Abstract
Directional control of tip-growing cells is essential for proper tissue organization and cell-to-cell communication in animals and plants1,2. In the sexual reproduction of flowering plants, the tip growth of the male gametophyte, the pollen tube, is precisely guided by female cues to achieve fertilization3. Several female-secreted peptides have recently been identified as species-specific attractants that directly control the direction of pollen tube growth4,5,6. However, the method by which pollen tubes precisely and promptly respond to the guidance signal from their own species is unknown. Here we show that tip-localized pollen-specific receptor-like kinase 6 (PRK6) with an extracellular leucine-rich repeat domain is an essential receptor for sensing of the LURE1 attractant peptide in Arabidopsis thaliana under semi-in-vivo conditions, and is important for ovule targeting in the pistil. PRK6 interacted with pollen-expressed ROPGEFs (Rho of plant guanine nucleotide-exchange factors), which are important for pollen tube growth through activation of the signalling switch Rho GTPase ROP1 (refs 7, 8). PRK6 conferred responsiveness to AtLURE1 in pollen tubes of the related species Capsella rubella. Furthermore, our genetic and physiological data suggest that PRK6 signalling through ROPGEFs and sensing of AtLURE1 are achieved in cooperation with the other PRK family receptors, PRK1, PRK3 and PRK8. Notably, the tip-focused PRK6 accumulated asymmetrically towards an external AtLURE1 source before reorientation of pollen tube tip growth. These results demonstrate that PRK6 acts as a key membrane receptor for external AtLURE1 attractants, and recruits the core tip-growth machinery, including ROP signalling proteins. This work provides insights into the orchestration of efficient pollen tube growth and species-specific pollen tube attraction by multiple receptors during male–female communication.
Similar content being viewed by others
Main
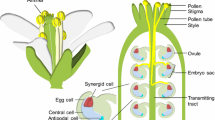
In the final step of pollen tube guidance, two synergid cells on the side of the egg cell are essential for the attraction of the pollen tube to the ovule9. We previously identified diffusible and species-specific attractants, defensin-like cysteine-rich LURE peptides, secreted from the synergid cell in the dicot plants Torenia fournieri and A. thaliana4,6. The attractants of A. thaliana, the AtLURE1 peptides, showed considerable attraction activity, but their knockdown partially impaired the precision of the pollen tube guidance around the ovule6. Moreover, various additional genes encoding secreted peptides, including many cysteine-rich peptides (CRPs), are likely to be expressed in the female gametophyte10, suggesting the existence of multiple ligand–receptor pairs for guidance. By focusing on receptor-like kinases (RLKs), which form a large gene family11 and consist of subfamilies with severalphylogenetically related genes, we screened T-DNA insertion lines for 23 genes, which encompass almost all pollen-specific RLK genes (see Methods), by a pollen tube attraction assay using the AtLURE1.2 peptide (a representative A. thaliana LURE1 peptide)6. Under semi-in-vivo conditions12,13, pollen tubes from each single mutant grew normally. We found that three independent insertion mutants for PRK6 completely lost their ability to react to AtLURE1.2, whereas all mutants of the other 22 genes, including seven other PRK genes, reacted to it (Fig. 1a, b). This semi-in-vivo result shows that, among the pollen-specific RLKs, PRK6 is essential for pollen tube reorientation towards the AtLURE1 attractant peptide.
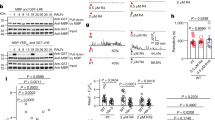
a, Pollen tube attraction assay for prk single mutants using AtLURE1.2. Ws, wild-type Arabidopsis ecotype Wassilewskija. b, Attracted wild-type (Columbia, Col-0) and insensitive prk6-1 mutant pollen tubes to AtLURE1 beads (asterisks). Arrowheads mark the tips of the pollen tubes. The data are representative of three images for each of Col-0 and prk6; in total, 12 or 10 tubes, respectively, showed similar growth properties. Scale bar, 20 μm. c, d, Complementation of the AtLURE1-insensitive prk6 phenotype by PRK6–mRuby2. In pollen tubes from hemizygous plants, mRuby2-positive (+) but not mRuby2-negative (−) pollen tubes responded to AtLURE1 beads (asterisks). The images are representative of 14 or 3 images for −mRuby or +mRuby, respectively. Scale bar, 20 μm. e, Pollen tube tip localization of PRK6–mRuby2 in a single-plane confocal image (top) and a pseudocolour intensity image (bottom). The data are representative of more than ten tubes. Scale bar, 10 μm. f, Semi-in-vivo pollen tube growth/AtLURE1-responsive assay for prk mutants 8.5 h after pollination (HAP). The data are representative of at least three assays. Note that, in addition to prk6, prk1 prk3 pollen tubes showed an impaired response to AtLURE1. Scale bars, 100 μm (top) and 10 μm (bottom).
PRK6 is one of eight PRK genes14, which encode transmembrane leucine-rich repeat (LRR) RLKs and are expressed specifically in the pollen tube13 (Extended Data Fig. 1a–d). To investigate the subcellular localization of PRK6, we introduced the pPRK6::PRK6-mRuby2 transgene into the prk6-1 mutant. Pollen tubes expressing PRK6 tagged to the red fluorescent protein mRuby2 (PRK6–mRuby2) displayed a functional response to AtLURE1 (Fig. 1c, d). In growing pollen tubes, PRK6–mRuby2 was localized predominantly at the plasma membrane of the tip, and detected in cytoplasmic granules with cytoplasmic streaming (Fig. 1e and Supplementary Video 1). These semi-in-vivo results show that PRK6 could contribute to the reception of an external AtLURE1 peptide at the pollen tube tip.
Studies of tomato LePRKs have suggested that PRK proteins act as signal-transducing receptors through association/dissociation of two PRK proteins15, and through interaction with several CRPs secreted from pollen and the pistil for pollen germination and growth stimulation16,17. We thus investigated whether other PRK family proteins function in pollen tube growth and attraction in combination with PRK6. Each prk single mutant and most prk multiple mutants of various combinations showed near-normal fertility (mean values, 85–100%), whereas triple mutants for PRK3, PRK6 and PRK8, which formed a single subclade (PRK3 subclass; Extended Data Fig. 1a), had a reduced seed set (mean values, 52–74%), and an additional mutation for PRK1, which has a gene structure similar to that of the PRK3 subclass genes, markedly reduced the seed set to ~10% (Extended Data Fig. 1e–g). We then analysed growth and responsiveness to AtLURE1 using a newly developed semi-in-vivo assay, in which pollen tubes grown on medium containing AtLURE1.2 peptide showed wavy and swollen tip growth in a concentration- and PRK6-dependent manner (Extended Data Fig. 2a–f and Supplementary Video 2). The AtLURE1-induced wavy morphology indicates a normal physiological response dependent on pollen tube competency13, because in vitro pollen tubes or semi-in-vivo chx21 chx23 mutant pollen tubes, which show a defect in ovule targeting but not in growth18, did not respond (Extended Data Fig. 2g, h). In this assay, we revealed that pollen tubes from multiple prk mutants that possess both prk3 and prk6 mutations exhibited a defect in growth, and more interestingly that prk1 prk3 double mutations as well as a prk6 single mutation impaired the response to AtLURE1.2 (Fig. 1f and Extended Data Fig. 2i). Expression of PRK3–mClover in prk3 prk6, which showed similar tip localization to that of PRK6, restored the growth defect but not the wavy response to AtLURE1 (Extended Data Fig. 2j, k), suggesting that they regulate different signalling pathways for pollen tube growth rather than redundant signalling pathways.
