Abstract
High-temperature stress is an important abiotic stress that significantly affects the growth and yield of agriculture crops, including cowpea, and is thus a great concern for global food security. Harnessing variability in crop germplasm through various breeding techniques could be a viable option for improving tolerance to various abiotic stresses, including heat stress. Classical breeding approaches have been used to determine the genetics of heat tolerance in cowpea. However, progress has been hampered as heat tolerance is governed by multiple genes and is highly influenced by G × E effects. During the last decade, the increase in cowpea genomic resources has played a key role in elucidating the QTLs controlling heat stress tolerance. Advances in transcriptome resources have also uncovered plausible candidate gene(s) in response to heat tolerance. Likewise, proteomic and metabolomic approaches have offered novel insights into the response of various heat shock and other related proteins and metabolites involved in heat stress tolerance. Most importantly, increasing the precision of phenotyping approaches would enable us to close the genotype–phenotype gap to better understand the heat tolerance response in cowpea. Next-generation breeding techniques, such as MAGIC, genomic selection, speed breeding, and genome editing tools, have the potential to accelerate the creation of heat-tolerant cowpea genotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cowpea [Vigna unguiculata (L.) Walp.]—a vital protein-rich nutritious crop—is an important grain legume that is grown widely across Africa, Southeast Asia, and Latin America, which provides valuable farmer income, especially in sub-Saharan Africa and other developing countries (Ehlers and Hall 1997; Langyintuo et al. 2003; Timko and Singh 2008; Muñoz-Amatriaín et al. 2017). Cowpea is a multifunctional legume providing food (grain), vegetables (green pods and tender levaes), fodder and is also used as green manure and cover crop globally (Alemu et al. 2016; Gonçalves et al. 2016). Globally, 5.8 million tons of cowpea is harvested from 11 million hectares each year (Xiong et al. 2016). Nigeria produces the most cowpea, contributing 2.24 million tons to the global cowpea basket (Boukar et al. 2013). In light of global climate change, heat stress-related events are becoming a serious problem, and may be of great concern for global food security (Bita and Gerats 2013). Cowpea growth and yield are significantly affected by heat stress in the major cowpea growing areas (Jha et al. 2014, 2017). Thus, to sustain cowpea yield under increasing high-temperature stress, cowpea germplasm needs to be screened and to identify ‘stress adaptive’ traits across various gene pools. Numerous classical genetic studies on heat stress tolerance effects on yield and yield parameters in cowpea have been reported; however, low heritability of these traits could not fully explain the genetic inheritance of heat tolerance (Patel and Hall 1988; Marfo and Hall 1992). Advances in cowpea genomics have enabled precise mapping of various trait QTLs of breeding importance, including heat stress tolerance across various linkage groups (Muchero et al. 2009; Lucas et al. 2013a). Likewise, decoding the cowpea genome sequence has enabled the use of SNP markers to conduct genome-wide association analysis for capturing novel genomic regions that contribute to various abiotic stresses, including heat stress tolerance (Muñoz-Amatriaín et al. 2017; Spriggs et al. 2018). Current advances in functional genomics could play an important role in underpinning the relevant candidate genes and QTLs associated various abiotic stress tolerance traits, including heat stress (Yao et al. 2016; Chen et al. 2017). Similarly, proteomics and metabolomics could unravel the role of various heat-shock proteins (HSPs) and metabolites involved in heat stress tolerance in cowpea. Several innovative breeding techniques—including multi-parent advanced generation intercross (MAGIC), speed breeding, genomic selection, and genome editing tools—have been introduced to expedite the genetic gain in various legumes, including cowpea (Meuwissen et al. 2001; Huang et al. 2015; Huynh et al. 2018; Hickey et al. 2019; Olatoye et al. 2019; Jha et al. 2020). Collectively, these innovative breeding technologies, along with various interdisciplinary approaches, could be used to improve the tolerance of cowpea to heat stress and thus support the increasing global demand for cowpea.
Anomalies associated with heat stress in cowpea at various growth phases
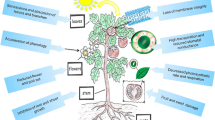
Heat stress challenges key physiological, biochemical, and other essential metabolic processes in plants, resulting in substantial yield losses (Bita and Gerats 2013; Jha et al. 2014, 2017). Intense heat stress impairs photosynthetic mechanism and machinery and respiration process, reduces stomatal conductance, inhibits TCA cycle activity, and increases reactive oxygen species (Wahid et al. 2007; Way and Yamori 2014). The reproductive phase remains the most vulnerable stage to high-temperature stress, which affects flower bud formation and opening, pollen viability and germination, pistil viability, and embryo development, and causes excessive flower abscission and anther dehiscence (Warrag and Hall 1983, 1984; Mutters et al. 1989; Hall 2004). The inhibition of such reproductive processes due to heat stress results in low pod set and pod yield, and early leaf senescence, leading to significant yield losses (Bagnall and King 1987; Ahmed et al. 1992). Notably, reproductive processes are critically affected when night temperatures reach > 20 °C (Ehlers and Hall 1998). Ahmed et al. (1993) reported that high night-time temperatures reduced the supply of sugars in the peduncles of heat-sensitive cowpea lines, resulting in poor pod set. Similarly, Mutters et al. (1989) claimed that heat-sensitive cowpea genotypes restrict proline translocation from anther walls to pollen under heat stress, which damaged the reproductive organs. Ismail and Hall (1998) indicated that a 1 °C increase in night-time temperature beyond 16.5 °C reduced grain yield in cowpea by 13.5%. Likewise, heat stress challenges the nitrogen fixation process via nodule-inhabiting rhizobacteria, which significantly reduces cowpea yield (Michiels et al. 1994). The impacts of heat stress on various reproductive and yield traits and biological processes are depicted in Fig. 1.
