Abstract
Many plant species translocate maternally synthesized specialized metabolites to the seed to protect the developing embryo and later the germinating seedling before it initiates its own de novo synthesis. While the transport route into the seed is well established for primary metabolites, no model exists for any class of specialized metabolites that move from maternal source tissue(s) to embryo. Glucosinolate seed loading in Arabidopsis depends on plasma membrane localized exporters (USUALLY MULTIPLE AMINO ACIDS MOVE IN AND OUT TRANSPORTERs, UMAMITs) and importers (GLUCOSINOLATE TRANSPORTERs, GTRs), but the critical barriers in the seed loading process remain unknown. Here we dissect the transport route of glucosinolates from their source in the reproductive organ to the embryo by re-introducing the transporters at specific apoplastic barriers in their respective mutant backgrounds. We find that UMAMIT exporters and GTR importers form a transporter cascade that is both essential and sufficient for moving glucosinolates across at least four plasma membrane barriers along the route. We propose a model in which UMAMITs export glucosinolates out of the biosynthetic cells to the apoplast, from where GTRs import them into the phloem stream, which moves them to the unloading zone in the chalazal seed coat. From here, the UMAMITs export them out of maternal tissue and ultimately, the GTRs import them into the embryo symplasm, where the seed-specific glucosinolate profile is established by enzymatic modifications. Moreover, we propose that methylsulfinylalkyl glucosinolates are the predominant mobile form in seed loading. Elucidation of the seed loading process of glucosinolates identifies barrier-specific targets for transport engineering strategies to eliminate or over-accumulate a specialized metabolite in seeds with minimal interruption of other cellular processes.



Similar content being viewed by others
Data availability
Expression data were obtained from the supporting information in ref. 16. Data on individual glucosinolate content in all experiments are supplied as Supplementary Data. Source data are provided with this paper.
Code availability
The R code for statistical analysis and plot generation are supplied as Supplementary Code.
References
Linkies, A., Graeber, K., Knight, C. & Leubner-Metzger, G. The evolution of seeds. New Phytol. 186, 817–831 (2010).
Corso, M. et al. in Plant Metabolomics in Full Swing Vol. 98 (eds Petriacq, P. & Bouchereau, A.) 35–70 (Elsevier, 2021).
Karmann, J., Müller, B. & Hammes, U. Z. The long and winding road: transport pathways for amino acids in Arabidopsis seeds. Plant Reprod. 31, 253–261 (2018).
Chen, L.-Q. et al. A cascade of sequentially expressed sucrose transporters in the seed coat and endosperm provides nutrition for the Arabidopsis embryo. Plant Cell 27, 607–619 (2015).
Braun, D. M. Phloem loading and unloading of sucrose: what a long, strange trip from source to sink. Annu. Rev. Plant Biol. 73, 553–584 (2022).
Nour-Eldin, H. H. & Halkier, B. A. Piecing together the transport pathway of aliphatic glucosinolates. Phytochem. Rev. 8, 53–67 (2009).
Xu, D. et al. Export of defensive glucosinolates is key for their accumulation in seeds. Nature 617, 132–138 (2023).
Nour-Eldin, H. H. et al. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 488, 531–534 (2012).
Müller, B. et al. Amino acid export in developing Arabidopsis seeds depends on umamit facilitators. Curr. Biol. 25, 3126–3131 (2015).
Horák, J. et al. The Arabidopsis thaliana response regulator ARR22 is a putative AHP phospho-histidine phosphatase expressed in the chalaza of developing seeds. BMC Plant Biol. 8, 77 (2008).
Mikkelsen, M. D., Naur, P. & Halkier, B. A. Arabidopsis mutants in the C-S lyase of glucosinolate biosynthesis establish a critical role for indole-3-acetaldoxime in auxin homeostasis. Plant J. 37, 770–777 (2004).
Brady, S. M. et al. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318, 801–806 (2007).
Stadler, R. & Sauer, N. The Arabidopsis thaliana AtSUC2 gene is specifically expressed in companion cells. Botanica Acta 109, 299–306 (1996).
Shimada, T. L., Shimada, T. & Hara-Nishimura, I. A rapid and non-destructive screenable marker, FAST, for identifying transformed seeds of Arabidopsis thaliana. Plant J. 61, 519–528 (2010).
Brown, P. D., Tokuhisa, J. G., Reichelt, M. & Gershenzon, J. Variation of glucosinolate accumulation among different organs and developmental stages of Arabidopsis thaliana. Phytochemistry 62, 471–481 (2003).
