Abstract
Exosomes are very small extracellular vesicles secreted by cells to local and distant tissues. These mini signal transporters elicit acute and chronic effects on recipient cells. Studies regarding exosomes and their relationship to disease, as well as healthy functions, are eliciting extraordinary excitement as data pours in from groups around the world. Reporting of exosome biogenesis, selective loading of cargo, directed release, and resulting changes in adjacent and distal cells are providing information that is changing the way we view cancer progression and treatment. As a result, the properties of exosomes are being exploited for diagnostic, prognostic, and therapeutic applications. First, by referring to the signaling molecules carried by exosomes, they are being tested as indicators of the presence of transformed cells in early stages of cancer. Secondly, the cargo of exosomes secreted from tumors have been linked to prognostic factors and metastatic properties. Thirdly, exosome-based therapies are being developed which utilize the inherent properties of these mini-transporters to affect and interfere with cancer. Exosome creation, loading, and release plays an important role in cancer formation, progression and organotropic metastasis. The developed and developing therapies should be considered with understanding of their advantages and pitfalls, as well as the various roles exosomes play in normal and pathogenic processes. The combination of previously discovered attributes of exosomes with new discoveries occurring daily provide valuable and additive relevant factors to be considered as we embark on the continued discovery of exosomes and their relationship to cancer diagnostics and therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Exosome processing and role in cancer formation and progression is varied between cells and tissue states. |
Exosomes relate to diagnostic, prognostic, and therapeutic potential on many levels. |
Several promising approaches to using exosomes for biomarkers, delivery vehicles, and therapeutics are changing the landscape of cancer diagnosis and treatment. |
1 Introduction
In this review, we discuss exosomes as related to their role in cancer. In vitro and animal studies have demonstrated that many cell types communicate via intercellular signaling using soluble factors and secreted extracellular vesicles (EVs) [1,2,3]. While they are secreted by all cells, cancer cells release significantly more EVs than normal cells, and these can be found in most bodily fluids, including urine, ascites, serum, and plasma [4,5,6,7,8]. Exosomes are a type of EV that delivers cargo containing bioactive lipids, cytokines, growth factors, receptors, transcription factors, DNA, non-coding regulatory RNAs, and messenger RNAs (mRNAs) into recipient cells. This cargo can have profound effects on recipient cells and reflect the cell of origin. Intercellular communication by exosomes contributes to the regulatory signaling of both normal and pathological processes, including cancer [1, 3, 9, 10]. Exosomes have a common morphology, size distribution, and specific marker expression, regardless of tissue origin. Although there are common exosome characteristics, the markers and cargo they carry differ with the original tissue, cell, and cell state. Additionally, the cargo content of exosomes can reflect the origin of the cell and may even indicate pre-metastatic niche formation. With these considerations, exosomes have been recognized as potential diagnostic and prognostic markers for various diseases [11, 12] as well as the assessment of treatment efficiency. Eventually, the data from exosome studies may be used to discover new ways to detect and eliminate malignant precursor cells.
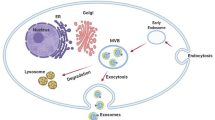
Exosomes were first described as EVs released by tumor cell lines [13]. Since then, they have been described as small, round, and cup-shaped vesicles that collect in multivesicular bodies (MVBs) with size ranges reported from 20 to 200 nm. Exosome biogenesis involves inward budding at the plasma membrane which produces intracellular early endosomes [14]. These bud inward, forming small vesicles termed intraluminal vesicles (ILVs) within a MVB [14]. There are several processes that are believed to facilitate the biogenesis and secretion of exosomes (Fig. 1). These include the endosomal soring complex required for transport (ESCRT) pathway [15,16,17,18] and a ceramide-dependent pathway [16, 19, 20]. Their formation may be controlled by the interaction of syndecan heparin sulphate chains with syntenin-1 and ALG-2 (apoptosis-linked gene 2)-interacting protein X (Alix) for recruitment of the ESCRT machinery [21,22,23]. In addition, protein cargo sorting [24, 25] and release [26] involve the ESCRT complex [19, 27] and other associated proteins such as Alix [21, 28] and Tsg101 [29]. The four components of the ESCRT pathway (ESCRT-O, -I, -II, and -III) aid in the movement of ubiquitinylated proteins [16, 30, 31]. Biogenesis has also been shown to occur in an ESCRT-independent manner through ceramide, flotillin [21], phospholipase D2 (PLD2)/ADP ribosylation factor 6 (ARF6) [32], and CD81 [33]. The ceramide-dependent pathway depends on lipid raft modification to ceramide [19, 20], which ultimately prompts the formation of ILVs in the MVB [16].
Exosome biogenesis. a Inward budding of the plasma membrane creates early endosomes. b Invagination of the early endosome membrane results in ILVs. This may be accomplished by several mechanisms, including: c Lipid rafts may be converted to ceramide, prompting inward budding of the membrane. d Syntenin may bind to ARF6 through the action of PLD2, promoting inward budding. e Ubiquitinated proteins may bind to ESCRT O, which recruits ESCRT I, II, and III. Cooperative interaction of sydecan chains with syntenin, the CD63 tetraspanin, and Alix promotes invagination, followed by interaction of Alix with ESCRT III. f The endosome with the ILVs is referred to as a MVB. The ILVs that remain for secretion to the extracellular space are exosomes. Note that the process by which exosomes are created results in the same orientation of surface markers as the cell membrane. Alix ALG-2 (apoptosis-linked gene 2)-interacting protein X, ARF6 ADP ribosylation factor 6, ESCRT endosomal soring complex required for transport, ILVs intraluminal vesicles, MVB multivesicular body, PLD2 phospholipase D2
Exosomes are secreted from the cell when the MVB fuses with the plasma membrane instead of fusing with a lysosome, releasing their contents extracellularly [14, 34, 35]. Exosomes may also be secreted by direct budding from the plasma membrane. Exosome release involves contributions from Rab proteins [23, 36,37,38], the actin cytoskeleton [39], the microtubule network [40], and cholesterol [41]. Importantly, Rab-mediated release has been shown to occur in a cell-specific manner [38, 42]. This process has been confirmed in antigen-presenting cells (APCs) [43, 44], epithelial cells [45], and tumor cells [46], among others. Due to the mechanism of intake and release, exosomes contain proteins from the plasma membrane and cytosol, as well as the extracellular domain of surface receptors from the cell [47], with very little protein from other organelles [48]. As a result, they bear both intraluminal and transmembrane proteins with the same orientation as the plasma membrane.
