Abstract
In this study, 10 lactic acid bacteria were isolated from Turkish fermented sausage (sucuk) and identified as 5 Lactobacillus plantarum, 1 Pediococcus acidilactici, 1 Weissella hellenica, 1 Lactobacillus pentosus, and 2 Lactobacillus sakei. PCR screening of genes encoding plantaricin A and pediocin showed the presence of plantaricin A gene in 9 and pediocin gene in 3 of strains. All isolates showed antibacterial and antifungal effect on most of the tested microorganisms. gad gene, encoding glutamic acid decarboxylase enzyme, was detected in all isolates except Weisella hellenica KS-24. Eight of isolates were determined as gamma-amino butyric acid (GABA) producer in the presence of 53 mM mono sodium glutamate (MSG) by HPLC and TLC analysis. DPPH scavenging activity was observed for all isolates. Additionally, isolates were able to produce exopolysaccharide in the presence of sucrose. The best exopolysaccharide (EPS) production was achieved with L. plantarum KS-11 and L. pentosus KS-27. As a result, this study characterized some techno-functional properties of LAB isolates from sucuk. It was concluded that the isolates studied have the potential to be used in obtaining functional products in meat industry, as well as strain selection may be effective in providing the desired properties in the product.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Turkish fermented sausage locally known as sucuk is a fermented dry-cured meat product which contains sheep/beef or buffalo meat. It is produced by using meat /tail fat together with various spices [1]. Sucuk manufacturing is made in two different ways, traditionally and commercially. Traditional method is differed by spontaneous fermentation (fermentation is carried out without starter culture) (Gökalp, Kaya, & Zorba, 2002). Lactic acid bacteria (LAB) are the main group of microbiota in fermented sausages. Lactobacillus is the most common genera isolated from sucuk [2, 3] similar to dry fermented sausages [4, 5]. Pediococcus, Weisella, Lactococcus, and Leuconostoc were generally isolated the other LAB genera in sucuk [2, 3, 6, 7]. Lactobacillus sakei, L. curvatus, L. plantarum, Lactobacillus pentosus, Lactobacillus casei, Pediococcus pentosaceus, and P. acidilactici are the most commonly used species in commercially presented culture preparations, for meat products [8]. However, in order to provide the taste and aroma of sucuk produced with traditional methods in industrial scale, commercial starter cultures specific to sucuk are needed. Preparation of starter culture for sucuk can be achieved by using strains isolated from the product.
The technological properties of fermented sausages can be altered depending on the biochemical capabilities of LAB strains [3, 9,10,11]. Fermented sausages are primarily characterized by increased acidity and distinctive aroma provided by fermentation [12]. LAB is effective in providing product-specific aroma with their technological features, as well as highly effective on texture, shelf life, and safety of product. Lactic acid bacteria contribute to product safety with antibacterial and antifungal effects exhibited by bacteriocin/bacteriocin-like metabolites and organic acids etc. [13,14,15]. Moreover, some lactic acid bacteria have antioxidative effects and decrease the accumulation of reactive oxygen species (ROS), as functional properties [16]. In the recent years, interests have been drawn to the ability of lactic acid bacteria to produce gamma-amino butyric acid (GABA), which is a non-protein amino acid structure further supporting their functionality [17]. GABA has many structural roles known for human metabolism such as cardiovascular functions and neurotransmitter in the brain [18,19,20]. GABA can be produced by α-decarboxylation of glutamic acid in the presence of glutamic acid decarboxylase (GAD) enzyme [21, 22]. And also, lactic acid bacteria can produce exopolysaccharide (EPS) in various composition and functionality [23, 24]. Lactic acid bacteria exopolysaccharides enhanced oxidative stability, color [25], and textural properties [26] of sausages. EPS can be showed prebiotic activity that has beneficial effects on human health [27].
Several studies on the characterization of lactic acid bacteria from sucuk have been performed [2, 3, 7, 28, 29] but the techno-functional properties of LAB isolates have not yet been investigated and detailed till now.
In this study, 10 different lactic acid bacteria strains were isolated and identified from traditionally produced sucuk. While the functional aspects of these identified lactic acid bacteria such as GABA and EPS production capabilities and antioxidative properties were examined, their antimicrobial activities were also determined. This study has showed that lactic acid bacteria isolated from sucuk, a fermented meat product, have potential to provide important functional properties to the product.
Materials and methods
Isolation of lactic acid bacteria
A total of 10 sucuk samples were collected from butchers in Bayburt province in summer season and used for isolation and identification purposes of lactic acid bacteria. All manufacturers used traditional methods (without starter culture).
For isolation and identification purposes, 10 g of sample was initially taken into sterile stomacher bags under aseptic conditions and then 90 mL of sterile saline solution (0.85%) was added and homogenized. The same procedure was repeated for each sample. The spread plate technique on MRS (de Man, Rogosa and Sharpe) plates was used for proper dilutions and incubated at 30 °C for 48 h in anaerobic conditions. Following incubation, typical colonies were selected and growth in MRS broth at 30 °C for 24 h. Same procedures were continuously repeated until pure culture was obtained from single colonies. For the obtained colonies, various properties such as morphological structures, microscopic views, Gram staining, and catalase properties were determined. Considering that lactic acid bacteria are Gram positive, catalase negative, and cocci or rod-shaped, the cultures selected were stored at − 80 °C in tubes containing glycerol (30%), glass beads, and MRS broth for further diagnostic tests.