We investigated in vivo PRK functions. Consistent with semi-in-vivo results, the prk3 prk6 double mutant showed slow pollen tube growth as it reached the bottom of the transmitting tract at 24 h after pollination (HAP), compared with 12 HAP in wild-type and prk6 pollen tubes (Extended Data Fig. 3). We then observed pollen tube attraction towards wild-type ovules on the septum surface. Some pollen tubes of prk6 single and prk3 prk6 double mutants, but not the wild type, failed to target nearby ovules (Fig. 2a–c and Extended Data Fig. 4a–d), suggesting that these mutants are less sensitive to ovular attractants in vivo. Furthermore, these mutant pollen tubes showed slightly wandering phenotypes after reaching the ovules, although most ovules eventually attracted mutant pollen tubes (Fig. 2d, dark grey bars), as observed in the AtLURE1-deficient ovules6. More severe defects in growth and attraction in vivo were observed in prk3 prk6 prk8 and prk1 prk3 prk6 triple mutants (Fig. 2d and Extended Data Fig. 4e–g) and were correlated with their fertility. Our physiological analyses demonstrate that the PRK3 subclass and PRK1 could act together as signal-transducing receptors for efficient growth and attraction through sensing of external signalling molecules, including the AtLURE1 attractant peptide.
a–c, Ovule-targeting of wild-type (a), prk6 (b), and prk3 prk6 (c) pollen tubes on the septum surface in wild-type pistils. Asterisks mark ovules that did not attract near pollen tubes passing through them (arrowheads). The data are representative of 1–3 images for each genotype. Similar growth properties were observed in a total of 4 samples. Scale bar, 100 μm. For entire images of pistils, see Extended Data Fig. 4. d, Quantitative analysis of guidance on the ovule. White and light-grey stacked bars show wild-type guidance, in which one pollen tube grows straight on the funiculus and enters the micropyle (white), or an additional pollen tube(s) is associated on the funiculus (light grey). Dark grey bars show abnormal guidance on the ovule, in which a pollen tube takes a 180° turn back on the funiculus and then enters the micropyle. Black bars show ovules that are not associated with a pollen tube. Data are mean and s.d. of four pistils.
Next, we examined the intracellular signal transduction mechanism of PRK6. It has been reported that tomato LePRK1 and LePRK2 and A. thaliana PRK2 interact with ROPGEF family proteins7,8. ROPGEFs activate intracellular signalling switches, ROPs (Rho-like GTPases from plants), that control various cellular responses19,20,21,22. In bimolecular fluorescence complementation (BiFC) assays in tobacco leaf epidermal cells, PRK6 interacted with pollen-expressed ROPGEFs at the plasma membrane (Extended Data Fig. 5a–c). Furthermore, the BiFC assay showed that PRK6 interacted with itself, PRK3, and receptor-like cytoplasmic kinases, LIP1 and LIP2, which are involved in pollen tube growth and attraction and partly in AtLURE1 signalling23 (Extended Data Fig. 5d). These results indicated that PRK6 forms a complex with factors for proper tip-growth at the plasma membrane.
We then investigated the essential domain of PRK6 for interaction with ROPGEFs and signal transduction using truncated PRK6 proteins (Extended Data Fig. 6a). A co-immunoprecipitation assay demonstrated that a kinase-domain-deleted PRK6 mutant (K-del) as well as full-length PRK6 were associated with ROPGEF12 in planta, whereas a cytosolic domain-deleted PRK6 mutant (cyto-del-1) was not (Extended Data Fig. 5e). Corresponding to this, PRK6 (K-del), but not a modified cytosolic domain-deleted PRK6 (cyto-del-2), complemented the AtLURE1-insensitive phenotype of the prk6 single mutant (Extended Data Fig. 7). Interestingly, PRK6 (K-del) did not restore the growth defect caused by prk3 prk6 mutations (Extended Data Fig. 7). These results suggest that the membrane-spanning PRK6 interacts with the downstream ROPGEFs via the juxtamembrane domain (the region between the transmembrane and kinase domains) for sensing of AtLURE1, and that the kinase domain of PRK6 has an important role together with PRK3 in pollen tube growth.
Flowering plants have several PRK proteins. Eight orthologous PRK genes were found in the close relatives Arabidopsis lyrata and C. rubella (Extended Data Fig. 6c). To determine whether PRK6 is sufficient to confer pollen tube responsiveness to a species-specific AtLURE1 peptide, we generated C. rubella plants expressing mRuby2-fused A. thaliana PRK6, which has a diverged ectodomain compared with the C. rubella PRK6 orthologue (CrPRK6) (Extended Data Fig. 6d). In a semi-in-vivo assay using C. rubella pistils and pollen from T1 hemizygous C. rubella plants, which produce haploid wild-type and transgenic pollen tubes, wild-type C. rubella pollen tubes did not react to AtLURE1.2, whereas C. rubella pollen tubes expressing A. thaliana PRK6 acquired the ability to respond to AtLURE1.2 (Fig. 3a, b). When we used an A. thaliana pistil for semi-in-vivo growth of C. rubella pollen tubes, a similar ability was acquired (4%, n = 49 for wild type; 50%, n = 10 for PRK6–mRuby2 C. rubella in lines #2 and #3). In an opposite manner, we assessed the ability of CrPRK6 to perceive AtLURE1 in A. thaliana. Although expression of CrPRK6 restored the AtLURE1-insensitive phenotype of the prk6 single mutant, it only partially restored the AtLURE1-insensitive phenotype in the prk multiple mutants, unlike PRK6 of A. thaliana (Extended Data Fig. 7). All of our genetic data indicate that PRK6 acts as a key receptor for sensing of species-specific AtLURE1 attractants in cooperation with other PRKs of A. thaliana.
a, b, Attraction assay for C. rubella pollen tubes by AtLURE1.2 beads (asterisks). In pollen tubes from T1 hemizygous C. rubella plants, wild-type pollen tubes (top images in a; ‘−’ in b) did not respond to AtLURE1, whereas pollen tubes containing PRK6–mRuby2 (bottom images in a; ‘+’ in b) did respond. Arrowheads mark the tips of the pollen tubes. The data are representative of 12 or 9 images for −mRuby or +mRuby, respectively. Scale bar, 20 μm.
