Abstract
Background and Objectives
Pharmacological control against ovarian serous cystadenocarcinoma has received increasing attention. The purpose of this study was to investigate multi-drug treatments as synergetic therapy for ovarian serous cystadenocarcinoma and to explore their mechanisms of action by the network pharmacology method.
Methods
Genes acting on ovarian serous cystadenocarcinoma were first collected from GEPIA and DisGeNET. Gene Ontology annotation, Kyoto Encyclopedia of Genes and Genomes pathway, Reactome pathway, and Disease Ontology analyses were then conducted. A connectivity map analysis was employed to identify compounds as treatment options for ovarian serous cystadenocarcinoma. Targets of these compounds were obtained from the Search Tool for Interacting Chemicals (STITCH). The intersections between the ovarian serous cystadenocarcinoma-related genes and the compound targets were identified. Finally, the Kyoto Encyclopedia of Genes and Genomes and Reactome pathways in which the overlapped genes participated were selected, and a correspondence compound-target pathway network was constructed.
Results
A total of 541 ovarian serous cystadenocarcinoma-related genes were identified. The functional enrichment and pathway analyses indicated that these genes were associated with critical tumor-related pathways. Based on the connectivity map analysis, five compounds (resveratrol, MG-132, puromycin, 15-delta prostaglandin J2, and valproic acid) were determined as treatment agents for ovarian serous cystadenocarcinoma. Next, 48 targets of the five compounds were collected. Following mapping of the 48 targets to the 541 ovarian serous cystadenocarcinoma-related genes, we identified six targets (PTGS1, FOS, HMOX1, CASP9, PPARG, and ABCB1) as therapeutic targets for ovarian serous cystadenocarcinoma by the five compounds. By analysis of the compound-target pathway network, we found the synergistic anti-ovarian serous cystadenocarcinoma potential and the underlying mechanisms of action of the five compounds.
Conclusion
In summary, latent drugs against ovarian serous cystadenocarcinoma were acquired and their target actions and pathways were determined by the network pharmacology strategy, which provides a new prospect for medicamentous therapy for ovarian serous cystadenocarcinoma. However, further in-depth studies are indispensable to increase the validity of this study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Although ovarian cancer accounts for only 2.5% of new tumor cases in women, it causes 5% of associated deaths as a result of the absence of symptomatology in its early phase, its easy recurrence, and its metastasis in the advanced stage. It ranks as the fifth leading cause of cancer-related mortality [1, 2]. The most common subtype of ovarian carcinoma, ovarian serous cystadenocarcinoma (OSC), constitutes 60–80% of ovarian epithelial neoplasms [3, 4]. Most patients are already in an advanced stage when diagnosed with OSC as a result of the lack of symptoms in the early stage [5], which leads to a heavy tumor burden [6]. Because of its low surgical resection rate and easy recurrence and metastasis, patients with advanced OSC usually have very poor survival outcomes [7]. Effective adjuvant medical treatment could control the disease for 5 years or more [8]. However, most cases relapse within 18 months and develop chemoresistance, which hampers further therapy [8]. Thus, developing a novel and efficacious medicamentous therapy strategy is a necessary and urgent task for the treatment of OSC.
Network pharmacology, first described by Hopkins [9], has been widely utilized in the exploration of new drugs and the re-purposing of existing drugs because of its systematic and integral perspective [10]. Network pharmacology has advantages in unraveling the underlying mechanisms of complex formulas based on a transformation in the development pattern from the contemporary “one target, one drug” mode to an advanced “multi-target, multi-component” mode [11]. In recent years, increasing attention has been paid to the application of the network pharmacology methodology to discover effective medications that can be used for disease therapy and to explore mechanisms of drug action [12,13,14].
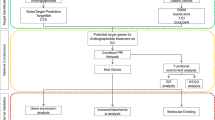
In this study, we employed the network pharmacology method to investigate multi-drug treatments to assist in the therapy for OSC and to explore their mechanisms of action. The workflow of the study is shown in Fig. 1 as follows. First, we obtained genes acting on OSC from GEPIA and DisGeNET. Next, we conducted Gene Ontology (GO) annotation, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway, Reactome pathway, and Disease Ontology (DO) analyses based on these genes to elucidate the potential pathogenesis of OSC. Subsequently, a connectivity map (CMap) approach was adopted to determine possible anti-OSC drugs. The targets of these agents were obtained from the Search Tool for Interacting Chemicals (STITCH). The intersections between the OSC-related genes and the drug targets were identified. Finally, the KEGG and Reactome pathways in which the overlapped genes participated were selected, and a correspondence compound-target pathway network was constructed and analyzed.
