Abstract
The objective of this research was to explore the candidate genes involved in calcium absorption of R. roxburghii plants, as well as their relationship with the bioactive substance accumulation in the fruit. RNA-seq and qRT-PCR were used to analyze the transcriptomic profiles of the roots and leaves of plants during calcium absorption. Based on the correlation between gene expression and calcium absorption rate, ion channels and carrier proteins were selected to further verify their response to exogenous calcium supplement and functions on the accumulation of ascorbate (AsA), flavonoid, and triterpenoid in the fruit. Transcriptome and qRT-PCR analysis revealed that a total of 25 candidate genes from seven families have been found to be closely associated with Ca2+ absorption, namely, cyclic nucleotide-gated ion channel (CNGC), zinc transporter (ZTP), metal tolerance protein (MTP), ER-type Ca2+ ATPase (ECA), auto-inhibited Ca2+ ATPase (ACA), glutamate receptor (GLR), and natural resistance-associated macrophage protein (NRAMP). Among these, RrCNGC1/7/10/12 and RrMTP2 play a vital role in responding to changes in external calcium levels. Moreover, exogenous Ca2+ promoted the accumulation of bioactive compounds such as AsA, total flavonoid, and total triterpenoid in fruits. RrCNGC2/3/12, RrZTP1, and RrGLR1 significantly responded to exogenous Ca2+ supplement, of which RrCNGC12 positively correlating with the levels of the above three types of bioactive substances. Seven families of ion channels and carrier proteins were jointly involved in calcium absorption and homeostasis in R. roxburghii plants, and RrCNGC12 play a critical role in the calcium-mediated regulation of fruit quality formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Calcium (Ca2+) is an essential nutrient for plant growth and a second messenger in plants. The cytosolic concentration of Ca2+ ([Ca2+]cyt) significantly influences crop growth and development (Hirschi 2004). Various environmental stimuli can cause changes of [Ca2+]cyt, and the increase of free Ca2+ in cellular is an early and crucial event of plant defense signaling (Lecourieux et al. 2006). Studies have shown that Ca2+ can promote the synthesis of multiple secondary metabolites by transmitting secondary signals to downstream target enzymes and thus enhances plant resistance to adverse external environments (Zhao et al. 2005; Ahmad et al. 2016; Martins et al. 2021; Michailidis et al. 2019; Juric et al. 2020). For example, melatonin induces the production of phenolic compounds by causing the influx of extracellular Ca2+ into cells and activating Ca2+/calmodulin (CaM) signaling (Vafadar et al. 2020). Under cold stress, Ca2+ strengthen the cold resistance of plant by maintaining soluble sugar and protein content in cells and increasing the content of unsaturated fatty acid in membrane (Shi et al. 2017). Wang et al. (2022) discovered that CaM and calmodulin-like (CML) genes contribute to improving drought resistance in maize. Therefore, the level of Ca2+ in the cytoplasm and the combination of Ca2+ sensors and downstream proteins participate in the growth and development process of plants.
Plant signals and responses are related to variations in [Ca2+]cyt, which is tightly regulated by a network of channels and carriers responsible for Ca2+ homeostasis and the production of Ca2+ signals (Tang and Luan 2017). These carriers and channels include passive fluxes (Ca2+ channels) and active transport (Ca2+-ATPases and Ca2+-antiporters) across the plasma membrane or endomembranes (Sanders et al. 1999, 2002; Demidchik et al. 2018; Zhang et al. 2019). A variety of Ca2+ channels have been found in plant cell membranes, primarily divided into three categories: depolarization-activated Ca2+-permeable channels (DACCs) and hyperpolarization-activated Ca2+-permeable channels (HACCs), and nonselective cation channels (NSCCs). Among these, HACCs in root hairs are highly selective for Ca2+ (Véry and Davies 2000), although some studies suggest that plant Ca2+ conductances are primarily mediated by NSCCs (Liu et al. 2018). These ion channels are closely related to the production of calcium signals and response to abiotic stress. Gao et al. (2016) reported that R491Q or R578K point mutations in cyclic nucleotide-gated channel 18 (CNGC18) resulted in abnormal Ca2+ gradients and defects in pollen tube guidance by impairing the activity of CNGC18 in Arabidopsis. The upregulation of Ca2+ influx channel protein mid1-complementing activity 1 (MCA1) gene and the downregulation of efflux channel protein cation exchanger 1 (CAX) and autoinhibited Ca2+-ATPase 1 (ACA1) genes in the cytoplasmic membrane jointly facilitated the increase of Ca2+ in the cytoplasm (Yang et al. 2022). In conclusion, channels and carriers of calcium ions and other cations jointly mediate the transport of Ca2+.
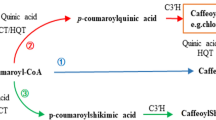
Rosa roxburghii Tratt. is a perennial shrub of the Rosaceae family, which is widely distributed in the karst region of southwest China with calcium-rich soil. The fruits of this species are valued for their nutritional and medicinal characteristics, especially their high ascorbic acid (AsA), triterpenoid, and flavonoid levels, and therefore believed to have valuable senescence-retarding and cancer-preventing effects (Ojo et al. 2022). Various studies have shown that Ca2+ plays a vital regulatory role in the growth and the formation of quality of R. roxburghii fruits (Luo et al. 2004). Increasing exogenous Ca2+ can induce the expression of GalLDH and GPP genes involved in the AsA synthesis pathway, as well as DHAR and MDHAR genes involved in the AsA regeneration pathway (Li and An 2016). This process promotes the synthesis and accumulation of AsA in R. roxburghii fruits (Zhang et al. 2012). Ca2+ needs to rely on the coordinated transport of ion channels and carriers to function, but the absorption and transport mechanism of Ca2+ in R. roxburghii have not been thoroughly studied. In this study, we analyzed the transcriptome characteristics during calcium absorption in R. roxburghii cv. ‘guinong 5’ using RNA-Seq technology and screened for candidate carrier or channel protein related to calcium absorption. And we also explored their possible relationship with the accumulation of main bioactive substances (AsA, flavonoids, and triterpenoids) in fruits. The results of this experiment have provided valuable candidate genes to help understand the calcium absorption mechanism of R. roxburghii and explored its relationship with the accumulation of bioactive substances.
2 Material and Methods
2.1 Material Selection and Cultivation
The R. roxburghii cv. ‘guinong 5’ saplings propagated from cuttings were planted in R. roxburghii resource nursery of Guizhou University. Saplings with 3-month age which showed similar growth and size were selected and transferred to a 1/8 concentration of Hoagland and Aron nutrient solution from soil for pre-cultivation. The nutrient solution was changed every 3 days, and its pH value was adjusted to 6.5 using 0.1 mol L−1 H2SO4 or NaOH. The culture temperature was maintained at 20 ℃.
2.2 Determination of Different Periods of Ca 2+ Uptake in R. roxburghii Plants
Yang and Fan (2022) found that the growth of R. roxburghii plants was most optimal at the Ca2+ concentration of 50–100 mg L−1. After conducting pre-experiments (Fig. S1), we selected the Ca2+ concentration of 2 mmol L−1 for the uptake experiment. The saplings were starved in deionized water for 24 h after being pre-cultured for 2 months. Following this, they were transferred to an culture solution consisting of 2 mmol L−1 Ca(CH3COO)2. Two time intervals, either every 1 h or every 10 min, were used to change the culture solution and determine the concentration of Ca2+ in the culture solution. This was conducted to determine the rapid Ca2+ uptake period and Ca2+ saturation period of the saplings. The uptake rate (μg g−1 h−1) was calculated using the equation: reduced Ca2+ concentration (μg mL−1) × total volume of nutrient solution (mL) / (fresh root weight (g) × time (h)).
2.3 Transcriptome Sequencing Analysis
2.3.1 Sampling and RNA Extraction
At the Ca2+ starvation, rapid Ca2+ uptake, and Ca2+ saturation stages, white aquatic roots and leaves of R. roxburghii saplings were collected separately to extract total RNA for transcriptome sequencing and qRT-PCR. Three biological replicates were used. The collected samples were stored at – 80 °C after being treated with liquid nitrogen for the extraction of total RNA. Each RNA sample was divided into two aliquots: one for RNA-seq and the other for qRT-PCR.