Finally, we tested the hypothesis that tip-focused PRK6 re-localizes to direct tip growth direction towards the AtLURE1 attractant. We performed time-lapse observation of the PRK6–mRuby-expressed pollen tube during reorientation towards Alexa488-labelled AtLURE1.2 (Supplementary Videos 3 and 4). Before and at the time of applying an AtLURE1 bead, PRK6 was observed symmetrically at the tip (Fig. 4a, g). Interestingly, PRK6 accumulated asymmetrically on the AtLURE1 bead side of the tip just before the pollen tube tip growth changed direction (Fig. 4b, c, g). The tip subregion where PRK6 had accumulated expanded gradually to change the growth direction towards AtLURE1 (Fig. 4d–f).
a–f, Time-lapse images of a PRK6–mRuby2 pollen tube during reorientation towards Alexa488-labelled AtLURE1.2. A gelatine bead containing Alexa488–AtLURE1.2 was placed on the medium at 0 s. Higher-magnification fluorescence intensity images of PRK6–mRuby2 are shown in the right panels in a–c. Asymmetric accumulation of PRK6–mRuby2 at the tip membrane was observed before a morphological change in the tip towards the AtLURE1 source (b, c, right panels). Fluorescence intensity (FI) along a white dashed line (2.5 μm from the front edge) is shown at the right. The sequential data are representative of 10 samples. Scale bars, 5 μm. For full time-lapse images, see Supplementary Videos 3 and 4. g, Fluorescence intensity of PRK6–mRuby2 along a line normal to the tip growth axis as shown in the right panels in a–c. Data for a further nine pollen tube samples and their mean lines are shown.
Here, we have shown that pollen tube tip-localized PRK6 regulates the direction of pollen tube tip growth as an essential receptor for AtLURE1 signalling. The pollen tube tip marker PRK6 could be re-localized asymmetrically by the external AtLURE1 peptide and may recruit the intracellular core tip growth machinery, such as ROPGEFs and ROP1 (ref. 2), for pollen tube reorientation. Furthermore, our results demonstrate that PRK6, in cooperation with other PRK family receptors, has a central role in the response to species-specific AtLURE1 and mediates efficient pollen tube growth in the pistil. In addition to studies on LePRK2 (refs 15, 16, 17), our genetic and physiological data suggest that pollen tube growth and attraction are fine-tuned via interactions among many receptors and multiple stimulants/attractants for successful reproduction. Although PRKs can potentially interact with CRPs16,17, the specific interaction between AtLURE1 and PRKs cannot be established because of AtLURE1 stickiness, which is mediated by a basic amino acid patch of AtLURE1 essential for its activity (Extended Data Fig. 8 and Supplementary Discussion). The sticky property and the attraction activity of AtLURE1 cannot be separated at present. Plants encode many CRPs (>800 genes in A. thaliana24) and RLKs with an extracellular domain (>450 genes in A. thaliana11). Plant surface receptors have evolved to recognize a variety of CRPs not only for cell differentiation25,26, expansion27 and self-recognition in the pollen–pistil interaction28, but also for positional signals for actively polarizing cells. It will be exciting to explore the molecular basis by which an assembly of receptors determines ligand specificity and to conduct real-time monitoring of ligand-induced signal transduction using the LURE attractant peptide.
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized, and investigators were not blinded to allocation during experiments and outcome assessment.
Plant materials
Arabidopsis thaliana accession Columbia (Col-0) was used as the wild type. Seeds of T-DNA insertion lines were obtained from ABRC and NASC, and T-DNA insertions were confirmed by genomic PCR (Extended Data Table 1). The insert sites were determined by sequencing of the PCR products, as described in Extended Data Fig. 1c. Plant growth conditions and transformation methods were described previously6. C. rubella seeds were obtained from ABRC (accession CS22697; ref. 29), and C. rubella plants in the rosette stage were subjected to vernalising cold treatment (8-h photoperiod at 4 °C for about 1 month) for flowering induction.
Collection of T-DNA insertion mutants of pollen-expressed RLKs
To investigate candidate RLKs responsible for AtLURE1 signalling, RLK genes encoding proteins with extracellular domains and displaying notable and specific expression in the pollen tube were selected as follows. Whether the more than 80 genes expressed in dry pollen or pollen tubes13 were expressed predominantly in the mature pollen was determined using the Arabidopsis eFP Browser (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi)30. Twenty-three pollen-dominant genes and their related genes were selected: PRK1–8 (see Extended Data Table 1), AT2G18470 (PROLINE-RICH EXTENSIN-LIKE RECEPTOR KINASE 4, PERK4), AT4G34440 (PERK5), AT3G18810 (PERK6), At1g49270 (PERK7), AT1G10620 (PERK11), AT1G23540 (PERK12), AT4G29450, AT3G13065 (STRUBBELIG-RECEPTOR FAMILY 4, SRF4), AT1G78980 (SRF5), AT4G18640 (MORPHOGENESIS OF ROOT HAIR 1, MRH1), AT5G45840, AT1G29750 (RECEPTOR-LIKE KINASE IN FLOWERS 1, RKF1), AT3G23750 (BAK1-ASSOCIATING RECEPTOR-LIKE KINASE 1, BARK1; or TMK4), AT1G19090 (CYSTEINE-RICH RLK 1, CRK1) and AT4G28670. A further five RLK genes of a subclass of the CrRLK1L family (AT3G04690 (ANXUR1), AT5G28680 (ANXUR2), AT4G39110, AT2G21480 and AT5G61350) were also pollen-dominant but were not examined in this study. T-DNA insertions in the coding or promoter regions of these selected 23 genes were identified by genomic PCR and sequencing of the PCR products. Semi-in-vivo pollen tubes from one or more lines for each gene were assessed by an attraction assay using the AtLURE1.2 peptide, as described below.
