Abstract
Objective:
The gut microbiota associates with obesity and related disorders, but recent meta-analyses have found that this association is, at best, of small effect. We argue that such analyses are flawed by the use of body mass index (BMI) as sole proxy for disease, and explore a classification method that distinguishes the cardiometabolic health status of individuals to look for more comprehensive associations between gut microbes and health.
Design:
We analyzed a 441 community-dwelling cohort on which we obtained demographic and health information, anthropometry and blood biochemistry data that served to categorize participants according to BMI, cardiometabolic health status and body size phenotypes. In addition, the participants donated fecal samples from which we performed 16S rRNA gene sequencing to analyze the gut microbiota.
Results:
We observed that health-related variables deteriorate with increased BMI, and that there are further discrepancies within a given BMI category when distinguishing cardiometabolically healthy and unhealthy individuals. Regarding the gut microbiota, both obesity and cardiovascular disease associate with reductions in α-diversity; having lean, healthy individuals the most diverse microbiotas. Moreover, the association between the gut microbiota and health stems from particular consortia of microbes; the prevalence of consortia involving pathobionts and Lachnospiraceae are increased in obese and cardiometabolically abnormal subjects, whereas consortia including Akkermansia muciniphila and Methanobrevibacter, Oscillospira and Dialister have higher prevalence in cardiometabolically healthy and normoweight participants.
Conclusions:
The incorporation of cardiometabolic data allows a refined identification of dissimilarities in the gut microbiota; within a given BMI category, marker taxa associated with obesity and cardiometabolic disease are exacerbated in individuals with abnormal health status. Our results highlight the importance of the detailed assessment and classification of individuals that should be carried out prior to the evaluation of obesity treatments targeting the gut microbiota.
Similar content being viewed by others
Introduction
Obesity and its associated diseases, such as type 2 diabetes and cardiovascular disease, are some of the most prevalent chronic diseases worldwide, accounting for 19.3 million deaths every year.1 While obesity research has focused primarily on nutritional, behavioral and genetic factors,2, 3 an additional element, which affects both food intake and metabolism, has gained notoriety: the gut microbiota, that is, the set of microorganisms that colonize the gastrointestinal tract.4
The association between the gut microbiota and obesity has been most studied,5 although it also exists with diabetes and cardiovascular disease.6, 7 Regarding obesity, early research showed that the gut microbiota of obese mice is enriched in Firmicutes and reduced in Bacteroidetes relative to lean animals.8 However, in humans this pattern has been inconsistent: some studies show similar findings,9 others reveal no association10 or contradict mouse studies.11 With the increase of available data, several meta-analyses have revisited the association between obesity and gut microbes, including analysis of α-diversity (that is, intra-subject diversity), of the abundance of Firmicutes and Bacteroidetes, and of the ratio of these abundances. In general, they have found either significant associations with small effect or no association.12, 13, 14
There are, however, two caveats with these meta-analyses. First, they have focused on finding differences in broad taxonomic groups (for example, Firmicutes, Bacteroidetes); while discrepancies may exist at such level, their biological and clinical relevance is hard to interpret because the vast diversity of these groups. Rather, alterations in the microbiota associated with changes in the nutritional status are observed on particular microorganisms from lower taxonomic ranks (for example, genus, species). For instance, Akkermansia muciniphila15 and Christensenella16 have received attention in recent years because of their association with reduced body mass index (BMI). Moreover, the fact that classification models trained to distinguish lean and obese individuals based on the abundance of the whole microbial community tend to perform poorly,14 while the use of selected bacterial groups increases accuracy,17 further strengthens the idea that few microorganisms drive the disease-associated changes in the microbiota.
Second, these studies used BMI as their sole explanatory variable to assess obesity. BMI is an easy-to-measure, convenient and noninvasive tool widely used in epidemiological and clinical contexts.18 Nevertheless, it fails to differentiate physiological, metabolic or health status by itself, as it aggregates individuals based only on their height and weight.19 Recent studies have addressed this issue by further discriminating subjects according to their cardiometabolic health status (CHS), generating a classification that takes into account the heterogeneity in the obesity-associated disease risk. Thus, an individual is categorized as cardiometabolically healthy or abnormal according to a series of clinical variables, in addition to being classified within a given BMI category,20, 21 obtaining six body size phenotypes (BSP; Figure 1).
Schematic view of the classification into body size phenotypes. BSP are obtained by subdividing each BMI category according to the cardiometabolic health status (CHS). CHS is obtained by considering abnormalities in blood pressure, HDL cholesterol, triglycerides, fasting glucose, insulin resistance and high sensitive C-reactive protein levels.