Genetic variability for heat stress tolerance in cowpea
Among the various approaches for mitigating heat stress, harnessing genetic variability in various gene pools remains one of the cheapest and most viable options (Mickelbart et al. 2015). Crop germplasm is a natural reservoir of various adaptive traits that enable plants to acclimate under stress conditions, including heat stress (Mickelbart et al. 2015). Considerable genetic variability for heat stress tolerance in cowpea has been captured for plant phenological and yield-related traits (Patel and Hall 1990; Ehlers and Hall 1996; Ehlers et al. 2000; Hall 2004; Timko and Singh 2008). Ehlers and Hall (1996) assessed juvenility, photoperiod and temperature response of 44 cowpea genotypes grown under moderate (31.5/16.5 °C day/night) and high temperatures stress (38/22.5 °C day/night) in the field (under long day condition) and moderate temperature stress (32/18 °C day/night) and high temperatures stress (35/26 °C day/night) in greenhouse (under short and long days) at Riverside and Coachella Valley in California. Based on the juvenility, photoperiod and temperature response these genotypes were categorized into 11 groups, which could allow cowpea breeders to select appropriate phenology and genotype for developing heat tolerant cowpea cultivars (Ehlers and Hall 1996). The cowpea genotype Mouride was registered as a heat-tolerant genotype (Cisse et al. 1995) based on early phenology traits related to the heat escape mechanism. Warrag and Hall (1983) reported that cowpea genotype TVu 4552 had better heat tolerance than California Blackeye No. 5 at the flowering stage. Yet, California Blackeye No. 27 performed well under heat stress, in terms of yield at the reproductive stage (Ehlers et al. 2000). Cowpea genotype Prima was identified as a promising donor parent for improving heat tolerance due to its high pod set under heat stress (Craufurd et al. 1998). In another study, TVu 4552 and Prima were selected as heat-tolerant genotypes, based on their small yield reductions under heat stress (Nielsen and Hall 1985a, b). Timko and Singh (2008) explored the potential of cowpea genotypes IT93K-452-1, IT98K-1111-1, IT93K-693-2, IT97K-472-12 as donors for improving heat stress tolerance based on yield and yield-related traits at the International Institute of Tropical Agriculture (IITA), Ibadan, Nigeria (see Table 1). However, advances in the development of heat-tolerant cowpea genotypes has been slower than for other grain legumes. Therefore, global cowpea germplasm, landraces, and crop wild relatives from gene banks should be screened on a large scale to accelerate the development of climate-resilient cowpea genotypes and increase genetic gain.
Genetics of heat stress tolerance in cowpea
Classical breeding-based studies have offered a preliminary insight into the genetic inheritance of heat stress tolerance in cowpea. An evaluation of heat-tolerant (Prima and TVu 4552) and heat-sensitive (Bambey 23, Magnolia, and 7964) cowpea genotypes in F1, F2, and backcross (BC) populations in the Imperial Valley, California, based on pod number per peduncle and proportion of tolerant plants, revealed that heat tolerance was controlled by a single dominant gene (Marfo and Hall 1992). However, Hall (1993) reported that a single recessive gene controlled heat tolerance during flower bud development. Based on the phenotypic observation in F1, F2, F3, and BC populations derived from TVu 4552 × California Blackeye #5 and Bambey 21 × PI 204,647 crosses, Patel and Hall (1988) reported that the heat‐induced browning of seed coat trait was controlled by single Hbs gene dominant to normal seed coat. However, Mendelian genetics alone does not address the complex genetic inheritance of heat tolerance, as it is governed by multiple genes or minor QTLs and profoundly affected by G × E interactions.
Genomic resources for heat stress tolerance in cowpea
In the last decade, cowpea genomic research has made significant progress with the development of various bi-parental cross-based linkage maps for elucidating the chromosomal location and genetic dissection of various traits of economic importance, including heat stress tolerance (Muchero et al. 2009; Lucas et al. 2013a, b; Pottorff et al. 2012, 2014a, b; Huynh et al. 2016; Lo et al. 2018; Santos et al. 2018; Herniter et al. 2018, 2019). Lucas et al. (2013a) reported five genomic regions in a CB27 × IT82E−18 RIL population that explained 11.5–18.1% of the phenotypic variation controlling heat tolerance in cowpea (Table 2). A comparative genome analysis indicated that these genomic regions are syntenic with the soybean genome and encode HSPs, heat-shock transcription factors, and proline transporters that contribute to enhancing heat tolerance in cowpea (Lucas et al. 2013a). Likewise, Potoroff et al. (2014b) reported three QTLs (Hbs-1, Hbs-2, and Hbs-3) developed from two RIL populations controlling heat stress-induced seed coat browning in cowpea. The identified QTLs explained 9.5–77.3% of the phenotypic variation, and the underlying candidate genes encoding ACC oxidase 2 and ethylene-responsive element-binding factor 3 (ERF3) were elucidated. Thus, SNP markers—SNP 1_0032, 1_1128 linked with Hbs-1, SNP 1_0280, 1_1534, and SNP 1_1404 linked with Hbs3 and SNP 1_1342 linked with Hbs-2—could be used in marker-assisted breeding to screen for heat stress-tolerant cowpea lines (Potoroff et al. 2014b).
