Abstract
Background
Prostate cancer (PrCa) is one of the most hereditable human cancers, however, only a small fraction of patients has been shown to carry deleterious variants in known cancer predisposition genes.
Methods
Whole-exome sequencing was performed in multiple affected members of 45 PrCa families to select the best candidate genes behind part of the PrCa missing hereditability. Recurrently mutated genes were prioritised, and further investigated by targeted next-generation sequencing in the whole early-onset and/or familial PrCa series of 462 patients.
Results
PRUNE2 stood out from our analysis when also considering the available data on its association with PrCa development. Ten germline pathogenic/likely pathogenic variants in the PRUNE2 gene were identified in 13 patients. The most frequent variant was found in three unrelated patients and identical-by-descent analysis revealed that the haplotype associated with the variant is shared by all the variant carriers, supporting the existence of a common ancestor.
Discussion
This is the first report of pathogenic/likely pathogenic germline variants in PRUNE2 in PrCa patients, namely in those with early-onset/familial disease. Importantly, PRUNE2 was the most frequently mutated gene in the whole series, with a deleterious germline variant identified in 2.8% of the patients, representing a novel prostate cancer predisposition gene.
Similar content being viewed by others
Background
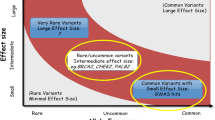
Prostate cancer (PrCa) is among the top five incident and deadliest cancers in men worldwide, with nearly 1.4 million new cases and 375,000 deaths in 2020, ranking second and fifth regarding incidence and mortality, respectively [1]. Despite being such a common disease, its aetiology remains poorly understood, with only three well-established risk factors, namely, advancing age, African ancestry, and a family history of the disease [1, 2]. Concerning family history, the relative risk of PrCa increases two- to threefold for men having a first-degree relative with PrCa and increases three to fivefold if the affected relative is diagnosed with PrCa before age 60 [3, 4]. In fact, the heritability of PrCa is one of the highest among human cancers, with inherited predisposition estimated to account for 5–15% of all cases [5,6,7], but, contrarily to other common cancers, very little is known about the causal genetic factors. The scarce knowledge, however, reveals significant genetic heterogeneity, with the contribution of both rare and common germline variants in moderate- to high-risk alleles and in low-risk alleles, respectively [2, 8].
Of the patients presenting familial aggregation and/or early-onset disease, only a few have been shown to carry rare deleterious variants in the HOXB13 gene [9,10,11], in genes predisposing to Hereditary Breast and Ovarian Cancer and Lynch Syndromes [12,13,14], namely, BRCA1, BRCA2 and the mismatch repair (MMR) genes, or in additional DNA repair genes, such as ATM, CHEK2 and PALB2 [15,16,17,18]. We have previously reported that, altogether, mutations in the BRCA2, MSH2 and HOXB13 genes account for only around 1.5% of our patients with early-onset and/or familial PrCa [11, 14]. Furthermore, using targeted next-generation sequencing (T-NGS) of 94 genes known to be involved in inherited cancer predisposition, we have identified pathogenic and “likely/potentially pathogenic” germline mutations in 14.9% of 121 PrCa cases selected for the high likelihood of hereditary disease [15]. Notwithstanding, most of our patients with early-onset or/and familial PrCa remain without a molecular diagnosis, and, thus, unveiling the missing heritability of PrCa may require the use of a more comprehensive and genome-wide approach. Populations with a high prevalence of founder effects, which is the case of the population from the north of Portugal, may simplify the identification of new cancer predisposition genes, especially when associated with homogeneous environmental exposures that reduce the non-genetic variance. We have previously identified common founder mutations in BRCA1, BRCA2 and in MMR genes [19,20,21], illustrating well how the structure of our population may help to identify rare variants and prioritise candidate genes involved in inherited cancer predisposition.
Aiming to identify new deleterious germline mutations behind a significant part of the missing PrCa hereditability, we performed whole-exome sequencing (WES) in duos and trios of PrCa patients from 45 of the 462 families previously recruited [11]. The best candidate genes were included in a customised NGS gene panel to access the prevalence of germline variants in a validation series of 417 additional PrCa families.
Methods
Patients and samples
This study enrolled 462 index cases from a previously described early-onset and/or familial PrCa series, which were recruited based on two criteria: an early-onset disease with PrCa diagnosis before the age of 56, and/or familial/hereditary PrCa with more than one case with PrCa and at least one family member diagnosed before the age of 66 [11]. The peripheral blood sample obtained from each patient recruited was labelled HPC (denoting the aim of the study to identify hereditary prostate cancer) followed by a sequential number. Regarding the histopathological diagnosis, all patients had prostate adenocarcinoma except one diagnosed with prostatic basal cell carcinoma and one with carcinosarcoma. The average age at onset was 56.6 years.