Classification methods that result in a more precise stratification of the surveyed population are most interesting in the study of microbiome-associated conditions. In the particular cases of obesity and related diseases, they may allow to better detect the signal among the noise introduced by the sole use of BMI and to disentangle the patterns associated with metabolic health. Here, we analyze data from a community-dwelling cohort to evaluate two hypotheses. (i) The incorporation of cardiometabolic data to subdivide three categories of BMI allows a better identification of dissimilarities in the gut microbiota. (ii) Within a given BMI category, marker taxa associated with obesity and cardiometabolic disease are more abundant in cardiometabolically abnormal than healthy individuals; marker taxa are groups of microbes that can be quantified to inform about a clinical condition.
Materials and methods
Study population
Sample size was calculated following a previous study performed by the same team.22 Accordingly, 28 subjects per BMI category were necessary to observe significant differences in the gut microbiota at the phylum level between lean and obese participants. Since the study considered three BMI categories and was aimed to be replicated in five cities, the required sample size was 420 subjects. We enrolled 441 adults in July–November 2014, with BMI⩾18.5 kg m−2, living in five Colombian cities. We excluded pregnant women, individuals who consumed antibiotics 3 months prior to enrollment, and individuals diagnosed with the following diseases: neurodegenerative disease; current or recent cancer (⩽1 year); and gastrointestinal diseases (Crohn’s disease, ulcerative colitis, short bowel syndrome, diverticulosis or celiac disease). Participants were enrolled in similar proportions by BMI (lean, overweight, obese), city, sex and age range (18–40 and 41–62 years). Written informed consent was obtained from all of them before beginning the study. The Bioethics Committee of SIU—University of Antioquia (Medellin, Colombia) approved the procedures described here.
Anthropometric evaluation and blood pressure
We measured weight, height, waist circumference and four skin folds (biceps, triceps, subscapular and ileocrestal). BMI was calculated as weight (kg)/height squared (m2); participants were classified as normoweight (18.5⩽BMI<25.0 kg m−2), overweight (25.0⩽BMI<30.0 kg/m2) or obese (BMI⩾30.0 kg/m2). Skinfolds served to calculate the fat percentage.23 Systolic and diastolic blood pressures were measured with electronic tensiometers. Each measure was evaluated twice and the average of the two measures was reported.
Caloric intake, physical activity, smoking and medicament use
These variables were self-reported using specific questionnaires. In the case of the caloric intake, we carried out 24-hour dietary recall interviews to quantify the calories in the habitual diet of participants using EVINDI 4.0(ref.24) and PC-SIDE (http://www.side.stat.iastate.edu/pc-side.php). Physical activity was measured with the international physical activity questionnaire (IPAQ-S).25
Blood biochemical parameters
Total cholesterol, HDL, LDL, VLDL, triglycerides, glucose, insulin, glycated hemoglobin, adiponectin and high sensitive C reactive protein (hs-CRP) were measured in fasting peripheral venous blood using standard techniques employed by a clinical laboratory (Dinámica IPS). Blood insulin served to calculate the insulin resistance index using the homeostasis model assessment (HOMA-IR).26
Body size phenotypes
To categorize participants according to their body size phenotypes, we used the definition proposed by Wildman et al.,20 where BMI categories are subdivided by the cardiometabolic health status of individuals using the strict criteria implemented by Tomiyama et al.21 That is, individuals are considered metabolically abnormal when they have ⩾2 of the following conditions: systolic/diastolic blood pressure ⩾130/85 mm Hg or consumption of antihypertensive medication; fasting triglycerides ⩾150 mg dl−1; HDL <40 mg dl−1 (men), <50 mg dl−1 (women) or consumption of lipid-lowering medication; fasting glucose ⩾100 mg dl−1 or consumption of antidiabetic medication; HOMA-IR >3, and hs-CRP >3 mg l−1. We selected HOMA-IR and hs-CRP cutoffs that better reflect the biochemical characteristics of the studied population.27
Gut microbiota characterization
A detailed description of laboratory and bioinformatics procedures can be found elsewhere.7 Briefly, each participant collected a fecal sample from which the total microbial DNA was extracted using the QIAamp DNA Stool Mini Kit. The V4 region of the 16 S rRNA gene was amplified with primers F515 and R806, and sequenced with the Illumina MiSeq platform.28 Raw reads were deposited at the SRA-NCBI under BioProject PRJNA417579. We processed amplicons using Mothur v.1.36 following its standard operating procedure as available on November 2015. Operational taxonomic units (OTUs) delimited at 97% identity were generated and classified using Greengenes 13_8_99. After bioinformatics curation, the number of sequences per individual ranged from 3,667 to 102,700, with a median of 28,561. Analyses that required absolute OTU frequencies were performed on sequence counts rarefied to 3,667 sequences per sample, otherwise, the relative OTU abundance was calculated for each sample.
