Abstract
Blood circulating cell-free DNA (ccfDNA) is a suggested biosource of valuable clinical information for cancer, meeting the need for a minimally-invasive advancement in the route of precision medicine. In this paper, we evaluated the prognostic and predictive potential of ccfDNA parameters in early and advanced breast cancer. Groups consisted of 150 and 16 breast cancer patients under adjuvant and neoadjuvant therapy respectively, 34 patients with metastatic disease and 35 healthy volunteers. Direct quantification of ccfDNA in plasma revealed elevated concentrations correlated to the incidence of death, shorter PFS, and non-response to pharmacotherapy in the metastatic but not in the other groups. The methylation status of a panel of cancer-related genes chosen based on previous expression and epigenetic data (KLK10, SOX17, WNT5A, MSH2, GATA3) was assessed by quantitative methylation-specific PCR. All but the GATA3 gene was more frequently methylated in all the patient groups than in healthy individuals (all p < 0.05). The methylation of WNT5A was statistically significantly correlated to greater tumor size and poor prognosis characteristics and in advanced stage disease with shorter OS. In the metastatic group, also SOX17 methylation was significantly correlated to the incidence of death, shorter PFS, and OS. KLK10 methylation was significantly correlated to unfavorable clinicopathological characteristics and relapse, whereas in the adjuvant group to shorter DFI. Methylation of at least 3 or 4 genes was significantly correlated to shorter OS and no pharmacotherapy response, respectively. Classification analysis by a fully automated, machine learning software produced a single-parametric linear model using ccfDNA plasma concentration values, with great discriminating power to predict response to chemotherapy (AUC 0.803, 95% CI [0.606, 1.000]) in the metastatic group. Two more multi-parametric signatures were produced for the metastatic group, predicting survival and disease outcome. Finally, a multiple logistic regression model was constructed, discriminating between patient groups and healthy individuals. Overall, ccfDNA emerged as a highly potent predictive classifier in metastatic breast cancer. Upon prospective clinical evaluation, all the signatures produced could aid accurate prognosis.
Similar content being viewed by others
Introduction
Breast cancer is by far the most frequently diagnosed malignancy in women globally, with an estimated 1.67 million new cases in 2012 [1]. Survival prediction and therapeutic strategies depend traditionally on tumor size, nodal status, hormonal receptor status, and the existence of metastatic lesions. However, breast cancer is an extensively heterogeneous disease leading to large variability in tumor evolution scenarios, often resulting in mortality due to drug resistance and metastasis. Currently, disease progression is monitored by imaging techniques and invasive tumor biopsies. Recent novel approaches implicating “liquid biopsies” such as blood circulating cell-free DNA (ccfDNA), have been considered to provide biosources of potential clinically relevant information, meeting the need for a convenient, minimally-invasive advancement in the route of precision medicine [2, 3]. However, although there are examples of FDA-approved circulating markers for some cancer types [4], the majority of them including ccfDNA are still experimental.
Low levels of ccfDNA exist in the blood of healthy individuals, whereas its amount is increased significantly in cancer [5, 6], liberated from tumor cells and carrying the mutation and methylation signatures of its malignant origin, thus dynamically mirroring its genetic and epigenetic profile [7]. Numerous studies from our and other groups attempted to validate specific cancer-related gene methylation detected in ccfDNA as a biomarker for early cancer diagnosis, accurate prognosis, and dynamic drug response monitoring [8,9,10]. The size distribution of ccfDNA fragments in cancer could also be another informative parameter, representing different releasing mechanisms and extracellular metabolic processes [11, 12]. Breast cancer patients have elevated levels of ccfDNA as compared to healthy women [13] and its methylation of cancer-related genes is similar to the primary tumor [14].
Our aim was to contribute to the knowledge of the biological characteristics of ccfDNA and extract information that could be of clinical value. We studied the ccfDNA fragment size distribution, levels, and methylation patterns of cancer-related genes in blood samples of early and advanced breast cancer patients. The panel of tumor-related genes was chosen based on previous expression and methylation data [14,15,16,17,18]. It consisted of Wnt Family Member 5 A (WNT5A), Spy-box 17 (SOX17), GATA Binding Protein 3 (GATA3), MutS protein homolog 2 (MSH2), and kallikrein 10 (KLK10). Data produced were analyzed by standard uni- and multi-variable statistics. In addition, an innovative, fully automated, machine learning pipeline for classification analysis was employed to produce classifiers and estimate their predictive performance (JADBio software, Gnosis Data Analysis) [19].
Results
Levels of ccfDNA in breast cancer patient groups and healthy volunteers
The concentration of ccfDNA was measured directly in plasma using the Qubit fluorometer. Levels of ccfDNA in the adjuvant and metastatic patient groups were significantly higher in relation to the healthy volunteer control group (p = 0.015 and <0.001, respectively) and between them, the metastatic group had greater levels than the adjuvant group (p = 0.009) (Fig. 1A, B). Receiver operating characteristic (ROC) curve analysis showed that ccfDNA levels could sufficiently discriminate healthy individuals from patients of the adjuvant group but not in the other groups, using the value of 425.5 ng/ml as a cut-off for ccfDNA concentration, AUC estimated at 0.776, sensitivity 80% and specificity 59% (p < 0.001) (95%CI 0.704–0.849) (Fig. 1C).