Germline DNA from all patients was extracted from peripheral blood leucocytes using the MagNA Pure LC DNA Isolation Kit —Large Volume (Roche Diagnostics GmbH, Penzberg, Germany). All DNA samples were quantified using Qubit dsDNA HS Assay (Thermofisher, Carlsbad, CA, USA), and their integrity was assessed by agarose gel electrophoresis.
Discovery series
We hypothesised that sequencing duos and trios of affected relatives would help to identify new PrCa predisposing genes, by allowing prioritisation of variants that segregate with the disease. Forty-nine of the 462 families provided DNA samples from two or three affected relatives (including the proband). Of those, 45 families, specifically, 6 with three available DNA samples and 39 with two available DNA samples, were selected for WES according to the following prioritisation: firstly, families with three available DNA samples; secondly, families with more than two family members diagnosed with PrCa; thirdly, families with probands diagnosed before the age of 61; and, lastly, families with the lowest average age at onset. The initial genetic screening with whole-exome sequencing (the “discovery series”) therefore included 96 patients from 45 families of the 462 families recruited. All the selected families had between two and six affected relatives (Supplementary Table 1), with an average number of 3.3 PrCa diagnoses per family. The degree of relationship between probands and affected relatives is shown in Supplementary Table 2. Demographic and clinicopathological characteristics are shown in Supplementary Table 3.
Validation series
To validate the involvement of the most promising candidate genes identified in the discovery series, the germline DNA from the remaining 417 index cases, regardless of the carrier status for variants in known DNA repair genes [14, 15], was sequenced using a customised NGS gene panel. The demographic and clinicopathological characteristics of these patients have already been described [11].
Sequencing
Whole-exome sequencing
Capture and sequencing
Approximately 1 μg of genomic DNA was enriched for exonic regions using the SureSelectXT2 Human All Exon v5 kit (Agilent Technologies, Santa Clara, CA, USA), according to the SureSelectXT2 Target Enrichment System for Illumina Multiplexed Sequencing protocol, at the Carvajal-Carmona Laboratory, Genome Center & Department of Biochemistry and Molecular Medicine, University of California, Davis, CA, USA. Sequencing of the pooled enriched libraries was an outsourced service, provided by the Beijing Genomics Institute sequencing facility at the UC Davis campus in Sacramento, CA, USA, and was performed with 100 bp paired-end reads using the Illumina HiSeq 2000 platform (Illumina Inc., San Diego, CA, USA).
Data processing
Raw sequence data was received in paired-end FASTQ format and processed at the Carvajal-Carmona Laboratory. Data were demultiplexed and converted to Sanger encoding using seqtk (https://github.com/lh3/seqtk). Fastq quality was assessed and checked with FastqQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Quality trimming was performed by SICKLE (https://github.com/najoshi/sickle). Sequence reads were trimmed and aligned to the Genome Analysis Toolkit (GATK) bundle human genome build GRCh37_decoy/hg19 using Burrows-Wheeler Alignment (BWA-mem, version 0.7.8-r455) [22, 23] and PCR duplicates were removed using Picard (1.118). GATK (v3.2–2) IndelRealigner and BaseRecalibrator were used for indel realignment and base quality score recalibration, and variants were called with five different callers: Freebayes [24], GATK HaplotypeCaller algorithm [25, 26], SAMtools [27], SNVer [28] and VarScan [29].
Variant annotation and prioritisation
Variants for all callers were combined and filtered according to the following filters: coverage ≥10; variant counts ≥5; variant frequency ≥10%; average single nucleotide variants (SNV) base quality ≥22; and ≥10% of variant reads on both strands. Variant annotation was carried out with ANNOVAR [30]. For secondary variant filtering, intronic variants at more than 2-bp away from exon-intron boundaries, synonymous, UTR variants, and variants present in more than 10% of the samples were excluded. In addition, an in-house perl-based script adapted from the ANNOVAR software was created to gather information for MAF (minor allele frequency) in non-Finnish European (NFE) and Iberic (IBS) populations, from the Genome Aggregation Database [gnomAD v2.1.1, June 2021] and the 1000 Genomes Project [1000 G, Phase 3 data], and for the pathogenicity predictors available at dbNSFP (v3.5). NFE data from 1000G was obtained by subtracting Finnish (FIN) data from the total of populations with European ancestry (FIN, GBR, CEU, and IBS). Frameshift, nonsense, or splicing variants with MAF ≤ 0.15% in NFE and IBS were retained in order to reduce the chance of filtering out eventual deleterious founder variants that could be relatively common in our population. Missense variants with MAF ≤ 0.15% and predicted to be damaging/deleterious by at least 9 of 11 pathogenicity predictors and supported by at least 3 of 4 conservation predictors (Supplementary Table 7), were considered “potentially pathogenic” and were retained, while variants classified as “likely benign” according to the guidelines of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG-AMP, source: InterVar [31]) were filtered out.