To detect changes in the composition of the gut microbiota at fine scale, we first defined co-abundance groups of microbes (CAGs), that is, OTUs that are found more frequently together, hence exhibit positive abundance correlations, and that reflect the underlying microbial structure.29 CAGs were defined by calculating the Spearman’s correlation coefficients between all OTUs that had median abundance ⩾0.01% across all samples and by applying the hierarchical clustering with Ward’s linkage. CAGs were validated by randomly splitting the OTU table and computing two separate correlation matrices; the correlation between these two matrices was tested with the Mantel test. In addition, we confirmed correlations using SparCC,30 an alternative method for computing correlations in compositional data. Note that CAGs are not expected to gather microbes by their taxonomy or phylogenetic relationships (this can occur but is not a direct consequence of the clustering). OTUs reunite within a CAG because the presence of OTU W is positively correlated with the presence of OTU Z (that is, they co-abound).
Statistical analyses
Mean and s.d. of health-related variables were obtained in the complete dataset (441 individuals). We also obtained these values in the subset of participants that did not report smoking or were under no pharmacological treatment (to the exception of over-the-counter vitamin and mineral supplements, phytotherapeutics and contraceptives) as a way to test the sensitivity of results to cigarette and medicament consumption. This subset was composed of 50% of the original participants (220 individuals); the obese group was the one where more participants were removed (61.4%) followed by overweight (49.1%) and normoweight (40.6%) (Supplementary Table S1).
Linear models were fitted in the complete dataset for the collection of variables that were not used in the BSP definition (namely, waist circumference, body fat, LDL, VLDL, insulin, and adiponectin; non-normally distributed variables were transformed with natural logarithm or arcsine square-root). In this way, we estimated the goodness of fit of models considering BMI, CHS or BSP by means of the Akaike’s information criterion (AIC). We controlled for potential confounding (age, sex, city of origin, calorie intake and physical activity) in all models.
For the gut microbiota, we computed α-diversity using the Shannon index and species richness, and tested for differences among BMI, CHS and BSP using ANOVA. In addition, linear models using BMI as a continuous variable and α-diversity were constructed to test the interaction between BMI and cardiometabolic health status. Where necessary, data were log-transformed. Next, we assessed differences in β-diversity (that is, inter-subject diversity) using the adonis function of the permutational multivariate analysis of variance (PERMANOVA) on phylogeny-based weighted and unweighted UniFrac matrices, as implemented in the GUniFrac package of R.31 Also, we tested for differences in the relative abundances of all OTUs classified as Bacteroidetes and Firmicutes, the ratio of the relative abundances of Bacteroidetes and Firmicutes (B/F ratio), and the abundance of CAGs among categories of the evaluated groupings using the Kruskal-Wallis test.
Finally, we used linear discriminant analysis (LDA) effect size (LEfSe)32 to agnostically identify microbial taxa that have a differential abundance between normoweight and obese, between cardiometabolically healthy and abnormal, and between normoweight-healthy and obese-abnormal subjects. We retained OTUs that showed strong associations in the LDA (OTUs with [log10] LDA scores ⩾3). While this may result in few retained OTUs, these are more likely to be biologically relevant. In addition, a quasipoisson generalized linear model on rarefied sequence counts33 was fitted to test the association of individual OTUs with a median abundance ⩾0.001% and log(BMI) as a continuous variable. P-values were corrected for multiple comparisons.
Code availability
The employed R code is available at https://github.com/jsescobar/bsp.
Results
Disease risk is better assessed by BSP compared to BMI or CHS
In order to capture the heterogeneity of cardiometabolic disease risk, we first classified participants according to BMI into normoweight, overweight and obese. These categories were then subdivided according to the cardiometabolic health status as either healthy or abnormal.20, 21 Importantly, AIC from linear models indicate that disease risk is better assessed by BSP than BMI or CHS (Supplementary Table S2). The biochemical and anthropometric characteristics of each resulting group are summarized in Table 1. Overall, we observed that the measured variables deteriorate with increased BMI, and that there are further discrepancies within a given BMI category when distinguishing cardiometabolically healthy and abnormal individuals. Cardiometabolic abnormalities gathered groups with older age, at least among normoweight and overweight. Noteworthy, women made most of the obese healthy individuals; in general, males tended to have more abnormalities than females (Figure 2). Self-reported food intake and physical activity are likely biased, as evinced by the decrease in the mean caloric intake and the increase in the number of metabolic equivalents with BMI. Still, cardiometabolically abnormal subjects displayed lower physical activity, higher caloric intake, and increased cigarette and medicament use compared with healthy participants. Importantly, BSP form a ladder-like pattern, on which each step is associated with a different level of disease risk. This pattern is robust and is not biased by cigarette and medicament consumption, as demonstrated by the analysis of the subset of non-smoker and non-medicated participants (Supplementary Table S3).