Semi-in-vivo attraction assay
Recombinant His-tagged AtLURE1.2 peptide was expressed in Escherichia coli, purified and refolded, as described previously6. The refolded His–AtLURE1.2 peptide was suggested to be a conformational isomer by reverse-phase high-pressure liquid chromatography (HPLC) using a Phenomenex Jupiter C18 column and a Jasco analytical instrument equipped with a UV-2077 plus detector and PU-2080 plus pumps. A construct for His–AtLURE1.2(GGGG) was generated from pET-28a-AtLURE1.2 by site-directed mutagenesis using the primers 5′-GTATGgGAgGGGGTggGTATATTC-3′ and 5′-cACCCCcTCcCATACAAGCTC-3′ (lowercase bases denote mutated bases from the original AtLURE1.2). No aggregation due to inappropriate folding was observed during refolding or concentration of the His–AtLURE1.2(GGGG) peptide. Alexa488-labelled His–AtLURE1.2 was produced using the refolded His–AtLURE1.2 peptide and the Alexa Fluor 488 Protein Labelling Kit (Thermo Fisher Scientific), according to the manufacturer’s protocol. For the semi-in-vivo attraction assay, pollen tubes were grown through cut styles of A. thaliana on solid pollen germination medium poured into a mould made with 2-mm thick silicone rubber and cover glasses31. About 4–5 h after hand-pollination, the topside cover glass was removed and the medium was covered with hydrated silicone oil (KF-96-100CS; Shin-Etsu). The assay for T1 hemizygous C. rubella plants was performed similarly using A. thaliana or C. rubella pistils as pollen acceptors. Attraction of pollen tubes towards the peptide was evaluated using gelatine beads (5% (w/v) gelatine (Nacalai) in the pollen medium without agar) containing 5 μM His-tagged AtLURE1.2 peptide under an inverted microscope (IX71, Olympus) equipped with a micro-manipulator (Narishige), as described previously6. The percentages of attracted pollen tubes are shown for the total number of pollen tubes in at least two assays. In the assay using hemizygous plants, the presence of the transgene in the pollen tube containing the transgene was confirmed by fluorescence observations after assessment of pollen tube responsiveness as a simple blind test. For the AtLURE1-responsive wavy assay, the purified AtLURE1.2 peptide was added to solid pollen germination medium, which was melted at 70 °C and then cooled to a certain degree. The mixture was mixed by vortexing and poured into the mould. Pollen tubes of each genotype were grown through cut styles, as described earlier.
Binary vector construction, genetic transformation, and selection of transformants
Plasmids encoding green and red fluorescent proteins, pcDNA3-Clover and pcDNA3-mRuby2 (gifts from M. Lin, Addgene plasmids 40259 and 40260)32, respectively, were used as templates to prepare binary vectors as follows. The original Clover was converted to A206K mutant form to prevent potential dimerization, and a restriction site KpnI in the nucleotide sequence was eliminated by a silent mutation, designated as monomeric Clover (mClover). Modified binary vectors pPZP211, pPZP221 (ref. 33) and pMDC99 (ref. 34) derivatives, pPZP211G (ref. 35), pPZP221G, and pMDC99G, were used for cloning of the mClover and mRuby2. pPZP221G was produced by the same procedure as that used for pPZP211G (ref. 35), and pMDC99G was produced by removal of ccdB by EcoRI digestion and self-ligation31 and by inserting multiple cloning sites, green fluorescent protein (GFP), and the NosT cassette of pPZP211G via HindIII and EcoRI sites. To add linkers to both the amino-terminal and carboxy-terminal of mClover and mRuby2, three rounds of PCR were performed with DNA templates for mClover and mRuby2, respectively, using three sets of primers: (5′-aggtggaggtggaATGGTGAGCAAGGGCGA-3′ and 5′-tccacctccacctgaCTTGTACAGCTCGTCCA-3′; 5′-tctggaggtggaggttcAGGTGGAGGTGGA-3′ and 5′-cggggtacccactagtttaattaagaattcTCCACCTCCACCTG-3′; 5′-aggcgcgccTCTGGAGGTGGAG-3′ and 5′-cggggtacccactagtttaattaagaattcTCCACCTCCACCTG-3′) (lowercase bases denote additional nucleotides for template DNAs). The PCR fragments were digested with AscI and KpnI and ligated into pPZP211G, pPZP221G and pMDC99G by replacing the GFP sequence, resulting in pPZP211Clo, pPZP221Clo, pPZP211Ru, pPZP221Ru, pMDC99Clo and pMDC99Ru vectors.
For the expression of full-length PRK6, kinase domain-deleted PRK6 (K-del), cytosolic domain-deleted PRK6 (cyto-del-2) and PRK6 orthologue of C. rubella (CrPRK6) as mRuby2-fusion protein under the control of their own promoter, genomic sequences of PRK6 or CrPRK6 containing promoter and coding regions were amplified and were cloned into the pPZP221Ru using SalI and AscI sites, resulting in pPZP221-pPRK6::PRK6-mRuby2, -pPRK6::PRK6 (K-del)-mRuby2, -pPRK6::PRK6 (cyto-del-2)-mRuby2, and -pCrPRK6::CrPRK6-mRuby2 vectors. These constructs were introduced into prk6-1, prk3-1 prk6-1, prk3-1 prk6-1 prk8-2 and prk1-2 prk3-1 prk6-1 plants by the floral dip method. For the heterologous expression of PRK6 in C. rubella, the pPZP221-pPRK6::PRK6-mRuby2 vector was used for C. rubella transformation by the floral dip method after flowering induction. Genomic sequences of PRK6 or PRK3 containing promoter and coding regions were also cloned into pMDC99Clo using SalI and AscI sites, and these constructs were introduced into prk3-1 prk6-1. Primers used for these constructs are listed in Supplementary Table 1.
For all transgenic lines expressing PRK proteins, T1 transformants were screened by moderate or weak fluorescence intensity in approximately half of the pollen grains, implying single insertion. Note, when pollen grains showing mid to strong fluorescence intensity were used for the semi-in-vivo pollen tube growth assay, few or no fluorescent pollen tubes emerged from the cut end, probably owing to the growth defect caused by excess PRK expression. T2 homozygous plants obtained from several selected T1 lines were used for the semi-in-vivo AtLURE1-responsive wavy assay.
BiFC assay
To prepare constructs for the BiFC assay in the leaf epidermal cells of Nicotiana benthamiana, cauliflower mosaic virus 35S promoter was introduced to the binary vector pPZP211G (ref. 35) using HindIII and PstI sites. Then, the GFP sequence was replaced by nucleotide sequences encoding each of amino acids 1–174 and 175–239 of enhanced yellow fluorescent protein (nYFP and cYFP, respectively) with the same linkers as the mClover and mRuby2 constructs, described above, resulting in pPZP211-p35SnY and pPZP211-p35ScY vectors. Genomic PRK2 and PRK6 were amplified and connected upstream of the cYFP sequence of pPZP211-p35ScY. The genomic sequences of PRK6, PRK3, LIP1 and LIP2 were connected upstream of the nYFP sequence of pPZP211-p35SnY. Genomic ROPGEF8, ROPGEF9, ROPGEF12, ROPGEF13 and ROPGEF12ΔC (encoding amino acids 1–443 of ROPGEF12 (ref. 8)) were amplified and connected downstream of the nYFP sequence in pPZP211-p35SnY. Primers used for these constructs are listed in Supplementary Table 1.