Khan, D. et al. Transcriptome atlas of the Arabidopsis funiculus—a study of maternal seed subregions. Plant J. 82, 41–53 (2015).
Kliebenstein, D. J. et al. Characterization of seed-specific benzoyloxyglucosinolate mutations in Arabidopsis thaliana. Plant J. 51, 1062–1076 (2007).
Lee, S. et al. Benzoylation and sinapoylation of glucosinolate R-groups in Arabidopsis. Plant J. 72, 411–422 (2012).
Meyer, L., Crocoll, C., Halkier, B. A., Mirza, O. A. & Xu, D. Identification of key amino acid residues in AtUMAMIT29 for transport of glucosinolates. Front. Plant Sci. 14, 1219783 (2023).
Madsen, S. R., Kunert, G., Reichelt, M., Gershenzon, J. & Halkier, B. A. Feeding on leaves of the glucosinolate transporter mutant gtr1gtr2 reduces fitness of Myzus persicae. J. Chem. Ecol. 41, 975–984 (2015).
Kim, J.-Y. et al. Cellular export of sugars and amino acids: role in feeding other cells and organisms. Plant Physiol. 187, 1893–1914 (2021).
Besnard, J. et al. Arabidopsis UMAMIT24 and 25 are amino acid exporters involved in seed loading. J. Exp. Bot. 69, 5221–5232 (2018).
Jørgensen, M. E. et al. Origin and evolution of transporter substrate specificity within the NPF family. eLife 6, e19466 (2017).
Peña-Varas, C. et al. Structural insights into the substrate transport mechanisms in GTR transporters through ensemble docking. Int. J. Mol. Sci. 23, 1595 (2022).
Chung, Y.-C., Cheng, H.-Y., Wang, W.-T., Chang, Y.-J. & Lin, S.-M. Transport efficiency of AtGTR1 dependents on the hydrophobicity of transported glucosinolates. Sci. Rep. 12, 5097 (2022).
Sanders, A. et al. AAP1 regulates import of amino acids into developing Arabidopsis embryos. Plant J. 59, 540–552 (2009).
Le, B. H. et al. Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc. Natl Acad. Sci. USA 107, 8063–8070 (2010).
Belmonte, M. F. et al. Comprehensive developmental profiles of gene activity in regions and subregions of the Arabidopsis seed. Proc. Natl Acad. Sci. USA 110, E435–E444 (2013).
Gottwald, J. R., Krysan, P. J., Young, J. C., Evert, R. F. & Sussman, M. R. Genetic evidence for the in planta role of phloem-specific plasma membrane sucrose transporters. Proc. Natl Acad. Sci. USA 97, 13979–13984 (2000).
Geu-Flores, F., Nour-Eldin, H. H., Nielsen, M. T. & Halkier, B. A. USER fusion: a rapid and efficient method for simultaneous fusion and cloning of multiple PCR products. Nucleic Acids Res. 35, e55 (2007).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Kurihara, D., Mizuta, Y., Sato, Y. & Higashiyama, T. ClearSee: a rapid optical clearing reagent for whole-plant fluorescence imaging. Development 142, 4168–4179 (2015).
Crocoll, C., Halkier, B. A. & Burow, M. Analysis and quantification of glucosinolates.Curr. Protoc. Plant Biol. 1, 385–409 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2020).
Acknowledgements
We thank L. Svenningsen, V. Kjær, E. Pupi, S. J. Heilmann-Clausen and J. Skytte Thorsen for laboratory assistance; the support staff at the Centre for Advanced Bioimaging and growth facilities at the Department of Plant and Environmental Sciences at the University of Copenhagen for technical assistance. We acknowledge the Danish National Research Foundation (DNRF99) for its financial support. H.H.N.-E. also acknowledges financial support from the Novo Nordisk Foundation (NNF19OC0056580).
Author information
Authors and Affiliations
Contributions
D.X., N.C.H.S. and B.A.H. conceptualized the project. D.X. and N.C.H.S. designed the experiments. N.C.H.S. performed most of the experiments, including line generation, bioimaging and glucosinolate extractions. D.X. and N.C.H.S. analysed data. C.K. generated native GTR2 complementation lines and performed initial screening of seed glucosinolates in these lines. C.C. performed LC–MS/MS analyses. A.S. supervised bioimaging experiments. N.C.H.S., D.X., B.A.H., H.H.N.-E. and A.S. interpreted data. N.C.H.S. and D.X. wrote the original draft of the manuscript. D.X., N.C.H.S., B.A.H., H.H.N.-E. and A.S. reviewed and edited the paper. All authors commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Peter Dörmann, Sakiko Okumoto and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
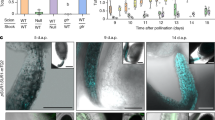
Extended Data Fig. 1 Crossing scheme for tissue-specific re-introduction of glucosinolate export activity in umamit29.