The gut microbiota associates with disease risk
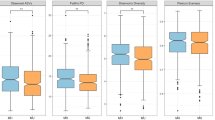
To determine whether the used categorizations improve the assessment of differences in the gut microbiota, we contrasted several ecological indices. We first evaluated differences in α-diversity and found that obese individuals and those with abnormal cardiometabolic health have less diverse microbiotas, whereas normoweight, healthy individuals have the most diverse ones (Supplementary Table S4). We detected significant differences in the observed species richness in all evaluated groupings (ANOVA for BMI: F2,438=6.51, P=0.003; CHS: F1,439=10.69, P=0.002; BSP: F5,435=3.84, P=0.004) and in the Shannon index between BMI categories (ANOVA: F2,438=3.28, P=0.04); no differences in the latter index were found for CHS or BSP. Furthermore, a significant CHS × BMI interaction was found when a linear model using BMI as a continuous variable and species richness was constructed (ANOVA: F2,438=6.16, P=0.002; Figures 3a and b). Differences in the Shannon index were not significant, but showed the same trend (Figures 3c and d). Most interestingly, the observed pattern is reminiscent of that obtained with the anthropometric and biochemical variables: changes in α-diversity are mainly explained by differences in BMI categories; nevertheless, it is possible to further differentiate individuals within a given category by their cardiometabolic health status, being the loss in α-diversity exacerbated in cardiometabolically abnormal individuals (Supplementary Table S4).
Next, we evaluated the proportion of the variance in the gut microbiota composition explained by the body size phenotypes. For this, we assessed differences in β-diversity using PERMANOVA on weighted and unweighted UniFrac distances. For both metrics, body size phenotypes better explained shifts in the gut microbiota than either BMI or cardiometabolic health status alone. The proportion of the variance explained by BSP (weighted: R2=0.017, P=0.015; unweighted: R2=0.014, P=0.007) was almost twice that of BMI (weighted: R2=0.010, P=0.001; unweighted: R2=0.007, P=0.002) and more than twice that of CHS (weighted: R2=0.007, P=0.002; unweighted: R2=0.004, P=0.003). Sensitivity analyses on α and β diversities using the subset of participants that did not smoke or consume medicaments showed similar results as the complete dataset (Supplementary Figure S1; Supplementary Tables S4 and S5).
Analyses of β diversity indicate that the gut microbiota of individuals categorized by different methods is, as a whole, somewhat different. However, since changes at this level are small (explaining ~1.5% of the total variance), we next assessed differences associated with obesity and cardiovascular risk at finer scales. We first tested whether differences in Firmicutes and Bacteroidetes were responsible of the above differences, and found small yet significant differences in the abundance of Bacteroidetes and the B/F ratio by BMI (P=0.04 in both cases; Supplementary Table S4). We found no significant differences in phylum-level microbial markers between cardiometabolic status or body size phenotypes.
Next, we looked into changes in specific networks of co-abundant microbes.29 The gut microbiota of our cohort forms the following five CAGs (Supplementary Figure S2). (i) The Pathobiont-CAG aggregated OTUs of Escherichia coli, Enterobacter hormaechei and several genera of opportunistic bacteria that contribute to various diseases under specific environmental conditions. (ii) The Lachnospiraceae-CAG consisted of OTUs from the Lachnospiraceae, Ruminococcaceae and Clostridiaceae families, including Faecalibacterium prausnitzii, as well as Actinobacteria such as Bifidobacterium adolescentis, Collinsella aerofaciens and Actinomyces sp. (iii) The Akkermansia-Bacteroidales-CAG comprised Akkermansia muciniphila, OTUs of the Bacteroidales and Clostridiales orders, and two Deltaproteobacteria (Bilophila and Desulfovibrio). (iv) The Prevotella-CAG included OTUs of Prevotella, Coriobacteriaceae and Erysipelotrichaceae. (v) Finally, the Ruminococcaceae-CAG gathered the archaeon Methanobrevibacter, many microbes of the Ruminococcaceae family, including the genus Oscillospira, and other Clostridiales such as Dialister. The complete list of OTUs gathering within each CAG is given as Supplementary Table S6.
Interestingly, the median abundance of the Lachnospiraceae-CAG and Pathobiont-CAG significantly increased with BMI and was higher in abnormal than healthy individuals according to CHS, although significance was only reached for the Pathobiont-CAG in the latter classification. In contrast, the abundance of the Akkermansia-Bacteroidales-CAG and Ruminococcaceae-CAG significantly decreased with BMI and was lower in healthy than abnormal individuals, though the difference was only significant in the Ruminococcaceae-CAG. The prevalence of the Prevotella-CAG tended to increase with BMI and to be higher in cardiometabolically abnormal than healthy individuals, but with no statistical significance (Figures 4a and b). Similar trends were seen when classifying individuals by body size phenotypes (Figure 4c). While statistical significance was not reached in all cases, it is clear that the median abundance of CAGs follows a stepped pattern, whose direction depends on the association of a given CAG with variables informing about obesity and cardiovascular health. Similar results were obtained in the analysis of the relative abundance of the most abundant OTUs of each CAG instead of the whole CAG (Supplementary Table S7).