Advances in high density genotyping assays has increased cowpea genomic resources, especially high throughput SNP markers (Carvalho et al. 2017; Pan et al. 2017; Xu et al. 2017; Lo et al. 2019), which have advanced the development of high-density genetic maps for narrowing down the QTLs of interest (Muñoz-Amatriaín et al. 2017; Pan et al. 2017; Huang et al. 2018). Likewise, the growing number of SNP markers has been useful to identify causal gene(s)/candidate gene(s) for various traits using marker-trait association analysis based on evolutionary recombination events at the whole genome level (Burridge et al. 2017; Xu et al. 2017; Lo et al. 2019). However, this approach has not been used to pinpoint heat-tolerant genomic regions in cowpea.
Potential of the cowpea genome sequence for exploring heat-tolerant genomic regions
Given the current progress of genome sequencing technologies, Muñoz-Amatriaín et al. (2017) assembled 323 Mb of the cowpea genome (variety IT97K-499-35, an African cultivar) using a shot-gun genome sequencing approach. Spriggs et al. (2018) later assembled 568 Mb (variety IT97K-499-35) and 609 Mb (variety IT86D-1010) of the cowpea genome, and Lonardi et al. (2019) released a draft genome sequence measuring 519.4 Mb (variety IT97K-499-35). Subsequently, Xia et al. (2019) assembled 632.8 Mb (V. unguiculata ssp. sesquipedialis) of the cowpea genome. These genome sequences will allow us to investigate copy number variations, presence-absence variations, and other structural variations that contribute to heat stress tolerance across the cowpea genome.
Progress of functional genomics for heat tolerance in cowpea
Functional genomics is a promising approach for investigating candidate genes and their responses to various biotic and abiotic stresses (Jha et al. 2015). Unprecedented advances in RNA-seq-driven transcriptome profiling have elucidated candidate gene(s) contributing to heat stress tolerance in various grain legumes, including chickpea, soybean, and common bean (Lopes-Caitar et al. 2013; Zhang et al. 2015; Agarwal et al. 2016). However, development of the functional genomics repertoire in cowpea for underpinning candidate genes that contribute to heat stress tolerance has received little attention (Chen et al. 2017; Misra et al. 2017). The ‘Vigna unguiculata Gene Expression Atlas (VuGEA),’ was developed using RNA-seq analysis to gain an insight into the regulation of drought stress and seed development at the molecular level (Yao et al. 2016). Similarly, a transcriptomic approach has been used to establish six tissue-specific transcriptome data sets from cowpea genotypes IT97K-499-35 and IT86D-1010, which has enriched the functional genomic resources in cowpea (Spriggs et al. 2019). The participatory role of miRNAs and their possible target genes contributing to chilling tolerance has been elucidated in cowpea (Zuo et al. 2018).
Proteomics is an important approach for investigating various proteins and complex stress-sensing and signal molecules involved in signal transduction pathways related to heat stress (Kosová et al. 2011; Arbona et al. 2013). Proteomic studies also improve our understanding of the various roles of proteins involved in alleviating the reactive oxygen species-induced toxicity effect under heat stress (Kosová et al. 2011). Previously, Heuss-LaRosa et al. (1987) reported the contribution of two proteins (70 and 80 kDa) involved in heat tolerance in cowpea. Notably, the functional roles of HSPs—HSP90, HSP100, HSP70, HSP60—are worth-mentioning for preventing the misfolding of cellular proteins under heat stress in plant (Al-Whaibi 2011). However, role of these HSPs contributing in heat tolerance has not yet been investigated in cowpea. Similarly, advances in metabolomics could offer insight into the various metabolites generated in response to heat stress, and thus identify the complex pathways controlling heat stress tolerance in cowpea (Shulaev et al. 2008; Ramalingam et al. 2015). In response to the heat stress response, plants reprogram their metabolic processes to adapt and survive under stress conditions. Metabolomics could shed light on the various novel metabolites and complex signaling networks contributing to heat stress adaptation (Bueno and Lopes 2020). Such studies are largely lacking in cowpea. Thus, identification of the expression pattern of various proteins and metabolites under heat stress using proteomics and metabolomics could assist in screening cowpea lines for heat tolerance.
Scope of phenomics approaches for resolving heat stress tolerance in cowpea
Considering the complex genetic inheritance and strong influence of G × E interactions, the precise measurement of the heat stress response remains challenging in cowpea (Furbank and Tester 2010; Houle et al. 2010). Emerging high throughput phenotyping approaches—including automated phenotyping systems involving RGB imaging, multi-sensor based high throughput phenotyping (HTP) platforms, thermal imaging, and hyperspectral reflectance-based HTP platforms, and both manned aircraft and unmanned aerial vehicles, have offered an excellent opportunity for precision phenotyping of various traits in various crop plants, both spatially and temporally (Busemeyer et al. 2013; Andrade-Sanchez et al. 2014; Deery et al. 2014; Crain et al. 2016; Gao et al. 2020). Spectral reflectance-based HTP has been used to build vegetation indices, viz., normalized difference vegetation index (NDVI) for measuring plant green biomass (Araus 2002). In wheat, the NDVI had a positive correlation with grain yield under heat stress (Cossani and Reynolds 2012; Lopes and Reynolds 2012; Juliana et al. 2019). These HTP approaches have not been used to interrogate the heat stress response dynamics in cowpea.