Experimental details and additional data points to support main Fig. 1c . a, Schematic representation of the crossing strategy. To ensure equal allelic load in the cross between pARR22:: UTMAMIT29-mVenus (pARR22::UT29V) and pSUR1::UTMAMIT29-mVenus (pSUR1::UT29V) and control and parent lines, the latter (Col-0, pARR22::UT29V and pSUR1::UT29V) were crossed to umamit29-1 (ut29-1) and the heterozygous progeny was used in the experiment. b, Total levels of glucosinolates in seeds harvested from F1 progeny of crosses between pSUR1::UT29V and pARR22::UT29V. “Set 2” and “Set 3” are additional replicates supporting the set shown in main Fig. 1c. Letters indicate statistical significant difference between groups (n = 9(Col-0), 12(ut29-1, pSUR1::UT29V-2, pARR22::UT29V-2 and -3 and pSUR1::UT29V-3 X pARR22::UT29V-3), 11(pSUR1::UT29V-2) and 6(pSUR1::UT29V-2 X pARR22::UT29V-2), p < 0.05, ANOVA, Games-Howell post-hoc test). The center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information. Note that: 1) in “Set 2”, we obtained only six pSUR1::UT29V X pARR22::UT29V plants, which impacts the power of the statistical analysis; and 2) in “Set 3” the pSUR1::UT29V X ut29-1 cross yielded no viable seeds due to technical issues. Instead, we analyzed seeds from uncrossed siliques of the homozygous pSUR1::UT29V parent. The data are similar between “Set 3” and the other replicates, suggesting that the pSUR1::UT29V allelic load has a negligible contribution to the observed effects.
Extended Data Fig. 2 Complementation of the seed glucosinolate phenotype of gtr1gtr2gtr3 and identification of single copy transformants.
a, Total glucosinolates in seeds from individual primary transformants harbouring different GTR2 expressed under various lengths of its native promoters in the gtr1gtr2gtr3 mutant background. Lines indicate mean levels. CDS: coding sequence. Genomic: full genomic gene fragment including introns and untranslated regions. Letters indicate significant difference between groups (p < 0.05, n = 10(Col-0 and pGTR2(2 kb)::GTR2(CDS)), 11(gtr1/2/3, gtr1/3 and pGTR2(6 kb)::GTR2(genomic)), 16(pGTR2(2 kb)::GTR2(genomic)), 8(pGTR2(4 kb)::GTR2(genomic) and 5(pGTR2(8 kb)::GTR2(genomic), ANOVA, Games-Howell post-hoc test). Dots represent individual data points. The center line indicates the mean. Details of the statistical analysis are provided as Supplementary Information. b, Segregation analysis of the three highest accumulating pGTR2(8 kb)::GTR2(genomic) lines in the T2 generation. The heterozygous lines #1 and #2 (ie. GTR2C-1 and GTR2C-2) were selected for downstream experiments. Line #3 likely contains multiple insertions. Scoring based on presence (heterozygous)/absence(homozygous) of red fluorescence in seeds. The analysis was done on plants grown from FastRed positive seeds, that is segregating mutant background seeds were removed from the population. The expected frequency is thus 2 heterozygotes:1 homozygote. Chi-square test performed in Microsoft Excel (n = 23, Expected frequency 2:1).
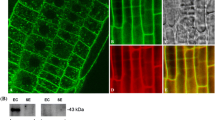
Extended Data Fig. 3 Effects of embryonic pSUC2::GTR2-mVenus and pUMAMIT29::UMAMIT29 expression on seed glucosinolate levels.
Seeds containing GTR2 expressed under the SUC2 promoter and seeds expressing UMAMIT29 under its native promoter were analyzed based on transgene presence in filial tissues (FastRed positive: maternal + filial expression, FastRed negative: maternal expression). a, pSUC2::GTR2-mVenus in gtr1gtr2gtr3 (pSUC2::GTR2V) expression in mature green stage embryo. Scale bar = 100 µm. Representative image of five embryos collected from two independent lines. b, Total glucosinolates in pSUC2::GTR2V sorted seeds (n = 10 for pSUC2::GTR2V; n = 5 for Col-0 and gtr1gtr2gtr3). c, Total glucosinolates in sorted seeds from pUMAMIT29::UMAMIT29-mVenus (UT29C) in the umamit29/umamit30/umamit31 triple mutant (ut29/30/31) (n = 6(Col-0 and ut29/30/31) and 10(UT29C Mat/Full)). Letters denote statistically significant differences between groups (p < 0.05, ANOVA, Games-Howell post-hoc test). “ns” in panel (c) indicates no significant difference between FastRed positive and FastRed negative seeds (Paired T-test (two-tailed), pairing based on parent plant, p = 0.561). In box plots, the center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information. Abbreviations: Full: FastRed positive; Mat: Maternal only, FastRed negative.
