAT1 were not modelled as the corresponding densities were not visible on the map. Eight sugar moieties and two lipid moieties were assigned according to map density. A downloaded 3D conformer of BCH (PubChem substance ID: 24891291) was processed with ‘phenix.elbow’ in PHENIX36, and the resulting structure was rigid-body fitted into the density map using Coot. The model for the LAT1–4F2hc + JPH203 complex was generated based on the 3.3 Å cryo-EM map of LAT1–4F2hc + JPH203, using the model of the LAT1–4F2hc + BCH complex as a starting model. JPH203 was not modelled as its corresponding density was not visible.
Structure refinement was performed with ‘phenix.real_space_refine’ in PHENIX, with secondary structural and geometric restraints to prevent structure overfitting. The final atomic model was evaluated using MolProbity37. To monitor the overfitting of the model, the model was refined against one of the two independent half maps from the gold-standard 3D refinement approach. Then, the refined model was tested against the other map38. For statistics associated with data collection, 3D reconstruction, and model refinement, see Extended Data Table 1.
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The atomic coordinates of the BCH-bound LAT1–4F2hc complex and the JPH203-incubated LAT1–4F2hc complex have been deposited in the PDB (http://www.rcsb.org) under the accession codes 6IRT and 6IRS, respectively. The overall electron microscopy map and the BCH-focused refined electron microscopy map of the BCH-bound LAT1–4F2hc complex, the electron microscopy map of the JPH203-incubated LAT1–4F2hc complex, and the electron microscopy map of the apo wild-type LAT1–4F2hc complex have been deposited in the Electron Microscopy Data Bank (https://www.ebi.ac.uk/pdbe/emdb/) under the accession codes EMD-9722, EMD-0679, EMD-9721, and EMD-0678, respectively. Source Data for Figs. 1b–e, 3d, 4c, f and Extended Data Figs. 1a, d, 8f, 9b are available online. All other data are available from the corresponding author upon reasonable request.
References
Kanai, Y. et al. Expression cloning and characterization of a transporter for large neutral amino acids activated by the heavy chain of 4F2 antigen (CD98). J. Biol. Chem. 273, 23629–23632 (1998).
del Amo, E. M., Urtti, A. & Yliperttula, M. Pharmacokinetic role of L-type amino acid transporters LAT1 and LAT2. Eur. J. Pharm. Sci. 35, 161–174 (2008).
Fotiadis, D., Kanai, Y. & Palacín, M. The SLC3 and SLC7 families of amino acid transporters. Mol. Aspects Med. 34, 139–158 (2013).
Kageyama, T. et al. The 4F2hc/LAT1 complex transports l-DOPA across the blood–brain barrier. Brain Res. 879, 115–121 (2000).
Friesema, E. C. et al. Thyroid hormone transport by the heterodimeric human system L amino acid transporter. Endocrinology 142, 4339–4348 (2001).
Tărlungeanu, D. C. et al. Impaired amino acid transport at the blood brain barrier is a cause of autism spectrum disorder. Cell 167, 1481–1494.e18 (2016).
Kim, D. K. et al. Characterization of the system L amino acid transporter in T24 human bladder carcinoma cells. Biochim. Biophys. Acta 1565, 112–122 (2002).
Yanagida, O. et al. Human L-type amino acid transporter 1 (LAT1): characterization of function and expression in tumor cell lines. Biochim. Biophys. Acta 1514, 291–302 (2001).
Verrey, F. et al. CATs and HATs: the SLC7 family of amino acid transporters. Pflugers Arch. Eur. J. Physiol. 447, 532–542 (2004).
Scalise, M., Galluccio, M., Console, L., Pochini, L. & Indiveri, C. The human SLC7A5 (LAT1): the intriguing histidine/large neutral amino acid transporter and its relevance to human health. Front. Chem. 6, 243 (2018).
Kandasamy, P., Gyimesi, G., Kanai, Y. & Hediger, M. A. Amino acid transporters revisited: new views in health and disease. Trends Biochem. Sci. 43, 752–789 (2018).
Fort, J. et al. The structure of human 4F2hc ectodomain provides a model for homodimerization and electrostatic interaction with plasma membrane. J. Biol. Chem. 282, 31444–31452 (2007).
Meury, M. et al. Detergent-induced stabilization and improved 3D map of the human heteromeric amino acid transporter 4F2hc-LAT2. PLoS ONE 9, e109882 (2014).
Rosell, A. et al. Structural bases for the interaction and stabilization of the human amino acid transporter LAT2 with its ancillary protein 4F2hc. Proc. Natl Acad. Sci. USA 111, 2966–2971 (2014).
Jungnickel, K. E. J., Parker, J. L. & Newstead, S. Structural basis for amino acid transport by the CAT family of SLC7 transporters. Nat. Commun. 9, 550 (2018).
Gao, X. et al. Structure and mechanism of an amino acid antiporter. Science 324, 1565–1568 (2009).
Gao, X. et al. Mechanism of substrate recognition and transport by an amino acid antiporter. Nature 463, 828–832 (2010).