A. Mean ccfDNA concentration as quantified by Qubit directly in the plasma of breast cancer patients and healthy volunteers (control) and B. corresponding box plots. $ indicates statistically significant differences in relation to control, * to adjuvant, # to the metastatic group. C. ROC curve analysis shows a sensitivity of 80% and specificity of 59% for discrimination between healthy individuals and patients of the adjuvant group, using 425.5 ng/ml as a cut-off value
Levels of ccfDNA showed no correlations to the clinicopathological characteristics of the tumor and the patients’ demographic data. However, statistical analysis showed that levels were significantly higher in the metastatic patients who died (median quartile: 569.0, interquartile range (IQR): 455.0–780.0) than in those who survived (median quartile: 439.0, IQR: 405.0–530.0) (p = 0.03) (Fig. 2A), a specific finding for this group as it was not observed in the others. Kaplan–Meier revealed that metastatic patients who had ccfDNA levels over the median value of 496.5 ng/ml had significantly shorten progression-free survival (PFS) than those who had below (Fig. 2B, p = 0.036), strengthening its significance as a prognostic parameter in this patient group. We then evaluated the predictive potential of ccfDNA levels for the treatment response of metastatic patients to first-line chemotherapy. Interestingly, the median value of ccfDNA of the “non-responders” was 970.0 ng/ml (min–max: 493.0–3000.0) and was significantly higher from the median value of the “responders” (465.0 ng/ml, min–max: 316.0–934.0) (p = 0.026) (Fig. 2C). ROC curve analysis showed that ccfDNA levels had statistically significant power to discriminate patients with metastatic cancer who had progressive disease (PD) at first clinical evaluation. The area under the ROC curve was 0.845 with sensitivity 83.3%, specificity 61.0%, and 95% CI 0.877–1.000, using as a cut-off concentration for ccfDNA the concentration value 513.5 ng/ml (p = 0.009) (Fig. 2D).
Prognostic and predictive correlations of ccfDNA levels in the metastatic group of patients. A. Box-plots represent the levels of ccfDNA in relation to survival. B. Kaplan–Meier depicts PFS in relation to the median value of ccfDNA concentration. C. Box-plots show ccfDNA plasma concentrations in responders and non-responders to first-line chemotherapy. D. ROC curve analysis shows sensitivity 83% and specificity 61% in discriminating patients with metastatic cancer who had disease progression at first clinical evaluation
ccfDNA fragment profiling
Following isolation of ccfDNA from plasma, DNA fragment analysis was performed by capillary electrophoresis. In the group of healthy individuals, 43.2% of the samples contained a DNA peak of ̴160–200 bp which indicates release during apoptosis, as well as peaks of larger DNA fragments, i.e., around 2000 bp (58.6% of samples) and peaks above 10,000 bp (45.7% of samples), indicating possible active release and necrosis, respectively. Similar DNA peaks appeared in all groups of the breast cancer patients (~160–200 bp in 60.0%, ~2000 bp in 37.3%, and >10,000 in 52.6%), but we also observed additional peaks between ~200 and 500 bp in 31.8% of the patients. Furthermore, 38.5% of metastatic and 43.0% of the neo-adjuvant samples contained peaks of smaller than 160 bp size, ranging from 22 to 160 bp. Statistical analysis revealed that cancer patients who had elevated total levels of ccfDNA (over their median value of 635.5 ng/ml used as a cut-off) contained a larger number of short fragments (<160 bp) than those patients with lower total levels of ccfDNA (p = 0.011). Interestingly, the presence of the estrogen receptor (ER+) was correlated to the presence of 2000 bp fragments (p = 0.030) in patients. Tumor size and the incidence of death were also statistically correlated to a greater number of total fragments (p = 0.035 and p = 0.040, respectively). No other correlation to age, DNA methylation, disease-free interval (DFI), overall survival (OS), or other clinicopathological features and DNA fragment distribution was found. Representative results are shown in Fig. 3.
Representative capillary electropherograms showing DNA fragment size distribution in ccfDNA isolated from plasma of healthy volunteers and breast cancer patients. A. A healthy volunteer sample containing a peak at 163 bp (representing 23.2% of total ccfDNA) indicative of apoptosis and peaks at around 2000 (representing 55.8% of total ccfDNA) indicative of active release. B. A sample from the adjuvant patient group, showing a peak at 160 bp (representing 99% of total ccfDNA) and a lower at 300 bp (0.5% of total ccfDNA). C. A sample from the metastatic group with a peak at 139 bp (representing 79.6% of total ccfDNA) and additional peaks at 270–400 bp (20.3% of total ccfDNA). D. A sample from the neo-adjuvant group with extensively fragmented ccfDNA. Peaks above 10,380 indicative of necrosis were detected in nearly half of the samples (concentrations above measurable levels). Peaks at 35 and 10,380 bp in all electropherograms represent high and low ladders, respectively
Methylation status of cancer-related genes in ccfDNA of healthy volunteers and patient groups
The methylation status of SOX17, WNT5A, KLK10, MSH2, and GATA3 was addressed by qMSP in isolated ccfDNA samples (Fig. 4A). Methylation of SOX17, WNT5A, KLK10 or the simultaneous methylation of at least three genes was detected more frequently in the 3 patient groups than in the controls (for p values see Fig. 4B). MSH2 was more frequently methylated in the adjuvant and metastatic groups than in the controls. GATA3 was more frequently methylated in the neo-adjuvant group than in the control and adjuvant groups. Median RQ values (methylation levels) and corresponding box-plots are shown in Fig. 4, C. Levels of methylation of KLK10 and GATA3 were higher in the neo-adjuvant group in relation to other groups. (for KLK10, neo-adjuvant vs control or adjuvant p = 0.001, for GATA3, neo-adjuvant vs control or adjuvant or metastatic p < 0.001).