2 Materials and Methods
2.1 Collection of Ovarian Serous Cystadenocarcinoma (OSC)-Related Genes
With filter conditions of |log2(fold change)| ≥ 1.5 and an adjusted p value of < 0.05, differentially expressed genes in OSC were obtained from GEPIA (http://gepia.cancer-pku.cn/), an online interactive website for analysis of RNA-sequencing data of 9736 cancer and 8587 non-cancer samples from the Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression (GTEx) projects [15]. Additionally, genes involved in OSC were collected from DisGeNET (http://www.disgenet.org/web/DisGeNET/menu;jsessionid=1mitof7oobjzml489rogrsth), a web server providing genes and variants associated with various diseases [16]. The overlapped genes in both datasets mentioned above were determined as target genes acting on OSC.
2.2 Functional Enrichment Analysis of OSC-Related Genes
Gene Ontology functional annotation, KEGG pathway, and DO analyses were performed by clusterProfiler, an R package used for enrichment analysis of gene clusters [17]. Reactome pathway enrichment analysis was carried out by Reactome FI, a plugin of Cytoscape for pathway and network analysis [18].
2.3 Connectivity Map (CMap) Analysis
Ovarian serous cystadenocarcinoma-related genes with two cohorts of up- and down-regulated genes were imported into CMap (https://portals.broadinstitute.org/cmap/) [19] and were compared with over 7000 gene-expression profiles following treatment by 1309 active compounds in human cell lines. A connectivity score from − 1 to 1 was used to assess the similarity between our query genes and CMap signatures. A positive score indicates an inducement effect of a compound on the query signatures. A negative score reflects a repression effect of a compound on the query signatures. In this study, compounds with a connective score of < − 0.9 were adopted as possible drugs for therapy for OSC.
2.4 Collection of Drug Targets
STITCH (http://stitch.embl.de/), a database of protein-chemical interaction networks [20], was applied to acquire drug targets. Subsequently, a disease-compound-target network was established by Cytoscape 3.6.1 software [21].
2.5 Compound-Target Pathway Network Establishment
An intersection between the chemical targets and OSC-associated genes was processed. The KEGG pathways and Reactome pathways that the resulting genes were involved in were respectively retrieved to build the compound-target-KEGG network and compound-target-Reactome network with Cytoscape 3.6.1.
3 Results
3.1 Collection of 541 Genes Acting on OSC from GEPIA and DisGeNET
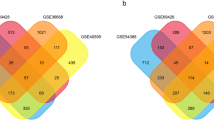
A total of 4237 differentially expressed genes, including 1502 up-regulated genes and 2735 down-regulated genes, were collected from GEPIA, an online tool providing a comparison between 426 OSC samples and 88 normal ovarian specimens (Fig. 2a). Additionally, 2113 genes involved in OSC were collected from the DisGeNET database. After determination of the intersections between the 4237 differentially expressed genes and the 2113 genes corresponding with OSC, 541 candidate genes, including 279 up-regulated and 262 down-regulated genes acting on OSC, were ultimately identified (Fig. 2b). The expression patterns of the 541 genes are shown in Fig. 3.
3.2 Functional Enrichment and Pathway Analyses of the 541 OSC-Related Genes
A GO enrichment analysis was conducted to illustrate the functional annotations of the 541 genes, and the top ten enriched biological processes, cellular components, and molecular functions are exhibited in Fig. 4. According to our results, the highly enriched biological processes were “positive regulation of cell motility”, “positive regulation of cell migration”, and “angiogenesis.” The highly enriched cellular components were “proteinaceous extracellular matrix”, “extracellular matrix component”, and “receptor complex.” The highly enriched molecular functions were “cytokine-receptor binding”, “growth-factor binding”, and “transmembrane receptor protein kinase activity.”