Novel breeding techniques for designing heat resilient cowpea
Several innovative breeding schemes have been introduced in cowpea to increase genetic gains. A MAGIC population involving eight founder parents has been developed to expand the cowpea genetic base and discover novel gene(s) and QTLs contributing to various abiotic stresses in cowpea (Huynh et al. 2018, 2019; Olatoye et al. 2019). This breeding scheme could be used to obtain better recombinants for developing heat-resilient cowpea genotypes.
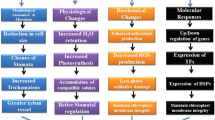
The increasing repertoire of SNP markers derived from high density genotyping platforms provides an excellent opportunity for genomic selection in cowpea. Genomic selection can predict the genomic-estimated breeding value of unobserved individuals using a statistical model that relies on the ‘training population’ with known genotypic and phenotypic information (Meuwissen et al. 2001). Thus, genomic selection provides an opportunity to select individuals for improving genetic gain. Among the novel breeding techniques, genome editing is the most promising for the rapid engineering of heat stress tolerance in various crop species, including cowpea (Ran et al. 2013). Recently, the CRISPR/Cas9 genome editing tool was used to disrupt the VuSYMRK gene responsible for nitrogen fixation in nodule and arbuscular mycorrhizal symbiosis in cowpea (Ji et al. 2019), indicating that it has potential for manipulating stress responses too. Thus, interdisciplinary approaches, including breeding, physiological, genomics, and modern emerging breeding tools, could assist in the development of heat-tolerant cowpea genotypes for future changing climate (Boukar et al. 2016) (see Fig. 2).
Conclusion and perspective
Cowpea production is challenged by incidence of various biotic and abiotic stresses, which have been causing considerable yield losses. The incorporation of crop wild relatives and landraces could help to introgress key adaptive traits contributing to heat tolerance. Along with conventional breeding approaches, molecular breeding approaches could accelerate the transfer of favorable alleles/genomic regions controlling heat stress tolerance in high-yielding, heat-sensitive elite cowpea cultivars. Moreover, increasing genotyping assays and the availability of cowpea genome sequence information should allow us to identify the haplotypes/genomic regions controlling heat stress tolerance for exploitation in cowpea breeding programs.
Further, efforts to resequencing global cowpea germplasm could underpin the genomic regions controlling adaptive traits contributing to heat stress adaptation existing across the whole genome (Muñoz-Amatriaín et al. 2017). Advances in functional genomics could strengthen our understanding of the complex gene networks and the function of candidate gene(s) governing the heat stress response and tolerance. Emerging breeding techniques, including genome editing tools, rapid generation advancement/speed breeding, and genomic selection, could expedite the development of heat-tolerant cowpea genotypes. Heat-tolerant lines should be tested rigorously across various field locations and multiple years. Moreover, the physiological basis for heat tolerance in cowpea is not known, and should be investigated based on various traits reported in other crops like membrane stability, photosynthesis, chlorophyll, and transpirational cooling, to augment the screening process of existing cowpea germplasm for heat tolerance.
References
Agarwal, G., Garg, V., Kudapa, H., Doddamani, D., Pazhamala, L. T., Khan, A. W., et al. (2016). Genome-wide dissection of AP2/ERF and HSP90 gene families in five legumes and expression profiles in chickpea and pigeonpea. Plant Biotechnology Journal, 14(7), 1563–1577.
Ahmed, F. E., Hall, A. E., & DeMason, D. A. (1992). Heat injury during floral development of cowpea (Vigna unguiculata, Fabaceae). American Journal of Botany, 79, 784–791.
Ahmed, F. E., Hall, A. E., & Madore, M. A. (1993). Interactive effects of high temperature and elevated carbon dioxide concentration on cowpea [Vigna unguiculata (L.) Walp.]. Plant, Cell and Environment, 16(7), 835–842.
Alemu, M., Asfaw, Z., Woldu, Z., Fenta, B. A., & Medvecky, B. (2016). Cowpea (Vigna unguiculata (L.) Walp.) (Fabaceae) landrace diversity in northern Ethiopia. International Journal of Biodiversity and Conservation, 8(11), 297–309.
Al-Whaibi, M. H. (2011). Plant heat-shock proteins: A mini review. Journal of King Saud University - Science, 23(2), 139–150.
Andrade-Sanchez, P., Gore, M. A., Heun, J. T., Thorp, K. R., Carmo-Silva, A. E., French, A. N., et al. (2014). Development and evaluation of a field-based high-throughput phenotyping platform. Functional Plant Biology, 41(1), 68–79.
Araus, J. L. (2002). Plant breeding and drought in C3 cereals: What should we breed for? Annals of Botany, 89(7), 925–940.
Arbona, V., Manzi, M., Ollas, C., & Gómez-Cadenas, A. (2013). Metabolomics as a tool to investigate abiotic stress tolerance in plants. International Journal of Molecular Sciences, 14(3), 4885–4911.