Genotypic characterization by RAPD-PCR analysis
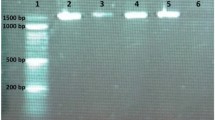
For strain differentiation of 100 LAB isolates, RAPD-PCR analysis was performed. Colonies of isolates were used as templates (without DNA isolation). Each PCR mix contained 0.75 µl template, 25 pmol of primer M13 (GAGGGTGGCGGTTCT), 5XPCR buffer for Taq polymerase (Promega, UK), 2.5 mM of deoxyribonucleoside triphosphates (dNTPs) (Bioline, UK), and 1.5 U Taq polymerase (Promega, UK). For the amplification with primer M13, denaturation was performed at 94 °C for 60 s, binding at 40 °C for 20 s, and 2 min for extension at 72 °C. PCR products were separated by electrophoresis in 1% agarose gel for 90 V 1.5 h [30].
Bacterial identification by 16S rRNA gene sequencing
Genomic DNA of isolates were extracted according to Barış [31]. For 16 sRNA gene sequences, primers AMP_F and AMP_R [32] were used. PCR mix contained 25 µL PCR Master Mix (Thermo Scientific, cat. No. K0171), 1 µL template DNA, 10 µM AMP_F, and 10 µM AMP_R primers and water in 50 µL total volume. The PCR program was run as 95 °C for 12 min, 20 cycles of 95 °C for 30 s, 55 °C for 20 s, and 72 °C for 30 s and 72 °C for 5 min final extension. Electrophoresis of PCR products were performed in gel prepared with 1% agarose to confirm the presence of amplification. Sequence analysis of PCR products were performed by GEN Plaza (Turkey). The 16S rRNA gene sequences of isolates were arranged in Bioedit and compared with NCBI database for the final identification by using Blast program (http://blast.ncbi.nlm.nih.gov). The 16S rRNA gene sequences were deposited in GenBank under accession numbers MK694773-MK694781 and MK845561 and were aligned in MEGAX 10.1.7. Phylogenetic tree was constituted by using neighbor join (NJ) method with 1000 bootstrap replicates [33]. Phylogenetic tree analyses were done with MEGAX [34].
Molecular detection of glutamic acid decarboxylase gene (gad) in LAB strains
The molecular detection of glutamic acid decarboxylase in LAB strains performed using primers CoreF (5′-CCTCGAGAAGCCGATCGCTTAGTTCG-3′) and CoreR (5′ TCATATTGACCGGTATAAGTGATGCCC-3′) [35]. PCR was performed with the following program: 10 min denaturation at 95 °C, followed by 20 cycles of 30 s at 95 °C, 30 s at 58 °C, and 1 min at 72 °C. PCR products were run on a gel (1.5%) to check the amplicon size 600 bp.
Determination GABA production levels of LAB strains
LAB isolates were grown in MRS broth containing 53 mM monosodium glutamate (MSG) for 96 h at 30 °C. It was then centrifuged at 6000 g for 5 min and the resulting supernatant was analyzed for the presence of GABA using thin layer chromatography (TLC). For this purpose, Silicagel 60 F254 TLC plates (Merck, Darmstadt, Germany) and 1-butanol: acetic acid: distilled water (4: 1:1) as mobile phase were used. After 2 h running, the plate was sprayed with 0.2% (w/v) ninhydrin solution and heated at 105 °C for 2 min. GABA producing isolates were detected by the spot loaded with MSG and GABA [36].
For quantitative evaluation, supernatant of GABA producer isolates were filtered by cellulose acetate syringe filter (0.45 µm). The GABA content of 72-h cell supernatants was determined by high-pressure liquid chromatography (HPLC) analysis. GABA derivatization was accomplished with dansyl chloride. Inertsil ODS column (13.5 µm, 4.6*250 mm) and 1:1 methanol: H2O (v:v) as mobile phase were used. The samples were separated at 40 °C and flow rate of 0.5 mL/min. The GABA content was given as “mM” based on standard curve prepared of GABA analytical standard (Sigma-Aldrich) [37].
Moleculer detection of bacteriocin encoding genes in LAB strains
The presence of gene encoding bacteriocin production was determined with different primer sets and reactions were performed at different annealing temperatures that were suitable for each primer sets (Table 1). PCR products were run on a gel (1% agarose) with electrophoresis at 100 V for 40 min to check the amplification. In the positive strains, appropriate amplicon sizes were detected.
Determination of antibacterial and antifungal activities of LAB strains
The antibacterial activities of the isolates against indicator pathogens (Bacillus cereus BC 6830, Escherichia coli BC 1402, Staphylococcus aureus ATCC 25,923, Salmonella typhimurium RSSK 95,091, and Yersinia enterocolitica ATCC 27,729) were determined according Schillinger, Lücke [38] by agar-spot and well diffusion tests. Briefly, 0.5 µl 24-h cultured isolates in MRS was dropped on a MRS agar plates with 4 dots at approximately 30 mm intervals. Plates were incubated under anaerobic conditions (Aneorocult A, Merck) at 30 °C for 24 h. The size of clear zones around the spots was measured. The isolates that gave positive results in the agar spot test incubated for 24 h at 30 °C in MRS broth. The cell free supernatant was separated by centrifugation at 7000 rpm for 10 min, and pH was adjusted to 6.6 by 1 N NaOH. Supernatant was filtered through a cellulose acetate filter (0.45 µm pore size) and used in the well diffusion test.