Top ten Gene Ontology (GO) enrichment annotations of the 541 genes related to ovarian serous cystadenocarcinoma. The GO analysis was conducted by R package clusterProfiler and visualized by R package GOplot. The cluster plot shows a circular dendrogram of the clustering of the 541 genes. The inner ring indicates the color-coded log2(fold change); the outer ring represents the assigned biological process [BP] (a), cellular component [CC] (c), and molecular function [MF] (e) terms. The cohort plot displays the 541 genes correlated via ribbons with their assigned BP (b), CC (d), and MF (f) terms
A KEGG pathway analysis was performed to explore the potential pathogenic mechanisms of these genes in the onset and progression of OSC. In total, 86 pathways were enriched by the 541 genes with an adjusted p-value of < 0.05 [Table S1 of the Electronic Supplementary Material (ESM)]. We found that these genes were closely related to cancer-associated pathways, such as the “p53 signaling pathway”, “PI3K-Akt signaling pathway”, “MAPK signaling pathway”, “cell cycle”, and “apoptosis” (Fig. 5). To further delineate the metabolic pathways in which these OSC-related genes participate, we then executed Reactome pathway analysis. With a false-discovery rate of < 0.05, a total of 75 pathways were identified (Table S2 of the ESM). The top 20 enriched pathways are illustrated in Fig. 6a.
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of the 541 genes related to ovarian serous cystadenocarcinoma. The KEGG analysis was conducted by R package clusterProfiler and visualized by R package ggplot2. a Significantly enriched KEGG pathways (adjusted p value < 0.05). b Top 15 enriched KEGG pathways
Reactome pathways and Disease Ontology (DO) analyses. The Reactome-pathway analysis was conducted by Reactome FI, a plugin of Cytoscape. The DO analysis was conducted by R package clusterProfiler. Both the most enriched Reactome pathways and DO terms were visualized by R package ggplot2. a Top 20 enriched Reactome pathways. b Top 20 enriched DO terms
Simultaneously, we also employed DO analysis, which provides disease-related annotations to genes, to investigate which diseases these genes were associated with and to explore diseases that may share similar pathogenesis with OSC. According to our results, these genes were also related to non-small-cell lung carcinoma, musculoskeletal-system cancer, renal-cell carcinoma, and breast carcinoma, in addition to being linked with ovarian malignant tumors (Fig. 6b).
3.3 Identification of Five Compounds as Treatment Options for OSC Based on the CMap Analysis
The CMap method was employed to screen drugs that could be used to treat OSC, and a total of five active compounds (resveratrol, MG-132, puromycin, 15-delta prostaglandin J2, and valproic acid) with connectivity scores of < − 0.9 were ultimately identified (Table 1). The chemical structures of the five compounds were obtained from PubChem [22] and are displayed in Fig. 7.
3.4 Identification of Targets for the Treatment of OSC by the Five Compounds
A total of 48 targets of the five compounds were collected from STITCH (Table 2), and an OSC-compound-target network with 53 nodes and 55 edges was constructed (Fig. 8). We then mapped the 48 targets to the 541 OSC-related genes to obtain common genes that may be targets for the treatment of OSC by these five compounds. Altogether, six genes (PTGS1, FOS, HMOX1, CASP9, PPARG, and ABCB1) were obtained (Table 3; Fig. 9). The expression levels of the six genes are shown in Fig. 10 with data from the GEPIA. Among the six genes, PPARG was a common target of resveratrol and 15-delta prostaglandin J2; PTGS1 was the target of resveratrol; FOS was the target of puromycin; both HMOX1 and CASP9 were the targets of 15-delta prostaglandin J2; and ABCB1 was the target of valproic acid. The relationships between the six genes and overall survival and disease-free survival in patients with OSC were explored using data from GEPIA. None of the six genes were prognostic biomarkers for patients with OSC (Fig. 11).
Venn diagrams for the intersections between genes acting on ovarian serous cystadenocarcinoma (OSC) and compound targets. The intersections between the 541 OSC-related genes and the 48 compound targets from the Search Tool for Interacting Chemicals (STITCH) were identified as targets for the treatment of OSC by the five compounds (resveratrol, MG-132, puromycin, 15-delta prostaglandin J2, and valproic acid)
Prognostic significance of genes PTGS1, FOS, HMOX1, CASP9, PPARG, and ABCB1 in patients with ovarian serous cystadenocarcinoma (OSC). Relationships between PTGS1 (a), FOS (b), HMOX1 (c), CASP9 (d), PPARG (e), and ABCB1 (f) expression and overall survival in patients with OSC. Relationships between PTGS1 (g), FOS (h), HMOX1 (i), CASP9 (j), PPARG (k), and ABCB1 (l) expression and disease-free survival in patients with OSC. HR hazard ratio
3.5 Construction and Analysis of the Compound-Target Pathway Network
To further elucidate the underlying pharmacological action of these candidate compounds for the treatment of OSC, we selected the KEGG and Reactome pathways in which the six genes participate to construct compound-target KEGG pathway and compound-target Reactome pathway networks (Table 3, Figs. 12, 13). According to our results, no related KEGG or Reactome pathway was found for gene PTGS1. Gene FOS was involved in 19 KEGG pathways and six Reactome pathways. Gene HMOX1 was involved in four KEGG pathways and three Reactome pathways. Gene CASP9 participated in 20 KEGG pathways and 13 Reactome pathways. Gene PPARG participated in two KEGG pathways and one Reactome pathway. Gene ABCB1 was related to only two KEGG pathways.