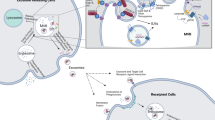
Exosomes enable intercellular communication without direct cell-to-cell contact. This communication can be antigen presentation, immune regulation, spread of infection, and initiation of altered signaling [49] through several mechanisms (Fig. 2). When juxtacrine signaling occurs, the exosomes bind a surface receptor on the recipient cell, initiating intracellular signaling. This signaling is markedly different from fusion and phagocytosis, which transfer the contents of the exosomes into the cell. In endocytosis, the entire exosome is transferred into the cell. The exosomes can also be internalized by endocytosis through several mechanisms reported to occur at acidic pH [41]. There are reports of endocytotic internalization of oligodendroglial and microglial exosomes by neurons [50, 51]. When direct fusion occurs, the contents of the exosomes are delivered to a recipient cell by fusion with the recipient cell membrane resulting in direct release of the cargo to the recipient cytoplasm. The binding of exosomes to the surface of recipient cells has been shown to be mediated by integrins, intercellular adhesion molecules [52], tetraspanins [53], specific glycoproteins [54], and heparan sulfate proteoglycans [55].
Mechanisms of exosome uptake by recipient cells. a The process by which exosomes are created results in the same orientation of surface markers as the cell membrane. b Exosome information is transferred to recipient cells through several mechanisms. Some cell types utilize multiple mechanisms. EBV Epstein-Barr virus, HIV human immunodeficiency virus, MHC myosin heavy chain, NK natural killer cells, Tregs regulatory T cells
The variety of mechanisms for the loading of proteins into exosomes, release mechanisms, and signal delivery routes are not necessarily contradictory. It points to the heterogeneity of exosome populations. In fact, the silencing of specific components of the exosome biogenesis machinery do not affect all exosomal markers in the same way. For example, breast cancer cells secrete exosomes with differing sizes, microRNA, and protein content [56]. In epithelial cells and a colon carcinoma cell line, exosomes released from the apical side of the cells are enriched in specific and different proteins from the basolateral side, even though both populations are identical on electron microscopy and typical exosome markers [57, 58].
Methods to isolate exosomes are a subject of constant debate [59,60,61]. There are many ways to isolate from any number of tissues and cells. The most widely used and reported technique is ultracentrifugation (UC) [4, 61,62,63,64,65]. It has been reported that UC accounts for 81% of all exosome isolation techniques [66]. UC involves spinning suspensions at very high speeds with or without density gradients. Density gradient UC separates exosomes based on their size and mass compared to the medium. This requires large, vacuum-fitted, cooled ultracentrifuges that need much maintenance and user training. UC suffers from high exosome loss [67] and at high velocities (> 100,000g), proteins aggregate and exosomes may clump [5, 59, 68]. Additionally, the viscosity of plasma and presence of lipoproteins, which have similar density and diameter to exosomes, co-isolate with the exosome component [59, 62, 63, 69, 70]. Isolation may also be accomplished using polymer-based precipitation of the exosomes by binding water surrounding the particles [71, 72] or immuno-affinity capture with antibodies against consensus exosome markers [73, 74]. Exosomes cannot be identified by using standard flow cytometry because they are too small to be distinguished from junk signals. But, the existence of markers on the surface of exosomes can be used to sort them [75] if attached to beads with consensus exosomal cell surface markers, such as CD63 and CD81 for distinction. The exosomes bound to beads are then probed with fluorescent antibodies against the same or other consensus exosomal cell surface markers. The distribution of these markers is varied amongst exosome populations from various cell types and tissue states. Microfluidics passes solution with exosomes through small channels on microchips that are treated with antibody [76, 77]. Ultrafiltration utilizes filter-based fractionation at specific speeds and membrane porosity to capture exosomes and other EVs [67, 78]. Commercially available kits have been developed using these techniques [79]. However, these kits have been criticized for low purity, contamination by other microvesicle types, contamination by proteins, loss of exosomes, and undesirable effects on the exosomes themselves [80]. Size exclusion chromatography (SEC) has been shown to isolate exosomes from blood plasma with no significant impurities [62, 67, 81,82,83]. The principle of SEC is separation based on size differences [84, 85]. SEC has been used to isolate vesicles from sera, ascites, and saliva, and was shown to effectively separate vesicles from proteins [62, 63, 86,87,88,89,90]. Böing et al. [62] showed that there is no risk of lipoprotein complex formation or vesicle aggregation as compared to differential centrifugation. With respect to the undesirable effects on the exosomes themselves [80], compared to UC, SEC results in a good recovery of vesicles with almost complete removal of contaminants [4, 62].
An enormous amount of information can be acquired by analysis of exosome contents. These analyses can include differential expression of genomes, transcriptomes, microRNA, lipid and proteome profiles, in addition to more detailed analysis of post-transcriptional or post-translational modifications. The analysis of exosome cargo is accomplished through numerous methodologies, including mass spectroscopy for complete protein screening [91,92,93], the more specific Western blot, next-generation sequencing for complete transcriptome screening [94,95,96], reverse transcription polymerase chain reaction (RT-PCR) for more specific RNA questions, and Nanostring for panels of RNA and microRNA probes. Data from such studies has led to the discovery of various exosome sources involved in cancer establishment and progression. Some of these sources are described in this review.
1.1 Exosomes from Cancer-Associated Fibroblasts
To avoid necrosis as tumor size increases, vascularization at the tumor begins, which establishes the entry point for metastatic cells [97]. Many of the cancer progression processes are influenced by cancer-associated fibroblasts (CAFs) within the tumor. CAFs stimulate new angiogenesis by secreting stromal-derived factor-1 [98], recruiting endothelial progenitor cells [99] and stimulating them to form tube-like structures [100]. It has been demonstrated that CAFs and tumor cells jointly invade blood vessels and implant themselves in metastatic sites [101]. Tumor-derived exosomes (TDEs) can convert mesenchymal stem cells into CAFs [102]. In breast cancer, exosomes from CAFs cross-talk with cancer cells to enhance migration [103] and stimulate epithelial to mesenchymal transition (EMT) [98] by secreting exosome-processed transforming growth factor (TGF)-1 [98]. The TGF-1 effect on stroma differentiation is abrogated by blocking of exosomes, even if the TGF-1 levels in the tumor stroma remain constant [98]. The CAFs also use enzymes (matrix metalloproteinases) to create tunnels through the extracellular matrix that the tumor cells can utilize [104]. It has been shown in sarcoma that exosome matrix metalloproteinases stimulate metastasis [105] and are prognostic for invasiveness in head and neck cancer [106]. In addition, exosomal CD81 (a common exosomal marker) was shown to work with CAFs to promote cancer cell mobility in breast cancer [103].
