Abstract
The extent to which low‐frequency (minor allele frequency (MAF) between 1–5%) and rare (MAF ≤ 1%) variants contribute to complex traits and disease in the general population is mainly unknown. Bone mineral density (BMD) is highly heritable, a major predictor of osteoporotic fractures, and has been previously associated with common genetic variants1,2,3,4,5,6,7,8, as well as rare, population‐specific, coding variants9. Here we identify novel non‐coding genetic variants with large effects on BMD (ntotal = 53,236) and fracture (ntotal = 508,253) in individuals of European ancestry from the general population. Associations for BMD were derived from whole‐genome sequencing (n = 2,882 from UK10K (ref. 10); a population‐based genome sequencing consortium), whole‐exome sequencing (n = 3,549), deep imputation of genotyped samples using a combined UK10K/1000 Genomes reference panel (n = 26,534), and de novo replication genotyping (n = 20,271). We identified a low‐frequency non‐coding variant near a novel locus, EN1, with an effect size fourfold larger than the mean of previously reported common variants for lumbar spine BMD8 (rs11692564(T), MAF = 1.6%, replication effect size = +0.20 s.d., Pmeta = 2 × 10−14), which was also associated with a decreased risk of fracture (odds ratio = 0.85; P = 2 × 10−11; ncases = 98,742 and ncontrols = 409,511). Using an En1cre/flox mouse model, we observed that conditional loss of En1 results in low bone mass, probably as a consequence of high bone turnover. We also identified a novel low‐frequency non‐coding variant with large effects on BMD near WNT16 (rs148771817(T), MAF = 1.2%, replication effect size = +0.41 s.d., Pmeta = 1 × 10−11). In general, there was an excess of association signals arising from deleterious coding and conserved non‐coding variants. These findings provide evidence that low‐frequency non‐coding variants have large effects on BMD and fracture, thereby providing rationale for whole‐genome sequencing and improved imputation reference panels to study the genetic architecture of complex traits and disease in the general population.
Similar content being viewed by others
Main
Recently, genetic discoveries have generally focused on common variants of small effect and rare coding variants identified through genome‐wide association studies (GWAS) and whole‐exome sequencing initiatives, respectively11,12. The effect of low‐frequency and rare non‐coding variants upon common diseases, and their underlying traits has been recently explored in an isolated population13,14, but has not been well‐studied to date in the general population. The UK10K project has generated a large whole‐genome sequence‐based resource to address this question in a general European‐ancestry population10, which is tenfold larger than the European subset of the 1000 Genomes project reference15.
Osteoporosis, diagnosed mainly through measurement of bone mineral density (BMD), is a common systemic skeletal disease characterized by an increased propensity to fracture. The narrow‐sense heritability of BMD has been estimated to be ∼85%, and GWAS have successfully identified numerous loci associated with BMD which in total explain ∼5% of the genetic variance for this trait16. However, these studies have been mainly unable to assess the role of low frequency (MAF 1–5%) and rare (MAF ≤ 1%) genetic variation, as these methods rely on testing common variants (MAF ≥ 5%). A recent sequencing‐based study identified a rare nonsense variant associated with BMD using 4,931 Icelandic subjects with low BMD and 69,034 population‐based controls9. This coding variant, which disrupts the function of LGR4, appears to be confined to the Icelandic population.
To investigate the role of rare and low‐frequency genetic variation on BMD in the general population of European descent, we first undertook whole‐genome sequencing in 2,882 subjects from two cohorts in the UK10K project and whole‐exome sequencing in 3,549 subjects from five cohorts (Supplementary Table 1) with BMD phenotypes. We then used a novel imputation reference panel generated by the UK10K and 1000 Genomes consortia to impute variants that were missing, or poorly captured, from previous GWAS studies in 26,534 subjects (Supplementary Table 1 and Extended Data Fig. 1a). The combined UK10K and 1000 Genomes reference panel, which contained 3,781 and 379 European individuals with whole‐genome sequences from UK10K and 1000 Genomes projects, respectively, enabled improved imputation, particularly of low‐frequency variants, when compared to the 1000 Genomes reference panel alone17. We then undertook de novo replication genotyping of lead variants in 13 cohorts for BMD, comprising 20,271 individuals of European descent.
We meta‐analyzed association results from all discovery cohorts (ntotal = 32,965, Supplementary Table 1) for BMD measured at the forearm, femoral neck and lumbar spine, the sites where osteoporotic fractures are most prevalent. We tested bi‐allelic single nucleotide variants (SNVs) with MAF ≥ 0.5% for association, declaring genome‐wide statistical significance at P ≤ 1.2 × 10−8 (accounting for all independent SNVs above this MAF threshold; Supplementary Methods)18. The sequence kernel association test (SKAT) was used to assess association of regions containing SNVs with MAF ≤ 5% and ≤1% (Supplementary Methods). All summary‐level meta‐analytic results are available for unrestricted download (http://www.gefos.org). Novel genome‐wide significant loci were then tested for their relationship with fracture in up to 508,253 individuals. Finally, functional genomics as well as cellular and animal models were used to investigate the relevance of these novel genetic associations to bone physiology.