4 Discussion
Network pharmacology, a systematic strategy based on the “multi-component, multi-target” model, has been exploited to decipher the interactions between drugs and human bodies in a holistic manner [9]. Increasing study has focused on the application of network pharmacology in drug discovery owing to its advantages in interpreting drug efficacy, toxicity, and metabolic properties [23,24,25].
In this study, we employed the network pharmacology method, concentrating on the exploration of effective drugs and druggable targets for the treatment of OSC. First, 541 genes acting on OSC were collected from GEPIA and DisGeNET. Functional enrichment and pathway analyses on the 541 genes were then conducted. The GO annotations showed that these genes were mainly enriched in an extracellular matrix, participating in the binding and activation of several cell receptors as well as in the subsequent triggering of certain signaling cascades. The KEGG pathway analysis is generally utilized for understanding the pathogenic mechanisms of various diseases at the molecular level [26]. In this study, following analysis of the KEGG pathway, we found that the OSC-related genes were closely associated with certain critical tumor-related pathways, such as the “p53 signaling pathway”, “MAPK signaling pathway”, “cell cycle”, and “apoptosis.” These findings indicate that the occurrence and development of OSC involve multiple pathways and multiple targets, which provide support for multi-drug therapy for OSC.
Increasing study has demonstrated that abnormalities of the immune system and changes in the immune environment contribute to cancer onset and progression [27,28,29]. However, the elucidation of the interaction between immune imbalance and OSC is still limited. In this study, the results of the Reactome pathway analysis showed that the genes acting on OSC were highly enriched in immune-related pathways, such as “Interleukin-4 and 13 signaling”, “signaling by interleukins”, and “cytokine signaling in the immune system”, revealing the intimate relationship between OSC and immune dysregulation. These findings provide not only an immunological view for interpreting the occurrence of OSC but also evidence for immunotherapy for OSC.
It is well known that a similar pathogenesis may exist in different diseases [30,31,32]. Study of the shared molecular mechanism is conducive to gaining insight into diverse diseases in an integral manner. Our results from the DO analysis showed that the 541 OSC-related genes were also linked with other malignant tumors, such as breast carcinoma, urinary-system cancer, pancreatic cancer, non-small-cell lung carcinoma, germ-cell cancer, and embryonal cancer, suggesting an inner similarity between OSC and these non-OSC cancers and suggesting a pan-cancerous treatment strategy for malignancies with the same pathogenic mechanism. Further studies are necessary to validate these findings.
CMap, founded in 2006, is a database containing over 7000 genome-wide transcriptional expression signatures from cultured human cells treated with 1309 compounds [33]. Unraveling the connections among gene expressions, diseases, and compounds [34], it is widely used for exploring potential novel drugs in various diseases [35,36,37]. In the present study, we implemented a CMap analysis to acquire drugs with therapeutic potential for OSC. A total of five active compounds (resveratrol, MG-132, puromycin, 15-delta prostaglandin J2, and valproic acid) were identified. Resveratrol is a phytoalexin derived from grapes and other food products with antioxidant and potential chemopreventive activities. Animal studies have reported that resveratrol may serve as an effective agent for cancer therapy in various malignancies, including OSC [38,39,40]. However, because of its poor bioavailability [41], clinical research on its anti-tumor effect in patients with OSC is still limited. Popat et al. [42] revealed that resveratrol (5 g/day) resulted in adverse events (nausea, diarrhea, fatigue, and renal toxicity) in a phase II clinical trial involving patients with relapsed or refractory multiple myeloma. Further in vivo and clinical experiments are needed to optimize the bioavailability of resveratrol and verify its anti-tumor effect in patients with OSC.