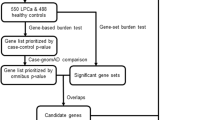
Since the goal of this study was to find new recurrently mutated genes associated with PrCa inherited predisposition, as a final candidate gene approach, we have prioritised genes with at least two variants identified (at least one frameshift, nonsense or affecting consensus splice sites) segregating with the disease in the families, which had been reported as having a role in PrCa development but not linked with DNA repair. To make sure that the prioritised variants were associated with the disease instead of being populational, all the variants found more frequently in healthy controls than in PrCa cases (described below), were discarded. PRUNE2 was the single gene fitting all these criteria.
Targeted next-generation sequencing (T-NGS)
DNA samples from the index patients of both the discovery and the validation series were sequenced using a customised gene panel designed with Agilent SureDesign, covering, among other genes, the coding and splicing regions of PRUNE2 (NM_015225.3).
For library preparation, ~50 ng of DNA were enriched for the custom primer regions using the SureSelectQXT protocol, following Agilent’s recommendations. Enriched libraries were pooled and sequencing outsourced by Health[in]Code (Spain, Coruña) in an Illumina HiSeq 2000 platform.
Data processing
The NextGENe software (v2.4.2.2; Softgenetics, State College, PA, USA) was used for sequence alignment to the reference genome (GRCh37/hg19) and variant calling. For analysis of single nucleotide variants (SNVs) and INDELs, both BAM and VCF files were imported into GeneticistAssistant (v1.8; Softgenetics) for quality control, variant annotation and primary filtering.
Variant annotation and prioritisation
Variants present in more than 10% of the samples in the whole dataset and variants with an alternative allele frequency (VAF) < 15% in any given patient were excluded to remove sequencing artefacts and low-level somatic mosaicism (which in any case are unlikely to be validated with Sanger sequencing). The performance of the SureSelect custom panel was evaluated by assessing per base coverage using an in-house python-based script, considering the full coding and splicing consensus regions. Variants in PRUNE2 were filtered as described above for WES data and were validated by Sanger sequencing.
All the variants detected by WES, as well as variants reported in previous studies [11, 14, 15] were also identified by T-NGS, further validating the performance of our custom NGS gene panel.
Genotyping by KASP technology
To understand if the most promising variants found in the PrCa patients are specifically associated with cancer development or, in opposition, are population-related, all frameshift, nonsense or splicing variants, as well as the most promising missense variants, were genotyped in 710 Portuguese control subjects, previously described [15]. For this purpose, the KASP SNP genotyping chemistry (LGC Genomics, Berlin, Germany) was used, according to the manufacturer’s instructions. Assay primers (Supplementary Table 4) were designed using the Primer-Blast [32] and the PCR reactions were run on a Roche LightCycler 480 Real-Time instrument. For data analysis, the LightCycler 480 Software 1.5.0 was used.
Haplotype analysis
To evaluate if recurrent variants identified in at least three families potentially resulted from founder effects in the population, identical-by-descent and phylogenetic analyses were performed for all carriers, from both the discovery and validation series.
The T-NGS data were phased using BEAGLE 4.1 [33], and identical-by-descent (IBD) haplotypes were determined using BEAGLE Refined IBD algorithm [34]. The lengths of the shared haplotype segments were estimated by the distance between the two last shared markers flanking the variants. Phylogenetic networks were reconstructed based on the median-joining algorithm [35] using PopART v1.7 [36].
A similar IBD and haplotype approach was applied to high-density SNP genotype data from the Portuguese early-onset and/or familial PrCa sample collection (354 PrCa cases and 180 controls) obtained with the Infinium OncoArray-500K BeadChip (Illumina), previously described [37], as part of the PRACTICAL consortium.
Results
Variants in PRUNE2
PRUNE2 was the only candidate gene fitting all the variant annotation and prioritisation criteria indicated above. Six different non-missense variants were identified in the PRUNE2 gene in eight families, two of which were in patients belonging to the discovery series, and four in patients from the validation series (Table 1). Regarding missense variants, 28 were found, all in patients from the validation series (Supplementary Table 5).
Non-missense variants
Two index patients from the discovery series were found to harbour PRUNE2 non-missense variants. One is an in-frame insertion variant, c.111_113dup, carried by patient HPC387 and his affected paternal cousin HPC516, the first having been diagnosed by the age of 50, and the latter by the age of 59 (Fig. 1a). The other is a splicing variant, c.7514-1G>A, shared by HPC164 and his brother, HPC486, who were diagnosed at the age of 51 and 60 years, respectively (Fig. 1b). They have two more affected brothers, however, further segregation analysis was not possible. None of these variants was found in our series of 710 healthy controls.
a Patient HPC387, harbouring the PRUNE2 variant c.111_113dup, which co-segregates with the disease in a paternal cousin, HPC516. b Patient HPC164, harbouring the PRUNE2 splicing variant c.7514-1G>A, segregating with the disease in his brother, HPC486. Males are represented by a square, females by a circle and unspecified gender by a diamond. A diagonal line through a symbol indicates that the individual is deceased, and coloured symbols represent affected individuals. The index case is indicated by an arrow and the cancer type and age at diagnosis are indicated when known.