Transient expression in N. benthamiana leaves was performed by agro-infiltration according to a method described previously20. In brief, Agrobacterium tumefaciens strains GV3101 (pMP90) containing each expression vector were cultured overnight in LB media. Equal amounts of Agrobacterium cultures for nYFP and cYFP constructs and the p19 silencing suppressor were mixed to a final A600 nm of 1.0 and collected and resuspended in infiltration buffer (10 mM MES, pH 5.6, 10 mM MgCl2 and 150 μM acetosyringone). The mixed suspensions were incubated at room temperature for ~3 h and infiltrated into leaves of N. benthamiana grown at 25 °C. Two to three days after infiltration, the leaves were cut into pieces for confocal microscope observation.
Analyses of pollen tube growth and guidance in pistils
To analyse pollen tube growth and guidance in the pistil, Col-0 pistils emasculated 1 day before were abundantly hand-pollinated with two or three fully dehiscent anthers from each genotype. Two types of aniline blue staining were performed 12 or 24 h after pollination as follows. For measurement of pollen tube growth inside the transmitting tract, aniline blue staining was performed, as described previously36. Pollinated pistils were dissected to remove a pair of ovary walls and then fixed in a 9:1 mixture of ethanol and acetic acid for more than 2 h. They were washed with 70% ethanol for ~30 min, treated with 1 N NaOH overnight, and stained with aniline blue solution (0.1% (w/v) aniline blue, 0.1 M K3PO4) for more than several hours. The pistils were observed under ultraviolet illumination using an upright microscope (DP71, Olympus). Multiple images for each pistil were combined using Adobe Photoshop CS4 (Adobe Systems), and lengths from the top of the stigma to the tip of the longest pollen tube were measured for maximum pollen tube length using the MacBiophotonics ImageJ software (http://www.macbiophotonics.ca/).
To evaluate pollen tube guidance after emergence on the septum surface of the pistil, dissected pistils were stained directly with modified aniline blue solution (5:8:7 (v/v) mixture of 2% aniline blue, 1 M glycerol, pH 9.5, and water), as described previously37, and observed under ultraviolet illumination using an upright microscope (DP71, Olympus). Quantitative analysis was performed by evaluating pollen tube growth on 10 upper ovules of both sides (total, 20 ovules per pistil) to eliminate bias in ovule number in a pistil.
Confocal microscopy
Confocal images were acquired using an inverted microscope (IX81, Olympus) equipped with a spinning disk confocal scanner (CSU-X1, Yokogawa Electric Corporation), 488 nm and 561 nm LD lasers (Sapphire, Coherent), and an EM-CCD camera (Evolve 512, Photometrics). For A. thaliana pollen tubes, a 60× silicone immersion objective lens (UPLSAPO60XS, Olympus) and a 1.6× intermediate magnification changer were used. For time-lapse imaging of PRK6–mRuby2 during pollen tube attraction towards a gelatine bead containing 5 μM Alexa488-labelled His-AtLURE1.2, sequential images using 488 nm and 561 nm lasers were acquired every 5 s. For the BiFC assay in N. benthamiana leaves, a 20× objective lens (UPLFLN20X, Olympus) was used. The confocal microscope system was controlled and time-lapse images were processed by MetaMorph (Universal Imaging). Images were edited with MacBiophotonics ImageJ.
Co-immunoprecipitation assay
To prepare transient expression vectors in N. benthamiana leaf cells, the cauliflower mosaic virus 35S promoter was introduced into the binary vector pPZP211Clo via HindIII and PstI sites, resulting in the pPZP211-p35SClo vector. The 3× Flag tag sequence was introduced into pPZP211-p35S using the AscI and SacI sites, resulting in the pPZP211-p35SFlag vector.
For co-immunoprecipitation of PRK-mClover and ROPGEF12-3 × Flag proteins, genomic sequences of full-length PRK3, full-length PRK6, PRK6 (K-del), PRK6 (cyto-del-1), and ROPGEF12 were inserted into the pPZP211-p35SClo or pPZP211-p35SFlag vectors. One of the PRK-mClover or mClover proteins plus the p19 silencing suppressor and ROPGEF12-3 × Flag were co-expressed in N. benthamiana leaves as described for the BiFC assay. The leaves were ground in mortars with liquid nitrogen and suspended in 3–3.5 × (w/v) extraction buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% glycerol, protease inhibitor cocktail (cOmplete EDTA-free, Roche)). The extracts were centrifuged twice at 10,000g for 10 min at 4 °C to remove precipitates. The supernatants, with the exception of the mClover sample, were ultracentrifuged at 100,000g for 30 min at 4 °C, and the pellets were solubilized in extraction buffer containing 0.5% Triton X-100. The solubilised membrane fraction samples and mClover sample plus 0.5% Triton X-100 were incubated with GFP-trap agarose beads (ChromoTek, gta-20) with rotation for 2 h at 4 °C. The beads were washed with buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl) four times. Then, the bound proteins were eluted with SDS sample buffer by heating at 70 °C for 5 min. The protein samples were separated on SDS–PAGE and subjected to immunoblot analysis. The immunoblot analysis was conducted on PVDF membranes (Immobilon-P, Millipore) using primary antibodies (anti-GFP (ab290, Abcam), or monoclonal anti-DYKDDDDK tag (Wako) for Flag tag) and secondary antibodies (goat anti-rabbit IgG peroxidase-labelled antibody or goat anti-mouse IgG peroxidase-labelled antibody (KPL)). Signals were visualized using Immobilon Western Chemiluminescent HRP Substrate (Millipore), detected with Light-Capture (ATTO).