1.2 Platelets, Exosomes, and Cancer
Under normal circumstances, leukocytes and especially natural killer (NK) cells are found in close relation to metastatic cells in the bloodstream and eliminate these cells [107]. Metastatic cells need to escape this immune surveillance. Platelets adhere to metastatic cells to shield them from NK cell activity [108]. In addition, direct signaling between platelets and cancer cells induces an EMT-like transition and promotes metastasis [109]. The depletion of platelets in mice inhibits metastasis, and platelet reconstitution restores the metastasis [110]. Platelet-derived exosomes assist in the maintenance of cancer cells in the tumor and the blood circulation by also accompanying cancer cells in the bloodstream and enabling their survival [111] through interaction with endothelial cells, leukocytes, and cancer cells [112, 113]. Here, they assist tumor cell adhesion to the endothelium of the recipient vessels and enable extravasation of tumor cells from the bloodstream into the pre-metastatic niche [114], especially via P-selectin [113].
2 Tumor-Derived Exosomes
Normal human blood contains about 1.5 billion exosomes/mL [72]. It has been reported that there are more exosomes secreted from tumor tissue than healthy tissue [115, 116], as well as into patient plasma from the tumor tissue [116, 117]. In one study, this increase was not shown to further increase with tumor progression and was not related to the immune response to the tumor [115], but in other studies there is a correlation to tumor progression [117, 118]. It is possible that the increase is due to local stress within the tumor, such as hypoxia [119] and acidic pH [120]. For example, exosome release from melanoma cells is significantly higher when the medium is not buffered [121] and plasma exosome concentrations of patients with stage III–IV melanoma were shown to be significantly higher than in the plasma of lower stages or healthy patients [118]. Exosomes secreted directly from tumors are referred to as TDEs. These serve to mediate the immune response against a tumor, promote tumor progression, promote migration and engraftment, and affect the response of the tumors to treatment.
Briefly, some mechanisms by which TDEs mediate the immune response against a tumor is by inducing apoptosis of T cells, inhibiting dendritic cell differentiation, inhibiting NK cell cytotoxicity, and inducing myeloid suppressor cells and regulatory T cells [122]. Additionally, the pro-tumorigenic inflammatory response created by TDEs has been shown to support neo-angiogenesis, invasion, and matrix remodeling [123]. This relationship between TDEs and tumorigenic inflammation has been shown in pancreatic ductal adenocarcinoma [9] and liver cancer. TDEs affect the surrounding stromal EMT [124], migration [125], invasion, and engraftment in part by stimulating tumor-associated macrophages [126]. The Toll-like receptors on the surface of the macrophages interact with the TDEs, activating nuclear factor (NF)-κB in the macrophages [127]. For example, in breast cancer, an increased concentration of macrophages correlates with poor prognosis [126].
2.1 Role of Microenvironmental Factors
Since tumor cells are continuously subjected to various forms of stress, there are increased secretions of exosomes by cancer cells [128]. The acidic microenvironment is characteristic of the tumor microenvironment and is considered an important phenotype of malignant tumors [129, 130]. This acidic microenvironment stimulates release of exosomes and enhances their cell fusion capabilities by affecting the lipid composition [121]. Logozzi et al. [131] showed definitively in 2017 that the acidic environment of prostate cancer cells increases the release of exosomes from tumor tissue into patient plasma and this does not occur under buffered conditions. Furthermore, in 2018, Logozzi et al. [132] reported that acidity, as occurs in the tumor microenvironment, increases the release of homogenous exosomes in five different cancer cell lines. This was compared to buffered condition, which resulted in a smaller release of heterogenous exosomes. Clearly, environmental acidity is a key factor in TDE characteristics and release. Heat is a stressor that increases release of immunosuppressive exosomes from B cells in leukemia and lymphoma [133]. Heat may also lead to antitumor functions by inducing the release of exosomes with heat shock proteins (HSPs). Hypoxia is a stressor in the tumor that induces the secretion of exosomes containing proteins associated with cell migration, extracellular matrix degradation, growth hormone signaling, endocytosis, and vascular endothelial growth factor (VEGF) signaling [104, 134]. These signals alter the microenvironment and facilitate angiogenesis and metastasis. For example, malignant brain tumor glioblastoma cell growth is increased and more angiogenic when exposed to exosomes derived under hypoxic conditions, as compared to normoxia-derived exosomes [135]. Exosomes from hypoxic tumor cells can impair NK cell function by secretion of TGF-β1 [136]. This leads to activation of EMT, which plays an important role in metastasis [109]. These stress-induced changes can also provide protective signals influencing the response of distant cells. For example, during oxidative stress or starvation in neuronal development, oligodendrocytes secrete exosomes that are taken up by neurons, promoting neuronal viability [51]. During ischemia/reperfusion injury, mesenchymal stem cells and cardiac progenitor cells release exosomes that have a cardio-protective effect [137] and reduce inflammation [138]. Recognizing the response of tumors to stress and their subsequent exosome release is important for directing discovery of diagnostic and prognostic biomarkers.
2.2 Cargo and Cancer
Exosomes, regardless of origin, contain proteins, lipids, and nucleic acids. The acquisition of cell membrane components and the specific sorting of cargo into exosomes are what distinguish exosomes as being from a particular cell, cell type, or cell state. These differences will distinguish the exosomes from one origin versus exosomes from another origin. The cargo of TDEs is varied and valuable to cancer progression. Consequently, understanding the cargo is important for research of tumor origination, as well as diagnostic, prognostic, and therapeutic uses. Since the exosomes contain lipids and receptors from the cell membrane, cytokines, growth factors, cytoplasmic receptors, transcription factors, non-coding regulatory RNAs, and mRNAs, they are good candidates for detection of the presence of abnormal cells using various body fluids. For example, significantly higher levels of macrophage migratory inhibition factor (MIF) were found in the serum exosomes of patients with pancreatic ductal adenocarcinoma with liver metastasis and progressive disease than in patients with full remission, suggesting MIF as prognostic marker in pancreatic carcinoma [9].
2.2.1 Proteins
Protein cargo of exosomes can be cytosolic, nuclear, transport-involved, adhesion-related, and plasma membrane-bound [48]. These can include cytokines, chemokines, growth factors, receptors, and hormones [139]. According to Exocarta [140], exosomes contain more than 41,860 different proteins. Commonly referenced proteins used to describe exosomes include integrins, tubulin and actin, sorting proteins Alix and Tsg101, major histocompatibility complex (MHC) class I (MHC-I) [141], HSP70, HSP90 [48], and tetraspanins CD9, CD63, CD81 and CD82. Differentially expressed protein cargo can be queried to indicate the status of the cells of origin. Transmembrane protein 256 in the urine has been shown to indicate prostate cancer with 94% sensitivity and 100% specificity [142]. Melo et al. [143] and Hu et al. [144] reported that Glypican-1 in serum exosomes can indicate tumor load/metastasis and stage, respectively, of pancreatic cancer patients with 100% sensitivity and specificity. Conversely, Yang et al. [145] reported much lower sensitivity and specificity for Glypican-1 in pancreatic cancer. Lai et al. [146] reported that, although their own previous studies show a relationship between Glypican-1 and pancreatic cancer, this marker alone is not sufficient and better results were seen using specific microRNAs. Hence, the use of Glypican-1 as a marker alone for pancreatic cancer is controversial. It is likely that more robust specificity will be achieved using a combination of proteins and microRNAs. Examples of other protein indicators found in exosomes are depicted in Table 1. Recent advances in proteomic analysis of exosomes by mass spectroscopy [147, 148] are sure to advance our understanding of exosome activity.