We identified four additional non-missense variants in patients belonging to the validation series, specifically, one nonsense, one frameshift, one in-frame and one splicing variant. The stop-gain variant, c.421C>T, which leads to a premature stop codon at codon 141, is carried by an early-onset patient, HPC403, diagnosed by the age of 53 and with no family history of PrCa. One healthy male was also found to carry the same variant. Patient HPC493 is the carrier of the frameshift variant c.2188dup and was diagnosed at 55 years, having a deceased paternal uncle with PrCa. This variant, which results in a premature stop codon at codon 735, was not found in non-cancer controls. Apart from the in-frame variant found in the discovery series, a second in-frame deletion variant (c.7669_7671del) found in a validation series’ patient, HPC433, was also absent in controls. The carrier was diagnosed at 58 years and his father also had PrCa, although his sample was not available to test for segregation. Lastly, three non-related patients, HPC119, HPC139, and HPC314, carry the same splicing variant, the c.509-2A>G transition, which was not found in any healthy control subject. The first patient (HPC119) fulfils the two recruitment criteria (Fig. 2a), with PrCa diagnosed at the age of 55, and with a strong family history of the disease, having five affected family members, specifically, the father, two brothers, one paternal cousin and one maternal uncle. Unfortunately, the relatives are either deceased or living abroad, making it unfeasible to perform additional segregation studies. The other carriers of this variant, patients HPC139 and HPC314, were also diagnosed at an early age, at 54 and 55 years, respectively, but had no family history of PrCa (Fig. 2b, c). We have tried to amplify the cDNA of the tumour sample from patient HPC314, the single available at our institution, however, the bad quality of the RNA sample did not allow to pursue additional studies on aberrant splicing events. Moreover, as PRUNE2 is not expressed in the blood (GTEx Portal accessed in 07/27/21), RNA analysis in a germline sample would not be informative.
To further address if the variants 7514-1G>A and c.509-2A>G, which occur in very conserved splicing regions [38] at −1 and −2 upstream of exon 5 and exon 9, respectively, can interfere with the normal splicing, we queried different in silico software. In fact, both splicing variants were predicted to be disease-causing by MutationTaster [39], and to affect the acceptor splice site, and, consequently, interfere with splicing, by the in silico tools Human Splicing Finder V3.0 [40] and MaxEntScan3ss [41] (Supplementary Table 6).
Missense variants
Of the 28 missense variants in PRUNE2 found in our patients, four variants occurred at highly conserved nucleotides (Supplementary Table 7), having been screened in healthy subjects to further clarify an eventual disease association (Table 2). The c.8725G>A variant, considered potentially pathogenic by 10 of the 11 in silico predictors (Supplementary Table 7), was found in HPC342, a patient diagnosed by the age of 53 without family history of PrCa. One healthy control with no family history of cancer was identified as a carrier of the same variant. The remaining three variants were predicted to be pathogenic by 9 of the 11 in silico predictors (Supplementary Table 7). The c.247G>A transition, identified in a patient with early-onset disease (HPC240) was not found in controls, the c.389C>T variant, identified in two patients fulfilling the family history criterion only (HPC178 and HPC268), was found in one male control, and the c.442G>A variant, carried by an early-onset patient with no family history of PrCa (HPC409), was found in one healthy female.
The remaining 24 missense variants were not further explored due to the unconserved nature of the involved nucleotides (Supplementary Table 7).
Founder effect of recurrent variants
Following the discovery of the recurrent splice variant c.509-2A>G in three unrelated patients, we performed a preliminary IBD analysis on chromosome 9 using T-NGS genotype data. The haplotype reconstruction revealed a shared ~195.3 kb segment flanking the variant (chr9:7932566–79520913) among all three carriers, highly suggesting a common founder origin (Fig. 3a). An extended shared haplotype spanning the entire targeted genotyped PRUNE2 gene region (~291.5 Kb) was identified in two of the three carriers (HPC119 and HPC139).
a Length of the shared haplotype flanking the PRUNE2 variant c.509-2 A > G in the three HPC carriers. The grey shading indicates the entire PRUNE2 gene. The blue region represents the shared haplotype identified by the analysis of T-NGS data (SureSelect gene panel), whereas the salmon region (which includes also the genes GCNT1, PCSK5 and RFK) represents the haplotype identified using the high-density SNP OncoArray genotype data. b Median-joining network reconstruction of the extended IBD haplotype region (chr9:78682426–79514465) for the Portuguese PRACTICAL dataset (374 cases and 180 controls). Each circle represents a haplotype, and circle size is proportional to the number of individuals carrying the specific haplotype. The haplotype harbouring the variant is coloured in red and highlighted in detail. Median-joining algorithm was implemented using PopArt software.