Μethylation status of breast cancer-related genes as detected by specific qMSP in ccfDNA of healthy individuals (control) and breast cancer patients. A. Percentages and numbers of samples found positive for gene methylation. B. Statistical significance of gene methylation between groups. C. Levels of gene methylation as calculated by qMSP (RQ values = 2−ΔΔCT×100). D. Box plots represent levels of methylation. $ indicates statistically significant differences in relation to control, * to adjuvant, # to metastatic. 1 = control, 2 = adjuvant, 3 = metastatic, 4 = neo-adjuvant
Analysis of ccfDNA methylation in respect to the clinicopathological characteristics of the tumor and the patients’ demographic data revealed several significant correlations. In specific, in both adjuvant and metastatic patient groups, the unmethylated status of WNT5A gene was correlated significantly to the presence of ER+, PR+, and HER2− phenotype (p = 0.040 and p = 0.016, respectively). HER2− women of the adjuvant group who were positive for the PR receptor (PR+) or had clear surgical limits or did not relapse had statistically significant more often unmethylated the KLK10 gene (p = 0.027, p = 0.021, and p = 0.004, respectively). Also, in the subgroup of triple-negative women in this patient group, the presence of KLK10 methylation was associated to recurrence (p = 0.014). When we analyzed the levels of methylation (RQ values), in the metastatic group, higher levels of WNT5A methylation were significantly correlated to larger tumor size (p = 0.022, r = 0.826). No other correlations were found in relation to age, menopause, or clinicopathological characteristics.
Survival analysis in the metastatic group of patients showed that those positive for SOX17 or WNT5A methylation or with at least 3 of any genes methylated had significantly shorter OS (p = 0.042, p = 0.043, and p = 0.048, respectively) (Fig. 5A–C). Especially in the subgroup of patients who were negative for the HER2/neu overexpression, the presence of SOX17 methylation was associated to higher risk of death (p = 0.017) and shorter OS (Fig. 5D, p = 0.011). Notably, the positive methylation status of at least 4 of any studied genes was correlated to the absence of chemotherapy response (p = 0.002). In the adjuvant group, patients positive for KLK10 methylation more often relapsed (p = 0.008) and had significantly shorter DFI (p = 0.013) as compared to others, indicating KLK10 as an adverse prognostic indicator.
Survival analysis in the metastatic group of patients of ccfDNA gene methylation. Overall survival (OS) in relation to A SOX17 methylation, B WNT5A methylation, C the methylation of at least 3 genes, D SOX17 methylation for the HER2− subgroup of patients. Data on the numbers of women still under observation in each time interval and the number of events seen are available in a supplementary file
Multivariate analysis using the JADBio tool
Our data were further analyzed by machine learning techniques in order to construct classifiers of predictive/prognostic value, combining the novel liquid biopsy-based experimental parameters emerged by our study and the established clinicopathological features of the study group. The JADBio tool employed for this analysis performs and compares automatically all standard, best-practices, and advanced machine learning techniques and produces the optimal along with the best interpretable model/signature. Data were analyzed against all relevant clinical end-points and we report here the classification tasks that resulted in predictive signatures.
For classification task the prediction of treatment response to first-line chemotherapy in the metastatic group of patients, clinical end-points were: progression disease (PD), partial response (PR) and stable disease (SD). The resulting best algorithm model was support vector machine (SVM) with AUC 0.740 and 95% confidence interval [0.622, 0.937]. Figure 6AI depicts the best interpretable model, a decision tree of 4 predictors, namely ccfDNA levels, the ER status, the number of metastatic sites and the levels of KLK10 methylation. The stability and individual feature contribution (IFC) values of each predictor of signature 1 are shown in Fig. 6AII. Analysis of the metastatic group data with patients defined as “responders” and “non-responders” resulted a uni-parametric logistic regression signature (equation) of AUC of 0.803 and 95% confidence interval [0.606, 1.000] with single feature the ccfDNA levels: signature 2: Probability (y = true) = 1/1 + exp(−M), M = −1.542 + 0.072 ccfDNAng/ml. This signature indicates that the use of ccfDNA blood concentration as a single parameter can predict response to therapy when used in this linear model.
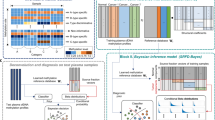
Best interpretable models emerging by JADBio data analysis A. For the prediction of response to first-line chemotherapy in the metastatic group. I. Decision Tree; 0 = progression disease; 1 = partial response; 2 = stable disease, II. Reference signature predictors and their stability and individual feature contribution (IFC). B. For overall survival (OS) in the metastatic group of patients. I. Reference signature, II. Kaplan–Meier curve predicting 3 levels of mortality according to survfit code which computes surviving for a Cox proportional hazard model. C. For discrimination between study groups. I. Reference signature, II. The 2 linear models of the multiple logistic regression model, III. Supervised Principal Component Analysis depicting discrimination between study groups
Survival analysis of the metastatic group data produced a Cox Regression model of concordance index 0.737 and 95% CI [0.593, 0.852]. The signature 3 consisted of 4 predictors, WNT5A methylation levels, response to treatment at first check, SOX17 methylation, and ccfDNA levels (Fig. 6BI). The Kaplan–Meier estimated OS categorized in 3 levels of mortality (see legend) (Fig. 6BII).
Finally, a Bagged Tree (Random Forest) has emerged as the best algorithm model for discriminating between healthy and patient groups, with AUC 0.844 and 95% CI [0.764, 0.908]. The best interpretable model was multiple logistic regression of 6 predictors (signature 4, Fig. 6CI), all ccfDNA based, namely SOX17, MSH2 and KLK10 methylation, KLK10 and WNT5A levels of methylation and ccfDNA levels. Fig. 6CII presents the 2 equations discriminating between metastatic patients over healthy individuals or over adjuvant respectively. The supervised principal component analysis figuring a bi-dimensional graphical representation of the distribution of the samples in the space, as defined by the constructed model, is shown in Fig. 6CIII.
