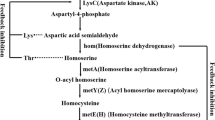
Abstract
Aspartate 4-decarboxylase (ASD) has been modified to obtain the catalytic ability of the unnatural substrate l-3-methylaspartate. However, the mechanism remains to be clarified. In the present study, the semi-rational modification was used to identify key residues of importance for the activity towards l-3-methylasparte. The ASD from Pseudomonas dacunhae 21192 (PdASD) was used as a template, which showed better activity than the other two ASDs. Four residues proved to be critical for the activity towards l-3-methylasparte, with three located in the active site and one on the surface. Combinatorial variants were constructed to analyze the role of each mutation. The enzymatic properties of the combined variants were determined and compared. The residue at the 17th position was a member of the substrate entrance gate and contributed to the activity by reducing the steric hindrance. The residue at the 37th position was necessary for activity. Two mutations, I288 and V382, exhibited strong epistatic interactions on the activity of ASD. Structural changes in the active site were analyzed by molecular dynamics simulations, and it is proposed that the increased activity of PdASD variants is related to a suitable binding pocket for the substrate. These results provide new evidence for the mechanism of β-decarboxylation, which lays the foundation for enhancing the activity of ASD.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
For decades, Pseudomonas dacunhae containing aspartate 4-decarboxylase (ASD) has been employed to produce l-alanine in the industry because of the stability and high catalytic efficiency of this enzyme [1,2,3]. Recently, more and more ASD genes have been cloned and characterized [4,5,6,7]. The crystal structures of ASD derived from P. dacunhae and Alcaligenes faecalis were resolved decades ago [8, 9]. Based on the crystal structure and characterization of enzymatic properties, molecular modifications were applied to ASDs to investigate the catalytic mechanism [8, 10, 11], leading to the development of new catalytic reactions [12,13,14].
The substrate channel and active site of an enzyme have always been the focus of catalytic mechanism research. The active site of an enzyme is the region where the enzyme binds to the substrate and catalyzes the reaction. Although the active site of an enzyme prefers the natural substrate, modification of the active site can effectively change the substrate specificity of the enzyme. For example, a chitosanase variant from Bacillus sp. MN accepted N-acetyl-d-glucosamine units as the substrate by modifying the enzyme activity site [15]. An ASD dual-point variant from P. dacunhae with remarkably improved activity towards 3(R)-benzyl-l-aspartate was constructed based on structure-guided engineering [14].
Moreover, tunnels and channels connecting the environment to the active site affect enzyme activity, specificity, stability and enantioselectivity [16]. A key tunneling residue in nitrile hydratase from Rhodococcus rhodochrous J1 to rac-mandelonitrile was identified and a variant with shifted stereoselectivity was obtained [17]. After changing the size and shape of the substrate tunnel, the regioselectivity of nitrile hydratases from Pseudomonas putida and Comamonas testosteroni switched between the selective hydration of two cyano groups of α,ω-dinitriles [18].
It has been reported that ASD from Acinetobacter radioresistens (ArASD) obtained catalytic activity towards L-3-methylasparte through molecular modifications of the active site and the substrate entrance gate [13]. In this study, the activity of ASD from P. dacunhae 21192 (PdASD) towards l-3-methylasparte was significantly enhanced by the semi-rational modification, and then the key residues of PdASD were identified. Molecular dynamics simulations and characterization of enzymatic properties were performed to analyze the effect of each residue on the activity and the structural changes.
Materials and methods
Construction of variants
The pET28a vector was used to express ASD genes. ASD genes originating from Acinetobacter radioresistens, Acinetobacter tandoii and Pseudomonas dacunhae 21192, respectively, were synthesized by AZENTA (Suzhou, China). Two hosts were employed: Escherichia coli JM109 was used to construct the plasmid (pET28a-ASD) and Escherichia coli BL21 (DE3) was used to express enzymes (ASDs).
The pET28a-pd21192-ASD was used as a template for site-directed mutagenesis, and the primers are listed in Table S1. DNA polymerase (Primer STAR Max, Takara Bio Inc., China) was used throughout the PCR procedure.