Through meta‐analysis of sequenced and imputed single‐SNV association tests from the discovery cohorts (Supplementary Table 1), we identified a novel locus at 2q14.2 harbouring variants associated with lumbar spine BMD (lead low‐frequency SNV rs11692564(T), MAF = 1.7%, effect size = +0.24 s.d., P = 4 × 10−9, Fig. 1 and Table 1). The direction of effect was consistent across all discovery cohorts (Extended Data Fig. 2) and the mean imputation information score for the imputed cohorts was 0.71 (Supplementary Table 2). This variant is located 53 kilobase pairs (kb) downstream from engrailed homeobox‐1 (EN1), which, to our knowledge, has not previously been associated with any osteoporosis‐related traits in humans. The rs11692564 variant was not present on HapMap imputation panels, nor on genotyping chips, underlining the importance of developing more comprehensive imputation reference panels.
a, A topological domain includes associated variants and EN1, and chromatin interaction analysis with paired‐end tag sequencing (ChIA‐PET for CTCF in MCF‐7 cell line) suggests a smaller interacting region containing EN1, and three genome‐wide significant variants for lumbar spine BMD (in red). hES cell, human embryonic stem cell. b, Association signals at the EN1 locus (green line at P = 1.2 × 10−8) for lumbar spine BMD. Red circles and triangles represent results from discovery and combined discovery and replication using fixed‐effects meta‐analysis (see Supplementary Information), respectively. c, Allele frequency versus absolute effect size for lumbar spine BMD for previously identified variants (blue)8 and the three EN1 novel variants (red). The red line denotes the mean of previously reported effect sizes.
To validate whole‐genome sequencing genotypes at rs11692564, we genotyped 1,853 whole‐genome sequenced subjects, and found all genotypes to be perfectly concordant (Supplementary Table 3). We validated imputation of rs11692564 in 3,601 imputed subjects through direct genotyping and observed that the association strengthened, and its statistical significance improved, as compared to imputed results (lumbar spine: imputed effect size = 0.22 s.d.; P = 0.05, genotyped effect size = 0.31 s.d.; P = 0.004) (Supplementary Table 4). We next sought additional evidence for the association at rs11692564 by performing additional de novo genotyping in 16,233 independent individuals and found a similarly large effect size in this population (effect size = +0.20 s.d.; P = 3 × 10−6). Meta‐analysis of the discovery and replication cohorts provided strong evidence for association (Pcombined‐meta = 2 × 10−14) (Table 1).
We also identified an additional association signal, arising from rs55983207 (MAF = 4%), 17 kb downstream of rs11692564 (r2 = 0.001) to be associated with femoral neck BMD from the combined meta‐analysis (Pmeta = 7.2 × 10−15, Table 1). A haplotype containing both effect alleles was not observed from within the UK10K whole‐genome sequenced cohort (total number of haplotypes = 7,562).
In addition to rs11692564, we also observed two additional novel genome‐wide significant variants for lumbar spine BMD near EN1, rs6542457 (MAF = 5.8%) and rs188303909 (MAF = 1.6%), which are 391 kb downstream and 67 kb upstream from rs11692564, respectively (Fig. 1b and Table 1). Variant rs188303909 was in moderate linkage disequilibrium (LD) with rs11692564 (r2 = 0.47), and conditional analysis demonstrated that these two association signals were not independent (Supplementary Table 5). However, rs6542457 was in low LD with rs11692564 (r2 = 0.002), and remained independent in conditional analyses (Supplementary Table 5). Overall, the EN1 locus harbours multiple non‐coding variants associated with lumbar spine and a single variant associated with femoral neck BMD. All three genome‐wide significant variants for lumbar spine BMD (Table 1) co‐localize solely with EN1 in a sub‐region of high interaction frequency within a single topologically associated domain19 (Fig. 1a).
The mean effect size of previously reported genome‐wide significant single nucleotide polymorphisms (SNPs) (MAF ≥ 5%) from the largest GWAS meta‐analysis to date for lumbar spine and femoral BMD was 0.048 s.d. and the largest effect size was 0.1 s.d.8. Hence, the observed effect size at rs11692564 is fourfold larger than this mean and twice that of the largest previously reported effect (Fig. 1c)8. For all genome‐wide significant variants, we observed larger effect sizes across decreasing MAF bins (Fig. 2a).
a, Box plots of the effect sizes of genome‐wide significant SNVs (P < 1.2 × 10−8), pruned for LD (r2 < 0.2) by MAF bin for discovery cohorts. Grey bars represent the values of β not observed and for which we lack statistical power to observe (at α ≤ 1.2 × 10−8 and power ≥ 0.8). P values per phenotype are from the non‐parametric trend test across MAF bins (see Supplementary Information). b, Proportion of single nucleotide variants (SNVs) passing a false discovery rate (FDR) q‐value of 0.05 across different annotation features in discovery cohorts (green) versus matched control variants (red). The three panels on the right‐hand side show enrichment across a range of evolutionary constraint scores (GERP++ score), in which green denotes SNVs above the threshold and red denotes variants below the threshold. Bars represent standard error (for Methods refer to the Supplementary Information). FA, forearm; FN, femoral neck, LS, lumbar spine.
An increase in BMD is associated with a decrease in risk of bone fracture. We therefore tested the association of rs11692564(T) (the low‐frequency allele at EN1 associated with the largest increase in BMD) in 18 cohorts comprising 508,253 individuals (98,742 cases and 409,511 controls, Supplementary Table 6). rs11692564(T) was strongly associated with a decreased risk of fracture (odds ratio (OR) = 0.85 (95% confidence interval (CI): 0.80–0.89); P = 2.0 × 10−11; I2 = 0.00) (Table 2 and Supplementary Table 7). Table 2 also shows clear associations between other variants near EN1 and risk of fracture. The fracture association at rs11692564 was 2.9‐fold larger than the mean of fracture associations detected in the largest GWAS to date, and 2.0‐fold larger than the largest previously identified fracture association8.