Bagnall, D. J., & King, R. W. (1987). Temperature and irradiance effects on yield in Cowpea (Vigna unguiculata). Field Crops Research, 16(3), 217–229.
Bita, C. E., & Gerats, T. (2013). Plant tolerance to high temperature in a changing environment: Scientific fundamentals and production of heat stress-tolerant crops. Frontiers in Plant Science, 4, 273.
Boukar, O., Bhattacharjee, R., Fatokun, C., Kumar, P. L., & Gueye, B. (2013). Cowpea and genomic resources. In M. Singh, H. D. Upadhyaya, & I. S. Bisht (Eds.), Genetic and genomic resources of grain legume improvement (pp. 137–156). London: Elsevier.
Boukar, O., Fatokun, C. A., Huynh, B.-L., Roberts, P. A., & Close, T. J. (2016). Genomic tools in cowpea breeding programs: Status and perspectives. Frontiers in Plant Science, 7, 757.
Bueno, P. C., & Lopes, N. P. (2020). Metabolomics to characterize adaptive and signaling responses in legume crops under abiotic stresses. ACS Omega, 5(4), 1752–1763.
Burridge, J. D., Schneider, H. M., Huynh, B.-L., Roberts, P. A., Bucksch, A., & Lynch, J. P. (2017). Genome-wide association mapping and agronomic impact of cowpea root architecture. Theoretical and Applied Genetics, 130(2), 419–431.
Busemeyer, L., Mentrup, D., Möller, K., Wunder, E., Alheit, K., Hahn, V., et al. (2013). BreedVision—A multi-sensor platform for non-destructive field-based phenotyping in plant breeding. Sensors, 13(3), 2830–2847.
Carvalho, M., Muñoz-Amatriaín, M., Castro, I., Lino-Neto, T., Matos, M., Egea-Cortines, M., et al. (2017). Genetic diversity and structure of Iberian Peninsula cowpeas compared to world-wide cowpea accessions using high density SNP markers. BMC Genomics, 18(1), 891.
Chen, H., Wang, L., Liu, X., Hu, L., Wang, S., & Cheng, X. (2017). De novo transcriptomic analysis of cowpea (Vigna unguiculata L. Walp.) for genic SSR marker development. BMC Genetics, 18(1), 65.
Cisse, N., Ndiaye, M., Thiaw, S., & Hall, A. E. (1995). Registration of ‘Mouride’ cowpea. Crop Science, 35(4), 1215–1216.
Cossani, C. M., & Reynolds, M. P. (2012). Physiological traits for improving heat tolerance in wheat. Plant Physiology, 160(4), 1710–1718.
Crain, J. L., Wei, Y., Barker, J., Thompson, S. M., Alderman, P. D., Reynolds, M., et al. (2016). Development and deployment of a portable field phenotyping platform. Crop Science, 56(3), 965–975.
Craufurd, P. Q., Bojang, M., Wheeler, T. R., & Summerfield, R. J. (1998). Heat tolerance in cowpea: Effect of timing and duration of heat stress. Annals of Applied Biology, 133(2), 257–267.
de Nunes, L. R., Pinheiro, P. R., Pinheiro, C. L., Lima, K. A. P., & Dutra, A. S. (2019). Germination and vigour in seeds of the cowpea in response to salt and heat stress. Revista Caatinga, 32(1), 143–151.
Deery, D., Jimenez-Berni, J., Jones, H., Sirault, X., & Furbank, R. (2014). Proximal remote sensing buggies and potential applications for field-based phenotyping. Agronomy, 4(3), 349–379.
Ehlers, J. D., & Hall, A. E. (1996). Genotypic classification of cowpea based on responses to heat and photoperiod. Crop Science, 36(3), 673–679.
Ehlers, J. D., & Hall, A. E. (1997). Cowpea (Vigna unguiculata L. Walp.). Field Crops Research, 53(1–3), 187–204.
Ehlers, J. D., & Hall, A. E. (1998). Heat tolerance of contrasting cowpea lines in short and long days. Field Crops Research, 55(1–2), 11–21.
Ehlers, J. D., Hall, A. E., Patel, P. N., Roberts, P. A., & Matthews, W. C. (2000). Registration of ‘California Blackeye 27’ cowpea. Crop Science, 40, 854–855.
Furbank, R. T., & Tester, M. (2011). Phenomics – technologies to relieve the phenotyping bottleneck. Trends in Plant Science, 16(12), 635–644.
Gao, G., Tester, M. A., & Julkowska, M. M. (2020). The use of high throughput phenotyping for assessment of heat stress-induced changes in Arabidopsis. Plant Phenomics. https://doi.org/10.34133/2020/3723916
Gonçalves, A., Goufo, P., Barros, A., Domínguez-Perles, R., Trindade, H., Rosa, E. A. S., et al. (2016). Cowpea (Vigna unguiculata L. Walp.), a renewed multipurpose crop for a more sustainable agri-food system: nutritional advantages and constraints: Nutritional aptitude of cowpea. Journal of the Science of Food and Agriculture, 96(9), 2941–2951.
Hall, A. E., (1993). Physiology and breeding for heat tolerance in cowpea, and comparison with other crops. In: Kuo, C.G. (Ed.), Proceedings of the international symposium on adaptation of food crops to temperature and water stress, Taiwan, 13–18 August. Asian Vegetable Research and Development Center, Shanhua (pp. 271–284), Taiwan. Publ. No. 93-410
Hall, A. E. (2004). Breeding for adaptation to drought and heat in cowpea. European Journal of Agronomy, 21(4), 447–454.