Extended Data Fig. 4 Levels of different glucosinolate species in complemented gtr1gtr2gtr3 seeds sorted based on the presence of transgene in filial tissues.
Glucosinolates in seeds from gtr1grt2gtr3 complemented with pGTR2(8 kb)::GTR2 (GTR2C) were sorted based om presence of the transgene in filial tissues (See Fig. 2, FastRed positive: maternal + filial expression, FastRed negative: maternal expression). Aggregated amounts were calculated based on glucosinolate side chain modification type. Letters indicate significant difference between seed pools within each glucosinolate type (n = 12, p < 0.05, ANOVA, Games-Howell post-hoc test). The center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information. Abbreviations: Full: FastRed positive; Mat: Maternal only, FastRed negative.
Extended Data Fig. 5 Effect of embryonic expression of GTR2 under the SUC2 promoter on seed glucosinolate profile.
Dihomo-methionine-derived glucosinolates in pSUC2::GTR2-mVenus in gtr1gtr2gtr3 (pSUC2::GTR2V) seeds sorted based on transgene presence in filial tissues (n = 10 for pSUC2::GTR2V; n = 5 for Col-0 and gtr1gtr2gtr3). Letters indicate statistically significant difference (p < 0.05, ANOVA, Games-Howell post-hoc test). The center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information. Abbreviations: Full: FastRed positive; Mat: Maternal only, FastRed negative.
Extended Data Fig. 6 4MSB levels in seed from umamit29 containing UMAMIT29 expressed by tissue-specific promoters.
4MSB levels in seeds harvested from F1 progeny of pSUR1::UMAMIT29-mVenus (pSUR1::UT29V) and pARR22::UMAMIT29-mVenus (pARR22::UT29V) crosses. Letters indicate significant difference between groups (n = 9(Col-0), 12(ut29-1, pSUR1::UT29V-2, pARR22::UT29V-2 and -3 and pSUR1::UT29V-3 X pARR22::UT29V-3), 13(pSUR1::UT29V-1), 14(pARR22::UT29V-1 pSUR1::UT29V-1 X pARR22::UT29V-1), 11(pSUR1::UT29V-2) and 6(pSUR1::UT29V-2 X pARR22::UT29V-2), p < 0.05, ANOVA, Games-Howell post-hoc test). The center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information.
Extended Data Fig. 7 Levels of different glucosinolate species in umamit29 containing UMAMIT29 expressed by tissue-specific promoters.
Glucosinolates in seeds harvested from F1 progeny of pSUR1::UT29V and pARR22::UT29V crosses. Total amounts were calculated based on glucosinolate side chain modification. Letters indicate significant difference between seed pools within each glucosinolate type (n = 9(Col-0), 12(ut29-1, pSUR1::UT29V-2, pARR22::UT29V-2 and -3 and pSUR1::UT29V-3 X pARR22::UT29V-3), 13(pSUR1::UT29V-1), 14(pARR22::UT29V-1 pSUR1::UT29V-1 X pARR22::UT29V-1), 11(pSUR1::UT29V-2) and 6(pSUR1::UT29V-2 X pARR22::UT29V-2), p < 0.05, ANOVA, Games-Howell post-hoc test). The center line indicates the median, the box limits denote the lower and upper quartiles, whiskers extend to the largest and smallest value no further than 1.5*IQR (inter-quartile range) from the hinge. Data beyond the end of the whiskers are plotted individually. Dots represent individual data points. Details of the statistical analysis are provided as Supplementary Information.
Supplementary information
Supplementary Information
List of oligonucleotides and statistical details for pairwise comparisons.
Supplementary Data
Glucosinolate profiles in all the samples in the study. The quantity of individual glucosinolate was shown in nmol per sample.
Supplementary Code
R-code (in R Markdown format) for statistical analysis and plot generation and the HTML output of this code.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sanden, N.C.H., Kanstrup, C., Crocoll, C. et al. An UMAMIT-GTR transporter cascade controls glucosinolate seed loading in Arabidopsis. Nat. Plants 10, 172–179 (2024). https://doi.org/10.1038/s41477-023-01598-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01598-4
- Springer Nature Limited