To obtain a better resolution of the variant haplotype and to verify if it extended beyond the PRUNE2 gene, we performed a similar IBD analysis approach using high-density genome-wide SNP data obtained for the Portuguese patients included in the PRACTICAL consortium [37], which included the three variant carriers. The IBD analysis of the genome-wide SNP data (Oncoarray) corroborated the existence of a smaller core shared haplotype, matching the conserved region previously identified in the analysis of the T-NGS data. Furthermore, it also revealed that the larger haplotype identified in the two carriers in the initial analysis extended to ~832 Kb (chr9: 78,682,426–79,514,465) (Fig. 3a). The phylogenetic network reconstruction of this extended haplotype region was consistent with the previous analysis, and revealed a high level of genetic variability, with none of the other genotyped participants of the Portuguese PRACTICAL dataset carrying the haplotype flanking the variant (Fig. 3b), further supporting a common ancestor.
Discussion
The goal of this work was to identify new genes associated with PrCa genetic susceptibility, so we hypothesised that using a genome-wide approach such as WES in duos and trios of affected relatives, in a relatively homogeneous population, could help us to prioritise candidate genes. In line with this, we have selected 96 patients from 45 PrCa families of our series of 462 families with early-onset and/or familial/hereditary disease. Recurrently mutated genes with at least one frameshift, nonsense or splicing variant segregating with the disease in the affected family members were prioritised. PRUNE2, so far not implicated in PrCa predisposition, stood out as a good candidate gene, due to the significant frequency of variants found in our series and its association with PrCa development [42].
PRUNE2, also known as BMCC1, was first described by Machida and colleagues, who correlated its increased expression with good prognosis in neuroblastoma patients, attributing to PRUNE2 a role in cellular transformation, differentiation, and also survival and aggressiveness of tumorigenic cells [43]. PRUNE2 gained relevance in PrCa due to its association with the well-known non-coding RNA PCA3 (the most specific PrCa biomarker), which was found to be embedded within intron 6 of PRUNE2 in the opposite orientation [44]. Later, it was shown that PCA3 and PRUNE2 negatively regulate each other through a complex post-transcriptional RNA editing mechanism, mediated by adenosine deaminase acting on RNA (ADAR) proteins [42]. The same study reported that PRUNE2 downregulation increases cell proliferation and transformation. Conversely, PCA3 knockdown decreases cell proliferation. Furthermore, by using data retrieved from The Cancer Genome Atlas (TCGA), the authors have also demonstrated that in human prostate cancer samples, both the mRNA levels and protein expression of PCA3 and PRUNE2, correlated inversely. High PRUNE2 and low PCA3 levels are detected in non-malignant prostate samples, whereas low PRUNE2 and high PCA3 are found in prostate carcinomas [42]. For this reason, PRUNE2 was proposed as a PrCa-specific tumour suppressor gene (TSG).
Here, we report the presence of germline monoallelic variants in PRUNE2 in early-onset and/or familial PrCa with deleterious and potentially pathogenic variants being present in 2.8% of the patients (13/462). While a small number of somatic variants have already been reported in PrCa patients in publicly available databases (Fig. 4), to the best of our knowledge, this is the first report of PRUNE2 germline variants in PrCa patients. Interestingly, most of the variants identified in our patients occur at the 5’- or 3’-terminal regions of the coding sequence (Fig. 4), raising the question to what extent they affect PRUNE2 expression. Due to the importance of PRUNE2 in the control of cellular transformation, specifically some of its domains (Fig. 4), it is plausible to assume that these variants may disrupt its normal tumour suppressor function. On the other hand, considering the mutual regulation between PRUNE2 and PCA3 [42], it would be interesting to investigate the impact of these germline variants in PCA3 expression, and how this regulation may contribute to prostate carcinogenesis.
Variants identified in our patients (marked by an asterisk), as well as variants found in the publicly available databases (source: BioMuta [54]) are shown. Variant circle colour code stands for deleteriousness: black, truncating; red, predicted truncating; yellow, potentially truncating (splicing); and green, missense. Main functional domains are shown: the DHHA2 domain (green) is located at the N-terminal of the protein and interacts with the metastasis suppressor Nm23-H1 [42]; the BNIP2 domain (blue) is important for apoptosis, through the interaction with pro- and anti-apoptotic molecules [55], and the CRAL-TRIO domain (orange), located at the C-terminal of the protein, is involved in actin remodelling, a fundamental process for both cell growth and migration [55].