15-Delta prostaglandin J2 is a natural, peroxisome proliferator-activated receptor gamma ligand. Study of the inhibitory effect of 15-delta prostaglandin J2 on tumors, including OSC, is currently limited. Only one study conducted by de Jong et al. [43] showed that 15-delta prostaglandin J2 induced apoptosis and inhibited migration of wild-type and doxorubicin-resistant ovarian cancer cells. Further animal and human studies are needed to corroborate the therapeutic application of 15-delta prostaglandin J2 and its potential side effects in patients with OSC.
Valproic acid is a fatty acid with anti-neoplastic and anti-angiogenesis potential. It is documented that valproic acid as a single agent inhibits tumor growth in animal models [44, 45]. It is also reported that co-treatment of valproic acid with classic anti-neoplastic drugs exerts synergistic effects, which have a broad-spectrum therapeutic application in cancer therapy [46]. Falchook et al. [47] included 32 cancer patients, of whom ten were diagnosed with ovarian carcinoma. All of the patients received co-administration of azacitidine, valproic acid, and carboplatin. The authors found that three patients with platinum-resistant ovarian cancer achieved stable disease for a longer duration on the clinical trial than on each patient’s pre-treatment. One patient achieved a 26% decrease in tumor size. The authors also observed some drug side effects (fatigue, neutropenia, anemia, thrombocytopenia, nausea, drowsiness, and altered mental status) during the clinical trial. In the present study, we explored the possibility of combining valproic acid with the other four compounds (resveratrol, MG-132, puromycin, and 15-delta prostaglandin J2) in the treatment of OSC. However, further in vitro, in vivo, and clinical studies are necessary to confirm these findings.
Puromycin is a protein-synthesis inhibitor and antimetabolite. Previous studies have reported its therapeutic application in breast cancer [48]. We know that both OSC and breast cancer are female reproductive-system tumors and that they may share similar genetic alterations, and our result from the DO analysis corroborates the similar pathogenesis of these two types of cancer. We thus speculate that puromycin may also act as a disincentive in OSC. Further experimental validation is needed. MG-132, a proteasome inhibitor, has been reported to have the ability to sensitize tumor cells to chemotherapy [49, 50]. Lu et al. [51] demonstrated that MG132 and suberoylanilide hydroxamic acid in vivo and in vitro synergistically suppressed gastric tumor growth. Its role in enhancing the chemosensitivity of cancer cells has also been observed in ovarian carcinoma [52]. However, the synergistic anti-tumor effects of combining MG-132 with the other four compounds (resveratrol, puromycin, 15-delta prostaglandin J2, and valproic acid) still need to be further studied with rigorous experiments.
To probe the pharmacological mechanism of the five compounds in OSC, we collected their targets from STITCH and subsequently integrated those pharmacological targets and the 541 OSC-related genes. Finally, six targets (PTGS1, FOS, HMOX1, CASP9, PPARG, and ABCB1) for the treatment of OSC by the five compounds were identified. Previous studies have reported that these genes could serve as therapeutic targets of anti-tumor agents. PTGS1, as a member of the cyclooxygenases, is inhibited by non-steroidal anti-inflammatory drugs such as aspirin, E-2-desmethylsulindac sulfides, isoxazole, triazole, benzamides, centaureidin, and bergenin [53]. Treatment of Herceptin® in breast cancer cells increases FOS expression, indicating that FOS may be a target of Herceptin® for cancer therapy [54]. The compound imidazole-dioxolane specifically inhibits the expression of HMOX1 in cancer therapy, but further experimental verification is needed [55]. Co-administration of ceranib-2 and carboplatin increases the expression of CASP9 in lung cancer cells [56], suggesting its potential as a target of ceranib-2 and carboplatin. However, for PPARG and ABCB1, few studies report their roles as targets of anti-cancer agents.
We mapped the six target genes to the KEGG and Reactome pathways to determine pathways in which the six genes participate. A network with a multi-compound, multi-target, and multi-pathway model was then constructed. We can see that multiple targets participated in the same pathway. For example, both genes FOS (target of puromycin) and CASP 9 (target of 15-delta prostaglandin J2) were related to apoptosis, indicating that puromycin and 15-delta prostaglandin J2 could exert synergistic anti-tumor effects by jointly regulating the apoptotic pathway. More interestingly, we found that both FOS (target of puromycin) and HOMX1 (target of 15-delta prostaglandin J2) were closely concerned with three immune-related pathways (interleukin-4 and interleukin-13 signaling, signaling by interleukins, and cytokine signaling in the immune system). These findings provide an immunological angle for the treatment of OSC by puromycin and 15-delta prostaglandin J2. Additionally, we found that the compounds resveratrol and 15-delta prostaglandin J2 together target one gene, PPARG, a member of the peroxisome proliferator-activated receptor family. This gene has been demonstrated to be correlated with several cancers, including OSC [57]. We hypothesize that resveratrol and 15-delta prostaglandin J2 may restrain the progression of OSC by co-targeting PPARG. More thorough studies are required to verify these findings.