2.3.2 Sequencing and Quality Control
mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in NEBNext First Strand Synthesis Reaction Buffer (5X). First-strand cDNA was synthesized using random hexamer primer and M-MuLV Reverse Transcriptase. Second-strand cDNA synthesis was subsequently performed using DNA Polymerase I and RNase H. The library fragments were purified with AMPure XP system (Beckman Coulter, Beverly, USA). Then 3 μl USER Enzyme (NEB, USA) was used with size-selected, adaptor-ligated cDNA at 37 °C for 15 min followed by 5 min at 95 °C before PCR. Then PCR was performed with Phusion High-Fidelity DNA polymerase, Universal PCR primers and Index (X) Primer. At last, PCR products were purified (AMPure XP system) and library quality was assessed on the Agilent Bioanalyzer 2100 system. The clustering of the index-coded samples was performed on a cBot Cluster Generation System using TruSeq PE Cluster Kit v4-cBot-HS (Illumia) according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina platform and paired-end reads were generated.
The raw reads were further processed with a bioinformatic pipeline tool, BMKCloud (www.biocloud.net) online platform. Raw reads of fastq format were firstly processed through fastp 0.21.0 (Chen et al. 2018). In this step, clean reads were obtained by removing reads containing adapter, reads containing ploy-N, and low-quality reads from raw data. At the same time, Q20, Q30, GC-content, and sequence duplication level of the clean data were calculated. All the downstream analyses were based on clean data with high quality. These clean reads were then mapped to the reference genome sequence. Hisat2 (Kim et al. 2015) tools soft were used to map with reference genome. Using StringTie (Pertea et al. 2015) to assemble reads been compared.
2.3.3 Gene Functional Annotation
Gene function was annotated using DIAMOND (Buchfink et al. 2015) based on the following databases: Nr (NCBI non-redundant protein sequences) (Deng et al. 2006); Pfam (Protein family) (Finn et al. 2014); KOG/COG (Clusters of Orthologous Groups of proteins) (Tatusov et al. 2000; Koonin et al. 2004); Swiss-Prot (A manually annotated and reviewed protein sequence database) (Apweiler et al. 2004); KO (KEGG Ortholog database) (Kanehisa et al. 2004); GO (Gene Ontology) (Ashburner et al. 2000).
2.3.4 Screening of Differentially Expressed Genes (DEGs) and Its GO and KEGG Enrichment Analysis
FPKM (fragments per kilobase of transcript per million fragments mapped) was used as a measure of transcript or gene expression level. DESeq2 software was used to perform differential gene expression analysis (Love et al. 2014), with DEGs screened based on the criteria of fold change ≥ 1.2 and FDR < 0.01. FDR was obtained by correcting the difference significance p-value. DEGs were annotated to the GO database, and their functional distribution statistics were counted, analyzed, and plotted (Young et al. 2010). The DEGs were also compared with the KEGG database to investigate the metabolic pathways and signaling pathways in which they are involved (Kanehisa et al. 2008).
2.4 R. roxburghii Saplings Treated with Different Levels of Calcium Supply
The 1/8 concentration of Hoagland and Arnon formula was used as the basic nutrient solution in the experiment, with slight modifications made to remove all Ca2+-containing components and to equalize nitrogen differences with 40 mg L−1 NH4NO3, while keeping other element concentrations unchanged. Calcium concentrations were controlled through Ca(CH3COO)2. According to the research of Yang and Fan (2022), we designed three calcium concentration treatments, namely 0 mmol L−1, 0.5 mmol L−1, and 2 mmol L−1. The pH of nutrient solution were adjusted to 6.5 using 0.1 mol L−1 NaOH and H2SO4. The cultivation temperature was 20 °C. R. roxburghii saplings were starved for 24 h and then transferred to the three nutrient solutions. Hydroponic roots and leaves of saplings in different treatments were collected after 0 day, 1 day, and 7 days of cultivation, respectively. A portion is dried to constant weight and stored at ordinary temperature, while another portion is treated with liquid nitrogen and stored at − 80 °C. The calcium content and the expression of candidate genes in these samples were determined. Each treatment was replicated biologically three times.
2.5 The Spatiotemporal Expressive Specificity of Candidate Genes
Excellent perennial asexual lines of R. roxburghii cv. ‘guinong 5’ were selected as test materials. Fruits, petals, and leaves from the upper and middle portions of the crown at 30 days after anthesis were used as samples for analyzing of the expression patterns in various organs. Fruit samples were taken at 30, 60, and 90 days after anthesis, at 10 am, with at least 300 g of fruits collected each time. The expression of candidate genes was determined using qRT-PCR in different developmental stages of fruits and in different tissue samples.
2.6 Spraying Plant Crown In Vivo with Exogenous Ca 2+
Good asexual lines of perennial R. roxburghii were sprayed with Ca2+ solution on the plant crown at 20 days, 50 days, and 80 days after anthesis. Thirty fruits were sampled at 30 days (30 May), 60 days (30 June), and 90 days (30 July) after anthesis. A portion is dried to constant weight and stored at ordinary temperature, while another portion is treated with liquid nitrogen and stored at − 80 °C. We set up four concentration treatments by Ca(CH3COO)2 solution, namely 100 mg L−1, 300 mg L−1, 500 mg L−1, and 700 mg L−1, with clear water as the control (Luo et al. 2004), and five trees grown in each treatment. The content of calcium, AsA, total flavonoid, and total triterpenoid content, as well as the expression of candidate genes, were determined in the samples from different treatments. Three biological replicates were analyzed for each indicator.
2.7 Soaking Fruit In Vitro with Exogenous Ca 2+
The R. roxburghii fruits of uniform development were collected in late July. According to the method of Zhang et al. (2012), each fruit was divided into three portions and placed in a culture solution with the following formulation: 2 mmol L−1 CaSO4 + 15 mmol L−1 glucose, 2 mmol L−1 CaSO4 + 15 mmol L−1 glucose + 5 mmol L−1 EGTA, and 15 mmol L−1 glucose as a control. Samples were taken after incubation in a light incubator for 0, 3, 6, 12, and 24 h, during which 54 μmol m−2 s−1 of light was provided from 8:00 to 19:00 at 25 ℃, with three biological replicates. The samples were taken to determine the content of calcium, AsA, total flavonoid, and total triterpenoid, as well as the expression of candidate genes, and the determination was repeated three times.
2.8 Measurement of Calcium, AsA, Total Flavonoid, and Total Triterpenoid
2.8.1 Determination of Calcium Content
The calcium content was determined by the HNO3-HClO4 digestion method and inductively coupled plasma emission spectrometry (ICP-AES). The samples were baked at 110 °C for 0.5 h and then placed in a thermostat at 70 °C until their weight became constant. The dried samples were ground and passed through a 100 mesh nylon sieve. The sample was weighed 0.5 g, incubated overnight in 15 mL of mixed acid (HNO3/HClO4 = 9:1, v/v), heated on an electric hot plate until white smoke was emitted, and continued to heat until about 1 mL of digest remained. After cooling, the digest was transferred to a clean volumetric flask, fixed to 50 mL, filtered, and the calcium content was determined by ICPE-9800 inductively coupled plasma emission spectrometer. Element content (mg g−1) was calculated as (D − B) × V/(W × 1000), where D is the detection value of the sample element concentration (mg L−1), B is the blank detection value (mg L−1), V is the total volume of the sample after digestion (mL), and W is the sample weight (g). All tests were performed in three biological and technical replicates.
2.8.2 Determination of AsA Content
Liquid chromatography was used for the determination of AsA content (Wang and An 2013). The tissue samples were ground evenly with 5 mL of 6% metaphosphoric acid solution in a pre-chilled mortar. The solution was then transferred to a 10-mL centrifuge tube and centrifuged for 15 min at 4 °C and 10,000 r/min. The supernatant was removed to a 10-mL centrifuge tube, and 3–4 mL of extract was added to the remaining residue, which was then centrifuged for an additional 10 min under the same conditions. The supernatant was removed and combined, and the volume was made constant to 10 mL. This solution was passed through a 0.45-μm filter membrane to obtain the AsA extract for determination. The AsA content was determined by high-performance liquid chromatography (HPLC) under the following conditions: Wondasil C18 column (4.6 mm × 150 mm, 5 μm); mobile phase, 0.2% metaphosphoric acid solution; flow rate, 1 mL/min; column temperature, 30 ℃; UV detector, 254 nm; injection volume, 20 μL. The AsA content was calculated based on the peak value of the HPLC spectrum (Fig. S2) using the following formula: AsA content (mg/100 g) = (m1 × v2 × 10−6 × 100)/ (m × v1), where m is the mass of the sample weighed in grams; m1 is the mass of AsA in the solution to be measured (g), calculated using the standard curve; v1 is the volume of the extraction solution used for determination in the sample (mL); and v2 is the total volume of the sample presented (mL).