Strain culture and protein purification
LB medium was used for bacterial growth and 2YT medium was used to express enzymes. The plasmid, which was correctly sequenced, was transformed into BL21. BL21 was then cultured overnight at 37 ℃ in LB solid medium (containing 50 μg/ml kanamycin). The colony was picked into 5 ml of LB liquid medium and cultured at 37 ℃ for 8 h. Then, 1 ml of bacterial broth was inoculated into 100 ml of 2YT medium. When OD600 reached the range of 0.6–0.8, IPTG (isopropyl-β-D-thiogalactoside) was added at a final concentration of 0.2 mM and the bacteria were induced at 25 ℃ for 20 h. Bacterial cells were collected and the enzyme was purified. The purification method was the same as in the previous article [7]. The His Trap HP column (GE Healthcare, USA) was used to purify enzymes. Binding buffer (including 20 mM imidazole) and washing buffer (including 500 mM imidazole) were used to balance the column and elute the enzyme, respectively. The Bradford method was used for determining the protein concentration.
Enzymatic activity assay and dual enzyme system catalytic assay
The appropriate amount of enzyme was dissolved in 10 mM sodium acetate–acetate buffer (pH 5.4) containing 0.5 mM PLP (Pyridoxal-5'-phosphate monohydrate) and 0.5 mM pyruvate and then incubated at 45 ℃ for 5 min. 0.1 M of l-3-Methylasparte was added to the reaction system. After 30 min, inactivation was performed at 100 ℃ for 10 min.
Reactions catalyzed by the dual enzyme system (1 mg/ml glutamate mutase and 1 mg/ml ASD) were carried out in potassium phosphate buffer (10 mM, pH 6.6, containing 5 mM PLP, 2 mM pyruvate, 1 mM DTT, 0.25 mM adenosylcobalamin and 100 mM l-glutamate) at 37 °C for 4 h. The inactivation was then performed at 100 °C for 10 min.
The inactivated mixture was centrifuged at 12 krpm for 10 min and the supernatant was used to detect the concentration of l-l-2-aminobutyrate. The detection of amino acids was performed as in the previous article [7]. Samples were derivatized by phenylisothiocyanate (PITC). A C18 column (250 × 4.6 mM, Dikma, China) was used to detect the concentration of l-l-2-aminobutyrate.
Molecular dynamics simulation
The structures of PLP and l-3-methylasparte were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov), and l-3-methylasparte was configured as (2S, 3S)-3-methylaspartate. l-3-methylasparte was covalently linked to PLP as a ligand that was docked to the wild-type ASD (PDB: 3fdd). Molecular docking was completed by the glide module of Schrodinger software, and molecular docking models were optimized by Schrodinger software. Molecular dynamics simulations were run using NAMD 2.14 software with the CHARmM36 force field. The topology and G3P parameters with the CHARmM force field were made using the web server SwissParam32 (https://www.swissparam.ch/). Kinetic simulations include 1 ns of water equilibration, 60 ps from 0 to 300 k and 50 ns of classical MD. TTcluster was used to analyze molecular dynamics simulation trajectories. DCCM was analyzed using Bio3D.
The FEP/HREMD method was used to calculate the binding free energy. CHARmM-GUI was used to generate FEP/HREMD input files. The size of the ligand binding pocket was calculated by POVME 3.0. The grid spacing was set to 1.0, and the distance cutoff was set to 1.09. Pockets were defined in all grid points within 3 Å of the ligand. The trajectory files of the 50-ns molecular dynamics simulations were extracted from 250 frames of PBD files at 0.2 ns intervals and used as input files for POVME 3.0.