5 Conclusion
We not only deciphered the pathogenesis of OSC in this study but also, by employing the network pharmacology strategy, identified multiple drugs that could be used for the treatment of OSC. Furthermore, we explored the anti-cancer mechanisms of these compounds in OSC from the perspective of a multi-compound, multi-target, and multi-pathway approach, which is conducive to tapping their potential for synergistic anti-tumor treatment. Further in-depth studies are indispensable to validate these findings.
References
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30.
Kaldawy A, Segev Y, Lavie O, Auslender R, Sopik V, Narod SA. Low-grade serous ovarian cancer: a review. Gynecol Oncol. 2016;143:433–8.
Li J, Fadare O, Xiang L, Kong B, Zheng W. Ovarian serous carcinoma: recent concepts on its origin and carcinogenesis. J Hematol Oncol. 2012;5:8.
Torre LA, Islami F, Siegel RL, Ward EM, Jemal A. Global cancer in women: burden and trends. Cancer Epidemiol Biomark Prev. 2017;26:444–57.
Bachmayr-Heyda A, Aust S, Auer K, Meier SM, Schmetterer KG, Dekan S, et al. Integrative systemic and local metabolomics with impact on survival in high-grade serous ovarian cancer. Clin Cancer Res. 2017;23:2081–92.
Bowtell DD, Bohm S, Ahmed AA, Aspuria PJ, Bast RC Jr, Beral V, et al. Rethinking ovarian cancer II: reducing mortality from high-grade serous ovarian cancer. Nat Rev Cancer. 2015;15:668–79.
Jayson GC, Kohn EC, Kitchener HC, Ledermann JA. Ovarian cancer. Lancet. 2014;384:1376–88.
Hopkins AL. Network pharmacology: the next paradigm in drug discovery. Nat Chem Biol. 2008;4:682–90.
Kibble M, Saarinen N, Tang J, Wennerberg K, Makela S, Aittokallio T. Network pharmacology applications to map the unexplored target space and therapeutic potential of natural products. Nat Prod Rep. 2015;32:1249–66.
Li S, Zhang B. Traditional Chinese medicine network pharmacology: theory, methodology and application. Chin J Nat Med. 2013;11:110–20.
Yu G, Zhang Y, Ren W, Dong L, Li J, Geng Y, et al. Network pharmacology-based identification of key pharmacological pathways of Yin-Huang-Qing-Fei capsule acting on chronic bronchitis. Int J Chron Obstruct Pulmon Dis. 2017;12:85–94.
Qi Q, Li R, Li HY, Cao YB, Bai M, Fan XJ, et al. Identification of the anti-tumor activity and mechanisms of nuciferine through a network pharmacology approach. Acta Pharmacol Sin. 2016;37:963–72.
Azmi AS. Adopting network pharmacology for cancer drug discovery. Curr Drug Discov Technol. 2013;10:95–105.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–102.
Pinero J, Bravo A, Queralt-Rosinach N, Gutierrez-Sacristan A, Deu-Pons J, Centeno E, et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2017;45:D833–9.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Wu G, Dawson E, Duong A, Haw R, Stein L. ReactomeFIViz: a Cytoscape app for pathway and network-based data analysis. F1000Res. 2014;3:146.
Musa A, Ghoraie LS, Zhang SD, Glazko G, Yli-Harja O, Dehmer M, et al. A review of connectivity map and computational approaches in pharmacogenomics. Brief Bioinform. 2017;18:903.
Szklarczyk D, Santos A, von Mering C, Jensen LJ, Bork P, Kuhn M. STITCH 5: augmenting protein-chemical interaction networks with tissue and affinity data. Nucleic Acids Res. 2016;44:D380–4.
Su G, Morris JH, Demchak B, Bader GD. Biological network exploration with Cytoscape 3. Curr Protoc Bioinform. 2014;47:8.13.1–24.