The antifungal activities of strains were determined against Aspergillus niger and Penicillium chrysogenum origined molded bread. Overnight LAB strain culture was spotted on MRS agar plates in 10 µl volume, under anaerobic conditions. After incubation at 30 °C for 24 h, the petri dishes were overlayed with PDA semi-solid medium (0.8% agar) containing 106 spores/mL. Spore-suspensions were prepared with sterile distilled water from fungal colonies incubated for 7 days in Potato Dextrose Agar (MERCK, Darmstadt) at 30 °C. The antifungal activity was determined by measuring the zone diameters observed in the petri plates that were incubated at 30 °C- 5 days, aerobically [39].
Determination of antioxidative activities of LAB strains
The strains were incubated in MRS medium at 30 °C for 24 h. Supernatant was removed after centrifugation at 2000 g for 5 min. Bacterial cells were re-suspended with distilled water. Antioxidative activity of cells was measured by 1-diphenyl-2-picrylhydrazyl (DPPH) scavenging activity. Two hundred µl of 0.2 mM DPPH solution (with methanol) was mixed with cell suspension. After incubation at dark conditions for 30 min, suspension was filtered through 0.45 µm pore syringe filter (Millipore). Absorbance value of samples was determined at 517 nm. Distilled water was used as blank. Scavenging activity was calculated as inhibition % = (1 − (Asample/Ablank)) *100 [16].
Determination of EPS production level
Isolates, previously growth in MRS broth, were inoculated at a volume of 1% into the 50 mL modified MRS/s medium containing of 100 g/L sucrose [40]. After 72 h incubation, supernatants of the samples (centrifuged for 30 min at 6000 g) and an equal volume of cold ethanol were mixed. The supernatant-ethanol mixture was kept for one night at 4 °C. The mixture (supernatant-ethanol) was centrifuged at 2000 g for 15 min in order to obtain EPS. It was afterwards suspended again with twofold volumes of ethanol and precipitated by centrifuge (at 2000 g for 15 min). Precipitates were dried at 55 °C. The dried samples were diluted with ultrapure water and EPS levels were determined as mg glucose/L by phenol–sulfuric acid method [41].
Statistical analysis
Data analysis was conducted by Minitab.19 package program. The differences between means were tested by Duncan’s multiple range test (p < 0.05). The results of statistical analysis were shown as mean value ± standard deviation in the tables.
Results and discussion
In this study, LAB strains were isolated from Turkish dry fermented sausage (sucuk). Sucuk samples were collected from 3 different producers. LAB counts of sucuk samples were determined and the results are shown in Table 2. The LAB counts of samples differed between 9.72 and 10.33 log cfu/g according to producers. There was no significant difference (p > 0.05) between producers. Total 100 lactic acid bacteria (Gram + , catalase −) isolates were selected for further analysis. With the use of RAPD-PCR analysis in genotypic identification, 10 isolates selected as a result of pre-screening were selected among 100 isolates for techno-functional characterization. Isolates were identified as Lactobacillus plantarum, Pediococcus acidilactici, Weissella hellenica, Lactobacillus pentosus, and Lactobacillus sakei by 16S rRNA gene sequence analysis. In spontaneously fermented sucuks, L plantarum [3, 42], Pediococcus acididilactici [43], and Lactobacillus sakei [44] were isolated as major LAB species. While Lactobacilli are generally included in the major flora, Weissella was detected as minor genera in sucuk [6, 42, 43]. Lücke [11] reported that while L. sakei and L. curvatus were dominant in sausages ripened at low temperatures (20–22 °C), L. plantarum was dominant at high temperatures (> 25 °C). Considering that the samples were collected in summer season (the ambient temperature above 25 °C), it can be said that the high number of isolates, identified as L. plantarum is due to the high ripening temperatures.
The phylogenetic tree obtained by evaluating the 16 s rRNA gene of the isolates is presented in Fig. 1. The isolates constituted 6 subgroups. The cluster analysis showed that 16 s rRNA gene sequences of L. plantarum strains were close to L. pentosus KS-27 strain while Pediococcus acidilactici, Weisella hellenica, and Lactobacillus sakei strains placed separately from other isolates. L. plantarum isolates fell into different subgroups. While L. plantarum KS-12 and KS-25 were close to the L. pentosus KS-27, L. plantarum KS-3 and KS-17 formed different subgroup.
A phylogenetic tree was constructed by using 16S rRNA gene sequences. Phylogenetic tree was created by Neighbor-Joining method (Saitou and Nei, 1987). Phylogenetic distances were calculated for 1200nt. The percentage of replicate trees (1000 replicates) was shown near the branches. Optimal tree with the sum of branch length = 0.20777641 is shown (Felsenstein, 1985). The evolutionary distances were computed using the p-distance method (Nei and Kumar, 2000)
Genes responsible for the production of pediocin, plantaricin, and the gene responsible for GAD production were amplified by PCR with specific primers (Table 3.).