Herniter, I. A., Muñoz-Amatriaín, M., Lo, S., Guo, Y.-N., & Close, T. J. (2018). Identification of candidate genes controlling black seed coat and pod tip color in cowpea (Vigna unguiculata [L.] Walp.). Genes Genomes Genetics, 8(10), 3347–3355. https://doi.org/10.1534/g3.118.200521
Herniter, I. A., Lo, R., Muñoz-Amatriaín, M., Lo, S., Guo, Y.-N., Huynh, B.-L., et al. (2019). Seed coat pattern QTL and development in cowpea (Vigna unguiculata [L.] Walp.). Frontiers in Plant Science, 10, 1346.
Heuss-LaRosa, K., Mayer, R. R., & Cherry, J. H. (1987). Synthesis of only two heat shock proteins is required for thermo-adaptation in cultured cowpea cells. Plant Physiology, 85(1), 4–7.
Hickey, L. T., Hafeez, A. N., Robinson, H., Jackson, S. A., Leal-Bertioli, S. C. M., Tester, M., et al. (2019). Breeding crops to feed 10 billion. Nature Biotechnology, 37(7), 744–754.
Houle, D., Govindaraju, D. R., & Omholt, S. (2010). Phenomics: The next challenge. Nature Reviews Genetics, 11(12), 855–866.
Huang, B. E., Verbyla, K. L., Verbyla, A. P., Raghavan, C., Singh, V. K., Gaur, P., et al. (2015). MAGIC populations in crops: Current status and future prospects. Theoretical and Applied Genetics, 128(6), 999–1017.
Huang, H., Tan, H., Xu, D., Tang, Y., Niu, Y., Lai, Y., et al. (2018). High-density genetic map construction and comparative genome analysis in asparagus bean. Scientific Reports, 8(1), 4836.
Huynh, B., Ehlers, J. D., Huang, B. E., Muñoz-Amatriaín, M., Lonardi, S., Santos, J. R. P., et al. (2018). A multi-parent advanced generation inter-cross (MAGIC) population for genetic analysis and improvement of cowpea (Vigna unguiculata L. Walp.). The Plant Journal, 93(6), 1129–1142.
Huynh, B. L., Ehlers, J. D., Close, T. J., & Roberts, P. A. (2019). Registration of cowpea (Vigna unguiculata L. Walp.) multiparent advanced generation intercross (MAGIC) population. Journal of Plant Registration, 13, 281–286.
Huynh, B.-L., Matthews, W. C., Ehlers, J. D., Lucas, M. R., Santos, J. R. P., Ndeve, A., et al. (2016). A major QTL corresponding to the Rk locus for resistance to root-knot nematodes in cowpea (Vigna unguiculata L. Walp.). Theoretical and Applied Genetics, 129(1), 87–95.
Ismail, A. M., & Hall, A. E. (1998). Positive and potential negative effects of heat-tolerance genes in cowpea. Crop Science, 38(2), 381–390.
Jha, U. C., Bhat, J. S., Patil, B. S., Hossain, F., & Barh, D. (2015). Functional genomics: applications in plant science. In D. Barh, M. Khan, & E. Davies (Eds.), Plant omics: the omics of plant science. New Delhi: Springer. https://doi.org/10.1007/978-81-322-2172-2_4.
Jha, U. C., Bohra, A., & Singh, N. P. (2014). Heat stress in crop plants: Its nature, impacts and integrated breeding strategies to improve heat tolerance. Plant Breeding, 133(6), 679–701.
Jha, U. C., Bohra, A., Parida, S. K., & Jha, R. (2017). Integrated “omics” approaches to sustain global productivity of major grain legumes under heat stress. Plant Breeding, 136(4), 437–459.
Jha, U. C., Bohra, A., & Nayyar, H. (2020). Advances in “omics” approaches to tackle drought stress in grain legumes. Plant Breeding, 139(1), 1–27.
Ji, J., Zhang, C., Sun, Z., Wang, L., Duanmu, D., & Fan, Q. (2019). Genome editing in cowpea Vigna unguiculata using CRISPR-Cas9. International Journal of Molecular Sciences, 20(10), 2471.
Juliana, P., Montesinos-López, O. A., Crossa, J., Mondal, S., González Pérez, L., Poland, J., et al. (2019). Integrating genomic-enabled prediction and high-throughput phenotyping in breeding for climate-resilient bread wheat. Theoretical and Applied Genetics, 132(1), 177–194.
Kosová, K., Vítámvás, P., Prášil, I. T., & Renaut, J. (2011). Plant proteome changes under abiotic stress—Contribution of proteomics studies to understanding plant stress response. Journal of Proteomics, 74(8), 1301–1322.
Langyintuo, A. S., Lowenberg-DeBoer, J., Faye, M., Lambert, D., Ibro, G., Moussa, B., et al. (2003). Cowpea supply and demand in West and Central Africa. Field Crops Research, 82(2–3), 215–231.
Lo, S., Muñoz-Amatriaín, M., Boukar, O., Herniter, I., Cisse, N., Guo, Y.-N., et al. (2018). Identification of QTL controlling domestication-related traits in cowpea (Vigna unguiculata L. Walp.). Scientific Reports, 8(1), 6261.