Discussion
Our study’s ambition is to enrich the knowledge about ccfDNA in breast cancer and to illuminate relevant information that could be of clinical value, with ultimate goal the production of predictive/prognostic classifiers. To the best of our knowledge, this is the first study which evaluated ccfDNA-based experimental parameters in a multiparametric approach in breast cancer. By directly measuring ccfDNA in plasma samples, our data showed that ccfDNA levels were higher in the adjuvant and metastatic group of patients in relation to healthy individuals, patients with metastatic disease showing the highest concentrations, in accordance to previous studies [13, 20, 21]. We could not observe any correlations of ccfDNA levels to tumor size and nodal involvement like others [22, 23], possibly due to different quantification methods and patient classification criteria. Elevated levels of ccfDNA were however statistically correlated to the incidence of death and shorter PFS in the metastatic group indicating a strong prognostic potential in this patient category, in concordance to previous studies [24, 25]. When evaluating the predictive value in breast cancer, Dawson et al. showed that high ccfDNA levels were correlated negatively to treatment response in metastatic disease, and that it was the earliest between circulating biomarkers [26]. In accordance, we also demonstrated that metastatic patients who achieved PD at the first clinical check had twice as much ccfDNA than patients who demonstrated SD or PR, considered as responders. Taking our analysis one step forward, using a machine learning approach by the JADBio software, a single-parametric linear model arose with great discriminating power and ccfDNA emerged as a highly potent predictive classifier.
There are several suggested cellular processes responsible for ccfDNA release and the size of fragment content is indicative for each one of them. Apoptosis results in samples enriched by fragments size ~160 bp and multiples, necrosis delivers fragments larger than 10,000 bp and active release from viable cells gives 2000 bp size fragments [27,28,29]. Size profiling by capillary electrophoresis showed fragments of all three types, i.e., ~160 bp, 2000 bp, and above 10,000 bp. Patients with increased tumor burden in the metastatic and neo-adjuvant groups often were abundant in shorter fragments and a more fragmented pattern of distribution in relation to adjuvant and control groups, in accordance to others [30]. Previous studies claimed that short fragments (<166 bp) of ccfDNA represent the tumor-originated DNA [12, 31]. This is supported by our data, showing that elevated levels of ccfDNA were correlated to more short fragments, as shown previously in hepatocellular carcinoma [11]. In contrast, others claim that the integrity of ccfDNA is greater in breast cancer than in healthy individuals [32].
A panel of five cancer-related genes was chosen for methylation analysis of ccfDNA. In specific, this is the first study investigating the methylation status of WNT5A, KLK10, MSH2, and GATA3 in plasma ccfDNA of breast cancer patients in addition to SOX17. All but the GATA3 gene were found more frequently methylated in all the patient groups than in healthy individuals. The unmethylated status of the tumor suppressor gene WNT5A [15], was associated to ER+PR+HER2− phenotype, i.e., with a less aggressive cancer longer OS in the metastatic group. Overall, consistent to expression findings [15, 33, 34], our data indicate for the first time that the methylation of WNT5A as detected in ccfDNA is a poor prognostic factor in advanced stage breast cancer. Similarly, we demonstrated here for the first time an association of SOX17 methylation to the incidence of death, shorter PFS and OS in metastatic patients, consolidating a poor prognostic value indicated before [14, 35, 36]. KLK10 hypermethylation and downregulation has been detected in breast cancer tumor tissues and correlated to shorter DFI and OS [16, 37]. We demonstrated here that this negative prognostic value can also be detected in liquid biopsy material, i.e., in ccfDNA. When we analyzed the methylation score of any of the cancer-related genes of our panel, we demonstrated that all patient groups had score over 3 more frequently than healthy volunteers. The metastatic patients showed this high score more often and it was significantly correlated to shorter OS. Notably, methylation score over 4 was predictive for the absence of pharmacotherapy response.
For the purpose of our analysis, we introduced the JADBio tool for multivariate predictive or diagnostic analysis, using an automated machine learning pipeline, employing both standard, best-practices, and advanced machine learning techniques. All these approaches are incorporated and tested throughout its pipeline and the outcome (resulting model/signature) is based on the best selected one. The huge advantage of the use of this tool is that in an automated way, it performs and compares all potential ways of analysis, that it would be practically impossible to be done by standard analysis. In addition, it produces an estimate of its performance on new patient groups, a great advantage in mature levels of biomarker development, therefore thought to achieve the best possible exploitation for the construction of classifiers with high-performance metrics to be forwarded to clinical validation. In fact, this is the first time that this powerful methodology is used for this type of datasets, although it has already produced signatures for other clinical datasets to predict for example development of lung cancer between smokers [38] or suicide amongst depressive patients (in press) and for the prediction of secreted proteins from their mature domain features [39]. Analysis by JADBio has confirmed our standard statistics results and further strengthened the prognostic/predictive capacity of our ccfDNA-related experimental parameters. Three more signatures have emerged, utilizing multiple features: (A) a predictive decision tree of favorable performance metrics for early discrimination of metastatic patients who would achieve PD, PR, or SD, (B) a potent prognostic signature for survival in metastatic patients, (C) a classification signature of 6 features, all related to ccfDNA and its methylation with sufficient discriminating capacity between control, adjuvant, and metastatic patient groups. Upon prospective clinical evaluation, this signature could aid early and accurate diagnosis. Few previous studies have addressed the building of effective classifiers based on gene methylation patterns in breast cancer [40, 41]. Our study was the first to use machine learning approaches, combining liquid biopsy experimental data and clinicopathological parameters for producing predictive/prognostic signatures.