Results and discussion
Activity comparison of ASDs from different organisms
The molecular modification was applied to ArASD to improve its ability to convert l-3-methylasparte to l-2-aminobutyrate, and l-glutamate was catalyzed to L-2-aminobutyrate by coupling a variant of ArASD to a glutamate mutase [13]. Subsequently, a variant, Ar-K18A/R38K/V287I, with significantly high activity towards l-3-methylasparte was obtained [19]. Sequence alignment was carried out between ArASD, PdASD and ASD from Acinetobacter tandoii (AtASD), and the corresponding variants of PdASD and AtASD, At-K18A/R38K/V287I and Pd-K17A/R37K/V288I, were constructed. The catalytic abilities of three variants, Ar-K18A/R38K/V287I, At-K18A/R38K/V287I and Pd-K17A/R37K/V288I, were compared (Fig. 1). The highest concentration of l-2-aminobutyrate was synthesized by coupling PdASD with glutamate mutase. Obviously, the catalytic ability of PdASD was superior to that of AtASD and ArASD, a result that is consistent with our previous study [7]. Additionally, β-decarboxylation activities of l-3-methylasparte were detected for ArASD, PdASD and AtASD (data not shown), which is consistent with the results of the dual enzyme system.
The β-decarboxylation activity of variants from different organisms towards l-3-methylasparte. The activity was assessed by the concentration of l-2-aminobutyrate synthesized from L-glutamate using purified ASD and purified glutamate mutase. The reactions were performed at pH 6.6 for 4 h. Ar: Acinetobacter radioresistens; At: Acinetobacter tandoii; Pd: Pseudomonas dacunhae 21192
Screening of key residues for the activity of ASD towards L-3-methylasparte
The substrate channel of PdASD was predicted to identify key residues for the activity of PdASD towards the larger substrate l-3-methylasparte. Three substrate channels were predicted by the software Caver (Fig S1). The shortest assumed channel has a bottleneck, the longest assumed channel has a few bottlenecks, and the middle assumed channel has two bottlenecks. The shortest assumed channel and the middle assumed channel were selected for alanine scanning of the constituent residues of the three bottlenecks. After excluding nine highly conserved residues and small residues (alanine, glycine), six of the fifteen residues were mutated into alanine to optimize the room for the narrowest part of the channel. Since wild-type activity towards l-3-methylasparte was not detected and valine at the 288th position was located at the active site, variant K17A/R37K was used as a template instead of variant K17A/R37K/V288I.
One candidate, threonine at the 382nd position, was screened out. The variant K17A/R37K/T382A showed higher activity than K17A/R37K and K17A/R37K/V288I (Fig. 2a). Since the threonine at the 382nd position played an important role in the activity towards l-3-methylasparte, the residue was subjected to site-saturation mutagenesis. The three variants, T382A, T382V and T382L, exhibited better catalytic ability, with variant K17A/R37K/T382V showing the highest activity towards l-3-methylasparte (Fig. 2b). The polar residues at this position have been found to be unfavorable for the activity towards l-3-methylasparte, while the aliphatic residues at this position are beneficial for its activity. The T382 residue was also considered important for the activity of 3(R)-benzyl-l-aspartate; however, the variant T382G was the best [14]. This suggests that the 382nd residue of PdASD is essential for substrate specificity, but the effect of residues depends on the substrate.
Screening of variants. a Alanine scanning of residues located at the substrate of the assumed channels. b Site saturation mutation of the 382nd residue. Activity towards L-3-methylasparte was determined at pH 5.4 using purified enzymes with the activity of K17A/R37K/V288I and K17A/R37K/T382A defined as 100%, respectively
Key residues for the β–decarboxylation activity
So far, four residues of PdASD have been shown to be important for the activity of l-3-methylasparte. Three (R37, V288 and T382) are located in the active site and one (K17) on the surface, which has been resolved by crystal structure analysis [9]. Combinatorial variants were constructed and their catalytic abilities were compared via two examinations: specific activity towards l-3-methylasparte (Fig. 3a) and synthesis of l-2-aminobutyrate from l-glutamate using a dual enzyme system containing a variant of PdASD and glutamate mutase (Fig. 3b). The results of the two experiments are consistent and include variants K17A/R37K and K17A/T382V with very little specific activity, but synthesize some l-2-aminobutyrate. We attribute this to their extremely low activity and excellent stability. Two variants, K17A/R37K/V288I/T382V and K17A/V288I/T382V, exhibited significantly higher catalytic abilities than the other variants (Fig. 3a and b).