Kim S, Thiessen PA, Bolton EE, Chen J, Fu G, Gindulyte A, et al. PubChem substance and compound databases. Nucleic Acids Res. 2016;44:D1202–13.
de Anda-Jauregui G, Guo K, McGregor BA, Hur J. Exploration of the anti-inflammatory drug space through network pharmacology: applications for drug eepurposing. Front Physiol. 2018;9:151.
Cao H, Li S, Xie R, Xu N, Qian Y, Chen H, et al. Exploring the mechanism of dangguiliuhuang decoction against hepatic fibrosis by network pharmacology and experimental validation. Front Pharmacol. 2018;9:187.
Liu H, Zeng L, Yang K, Zhang G. A network pharmacology approach to explore the pharmacological mechanism of xiaoyao powder on anovulatory infertility. Evid Based Compl Alternat Med. 2016;2016:2960372.
Du J, Yuan Z, Ma Z, Song J, Xie X, Chen Y. KEGG-PATH: Kyoto Encyclopedia of Genes and Genomes-based pathway analysis using a path analysis model. Mol BioSyst. 2014;10:2441–7.
Fagundes CP, Glaser R, Johnson SL, Andridge RR, Yang EV, Di Gregorio MP, et al. Basal cell carcinoma: stressful life events and the tumor environment. Arch Gen Psychiatry. 2012;69:618–26.
Flecken T, Spangenberg HC, Thimme R. Immunobiology of hepatocellular carcinoma. Langenbecks Arch Surg. 2012;397:673–80.
Sprinzl MF, Galle PR. Immune control in hepatocellular carcinoma development and progression: role of stromal cells. Semin Liver Dis. 2014;3:376–88.
Kalaria R. Similarities between Alzheimer’s disease and vascular dementia. J Neurol Sci. 2002;203–204:29–34.
Goh KI, Cusick ME, Valle D, Childs B, Vidal M, Barabasi AL. The human disease network. Proc Natl Acad Sci USA. 2007;104:8685–90.
Hu G, Agarwal P. Human disease-drug network based on genomic expression profiles. PLoS One. 2009;4:e6536.
Qu XA, Rajpal DK. Applications of connectivity map in drug discovery and development. Drug Discov Today. 2012;1:1289–98.
Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell. 2017;171(1437–52):e17.
Malcomson B, Wilson H, Veglia E, Thillaiyampalam G, Barsden R, Donegan S, et al. Connectivity mapping (ssCMap) to predict A20-inducing drugs and their antiinflammatory action in cystic fibrosis. Proc Natl Acad Sci USA. 2016;113:E3725–34.
Walf-Vorderwulbecke V, Pearce K, Brooks T, Hubank M, van den Heuvel-Eibrink MM, Zwaan CM, et al. Targeting acute myeloid leukemia by drug-induced c-MYB degradation. Leukemia. 2018;32:882–9.
Chien W, Sun QY, Lee KL, Ding LW, Wuensche P, Torres-Fernandez LA, et al. Activation of protein phosphatase 2A tumor suppressor as potential treatment of pancreatic cancer. Mol Oncol. 2015;9:889–905.
Sengottuvelan M, Deeptha K, Nalini N. Influence of dietary resveratrol on early and late molecular markers of 1,2-dimethylhydrazine-induced colon carcinogenesis. Nutrition. 2009;25:1169–76.
Tan L, Wang W, He G, Kuick RD, Gossner G, Kueck AS, et al. Resveratrol inhibits ovarian tumor growth in an in vivo mouse model. Cancer. 2016;122:722–9.
Piotrowska-Kempisty H, Rucinski M, Borys S, Kucinska M, Kaczmarek M, Zawierucha P, et al. 3′-hydroxy-3,4,5,4′-tetramethoxystilbene, the metabolite of resveratrol analogue DMU-212, inhibits ovarian cancer cell growth in vitro and in a mice xenograft model. Sci Rep. 2016;6:32627.
Carter LG, D’Orazio JA, Pearson KJ. Resveratrol and cancer: focus on in vivo evidence. Endocr Relat Cancer. 2014;21:R209–25.
Popat R, Plesner T, Davies F, Cook G, Cook M, Elliott P, et al. A phase 2 study of SRT501 (resveratrol) with bortezomib for patients with relapsed and or refractory multiple myeloma. Br J Haematol. 2013;160:714–7.
de Jong E, Winkel P, Poelstra K, Prakash J. Anticancer effects of 15d-prostaglandin-J2 in wild-type and doxorubicin-resistant ovarian cancer cells: novel actions on SIRT1 and HDAC. PLoS One. 2011;6:e25192.