At least one of the genes encoding bacteriocin was found in all isolates. Pediocin and plantaricin genes were not amplified in W. hellenica KS-24. L. plantarum KS-11, L. plantarum subp, plantarum KS-12, and L. pentosus KS-27 contained the gene responsible for the pediocin production. In the well diffusion test, antimicrobial activity was not detected with the neutralized supernatant. The antibacterial effects of isolates on Gram positive (B. cereus, S. aureus) and Gram negative (S. typhimurium, Y. enterolitica, and E. coli) food pathogens in agar-spot test are presented in Table 4. All isolates inhibited the test pathogens at different levels. In fact, W. hellenica KS-24 and L. sakei KS-30 isolates did not inhibit the growth of S. aureus. The highest inhibitions were observed against to Gram negative, S. typhimurium, E. coli, and Y. enterocolitica pathogens, while the effect on B. cereus and S. aureus was minor (negligible small). Similarly, Bartkiene et al. [45] stated in their study that L. sakei and P. acidilactici strains have antimicrobial effects on E. coli, S. typhimurium, Y. enterolitica, B. cereus, and S. aureus strains. Gao et al. [46] reported that many LAB strains have antagonistic effects against some food borne pathogens such as Salmonella species, E. coli, Bacillus cereus, and S. aureus strains. The inhibition effects of lactic acid bacteria on pathogens may be caused by organic acids, as well as by hydrogen peroxide and antimicrobial peptides such as bacteriocins [47]. By evaluating the presence of bacteriocin encoding genes and antimicrobial activity, it can be said that the antimicrobial activity in LAB isolates may be caused by bacteriocin. However, the lack of antimicrobial activity in neutralized supernatants suggested other bacteriostatic effects. It can be said that the antimicrobial activity detected with W. hellenica KS-24 was not due to pediocin or plantaricin.
Another functional feature desired in lactic acid bacteria is antifungal behavior. Fungal growth is an undesirable and frequently encountered problem in many fermented meat products such as sausages [48]. Aspergillus and Penicillium are fungal species that are frequently isolated from meat and meat products [49,50,51]. The antifungal activities of strains are presented in Table 5. L. plantarum KS-17, KS-25, and KS-27 had a strong antifungal effect on A. niger and P. chrysogenum and L. sakei isolates showed weak effect (Table 5). While L. sakei strains exhibited an inhibitory effect on fungal growth, other isolates inhibited both spore formation and mycelial development. Cizeikiene et al. [47] reported that L. sakei KTU05-6 strain inhibited A. niger MD-029 growth, inhibited spore formation of P. acidilactici KTU05-7 and did not affect P. chrysogenum 48-L growth. The antifungal effects of lactic acid bacteria can be caused by many compounds such as organic acids, reuterine, phenyl lactic acid, cyclic dipeptides, fatty acids, and proteinaceous substances [52]. Besides the antibacterial effects of the isolates, the presence of antifungal effects is promising in that they can be used as a protective culture.
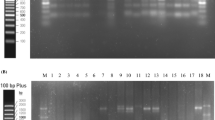
Lactic acid bacteria can produce GABA by decarboxylation of glutamic acid. Core F and Core R primer sets were used in the isolates to determine the presence of GAD enzyme production ability that provides GABA accumulation. All tested strains except W. hellenica KS-24 harbored gad gene responsible for GABA production. It was amplified to approximately 540 bp with Core F and Core R primers. The GABA production capabilities of LAB strains in 53 mM MSG concentration were determined primarily by TLC (Fig. 2). TLC analysis showed KS-20, KS-24, KS-30, and KS-82 isolates did not produce GABA. The presence of GABA in samples obtained using the same medium was quantitatively determined by HPLC analysis. GABA production amounts of strains are given in Table 6. When evaluated by species, it was determined that L. plantarum strains produced high amounts of GABA. Contrary to other studies, it was reported that P. acidilactici KS-20, W. hellenica KS-24, and L.sakei (KS-30, KS-82) were not capable of producing GABA in the presence of 53 mM L-glutamate in accordance with TLC. The highest GABA production was determined to be 1.657 mM with L. plantarum KS-25 (p < 0.05). Lactobacilli are already known as GABA producers [53]. Demirbaş et al. [39] found 4.92 mM GABA production in the medium containing 53 mM MSG with L. plantarum isolate. In another study, Zhang et al. [54] detected GABA production ranging from 3.5 to 14 mM with L. plantarum isolates in the presence of 97 mM MSG. W. hellenica KS-24, which is the only isolate among all the strains that does not contain the GAD gene, did not produce GABA as expected. However, P. acidilactici and L. sakei isolates contained GAD gene in their structures, they did not produce GABA. GABA production levels of the isolates are in similar intervals with other previous studies [55]. Recently, L. plantarum [54] W. hellenica (Barla et al., 2016), P. acidilactici [55], L. sakei [56], and L. pentosus [57] were isolated as GABA producer species from fermented foods. The presence of GABA production ability enables the isolates to be evaluated as functional starter cultures instead of classical starter cultures [55]. In this study, the production abilities were investigated with L-glutamate addition. However, the glutamic acid content of meat offers lactic acid bacteria the possibility of producing GABA. For example, Ratanaburee et al. [17] also showed the presence of GABA in samples that did not add monosodium glutamate in the production of “nham,” a fermented meat product.