In conclusion, our data support the value of ccfDNA, in terms of plasma concentration and methylation patterns, as a liquid-biopsy biomaterial carrying important clinical information for breast cancer prognosis and monitoring. Overall, in our study ccfDNA emerged as a highly potent predictive classifier in metastatic breast cancer. Upon prospective clinical evaluation, all the signatures produced, based on ccfDNA innovative parameters in combination with established clinicopathological features, could aid early and accurate diagnosis and prognosis, meeting the need for a minimally-invasive advancement in the route of precision medicine.
Methods
Study groups and clinical samples
Breast cancer patients who visited the Department of Medical Oncology of PGNA between 2009 and 2017 were included in the study. Blood samples were collected following diagnosis from three patient groups: (a) 150 patients having recently (within the previous month) undergone surgery for primary breast cancer, exactly before the initiation of adjuvant therapy (adjuvant group), (b) 16 patients upon diagnosis for breast cancer, having no previous surgery, before the initiation of neo-adjuvant therapy (neo-adjuvant group), (c) 34 patients upon diagnosis for metastatic disease before the initiation of first-line chemotherapy (a combination of Taxane/Anthracyclines) (metastatic group). The clinicopathological features for all patient groups are presented in Table 1. Follow-up data until November 2017 were also available. The median follow-up period for the adjuvant group of patients was 60 months (min–max: 2–98 months): in this period 26 (17.93%) patients have died as a consequence of their disease progression, having a median follow up period of 44 months (min–max: 2–96). The median follow up time for the metastatic breast cancer group was 43 months (min–max: 1–78) at the time of which 21 (61.76%) patients died, having a median follow up period of 24 months (min–max: 1–77). The median follow up period for patients who started neo-adjuvant therapy was 61 months (min–max: 23–86), 6 (37.5%) of them deceasing during that period, with a median follow up period of 39 (min–max: 23–53). Peripheral blood was collected in EDTA before treatment and processed immediately for plasma isolation. In parallel, blood samples from 35 healthy donors were included in our study [mean age: 47.3(±SD) (±6.8), median: 48.0 (range: 27.0–59.0)] (control group). All blood samples were centrifuged immediately twice at 3000×g and then at 14,000×g for 10 min and plasma was stored at −80 °C until further use.
ccfDNA quantification
ccfDNA was quantified directly in unpurified plasma using a Qubit fluorometer 3.0 (Invitrogen Ltd., Life Technologies, UK) and a Qubit dsDNA HS Assay kit (Invitrogen Ltd., Thermo Fisher Scientific, UK) according to manufacturer’s instructions. The detection range of the kit was 10 pg/µl to 100 ng/µl.
DNA extraction and qualitative assessment of ccfDNA
ccfDNA from plasma were extracted using the QIAamp DNA Blood Mini kit (Qiagen, Germany). Specifically, DNA was eluted from 500 μl of plasma in 25 μl elution buffer and then stored in −20 °C until further use. Quality of the extracted DNA was assessed by quantitative PCR for the GAPDH gene using the KAPA SYBR Fast Master Mix (KapaBiosystems, EU). Primer sequences, annealing temperatures and related references are shown in Table 2. Samples with a quantification cycle (Ct) > 35 were excluded from further analysis. The efficiency (expressed as E = 10−1/slope−1) of assays was evaluated by using serial dilutions of placental DNA (Sigma Co., USA) in H2O (100–0.01 ng). Results were calculated using the MxPro QPCR software.
Fragment size profiling of ccfDNA
The fragment distribution of the extracted ccfDNA was analyzed by capillary electrophoresis using the High Sensitivity DNA kit and an Agilent 2100 Bioanalyzer (Agilent Technologies Inc., Santa Clara, CA) equipped with Expert 2100 software. The assay was performed according to manufacturer’s instructions using 1 μl of ccfDNA sample.
Sodium bisulfite conversion
Bisulfite conversion was performed by EZ DNA Methylation-Gold™ Kit (ZYMO Research Co., Orange, CA) as described by the manufacturer. During conversion, all unmethylated but not the methylated-cytosines of ccfDNA were converted to uracil. DNA was then eluted in 10 μl elution buffer and stored at −80 °C until use. In each experiment, CpGenome Human methylated and non-methylated DNA standards (Merck Millipore, Germany) or H2O were included as positive and negative controls respectively.
Quantitative methylation analysis (qMSP)
Promoter methylation of WNT5A, GATA3, MSH2, SOX17, and KLK10 exon 3 methylation were analyzed by qMSP [9, 42,43,44,45,46,47]. A methylation-independent assay with non-CpG including primers for the β-actin gene (ACTB) was used in order to verify DNA quality and to normalize results. Specificity and cross-reactivity of methylated and unmethylated primers (Table 2, TIB MOLBIOL, Germany) were evaluated by using unconverted gDNA, SB-converted methylated and non-methylated DNA standards. Analytical sensitivity of qMSP assays was evaluated by using serial dilutions of SB-converted methylated and non-methylated DNA standards and was found to be 0.1%. The assay efficiency (expressed as E = 10−1/slope−1) was evaluated by using serial dilutions of the SB-converted methylated DNA standards in H2O (100–0.01 ng) and was in the range of 91–105%. The analysis was performed according to the RQ sample (Relative Quantification) = 2−ΔΔCT method [48]. Specifically, ΔΔCT values were generated for each target after normalization by ACTB values and using 1% methylation as calibrator and then were multiplied by 100 (RQ = 2−ΔΔCT×100). Amplification signal >40 cycle was considered negative.