Finally, we found a strong correspondence between the analysis of individual OTUs (LEfSe and GLM) and results by CAGs (Supplementary Table S8). OTUs classified as Methanobrevibacter, Oscillospira, Dialister, Clostridium hathewayi, Propionispora hippei, 02d06 (Clostridiaceae) and unclassified Christensenellaceae, among others, most of which belong to the Ruminococcaceae-CAG and Akkermansia-Bacteroidales-CAG, are associated with beneficial health states. On the other hand, detrimental categories are associated with OTUs clustered into the Pathobiont-CAG and Lachnospiraceae-CAG, such as E. coli, Streptococcus sp., Blautia sp., Gemmiger formicilis and SMB53 (Clostridiaceae).
Discussion
BMI is the tool par excellence to diagnose obesity.1 There is a strong association between high BMI and cardiovascular disease, hypertension, glucose intolerance, type 2 diabetes and dyslipidemia.34 However, it has been demonstrated that BMI has a low discriminatory power to distinguish body fat and lean mass.35 While flawed, its ease of use and pervasiveness make it reasonable to expect BMI to still be used in the years to come.36 It is possible, nevertheless, to compensate some of the problems associated with BMI by incorporating measurements that provide complementary information,20 such as the approach herein implemented.
The BSP classification considers the non-uniformity of disease risk associated to BMI categories by further discriminating individuals by cardiometabolic health,21 and permits the individual assessment of groups of particular interest, such as the cardiometabolically healthy–obese phenotype37 or the establishment of reference values by considering ‘truly healthy’ individuals, such as the cardiometabolically healthy–normoweight. The method is not without flaws, however. Body size phenotypes have not been standardized; clinical baseline values might vary among populations; the clinical cutoffs are to some extent arbitrary, making borderline values difficult to treat; and BSP prevalence depends upon the number of alterations required to classify individuals as cardiometabolically healthy or abnormal.21 In addition, this method requires measurement of additional clinical variables. Despite these limitations, we underscore the heterogeneity of cardiovascular risk within the different categories of BMI, demonstrating that BSP better assess disease risk compared with BMI or CHS, as demonstrated by AIC-based model selection, and highlight hidden groups of individuals. Approximately 34% of normoweight individuals were cardiometabolically abnormal, while 35% and 16% of overweight and obese participants were cardiometabolically healthy, respectively. Hence, the BSP categorization has the advantage of providing information about both the health and nutritional status of individuals, calling for more sophisticated methods for assessing cardiometabolic disease risk, as this is clearly not appropriately gauged by BMI alone.
We observed a tiered pattern in the measured variables, with the tendency to reach detrimental values with increased BMI and, within a BMI category, with abnormal cardiometabolic health. Our sensitivity analysis suggests that this pattern is not much affect by the cigarette and medicament consumption. Interestingly, differences within a given BMI category were also evidenced in the caloric intake and physical activity indicators. Although these two self-reported variables suffer of social approval bias,38 which we corroborated with higher physical activity and lower caloric intake in obese compared to normoweight subjects, it is nonetheless noteworthy that, when considering each BMI category separately, we consistently observed values that better reflected the status of the individuals in the context of energy balance.
Microbiota and health
Body size phenotypes allow detailed examinations of the gut microbiota in the studied cohort. The gut microbiota has been associated with obesity and cardiovascular health,39 albeit with small effect sizes.14 While our results show that the proportion of the variance in the gut microbiota explained by any of the explored categorizations is low, the use of BSP allows better distinguishing dissimilarities in the microbial community, confirming our first hypothesis. The fact that differences in the overall community are small reinforces the idea that changes in the abundance of a reduced number of taxa highly associated with different clinical markers are the drivers of this divergence.