Screening of GABA producing LAB isolates by TLC plates (all strains (L. plantarum KS-3, KS-11, KS-17, KS-25, L. plantarum subp. plantarum KS-12, P. acidilactici KS-20, W.hellenica KS-24, L. pentosus KS-27, L.sakei KS-30, KS-82) were grown in MRS broth containing 53 mM MSG for 96 h at 30 °C, GABA GABA standard, MSG MSG standard)
One other biological function of lactic acid bacteria is their antioxidant activity. DPPH radical scavenging activity is one of the most preferred methods for determining antioxidative activity of lactic acid bacteria [58,59,60,61]. Ten LAB isolates were tested for antioxidative abilities. DPPH scavenging activities of supernatant free cells of strains are presented in Table 7. Radical scavenging abilities were not different from each other (p < 0.05). Regardless of species, all isolates showed approximately 20% DPPH scavenging activity. In previous studies, L. plantarum strains showed approximately 10–20% DPPH scavenging activity at 109 cfu/ml concentration [61], P. acidilactici strain showed approximately 40% inhibition at 109 cfu/ml concentration [62]. Contrary to these results, Meira et al. (2012) reported none of studied Lactobacillus species showed DPPH scavenging activity (60 µM). Considering the studies, it can be said that LAB isolates show a moderate antioxidant activity. Antioxidant activity demonstrated by LAB cells can be due to cell surface proteins [61] or exopolysaccharides (EPS) [63]. EPS can exhibit bioactive properties (antiviral, anticarcinogen, antioxidative, etc.) and can contribute to probiotic activity by providing colonization [64, 65]. There are many microorganisms reported as EPS producers [66,67,68]. Since most LAB species have been generally recognized as safe (GRAS) status [69], species that can produce EPS attract attention. The fact that EPS producer LAB isolates improve the texture properties of meat products [26] and delay color and oxidation [25] show that EPS production is also an important factor technologically. EPS production levels determined after 72 h incubation of strains in sucrose containing medium are indicated in Table 8. L. plantarum KS-11 (991.75 ± 94.81) and L. pentosus KS-27 (991.75 ± 37.55) exhibited the highest EPS production and lower EPS production levels were determined with KS-20, KS-24, KS-30, and KS-82 strains. L plantarum [70] and L. pentosus (Rodríguez-Carvajal et al., 2008) are lactobacilli that are commonly capable of producing EPS. L. plantarum strains provided higher EPS production in this study. Mıdık et al. [71] determined 512.81 mg/L EPS production with L. plantarum isolate. Their LABs show EPS production at low levels (50–400 mg/L) [72] as well as at higher levels (10.78 g/L) in optimized conditions such as L. rhamnosus strain [73]. EPS production is also found in the literature.
Conclusion
In this study, 10 lactic acid bacteria isolated from sucuk were genotypically identified. Furthermore, some functional properties of the isolates were evaluated. It was observed that some isolates are capable of producing GABA, which is known to have positive effects on human health. The highest GABA production was achieved with L. plantarum KS-25 strain. Additionally, it was determined that a significant part of the isolates is capable of producing EPS and all isolates show antioxidant activity. Most of the isolates inhibited five food-borne pathogens, while all isolates showed antagonistic effect on A. niger and P. chrysogenum. In these isolates, substantially antifungal, antibacterial, antioxidant activity, and ability to produce EPS and GABA have been determined, although it varies according to the strain representing some technological and functional properties of autochthonous LAB strains from sucuk. Further work will investigate these functional properties of LAB isolates in vitro conditions such as fermented sausages and meat products.
References
Erkmen O, Bozkurt H (2004) Quality characteristics of retailed sucuk (Turkish dry-fermented sausage). Food Technol Biotechnol 42(1):63–69
Erginkaya Z, Yalanca İ, Ünal Turhan E (2019) Geleneksel et ürünlerindeki laktik asit bakterilerinin antibiyotik direnç profili. Pamukkale University J Eng Sci 25 (7):834–838
Kaban G, Kaya M (2008) Identification of lactic acid bacteria and Gram-positive catalase-positive cocci isolated from naturally fermented sausage (sucuk). J Food Sci 73(8):M385–M388
Drosinos EH, Paramithiotis S, Kolovos G, Tsikouras I, Metaxopoulos I (2007) Phenotypic and technological diversity of lactic acid bacteria and staphylococci isolated from traditionally fermented sausages in Southern Greece. Food Microbiol 24(3):260–270