Comparison of catalytic abilities of the combinatorial variants. a The specific activities of the combinatorial variants towards L-3-methylasparte. Reactions were performed at pH 5.4 for 30 min. b Concentration of L-2-aminobutyrate catalyzed by purified combinatorial variants and purified glutamate mutase. Reactions were performed at pH 6.6 for 4 h
Only one variant, K17A/T382V/ΔR37, was deprived of the activity, in which the 37th residue was deleted. No activity was detected for K17A/T382V/ΔR37 compared to the slightly more active variant K17A/T382V, indicating that the residue at the 37th position is an essential residue for the activity of l-3-methylasparte. In addition, no activity of K17A/T382V/ΔR37 towards the natural substrate, L-aspartate, was detected (Fig S2). Our results support that the residue at the 37th position is critical for regulating the conformation of the internal aldimine of PLP-dependent enzymes [19]. However, our results contradict a report suggesting that R37 only plays a role in optimizing catalytic efficiency, but is not essential for the activity of ASD towards l-aspartate [8].
Variant K17A/V288I exhibited better catalytic ability than variant K17A/T382V, indicating a greater effect of mutation I288 than V382 on the activity of l-3-methylasparte. Variant K17A/V288I/T382V presented better catalytic ability than both K17A/V288I and K17A/T382V, indicating that the best activity is achieved through collaborative cooperation of I288 and V382. The results suggest that mutations in V288I and T382V exhibit an epistatic effect, which has been shown to exist in proteins [20]. The same phenomenon was also observed in variants K17A/R37K/T382V, K17A/R37K/V288I and K17A/R37K/V288I/T382V. However, the activity of K17A/V288I and K17A/T382V towards l-aspartate showed opposite results (Fig S2), implying that a mutation may be beneficial for an enzyme-catalyzed reaction but detrimental for the other one. This conclusion was also reached by comparing the activities of two variants with only one different residue towards l-aspartate and l-3-methylasparte, such as K17A/R37K and K17A/R37K/T382V, K17A/R37K and K17A/R37K/V288I, K17A/R37K/V288I and K17A/R37K/T382V and K17A/R37K/V288I/T382V and K17A/R37K/V288I.
The residue K17 has been predicted to be a member of the gate of the substrate entrance [9]. Comparing the catalysis of K17A/V288I/T382V and V288I/T382V (Fig. 3a and b), our results also agree that K17A improves the activity toward L-3-methylasparte by reducing the steric hindrance of the 17th residue.
Structural changes in the active site of V288I/T382V
Variant V288I/T382V was targeted to analyze the conformational differences in the active site, as only a slight difference was observed between the activities of K17A/V288I/T382V and K17A/R37K/V288I/T382V, and the residue at the 17th position was located on the surface. Molecular dynamics simulations were performed to gain insight into the conformational changes resulting from the V288I and T382V mutations. The 50-ns molecular dynamics simulation was run for the wild type, two single point variants V288I and T382V, and the dual-point variant V288I/T382V, respectively. The results of the RMSD showed that the dual-point variant V288I/T382V was more stable than the other three ASDs (Fig S3). This conclusion was also drawn from the results of the trajectory analysis (Fig. 4). The dynamics cross-correlation map shows that the region is dynamically correlated with flexibility [21]. The correlation of the variant V288I/T382V was better than the other three ASDs, supporting the V288I/T382V variant as the most stable one.
The size of the substrate-binding pocket was calculated and compared. The pocket of the wild type was observed to be much larger than that of the three variants, while the pocket of V288I/T382V was the smallest (Fig. 5a). The ligand covalently bound by l-3-methylasparte and PLP was docked at the active site of the wild type and variants. The free energy of the enzyme–ligand complex was then calculated. As shown in Fig. 5b, the complex of V288I/T382V with the ligand had the lowest free energy, indicating that V288I/T382V has the strongest binding affinity to l-3-methylasparte [22, 23]. The free energy of V288I was lower than that of T382V, suggesting that V288I has a stronger binding affinity to l-3-methylasparte. These results support the activity assay with activity in the order K17A/V288I/T382V > K17A/V288I > K17A/T382V (Fig. 3). Focusing on the active site, the substrate-binding pocket of the wild type was remarkably smaller than that of V288I/T382V (Fig. 6). Recent studies have shown that the enzyme will obtain a significant rate enhancement if the reactivity region is populated [24]. The side chains of I288 and V382 are larger than those of V288 and T382. Presumably, the side chains of I288 and V382 only fill a portion of the substrate-binding pocket, so l-3-methylasparte can fit comfortably into the pocket. A suitable combination of l-3-methylasparte and V288I/T382V could produce better catalysis.