We tested whether differences in the gut microbiota were due to shifts in well-known taxonomic groups; while we found changes in the abundance of Bacteroidetes and in the Bacteroidetes/Firmicutes ratio associated with BMI, the magnitude of these differences is small. In contrast, our more comprehensive approach using networks of microbes gives greater insight into the biological association between disease risk and gut microorganisms. In particular, higher risk was associated with higher prevalence of the Pathobiont-CAG whereas lower risk was associated with higher prevalence of the Akkermansia–Bacteroidales–CAG and Ruminococcaceae–CAG. The Pathobiont-CAG and Lachnospiraceae-CAG were associated with cardiometabolic disease and obesity; members of said consortia have been previously shown to be linked with these conditions. For instance, Enterobacter and Escherichia, both Gram-negative opportunistic pathogens members of the Pathobiont-CAG, may be key in obesity as lipopolysaccharide induces inflammation and endotoxemia, resulting in obesity and insulin resistance.40 Likewise, some members of Lachnospiraceae have been shown to be associated with diabetes and obesity.41 On the other hand, OTUs composing the Akkermansia–Bacteroidales–CAG and Ruminococcaceae–CAG have been linked to improved health outcomes. Akkermansia muciniphila has become one of the main targets of research and intervention in the human gut,42 as it has been consistently linked with metabolic health and leanness.43 Similarly, other members of this CAG, such as Alistipes and Bacteroides from lean mice, quickly invade the microbiota of co-housed obese mice;44 members of the Ruminococcaceae-CAG, such as Methanobrevibacter, Oscillospira and Dialister are associated with lower BMI45, 46, 47 and weight reduction.48 Analyses by OTUs showed strong agreement with those obtained with CAGs, corroborating that CAGs are valuable tools that thoroughly capture the complex and multivariate nature of the gut microbiota.29
A key parameter linking health and the gut microbiota is α-diversity. A healthy microbiota is thought to be characterized by high microbial and gene diversity, presumed to reflect ecosystem stability and resilience.17, 49 In agreement with our second hypothesis, we found a significant reduction in species richness with BMI, with a similar tendency for the Shannon index, and diminished values of these indices in subjects with abnormal CHS. These results are consistent with previous studies showing that individuals with higher metabolic risk have less diverse microbiotas.17, 50 Differences in α-diversity within a given BMI category (or any other targeted condition for that matter) should be taken into account, as the success of gut microbiota interventions, particularly dietary interventions, are dependent on the baseline α-diversity.51, 52 In mice, it has been shown that a diet low in microbiota-accessible carbohydrates induces the depletion of multiple microbes, resulting in a progressive loss of diversity over generations. Furthermore, the sole reintroduction of these carbohydrates into the diet does not restore the gut microbiota to its original state.53 Similarly, diet-induced weight loss and stabilization interventions in humans can improve health-associated parameters but their efficiency is diminished in individuals with lower gene richness.50
Interestingly, the values observed when comparing the α-diversity between body size phenotypes echoes the ladder-like pattern formed by the anthropometric and biochemical variables. This was also observed in the abundance of several of the microbial consortia evaluated here. Of the four CAGs that significantly differ between BMI categories, two also differ by CHS. This may indicate that, while the association between the Akkermansia–Bacteroidales–CAG and Lachnospiraceae–CAG with markers of cardiovascular disease is driven by their association with the nutritional status, the Pathobiont–CAG and Ruminococcaceae–CAG are associated with cardiovascular health beyond BMI.
In synthesis, it is clear that catchall methods undertaken to tackle the increasing incidence of obesity and associated diseases have been largely ineffective. Our results highlight the importance of refined classification methods that appropriately gauge individual disease risk. The use of body size phenotypes, or similar approaches that disentangle obesity and cardiometabolic disease, seems of particular importance in the context of gut microbiota-based personalized nutrition and medicine.
References
World Health Organization Global Status Report On Noncommunicable Diseases 2014 2014.
Ichihara S, Yamada Y . Genetic factors for human obesity. Cell Mol Life Sci 2008; 65: 1086–1098.
Leech RM, McNaughton SA, Timperio A . Clustering of children’s obesity-related behaviours: associations with sociodemographic indicators. Eur J Clin Nutr 2014; 68: 623–628.
The Human Microbiome Project Consortium The Human Microbiome Project Consortium Huttenhower C The Human Microbiome Project Consortium Gevers D The Human Microbiome Project Consortium Knight R The Human Microbiome Project Consortium Abubucker S The Human Microbiome Project Consortium Badger JH et al. Structure, function and diversity of the healthy human microbiome. Nature 2012; 486: 207–214.
Dugas LR, Fuller M, Gilbert J, Layden BT . The obese gut microbiome across the epidemiologic transition. Emerg Themes Epidemiol 2016; 13: 2.
Tang WHW, Kitai T, Hazen SL . Gut microbiota in cardiovascular health and disease. Circ Res 2017; 120: 1183–1196.
de la Cuesta-Zuluaga J, Mueller NT, Corrales-Agudelo V, Velásquez-Mejía EP, Carmona JA, Abad JM et al. Metformin is associated with higher relative abundance of mucin-degrading akkermansia muciniphila and several short- chain fatty acid – producing microbiota in the gut. Diabetes Care 2017; 40: 54–62.
Ley RE, Backhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI et al. Obesity alters gut microbial ecology. Proc Natl Acad Sci USA 2005; 102: 11070–11075.
Ley RE, Turnbaugh PJ, Klein S, Gordon JI . Microbial ecology: human gut microbes associated with obesity. Nature 2006; 444: 1022–1023.
Schwiertz A, Taras D, Schafer K, Beijer S, Bos NA, Donus C et al. Microbiota and SCFA in lean and overweight healthy subjects. Obesity 2010; 18: 190–195.