References
Itofusa, R. & Kamiguchi, H. Polarizing membrane dynamics and adhesion for growth cone navigation. Mol. Cell. Neurosci. 48, 332–338 (2011)
Yang, Z. Cell polarity signaling in Arabidopsis. Annu. Rev. Cell Dev. Biol. 24, 551–575 (2008)
Higashiyama, T. & Takeuchi, H. The mechanism and key molecules involved in pollen tube guidance. Annu. Rev. Plant Biol. 66, 393–413 (2015)
Okuda, S. et al. Defensin-like polypeptide LUREs are pollen tube attractants secreted from synergid cells. Nature 458, 357–361 (2009)
Márton, M. L., Fastner, A., Uebler, S. & Dresselhaus, T. Overcoming hybridization barriers by the secretion of the maize pollen tube attractant ZmEA1 from Arabidopsis ovules. Curr. Biol. 22, 1194–1198 (2012)
Takeuchi, H. & Higashiyama, T. A species-specific cluster of defensin-like genes encodes diffusible pollen tube attractants in Arabidopsis. PLoS Biol. 10, e1001449 (2012)
Kaothien, P. et al. Kinase partner protein interacts with the LePRK1 and LePRK2 receptor kinases and plays a role in polarized pollen tube growth. Plant J. 42, 492–503 (2005)
Zhang, Y. & McCormick, S. A distinct mechanism regulating a pollen-specific guanine nucleotide exchange factor for the small GTPase Rop in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 104, 18830–18835 (2007)
Higashiyama, T. et al. Pollen tube attraction by the synergid cell. Science 293, 1480–1483 (2001)
Jones-Rhoades, M. W., Borevitz, J. O. & Preuss, D. Genome-wide expression profiling of the Arabidopsis female gametophyte identifies families of small, secreted proteins. PLoS Genet. 3, e171 (2007)
Shiu, S. H. & Bleecker, A. B. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl Acad. Sci. USA 98, 10763–10768 (2001)
Palanivelu, R. & Preuss, D. Distinct short-range ovule signals attract or repel Arabidopsis thaliana pollen tubes in vitro. BMC Plant Biol. 6, 7 (2006)
Qin, Y. et al. Penetration of the stigma and style elicits a novel transcriptome in pollen tubes, pointing to genes critical for growth in a pistil. PLoS Genet. 5, e1000621 (2009)
Chang, F., Gu, Y., Ma, H. & Yang, Z. AtPRK2 promotes ROP1 activation via RopGEFs in the control of polarized pollen tube growth. Mol. Plant 6, 1187–1201 (2013)
Wengier, D. et al. The receptor kinases LePRK1 and LePRK2 associate in pollen and when expressed in yeast, but dissociate in the presence of style extract. Proc. Natl Acad. Sci. USA 100, 6860–6865 (2003)
Tang, W., Ezcurra, I., Muschietti, J. & McCormick, S. A cysteine-rich extracellular protein, LAT52, interacts with the extracellular domain of the pollen receptor kinase LePRK2. Plant Cell 14, 2277–2287 (2002)
Tang, W., Kelley, D., Ezcurra, I., Cotter, R. & McCormick, S. LeSTIG1, an extracellular binding partner for the pollen receptor kinases LePRK1 and LePRK2, promotes pollen tube growth in vitro. Plant J. 39, 343–353 (2004)
Lu, Y. et al. Pollen tubes lacking a pair of K+ transporters fail to target ovules in Arabidopsis. Plant Cell 23, 81–93 (2011)
Berken, A. Thomas. C. & Wittinghofer. A. A new family of RhoGEFs activates the Rop molecular switch in plants. Nature 436, 1176–1180 (2005)
Oda, Y. & Fukuda, H. Initiation of cell wall pattern by a Rho- and microtubule-driven symmetry breaking. Science 337, 1333–1336 (2012)
Li, H., Lin, Y., Heath, R. M., Zhu, M. X. & Yang, Z. Control of pollen tube tip growth by a Rop GTPase-dependent pathway that leads to tip-localized calcium influx. Plant Cell 11, 1731–1742 (1999)
Gu, Y. et al. A Rho family GTPase controls actin dynamics and tip growth via two counteracting downstream pathways in pollen tubes. J. Cell Biol. 169, 127–138 (2005)
Liu, J. et al. Membrane-bound RLCKs LIP1 and LIP2 are essential male factors controlling male-female attraction in Arabidopsis. Curr. Biol. 23, 993–998 (2013)
Silverstein, K. A. et al. Small cysteine-rich peptides resembling antimicrobial peptides have been under-predicted in plants. Plant J. 51, 262–280 (2007)
Lee, J. S. et al. Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes Dev. 26, 126–136 (2012)
Lee, J. S. et al. Competitive binding of antagonistic peptides fine-tunes stomatal patterning. Nature 522, 439–443 (2015)
Haruta, M., Sabat, G., Stecker, K., Minkoff, B. B. & Sussman, M. R. A peptide hormone and its receptor protein kinase regulate plant cell expansion. Science 343, 408–411 (2014)
Takayama, S. et al. Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature 413, 534–538 (2001)
Slotte, T. et al. The Capsella rubella genome and the genomic consequences of rapid mating system evolution. Nature Genet. 45, 831–835 (2013)
Winter, D. et al. An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007)
Hamamura, Y. et al. Live imaging of calcium spikes during double fertilization in Arabidopsis. Nature Commun. 5, 4722 (2014)
Lam, A. J. et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nature Methods 9, 1005–1012 (2012)
Hajdukiewicz, P., Svab, Z. & Maliga, P. The small, versatile pPZP family of Agrobacterium binary vectors for plant transformation. Plant Mol. Biol. 25, 989–994 (1994)
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003)
Susaki, D., Takeuchi, H., Tsutsui, H., Kurihara, D. & Higashiyama, T. Live imaging and laser disruption reveal the dynamics and cell-cell communication during Torenia fournieri female gametophyte development. Plant Cell Physiol. 56, 1031–1041 (2015)
Maruyama, D. et al. Independent control by each female gamete prevents the attraction of multiple pollen tubes. Dev. Cell 25, 317–323 (2013)
Nagahara, S., Takeuchi, H. & Higashiyama, T. Generation of a homozygous fertilization-defective gcs1 mutant by heat-inducible removal of a rescue gene. Plant Reprod. 28, 33–46 (2015)
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30, S162–S173 (2009)
Miyazaki, S. et al. ANXUR1 and 2, sister genes to FERONIA/SIRENE, are male factors for coordinated fertilization. Curr. Biol. 19, 1327–1331 (2009)
Wrzaczek, M. et al. GRIM REAPER peptide binds to receptor kinase PRK5 to trigger cell death in Arabidopsis. EMBO J. 34, 55–66 (2015)
Acknowledgements
We thank M. Hasebe and S. Miyazaki for seeds of a part of mutant lines; Y. Matsubayashi, H. Shinohara, D. Maruyama, M. Ohtsu and K. Motomura for valuable comments and discussions; D. Kurihara, Y. Hamamura and S. Nagahara for technical assistance with confocal microscopy and physiological analyses; M. M. Kanaoka for agro-infiltration method using tobacco leaf; S. Oishi for reverse-phase high-pressure liquid chromatography (HPLC); and the Japan Advanced Plant Science Network for use of some microscopes. This work was supported by grants from the Japan Science and Technology Agency (ERATO project to T.H.) and the Japan Society for the Promotion of Science Fellowships (no. 5834 to H.T.).