2.2.2 Lipids
Exosomes also contain lipids from membrane lipid rafts including ceramides, sphingolipids, cholesterol, and glycerophospholipids [48]. The localization of many cell membrane components to lipid rafts, which preferentially transfer to exosomes, allows selective omission of some signaling molecules that would otherwise lead to elimination of the exosomes within the extracellular space. Exosomal lipids have also been used for therapeutic targeting. For example, blocking ceramides to reduce the shedding of TDEs.
2.2.3 Nucleic Acids
Importantly, exosomes contain nucleic acids such as microRNA [69], mRNA, DNA, and long non-coding RNAs (lncRNAs). Mitochodrial DNA [161], single-stranded DNA, and chromosomal segments (transposons) [162] have been identified in tumor microvesicles. In 2014, double-stranded DNA was found in exosomes from the serum of pancreatic cancer patients that reflected the mutations in the cells of origin [163]. Since the genetic material can be processed and expressed within the recipient cells, the presence of mutated DNA in TDEs is an important consideration. This cargo is not researched as extensively as RNA content and should not be overlooked.
mRNAs are selectively enriched in exosomes [69]. Microvesicles from embryonic stem cells were able to reprogram hematopoietic progenitors [164] and endothelial microvesicles activated endothelial cells [165] by ‘horizontal transfer’ of mRNA. The sorting of mRNAs normally occurs through coding in the 3'-untranslated region (UTR), as previously described [166]. Trafficking of specific mRNA into exosomes may also be due to coding and potentially subsequent folding within the 3ʹUTR fragments [167]. In an interesting 2013 study, Batagov et al. [168] reported that mRNAs in exosomes exhibit enrichment of 3ʹUTR fragments. These are potential insights into the specific sorting mechanism for mRNA secretion to exosomes, although considerably more information is still sorely needed. The 3ʹUTRs have many regulatory sequences for RNA-binding proteins and microRNAs [167, 169] and the major RNA content of exosomes is microRNAs. MicroRNAs in TDEs are representative of the altered microRNA profile of the tumor. After transport to local and distant sites, the result is influence of gene expression in the target cells of the surrounding stroma and the distant pre-metastatic niche [170]. MicroRNAs packaged in exosomes are very stable in serum and plasma, making the use of exosomes for cancer microRNA profiles very appealing. The idea that mature microRNA, which is normally quickly degraded, is present outside of the cell in the plasma is fascinating. Mitchell et al. [116] showed that these are surprisingly stable and claims that they can be at room temperature for 24 h and can tolerate up to eight freeze/thaw cycles.
Interestingly, processing of pre-microRNAs to mature-microRNAs can occur within exosomes, which allows for direct effects on the recipient cells without further processing [171]. In 2007, Kawahara et al. [172] reported the importance of adenosine-to-inosine (A-I) editing of microRNA. Since, A-I microRNA editing has been related to several human cancers [173, 174]. In fact, microRNA editing has been related to human cancer prognosis with an exhaustive characterization of microRNA sequences for 20 cancer types [175]. These data were further explored to show that A-I microRNA editing is repressed in cancer tissue as compared to normal tissue [176]. Importantly, a recent study has reported, for the first time, that microRNA editing occurs within exosomes which may be predictive of cancer pathogenesis [177]. MicroRNA has become a highly investigated component of cancer at all stages and microRNA in exosomes is becoming more illustrative by the day. For example, in prostate cancer, compared to patients with non-recurrent disease, both miR-141 and miR-375 are increased in the plasma exosomes with recurrent disease after radical prostatectomy [178]. Mitchell et al. [116] also showed increased miR-141 indicative of prostate cancer metastasis with 60% sensitivity and 100% specificity. Table 2 presents some examples of published exosomal microRNA research related to cancer.
The second most abundant RNA in exosomes is lncRNA [220]. This RNA is responsible for regulating gene expression by binding to DNA, but does not code for proteins, or at least does not demonstrate more than 100 amino acid reading frames. lncRNA has been shown to play a role in tumorigenesis. With consideration of the DNA content in exosomes mentioned previously, this component is an interesting part of the signaling carried by TDEs. It is likely the lncRNA studies will reveal more in-depth information about the transfer of mutated DNA to healthy cells, as well as the protection of that DNA during transport. Some studies of lncRNA have been included in Table 2.
3 Exosomes and Cancer Progression
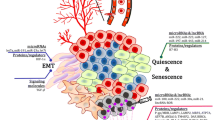
Exosomes contribute to all phases of tumor progression and metastasis (Fig. 3).
The contributions of exosomes to cancer progression. Exosomes contribute to all phases of tumor progression and metastasis. EMT: tumor cells secrete exosomes to surrounding stromal cells and nearby vasculature, signaling EMT in stromal cells and increasing the porosity of the vessels. Migration: exosomes from different tumors are attracted to specific distal tissue types. Preparation: at the distal sites, exosomes alter the local signaling to suppress the immune response and send inflammatory signals, increasing vascular leakiness for incoming cells (arriving tumor cells). Metastasis: as tumor cells arrive at the prepared sites, their exosomes continue to suppress the immune response and alter the new environment. EMT epithelial to mesenchymal transition, TDEs tumor-derived exosomes
3.1 Epithelial to Mesenchymal Transition
It is believed that the tumor-stroma interaction initiates the EMT, which frequently begins the metastatic process. The tumor cells lose their polarity and junctions, reduce proliferation, and increase migration and invasion [221]. This is reversed when the tumor cells reach their destination niche. TDEs can force other cancer cells to follow this process as well. TDEs, but not exosomes in general, are able to induce EMT and enhance migratory activity. Exosome-depleted cancer cells fail to gain a stroma-mediated growth advantage in vivo [98]. Exosomes from nasopharyngeal carcinoma induce EMT by high hypoxia inducible factor 1a (HIF1a) and high latent membrane protein 1 (LMP1; which reduces degradation of HIF1a) [222]. Exosomes from invasive bladder cancer caused urothelial cells to exhibit EMT features, but exosomes from embryonic kidney cells did not cause these changes [223, 224]. This has also been seen in glioblastoma, lung carcinoma, and gastric cancer cells [55, 225, 226]. In lung carcinoma and melanoma, exosomal miR-23a leads to EMT promotion [227, 228]. Exosomes from melanoma, gastric, and colorectal cancer contain miR-191 and let7a, which modulate EMT [229,230,231]. These examples leave little doubt that stromal EMT, facilitating migration of tumor cells, is controlled in part by TDEs.