EN1 encodes a homeobox gene central to mouse limb development20, which has been shown to be involved in Wnt signalling interaction with Dkk1 (ref. 21). Studies of calvarial bone development and fracture healing of long bones in mice have shown that perinatal En1−/− mutants display osteopenia and enhanced skull bone resorption22, whereas in normal adult mice En1 is upregulated in the bone callus post‐fracture22. Investigating the functional role of EN1, we detected En1 expression during osteoblastogenesis in developing and mature cultured murine calvarial osteoblasts, but not in marrow‐derived osteoclasts, or in human primary osteoclast cultures (Fig. 3a and Extended Data Fig. 3). To determine where En1 is active in adult bones, we analysed vertebrae from En1lacZ/+ knock‐in mice23 and detected LacZ expression in proliferative and hypertrophic chondrocytes, osteogenic cells in the periosteum and trabecular bone surface, and in osteocytes of cortical and trabecular bone (Fig. 3b and Extended Data Fig. 4).
a, Left, quantitative expression of En1 and its temporal pattern (RNA‐seq) in cultured calvarial murine osteoblasts (n = 3 per time point). Right, confirmation of the expression of En1 in a separate RT–PCR experiment of cultured calvarial murine osteoblasts and lack of expression in osteoclasts matured from bone-marrow-derived precursor cells (positive controls for osteoblasts (osteocalcin) and osteoclast (RANK) are also shown). TPM, transcripts per million. b, Representative sections from lumbar vertebra 2 show the growth plate and bone marrow (GP and BM, left), cortical bone (CB, middle), and trabecular bone (TB, right) at ×40 magnification from En1lacZ/+ adult mice (n = 2) stained for β‐gal activity (LacZ blue, En1+ cells) and alkaline phosphatase (AP, red late chondrocytes and actively calcifying tissues). In the periosteum (PO), all the LacZ+ cells were AP+; some AP− BM cells expressed LacZ. Some AP− proliferative chondrocytes in the GP expressed LacZ+, whereas most AP+ hypertrophic chondrocytes expressed LacZ. Some AP− osteocytes (Ocy) in CB and TB were LacZ+. c, Left, histomorphometry images of lumbar vertebrae 5 show decreased trabecular bone volume and increased bone surface area occupied by osteoclast cells when comparing En1cre/flox (self‐deleted En1, sdEn1) mutants and En1flox/+ control mice. Right, reconstructed micro‐CT images show the mineral density in a control and an sdEn1 animal. d, Micro‐CT and histomorphometry measures within sdEn1 (n = 5) and controls (En1lox/+, n = 6). By micro‐CT, sdEn1 mutants exhibit decreased L5 trabecular number (Tb.N) and thickness (Tb.Th), as well as deceased bone volume fraction (BV/TV). Using histomorphometry, sdEn1 mutants exhibit increased osteoclastic area (TRAP/BS). BS, bone surface; TRAP, tartrate acid staining. Average for each measure denoted by the solid horizontal line. For each group, P value between control and sdEn1 is noted below label and was computed using paired t‐test.
Using En1cre/+; R26lox‐STOP‐lox–EYFP reporter mice to genetically tag cells for which the En1 promoter was active at any point within a cell lineage, we confirmed that En1 expression was only observed in osteogenic lineages (Extended Data Fig. 4). As most En1−/− animals die soon after birth, we generated En1cre/flox self‐deleted En1 (sdEn1) conditional mutants24 (n = 5) and demonstrated by X‐ray micro‐computed tomography (micro‐CT) that mutants have lower trabecular bone volume fraction (BV/TV), trabecular number, and trabecular thickness in both the lumbar L5 vertebrae (Fig. 3c, d and Extended Data Fig. 5) and the femur (Extended Data Fig. 5) as compared to littermate controls (n = 6). A decrease in femoral cortical thickness was also observed (Extended Data Fig. 5). By histomorphometry (Fig. 3c), we observed that the sdEn1 mice had a statistically higher proportion of osteogenic and osteoclastic cells compared to littermate controls (Fig. 3d and Supplementary Table 8). The driving force for the low bone mass would appear to be an increase in osteoclastic activity induced by En1 null osteogenic cells. This in turn initiates the expected coupled increase in mineralizing bone formation (Fig. 3b, d) mediated by an increased number of osteogenic cells and thus conforms to a high turnover osteoporosis‐like phenotype, although dynamic histomorphometry and evidence from bone turn‐over markers would be required to confirm an increased rate of bone formation (Extended Data Fig. 4). Genetic evidence from homologous regions in mice also supported a role for En1 in bone, as the homologous region contained a quantitative trait loci (QTL) peak for femur BMD (Supplementary Table 9)25. These findings, together with an earlier study focusing on En1 function in calvarial bone development22 implicate this gene as an important mediator in skeletal biology.
Together, these findings suggest that EN1 plays an important role in bone physiology and that low‐frequency non‐coding variants mapping near EN1 have large effects on BMD and risk of fracture in the general European population.
We also identified a novel SNV at 7q31.31 within the intron of CPED1 (rs148771817(T), MAF = 1.2%, effect size = +0.47 s.d., Pdiscovery = 9.31 × 10−9) associated with forearm BMD (Table 1, Supplementary Table 10 and Extended Data Fig. 6). We replicated the association at rs148771817 in 2,539 independent individuals and found a similar effect size (effect size = +0.41 s.d., P = 6 × 10−4), and combined meta‐analysis of the discovery and replication cohorts for further improved statistical evidence for association (+0.46 s.d., P = 1 × 10−11) (Table 1). This variant had an effect size 2.2‐fold larger than the mean of previously reported effects for common variants associated with forearm BMD (Extended Data Fig. 6)26.