Duncan SH, Lobley GE, Holtrop G, Ince J, Johnstone AM, Louis P et al. Human colonic microbiota associated with diet, obesity and weight loss. Int J Obes 2008; 32: 1720–1724.
Walters WA, Xu Z, Knight R . Meta-analyses of human gut microbes associated with obesity and IBD. FEBS Lett 2014; 588: 4223–4233.
Finucane MM, Sharpton TJ, Laurent TJ, Pollard KS . A taxonomic signature of obesity in the microbiome? Getting to the guts of the matter. PLoS ONE 2014; 9: e84689.
Sze MA, Schloss PD . Looking for a signal in the noise: revisiting obesity and the microbiome. MBio 2016; 7: e01018–e01116.
Plovier H, Everard A, Druart C, Depommier C, Van Hul M, Geurts L et al. A purified membrane protein from Akkermansia muciniphila or the pasteurized bacterium improves metabolism in obese and diabetic mice. Nat Med 2016; 23: 107–113.
Goodrich JK, Davenport ER, Beaumont M, Jackson MA, Knight R, Ober C et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 2016; 19: 731–743.
Le Chatelier E, Nielsen T, Qin J, Prifti E, Hildebrand F, Falony G et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013; 500: 541–546.
Nicholls SG . Standards and classification: a perspective on the ‘obesity epidemic’. Soc Sci Med 2013; 87: 9–15.
Romero-Corral A, Somers VK, Sierra-Johnson J, Thomas RJ, Collazo-Clavell ML, Korinek J et al. Accuracy of body mass index in diagnosing obesity in the adult general population. Int J Obes 2008; 32: 959–966.
Wildman RP, Muntner P, Reynolds K, McGinn AP, Rajpathak S, Wylie-Rosett J et al. The obese without cardiometabolic risk factor clustering and the normal weight with cardiometabolic risk factor clustering: prevalence and correlates of 2 phenotypes among the US population (NHANES 1999-2004). Arch Intern Med 2008; 168: 1617–1624.
Tomiyama AJ, Hunger JM, Nguyen-Cuu J, Wells C . Misclassification of cardiometabolic health when using body mass index categories in NHANES 2005-2012. Int J Obes 2016; 40: 883–886.
Escobar JS, Klotz B, Valdes BE, Agudelo GM . The gut microbiota of Colombians differs from that of Americans, Europeans and Asians. BMC Microbiol 2014; 14: 311.
Siri WE . Body composition from fluid spaces and density: analysis of methods. Tech Meas body Compos 1961; 61: 223–244.
Manjarrés L, Manjarrés S Programa de Evaluación de Ingesta Dietética EVINDI v4. Universidad de Antioquia. 2008.
Craig CL, Marshall AL, Sjostrom M, Bauman AE, Booth ML, Ainsworth BE et al. International physical activity questionnaire: 12-country reliability and validity. Med Sci Sports Exerc 2003; 35: 1381–1395.
Matthews DR, Hosker JP, Rudenski AS, Naylor BA, Treacher DF, Turner RC . Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 1985; 28: 412–419.
Gallo J, Aristizábal D, Segura Á, Correa M, Zapata N . Relationship of insulin resistance with heart structure, function, and metabolism in young, nonobese adults. Acta Medica Colomb 2008; 33: 117–126.
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD . Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl Environ Microbiol 2013; 79: 5112–5120.
Claesson MJ, Jeffery IB, Conde S, Power SE, O’Connor EM, Cusack S et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 2012; 488: 178–184.
Friedman J, Alm EJ . Inferring Correlation Networks from Genomic Survey Data. PLoS Comput Biol 2012; 8: e1002687.
Chen J, Bittinger K, Charlson ES, Hoffmann C, Lewis J, Wu GD et al. Associating microbiome composition with environmental covariates using generalized UniFrac distances. Bioinformatics 2012; 28: 2106–2113.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS et al. Metagenomic biomarker discovery and explanation. Genome Biol 2011; 12: R60.
O’Hara RB, Kotze DJ . Do not log-transform count data. Methods Ecol Evol 2010; 1: 118–122.
McGee DL . Body mass index and mortality: a meta-analysis based on person-level data from twenty-six observational studies. Ann Epidemiol 2005; 15: 87–97.
Romero-Corral A, Montori VM, Somers VK, Korinek J, Thomas RJ, Allison TG et al. Association of bodyweight with total mortality and with cardiovascular events in coronary artery disease: a systematic review of cohort studies. Lancet (London, England) 2006; 368: 666–678.
Ortega FB, Sui X, Lavie CJ, Blair SN . Body mass index, the most widely used but also widely criticized index would a criterion standard measure of total body fat be a better predictor of cardiovascular disease mortality? Mayo Clin Proc 2016; 91: 443–455.
Denis GV, Hamilton JA . Healthy obese persons: How can they be identified and do metabolic profiles stratify risk? Curr Opin Endocrinol Diabetes Obes 2013; 20: 369–376.