3.2 Migration and Metastasis
When tumor cells lose their adhesion to the stroma they may migrate into the bloodstream (migration/metastasis). TDEs have been shown to play a role in this process as well. TDEs promote migration and invasion in oral and esophageal squamous carcinoma [188]. In breast cancer, miR-105 in TDEs influences invasion into blood vessels. In brain cancer, blocking TDEs from astrocytes inhibits metastasis [23]. Zhou et al. [170] showed that miR-105 in patient serum is prognostic for metastasis and miR-105 in the exosomes correlates with miR-105 levels in the primary tumor [170]. Further, miR-21 in serum exosomes correlates with migration, invasion, recurrence, and metastasis of esophageal cancer [232].
The tumor cells must recognize the prepared sites for invasion and, upon arrival to these distant sites, exit the circulation and invade the tissue. Prior to invasion and engraftment, the host organ is prepared by TDEs. Hence, the metastatic site is also a consequence of intricate tumor–stroma interactions [233]. Paget’s [235] 1989 ‘soil and seed’ hypothesis expanded upon Fidler [234], suggesting that circulating tumor cells form metastatic nucleation sites only if the seed (tumor cell) and host (organ) are compatible. A pre-metastatic niche formation is required for tumor cells to engraft a distant organ. It was shown in vivo and in vitro with malignant melanoma that this pre-metastatic niche is partially signaled through TDEs. In fact, TDEs are sufficient to direct tumor cells to a specific organ [155]. In malignant melanoma, melanoma exosomes home to sentinel lymph nodes in vivo and recruit melanoma cells [236]. An elegant study of an organotropic metastasis process in pancreatic cancer by Costa-Silva et al. [9] presented that the TDEs cause induction of inflammatory and fibrotic signaling, which activates migration inhibitory factor, which supports TGF-1 production, which enhances production of fibronectin, which attracts bone marrow dendritic cells to the liver. In support of this process, high levels of TGF-1 have been associated with poor prognosis in pancreatic ductal adenocarcinoma [237] and higher levels of migration inhibitory factor (supports TGF-1 function) were found in the serum of patients with pancreatic ductal adenocarcinoma with liver metastasis and progressive disease than in patients with full remission [9]. Additionally, TGF-1 inhibitors are having positive results in clinical trials [238]. In 2016, Lugini et al. [239] showed that exosomes from tumor cells induce transformation of mesenchymal stem cells, supporting the growing evidence that exosomes are important for metastasis.
3.3 Engraftment
When exosomes reach their pre-metastatic niche, they either become dormant tumor cells or proliferate. The TDEs recruit bone marrow dendritic cells by upregulating inflammation, which enhances vascular leakiness at the metastatic sites [240]. Hood et al. [236] showed that exosomes from highly metastatic melanoma cell lines localize to the lungs and increase the permeability of lung endothelium (likely by enhancing the expression of HSP90 and HSP70), which promotes invasion of tumor cells. These exosomes were also seen to localize to bone marrow, kidney, liver, and spleen, establishing metastatic niches, but do not remain in the blood. Bone marrow that has been exposed to metastatic melanoma exosomes had three times more metastatic spread in a mouse model. In pancreatic cancer, the exosomes from the pancreatic tumor interact with the liver Kupffer cells to promote engraftment of tumor cells in the liver [9]. Engraftment begins with reversion of EMT to an epithelial, proliferative, less migratory phenotype with establishment of stable cell junctions. In the recipient organ, TDEs help maintain the tumor cells by promoting immunosuppression. They inhibit effector cells and stimulate regulatory T cells [241, 242], reduce NK cell cytotoxicity [243, 244], and suppress T cell activation by activating myeloid-derived suppressor cells [245, 246].
4 Acquired Resistance Through Exosomes
Recognizing the response of tumors to stress and their subsequent exosome release is not only important for directing discovery, but also for understanding therapeutic responses. Hepatocellular carcinoma cells respond to anticancer drugs by releasing HSP-bearing exosomes, which stimulate the NK cell response [247]. Exosomes may also be involved in removing chemotherapeutic agents from the tumor cells. Exosomes from drug-resistant cells have been shown to contain multidrug resistance-related proteins [248] which help sequester the drugs into vesicles. P-glycoprotein and multidrug resistance protein-1 can be transferred from drug-resistant to drug-sensitive cells through exosomes [249, 250], resulting in an increase in the substrates efflux. For example, after treatment with cisplatin, ovarian carcinoma cells show an increase in the secretion of exosomes carrying cisplatin [251]. Irradiation can induce a senescent phenotype associated with increased production and release of exosome-like vesicles into the microenvironment that can potentially influence tumor progression [252]. For example, proton irradiation has the potential to increase the release of exosomes with survivin, an anti-apoptotic protein involved in cellular proliferation, survival, and tumor cell invasion [253]. The transfer of multidrug resistance-related mRNAs and microRNAs from drug-resistant to drug-sensitive cells may also play a role in the spread of resistance [254]. Interestingly, shear stress has been shown to induce the transfer of athero-protective microRNA from endothelial cells to smooth muscle cells, where they reduce atherosclerotic lesion formation [255].
5 Exosomes for Therapeutics
A significant response against cancer cells is antibody-dependent cytotoxicity, a process that is the basis for therapeutic antibodies [256]. Unfortunately, TDEs have been shown to neutralize antibodies against tumors by forming a complex at the surface [257], potentially reducing the effects of anticancer drugs [258]. For example, in breast cancer it was shown that TDEs express HER2 (human epidermal growth factor receptor 2) and epithelial cell adhesion molecule (EpCAM) antigens. These bind antibodies and render them ineffective [258]. B cell lymphoma secretes exosomes with CD20. The rituximab CD20 antibody therapy for B cell lymphoma is neutralized by these exosome–antigens. In support of this finding, depleting exosomes from the plasma of B cell lymphoma patients increased the efficacy of rituximab [257].