We previously identified rs7776725 to be associated with BMD at WNT16, a gene neighbouring CPED1, (Extended Data Fig. 6) and demonstrated that knockout of Wnt16 in mice confers a 50% decrease in bone strength (P = 7 × 10−13)26,27. We have recently shown that osteoblast‐derived Wnt16 represses osteoclastogenesis28. As a result, we undertook conditional analysis of rs148771817 upon rs7776725. The rs148771817 variant remained associated after conditioning, albeit with lower statistical significance (effect size = 0.35 s.d.; Pmeta = 1 × 10−7; Extended Data Fig. 6d). Similarly, conditional analysis of the common variant upon rs148771817 revealed little change in the effect size or the statistical significance (Supplementary Table 5). Although we acknowledge that both variants may be causal, our data does not permit us to distinguish if one or both of these variants have distinct biologic effects.
While rs148771817 is intronic in CPED1, we found that DNA accessibility at this region, as measured by DNase I hypersensitivity data from ENCODE studies, was moderately correlated with DNA accessibility at the WNT16 promoter in 305 cell types (maximum r2 = 0.4, P = 2.2 × 10−15, Supplementary Table 11), whereas correlation to the promoter of CPED1 was lower (maximum r2 = 0.1, P = 0.06). Moreover, analysis of chromosome conformation capture Hi‐C interaction frequencies from human H1 embryonic stem cells shows elevated interaction frequency between rs148771817 and WNT16 (Extended Data Fig. 6), though we also observed stronger interactions between these loci and their immediate neighbouring regions.
We assessed whether association signals were enriched for deleterious coding SNVs or SNVs with increased evolutionary constraint (see Supplementary Methods). These two groups of SNVs were matched to control SNVs by MAF and distance to gene (Supplementary Methods and Supplementary Table 12), followed by LD pruning (r2 < 0.2). We observed enrichment of association signal across the spectrum of positive evolutionary constraint thresholds, which was comparable to deleterious coding variants (Fig. 2b).
In total, we have identified multiple variants associated with BMD, including 3 genome‐wide significant loci for forearm BMD, 14 for femoral neck and 19 for lumbar spine (Supplementary Tables 10, 13, 14, 15, and Extended Data Figs 7 and 8). A common variant not on previous HapMap imputation panels, near the SOX6 gene was also identified (rs11024028, MAF = 20%) (Table 1), and was found to be an independent signal from a previously reported signal at this locus (rs7108738, r2 = 0.002)8. Consistent with recent experiments29,30, region‐based collapsing methods did not identify any convincing novel associations that were not already identified as genome‐wide significant through single SNV associations. This included collapsing variants below 1% and 5% MAF thresholds, including all variants, only variants with increased GERP++ scores or those from protein‐coding regions (Supplementary Table 16 and Extended Data Figs 9 and 10).
We have identified low‐frequency, non‐coding genetic variants of large effect that are present in the general population and associate with BMD and fracture. These variants have effect sizes up to fourfold larger than the mean effect described for common variants associated with BMD and approximately threefold larger than those for fracture. Our study illustrates that larger reference panels, covering relevant ethnicities, will facilitate the discovery of low frequency and rare variants. This was enabled here by a large imputation reference panel (UK10K and 1000 Genomes) which offered tenfold more European samples than the 1000 Genomes reference panel available at the time of analysis (phase I version 3). Although we did not identify coding low‐frequency or rare variants associated with BMD at a genome‐wide significant level, we did observe that deleterious coding variants were enriched for association as a group. This suggests the existence of as yet undiscovered coding variants influencing BMD. Importantly, we have also generated new functional evidence for a central role of the homeobox protein engrailed-1 gene in regulation of BMD and identified EN1 as a critical protein in bone biology. Our findings demonstrate the utility of whole‐genome sequencing‐based discovery and deep imputation to enable the identification of novel genetic associations. These discoveries provide an improved understanding of the pathophysiology of osteoporosis and suggest that more comprehensive sets of whole‐genome sequenced individuals, covering relevant ethnicities, will enable accurate imputation and thus facilitate discovery of low frequency and rare variants influencing complex traits and common disease.
Methods
More details for the Methods are in the Supplementary Information. All human studies were approved by their institutional ethics review committees, and all participants provided written informed consent.
Data reporting
No statistical methods were used to predetermine sample size. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment, except the teams undertaking micro‐CT and histomorphometry experiments were blinded to each other’s results.
Whole‐genome sequencing
ALSPAC and TwinsUK cohorts were sequenced at an average read depth of 6.7× through the UK10K program (http://www.UK10K.org) using the Illumina HiSeq platform, and aligned to the GRCh37 human reference using BWA31. SNV calls were completed using samtools/bcftools and VQSR and GATK were used to recall these calls.
Whole‐exome sequencing
The AOGC, FHS, RS‐I, ESP and ERF cohorts were whole‐exome sequenced as described in the Supplementary Information.
Whole‐genome genotyping
All remaining discovery cohorts were genome‐wide genotyped and imputed to the UK10K/1000 Genomes reference panel, as described in the Supplementary Information.
Association testing for BMD
Single variants with a MAF > 0.5% were tested for an additive effect on lumbar spine, femoral neck and forearm BMD, adjusting for sex, age, age2, weight and standardized to have a mean of zero and a standard deviation of one. Meta‐analysis of cohort‐level summary statistics was undertaken using GWAMA32. Conditional analyses for significant SNVs was performed using GCTA33. Region‐based collapsing tests were performed using skatMeta34, an implementation of the SKAT method35 that enables the meta‐analysis of multiple cohorts. For each cohort, variants with MAF ≤ 5% or ≤1% were collected and meta‐analysis using skatMeta was conducted for windows of 30 SNVs within each region, overlapping by 10 SNVs.
Replication genotyping
Lead SNVs were selected for replication genotyping, which was performed at LGC Genomics, Erasmus MC and deCODE Genetics using KASP genotyping. Association testing for replication genotyping was undertaken using the same additive model, using the same covariates for BMD, as above.