Papamanoli E, Tzanetakis N, Litopoulou-Tzanetaki E, Kotzekidou P (2003) Characterization of lactic acid bacteria isolated from a Greek dry-fermented sausage in respect of their technological and probiotic properties. Meat Sci 65(2):859–867
Kesmen Z, Yetiman AE, Gulluce A, Kacmaz N, Sagdic O, Cetin B, Adiguzel A, Sahin F, Yetim H (2012) Combination of culture-dependent and culture-independent molecular methods for the determination of lactic microbiota in sucuk. Int J Food Microbiol 153(3):428–435. https://doi.org/10.1016/j.ijfoodmicro.2011.12.008
Çon AH, Gökalp HY (2000) Production of bacteriocin-like metabolites by lactic acid cultures isolated from sucuk samples. Meat Sci 55(1):89–96
Ammor MS, Mayo B (2007) Selection criteria for lactic acid bacteria to be used as functional starter cultures in dry sausage production: an update. Meat Sci 76(1):138–146. https://doi.org/10.1016/j.meatsci.2006.10.022
Aymerich T, Martin B, Garriga M, Hugas M (2003) Microbial quality and direct PCR identification of lactic acid bacteria and nonpathogenic staphylococci from artisanal low-acid sausages. Appl Environ Microbiol 69(8):4583–4594
Lücke F-K (2000) Utilization of microbes to process and preserve meat. Meat Sci 56(2):105–115
Lücke F (1985) Mikrobiologische Vorgänge bei der Herstellung von Rohwurst und Rohschinken. Mikrobiologie und qualität von rohwurst und rohschinken 1:85-102
Getty KJK, Phebus RK, Marsden JL, Fung DYC, Kastner CL (2000) escherichia coli O157:H7 and fermented sausages: a review1. J Rapid Methods Autom Microbiol 8(3):141–170. https://doi.org/10.1111/j.1745-4581.2000.tb00215.x
Dalié D, Deschamps A, Richard-Forget F (2010) Lactic acid bacteria–Potential for control of mould growth and mycotoxins: a review. Food Control 21(4):370–380
Kamiloğlu A, Kaban G, Kaya M (2019) Effects of autochthonous Lactobacillus plantarum strains on Listeria monocytogenes in sucuk during ripening. J Food Saf 39(3):e12618
Smaoui S, Elleuch L, Bejar W, Karray-Rebai I, Ayadi I, Jaouadi B, Mathieu F, Chouayekh H, Bejar S, Mellouli L (2010) Inhibition of fungi and gram-negative bacteria by bacteriocin BacTN635 produced by Lactobacillus plantarum sp. TN635. Appl Biochem Biotechnol 162(4):1132–1146
Zhang S, Liu L, Su Y, Li H, Sun Q, Liang X, Lv J (2011) Antioxidative activity of lactic acid bacteria in yogurt. Afr J Microbiol Res 5(29):5194–5201
Ratanaburee A, Kantachote D, Charernjiratrakul W, Sukhoom A (2013) Selection of γ-aminobutyric acid-producing lactic acid bacteria and their potential as probiotics for use as starter cultures in T hai fermented sausages (N ham). Int J Food Sci Technol 48(7):1371–1382
Mody I, De Koninck Y, Otis TS, Soltesz I (1994) Bridging the cleft at GABA synapses in the brain. Trends Neurosci 17(12):517–525
Sarasa SB, Mahendran R, Muthusamy G, Thankappan B, Selta DRF, Angayarkanni J (2020) A brief review on the non-protein amino acid, gamma-amino butyric acid (GABA): its production and role in microbes. Curr Microbiol 77(4):534–544. https://doi.org/10.1007/s00284-019-01839-w
Shimada M, Hasegawa T, Nishimura C, Kan H, Kanno T, Nakamura T, Matsubayashi T (2009) Anti-hypertensive effect of γ-aminobutyric acid (GABA)-rich Chlorella on high-normal blood pressure and borderline hypertension in placebo-controlled double blind study. Clin Exp Hypertens 31(4):342–354
Li H, Cao Y (2010) Lactic acid bacterial cell factories for gamma-aminobutyric acid. Amino Acids 39(5):1107–1116
Yogeswara IBA, Maneerat S, Haltrich D (2020) A brief review on glutamate decarboxylase from lactic acid bacteria—A Key Enzyme in GABA Synthesis. Microorganisms 8:1923
Laws A, Gu Y, Marshall V (2001) Biosynthesis, characterisation, and design of bacterial exopolysaccharides from lactic acid bacteria. Biotechnol Adv 19(8):597–625
Xu Y, Cui Y, Yue F, Liu L, Shan Y, Liu B, Zhou Y, Lü X (2019) Exopolysaccharides produced by lactic acid bacteria and Bifidobacteria: structures, physiochemical functions and applications in the food industry. Food Hydrocolloids 94:475–499
Trabelsi I, Ktari N, Triki M, Bkhairia I, Slima SB, Aydi SS, Aydi S, Abdeslam A, Salah RB (2018) Physicochemical, techno-functional, and antioxidant properties of a novel bacterial exopolysaccharide in cooked beef sausage. Int J Biol Macromol 111:11–18
Dertli E, Yilmaz MT, Tatlisu NB, Toker OS, Cankurt H, Sagdic O (2016) Effects of in situ exopolysaccharide production and fermentation conditions on physicochemical, microbiological, textural and microstructural properties of Turkish-type fermented sausage (sucuk). Meat Sci 121:156–165
Patel S, Majumder A, Goyal A (2012) Potentials of exopolysaccharides from lactic acid bacteria. Indian J Microbiol 52(1):3–12