2.8.3 Determination of Total Triterpenoid Content
A standard curve was prepared using ursolic acid as the standard. The tissue sample weighing 0.5 g was added to 8 mL of 75% ethanol solution, then centrifuged at 7000 r/min for 10 min, after 50 min of ultrasonic extraction at 50 ℃. The supernatant was aspirated and fixed to a volume of 10 mL, after which it was shaken well. For determination, 0.2 mL of the extract was taken, and 0.5 mL of 5% vanillin-acetic acid solution and 0.8 mL of perchloric acid were added. The resulting mixture was shaken well and heated in a water bath at 60 ℃ for 20 min, and then immediately cooled in ice water. Finally, 3.5 mL of glacial acetic acid was added, shaken well and the absorbance was measured at a wavelength of 545 nm. The total triterpenoid content was calculated using the following formula: total triterpenoid content (mg/100 g) = (m1 × v2 × 10−6 × 100)/ (m × v1), where m is the mass of the sample weighed in grams; m1 is the mass of total triterpenoid in the solution to be measured (g), calculated using the standard curve; v1 is the volume of the extraction solution used for determination in the sample (mL); and v2 is the total volume of the sample presented (mL).
2.8.4 Determination of Total Flavonoid Content
The NaNO2-Al(NO3)3 colorimetric method was employed to determine the total flavonoid content. The rutin standard was used to prepare the standard curve. 0.5 g of the sample was weighed and sonicated for 2 h in 15 mL of 30% methanol. The mixture was then centrifuged at 8000 r/min for 10 min, and the supernatant was extracted. The extracted supernatant was fixed to a volume of 10 mL, after which 2 mL of the extract was pipetted into a 10-mL centrifuge tube. To this solution, 0.4 mL of 5% NaNO2, 0.6 mL of 10% Al (NO3)3, and 4 mL of 4% NaOH were added. The resulting mixture was shaken well at each step, left at ordinary temperature for 6 min, fixed to a volume of 10 mL with 30% methanol, and left standing for 15 min. The absorbance value at 510 nm was measured, and each sample was replicated three times. The total flavonoid content was calculated using the following formula: total flavonoid content (mg/100 g) = (m1 × v2 × 10−6 × 100)/ (m × v1), where m is the mass of the sample weighed (g); m1 is the mass of total flavonoid in the solution measured (g) using the standard curve; v1 is the volume of the extraction solution used for the determination in the sample (mL); and v2 is the total volume of the sample presented (mL).
2.9 RNA Extraction and qRT-PCR
Total RNA was extracted using the Tiangen RNAprep Pure Polysaccharide Polyphenol Plant Total RNA Extraction Kit based on the manufacturer’s instructions. The RNA was reverse transcribed using the RNA PCR Kit Ver. 2.1 (TaKaRa, Dalian, China). Primers were synthesized according to Table S1, and qRT-PCR was performed using TB Green Premix Ex Taq II (Tli RNaseH Plus) (Code No. RR820A/B). Ct values were calculated based on three replicate technical experiments performed on three biological replicates. The UBQ gene was used as an internal reference gene for normalization. The relative expression levels of target genes were determined using the 2−ΔΔCt method.
2.10 Statistical Analysis
Data were processed and plotted using Excel 2016 data processing software. Heat maps were plotted using TBtools, and correlation and significance of differences were analyzed using SPSS 18.0. Data were presented as mean values ± SEs, and statistical analyses were performed using the Duncan test (p < 0.05).
3 Results
3.1 Determination of Different Periods of Ca 2+ Uptake in R. roxburghii
In this experiment, a 2 mmol L−1 Ca(CH3COO)2 solution was used as the absorption solution and the Ca2+ absorption characteristic of R. roxburghii at different times was analyzed (Fig. 1). Calcium content in absorption solution had the most reduction within the first hour of absorption. The calcium saturation was reached after 5–6 h. Next, we conducted experiments at 10-min intervals, and the rapid Ca2+ uptake period of R. roxburghii occurred within 0–10 min. Therefore, we chose 5 min as the rapid Ca2+ uptake period, 6 h as the Ca2+ saturation period, and 0 min as the starvation period.
3.2 Quality Assessment of Transcriptome Sequencing and Gene Annotation
The main site of Ca2+ uptake in plants is the root, driven by transpiration pull. Accordingly, the transcriptome of R. roxburghii roots and leaves was sequenced at different calcium uptake periods (0 min, 5 min, and 6 h). After quality control, a total of 116.13 Gb of clean data were obtained. Each sample’s clean data reached 5.71 Gb, Q30 was 94.06% and above, and the GC content ranged from 46.24 to 48.34% (Table S2). The check of base type distribution showed stable content of GC and AT (Fig. S3), and the sequencing error rate was below 0.0005 (Fig. S4). The results of sample replication test demonstrated good sample reproducibility (Fig. 2A). Referring to the selected reference genome sequences, the number of annotated unigene was 37,241, and 86.1% of unigene was compared to the NR database (Table S3). Statistical analysis of the annotation results from NR database revealed (Fig. 2B) that R. roxburghii has a closer relationship with Rosa chinensis, followed by Fragaria vesca.
3.3 Identification of DEGs and Their GO and KEGG Enrichment
Using the screening criteria of fold change ≥ 1.2 and FDR < 0.01, a total of 12,314 DEGs were identified in the experiment. Of these, 1323 genes in roots and 1546 genes in leaves were upregulated in expression during the rapid Ca2+ uptake period compared to the starvation period. The comparison between groups demonstrated more DEGs in roots than in leaves (Fig. S5), suggesting a more active response in roots during Ca2+ uptake. GO functional enrichment contained three main branches (Fig. 3A), including biological process, molecular function, and cellular component. DEGs were mostly involved in cellular processes and metabolic processes within the biological process module, membrane and membrane part within the cellular component module, and catalytic activity, binding, and transporter activity within the molecular function module. The KEGG pathway classification grouped all DEGs into five major categories, with a total of 134 pathways (Fig. 3B). The DEGs of MAPK signaling pathway, plant hormone signal transduction, and plant-pathogen interaction accounted for a higher percentage, up to 5.61%, 7.61%, and 11.05%, respectively. This finding demonstrates that Ca2+ is involved in signal transduction, multiple metabolic processes, and defense processes in R. roxburghii.
DEG enrichment analysis. A Statistical diagram of GO annotation classification of DEG; B enrichment map of DEG KEGG pathway. Note: A The abscissa is the go classification, the left side of the ordinate is the percentage of the number of genes, and the right side is the number of genes; B the ordinate is the name of KEGG channel; the abscissa is the enrichment factor, representing the significance of DEG enrichment level in this pathway; the color of the circle represents Q value. The smaller this value is, the more reliable the enrichment significance of DEG in this pathway is; the size of the circle indicates the number of genes enriched in the pathway. The larger the circle, the more DEG
3.4 Identification and Expression Validation of Genes Involved in Calcium Uptake and Transport in R. roxburghii
Based on previous research literature, we screened the DEGs of ion channels and carrier that were related to ion transport, combined with the results of GO annotation, COG function enrichment, and KEGG pathway enrichment. We further correlated these findings with the changes in the Ca2+ uptake rate of R. roxburghii saplings (Table 1). We used the screening criteria of correlation coefficient absolute value > 0.7 and FPKM values > 10 to identify candidate genes involved in calcium uptake and transport. By these methods, we screened 25 candidate genes in leaves and roots (Fig. 4). These genes belong to seven gene families, which include the cyclic nucleotide-gated ion channel (CNGC), zinc transporter (ZTP), metal tolerance protein (MTP), ER-type Ca2+ ATPase (ECA), Autoinhibited Ca2+ ATPase (ACA), glutamate receptor (GLR), and natural resistance-associated macrophage protein (NRAMP). The expression of RrACA1/2/3, RrCNGC1/2/3/4/5/6/7/8, RrECA1, RrGLR1, and RrMTP1/2 in leaves was positively correlated with the rate of Ca2+ uptake. This indicates that they facilitate the upward transport of Ca2+ and promote its uptake. In the root, the expression of RrCNGC9/10, RrECA2, and RrGLR2 was positively correlated with the rate of Ca2+ uptake, suggesting that these genes play a positive regulatory role in Ca2+ uptake. Interestingly, we found that the expression of genes RrCNGC11/12, RrMTP1/3, RrNRAMP1, and RrZTP1/2 in roots exhibited a negative correlation with the rate of Ca2+ uptake, suggesting that these genes may regulate ion homeostasis during starvation and calcium saturation periods. qRT-PCR results confirmed the reliability of the transcriptome sequencing results (Figs. S6 and S7).