As reported in Sect. 3.1 and 3.2, inhibiting tumor exosome release may be a therapeutic approach in cancer treatment. The synthesis of ceramide (a component of the exosomal membrane) or the secretion of microRNAs could be chemically blocked. GW4869, a chemical blocker, impairs release of both exosomes and pro-inflammatory cytokines through blockage of neutral sphingomyelinase 2, which regulated both ceramide and microRNA secretion. GW4869 has been shown to inhibit tumor exosome release in vitro [259]. Unfortunately, concerns remain about possible systemic off-target effects of inhibiting exosome release or potential undesirable effects of altering microRNA secretion. It would be a lofty aim, and potentially beneficial, to find/develop exosome blockades that are cancer specific or only elicit local inhibition. With evidence emerging regarding the risk linked to pathway inhibitor metagenomics, and assays designed to detect associated exosome profiles, the future of synthetic nucleotides, encapsulation, and vesicle-based gain-of-function strategies would seem both opportune and obvious from the perspective of minimalist bacterial genomes and cells that have been built since 2010 by Craig Venter and his team at Synthetic Genomics (https://www.syntheticgenomics.com/).
APC-derived exosomes carry MHC-I molecules loaded with antigen. These are presented to dendritic cells, mediating T cell activation and T cell-dependent immune responses against the tumor cells. These properties with exosomes as antitumor vaccines support the added benefit of being cell-free [44]. The difficulty with this method is that APCs are very difficult to culture. The specific tumor antigen must be MHC-I haplotype matched and the specific tumor antigens need to be transferred to the APCs [260]. Hence, the exosomes would need to be prepared and collected from APCs for each individual patient based on their MHC-I haplotype and their specific cancer. TDEs can carry shared tumor antigens, not just antigens specific to one tumor [46], which may provide cross-protection against various cancer types. Importantly, these do not require MHC-I haplotyping. For example, human Mucin 1 (hMUC1) has been shown to indicate aggression and poor outcome. Exosomes carrying hMUC1 could be used as a vaccine against MUC1-expressing tumors independent of MHC-I [261]. Another example is the heat treatment of tumor cells to induce HSP70 release in the tumor exosomes. HSP70 stimulates monocytes and dendritic cells, amplifying the immune response further [260]. The HSP70-loaded exosomes could also directly activate NK cells to initiate the apoptosis of tumor cells [10], which has been shown to decrease the size of primary tumors and reduce metastasis in a mouse model [262]. Andre et al. [6] showed that exosomes from melanoma patient ascites activated dendritic cells to produce an antitumor T cell response [6]. Phase I clinical trials are showing promising results for stage III/IV malignant melanoma [263] and non-small lung carcinoma [264].
Competitive implant (metastatic trap [M-Trap]) is an interesting approach where exosomes are embedded in a three-dimensional scaffold and implanted into the peritoneum to establish an artificial pre-metastatic niche. This implant competes with the natural peritoneum for metastatic cells. This process has demonstrated significant efficacy in increasing survival of ovarian cancer patients [265]. An additional mode of treatment currently being investigated is transport of chemo-, microRNA, and biological therapeutics to the tumor cells using natural or synthetic nanoparticles. An example of natural nanoparticle transportation is the packing of paclitaxel into exosomes, which was shown to be 50 times more cytotoxic to drug-resistant lung cancer than conventional paclitaxel [266]. Synthetic nanoparticles could transport biologicals or tumor-suppressive microRNA to tumor cells via liposome preps or mimetics [267, 268] to reduce the toxicity of the chemotherapeutic agents [269]. For example, in a phase III metastatic breast cancer study, administration of liposome-encapsulated doxorubicin HCl with cyclophosphamide allowed a higher median cumulative dose (> 1260 vs. 480 mg/mL) to be safely administered [270, 271]. This methodology allows the use of higher concentrations for release at the tumor sites and has also been investigated in metastatic breast cancer and non-Hodgkin lymphoma. Cells can be transfected with therapeutic microRNAs to change the exosome cargo profile. This re-program could induce apoptosis or increase p53 signaling or direct macrophages [272], to name a few. The exosomes collected from these transfected cells may be used induce desired effects on recipient tissues. Alternatively, the use of anti-acidic approaches for targeting as well as treatment may be combined with known properties of exosome release and delivery [129, 130]. For example, the delivery of proton pump inhibitors (which correct the acidic pH) to reduce exosome release from tumor tissue [130] has been shown to re-sensitize melanoma to cisplatin treatment both in vitro and in vivo [273]. Another approach might utilize pH to control the release of therapeutic substances, such as CO2 [129] or acridine orange [274], to create a hostile tumor environment or targetable environment (e.g., photosensitization) [275].
6 Conclusion
A significant response against cancer cells is offered in antibody-dependent cytotoxicity. As a basis for therapeutics, this process embodies the specificity of the tumor that might accentuate the host immune systems to recognition, regulation, and enhancement against the hostile defense of transitional tumor cells. In this review, the exosomal contribution to cell reaction, immune dodging, and metastatic transition is explored with details, generalities, and specificities. Therein lies the challenge: specificity is a multi-gene transcriptional process, and while there are clearly checkpoint inhibition points that might be exploited in treatment, the accountable differences existing within distinct populations underpin the precision medicine that is being sought.
As a case in point comparing populations of European descent with African Americans, 2210 genes express more than two-fold increases or decreases in non-small cell lung cancer (NSCLC) from African Americans compared with matched normal tissue, while for European American samples, 2921 genes were differentially expressed by more than two-fold simply in NSCLC and normal tissue. These numbers indicate both overlap and non-overlapping specificity. Interestingly, 637 genes were differentially expressed only in African Americans and 1844 in European Americans. These numbers are likely different when derived from the cargo of TDEs, but the questions for precision medicine remain, including “should we concentrate on the overlapping signals for pan-tumor indicators?” or “should we concentrate on the patient-specific indicators?” I suggest that both are equally important approaches for different purposes. Pan-tumor indicators for specific cancer groups throughout ethnicities greatly simplifies the diagnostic biomarker process and could lead to more timely indications of developing cancers. The development of patient-specific indicators is crucial to prognostics and treatments for obvious reasons. Further, does the use of absolute genomics and a full read make sense, or do the transitional states of epitope morphogenesis noted herein bear a possible diferent strategy for exosomal seeding and membrane delivery?