Lo, S., Muñoz-Amatriaín, M., Hokin, S. A., Cisse, N., Roberts, P. A., Farmer, A. D., et al. (2019). A genome-wide association and meta-analysis reveal regions associated with seed size in cowpea [Vigna unguiculata (L.) Walp.]. Theoretical and Applied Genetics, 132(11), 3079–3087.
Lonardi, S., Muñoz-Amatriaín, M., Liang, Q., Shu, S., Wanamaker, S. I., Lo, S., et al. (2019). The genome of cowpea (Vigna unguiculata [L.] Walp.). The Plant Journal, 98(5), 767–782.
Lopes, M. S., & Reynolds, M. P. (2012). Stay-green in spring wheat can be determined by spectral reflectance measurements (normalized difference vegetation index) independently from phenology. Journal of Experimental Botany, 63(10), 3789–3798.
Lopes-Caitar, V. S., de Carvalho, M. C. C. G., Darben, L. M., Kuwahara, M. K., Nepomuceno, A. L., Dias, W. P., & Marcelino-Guimaraes, F. C. (2013). Genome-wide analysis of the Hsp20 gene family in soybean: Comprehensive sequence, genomic organization and expression profile analysis under abiotic and biotic stresses. BMC Genomics, 14, 577.
Lucas, M. R., Ehlers, J. D., Huynh, B.-L., Diop, N.-N., Roberts, P. A., & Close, T. J. (2013a). Markers for breeding heat-tolerant cowpea. Molecular Breeding, 31(3), 529–536.
Lucas, M. R., Huynh, B.-L., da Silva Vinholes, P., Cisse, N., Drabo, I., Ehlers, J. D., et al. (2013b). Association studies and legume synteny reveal haplotypes determining seed size in Vigna unguiculata. Frontiers in Plant Science, 4, 95.
Marfo, K. O., & Hall, A. E. (1992). Inheritance of heat tolerance during pod set in cowpea. Crop Science, 32(4), 912–918.
Meuwissen, T. H. E., Hayes, B. J., & Goddard, M. E. (2001). Prediction of total genetic value using genome-wide dense marker maps. Genetics, 157, 1819–1829.
Michiels, J., Verreth, C., & Vanderleyden, J. (1994). Effects of temperature stress on bean-nodulating rhizobium strains. Applied and Environmental Microbiology, 60(4), 1206–1212.
Mickelbart, M. V., Hasegawa, P. M., & Bailey-Serres, J. (2015). Genetic mechanisms of abiotic stress tolerance that translate to crop yield stability. Nature Reviews Genetics, 16(4), 237–251.
Misra, V. A., Wang, Y., & Timko, M. P. (2017). A compendium of transcription factor and Transcriptionally active protein coding gene families in cowpea (Vigna unguiculata L.). BMC Genomics, 18(1), 898.
Muchero, W., Diop, N. N., Bhat, P. R., Fenton, R. D., Wanamaker, S., Pottorff, M., et al. (2009). A consensus genetic map of cowpea [Vigna unguiculata (L.) Walp.] and synteny based on EST-derived SNPs. Proceedings of the National Academy of Sciences, 106(43), 18159–18164.
Muñoz-Amatriaín, M., Mirebrahim, H., Xu, P., Wanamaker, S. I., Luo, M., Alhakami, H., et al. (2017). Genome resources for climate-resilient cowpea, an essential crop for food security. The Plant Journal, 89(5), 1042–1054.
Mutters, R. G., Ferreira, L. G. R., & Hall, A. E. (1989). Proline content of the anthers and pollen of heat-tolerant and heat-sensitive cowpea subjected to different temperatures. Crop Science, 29(6), 1497–1500.
Nielsen, C. L., & Hall, A. E. (1985a). Responses of cowpea (Vigna unguiculata (L.) Walp.) in the field to high night air temperature during flowering. II Plant responses. Field Crops Research, 10, 181–196.
Nielsen, C. L., & Hall, A. E. (1985b). Responses of cowpea [Vigna unguiculata (L.) Walp.] in the field to high night air temperature during flowering: I. Thermal regimes of production regions and field experiment system. Field Crops Research, 10, 167–179.
Ntare, B. N. (1992). Variation in reproductive efficiency and yield of cowpea under high temperature conditions in Sahelian environment. Euphytica, 59, 27–32.
Olatoye, M. O., Hu, Z., & Aikpokpodion, P. O. (2019). Epistasis detection and modeling for genomic selection in cowpea (Vigna unguiculata L. Walp.). Frontiers in Genetics, 10, 677.
Pan, L., Wang, N., Wu, Z., Guo, R., Yu, X., Zheng, Y., et al. (2017). A high density genetic map derived from RAD sequencing and its application in QTL analysis of yield-related traits in Vigna unguiculata. Frontiers in Plant Science, 8, 1544.
Patel, P. N., & Hall, A. E. (1988). Inheritance of heat-induced brown discoloration in seed coats of cowpea. Crop Science, 28, 929–932.
Patel, P. N., & Hall, A. E. (1990). Genotypic variation and classification of cowpea for reproductive responses to high temperature under long photoperiods. Crop Science, 30, 614–621.