3.5 Changes of Calcium Content in R. roxburghii Under Different Calcium Concentrations and the Response of Genes Related to Ca 2+ Uptake and Transport
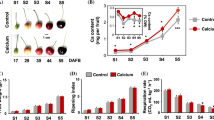
To further identify genes that respond to different levels of calcium, the calcium content of roots and leaves was measured under different calcium concentrations (0 mmol L−1, 0.5 mmol L−1, and 2 mmol L−1), and the expression of 25 candidate genes was assessed in this experiment (Fig. 5). The results revealed that the calcium content in leaves and roots increased rapidly after 0.5 mmol L−1 and 2 mmol L−1 Ca2+ treatments from 0 to 1 day compared with the control (0 mmol L−1 Ca2+). After that, the calcium content in roots kept increasing, while the calcium content in leaves showed a slight decreasing trend, indicating that calcium migrated from leaves to roots in the later stages of calcium uptake. The qRT-PCR analysis and correlation analysis indicated that the expression of RrCNGC7 in the roots was significantly and positively correlated with the calcium content, while the expression of RrCNGC1/10/12 and RrMTP2 in leaves was significantly and positively correlated with the calcium content. When the external calcium concentration changes, these genes played a key role in calcium accumulation within roots or leaves.
Changes in calcium content of R. roxburghii leaves (A) and roots (B) under different calcium concentration treatments and changes in candidate gene expression in leaves (C) and roots (D). Note: Different letters in the figure indicate that the calcium content of different treatments has significant differences. The numbers in the heat map indicate the correlation coefficient between the expression levels of candidate genes and calcium content. *indicates that there is a significant correlation between gene expression and calcium content at the level of 0.05; **indicates that there is a very significant correlation at the level of 0.01
3.6 Spatiotemporal Expression Characteristics of Candidate Genes
In this experiment, the expression patterns of 25 candidate genes were assessed in various tissues of R. roxburghii, including fruits at 30 days after anthesis, leaves, and petals, as well as in fruits at different developmental stages (30, 60, and 90 days after anthesis). The results demonstrated that the expression of these genes was tissue-specific and time-specific (Fig. 6). The expression levels of most genes were significantly higher in leaves and fruits than in petals, and they were also higher in the later stages of fruit development compared to the early stages.
3.7 Relationship Between Calcium Uptake and Transport Genes and Major Active Substances in R. roxburghii Fruits
The R. roxburghii has a significant advantage compared to other plants in active substances of fruit, namely AsA, triterpenoid, and flavonoid. In Fig. 6, it was observed that the majority of candidate genes exhibited high expression levels in middle and late stage fruits. Consequently, the question arises: does Ca2+ regulate the synthesis of these substances? To seek answers, we conducted two treatments on R. roxburghii, namely exogenous Ca2+ soaking of friuts in vitro and spraying plant crown in vivo.
Using the common extracellular Ca2+ chelator, ethylenebis (oxyethylenenitrilo) tetraacetic acid (EGTA) to soak fruit, we observed that the treatment of Ca2+ add to glucose significantly increased the calcium content of the fruits, while the inclusion of EGTA led to a decrease in calcium levels. The trends of AsA, total triterpenoid, and total flavonoid were consistent with the trends observed for calcium content. After the treatment of spraying Ca(CH3COO)2 at concentrations ranging from 100 to 700 mg L−1, it was observed that Ca2+ promoted the accumulation of the three active substances in the fruits at 30 and 60 days after anthesis. However, the optimal Ca2+ concentration for accumulation of these three substances was different at 90 days after anthesis, suggesting a dosage effect of calcium ions on these substances (Fig. 7).
Our correlation analyses and gene expression patterns revealed that the calcium content in fruits was significantly and positively correlated with the content of AsA, total triterpenoid, and total flavonoid after the treatment of Ca2+ soaking fruits in vitro. And RrCNGC3/12 and RrGLR1 played a crucial role in this process (Fig. 8). The correlation coefficient between calcium content and the content of total triterpenoid and total flavonoid in the fruits was greater than 0.8 and showed a highly significant positive correlation with AsA after the Ca2+ spraying (Fig. 9). The expression of RrCNGC2 and AsA content showed a highly significant positive correlation, and RrCNGC12 was significantly correlated with the total triterpenoid and total flavonoid contents and clustered into one group at 30 days after anthesis. The correlation coefficients of calcium content with AsA and total triterpenoid in fruits at 60 days after anthesis were high, and the expression of RrZTP1 was significantly and positively correlated with total flavonoid content. However, as the fruit develops, the correlation coefficients between calcium content in the fruits and the three substances gradually decreased, indicating that Ca2+ mainly regulates the formation of these substances in the early and middle stages of fruit development.
Expression of candidate genes and its correlation analysis with calcium, AsA, total flavonoid, and total triterpenoid content after Ca2+ soaking fruits in vitro. Note: CK, 15 mmol L−1 glucose soaking treatment; Ca, 2 mmol L−1 CaSO4 + 15 mmol L−1 glucose; EGTA, 2 mmol L−1 CaSO4 + 15 mmol L−1 glucose + 5 mmol L−1 EGTA; 0 h, 3 h, 6 h, 12 h, and 24 h represent soaking time. The graph is divided into two parts: the left shows the heat map of gene expression, and the right shows the correlation analysis of gene expression with calcium, AsA, total flavonoid, and total triterpenoid content
Expression of candidate genes and its correlation analysis with calcium, AsA, total flavonoid, and total triterpenoid contents in fruits of A (30 days after anthesis), B (60 days after anthesis), and C (90 days after anthesis) after Ca2+ spraying plant crown in vivo. Note: CK, the water treatment, and the concentration in the figure is the concentration of Ca(CH3COO)2 used for spraying. Each graph is divided into two parts: the left shows the heat map of gene expression, and the right shows the correlation analysis of gene expression with calcium, AsA, total flavonoid, and total triterpenoid content
4 Discussion
4.1 The Uptake of Calcium by R. roxburghii Is a Rapid Process That May Be Involved in Plant Defense Against Pathogens
Compared to other plants, R. roxburghii has the highest content of AsA and is rich in flavonoids, triterpenoids, and other active substances. It grows in karst regions, where the soil calcium content is considerably high. Ca2+ uptake, transport, and function have been a hot topic of research in the field of plant nutrition. However, the calcium nutrition study of R. roxburghii is the early stages of physiological research.
In this experiment, we tried to explore the calcium absorption and transport mechanisms in R. roxburghii. Transcriptome sequencing is a common tool for molecular biology research now, with RNA-Seq being preferentially applied due to its high throughput, broad detection range, and accurate quantification (Bräutigam and Gowik 2010). To select suitable sequencing materials, we performed Ca2+ uptake tests in R. roxburghii. Our results revealed that the rapid uptake period occurred within 0–10 min, and the Ca2+ saturation state was reached after 5–6 h. Similarly, the fastest period of nutrient uptake for iron, phosphorus, potassium, and calcium was 1 h, after which the uptake rate decreased gradually until stabilization in Malus hupehensis (Fan and Yang 2014). Additionally, phosphorus-deficient maize plants sprayed with 200 mmol/L KH2PO4 showed that phosphorus was primarily absorbed within the first 6 h (Görlach et al. 2021), indicating that nutrient uptake is a rapid process.