Fang, Y. et al. Structure of a prokaryotic virtual proton pump at 3.2 Å resolution. Nature 460, 1040–1043 (2009).
Ma, D. et al. Structure and mechanism of a glutamate-GABA antiporter. Nature 483, 632–636 (2012).
Garcia, M. L., Viitanen, P., Foster, D. L. & Kaback, H. R. Mechanism of lactose translocation in proteoliposomes reconstituted with lac carrier protein purified from Escherichia coli. 1. Effect of pH and imposed membrane potential on efflux, exchange, and counterflow. Biochemistry 22, 2524–2531 (1983).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl−-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Napolitano, L. et al. Novel insights into the transport mechanism of the human amino acid transporter LAT1 (SLC7A5). Probing critical residues for substrate translocation. Biochim. Biophys. Acta Gen. Subj. 1861, 727–736 (2017).
Dickens, D. et al. Modulation of LAT1 (SLC7A5) transporter activity and stability by membrane cholesterol. Sci. Rep. 7, 43580 (2017).
Gong, D. et al. Structure of the human plasma membrane Ca2+-ATPase 1 in complex with its obligatory subunit neuroplastin. Nat. Commun. 9, 3623 (2018).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zhou, N., Wang, H. & Wang, J. EMBuilder: a template matching-based automatic model-building program for high-resolution cryo-electron microscopy maps. Sci. Rep. 7, 2664 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Acknowledgements
We thank the Tsinghua University branch of China National Center for Protein Sciences (Beijing) for cryo-EM and computational facility support, and X. Li for technical support in electron microscopy data acquisition. This work was funded by the National Key Basic Research (973) Program (2015CB910101) and the National Key R&D Program (2016YFA0500402) from the Ministry of Science and Technology of China, and by the National Natural Science Foundation of China (projects 31621092, 31630017, and 81861138009).
Reviewer information
Nature thanks David Dickens, Simon Newstead and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Q.Z. and R.Y. conceived the project. R.Y. and X.Z. prepared the samples. Q.Z., R.Y., X.Z., and J.L. collected the electron microscopy data. Q.Z. processed the electron microscopy data and built the atomic model. Q.Z., R.Y., and X.Z. designed, analysed, and performed the biochemical experiments. Q.Z., R.Y., and X.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization of the LAT1–4F2hc complex.
a, IC50 measurement of eight amino acid substrates (Phe, Trp, Leu, Tyr, l-DOPA, His, Met, and Ile) and two inhibitors (JPH203and BCH) for the inhibition of Leu uptake into the LAT1–4F2hc proteoliposomes. Owing to the low solubility of JPH203, its inhibitory effect could not be measured above 2 µM. b, The IC50 values for the listed inhibitors. NA, not available. c, SEC purification. Left, LAT1 alone was eluted ~1.5 ml after the LAT1–4F2hc complex in the presence of 0.01% LMNG and 0.001% cholesteryl hemisuccinate (CHS). Right, the peak fractions were subjected to SDS–PAGE and visualized by Coomassie blue staining. The experiment was repeated twice independently with similar results. d, The LAT1(A36E) variant has similar transport activity to the wild type in the presence of 4F2hc. The measured Km and Vmax values for the LAT1(A36E)–4F2hc complex were approximately 162.3 ± 23.6 µM and 82.5 ± 5.4 nmol mg−1 min−1, respectively. Data in a and f are mean ± s.d. of three technical independent experiments.
Extended Data Fig. 2 Cryo-EM analysis of the LAT1–4F2hc complex.
a, Representative cryo-EM micrograph and 2D class averages. The raw images are similar for all three samples; HAT + BCH is shown. The scale bar for the 2D class averages (right images) is 10 nm. b, Euler angle distribution of the 3D reconstruction for LAT1–4F2hc in the presence of BCH. c, Local-resolution maps for HAT + BCH. d, Fourier shell correlation (FSC) curves of the model refined against the overall 3.5 Å map for HAT + BCH (black); against the first half map versus the same map (red); and against the first half map versus the second half map (green). The small difference between the red and green curves indicates that the refinement of the atomic coordinates was not affected by overfitting. e–g and h–j are the same as b–d, but for HAT + JPH203 and apo-HAT, respectively. For apo-HAT, only the FSC curve of the atomic model versus the electron microscopy map was calculated. k, Gold standard FSC curves for the 3D refinement of HAT + BCH (blue), BCH-focused 3D refinement of HAT + BCH (red), HAT + JPH203 (green), and apo-HAT (cyan). l, Overall structural comparison of HAT + BCH (cyan), HAT + JPH203 (yellow), and apo-HAT (pink).
Extended Data Fig. 3 Flow chart for cryo-EM data processing.
For details, see ‘Data processing’ in the Methods section. K, number of classifications.
Extended Data Fig. 4 Electron microscopy maps of representative segments.
a, The density for the transmembrane helix of 4F2hc. b, The densities of the 12 transmembrane segments of LAT1 in the HAT + BCH complex. c, The BCH density in the electron microscopy map (marked with an asterisk) is observed in the electron microscopy map of HAT + BCH (cyan mesh), but is absent in apo-HAT (pink mesh). All of the maps are contoured at 6σ.