Nagai H, Fujioka-Kobayashi M, Ohe G, Hara K, Takamaru N, Uchida D, et al. Antitumour effect of valproic acid against salivary gland cancer in vitro and in vivo. Oncol Rep. 2014;31:1453–8.
Mattheolabakis G, Wang R, Rigas B, Mackenzie GG. Phospho-valproic acid inhibits pancreatic cancer growth in mice: enhanced efficacy by its formulation in poly-(l)-lactic acid-poly(ethylene glycol) nanoparticles. Int J Oncol. 2017;51:1035–44.
Cincarova L, Zdrahal Z, Fajkus J. New perspectives of valproic acid in clinical practice. Expert Opin Investig Drugs. 2013;22:1535–47.
Falchook GS, Fu S, Naing A, Hong DS, Hu W, Moulder S, et al. Methylation and histone deacetylase inhibition in combination with platinum treatment in patients with advanced malignancies. Invest New Drugs. 2013;31:1192–200.
Jung JH, Sohn EJ, Shin EA, Lee D, Kim B, Jung DB, et al. Melatonin suppresses the expression of 45S preribosomal RNA and upstream binding factor and enhances the antitumor activity of puromycin in MDA-MB-231 breast cancer cells. Evid Based Complement Alternat Med. 2013;2013:879746.
Singh SV, Ajay AK, Mohammad N, Malvi P, Chaube B, Meena AS, et al. Proteasomal inhibition sensitizes cervical cancer cells to mitomycin C-induced bystander effect: the role of tumor microenvironment. Cell Death Dis. 2015;6:e1934.
Li W, Zhang X, Olumi AF. MG-132 sensitizes TRAIL-resistant prostate cancer cells by activating c-Fos/c-Jun heterodimers and repressing c-FLIP(L). Cancer Res. 2007;67:2247–55.
Lu H, Yang XF, Tian XQ, Tang SL, Li LQ, Zhao S, et al. The in vitro and vivo anti-tumor effects and molecular mechanisms of suberoylanilide hydroxamic acid (SAHA) and MG132 on the aggressive phenotypes of gastric cancer cells. Oncotarget. 2016;7:56508–25.
Guo N, Peng Z, Zhang J. Proteasome inhibitor MG132 enhances sensitivity to cisplatin on ovarian carcinoma cells in vitro and in vivo. Int J Gynecol Cancer. 2016;26:839–44.
Vitale P, Panella A, Scilimati A, Perrone MG. COX-1 inhibitors: beyond structure toward therapy. Med Res Rev. 2016;36:641–71.
Lubig J, Lattrich C, Springwald A, Haring J, Schuler S, Ortmann O, et al. Effects of a combined treatment with GPR30 agonist G-1 and herceptin on growth and gene expression of human breast cancer cell lines. Cancer Invest. 2012;30:372–9.
Hjortso MD, Andersen MH. The expression, function and targeting of haem oxygenase-1 in cancer. Curr Cancer Drug Targets. 2014;14:337–47.
Yildiz-Ozer M, Oztopcu-Vatan P, Kus G. The investigation of ceranib-2 on apoptosis and drug interaction with carboplatin in human non small cell lung cancer cells in vitro. Cytotechnology. 2018;70:387–96.
Ivan C, Hu W, Bottsford-Miller J, Zand B, Dalton HJ, Liu T, et al. Epigenetic analysis of the Notch superfamily in high-grade serous ovarian cancer. Gynecol Oncol. 2013;128:506–11.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by a Medical Excellence Award funded by a Creative Research Development Grant from the First Affiliated Hospital of Guangxi Medical University.
Conflict of interest
Dan-dan Xiong, Yue Qin, Wen-qing Xu, Rong-quan He, Hua-yu Wu, Dan-min Wei, Jing-jing Zeng, Yi-wu Dang, and Gang Chen have no conflicts of interest directly relevant to the contents of this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xiong, Dd., Qin, Y., Xu, Wq. et al. A Network Pharmacology-Based Analysis of Multi-Target, Multi-Pathway, Multi-Compound Treatment for Ovarian Serous Cystadenocarcinoma. Clin Drug Investig 38, 909–925 (2018). https://doi.org/10.1007/s40261-018-0683-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40261-018-0683-8