Calmodulin (CaM) and calmodulin-like (CML) proteins serve as primary Ca2+ sensors and regulate various cellular functions by modulating the activity of diverse target proteins (Cheval et al. 2013). Our assay of GO functional enrichment discovered that DEGs were enriched in functions like catalytic activity, binding, and transport activity, revealing that calcium regulates multiple cellular functions in R. roxburghii cells. Intriguingly, three pathways, namely, plant-pathogen interaction, MAPK signaling pathway, and plant hormone signal transduction, showed the highest percentage of DEGs in the KEGG pathway enrichment. Prior research has demonstrated that calcium signaling through Ca2+ influx in plant apoplast is a crucial stage for disease resistance proteins to function. A calmodulin-binding transcription factor links calcium signaling to antiviral RNAi defense in plants (Wang et al. 2021b). The ZAR1 protein has Ca2+-selective channel activity and induces immunity and cell death (Bi et al. 2021). AtNLRs can create Ca2+-permeable cation channels that modulate cytoplasmic Ca2+ levels and directly mediate cell death signals (Jacob et al. 2021). Intracellular immune receptors form atypical calcium-permeable cation channels in the PM and mediate a prolonged calcium influx, which overcomes the negative effects of pathogen effectors and enhances plant immune responses (Kim et al. 2022). Thus, we suggest that the mechanism of calcium uptake and translocation has a similar network to that of plant disease resistance.
4.2 Seven Gene Families Directly or Indirectly Regulate Calcium Transport Processes in R. roxburghii and Respond to Changes in Calcium Concentration
The treatment of sparying Ca2+ solution in vivo in grape triggers a tremendous number of DEGs associated with calcium transport, affecting the plant’s calcium uptake (Yu et al. 2020). The transport and buffering processes of calcium depend on the participation of multiple ion channels and carrier proteins (Zhang et al. 2019). There were 25 genes involved in calcium uptake and transport process in this trial, covering seven gene families, including ECA, ACA, GLR, CNGC, ZTP, MTP, and NRAMP.
There are two types of Ca2+-pumping ATPase in plant cells, the ECAs and ACAs (Bonza and De Michelis 2011). The ACA is usually located on the plasma membrane, and it is highly selective, only transporting Ca2+ and promoting Ca2+ into the vacuole or the apoplast (Boursiac et al. 2010; Huda et al. 2013; Costa et al. 2017). ECA located in the endoplasmic reticulum (Dodd et al. 2010). Moeder et al. (2019) suggested that two ligand-gated ion channel families, GLRs and CNGCs, play significant roles in immune responses. GLRs are located on the plasma membrane as non-selective Ca2+ channels that can mediate long-distance cytoplasmic Ca2+ signaling (Vincill et al. 2013; Suda and Toyota 2022). CNGCs are distributed across various organelle membranes, and it is usually activated by cAMP and cGMP to promote Ca2+ influx (Gobert et al. 2006; James and Zagotta 2018). Wang et al. (2021a) discovered that CNGC15 and the nitrate transceptor NRT1.1 as a molecular switch together to regulate calcium influx based on nitrate levels. This suggests that non-calcium transporter proteins could also influence calcium transport. Our study found a correlation between the expression of RrMTPs, RrNRAMPs, and RrZTPs families and the rate of calcium uptake. ZTPs are located in the endoplasmic reticulum membrane (Wang et al. 2010), increasing the zinc concentration in the cytosol (Baltaci and Yuce 2018). NRAMP is a type of bivalent metal transporter protein that transports ions like Fe2+, Mn2+, and Zn2+ from the apoplast or organelle to the cytosol (Nevo and Nelson 2006; Mani and Sankaranarayanan 2018), which located in the plasma membrane, endoplasmic reticulum membrane, and golgi membrane (Gao et al. 2018; Li et al. 2022). MTPs located in golgi, vacuole, and mitochondria can transfer Zn, Mn, and Fe into the organelle (Farthing et al. 2017; Gu et al. 2021; Migocka et al. 2018). Some MTPs located at the plasma membrane also contribute to the efflux of excess metal ions from plants and maintenance of ions homeostasis, like MTP10 (Ge et al. 2022a, b; Zhang et al. 2020; Migocka et al. 2015). Interestingly, in previous studies, MTPs and NRAMPs were mainly implicated in the transportation of Mg2+ and facilitated the transport of cations from the subsurface part to the aboveground part (He et al. 2021), and NRAMPs did not have the protein structure to transport Ca2+ (Ehrnstorfer et al. 2014). Ge et al. (2022a, b) found Ca2+ and Mg2+ have antagonistic effects, and MTP10 can maintain the homeostasis between Mg and Ca. We presume that NRAMPs, ZTPs, and MTPs mutually regulate each other, thereby coordinating ion homeostasis in R. roxburghii. However, further testing is required to support this theory.
The calcium uptake is a complex process, which requires different ion channels at various stages of uptake. Both leaves and roots serve as organs for plants to absorb nutrients. Unlike soil fertilization, nutrients sprayed on the leaf surface are mainly absorbed by crops through leaf cuticles and stomata (Niu et al. 2021). In this paper, only the mode of calcium uptake by the roots is discussed, and other modes need to be further explored. Based on previous literature, we predicted the functional pattern of each gene in a hypothetical R. roxburghii cell (Fig. 10). Ca2+ must cross the casparian strip and enter the cytoplasm through Ca2+ channels such as CNGCs and GLRs in the plasma membrane after contacting the epidermis of roots (Alcock et al. 2021). Subsequently, channel proteins such as MTPs, NRAMPs, ZTPs, and CNGCs play a dual role. On the one hand, they carry out long-distance signal transduction and promote upward transport of Ca2+ with the help of transpiration pull. On the other hand, they actively regulate ion homeostasis during calcium uptake. Finally, Ca2+ stored in the organelles of the leaf through ECAs and ACAs. Calcium absorption is affected by abiotic stress factors (Gong et al. 2020). The process correlates with calcium concentration in this experiment. We obtained different candidate genes in different concentrations of the culture solution, reflecting the fact that the uptake pattern changes depending on the calcium concentration. The reason for changes in transportation modes may be that calcium also plays roles as a second messenger coupling a wide range of extracellular stimuli with intracellular responses (Shao et al. 2008).
4.3 The Expression of Transporter Proteins Is Spatiotemporally Specific and Involves the Synthesis of Bioactive Substances
The expression abundance of genes is not alike in different tissues. For example, GLRs preferentially expressed in root (Price et al. 2013). However, out of the 25 candidate genes, most show high expression levels in leaves and fruits (Fig. 6). Calcium oxalate crystals are found exclusively in the phloem of R. roxburghii leaves and stems, as well as in fruits under high-calcium conditions, but not in roots (Meng and Fan 2022). We hypothesize that the main role of transporter proteins in roots is to transport Ca2+ rather than store Ca2+. As the fruit matured, the calcium content in fruit gradually decreased and differences in the expression pattern of ion channels appeared (Fig. 6). These results suggest that calcium absorption is also influenced by tissue and developmental stage.
Wang et al. (2016) discovered that the addition of Ca2+ increased the levels of gamma-aminobutyric acid, isoflavones, phenolics, and vitamins. When plants faced to Ca(NO3)2 stress, the levels of AsA and glutathione decreased significantly, along with the activities of antioxidant enzymes involved in the AsA-GSH cycle (Yuan et al. 2013). Similarly, we found that Ca2+ increased the accumulation of three active substances, including AsA, total triterpenoid, and total flavonoid in R. roxburghii fruits. Research shows that exogenous Ca2+ can alleviate SAR-induced oxidative damage to the cell membrane by enhancing antioxidative capacity (Liang et al. 2021). PMM interacts with CML10 to regulate AsA production in the presence of Ca2+ (Cho et al. 2016). Thus, Ca2+ is involved in the synthesis of active substances. Interestingly, as the fruit continued to develop, the Ca2+ content decreased, while the AsA content consistently increased. Exogenous AsA triggers a transient increase in [Ca2+]cyt in Arabidopsis roots (Makavitskaya et al. 2018). Ca2+ may be regulated by feedback from substances such as AsA.
The expression of RrCNGC12 and RrGLR1 showed a significant and positive correlation with all three substances after Ca2+ soaking treatment (Fig. 8), indicating their potential role in the common pathway for these substances’ synthesis. Additionally, the expression of RrCNGC2 and AsA content in the fruits at 30 days after anthesis showed a highly significant and positive correlation, while the expression of RrZTP1 was significantly and positively correlated with the total flavonoid content in the fruits at 60 days after anthesis (Fig. 9). The disparity in results between the two treatments could be ascribed to the intricacy of the calcium uptake process. The transport process entailed the movement of Ca2+ across various tissues in the Ca2+ spraying treatment. Previous research has demonstrated that exogenous Si boosts the tolerance of cucumber seedlings to cinnamic acid by upregulating the transcript levels of several enzyme genes involved in the AsA-GSH cycle pathway (Meng et al. 2021). Similarly, Ca2+ can transmit the secondary signals to downstream target enzymes and activate them to synthesize various secondary metabolites (Zhao et al. 2005). Based on these findings, we propose that RrCNGC2/3/12, RrGLR1, and RrZTP1 in R. roxburghii fruits influence the accumulation of AsA, triterpenoid, and flavonoid from the protein level and transcriptional level by regulating the Ca2+ level. However, the specific mechanism needs further investigation.