Fracture association testing
Fractures were defined as those occurring at any site, except fingers, toes and skull, after age 18. Both incident and prevalent fractures were included and were verified by either radiographic, casting, physician, or subject reporting. Fractures resulting from any type of trauma were considered. Covariates included in the additive model were age, age2, sex, height, weight, oestogen/menopause status (when available), ancestral genetic background and cohort‐specific covariates (such as clinical centre). Association testing was done in two phases. The first involved all 1,482 genome‐wide significant SNVs for BMD. In the second phase of fracture association testing, variants at EN1 were assessed in 18 cohorts, comprising 98,467 cases and 409,736 controls. Meta‐analysis of cohort‐level summary statistics was performed using GWAMA32.
Functional genomics
We tested whether variants with increasing GERP++ scores36 were more strongly associated with BMD than SNVs matched for distance to gene and MAF, after LD pruning using PLINK37 at an r2 of <0.2, using windows of 100 kb and a step of 20 kb. Coding variants were partitioned as deleterious using Variant Effect Predictor38 LD pruned (r2 < 0.2). The proportion of variants passing an FDR q‐value of ≤0.05 were reported.
En1 murine expression experiments
Pre‐osteoblast‐like cell were differentiated to osteoblasts from calvaria of C57BL/6J mice and expression levels of each gene was quantified using RNA‐seq. The temporal expression of En1 in cell culture experiments of these osteoblasts and bone-marrow-derived osteoclasts (isolated from long bones of six‐week‐old mice) was measured by PCR, with Bglap (osteocalcin) and Tnfrsf11a (RANK), serving as controls. Total mRNA for En1 in osteoblasts was quantified using real‐time PCR.
Micro‐CT and histomorphometry
Mouse husbandry and all experiments were performed in accordance with Memorial Sloan‐Kettering Cancer Center Institutional Animal Care and Use Committee‐approved protocols. Bone characteristics of self‐deleted conditional En1(sdEn1) mutants were compared to En1+/flox littermates using micro‐CT. The same animals were assessed for histomorphometry (and laboratories performing micro‐CT and histomorphometry were blinded to each other’s results). After tissue sectioning, samples were stained for calcification (calcein blue), tartrate acid (TRAP) to assess for osteoclasts and alkaline phosphatase to assess for osteoblasts.
Murine histology
Two‐month‐old En1lacZ/+ mice39 were sectioned at bone sites and stained for X‐gal and/or alkaline phosphatase and imaged at ×400.
References
Richards, J. B. et al. Bone mineral density, osteoporosis, and osteoporotic fractures: a genome‐wide association study. Lancet 371, 1505–1512 (2008)
Styrkarsdottir, U. et al. Multiple genetic loci for bone mineral density and fractures. N. Engl. J. Med. 358, 2355–2365 (2008)
Styrkarsdottir, U. et al. New sequence variants associated with bone mineral density. Nature Genet. 41, 15–17 (2009)
Rivadeneira, F. et al. Twenty bone‐mineral‐density loci identified by large‐scale meta‐analysis of genome‐wide association studies. Nature Genet. 41, 1199–1206 (2009)
Duncan, E. L. et al. Genome‐wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet. 7, e1001372 (2011)
Koller, D. L. et al. Genome‐wide association study of bone mineral density in premenopausal European‐American women and replication in African‐American women. J. Clin. Endocrinol. Metab. 95, 1802–1809 (2010)
Xiong, D.‐H. et al. Genome‐wide association and follow‐up replication studies identified ADAMTS18 and TGFBR3 as bone mass candidate genes in different ethnic groups. Am. J. Hum. Genet. 84, 388–398 (2009)
Estrada, K. et al. Genome‐wide meta‐analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nature Genet. 44, 491–501 (2012)
Styrkarsdottir, U. et al. Nonsense mutation in the LGR4 gene is associated with several human diseases and other traits. Nature 497, 517–520 (2013)
The UK10K Consortium The UK10K project identifies rare variants in health and disease. Nature http://dx.doi.org/10.1038/nature14962 (this issue)
Hindorff, L. A. et al. Potential etiologic and functional implications of genome‐wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009)
Kiezun, A. et al. Exome sequencing and the genetic basis of complex traits. Nature Genet. 44, 623–630 (2012)
Gudbjartsson, D. F. et al. Large‐scale whole‐genome sequencing of the Icelandic population. Nature Genet. 47, 435–452 (2015)
Sulem, P. et al. Identification of a large set of rare complete human knockouts. Nature Genet. 47, 448–444 (2015)
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Richards, J. B., Zheng, H.‐F. & Spector, T. D. Genetics of osteoporosis from genome‐wide association studies: advances and challenges. Nature Rev. Genet. 13, 576–588 (2012)
Huang, J. et al. Improved imputation of low‐frequency and rare variants using the UK10K haplotype reference panel. Nature Comm. 6, 8111 (2015)
Xu, C., Tachmazidou, I., Walter, K., Ciampi, A., Zeggini, E. & Greenwood, C. M. T. Estimating genome‐wide significance for whole‐genome sequencing studies. Genet. Epidemiol. 38, 281–290 (2014)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Loomis, C. A. et al. The mouse Engrailed‐1 gene and ventral limb patterning. Nature 382, 360–363 (1996)