Çon AH, Gökalp HY, Kaya M (2001) Antagonistic effect on Listeria monocytogenes and L. innocua of a bacteriocin-like metabolite produced by lactic acid bacteria isolated from sucuk. Meat Sci 59(4):437–441
Toksoy A, Beyatli Y, Aslim B (1999) Studingon metabolic and antimicrobial activities of some L plantarum strains isolated from sausages. Turk J Vet Anim Sci 23(6):533–540
Dertli E, Mercan E, Arıcı M, Yılmaz MT, Sağdıç O (2016) Characterisation of lactic acid bacteria from Turkish sourdough and determination of their exopolysaccharide (EPS) production characteristics. LWT-Food Sci Technol 71:116–124
Barış Ö (2009) Erzurum İlindeki Mağaralarda Damlataşı Oluşumunda Etkili Bakterilerin İzolasyonu Karakterizasyonu Ve Tanısı. Fen Bilimleri Enstitüsü, Doktora Tezi, Erzurum:135
Baker G, Smith JJ, Cowan DA (2003) Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55(3):541–555
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549
Park K-B, Oh S-H (2007) Cloning, sequencing and expression of a novel glutamate decarboxylase gene from a newly isolated lactic acid bacterium, Lactobacillus brevis OPK-3. Biores Technol 98(2):312–319
Villegas JM, Brown L, de Giori GS, Hebert EM (2016) Optimization of batch culture conditions for GABA production by Lactobacillus brevis CRL 1942, isolated from quinoa sourdough. LWT-Food Sci Technol 67:22–26
Tuberoso CIG, Congiu F, Serreli G, Mameli S (2015) Determination of dansylated amino acids and biogenic amines in Cannonau and Vermentino wines by HPLC-FLD. Food Chem 175:29–35
Schillinger U, Lücke FK (1989) Antibacterial activity of Lactobacillus sake isolated from meat. Appl Environ Microbiol 55(8):1901–1906
Demirbaş F, İspirli H, Kurnaz AA, Yilmaz MT, Dertli E (2017) Antimicrobial and functional properties of lactic acid bacteria isolated from sourdoughs. LWT Food Sci Technol 79:361–366. https://doi.org/10.1016/j.lwt.2017.01.067
van Geel-Schutten G, Flesch F, Ten Brink B, Smith M, Dijkhuizen L (1998) Screening and characterization of Lactobacillus strains producing large amounts of exopolysaccharides. Appl Microbiol Biotechnol 50(6):697–703
Dubois M, Gilles KA, Hamilton JK, Pt R, Smith F (1956) Colorimetric method for determination of sugars and related substances. Anal Chem 28(3):350–356
Adiguzel G, Atasever M (2009) Phenotypic and genotypic characterization of lactic acid bacteria isolated from Turkish dry fermented sausage. Rom Biotechnol Lett 14(1):4130–4138
Yüceer Ö, Özden Tuncer B (2015) Determination of antibiotic resistance and biogenic amine production of lactic acid bacteria isolated from fermented Turkish sausage (Sucuk). J Food Saf 35(2):276–285. https://doi.org/10.1111/jfs.12177
Özdemir F, Nadeem HŞ, Akdoğan A, Dinçer C, Topuz A (2018) Effect of altitude, shooting period, and tea grade on the catechins, caffeine, theaflavin, and thearubigin of Turkish black tea. Turk J Agric For 42(5):334–340
Bartkiene E, Bartkevics V, Mozuriene E, Krungleviciute V, Novoslavskij A, Santini A, Rozentale I, Juodeikiene G, Cizeikiene D (2017) The impact of lactic acid bacteria with antimicrobial properties on biodegradation of polycyclic aromatic hydrocarbons and biogenic amines in cold smoked pork sausages. Food Control 71:285–292. https://doi.org/10.1016/j.foodcont.2016.07.010
Gao Z, Daliri EB-M, Wang J, Liu D, Chen S, Ye X, Ding T (2019) Inhibitory effect of lactic acid bacteria on foodborne pathogens: a review. J Food Prot 82(3):441–453
Cizeikiene D, Juodeikiene G, Paskevicius A, Bartkiene E (2013) Antimicrobial activity of lactic acid bacteria against pathogenic and spoilage microorganism isolated from food and their control in wheat bread. Food Control 31(2):539–545
Soncu ED, Arslan B, Ertürk D, Küçükkaya S, Özdemir N, Soyer A (2018) Microbiological, physicochemical and sensory characteristics of Turkish fermented sausages (sucuk) coated with chitosan-essential oils. LWT 97:198–204
Andersen S, Frisvad J (1994) Penicillin production by Penicillium nalgiovense. Lett Appl Microbiol 19(6):486–488
Matos T, Jensen B, Bernardo F, Barreto A, Hojberg Ø (2007) Mycoflora of two types of Portuguese dry-smoked sausages and inhibitory effect of sodium benzoate, potassium sorbate, and methyl p-hydroxybenzoate on mold growth rate. J Food Prot 70(6):1468–1474
Zohri A, Moharram A, Refaie R (2014) Mycobiota contaminating beef burger and sausage with reference to their toxins and enzymes. J Basic Appl Mycol (Egypt) 5:61
Crowley S, Mahony J, van Sinderen D (2013) Current perspectives on antifungal lactic acid bacteria as natural bio-preservatives. Trends Food Sci Technol 33(2):93–109
Thwe SM, Kobayashi T, Luan T, Shirai T, Onodera M, Hamada-Sato N, Imada C (2011) Isolation, characterization, and utilization of γ-aminobutyric acid (GABA)-producing lactic acid bacteria from Myanmar fishery products fermented with boiled rice. Fish Sci 77(2):279–288