Calculation of the substrate-binding pocket size and binding free energy of the enzyme–ligand complex. a Analysis of the pocket size of wild-type PdASD (WT) and its variants with different colors illustrated. b Determination of the conformational free energy of the enzyme–ligand complex. Enzymes include wild-type PdASD (WT) and its variants. The ligand is l-3-methylasparte covalently linked to PLP
Comparison of the active site of PdASD and V288I/T382V. PLP-bound l-3-methylasparte was used as the ligand for docking with the dimer of wild-type PdASD and V288I/T382V. Three loops containing three key residues located at the active site are presented. Loop1 (G36-N41) and loop2 (D286-G290) are located in the same subunit, and loop3 (R375-S386) is located in another subunit, which is shown in a different color. The three key residues are indicated by light purple sticks and light pink sticks, respectively. The ligand is shown by the green stick
Conclusion
In this study, four key residues for the activity of ASD towards l-3-methylaspartate were identified by semi-rational modifications. Molecular dynamics simulations and characterization of enzymatic properties were performed to analyze the effect of each residue on activity and structural changes. Two variants with higher catalytic efficiency towards l-3-methylaspartate were obtained, and their increased activities were attributed to a suitable binding pocket for the substrate. A trade-off relationship between the activity towards l-3-methylaspartate and towards the natural substrate (l-aspartate) was also supported. These results provide new ideas for the mechanism of β-decarboxylation.
Availability of data and materials
The data generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
References
Chibata I, Kakimoto T, Kato J. Enzymatic production of l-alanine by Pseudomonas dacunha. Appl Microbiol Biotechnol. 1965;13:638–45.
Shibatani T, Kakimoto T, Chibata I. Stimulation of l-asparate beta-decarboxylase formation by l-glutamate in Pseudomonas dacunhae and improved production of l-alanine. Appl Environ Microbiol. 1979;38:395–464.
Takamatsu S, Tosa T. Production of l alanine and d-aspartic acid. Bioprocess Technol. 1993;16:25–35.
Chen CC, Chou TL, Lee CY. Cloning, expression and characterization of l-aspartate β-decarboxylase gene from Alcaligenes faecalis CCRC 11585. J Ind Microbiol Biotechnol. 2000;25:132–40.
Lee DG, Song TY, Kim NY, Lee EJ. Cloning, purification and characterization of novel l-aspartate β-decarboxylase from Enterococcus faecalsis. J Life Sci. 2006;16:44–8.
Liu Z, Zhao H, Han L, Cui WC, Zhou Z, Zhou Z. Improvement of the acid resistance, catalytic efficiency, and thermostability of nattokinase by multisite-directed mutagenesis. Biotechnol Bioeng. 2019;116:1833–43.
Cui R, Liu Z, Yu P, Zhou L, Zhou Z. Biosynthesis of l-alanine from cis-butenedioic anhydride catalyzed by a triple-enzyme cascade via a genetically modified strain. Green Chem. 2021;23:7290.
Lima S, Sundararaju B, Huang C, Khristoforov R, Momany C, Phillips SR. The crystal structure of the Pseudomonas dacunhae aspartate-β-decarboxylase dodecamer reveals an unknown oligomeric assembly for a pyridoxal-5′-phosphate-dependent enzyme. J Mol Biol. 2009;388:98–108.
Chen HJ, Ko TP, Lee CY, Wang NC, Wang AH. Structure, assembly, and mechanism of a PLP-dependent dodecameric l-aspartate beta-decarboxylase. Structure. 2009;17:517–29.