Using available data from the TCGA Dataset, we also observed that PRUNE2 expression is decreased in PrCa samples in comparison with adjacent normal prostate (Supplementary Fig. 1), supporting its role as tumour suppressor [42]. However, we also observed that its expression pattern in tumour samples vs normal adjacent tissues is transversal to other tumour types. In fact, two of the 13 patients carrying PRUNE2 variants, HPC164 (Fig. 1b) and HPC240 (Supplementary Fig. 2), had second neoplasia (colon and kidney, respectively) and several carriers have relatives showing other cancer types (namely, breast and colon), consistent with the decreased PRUNE2 expression observed in tumour samples of most of the cancers (Supplementary Fig. 1). Moreover, a WES study in patients with parathyroid carcinoma identified one patient with a PRUNE2 germline variant, along with several other patients carrying somatic variants, altogether representing 18% of the patients [45]. Other authors have also reported inactivating somatic variants in PRUNE2, namely in a subset of patients with Merkel cell carcinoma [46], and in patients with solid papillary carcinoma with the reverse polarity of the breast [47]. Altogether, these observations strongly suggest that, as many other TSG, PRUNE2 might be a pan-cancer TSG.
The identification of recurrent founder variants in clinically relevant genes is important to improve risk assessment in specific populations and may contribute to the development of more cost-effective targeted genetic screening strategies. Therefore, in addition to a genetic screening of early-onset and familial/hereditary PrCa patients to identify pathogenic variants contributing to increased PrCa risk, we sought to investigate whether pathogenic/likely pathogenic variants present in more than two families originated independently or from a single founder mutational event. The haplotype and phylogenetic analyses of the three carriers of the c.509-2A>G PRUNE2 variant, the only likely pathogenic variant present in three families in this study, revealed a shared and rare haplotype flanking the variant, which corroborates a founder origin instead of multiple independent occurrences. Similarly, several other founder variants in cancer-predisposing genes have been described in distinct populations, including in the Portuguese population [19, 37, 48,49,50].
In an era of precision and personalised oncology, understanding the molecular basis of hereditary PrCa is paramount for predictive testing and targeted screening, risk evaluation, and ultimately for treatment decision-making [8, 51,52,53]. By demonstrating that PRUNE2 is the most frequently mutated gene in our series, occurring in a mutually exclusive manner with deleterious variants in known DNA repair genes (data not shown), we strengthened the association between PRUNE2 and PrCa development, thereby proposing PRUNE2 as a novel PrCa predisposition gene.
Data availability
Data supporting the results reported in this paper can be found at: https://hive.biochemistry.gwu.edu/biomuta/proteinview/P38936 and https://portal.gdc.cancer.gov/.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer J Clin. 2021. https://doi.org/10.3322/caac.21660
Brandão A, Paulo P, Teixeira M. Hereditary predisposition to prostate cancer: from genetics to clinical implications. Int J Mol Sci. 2020;21:1–23.
Bruner D, Moore D, Parlanti A, Dorgan JPE. Relative risk of prostate cancer for men with affected relatives: systematic review and meta-analysis. Int J Cancer. 2003;107:797–803.
Brandt A, Bermejo J, Sundquist J, Hemminki K. Age at diagnosis and age at death in familial prostate cancer. Oncologist. 2009;14:1209–17.
Verhage BAJ, Baffoe-Bonnie AB, Baglietto L, Smith DS, Bailey-Wilson JE, Beaty TH, et al. Autosomal dominant inheritance of prostate cancer: a confirmatory study. Urology. 2001;57:97–101.
Verhage BAJ, Aben KKH, Witjes JA, Straatman H, Schalken JA, Kiemeney LALM. Site-specific familial aggregation of prostate cancer. Int J Cancer. 2004;109:611–7.
Vietri MT, D’elia G, Caliendo G, Resse M, Casamassimi A, Passariello L, et al. Hereditary prostate cancer: genes related, target therapy and prevention. Int J Mol Sci. 2021;22; https://doi.org/10.3390/ijms22073753.
Raghallaigh HN, Eeles R. Genetic predisposition to prostate cancer: an update. Fam Cancer. 2021. https://doi.org/10.1007/S10689-021-00227-3
Ewing CM, Ray AM, Lange EM, Zuhlke KA, Robbins CM, Tembe WD, et al. Germline mutations in HOXB13 and prostate-cancer risk. N. Engl J Med. 2012;366:141–9.
Lin X, Qu L, Chen Z, Xu C, Ye D, Shao Q, et al. A novel germline mutation in HOXB13 is associated with prostate cancer risk in Chinese men. Prostate. 2013;73:169–75.
Maia S, Cardoso M, Pinto P, Pinheiro M, Santos C, Peixoto A, et al. Identification of two novel HOXB13 germline mutations in Portuguese prostate cancer patients. PLoS ONE. 2015;10:e0132728.
Grindedal EM, Møller P, Eeles R, Stormorken AT, Bowitz-Lothe IM, Landrø SM, et al. Germ-line mutations in mismatch repair genes associated with prostate cancer. Cancer Epidemiol Biomark Prev. 2009;18:2460–7.
Castro E, Eeles R. The role of BRCA1 and BRCA2 in prostate cancer. Asian J Androl. 2012;14:409–14.
Maia S, Cardoso M, Paulo P, Pinheiro M, Pinto P, Santos C, et al. The role of germline mutations in the BRCA1/2 and mismatch repair genes in men ascertained for early-onset and/or familial prostate cancer. Fam Cancer. 2016;15:111–21.