5 Conclusion
In this research, 12,314 DEGs took park in the course of calcium uptake in R. roxburghii. We screened 15 and 11 genes involved in Ca2+ uptake in leaves and roots, respectively. Apart from the Ca2+ transport genes such as CNGCs, ECAs, ACAs, and GLRs, other metal ion transport genes such as NRAMPs, ZTPs, and MTPs are also involved in the Ca2+ uptake process and jointly regulate the ions homeostasis. The CNGC and MTP gene families are particularly instrumental in responding to changes of the exogenous calcium concentration. We have also observed that the exogenous calcium promotes AsA, flavonoid, and triterpenoid accumulation in fruits, and RrCNGC12 play a critical role in this process. Overall, our findings preliminarily shed light on the molecular mechanisms of calcium uptake and the potential effects of Ca2+ on improving fruit quality formation.
Data Availability
All sequencing data are available through the NCBI Sequence Read Archive under the accession number PRJNA993289.
References
Ahmad P, Latef AAA, Abd Allah EF, Hashem A, Sarwat M, Anjum NA, Gucel S (2016) Calcium and potassium supplementation enhanced growth, osmolyte secondary metabolite production, and enzymatic antioxidant machinery in cadmium-exposed chickpea (Cicer arietinum L.). Front Plant Sci 7:513. https://doi.org/10.3389/fpls.2016.00513
Alcock TD, Thomas CL, Ó Lochlainn S, Pongrac P, Wilson M, Moore C, Reyt G, Vogel-Mikuš K, Kelemen M, Hayden R, Wilson L, Stephenson P, Østergaard L, Irwin JA, Hammond JP, King GJ, Salt DE, Graham NS, White PJ, Broadley MR (2021) Magnesium and calcium overaccumulate in the leaves of a schengen3 mutant of Brassica rapa. Plant Physiol 186:1616–1631. https://doi.org/10.1093/plphys/kiab150
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, Martin MJ, Natale DA, O’Donovan C, Redaschi N, Yeh LS (2004) UniProt: the Universal Protein knowledgebase. Nucleic Acids Res 32:D115-119. https://doi.org/10.1093/nar/gkh131
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. Gene Ontol Consortium Nat Genet 25:25–29. https://doi.org/10.1038/75556
Baltaci AK, Yuce K (2018) Zinc transporter proteins. Neurochem Res 43:517–530. https://doi.org/10.1007/s11064-017-2454-y
Bi GZ, Su M, Li N, Liang Y, Dang S, Xu JC, Hu MJ, Wang JZ, Zou MX, Deng YA, Li QY, Huang SJ, Li JJ, Chai JJ, He KM, Chen YH, Zhou JM (2021) The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling. Cell 184:3528–3541. https://doi.org/10.1016/j.cell.2021.05.003
Bonza MC, De Michelis MI (2011) The plant Ca2+-ATPase repertoire: biochemical features and physiological functions. Plant Biol 13:421–430. https://doi.org/10.1111/j.1438-8677.2010.00405.x
Boursiac Y, Lee SM, Romanowsky S, Blank R, Sladek C, Chung WS, Harper JF (2010) Disruption of the vacuolar calcium-ATPases in Arabidopsis results in the activation of a salicylic acid-dependent programmed cell death pathway. Plant Physiol 154:1158–1171. https://doi.org/10.1104/pp.110.159038
Bräutigam A, Gowik U (2010) What can next generation sequencing do for you? Next generation sequencing as a valuable tool in plant research. Plant Biol 12:831–841. https://doi.org/10.1111/j.1438-8677.2010.00373.x
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Cheval C, Aldon D, Galaud JP, Ranty B (2013) Calcium/calmodulin-mediated regulation of plant immunity. Bba-Mol Cell Res 1833:1766–1771. https://doi.org/10.1016/j.bbamcr.2013.01.031
Cho KM, Nguyen HT, Kim SY, Shin JS, Cho DH, Hong SB, Shin JS, Ok SH (2016) CML10, a variant of calmodulin, modulates ascorbic acid synthesis. New Phytol 209:664–678. https://doi.org/10.1111/nph.13612
Costa A, Luoni L, Marrano CA, Hashimoto K, Koster P, Giacometti S, De Michelis MI, Kudla J, Bonza MC (2017) Ca2+-dependent phosphor regulation of the plasma membrane Ca2+-ATPase ACA8 modulates stimulus-induced calcium signatures. J Exp Bot 68:3215–3230. https://doi.org/10.1093/jxb/erx162
Demidchik V, Shabala S, Isayenkov S, Cuin TA, Pottosin I (2018) Calcium transport across plant membranes: mechanisms and functions. New Phytol 220:49–69. https://doi.org/10.1111/nph.15266
Deng YY, Li JQ, Wu SF, Zhu YP, Zhu YP, Chen YW, He FC (2006) Integrated nr database in protein annotation system and its localization. Comput Eng 32:71–74. https://doi.org/10.3969/j.issn.1000-3428.2006.05.026
Dodd AN, Kudla J, Sanders D (2010) The language of calcium signaling. Annu Rev Plant Biol 61:593–620. https://doi.org/10.1146/annurev-arplant-070109-104628
Ehrnstorfer IA, Geertsma ER, Pardon E, Steyaert J, Dutzler R (2014) Crystal structure of a SLC11 (NRAMP) transporter reveals the basis for transition-metal ion transport. Nat Struct Mol Biol 21:990–996. https://doi.org/10.1038/nsmb.2904
Fan WG, Yang HQ (2014) Response of root architecture, nutrients uptake and shoot growth of Malus hupehensis seedling to the shape of root zone. Chinese Agricultural Science 47:3907–3913. https://doi.org/10.3864/j.issn.0578-1752.2014.19.020
Farthing EC, Menguer PK, Fett JP, Williams LE (2017) OsMTP11 is localised at the Golgi and contributes to Mn tolerance. Sci Rep 7:15258. https://doi.org/10.1038/s41598-017-15324-6
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222-230. https://doi.org/10.1093/nar/gkt1223
Gao QF, Gu LL, Wang HQ, Fei CF, Fang X, Hussain J, Sun SJ, Dong JY, Liu HT, Wang YF (2016) Cyclic nucleotide-gated channel 18 is an essential Ca2+ channel in pollen tube tips for pollen tube guidance to ovules in Arabidopsis. Proc Natl Acad Sci USA 113:3096–3101. https://doi.org/10.1073/pnas.1524629113
Gao H, Xie W, Yang C, Xu J, Li J, Wang H, Chen X, Huang CF (2018) NRAMP2, a trans-Golgi network-localized manganese transporter, is required for Arabidopsis root growth under manganese deficiency. New Phytol 217:179–193. https://doi.org/10.1111/nph.14783
Ge H, Shao Q, Chen J, Chen J, Li X, Tan Y, Lan W, Yang L, Wang Y (2022a) A metal tolerance protein, MTP10, is required for the calcium and magnesium homeostasis in Arabidopsis. Plant Signal Behav 17:2025322. https://doi.org/10.1080/15592324.2021.2025322
Ge H, Wang Y, Chen J, Zhang B, Chen R, Lan W, Luan S, Yang L (2022b) An Arabidopsis vasculature distributed metal tolerance protein facilitates xylem magnesium diffusion to shoots under high-magnesium environments. J Integr Plant Biol 64:166–182. https://doi.org/10.1111/jipb.13187
Gobert A, Park G, Amtmann A, Sanders D, Maathuis FJM (2006) Arabidopsis thaliana cyclic nucleotide gated channel 3 forms a non-selective ion transporter involved in germination and cation transport. J Exp Bot 57:791–800. https://doi.org/10.1093/jxb/erj064
Gong Z, Xiong L, Shi H, Yang S, Herrera-Estrella LR, Xu G, Chao DY, Li J, Wang PY, Qin F, Li J, Ding Y, Shi Y, Wang Y, Yang Y, Guo Y, Zhu JK (2020) Plant abiotic stress response and nutrient use efficiency. Sci China Life Sci 63:635–674. https://doi.org/10.1007/s11427-020-1683-x
Görlach BM, Sagervanshi A, Henningsen JN, Pitann B, Mühling KH (2021) Uptake, subcellular distribution, and translocation of foliar-applied phosphorus: short-term effects on ion relations in deficient young maize plants. Plant Physiol Bioch 166:677–688. https://doi.org/10.1016/j.plaphy.2021.06.028