Author information
Authors and Affiliations
Contributions
H.T. designed the study; H.T. performed experiments; H.T. and T.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Phylogenetic relationship, expression, gene structure and fertility of A. thaliana PRK family protein mutants.
a, A neighbour-joining (NJ) tree constructed using full-length amino-acid sequences of PRK1–PRK6 (ref. 14), PRK7 and PRK8 (assigned in this study). The bootstrap values as percentages and the scale for substitutions per site are shown. b, PRK expression during pollen germination and growth. Expression levels are shown using normalized values and standard deviation from microarray data (n = 4 for dry pollen, 30 min in vitro pollen tube (PT), and 4 h in vitro PT; n = 3 for semi-in-vivo PT)13. c, Structure and T-DNA insertion of PRK genes. Grey boxes show exons, and black boxes show introns or untranslated regions that are registered in The Arabidopsis Information Resource (TAIR). The T-DNA insertion sites determined by genomic PCR and sequencing are drawn on the gene structure and indicated in Extended Data Table 1. d, Reverse transcription PCR (RT–PCR) analysis of the prk single mutants. Anther cDNA was used for the analysis. Positions of the primers are indicated in the gene structure (c). ACT2 was used as the loading control. For gel source data, see Supplementary Fig. 1. e, f, The rate of developing seeds upon self-pollination of prk single mutants (e) and upon reciprocal crosses with Col-0 and prk multiple mutants (f). Asterisks in e indicate the mutants used for the prk multiple mutants in this study. Note that, in addition to multiple mutants of PRK1 and PRK3 subclass genes (shown in dark blue), multiple mutants of PRK1, PRK4 and PRK6, which are the top three most highly expressed in semi-in-vivo pollen tubes, and PRK1, PRK2, PRK4 and PRK5, which form another subclade, were analysed. The prk1 prk2 prk4 prk5 multiple mutant contains prk1 prk2 prk5 mutations that cause reduced pollen tube growth in vitro14. Data are mean and s.d. of three (all samples in e; Col-0 pistil × prk3-1 prk6-1 pr8-1 and prk3-1 prk6-1 pr8-2 in f) or four (other samples in f) pistils. g, Developing seeds in siliques 8 days after pollination with Col-0, prk6 and the prk1 prk3 prk6 triple mutant. The images are representative of four samples. Scale bar, 1 mm.
Extended Data Figure 2 Evaluation of AtLURE1-responsive wavy assay.
a–f, Semi-in-vivo pollen tubes on medium containing the indicated concentrations of AtLURE1.2 peptide. Entire (a–d) and magnified (e, f) images show wavy and swollen tip growth of wild-type, but not prk6-1 mutant, pollen tubes in a concentration-dependent manner. Scale bars, 200 μm (a–d) and 20 μm (e, f). g, Growth of PRK6–mRuby2 pollen tubes that directly germinated on medium (that is, in vitro pollen tubes) containing 1 μM AtLURE1.2 peptide. h, Growth of semi-in-vivo pollen tubes from chx21-s1/chx21-s1 chx23-4/CHX23 plant18 on medium containing 1 μM AtLURE1.2 peptide. Roughly half the pollen tubes showed wavy growth as in the wild type (arrowheads), but the rest did not (arrows). These results indicate that in vitro pollen tubes and chx21 chx23 double-mutant pollen tubes have no or less ability to respond to the external AtLURE1 peptide. Scale bars, 200 μm. i, Semi-in-vivo pollen tube growth and AtLURE1-responsive wavy assay for prk mutants additional to those shown in Fig. 1f. Scale bars, 100 μm (top) and 10 μm (bottom). j, Complementation of the growth defect in prk3 prk6 pollen tubes by expression of PRK3–mClover or PRK6–mClover. Note that PRK3–mClover expression restored the growth defect but not the wavy response. The images of a–j are representative of at least three assays. k, Pollen tube tip localization of PRK3–mClover in a single-plane confocal image (top) and its intensity image by pseudocolour (bottom). The data are representative of three samples. Scale bar, 10 μm.
Extended Data Figure 3 Pollen tube growth of prk mutants in the pistil.
a, Pollen tubes of Col-0, prk6, prk3 prk6 and prk1 prk3 prk6 growing in the Col-0 pistils. Aniline blue staining was performed 12 or 24 HAP. White arrows indicate the tip of the longest pollen tube in the transmitting tract. Data are representative of three samples for each genotype. Scale bar, 500 μm. b, Length from the top of the stigma to the tip of the longest pollen tube, 12 or 24 HAP with Col-0 and prk mutants. About 2,700 μm is the maximum limit for the length in this measurement. The data are the mean and s.d. of three pistils. ND, no data.
Extended Data Figure 4 Growth and ovule-targeting of prk mutant pollen tubes on the septum surface.
a–g, Entire images of growth and ovule-targeting of wild-type (a), prk6 (b), prk3 prk6 (c, d), prk3 prk6 prk8-2 (e, f), and prk1 prk3 prk6 (g) pollen tubes on the septum surface in wild-type pistils. Arrows indicate the tip of the longest pollen tube on the septum surface. Asterisks mark ovules that did not attract near pollen tubes. The regions shown in a, b and d are shown in Fig. 2e, f and g, respectively, as higher magnification images. Data are representative of 1–3 images for each genotype. Similar growth properties were observed in a total of 4 samples. Scale bar, 500 μm. Quantitative analysis is shown in Fig. 2d. No analysis was performed for the prk1 prk3 prk6 mutant because almost no pollen tube reached the ovule.
Extended Data Figure 5 Interaction of PRK6 with pollen-expressed ROPGEFs, PRKs and LIPs.
a, Gene expression of ROPGEFs during pollen germination and growth. The data are normalized expression values and standard deviation from microarray data (n = 4 for dry pollen, 30 min in vitro PT, and 4 h in vitro PT; n = 3 for semi-in-vivo PT)13 as noted in Extended Data Fig. 1b. ROPGEF8, ROPGEF9, ROPGEF11, ROPGEF12 and ROPGEF13 are expressed specifically in the dry pollen grain and pollen tube. b, BiFC assay showing the interaction between PRK6–cYFP and nYFP–GEF8, nYFP–GEF9, nYFP–GEF12, or nYFP–GEF13 (see Methods). c, A control experiment using C-terminal-deleted ROPGEF12 (ROPGEF12ΔC). The C-terminal domain is suggested to mediate the interaction with PRK2 (ref. 8). d, BiFC assay showing interaction between PRK6–cYFP and PRK6–nYFP, PRK3–nYFP, LIP1–nYFP or LIP2–nYFP. Scale bars, 50 μm. Images are representative of more than three experiments. e, Co-immunoprecipitation assay of PRK–mClover and ROPGEF12 proteins expressed in N. benthamiana leaf cells. ROPGEF12-3 × Flag protein was precipitated with full-length PRK3, PRK6 and kinase domain-deleted PRK6 (K-del), but not mClover control or cytosolic domain-deleted PRK6 (cyto-del-1). Data are representative of three experiments. For gel source data, see Supplementary Fig. 2.