Adamska, M., MacDonald, B. T., Sarmast, Z. H., Oliver, E. R. & Meisler, M. H. En1 and Wnt7a interact with Dkk1 during limb development in the mouse. Dev. Biol. 272, 134–144 (2004)
Deckelbaum, R. A., Majithia, A., Booker, T., Henderson, J. E. & Loomis, C. A. The homeoprotein engrailed 1 has pleiotropic functions in calvarial intramembranous bone formation and remodeling. Development 133, 63–74 (2006)
Matise, M. P. & Joyner, A. L. Expression patterns of developmental control genes in normal and Engrailed‐1 mutant mouse spinal cord reveal early diversity in developing interneurons. J. Neurosci. 17, 7805–7816 (1997)
Sgaier, S. K. et al. Genetic subdivision of the tectum and cerebellum into functionally related regions based on differential sensitivity to engrailed proteins. Development 134, 2325–2335 (2007)
Ackert‐Bicknell, C. L. et al. Mouse BMD quantitative trait loci show improved concordance with human genome‐wide association loci when recalculated on a new, common mouse genetic map. J. Bone Miner. Res. 25, 1808–1820 (2010)
Zheng, H.‐F. et al. WNT16 influences bone mineral density, cortical bone thickness, bone strength, and osteoporotic fracture risk. PLoS Genet. 8, e1002745 (2012)
Medina‐Gomez, C. et al. Meta‐analysis of genome‐wide scans for total body BMD in children and adults reveals allelic heterogeneity and age‐specific effects at the WNT16 locus. PLoS Genet. 8, e1002718 (2012)
Movérare‐Skrtic, S. et al. Osteoblast‐derived WNT16 represses osteoclastogenesis and prevents cortical bone fragility fractures. Nature Med. 20, 1279–1288 (2014)
Ladouceur, M., Dastani, Z., Aulchenko, Y. S., Greenwood, C. M. T. & Richards, J. B. The empirical power of rare variant association methods: results from Sanger sequencing in 1,998 individuals. PLoS Genet. 8, e1002496 (2012)
Tang, H. et al. A large‐scale screen for coding variants predisposing to psoriasis. Nature Genet. 46, 45–50 (2014)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Mägi, R. & Morris, A. P. GWAMA: software for genome‐wide association meta‐analysis. BMC Bioinformatics 11, 288 (2010)
Yang, J. et al. Conditional and joint multiple‐SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nature Genet. 44, 369–375 (2012)
Voorman, A. A., Brody, J. & Lumley, T. SkatMeta: an R package for meta‐analyzing region‐based tests of rare DNA variants. (http://cran.r-project.org/web/packages/skatMeta) (2013)
Wu, M. C. et al. Rare‐variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011)
Davydov, E. V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLOS Comput. Biol. 6, e1001025 (2010)
Chang, C. C. et al. Second‐generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015)
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP effect predictor. Bioinformatics 26, 2069–2070 (2010)
Hanks, M., Wurst, W., Anson‐Cartwright, L., Auerbach, A. B. & Joyner, A. L. Rescue of the En‐1 mutant phenotype by replacement of En‐1 with En‐2. Science 269, 679–682 (1995)
Acknowledgements
Full acknowledgements are listed in the Supplementary Information.
Author information
Authors and Affiliations
Consortia
Contributions
Principal Investigators: A.H., A.J., A.U., A.X.‐A., B.L., C.A.‐B., Ch.C., C.L., Cl.C., C.L.D., C.M.v.D., C.O., D.S.E., D.Ga., D.Go., D.Gr., D.H., D.Ki., D.M., E.D., E.O., F.Ri., F.Ro., G.D.S., J.B.R., J.D., J.Re., J.Ri., J.‐T.K., J.Tu., K.A., L.A.C., L.L., L.P.G.M.d.G., M.B., M.M.F., N.S., N.T., N.v.d.V., N.vS., P.R., R.D., R.D.J., R.L.P., S.G.W., S.H.R., T.H., T.P., T.S., U.P.‐K.,V.G., X.N.,Y.‐H.H. Genotyping: AOGC Consortium, A.U., B.M., B.W., C.L., C.M.v.D., C.N., C.O., C.W., D.C., D.Gr., E.D., E.O., F.Ri., G.G., G.Tr., J.Er., J.Jv.M., J.Re., J.Ri., J.‐T.K., J.v.R., M.B., M.C.F., M.J., M.Z., N.A., N.G.‐G., N.S., N.T., P.Ar., P.D., P.R., R.K., S.H.R., S.M., S.R., U.P.‐K., X.N. and Y.‐H.H. Phenotyping: A.E., A.H., A.L., AOGC Consortium, A.P.H., A.U., A.X.‐A., B.M., C.G., C.K., C.L., C.L.D., C.M.v.D., C.O., D.Go., D.Ka., D.Ki., D.M., E.D., E.N., E.O., F.E.M., F.K., F.Ri., F.Ro., G.H., J.B., J.C., J.Ei., J.O., J.Re., J.Ri., J.‐T.K., J.To., K.E., K.S., K.T., L.O., L.R., L.V., M.B., M.C.F., M.M.F., M.C.Z., M.K., M.Z., N.A., N.S., N.T., O.L., O.S., R.L.P., S.G.W., S.G., S.H.R., S.K., T.N., T.S. and U.P.‐K. Functional experiments: A.J., A.R.‐D., B.Ge, C.A.‐B., C.H., C.L., C.L.D., C.O., C.U., D.Ga., D.P., E.G., H.Y.P.‐M., J.D., J.F., K.Ch., Ma.M., M.H., N.S., O.S., S.B., S.C., S.‐H.C., St.W., T.K., T.P., U.P.‐K., W.C. and X.J. Data analysis: A.E., A.K., A.S., A.V.S., B.M., C.A.‐B., Ch.C., C.‐H.C., C.K., C.L., C.L.D., C.M.‐G., C.M.T.G., C.O., C.T.L., C.W., D.S.E., D.M.E., D.C., D.Ka., D.M., D.P., E.D., E.G., E.N., F.G., F.Ri., G.H., G.Th., H.‐F.Z., J.B.R., J.D., J.Er., J.F., J.Ha., J.Hu., J.K., J.v.R., K.Ch., K.E., K.W., L.A.C., L.H., L.M., L.O., L.R., L.V., M.B., M.C., M.H., M.K., N.A., N.S., N.T., O.L., P.Au., P.D., P.L., R.L., S.B., S.C., S.G.W., S.K., U.P., U.P.‐K., V.F., W.‐C.C., Y.‐H.H., Y.M. and Y.Z. Meta‐analysis: H.‐F.Z., V.F. and Y.‐H.H. Lead analysts: H.‐F.Z. and V.F. Wrote first draft: J.B.R.