Paulo P, Maia S, Pinto C, Pinto P, Monteiro A, Peixoto A, et al. Targeted next generation sequencing identifies functionally deleterious germline mutations in novel genes in early-onset/familial prostate cancer. PLoS Genet. 2018;14; https://doi.org/10.1371/journal.pgen.1007355.
Pritchard CC, Mateo J, Walsh MF, de Sarkar N, Abida W, Beltran H, et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N. Engl J Med. 2016;375:443–53.
Southey MC, Goldgar DE, Winqvist R, Pylkäs K, Couch F, Tischkowitz M, et al. PALB2, CHEK2 and ATM rare variants and cancer risk: data from COGS. J Med Genet. 2016;53:800.
Leongamornlert D, Saunders E, Wakerell S, Whitmore I, Dadaev T, Cieza-Borrella C, et al. Germline DNA repair gene mutations in young-onset prostate cancer cases in the UK: evidence for a more extensive genetic panel. Eur Urol. 2019;76:329–37.
Peixoto A, Santos C, Pinheiro M, Pinto P, Soares MJ, Rocha P, et al. International distribution and age estimation of the Portuguese BRCA2 c.156_157insAlu founder mutation. Breast Cancer Res Treat. 2011;127:671–9.
Peixoto A, Santos C, Pinto P, Pinheiro M, Rocha P, Pinto C, et al. The role of targeted BRCA1/BRCA2 mutation analysis in hereditary breast/ovarian cancer families of Portuguese ancestry. Clin Genet. 2015;88:41–48.
Pinheiro M, Pinto C, Peixoto A, Veiga I, Mesquita B, Henrique R, et al. The MSH2 c.388_389del mutation shows a founder effect in Portuguese Lynch syndrome families. Clin Genet. 2013;84:244–50.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. 2013; http://github.com/lh3/bwa (Accessed 28 Jul 2021).
Garrison E, Marth G. Haplotype-based variant detection from short-read sequencing. 2012; https://arxiv.org/abs/1207.3907v2 (Accessed 28 Jul 2021).
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297.
DePristo MA, Banks E, Poplin RE, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Wei Z, Wang W, Hu P, Lyon GJ, Hakonarson H. SNVer: a statistical tool for variant calling in analysis of pooled or individual next-generation sequencing data. Nucleic Acids Res. 2011;39; https://doi.org/10.1093/NAR/GKR599.
Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009;25:2283–5.
Wang K, Li M, Hakonarson H ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38; https://doi.org/10.1093/NAR/GKQ603.
Li Q, Wang K. InterVar: clinical interpretation of genetic variants by the 2015 ACMG-AMP guidelines. Am J Hum Genet. 2017;100:267–80.
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinforma. 2012;13:134.
Browning SR, Browning BL. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet. 2007;81:1084–97.
Browning BL, Browning SR. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics. 2013;194:459–71.
Bandelt HJ, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evolution. 1999;16:37–48.
Leigh JW, Bryant D. POPART: full-feature software for haplotype network construction. Methods Ecol Evolution. 2015;6:1110–6.
Brandão A, Paulo P, Maia S, Pinheiro M, Peixoto A, Cardoso M, et al. The CHEK2 variant C.349A>G is associated with prostate cancer risk and carriers share a common ancestor. Cancers. 2020;12:1–17.
Abramowicz A, Gos M. Splicing mutations in human genetic disorders: examples, detection, and confirmation. J Appl Genet. 2018;59:253–68.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361–2.
Desmet FO, Hamroun D, Lalande M, Collod-Bëroud G, Claustres M, Béroud C. Human splicing finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 2009;37; https://doi.org/10.1093/nar/gkp215.
Yeo G, Burge CB. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Comput Biol. 2004;11:377–94.
Salameh A, Lee AK, Cardó-Vila M, Nunes DN, Efstathiou E, Staquicini FI, et al. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc Natl Acad Sci USA. 2015;112:1–6.
Machida T, Fujita T, Ooo M, Ohira M, Isogai E, Mihara M, et al. Increased expression of proapoptotic BMCC1, a novel gene with the BNIP2 and Cdc42GAP homology (BCH) domain, is associated with favorable prognosis in human neuroblastomas. Oncogene. 2006;25:1931–42.
Clarke RA, Zhao Z, Guo A-Y, Roper K, Teng L, Fang Z-M, et al. New genomic structure for prostate cancer specific gene PCA3 within BMCC1: implications for prostate cancer detection and progression. PLoS ONE 2009;4:e4995.
Yu W, McPherson JR, Stevenson M, van Eijk R, Heng HL, Newey P, et al. Whole-exome sequencing studies of parathyroid carcinomas reveal novel PRUNE2 mutations, distinctive mutational spectra related to APOBEC-catalyzed DNA mutagenesis and mutational enrichment in kinases associated with cell migration and invasion. J Clin Endocrinol Metab. 2015;100:E360–E364.