It may be time to rethink how targeted therapy is initiated and monitored when managing cancer patients. Driven by multiple genetic changes and blood-based tests, a changing complex genomic landscape suggests that new first-line combination therapies target a fuller array of mutations to eradicate patient’s cancer before drug resistance can arise [276]. Theorizing that drugs targeting an EGFR mutation, as an example, may be able to wipe out the cells carrying that mutation, this approach risks leaving cells behind that might enhance additional mutations; contrasting the prevailing dogma that some cancers (e.g., NSCLC) are ‘driven’ by only one primary genetic mutation [186]. The single-driver view of cancer was buttressed by cell-free genomic studies to analyze patient blood samples for any mutations in 73 genes known to contribute to cancer. Overall, the researchers analyzed liquid biopsy data from 1122 patients whose tumors contained a mutated EGFR gene and 944 patients whose tumors did not have this mutation. Utilizing basic principles of the exosome platform of cell-free exchange and circulation, it was possible to determine that 92.9% of tumors from patients with advanced-stage lung cancer harbored multiple changes in cancer-related genes in addition to the EGFR driver mutation. On average, tumors contained two to three altered genes in addition to EGFR; however, some contained as many as 13 altered genes. What is the vector that might offer a successful incorporation of something akin to a manifold of identities so that each is seeded under the same tumor diagnostic? Perhaps it is first-line combination therapies capable of targeting multiple genetic pathways in patients’ tumors, rather than waiting for resistance to develop before initiating a subsequent drug. Alternatively, the use of more basic and global tumor properties to diagnose, target, and treat malignant tissue may be a preferred approach [277]. Examples include quantification of TDEs in the circulation and pH targeting or adjustment [132]. The International Society for Extracellular Vesicles (ISEV) has published a paper presenting the organizational opinions regarding clinical and regulatory foresight for EV-based therapeutics [278].
Recognition of the value of exosomes in the context of amplified release, early signaling to distal cells, harbingers of metastatic potential, and opportunity for multiple, minimally invasive access to tumor expression is increasing exponentially. Of note, the very simple property of increased secretion of exosomes from malignant tissue may be indicative as a stand-alone property. Furthermore, use of exosomes as a cell-free delivery of therapeutics might offer broader success and less advantage for tumor-evading genetic shifts. In some ways, the only meaningful strategy is to address the evolving genetic complexity with combinations of post-transcriptional and translational interception of undesirable cell–cell signaling.
References
Kucharzewska P, Belting M. Emerging roles of extracellular vesicles in the adaptive response of tumour cells to microenvironmental stress. J Extracell Vesicles. 2013. https://doi.org/10.3402/jev.v2i0.20304.
Meckes DG Jr. Exosomal communication goes viral. J Virol. 2015;89(10):5200–3. https://doi.org/10.1128/JVI.02470-14.
Barteneva NS, Maltsev N, Vorobjev IA. Microvesicles and intercellular communication in the context of parasitism. Front Cell Infect Microbiol. 2013;3:49. https://doi.org/10.3389/fcimb.2013.00049.
Baranyai T, Herczeg K, Onódi Z, Voszka I, Módos K, Marton N, et al. Isolation of exosomes from blood plasma: qualitative and quantitative comparison of ultracentrifugation and size exclusion chromatography methods. PLoS One. 2015;10(12):e0145686. https://doi.org/10.1371/journal.pone.0145686.
Bard MP, Hegmans JP, Hemmes A, Luider TM, Willemsen R, Severijnen LA, et al. Proteomic analysis of exosomes isolated from human malignant pleural effusions. Am J Respir Cell Mol Biol. 2004;31(1):114–21. https://doi.org/10.1165/rcmb.2003-0238OC.
Andre F, Schartz NE, Movassagh M, Flament C, Pautier P, Morice P, et al. Malignant effusions and immunogenic tumour-derived exosomes. Lancet. 2002;360(9329):295–305. https://doi.org/10.1016/S0140-6736(02)09552-1.
Keller S, Sanderson MP, Stoeck A, Altevogt P. Exosomes: from biogenesis and secretion to biological function. Immunol Lett. 2006;107(2):102–8. https://doi.org/10.1016/j.imlet.2006.09.005.
Lawson C, Vicencio JM, Yellon DM, Davidson SM. Microvesicles and exosomes: new players in metabolic and cardiovascular disease. J Endocrinol. 2016;228(2):R57–71. https://doi.org/10.1530/JOE-15-0201.
Costa-Silva B, Aiello NM, Ocean AJ, Singh S, Zhang H, Thakur BK, et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat Cell Biol. 2015;17(6):816–26. https://doi.org/10.1038/ncb3169.
Kahlert C, Kalluri R. Exosomes in tumor microenvironment influence cancer progression and metastasis. J Mol Med (Berl). 2013;91(4):431–7. https://doi.org/10.1007/s00109-013-1020-6.
Hornick NI, Huan J, Doron B, Goloviznina NA, Lapidus J, Chang BH, et al. Serum exosome MicroRNA as a minimally-invasive early biomarker of AML. Sci Rep. 2015;5:11295. https://doi.org/10.1038/srep11295.
De Toro J, Herschlik L, Waldner C, Mongini C. Emerging roles of exosomes in normal and pathological conditions: new insights for diagnosis and therapeutic applications. Front Immunol. 2015;6:203. https://doi.org/10.3389/fimmu.2015.00203.
Trams EG, Lauter CJ, Salem N Jr, Heine U. Exfoliation of membrane ecto-enzymes in the form of micro-vesicles. Biochim Biophys Acta. 1981;645(1):63–70.
Beach A, Zhang HG, Ratajczak MZ, Kakar SS. Exosomes: an overview of biogenesis, composition and role in ovarian cancer. J Ovarian Res. 2014;7:14. https://doi.org/10.1186/1757-2215-7-14.
Colombo M, Raposo G, Thery C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu Rev Cell Dev Biol. 2014;30:255–89. https://doi.org/10.1146/annurev-cellbio-101512-122326.
Kalra H, Drummen GP, Mathivanan S. Focus on extracellular vesicles: introducing the next small big thing. Int J Mol Sci. 2016;17(2):170. https://doi.org/10.3390/ijms17020170.
Hanson PI, Shim S, Merrill SA. Cell biology of the ESCRT machinery. Curr Opin Cell Biol. 2009;21(4):568–74. https://doi.org/10.1016/j.ceb.2009.06.002.
Mayers JR, Audhya A. Vesicle formation within endosomes: an ESCRT marks the spot. Commun Integr Biol. 2012;5(1):50–6.
Trajkovic K, Hsu C, Chiantia S, Rajendran L, Wenzel D, Wieland F, et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science. 2008;319(5867):1244–7. https://doi.org/10.1126/science.1153124.
Stuffers S, Sem Wegner C, Stenmark H, Brech A. Multivesicular endosome biogenesis in the absence of ESCRTs. Traffic. 2009;10(7):925–37. https://doi.org/10.1111/j.1600-0854.2009.00920.x.
Baietti MF, Zhang Z, Mortier E, Melchior A, Degeest G, Geeraerts A, et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat Cell Biol. 2012;14(7):677–85. https://doi.org/10.1038/ncb2502.
Roucourt B, Meeussen S, Bao J, Zimmermann P, David G. Heparanase activates the syndecan-syntenin-ALIX exosome pathway. Cell Res. 2015;25(4):412–28. https://doi.org/10.1038/cr.2015.29.