Miller TM, Abdel-Maksoud MF, Crane LA, Marcus AC, Byers TE . Effects of social approval bias on self-reported fruit and vegetable consumption: a randomized controlled trial. Nutr J 2008; 7: 18.
Fu J, Bonder MJ, Cenit MC, Tigchelaar EF, Maatman A, Dekens JAM et al. The gut microbiome contributes to a substantial proportion of the variation in blood lipidsnovelty and significance. Circ Res 2015; 117: 817–824.
Cani PD, Bibiloni R, Knauf C, Waget A, Neyrinck AM, Delzenne NM et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 2008; 57: 1470–1481.
Verdam FJ, Fuentes S, de Jonge C, Zoetendal EG, Erbil R, Greve JW et al. Human intestinal microbiota composition is associated with local and systemic inflammation in obesity. Obesity 2013; 21: E607–E615.
Derrien M, Belzer C, de Vos WM . Akkermansia muciniphila and its role in regulating host functions. Microb Pathog 2016; 106: 171–181.
Dao MC, Everard A, Aron-Wisnewsky J, Sokolovska N, Prifti E, Verger EO et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: relationship with gut microbiome richness and ecology. Gut 2016; 65: 426–436.
Ridaura VK, Faith JJ, Rey FE, Cheng J, Duncan AE, Kau L et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science (80-) 2013; 341: 1241214.
Million M, Angelakis E, Maraninchi M, Henry M, Giorgi R, Valero R et al. Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int J Obes 2013; 37: 1460–1466.
Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R et al. Human genetics shape the gut microbiome. Cell 2014; 159: 789–799.
Konikoff T, Gophna U . Oscillospira: a Central, Enigmatic Component of the Human Gut Microbiota. Trends Microbiol 2016; 24: 523–524.
Louis S, Tappu R-M, Damms-Machado A, Huson DH, Bischoff SC . Characterization of the gut microbial community of obese patients following a weight-loss intervention using whole metagenome shotgun sequencing. PLoS ONE 2016; 11: e0149564.
Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R . Diversity, stability and resilience of the human gut microbiota. Nature 2012; 489: 220–230.
Cotillard A, Kennedy SP, Kong LC, Prifti E, Pons N, Le Chatelier E et al. Dietary intervention impact on gut microbial gene richness. Nature 2013; 500: 585–588.
Tap J, Furet J-P, Bensaada M, Philippe C, Roth H, Rabot S et al. Gut microbiota richness promotes its stability upon increased dietary fibre intake in healthy adults. Environ Microbiol 2015; 17: 4954–4964.
Dao MC, Everard A, Clement K, Cani PD . Losing weight for a better health: Role for the gut microbiota. Clin Nutr Exp 2016; 6: 39–58.
Sonnenburg ED, Smits SA, Tikhonov M, Higginbottom SK, Wingreen NS, Sonnenburg JL . Diet-induced extinctions in the gut microbiota compound over generations. Nature 2016; 529: 212–215.
Acknowledgements
We thank the participants who took part in the study, and the Vidarium, EPS SURA and Dinámica IPS staff that helped with recruitment and field work; the APOLO Scientific Computing Center at EAFIT University hosted bioinformatics resources and the University of Michigan Medical School Host Microbiome Initiative for sequencing. This work was funded by Grupo Empresarial Nutresa, EPS SURA, and Dinámica I.P.S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
JdlCZ, VCA and JSE are employees of a food company. JAC and JMA are employees of health provider companies..
Additional information
DISCLAIMER
The funders of this work have not had any role in designing or conducting the study; in the collection, analysis, or interpretation of the data; in the preparation, review, or approval of the manuscript; or in the decision to submit the manuscript for publication.
Supplementary Information accompanies this paper on International Journal of Obesity website
Supplementary information
Rights and permissions
About this article
Cite this article
de la Cuesta-Zuluaga, J., Corrales-Agudelo, V., Carmona, J. et al. Body size phenotypes comprehensively assess cardiometabolic risk and refine the association between obesity and gut microbiota. Int J Obes 42, 424–432 (2018). https://doi.org/10.1038/ijo.2017.281
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2017.281
- Springer Nature Limited
This article is cited by
-
Gut microbiota in patients with obesity and metabolic disorders — a systematic review
Genes & Nutrition (2022)
-
Profile of gut microbiota and serum metabolites associated with metabolic syndrome in a remote island most afflicted by obesity in Japan
Scientific Reports (2022)
-
The hallmarks of dietary intervention-resilient gut microbiome
npj Biofilms and Microbiomes (2022)
-
The value of gut microbiota to predict feed efficiency and growth of rabbits under different feeding regimes
Scientific Reports (2021)
-
Chronic oral exposure to pesticides and their consequences on metabolic regulation: role of the microbiota
European Journal of Nutrition (2021)