Standard statistical analysis
The Kolmogorov–Smirnov test was applied to check for normality in distribution. In cases of lack of normality, appropriate non-parametric statistics were used, like Mann–Whitney and Kruskal–Wallis tests. The median value for age and ccfDNA concentration was used as a cut off in order to divide into subgroups for further statistical analysis of binary discrete outcome. For comparison between discrete variables, like the methylation status and clinicopathological features, the chi-square and the Fischer’s exact tests were used. An ANOVA test was used for comparisons of continuous variables among three or more different subgroups. Survival curves were calculated using the Kaplan–Meier and comparisons were performed using the log-rank test. We used OS, PFS, and DFI as end points in patient survival. Metastatic patients who showed PR to treatment or SD at the first clinical check after first-line treatment initiation were considered as “responders”, whereas those who showed clinical PD were considered as “non-responders” according to Response Evaluation Criteria in Solid Tumors (RESIST) criteria version 1.1 [49]. The predictive power of the ccfDNA levels and methylation status was tested using ROC curve analysis. Statistical significance was set at p-value < 0.05. Statistical analysis was performed using the IBM SPSS 19.0 statistical software (IBM Corp. 2010. IBM SPSS Statistics for Windows, Version 19.0. Armonk, NY, USA).
Multivariate analysis by JADBio
Automated predictive modeling was performed by the Just Add Data v0.6 tool (JADBio; Gnosis Data Analysis; www.gnosisda.gr). JADBio employs both standard, best-practices, and advanced machine learning techniques for analysis. JADBio works as follows: it first selects the appropriate algorithms to try for the task at hand, depending on the outcome type, predictor type, user preferences (e.g., importance of quality of analysis vs speed of analysis) using an artificial intelligence decision support system. The algorithms are selected to perform the following steps: data transformations, data preprocessing, imputation of missing values, feature selection, predictive modeling, and data visualization. The AI system also selects which tuning hyper-parameter values to try for each algorithm. All combinations of algorithms for each step and hyper-parameter values (called configurations) are applied using a 10-fold cross-validation protocol (or similar out-of-sample estimation protocol for large sample sizes) to produce thousands of predictive models and their corresponding estimates of performance. The performance estimate of the best model is known to be over-optimistic; this is a phenomenon conceptually equivalent to multiple hypothesis testing in statistics. JADBio applies a bootstrap-based adjustment to the final reported performance [19] to remove this optimism and to return slightly conservative estimates of performance. The same bootstrap-based algorithm is employed to produce the (adjusted) confidence intervals of performance.
JADBio performs biosignature discovery (feature selection) using the statistically equivalent signature or SES algorithm for feature selection that can address both classification and survival analysis outcomes. A biosignature is defined as a minimal-size subset of predictors (features, molecular quantities, biomarkers, risk factors), which collectively (multivariately) lead to an optimal predictive model, neglecting all other features as irrelevant or redundant for prediction given the selected features. It is possible that multiple equivalent signatures are present in an analysis problem. JADBio’s algorithm tries to find as many equivalent signatures as possible. For each feature in a signature, a stability metric is produced, interpreted as the probability that the feature would have been selected again had the same study was to be repeated with new subjects. High stability features indicate features that are robustly selected. JADBio also reports the added-predictive-value of each selected feature in a signature, denoted as IFC, defined as the predictive performance achieved when that feature is removed from the model, relative to the optimal.
For classification modeling, JADBio tries SVM [50] with full polynomial and Gaussian kernels, random forests [51], ridge logistic regression [52], and decision trees [53]. For censored time-to-event analysis, a.k.a. survival analysis, JADBio employs random survival trees and Ridge Cox regression models.
JADBio reports several metrics of predictive performance and their confidence intervals using the algorithm described in Tsamardinos et al. [19]. For classification, it reports, among others, the area under the receiver operating characteristic (ROC) curve (AUC) and the accuracy (percentage of correct predictions); for survival analysis outcomes it reports the concordance index (C-index); a C-index of 90% means that between a pair of randomly selected subjects, the model assigns higher risk to the individual that experienced the event first, 90% of the times.
As most modern machine learning models are completely incomprehensible to a human, JADBio reports not only the best-out-of-all model, but also the best-interpretable model (linear models or decision trees). The interpretable model may possibly be sacrificing some predictive performance to gain interpretability. In this manuscript, we report the predictive performance of the best models and depict the best interpretable. For survival analysis, post-analysis, JADBio automatically stratifies predictions to risk strata (e.g., low, medium, high) and produces the estimated Kaplan–Meier curves of the predictions. Well separated curves visually depict the success of the model in predicting survival.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2018;68:394–424.
Pantel K, Alix-Panabieres C. Real-time liquid biopsy in cancer patients: fact or fiction? Cancer Res. 2013;73:6384–8.
Alix-Panabieres C, Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016;6:479–91.
Lamb YN, Dhillon S. Epi proColon® 2.0 CE: a blood-based screening test for colorectal cancer. Mol Diagn Ther. 2017;21:225–32.
Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011;11:426–37.
Elshimali YI, Khaddour H, Sarkissyan M, Wu Y, Vadgama JV. The clinical utilization of circulating cell free DNA (CCFDNA) in blood of cancer patients. Int J Mol Sci. 2013;14:18925–58.
Lu JL, Liang ZY. Circulating free DNA in the era of precision oncology: pre- and post-analytical concerns. Chronic Dis Transl Med. 2016;2:223–30.