Gu D, Zhou X, Ma Y, Xu E, Yu Y, Liu Y, Chen X, Zhang W (2021) Expression of a Brassica napus metal transport protein (BnMTP3) in Arabidopsis thaliana confers tolerance to Zn and Mn. Plant Sci 304:110754. https://doi.org/10.1016/j.plantsci.2020.110754
He KM, Chen YH, Zhou JM (2021) The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling. Cell 184:3528–3541. https://doi.org/10.1016/j.cell.2021.05.003
Hirschi KD (2004) The calcium conundrum. Both versatile nutrient and specific signal. Plant Physiol 136:2438–2442. https://doi.org/10.1104/pp.104.046490
Huda KM, Banu MS, Tuteja R, Tuteja N (2013) Global calcium transducer P-type Ca2+-ATPases open new avenues for agriculture by regulating stress signalling. J Exp Bot 64:3099–3109. https://doi.org/10.1093/jxb/ert182
Jacob P, Kim NH, Wu FH, El Kasmr F, Chi Y, Walton WG, Furzer OJ, Lietzan AD, Sunil S, Kempthorn K, Redinbo MR, Pei ZM, Wan L, Dangl JL (2021) Plant “helper” immune receptors are Ca2+-permeable nonselective cation channels. Science 373:420–425. https://doi.org/10.1126/science.abg7917
James ZM, Zagotta WN (2018) Structural insights into the mechanisms of CNBD channel function. J Gen Physiol 150:225–244. https://doi.org/10.1085/jgp.201711898
Juric S, Stracenski KS, Krol-Kilinska Z, Zutic I, Uher SF, Dermic E, Topolovec-Pintaric S, Vincekovic M (2020) The enhancement of plant secondary metabolites content in Lactuca sativa L. by encapsulated bioactive agents. Sci Rep 10:3737. https://doi.org/10.1038/s41598-020-60690-3
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280. https://doi.org/10.1093/nar/gkh063
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36:D480-484. https://doi.org/10.1093/nar/gkm882
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Kim NH, Jacob P, Dangl JL (2022) Con-Ca2+-tenating plant immune responses via calcium-permeable cation channels. New Phytol 234:813–818. https://doi.org/10.1111/nph.18044
Koonin EV, Fedorova ND, Jackson JD, Jacobs AR, Krylov DM, Makarova KS, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, Rogozin IB, Smirnov S, Sorokin AV, Sverdlov AV, Vasudevan S, Wolf YI, Yin JJ, Natale DA (2004) A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol 5:R7. https://doi.org/10.1186/gb-2004-5-2-r7
Lecourieux D, Ranjeva R, Pugin A (2006) Calcium in plant defence-signalling pathways. New Phytol 171:249–269. https://doi.org/10.1111/j.1469-8137.2006.01777.x
Li L, Zhu Z, Liao Y, Yang C, Fan N, Zhang J, Yamaji N, Dirick L, Ma JF, Curie C, Huang CF (2022) NRAMP6 and NRAMP1 cooperatively regulate root growth and manganese translocation under manganese deficiency in Arabidopsis. Plant J 110:1564–1577. https://doi.org/10.1111/tpj.15754
Li LL, An HM (2016) Effects of Ca2+ and Cu2+ on the expression of genes related to AsA metabolism in Rosa roxburghii. Fruits. J Hortic 43:1377–1382. https://doi.org/10.16420/j.issn.0513-353x.2016-0013
Liang C, Zhang Y, Ren X (2021) Calcium regulates antioxidative isozyme activity for enhancing rice adaption to acid rain stress. Plant Sci 306:110876. https://doi.org/10.1016/j.plantsci.2021.110876
Liu JY, Niu YF, Zhang JJ, Zhou YQ, Ma Z, Huang X (2018) Ca2+ channels and Ca2+ signals involved in abiotic stress responses in plant cells: recent advances. Plant Cell Tiss Org 132:413–424. https://doi.org/10.1007/s11240-017-1350-0
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Luo C, Fan WG, Liu JP, An HM (2004) Effects of different treatments on fruit development of Rosa roxburghii Tratt. J Guizhou Normal Univ (Natl Sci Ed) 4:7–11. https://doi.org/10.3969/j.issn.1004-5570.2004.04.002
Makavitskaya M, Svistunenko D, Navaselsky I, Hryvusevich P, Mackievic V, Rabadanova C, Tyutereva E, Samokhina V, Straltsova D, Sokolik A, Voitsekhovskaja O, Demidchik V (2018) Novel roles of ascorbate in plants: induction of cytosolic Ca2+ signals and efflux from cells via anion channels. J Exp Bot 69:3477–3489. https://doi.org/10.1093/jxb/ery056
Mani A, Sankaranarayanan K (2018) In silico analysis of natural resistance-associated macrophage protein (NRAMP) family of transporters in rice. Protein J 37:237–247. https://doi.org/10.1007/s10930-018-9773-y
Martins V, Unlubayir M, Teixeira A, Geros H, Lanoue A (2021) Calcium and methyl jasmonate cross-talk in the secondary metabolism of grape cells. Plant Physiol Bioch 165:228–238. https://doi.org/10.1016/j.plaphy.2021.05.034
Meng QJ, Fan WG (2022) Calcium-tolerance type and adaptability to high-calcium habitats of Rosa roxburghii. Chin J Plant Ecol 46:1562–1572. https://doi.org/10.17521/cjpe.2022.0172
Meng X, Luo SL, Dawuda MM, Gao XQ, Wang SY, Xie JM, Tang ZQ, Liu ZC, Wu Y, Jin L, Lyu J, Yu JH (2021) Exogenous silicon enhances the systemic defense of cucumber leaves and roots against CA-induced autotoxicity stress by regulating the ascorbate-glutathione cycle and photosystem II. Ecotox Environ Safe 227:112879. https://doi.org/10.1016/j.ecoenv.2021.112879
Michailidis M, Karagiannis E, Tanou G, Sarrou E, Stavridou E, Ganopoulos I, Karamanoli K, Madesis P, Martens S, Molassiotis A (2019) An integrated metabolomic and gene expression analysis identifies heat and calcium metabolic networks underlying postharvest sweet cherry fruit senescence. Planta 250:2009–2022. https://doi.org/10.1007/s00425-019-03272-6
Migocka M, Papierniak A, Kosieradzka A, Posyniak E, Maciaszczyk-Dziubinska E, Biskup R, Garbiec A, Marchewka T (2015) Cucumber metal tolerance protein CsMTP9 is a plasma membrane H+-coupled antiporter involved in the Mn2+ and Cd2+ efflux from root cells. Plant J 84:1045–1058. https://doi.org/10.1111/tpj.13056
Migocka M, Małas K, Maciaszczyk-Dziubinska E, Posyniak E, Migdal I, Szczech P (2018) Cucumber Golgi protein CsMTP5 forms a Zn-transporting heterodimer with high molecular mass protein CsMTP12. Plant Sci 277:196–206. https://doi.org/10.1016/j.plantsci.2018.09.011
Moeder W, Phan V, Yoshioka K (2019) Ca2+ to the rescue - Ca2+ channels and signaling in plant immunity. Plant Sci 279:19–26. https://doi.org/10.1016/j.plantsci.2018.04.012
Nevo Y, Nelson N (2006) The NRAMP family of metal-ion transporters. Bba-Mol Cell Res 1763:609–620. https://doi.org/10.1016/j.bbamcr.2006.05.007
Niu JH, Liu C, Huang ML, Liu KZ, Yan DY (2021) Effects of foliar fertilization: a review of current status and future perspectives. J Soil Sci Plant Nut 21:104–118. https://doi.org/10.1007/s42729-020-00346-3
Ojo O, Mphahlele MP, Oladeji OS, Mmutlane EM, Ndinteh DT (2022) From wandering weeds to pharmacy: an insight into traditional uses, phytochemicals and pharmacology of genus Chromolaena (Asteraceae). J Ethnopharmacol 291:115155. https://doi.org/10.1016/j.jep.2022.115155
Pertea M, Pertea GM, Antonescu CM, Chang TC, Mendell JT, Salzberg SL (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33:290–295. https://doi.org/10.1038/nbt.3122