Harms PW, Vats P, Verhaegen ME, Robinson DR, Wu YM, Dhanasekaran SM, et al. The distinctive mutational spectra of polyomavirus-negative Merkel cell carcinoma. Cancer Res. 2015;75:3720–7.
Alsadoun N, MacGrogan G, Truntzer C, Lacroix-Triki M, Bedgedjian I, Koeb MH, et al. Solid papillary carcinoma with reverse polarity of the breast harbors specific morphologic, immunohistochemical and molecular profile in comparison with other benign or malignant papillary lesions of the breast: a comparative study of 9 additional cases. Mod Pathol. 2018;31:1367–80.
Pinto P, Peixoto A, Santos C, Rocha P, Pinto C, Pinheiro M, et al. Analysis of founder mutations in rare tumors associated with hereditary breast/ovarian cancer reveals a novel association of BRCA2 mutations with ampulla of vater carcinomas. PLoS ONE. 2016;11:e0161438.
Laitinen VH, Wahlfors T, Saaristo L, Rantapero T, Pelttari LM, Kilpivaara O, et al. HOXB13 G84E mutation in Finland: population-based analysis of prostate, breast, and colorectal cancer risk. Cancer Epidemiol Biomark Prev. 2013;22:452–60.
Varon R, Seemanova E, Chrzanowska K, Hnateyko O, Piekutowska-Abramczuk D, Krajewska-Walasek M, et al. Clinical ascertainment of Nijmegen breakage syndrome (NBS) and prevalence of the major mutation, 657del5, in three Slav populations. Eur J Hum Genet. 2000;8:900–2.
Bancroft EK, Page EC, Castro E, Lilja H, Vickers A, Sjoberg D, et al. Targeted prostate cancer screening in BRCA1 and BRCA2 mutation carriers: results from the initial screening round of the IMPACT study. Eur Urol. 2014;66:489–99.
Castro E, Goh C, Olmos D, Saunders E, Leongamornlert D, Tymrakiewicz M, et al. Germline BRCA mutations are associated with higher risk of nodal involvement, distant metastasis, and poor survival outcomes in prostate cancer. J Clin Oncol. 2013;31:1748–57.
Mateo J, Carreira S, Sandhu S, Miranda S, Mossop H, Perez-Lopez R, et al. DNA-repair defects and olaparib in metastatic prostate cancer. N Engl J Med. 2015;373:1697–708.
Dingerdissen HM, Torcivia-Rodriguez J, Hu Y, Chang TC, Mazumder R, Kahsay R. BioMuta and BioXpress: mutation and expression knowledgebases for cancer biomarker discovery. Nucleic Acids Res. 2018;46:D1128–36.
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 2021;49:D412–9.
Funding
This work was supported by IPO-Porto Research Center (CI-IPOP-16-2012) and by Fundação para a Ciência e a Tecnologia (FCT; PTDC/DTP-PIC/1308/2014—POCI-01-0145-FEDER-016889). The following authors were funded by FCT with scholarships or research contracts: MC (SFRH/BD/116557/2016), SM (SFRH/BD/71397/2010), AB (PTDC/DTP-PIC/1308/2014—POCI-01-0145-FEDER-016889; POCI-01-FEDER-028245; UIDP/DTP/00776/2020; 2021.03835.CEECIND), and PP (PEst-OE/SAU/UI0776/2014; UID/DTP/00776/2013/POCI-01-0145-FEDER-006868; CEECINST/00091/2018). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Luis Carvajal-Carmona receives funding from The Auburn Community Cancer Endowed Chair on Basic Science and from the National Cancer Institute of the National Institutes of Health (grant P30CA093373). The opinions expressed in this article are the author’s own and do not reflect the view of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conception and design: MT and LGC-C. Data acquisition: MC, SM, PP, AB, RS and NB. Analysis and interpretation of the data: MC, SM, PP, PL and AB. Drafting of the manuscript: MC. Critical revision of the manuscript: MT, PP, LGC-C and AB. Statistical analysis: AB and PL. Funding obtention: MT and LGC-C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Institutional Ethics Committee of the Portuguese Oncology Institute-Porto (approval number: 38.010) and performed in accordance with the Declaration of Helsinki. Written consent was obtained from all participants.
Consent to publish
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cardoso, M., Maia, S., Brandão, A. et al. Exome sequencing of affected duos and trios uncovers PRUNE2 as a novel prostate cancer predisposition gene. Br J Cancer 128, 1077–1085 (2023). https://doi.org/10.1038/s41416-022-02125-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-02125-6
- Springer Nature Limited
This article is cited by
-
Using gene and gene-set association tests to identify lethal prostate cancer genes
Prostate Cancer and Prostatic Diseases (2024)
-
Assessment of genetic and clinical factors in T2D susceptibility among patients with hypertension
Acta Diabetologica (2024)