Wang NC, Ko TP, Lee CY. Inactive S298R disassembles the dodecameric l-aspartate 4-decarboxylase into dimers. Biochem Biophys Res Commun. 2008;374:134–7.
Phillips SR, Lima S, Khristoforov R, Sudararaju B. Insights into the mechanism of Pseudomonas dacunhae aspartate beta-decarboxylase from rapid-scanning stopped-flow kinetics. Biochemistry. 2010;49:5066–73.
Wang NC, Lee CY. Enhanced transaminase activity of a bifunctional L-aspartate 4-decarboxylase. Biochem Biophys Res Commun. 2007;356:368–73.
Liu Y, Han L, Cheng Z, Liu ZZ, Zhemin. Enzymatic biosynthesis of L-2-aminobutyric acid by glutamate mutase coupled with l-aspartate-β-decarboxylase using l-glutamate as the sole substrate. ACS Catal. 2020;10:13913–7.
Zhang M, Hu P, Zheng Y-C, Zeng B-B, Chen Q, Zhang Z-J, Xu J-H. Structure-guided engineering of Pseudomonas dacunhae l-aspartate β-decarboxylase for l-homophenylalanine synthesis. Chem Commun. 2020;56:13876.
Regel EK, Weikert T, Niehues A, Moerschbacher BM, Singh R. Protein-Engineering of chitosanase from Bacillus sp. MN to alter its substrate specificity. Biotechnol Bioeng. 2018;115:863–73.
Kokkonen P, Bednar D, Pinto G, Prokop Z, Damborsky J. Engineering enzyme access tunnels. Biotechnol Adv. 2019;37:107386.
Cheng Z, Peplowski L, Cui W, Xia Y, Liu Z, Zhang J, Kobayashi M, Zhou Z. Identification of key residues modulating the stereoselectivity of nitrile hydratase toward rac-mandelonitrile by semi-rational engineering. Biotechnol Bioeng. 2018;115:524–35.
Cheng Z, Cui W, Liu Z, Zhou L, Wang M, Kobayashi M, Zhou Z. A switch in a substrate tunnel for directing regioselectivity of nitrile hydratases towards α, ω-dinitriles. Catal Sci Technol. 2016;6:1292.
Liu Y, Hao M, Zhou Z, Liu Z. Isolation and mechanism analysis of a catalytic efficiency improved l-aspartate β-decarboxylase toward 3-methylaspartic acid. Syst Microbiol Biomanuf. 2022;2:157–64.
Parera M, Martinez AM. Strong epistatic interactions within a single protein. Mol Biol Evol. 2014;31:1546–53.
Yu H, Dalbya AP. Exploiting correlated molecular-dynamics networks to counteract enzyme activity–stability trade-off. Proc Natl Acad Sci USA. 2018;115:E12192–200.
Swanson JM, Henchman RH, Mccammon JA. Revisiting free energy calculations: a theoretical connection to MM/PBSA and direct calculation of the association free energy. Biophys J. 2004;86:67–74.
Zhang W, Jia M, Yu M, Zhang T, Zhou L, Jiang B, Mu W. Improving the thermostability and catalytic efficiency of the d-Psicose 3-Epimerase from Clostridium bolteae ATCC BAA-613 using site-directed mutagenesis. J Agric Food Chem. 2016;64:3386–93.
Bonk MB, Weis WJ, Tidor B. Machine learning identifies chemical characteristics that promote enzyme catalysis. J Am Chem Soc. 2019;141:4108–18.
Funding
This work was supported by the National Natural Science Foundation of China (21878125).
Author information
Authors and Affiliations
Contributions
MH has made a major contribution and performed experiments in variant construction, characterization and molecular dynamics simulation. LH supervised the molecular dynamics simulation and structural analysis. ZL conceived the project and designed the experiments. The manuscript was drafted by MH and ZL, revised by ZZ and approved by all co-authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this article.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hao, M., Han, L., Zhou, Z. et al. Identification of key residues for the activity of aspartate 4-decarboxylase towards l-3-methylaspartate. Syst Microbiol and Biomanuf 3, 449–456 (2023). https://doi.org/10.1007/s43393-022-00143-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43393-022-00143-2