Corresponding author
Ethics declarations
Competing interests
Authors from deCODE Genetics are employees of deCODE Genetics. Authors from 23andMe are employees of 23andMe. Remaining authors declare no competing financial interests.
Additional information
Source code used in preparation of results is available at https://github.com/richardslab/gefos.seq/. BMD discovery meta‐analysis results are available from http://www.gefos.org. Information pertaining to UK10K can be obtained from http://www.uk10k.org.
Lists of participants and their affiliations appear in the Supplementary Information.
Lists of participants and their affiliations appear in the Supplementary Information.
Extended data figures and tables
Extended Data Figure 1 Discovery single variant meta‐analysis.
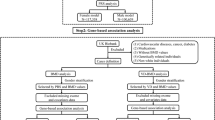
a, Overall study design. b, From top to bottom, quantile–quantile plots for the sex‐combined single SNV meta‐analysis, sex‐stratified single SNV meta‐analysis (forearm phenotype consists solely of female‐only cohorts), and sex‐combined single SNV conditional meta‐analysis Plots depicts P values prior (blue) and after (red) conditional analysis on genome-wide significant variants (see Supplementary Methods). c, From top to bottom, Manhattan plots for sex‐combined meta‐analysis for lumbar spine BMD, femoral neck BMD, and forearm BMD. Each plot depicts variants from the UK10K/1000G reference panel with MAF > 0.5% across the 22 autosomes (odd, grey; even, black) against the −log10 P value from the meta‐analysis of 7 cohorts (dots). Also depicted are the subset variants from the reference panel that are also present in ref. 8 with P value <5 × 10−6 (diamonds). Variants with MAF < 5% and P < 1.2 × 10−6 are also depicted (red). d, Quantile–quantile plots for the sex‐combined meta‐analysis of lumbar spine, femoral neck, and forearm BMD for SNVs present across both exome‐sequenced and genome-sequenced and imputed cohorts, that is, SNV present only in genome-sequenced or imputed cohorts are not shown. e, Manhattan plot for the meta‐analysis of sex‐combined results for lumbar spine BMD for SNVs present in exome‐sequenced and genome-sequenced and imputed cohorts, that is, SNV present only in genome-sequenced or imputed cohorts are not shown (from left to right: lumbar spine, forearm and femoral neck BMD).
Extended Data Figure 2 Forest plots by cohort for genome‐wide significant loci from discovery meta‐analysis.
Forest plots for three BMD phenotypes are shown. Title of each plot includes gene overlapping the SNV and its genomic position on build hg19. P values are from fixed‐effect meta‐analysis (see Supplementary Information).
Extended Data Figure 3 Gene expression in human and mouse.
a, Quantification of Dock8 expression and its temporal pattern through RNA‐seq in cultured calvarial murine osteoblasts across day 2 through to day 18 of osteoblast development. Shown for comparison is Bglap, which encodes osteocalcin, a critical protein in osteoblasts. b, Quantification of expression of genome‐wide significant genes and their temporal pattern through RNA‐seq in cultured calvarial murine osteoblasts across day 2 through to day 18 of osteoblast development. c, Expression of EN1 mRNA in human cells presented as per cent of GAPDH mRNA. d, Expression of En1 in control and sdEn1 mice in purified osteoblast culture. For osteoblast marker gene expression, total mRNAs were purified from osteoblast cultures at day 10 and measured using quantitative real‐time PCR. mRNA levels were normalized relative to GAPDH mRNA. e. Real‐time PCR expression of control and sdEn1 as compared to 18S mRNA in whole vertebral bone extract. All data are shown as mean ± s.e.m. Significance computed by Student’s unpaired t‐test.
Extended Data Figure 4 Histological assessment of En1cre‐expressing cells in skeletal cells of the vertebra.
a, Lineage history of En1cre‐expressing cells in skeletal cells of the vertebra. The En1cre allele was combined with the R26LSL‐YFP reporter allele and examined using frozen fluorescent immunohistochemistry and alkaline phosphatase (AP) staining. Cell nuclei were detected with DAPI. YFP‐expressing cells have expressed Cre (En1) at some time in their history. In subpanel A, control animals lacking the R26LSL‐YFP reporter show low background YFP signal (green). In subpanel B, En1cre/+; R26LSL‐YFP/+ mice YFP‐expressing cells are detected in the growth plate chondrocytes of the vertebra (asterisk), trabecular bone lining cells (arrow) and osteocytes (arrowhead). Note, high fluorescent background staining in the marrow space. In subpanel C, the same section is shown stained for AP activity using the Fast Red substrate. Strong activity is present in the hypertrophic chondrocytes of the growth plate and trabecular bone lining cells (arrow). In subpanel D, alignment of the AP and YFP images shows that the trabecular lining cells co‐express AP and YFP. b, Co‐localization of En1 and alkaline phosphatase expression. Images of lumbar vertebrae sections (growth plate and trabecular bone regions, ×40 magnification) from two‐month old En1lacZ/+ mice (see Fig. 3b), stained for LacZ and alkaline phosphatase (AP), false‐coloured as indicated. Double‐positive cells are indicated by arrows, single‐positive cells are indicated by arrowheads (LacZ+) or asterisks (AP+). Except for some chondrocytes, most AP+ cells are also LacZ+, that is, express En1. The bone marrow was digitally removed, as it contains no AP+ cells.