Pottorff, M. O., Wanamaker, S., Ma, Y. Q., Ehlers, J. D., Roberts, P. A., & Close, T. J. (2012). Genetic and physical mapping of candidate genes for resistance to Fusarium oxysporum f. sp. tracheiphilum Race 3 in cowpea [Vigna unguiculata (L.) Walp]. PLoS ONE, 7(7), e41600.
Pottorff, M. O., Li, G., Ehlers, J. D., Close, T. J., & Roberts, P. A. (2014a). Genetic mapping, synteny, and physical location of two loci for Fusarium oxysporum f. sp. tracheiphilum race 4 resistance in cowpea Vigna unguiculata (L.) Walp. Molecular Breeding, 33, 779–791.
Pottorff, M. O., Roberts, P. A., Close, T. J., Lonardi, S., Wanamaker, S., & Ehlers, J. D. (2014b). Identification of candidate genes and molecular markers for heat-induced brown discoloration of seed coats in cowpea [Vigna unguiculata (L.) Walp]. BMC Genomics, 15(1), 328.
Ramalingam, A., Kudapa, H., Pazhamala, L. T., Weckwerth, W., & Varshney, R. K. (2015). Proteomics and metabolomics: Two emerging areas for legume improvement. Frontiers in Plant Science, 6, 1116.
Ran, F. A., Hsu, P. D., Wright, J., Agarwala, V., Scott, D. A., & Zhang, F. (2013). Genome engineering using the CRISPR-Cas9 system. Nature Protocols, 8(11), 2281–2308.
Santos, J. R. P., Ndeve, A. D., Huynh, B.-L., Matthews, W. C., & Roberts, P. A. (2018). QTL mapping and transcriptome analysis of cowpea reveals candidate genes for root-knot nematode resistance. PLoS ONE, 13(1), e0189185.
Spriggs, A., Henderson, S. T., Hand, M. L., Johnson, S. D., Taylor, J. M., & Koltunow, A. (2018). Assembled genomic and tissue-specific transcriptomic data resources for two genetically distinct lines of cowpea (Vigna unguiculata (L.) Walp.). Gates Open Research, 2, 7.
Shulaev, V., Cortes, D., Miller, G., & Mittler, R. (2008). Metabolomics for plant stress response. Physiologia Plantarum, 132, 199–208.
Timko, M. P., & Singh, B. B. (2008). Cowpea, a multifunctional legume. In P. H. Moore & R. Ming (Eds.), Genomics of tropical crop plants (pp. 227–258). New York, NY: Springer.
Wahid, A., Gelani, S., Ashraf, M., & Foolad, M. (2007). Heat tolerance in plants: An overview. Environmental and Experimental Botany, 61(3), 199–223.
Warrag, M. O. A., & Hall, A. E. (1983). Reproductive responses of cowpea to heat stress: Genotypic differences in tolerance to heat at flowering 1. Crop Science, 23(6), 1088–1092.
Warrag, M. O. A., & Hall, A. E. (1984). Reproductive responses of cowpea (Vigna nguiculate (L.) Walp.) to heat stress. II. Responses to night air temperature. Field Crops Research, 8, 17–33.
Way, D. A., & Yamori, W. (2014). Thermal acclimation of photosynthesis: On the importance of adjusting our definitions and accounting for thermal acclimation of respiration. Photosynthesis Research, 119(1–2), 89–100.
Xia, Q., Pan, L., Zhang, R., Ni, X., Wang, Y., Dong, X., et al. (2019). The genome assembly of asparagus bean Vigna nguiculate ssp. Sesquipedialis. Scientific Data, 6(1), 124.
Xiong, H., Shi, A., Mou, B., Qin, J., Motes, D., Lu, W., et al. (2016). Genetic diversity and population structure of cowpea (Vigna nguiculate L. Walp.). PLoS ONE, 11(8), e0160941.
Xu, P., Wu, X., Muñoz-Amatriaín, M., Wang, B., Wu, X., Hu, Y., et al. (2017). Genomic regions, cellular components and gene regulatory basis underlying pod length variations in cowpea (V. unguiculate L. Walp.). Plant Biotechnology Journal, 15(5), 547–557.
Yao, S., Jiang, C., Huang, Z., Torres-Jerez, I., Chang, J., Zhang, H., et al. (2016). The Vigna unguiculate gene expression atlas (VuGEA) from de novo assembly and quantification of RNA-seq data provides insights into seed maturation mechanisms. The Plant Journal, 88, 318–327.
Zhang, L., Zhao, H.-K., Dong, Q.-L., Zhang, Y.-Y., Wang, Y.-M., Li, H.-Y., et al. (2015). Genome-wide analysis and expression profiling under heat and drought treatments of HSP70 gene family in soybean (Glycine max L.). Frontiers in Plant Science, 6, 773.
Zuo, J., Wang, Y., Zhu, B., Luo, Y., Wang, Q., & Gao, L. (2018). sRNAome and transcriptome analysis provide insight into chilling response of cowpea pods. Gene, 671, 142–151.
Acknowledgements
UCJ conceived the idea and wrote the MS with HN, RJ, PJP and KHMS. UCJ acknowledges support from the Indian Council of Agricultural Research (ICAR), New Delhi, India
Funding
No funding was used for writing this article.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jha, U.C., Nayyar, H., Jha, R. et al. Heat stress and cowpea: genetics, breeding and modern tools for improving genetic gains. Plant Physiol. Rep. 25, 645–653 (2020). https://doi.org/10.1007/s40502-020-00544-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-020-00544-2