Zhang Q, Zeng L, Tan X, Tang J, Xiang W (2017) An efficient γ-aminobutyric acid (GABA) producing and nitrite reducing ability of Lactobacillus plantarum BC114 isolated from Chinese Paocai. Food Sci Technol Res 23(5):749–755
Yu H-H, Choi JH, Kang KM, Hwang H-J (2017) Potential of a lactic acid bacterial starter culture with gamma-aminobutyric acid (GABA) activity for production of fermented sausage. Food Sci Biotechnol 26(5):1333–1341
Kook M-C, Seo M-J, Cheigh C-I, Pyun Y-R, Cho S-C, Park H (2010) Enhanced production of ${\gamma} $-aminobutyric acid using rice bran extracts by Lactobacillus sakei B2–16. J Microbiol Biotechnol 20(4):763–766
Kim M-J, Kim K-S (2012) Isolation and identification of γ-aminobutyric acid (GABA)-producing lactic acid bacteria from Kimchi. J Korean Soc Appl Biol Chem 55(6):777–785
Ayeni FA, Sánchez B, Adeniyi BA, Clara G, Margolles A, Ruas-Madiedo P (2011) Evaluation of the functional potential of Weissella and Lactobacillus isolates obtained from Nigerian traditional fermented foods and cow’s intestine. Int J Food Microbiol 147(2):97–104
Jain S, Verma R, Murdia L, Jain H, Sharma G (2011) Optimization of process parameters for osmotic dehydration of papaya cubes. J Food Sci Technol 48(2):211–217
Kang C-H, Han SH, Kim J-S, Kim Y, Jeong Y, Park HM, Paek N-S (2019) Inhibition of nitric oxide production, oxidative stress prevention, and probiotic activity of lactic acid bacteria isolated from the human vagina and fermented food. Microorganisms 7(4):109
Li S, Zhao Y, Zhang L, Zhang X, Huang L, Li D, Niu C, Yang Z, Wang Q (2012) Antioxidant activity of Lactobacillus plantarum strains isolated from traditional Chinese fermented foods. Food Chem 135(3):1914–1919
Chen Q, Kong B, Sun Q, Dong F, Liu Q (2015) Antioxidant potential of a unique LAB culture isolated from Harbin dry sausage: In vitro and in a sausage model. Meat Sci 110:180–188
Pan D, Mei X (2010) Antioxidant activity of an exopolysaccharide purified from Lactococcus lactis subsp. lactis 12. Carbohydr Polym 80(3):908–914
Abid Y, Casillo A, Gharsallah H, Joulak I, Lanzetta R, Corsaro MM, Attia H, Azabou S (2018) Production and structural characterization of exopolysaccharides from newly isolated probiotic lactic acid bacteria. Int J Biol Macromol 108:719–728
Garai-Ibabe G, Areizaga J, Aznar R, Elizaquivel P, Prieto A, Irastorza A, Dueñas MT (2010) Screening and selection of 2-branched (1, 3)-β-D-glucan producing lactic acid bacteria and exopolysaccharide characterization. J Agric Food Chem 58(10):6149–6156
Biswas J, Paul A (2017) Diversity and production of extracellular polysaccharide by halophilic microorganisms. Biodiversity Int J 1(2):00006
Nouha K, RD T, RY S (2015) EPS producing microorganisms from municipal wastewater activated sludge. J Pet Environ Biotechnol 7:255
Poli A, Di Donato P, Tommonaro G, Abbamondi GR, Finore I, Nicolaus B (2018) Exopolysaccharide-producing microorganisms from extreme areas: chemistry and application. In: Extremophiles in Eurasian ecosystems: ecology, diversity, and applications. Springer, pp 405–433
Rodríguez-Sánchez S, Ramos IM, Seseña S, Poveda JM, Palop ML (2021) Potential of Lactobacillus strains for health-promotion and flavouring of fermented dairy foods. LWT 143:111102
Gangoiti MV, Puertas A, Hamet MF, Peruzzo PJ, Llamas M, Medrano M, Prieto A, Dueñas MT, Abraham AG (2017) Lactobacillus plantarum CIDCA 8327: an α-glucan producing-strain isolated from kefir grains. Carbohyd Polym 170:52–59
Mıdık F, Tokatlı M, Bağder Elmacı S, Özçelik F (2020) Influence of different culture conditions on exopolysaccharide production by indigenous lactic acid bacteria isolated from pickles. Arch Microbiol 202(4):875–885. https://doi.org/10.1007/s00203-019-01799-6
Cirrincione S, Breuer Y, Mangiapane E, Mazzoli R, Pessione E (2018) ‘Ropy’ phenotype, exopolysaccharides and metabolism: study on food isolated potential probiotics LAB. Microbiol Res 214:137–145. https://doi.org/10.1016/j.micres.2018.07.004
Bhati A, Baghel AK, Singhal B (2021) Optimization of culture conditions for EPS production in Lactobacillus rhamnosus MTCC 5462 through Taguchi design methodology. In: Proceedings of International Conference on Scientific and Natural Computing. Springer, pp 253–260
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Rosane Freitas Schwan
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kamiloğlu, A. Functional and technological characterization of lactic acid bacteria isolated from Turkish dry-fermented sausage (sucuk). Braz J Microbiol 53, 959–968 (2022). https://doi.org/10.1007/s42770-022-00708-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-022-00708-2