Price MB, Kong D, Okumoto S (2013) Inter-subunit interactions between glutamate-like receptors in Arabidopsis. Plant Signal Behav 8:e27034. https://doi.org/10.4161/psb.27034
Sanders D, Brownlee C, Harper JF (1999) Communicating with calcium. Plant Cell 11:691–706. https://doi.org/10.1105/tpc.11.4.691
Sanders D, Pelloux J, Brownlee C, Harper JF (2002) Calcium at the crossroads of signalling. Plant Cell. 14:S401–S417. https://doi.org/10.1105/tpc.002899
Shao HB, Song WY, Chu LY (2008) Advances of calcium signals involved in plant anti-drought. Cr Biol 331:587–596. https://doi.org/10.1016/j.crvi.2008.03.012
Shi QW, Song QB, Wang HX, Bai CM, Wu D, Dong QP, Cheng X, Han XR, Liu YF (2017) Research progress of plant calcium nutrition and calcium/cold signal interaction. ABSR BBE 2017 4:99–102. https://doi.org/10.2991/bbe-17.2017.16
Suda H, Toyota M (2022) Integration of long-range signals in plants: a model for wound-induced Ca2+, electrical, ROS, and glutamate waves. Curr Opin Plant Biol 69:102270. https://doi.org/10.1016/j.pbi.2022.102270
Tang RJ, Luan S (2017) Regulation of calcium and magnesium homeostasis in plants: from transporters to signaling network. Curr Opin Plant Biol 39:97–105. https://doi.org/10.1016/j.pbi.2017.06.009
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33–36. https://doi.org/10.1093/nar/28.1.33
Vafadar F, Amooaghaie R, Ehsanzadeh P, Ghanadian M, Talebi M, Ghanati F (2020) Melatonin and calcium modulate the production of rosmarinic acid, luteolin, and apigenin in Dracocephalum kotschyi under salinity stress. Phytochemistry 177:112422. https://doi.org/10.1016/j.phytochem.2020.112422
Véry AA, Davies JM (2000) Hyperpolarization-activated calcium channels at the tip of Arabidopsis root hairs. Proc Natl Acad Sci USA 97:9801–9806. https://doi.org/10.1073/pnas.160250397
Vincill ED, Clarin AE, Molenda JN, Spalding EP (2013) Interacting glutamate receptor-like proteins in phloem regulate lateral root initiation in Arabidopsis. Plant Cell 25:1304–1313. https://doi.org/10.1105/tpc.113.110668
Wang M, Xu Q, Yu J, Yuan M (2010) The putative Arabidopsis zinc transporter ZTP29 is involved in the response to salt stress. Plant Mol Biol 73:467–479. https://doi.org/10.1007/s11103-010-9633-4
Wang XK, Yang RQ, Zhou YL, Gu ZX (2016) A comparative transcriptome and proteomics analysis reveals the positive effect of supplementary Ca2+ on soybean sprout yield and nutritional qualities. J Proteomics 143:161–172. https://doi.org/10.1016/j.jprot.2016.04.020
Wang XH, Feng CX, Tian LL, Hou CC, Tian W, Hu B, Zhang Q, Ren ZJ, Niu Q, Song JL, Kong DD, Liu LY, He YK, Ma LG, Chu CC, Luan S, Li LG (2021a) A transceptor-channel complex couples nitrate sensing to calcium signaling in Arabidopsis. Mol Plant 14:774–786. https://doi.org/10.1016/j.molp.2021.02.005
Wang Y, Gong Q, Wu Y, Huang F, Ismayil A, Zhang D, Li H, Gu H, Ludman M, Fátyol K, Qi Y, Yoshioka K, Hanley-Bowdoin L, Hong Y, Liu Y (2021b) A calmodulin-binding transcription factor links calcium signaling to antiviral RNAi defense in plants. Cell Host Microbe 29:1393–1406. https://doi.org/10.1016/j.chom.2021.07.003
Wang Z, Wang LH, Li JX, Yang W, Ci JB, Ren XJ, Wang W, Wang YB, Jiang LY, Yang WG (2022) Identification and expression analysis revealed drought stress-responsive Calmodulin and Calmodulin-like genes in maize. J Plant Interact 17:450–461. https://doi.org/10.1080/17429145.2022.2047235
Wang LL, An HM (2013) Optimization of a method for the determination of vitamin C in Rosa roxburghii fruits by HPLC. Modern Food Science and Technology 29: 397–400. https://doi.org/10.13982/j.mfst.1673-9078.2013.02.009
Yang HR, Fan WG (2022) Effects of different calcium supply levels on growth, mineral element absorption and related physiological and biochemical characteristics of Rosa roxburghii seedlings. J Fruit Sci 39:1891–1902. https://doi.org/10.13925/j.cnki.gsxb.20220101
Yang JS, Li WL, Xing C, Xing GN, Guo YX, Yuan HL (2022) Ca2+ participates in the regulation of microalgae triacylglycerol metabolism under heat stress. Environ Res 208:112696. https://doi.org/10.1016/j.envres.2022.112696
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14. https://doi.org/10.1186/gb-2010-11-2-r14
Yu J, Zhu MT, Wang MJ, Xu YS, Chen WT, Yang GS (2020) Transcriptome analysis of calcium-induced accumulation of anthocyanins in grape skin. Sci Hortic-Amsterdam 260:108871. https://doi.org/10.1016/j.scienta.2019.108871
Yuan LY, Du J, Yuan YH, Shu S, Sun J, Guo SR (2013) Effects of 24-epibrassinolide on ascorbate-glutathione cycle and polyamine levels in cucumber roots under Ca(NO3)2 stress. Acta Physiol Plant 35:253–262. https://doi.org/10.1007/s11738-012-1071-2
Zhang X, Yang M, An HM, Huang W, Liu W (2012) Effects of exogenous divalent cations Ca2+, Mg2+, and Cu2+ and acriflavine on ascorbate biosynthesis in Rosa roxburghii fruits. Chin Agric Sci 45:1144–1149. https://doi.org/10.3864/j.issn.0578-1752.2012.06.012
Zhang XM, Liu LX, Su ZM, Shen ZJ, Gao GF, Yi Y, Zheng HL (2019) Transcriptome analysis of Medicago lupulina seedlings leaves treated by high calcium provides insights into calcium oxalate formation. Plant Soil 444:299–314. https://doi.org/10.1007/s11104-019-04283-8
Zhang X, Li Q, Xu W, Zhao H, Guo F, Wang P, Wang Y, Ni D, Wang M, Wei C (2020) Identification of MTP gene family in tea plant (Camellia sinensis L.) and characterization of CsMTP8.2 in manganese toxicity. Ecotox Environ Safe 202:110904. https://doi.org/10.1016/j.ecoenv.2020.110904
Zhao J, Davis LC, Verpoorte R (2005) Elicitor signal transduction leading to production of plant secondary metabolites. Biotechnol Adv 23:283–333. https://doi.org/10.1016/j.biotechadv.2005.01.003
Funding
This work was supported by the Joint Fund of the National Natural Science Foundation of China and the Karst Science Research Center of Guizhou Province (No. U1812401) and the National Natural Science Foundation of China (grant no. 32260730).
Author information
Authors and Affiliations
Contributions
Huaming An: conceptualization, methodology, formal analysis, writing — review and editing, visualization, supervision, funding acquisition. Min Lu: resources, validation, investigation. Zhao Wang: software, formal analysis, investigation, visualization, writing — original draft.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• We conducted experiments using Rosa roxburghii saplings as materials in three stages: Ca2+ starvation, rapid Ca2+ uptake, and Ca2+ saturation.
• We investigated the Ca2+ uptake patterns and molecular mechanisms in Rosa roxburghii plants.
• Some genes associated with Ca2+ uptake have an impact on fruit bioactive substance accumulation.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Z., Lu, M. & An, H. Transcriptome Analysis Reveals Candidate Genes Involved in Calcium Absorption of Rosa roxburghii Plants and their Effects on the Bioactive Substance Accumulation in Fruit. J Soil Sci Plant Nutr 24, 732–748 (2024). https://doi.org/10.1007/s42729-023-01579-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-023-01579-8