Extended Data Figure 5 Micro‐CT results for control (En1flox/+) and self‐deleting En1 knockout (sdEn1, En1cre/flox) animals.
a, Trabecular bone micro‐CT images from lumbar vertebra 5. b, Morphological characteristics at lumbar vertebra 4, 5, and 6 (from bottom to top). c, d, Morphological characteristics of left femur trabecular bone (c) and left femur cortical bone (d). e, Micro‐CT parameter results for the comparison of control and sdEn1 animals at lumbar vertebra 5, femur trabecula, and femur cortical bone. Horizontal lines denote mean of observations. Significance between control and sdEn1 is calculated using an unpaired t‐test.
Extended Data Figure 6 Novel association from 7q31.3.
a, Chromatin interaction data from Hi‐C performed in H1 embryonic stem cells23 of a 2 Mb region encompassing rs148771817 (red and identified by arrow) and WNT16. b, The left axis denotes the association P value (red and green lines at P = 1.2 × 10−5 and 1.2 × 10−8, respectively). The novel genome‐wide significant SNV, rs148771817, within an intron of CPED1, and the lead genome‐wide significant SNV rs7776725 upstream to WNT16 (within FAM3C) are in low LD with each other. c, Allele frequency versus absolute effect size (in standard deviations) for forearm BMD of all previously identified genome‐wide significant variants (blue)8 and the novel variant within CPED1 (red), rs148771817 from replication meta‐analysis. The blue line denotes the mean of effect sizes for previously reported forearm BMD variants. d, Meta‐analysis summary statistics of rs148771817 conditioned on rs7776725.
Extended Data Figure 7 Regional plots of genome‐wide significant loci from single‐SNV association tests for forearm and femoral neck BMD.
Each regional plot depicts SNVs within 1 Mb of a locus’ lead SNV (x axis) and their associated meta‐analysis P value (−log10). SNVs are colour-coded according to r2 with the lead SNV (labelled, r2 calculated from UK10K whole‐genome sequencing data set). Recombination rate (blue line), and the position of genes, their exons and the direction of transcription are also displayed (below plot).
Extended Data Figure 8 Regional plots of genome‐wide significant loci from single‐SNV association tests from lumbar spine BMD.
Each regional plot depicts SNVs within 1 Mb of a locus’ lead SNV (x axis) and their associated meta‐analysis P value (−log10). SNVs are colour coded according to r2 with the lead SNV (labelled, r2 calculated from UK10K whole genome sequencing data set). Recombination rate (blue line), and the position of genes, their exons and the direction of transcription are also displayed (below plot).
Extended Data Figure 9 Region‐based association tests using skatMeta for windows of 30 SNVs and window step of 20 SNVs.
a, Left, quantile–quantile plots for forearm (FA) BMD, femoral neck (FN) BMD, and lumbar spine (LS) BMD. For each MAF range considered (<5% or < 1%), analysis was conducted across all variants, variant overlapping coding exons, and variants with GERP++ score >1. b, Right, Manhattan plots forearm BMD, femoral neck BMD, and lumbar spine BMD. For each MAF range considered (<5% or < 1%), analysis was conducted across all variants, variant overlapping coding exons, and variants with GERP++ score >1. Blue lines indicate genome‐wide suggestive (P = 1.2 × 10−6) thresholds and red lines indicate genome‐wide significant (P = 1.2 × 10−8) thresholds.
Extended Data Figure 10 Single variant analysis of signals from region‐based tests.
a, Drop‐one SNV (left) and drop‐one cohort (right) for genome‐wide significant 30 SNV windows for femoral neck and forearm BMD from skatMeta analysis. On left, for a given 30 SNV window, the −log10P of skatMeta test for 29 SNVs, excluding (that is, dropping) the SNV at position labelled on the x axis. On right, for given 30 SNV window on left, the −log10P of skatMeta test for all cohorts, excluding (that is, dropping) cohort labelled on x axis. b, Regional view of CPED1/WNT16 locus for forearm BMD. Significant SNVs from single variant meta‐analysis (rs148771817 and rs79162867, in blue) overlap significant regions found using region‐based test (red bars).
Supplementary information
Supplementary Information
This file contains Supplementary Text and Supplementary References. (PDF 1093 kb)
Supplementary Tables
This file contains Supplementary Tables 1-19. (XLSX 757 kb)
Rights and permissions
About this article
Cite this article
Zheng, H., Forgetta, V., Hsu, Y. et al. Whole‐genome sequencing identifies EN1 as a determinant of bone density and fracture. Nature 526, 112–117 (2015). https://doi.org/10.1038/nature14878
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14878
- Springer Nature Limited
This article is cited by
-
Genetic evidence of the causal relationship between chronic liver diseases and musculoskeletal disorders
Journal of Translational Medicine (2024)
-
Causal relationship between glycemic traits and bone mineral density in different age groups and skeletal sites: a Mendelian randomization analysis
Journal of Bone and Mineral Metabolism (2024)
-
Investigating the association between serum ADAM/ADAMTS levels and bone mineral density by mendelian randomization study
BMC Genomics (2023)
-
Osteoporosis and osteoarthritis: a bi-directional Mendelian randomization study
Arthritis Research & Therapy (2023)
-
Causal association of epigenetic aging and osteoporosis: a bidirectional Mendelian randomization study
BMC Medical Genomics (2023)