Matthaios D, Balgkouranidou I, Karayiannakis A, Bolanaki H, Xenidis N, Amarantidis K, et al. Methylation status of the APC and RASSF1A promoter in cell-free circulating DNA and its prognostic role in patients with colorectal cancer. Oncol Lett. 2016;12:748–56.
Balgkouranidou I, Chimonidou M, Milaki G, Tsaroucha E, Kakolyris S, Georgoulias V, et al. SOX17 promoter methylation in plasma circulating tumor DNA of patients with non-small cell lung cancer. Clin Chem Lab Med. 2016;54:1385–93.
Mastoraki S, Strati A, Tzanikou E, Chimonidou M, Politaki E, Voutsina A, et al. ESR1 methylation: a liquid biopsy-based epigenetic assay for the follow-up of patients with metastatic breast cancer receiving endocrine treatment. Clin Cancer Res. 2018;24:1500–10.
Jiang P, Chan CW, Chan KC, Cheng SH, Wong J, Wong VW, et al. Lengthening and shortening of plasma DNA in hepatocellular carcinoma patients. Proc Natl Acad Sci USA. 2015;112:E1317–25.
Mouliere F, Rosenfeld N. Circulating tumor-derived DNA is shorter than somatic DNA in plasma. Proc Natl Acad Sci USA. 2015;112:3178–9.
Leon SA, Shapiro B, Sklaroff DM, Yaros MJ. Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res. 1977;37:646–50.
Chimonidou M, Strati A, Malamos N, Georgoulias V, Lianidou ES. SOX17 promoter methylation in circulating tumor cells and matched cell-free DNA isolated from plasma of patients with breast cancer. Clin Chem. 2013;59:270–9.
Jonsson M, Dejmek J, Bendahl PO, Andersson T. Loss of Wnt-5a protein is associated with early relapse in invasive ductal breast carcinomas. Cancer Res. 2002;62:409–16.
Yousef GM, Yacoub GM, Polymeris ME, Popalis C, Soosaipillai A, Diamandis EP. Kallikrein gene downregulation in breast cancer. Br J Cancer. 2004;90:167–72.
Kappil MA, Liao Y, Terry MB, Santella RM. DNA repair gene expression levels as indicators of breast cancer in the Breast Cancer Family Registry. Anticancer Res. 2016;36:4039–44.
McCleskey BC, Penedo TL, Zhang K, Hameed O, Siegal GP, Wei S. GATA3 expression in advanced breast cancer: prognostic value and organ-specific relapse. Am J Clin Pathol. 2015;144:756–63.
Tsamardinos I, Greasidou E, Borboudakis G. Bootstrapping the out-of-sample predictions for efficient and accurate cross-validation. Mach Learn. 2018;107:1895–922.
Catarino R, Ferreira MM, Rodrigues H, Coelho A, Nogal A, Sousa A, et al. Quantification of free circulating tumor DNA as a diagnostic marker for breast cancer. DNA Cell Biol. 2008;27:415–21.
Salvi S, Gurioli G, De Giorgi U, Conteduca V, Tedaldi G, Calistri D, et al. Cell-free DNA as a diagnostic marker for cancer: current insights. Onco Targets Ther. 2016;9:6549–59.
Tangvarasittichai O, Jaiwang W, Tangvarasittichai S. The plasma DNA concentration as a potential breast cancer screening marker. Indian J Clin Biochem. 2015;30:55–8.
Agassi R, Czeiger D, Shaked G, Avriel A, Sheynin J, Lavrenkov K, et al. Measurement of circulating cell-free DNA levels by a simple fluorescent test in patients with breast cancer. Am J Clin Pathol. 2015;143:18–24.
Olsson E, Winter C, George A, Chen Y, Howlin J, Tang MH, et al. Serial monitoring of circulating tumor DNA in patients with primary breast cancer for detection of occult metastatic disease. EMBO Mol Med. 2015;7:1034–47.
Tan G, Chu C, Gui X, Li J, Chen Q. The prognostic value of circulating cell-free DNA in breast cancer: a meta-analysis. Medicine. 2018;97:e0197.
Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013;368:1199–209.
Anker P, Stroun M, Maurice PA. Spontaneous release of DNA by human blood lymphocytes as shown in an in vitro system. Cancer Res. 1975;35:2375–82.
Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313:139–42.
Laktionov PP, Tamkovich SN, Rykova EY, Bryzgunova OE, Starikov AV, Kuznetsova NP, et al. Extracellular circulating nucleic acids in human plasma in health and disease. Nucleosides Nucleotides Nucleic Acids. 2004;23:879–83.
Madhavan D, Wallwiener M, Bents K, Zucknick M, Nees J, Schott S, et al. Plasma DNA integrity as a biomarker for primary and metastatic breast cancer and potential marker for early diagnosis. Breast Cancer Res Treat. 2014;146:163–74.
Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN. Fragment length of circulating tumor. DNA. 2016;12:e1006162.
Umetani N, Giuliano AE, Hiramatsu SH, Amersi F, Nakagawa T, Martino S, et al. Prediction of breast tumor progression by integrity of free circulating DNA in serum. J Clin Oncol. 2006;24:4270–6.
Leris AC, Roberts TR, Jiang WG, Newbold RF, Mokbel K. WNT5A expression in human breast cancer. Anticancer Res. 2005;25:731–4.
Trifa F, Karray-Chouayekh S, Jmal E, Jmaa ZB, Khabir A, Sellami-Boudawara T, et al. Loss of WIF-1 and Wnt5a expression is related to aggressiveness of sporadic breast cancer in Tunisian patients. Tumour Biol. 2013;34:1625–33.