Extended Data Fig. 5 Comparison of the substrate-binding sites of LAT1, GkApcT, and AdiC.
The BCH-binding site in LAT1 is conserved in GkApcT and AdiC. Similar to the coordination of Arg by GkApcT and AdiC, the carboxyl and amino groups of BCH mainly form hydrogen bonds with atoms from TM1 and TM6, respectively. LAT1, GkApcT (PDB 6F34), and AdiC (PDB 3L1L) are coloured cyan, yellow, and green, respectively.
Extended Data Fig. 6 Sequence alignment of LAT1 homologues.
The sequences were aligned using CLUSTAL 2.1 (https://www.genome.jp/tools-bin/clustalw) with manual adjustments according to the 3D structures. The five aligned sequences are LAT1 and LAT2 from Homo sapiens, ApcT from Geobacillus kaustophilus, and AdiC and GadC from E. coli. The conserved residues that are involved in hydrophobic interactions with 4F2hc and the critical residues that are involved in the transport path are outlined with a black box. Amino acids that are identical or conserved in at least four sequences are coloured red or yellow. The secondary structural elements of LAT1 are indicated above the sequence alignment. Ala36 is marked with a solid black circle. Residues that are involved in 4F2hc interaction; that line the transport path; and that are related to autism spectrum disorders are indicated with solid magenta circles; solid blue circles; and solid blue triangles, respectively.
Extended Data Fig. 7 Sequence alignment of 4F2hc homologues.
a, Residues mediating the interactions with LAT1 on the extracellular interface are conserved in 4F2hc homologues. b, Alignment of the N-terminal segment and transmembrane helix. The residues involved in the interaction with LAT1 are outlined with a black box. The sequences of 4F2hc in eight vertebrates are aligned using CLUSTAL 2.1. The sequences, from top to bottom, are from Homo sapiens, Mus musculus, Rattus norvegicus, Oryctolagus cuniculus, Felis catus, Pelodiscus sinensis, Xenopus laevis, and Danio rerio, respectively. The residues that are invariant and highly conserved are coloured red and yellow, respectively. Phe190, Trp191, Trp194, Met197, Leu198, Ile205, Lys533, and Arg535—which are mapped to the interface of the two proteins—are indicated by solid magenta circles. Arg183 in the lipid-binding pocket is indicated by a solid blue circle.
Extended Data Fig. 8 Lipid molecules bound to LAT1.
a, Densities for the bound lipids. The densities, shown in blue mesh, are contoured at 5σ. b, Multiple hydrophobic amino acid residues (spheres) coordinate the two lipids. c, The hydrophobic tails of the lipids are bound to the hydrophobic surface of the LAT1–4F2hc complex. d, The hydrophilic heads of the two lipids are both surrounded by positively charged surface patches. e, A cholesterol-like density is located in a cleft surrounded by TM3, TM9, TM10, and TM12 of LAT1. The density, shown in blue mesh, is contoured at 7σ. f, The addition of 10% (wt/wt) cholesterol during proteoliposome preparation significantly enhanced the transport activity of the LAT1–4F2hc complex. EPL, E. coli polar lipid extract. Data are mean ± s.d. of three independent experiments. Each dot represents a technical replicate.
Extended Data Fig. 9 The LAT1 mutations A246V and P375L, which are associated with autism spectrum disorders, retain residual transport activity in vitro.
a, The mutants containing A246V or P375L displayed similar behaviour in solution to the wild-type complex when subjected to SEC. The experiment was repeated three times independently with similar results. b, The LAT1(A246V) and LAT1(P375L) mutants retained less than 5% transport activity compared with the wild type. The results of the counterflow assays are shown as mean ± s.d. of three independent experiments. Each dot represents a technical replicate. c, Ala246 of LAT1 is located at the beginning of TM6 and is involved in the interaction between TM1b and TM6a. d, Phe375 of LAT1 is located on the intracellular tip of TM9, and is surrounded by hydrophobic residues and the bound lipids. Mutation of Phe375 may affect the conformation of helix H4, which is involved in 4F2hc interaction.
Supplementary information
Supplementary Figures
This file contains uncropped blot scans.
Rights and permissions
About this article
Cite this article
Yan, R., Zhao, X., Lei, J. et al. Structure of the human LAT1–4F2hc heteromeric amino acid transporter complex. Nature 568, 127–130 (2019). https://doi.org/10.1038/s41586-019-1011-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1011-z
- Springer Nature Limited
This article is cited by
-
SLC7A5 correlated with malignancies and immunotherapy response in bladder cancer
Cancer Cell International (2024)
-
BCAA mediated microbiota-liver-heart crosstalk regulates diabetic cardiomyopathy via FGF21
Microbiome (2024)
-
Light-responsive adipose-hypothalamus axis controls metabolic regulation
Nature Communications (2024)
-
Cryo-EM structure of the human Asc-1 transporter complex
Nature Communications (2024)
-
The complete assembly of human LAT1-4F2hc complex provides insights into its regulation, function and localisation
Nature Communications (2024)