Extended Data Figure 6 PRK6 protein structure and PRK proteins of A. thaliana, A. lyrata and C. rubella.
a, Structures of the PRK6 protein and its deletion version used in this study. The PRK6 extracellular domain contains the N-terminal cap and six LRRs. JM, juxtamembrane domain; N-cap, N-terminal cap; SP, signal peptide; TM, transmembrane domain. The numbers indicate the amino acid ranges of each domain. b, A 3D ribbon model of the PRK6 extracellular domain, amino acid residues 28–231, was predicted using the homology modelling platform, SWISS-MODEL (http://swissmodel.expasy.org/), and the FLS2 crystal structure (Protein Data Bank (PDB) accession 4MN8) as a template, and was drawn using Swiss-PdbViewer (http://spdbv.vital-it.ch/)38. The PRK6 extracellular domain contains the N-terminal cap and six LRRs. c, A neighbour-joining tree constructed using PRK protein sequences from tomato (Lycopersicon esculentum, LePRK1-3), A. thaliana (AtPRK1–AtPRK8), A. lyrata (AlPRK1–AlPRK8), and C. rubella (CrPRK1–CrPRK8). The bootstrap values as percentages and the scale for substitutions per site are shown. Accession numbers for A. lyrata and C. rubella PRKs: AlPRK1 (XP_002868416), AlPRK2 (XP_002883746), AlPRK3 (XP_002877261), AlPRK4 (XP_002883234), AlPRK5 (XP_002891583), AlPRK6 (XP_002871954), AlPRK7 (XP_002867307, modified according to the genome sequence), AlPRK8 (XP_002887434), CrPRK1 (EOA19015), CrPRK2 (EOA32286, partial sequence), CrPRK3 (EOA25493), CrPRK4 (EOA31871), CrPRK5 (EOA37472), CrPRK6 (EOA23063), CrPRK7 (EOA18255), and CrPRK8 (EOA34527). d, A sequence alignment of AtPRK3, AtPRK6 and CrPRK6. Signal peptide, N-terminal cap, LRR1–LRR6, transmembrane domain and kinase domain are indicated beneath the alignment.
Extended Data Figure 7 Semi-in-vivo pollen tube growth and response to the AtLURE1 peptide of PRK6 variant mutants.
Pollen tubes of prk6, prk3 prk6, prk3 prk6 prk8-2 and prk1 prk3 prk6 mutants were assessed in this assay. Full-length PRK6, the PRK6 orthologue of C. rubella (CrPRK6), kinase-domain-deleted PRK6 (K-del), and cytosolic-domain-deleted PRK6 (cyto-del-2) were expressed as mRuby2 fusion proteins under the control of their own promoters. Upper differential interference contrast images show semi-in-vivo pollen tube growth in the medium containing the AtLURE1 peptide at 6 HAP. Yellow arrowheads mark some of the pollen tubes showing apparent wavy phenotype. The bottom two images are a blight field image and a confocal image for mRuby2 of a representative pollen tube in the wavy assay. The data are representative images of at least three assays for one or two lines of each genotype. Scale bars, 200 μm (top) and 20 μm (bottom).
Extended Data Figure 8 A conserved basic amino acid patch of LURE is essential for attraction.
a, The sequence of full-length AtLURE1.2 accompanied by lysine/arginine residues (yellow highlight) mutated to glycines for AtLURE1.2(GGGG). Cysteine residues in the mature peptide are shown in red. b, c, Semi-in-vivo attraction assay using gelatine beads containing 5 μM His–AtLURE1.2(GGGG) (b) and wavy assay using 10 μM His–AtLURE1.2(GGGG) in the medium (c). The AtLURE1.2(GGGG) peptide showed no activity in these assays. The data are representative of 14 or 3 samples for b or c, respectively. Scale bars, 20 μm (b) and 200 μm (c).
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and Supplementary Figures 1-2 which show uncropped scans with size marker indications. (PDF 289 kb)
Supplementary Table 1
This table contains a list of primers used in the study. (XLS 34 kb)
Time lapse imaging of PRK6 mRuby2 during pollen tube tip growth
PRK6 mRuby2 was observed predominantly at the plasma membrane of the tip and detected in cytoplasmic granules with cytoplasmic streaming. Scale bar, 10 μm. (MOV 1215 kb)
Time lapse imaging of semi-in vivo pollen tube growth and response to the AtLURE1 peptide
Pollen tubes of wild-type (Col-0), prk6, and prk6 expressing PRK6-mRuby2 (PRK6-mRuby2) grew straight in the absence of AtLURE1 peptides in the medium (-). In this observation, the pollen tube growth rate at 3-4 h after pollination (180-240 min) was 200 ± 25 µm/h in the wild type and 151 ± 18 µm/h in prk6. Col-0 and PRK6-mRuby2, but not prk6, showed the wavy growth phenotype in the presence of 1 µM AtLURE1.2 in the medium (+). (MOV 2319 kb)
Time lapse imaging of PRK6 mRuby2 and Alexa488-labeled AtLURE1.2 during pollen tube reorientation
Time-lapse images captured every 5 s shown in Fig. 4a-f. Overlaid images of PRK6-mRuby2 and Alexa488-labeled AtLURE1.2 (left) and intensity images for PRK6-mRuby2 (right) are shown. A gelatine bead containing 5 µM Alexa488-labeled AtLURE1.2 was used in this assay. Scale bar, 10 μm. (MOV 3680 kb)
An additional example of time lapse imaging of PRK6 mRuby2 and Alexa488-labeled AtLURE1.2 during pollen tube reorientation
An additional example of the assay shown in Supplementary Video 3. Scale bar, 10 μm. (MOV 5087 kb)
Rights and permissions
About this article
Cite this article
Takeuchi, H., Higashiyama, T. Tip-localized receptors control pollen tube growth and LURE sensing in Arabidopsis. Nature 531, 245–248 (2016). https://doi.org/10.1038/nature17413
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17413
- Springer Nature Limited
This article is cited by
-
Deep imaging reveals dynamics and signaling in one-to-one pollen tube guidance
EMBO Reports (2024)
-
Synergid cell calcium oscillations refine understanding of FERONIA/LORELEI signaling during interspecific hybridization
Plant Reproduction (2024)
-
Pollen tube emergence is mediated by ovary-expressed ALCATRAZ in cucumber
Nature Communications (2023)
-
Two aspartic proteases, BnaAP36s and BnaAP39s, regulate pollen tube guidance in Brassica napus
Molecular Breeding (2023)
-
Whole-mount RNA in situ hybridization technique in Torenia ovules
Plant Reproduction (2023)