Fu DY, Wang ZM, Li C, Wang BL, Shen ZZ, Huang W, et al. Sox17, the canonical Wnt antagonist, is epigenetically inactivated by promoter methylation in human breast cancer. Breast Cancer Res Treat. 2010;119:601–12.
Fu D, Ren C, Tan H, Wei J, Zhu Y, He C, et al. Sox17 promoter methylation in plasma DNA is associated with poor survival and can be used as a prognostic factor in breast cancer. Medicine. 2015;94:e637.
Kioulafa M, Kaklamanis L, Stathopoulos E, Mavroudis D, Georgoulias V, Lianidou ES. Kallikrein 10 (KLK10) methylation as a novel prognostic biomarker in early breast cancer. Ann Oncol. 2009;20:1020–5.
Markaki M, Tsamardinos I, Langhammer A, Lagani V, Hveem K, Roe OD. A validated clinical risk prediction model for lung cancer in smokers of all ages and exposure types: a HUNT study. EBioMedicine. 2018;31:36–46.
Orfanoudaki G, Markaki M, Chatzi K, Tsamardinos I, Economou A. MatureP: prediction of secreted proteins with exclusive information from their mature regions. Sci Rep. 2017;7:3263.
Li Y, Melnikov AA, Levenson V, Guerra E, Simeone P, Alberti S, et al. A seven-gene CpG-island methylation panel predicts breast cancer progression. BMC Cancer. 2015;15:417.
List M, Hauschild AC, Tan Q, Kruse TA, Mollenhauer J, Baumbach J, et al. Classification of breast cancer subtypes by combining gene expression and DNA methylation data. J Integr Bioinformatics. 2014;11:236.
van der Meide WF, Snellenberg S, Meijer CJ, Baalbergen A, Helmerhorst TJ, van der Sluis WB, et al. Promoter methylation analysis of WNT/beta-catenin signaling pathway regulators to detect adenocarcinoma or its precursor lesion of the cervix. Gynecol Oncol. 2011;123:116–22.
Li B, Goyal J, Dhar S, Dimri G, Evron E, Sukumar S, et al. CpG methylation as a basis for breast tumor-specific loss of NES1/kallikrein 10 expression. Cancer Res. 2001;61:8014–21.
Moura Lima E, Ferreira Leal M, Cardoso Smith Mde A, Rodriguez Burbano R, Pimentel de Assumpcao P, Bello MJ, et al. DNA mismatch repair gene methylation in gastric cancer in individuals from northern Brazil. Biocell. 2008;32:237–43.
Cooper SJ, Zou H, Legrand SN, Marlow LA, von Roemeling CA, Radisky DC, et al. Loss of type III transforming growth factor-beta receptor expression is due to methylation silencing of the transcription factor GATA3 in renal cell carcinoma. Oncogene. 2010;29:2905–15.
Hattermann K, Mehdorn HM, Mentlein R, Schultka S, Held-Feindt J. A methylation-specific and SYBR-green-based quantitative polymerase chain reaction technique for O6-methylguanine DNA methyltransferase promoter methylation analysis. Anal Biochem. 2008;377:62–71.
Miao F, Chen Z, Genuth S, Paterson A, Zhang L, Wu X, et al. Evaluating the role of epigenetic histone modifications in the metabolic memory of type 1 diabetes. Diabetes. 2014;63:1748–62.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45:228–47.
Boser BGI, Vapnik V. A training algorithm for optimal margin classifiers. ACM Digital Library. 1992:144–52.
Hoerl AE, Kennard RW. Ridge regression: biased estimation for nonorthogonal problems. Technometrics. 1970;12:55–67.
Breiman L. Random forests. Mach Learn. 2001;45:5–32.
Breiman LFH, Olshen RA, Stone CJ. Classification and regression trees. Wadsworth International Group; Belmont, California, 1984.
Acknowledgements
We would like to thank Dr. Vassilis Vassilakakis and Dr. Maria Markaki for preliminary data processing.
Funding
Ms Maria Panagopoulou received a scholarship for the implementation of her PhD Thesis, co-funded through the Act: “PROGRAM FOR THE GRANTING OF SCHOLARSHIPS FOR POSTGRADUATE STUDIES OF SECOND CYCLE STUDIES”. State Scholarship Foundation in Greece (IKY) (Operational Program “Human Resources Development—Education and Lifelong Learning”, Partnership Agreement PA 2014-2020).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The study was approved by the Scientific Board of the University General Hospital of Evros (PGNE), following assessment by Ethics Committee (decision 14/895/28.11.11), and was conducted according to the ethical principles of the 1964 Declaration of Helsinki and its later amendments.
Informed consent
All patients participated after signing a voluntary informed consent.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Panagopoulou, M., Karaglani, M., Balgkouranidou, I. et al. Circulating cell-free DNA in breast cancer: size profiling, levels, and methylation patterns lead to prognostic and predictive classifiers. Oncogene 38, 3387–3401 (2019). https://doi.org/10.1038/s41388-018-0660-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0660-y
- Springer Nature Limited
This article is cited by
-
Benchmarking UMI-aware and standard variant callers for low frequency ctDNA variant detection
BMC Genomics (2024)
-
Extensive review on breast cancer its etiology, progression, prognostic markers, and treatment
Medical Oncology (2023)
-
The double agents in liquid biopsy: promoter and informant biomarkers of early metastases in breast cancer
Molecular Cancer (2022)
-
Just Add Data: automated predictive modeling for knowledge discovery and feature selection
npj Precision Oncology (2022)
-
Functional regulations between genetic alteration-driven genes and drug target genes acting as prognostic biomarkers in breast cancer
Scientific Reports (2022)