Zhang J, Li S, Li L, Li M, Guo C, Yao J, et al. Exosome and exosomal MicroRNA: trafficking, sorting, and function. Genom Proteom Bioinf. 2015;13(1):17–24. https://doi.org/10.1016/j.gpb.2015.02.001.
de Gassart A, Geminard C, Hoekstra D, Vidal M. Exosome secretion: the art of reutilizing nonrecycled proteins? Traffic. 2004;5(11):896–903. https://doi.org/10.1111/j.1600-0854.2004.00223.x.
Raiborg C, Stenmark H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature. 2009;458(7237):445–52. https://doi.org/10.1038/nature07961.
Tamai K, Tanaka N, Nakano T, Kakazu E, Kondo Y, Inoue J, et al. Exosome secretion of dendritic cells is regulated by Hrs, an ESCRT-0 protein. Biochem Biophys Res Commun. 2010;399(3):384–90. https://doi.org/10.1016/j.bbrc.2010.07.083.
Colombo M, Moita C, van Niel G, Kowal J, Vigneron J, Benaroch P, et al. Analysis of ESCRT functions in exosome biogenesis, composition and secretion highlights the heterogeneity of extracellular vesicles. J Cell Sci. 2013;126(Pt 24):5553–65. https://doi.org/10.1242/jcs.128868.
Hurley JH, Odorizzi G. Get on the exosome bus with ALIX. Nat Cell Biol. 2012;14(7):654–5. https://doi.org/10.1038/ncb2530.
Nabhan JF, Hu R, Oh RS, Cohen SN, Lu Q. Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proc Natl Acad Sci USA. 2012;109(11):4146–51. https://doi.org/10.1073/pnas.1200448109.
Stahelin RV, Long F, Diraviyam K, Bruzik KS, Murray D, Cho W. Phosphatidylinositol 3-phosphate induces the membrane penetration of the FYVE domains of Vps27p and Hrs. J Biol Chem. 2002;277(29):26379–88. https://doi.org/10.1074/jbc.M201106200.
Teo H, Gill DJ, Sun J, Perisic O, Veprintsev DB, Vallis Y, et al. ESCRT-I core and ESCRT-II GLUE domain structures reveal role for GLUE in linking to ESCRT-I and membranes. Cell. 2006;125(1):99–111. https://doi.org/10.1016/j.cell.2006.01.047.
Ghossoub R, Lembo F, Rubio A, Gaillard CB, Bouchet J, Vitale N, et al. Syntenin-ALIX exosome biogenesis and budding into multivesicular bodies are controlled by ARF6 and PLD2. Nat Commun. 2014;5:3477. https://doi.org/10.1038/ncomms4477.
Perez-Hernandez D, Gutierrez-Vazquez C, Jorge I, Lopez-Martin S, Ursa A, Sanchez-Madrid F, et al. The intracellular interactome of tetraspanin-enriched microdomains reveals their function as sorting machineries toward exosomes. J Biol Chem. 2013;288(17):11649–61. https://doi.org/10.1074/jbc.M112.445304.
Harding C, Heuser J, Stahl P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. J Cell Biol. 1983;97(2):329–39.
Pan BT, Teng K, Wu C, Adam M, Johnstone RM. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. J Cell Biol. 1985;101(3):942–8.
Barr F, Lambright DG. Rab GEFs and GAPs. Curr Opin Cell Biol. 2010;22(4):461–70. https://doi.org/10.1016/j.ceb.2010.04.007.
Stenmark H. Rab GTPases as coordinators of vesicle traffic. Nat Rev Mol Cell Biol. 2009;10(8):513–25. https://doi.org/10.1038/nrm2728.
Ostrowski M, Carmo NB, Krumeich S, Fanget I, Raposo G, Savina A, et al. Rab27a and Rab27b control different steps of the exosome secretion pathway. Nat Cell Biol. 2010;12(1):19–30. https://doi.org/10.1038/ncb2000.
Hoshino D, Kirkbride KC, Costello K, Clark ES, Sinha S, Grega-Larson N, et al. Exosome secretion is enhanced by invadopodia and drives invasive behavior. Cell Rep. 2013;5(5):1159–68. https://doi.org/10.1016/j.celrep.2013.10.050.
Martin-Cofreces NB, Baixauli F, Sanchez-Madrid F. Immune synapse: conductor of orchestrated organelle movement. Trends Cell Biol. 2014;24(1):61–72. https://doi.org/10.1016/j.tcb.2013.09.005.
Record M, Carayon K, Poirot M, Silvente-Poirot S. Exosomes as new vesicular lipid transporters involved in cell-cell communication and various pathophysiologies. Biochim Biophys Acta. 2014;1841(1):108–20. https://doi.org/10.1016/j.bbalip.2013.10.004.
Hsu C, Morohashi Y, Yoshimura S, Manrique-Hoyos N, Jung S, Lauterbach MA, et al. Regulation of exosome secretion by Rab35 and its GTPase-activating proteins TBC1D10A-C. J Cell Biol. 2010;189(2):223–32. https://doi.org/10.1083/jcb.200911018.
Raposo G, Nijman HW, Stoorvogel W, Liejendekker R, Harding CV, Melief CJ, et al. B lymphocytes secrete antigen-presenting vesicles. J Exp Med. 1996;183(3):1161–72.
Zitvogel L, Regnault A, Lozier A, Wolfers J, Flament C, Tenza D, et al. Eradication of established murine tumors using a novel cell-free vaccine: dendritic cell-derived exosomes. Nat Med. 1998;4(5):594–600.
van Niel G, Raposo G, Candalh C, Boussac M, Hershberg R, Cerf-Bensussan N, et al. Intestinal epithelial cells secrete exosome-like vesicles. Gastroenterology. 2001;121(2):337–49.
Wolfers J, Lozier A, Raposo G, Regnault A, Thery C, Masurier C, et al. Tumor-derived exosomes are a source of shared tumor rejection antigens for CTL cross-priming. Nat Med. 2001;7(3):297–303. https://doi.org/10.1038/85438.
Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2(8):569–79. https://doi.org/10.1038/nri855.
Théry C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Current protocols in cell biology. New York: Wiley; 2001.
Steinbichler TB, Dudas J, Riechelmann H, Skvortsova II. The role of exosomes in cancer metastasis. Semin Cancer Biol. 2017;44:170–81. https://doi.org/10.1016/j.semcancer.2017.02.006.
Fitzner D, Schnaars M, van Rossum D, Krishnamoorthy G, Dibaj P, Bakhti M, et al. Selective transfer of exosomes from oligodendrocytes to microglia by macropinocytosis. J Cell Sci. 2011;124(Pt 3):447–58. https://doi.org/10.1242/jcs.074088.
Fruhbeis C, Frohlich D, Kuo WP, Kramer-Albers EM. Extracellular vesicles as mediators of neuron-glia communication. Front Cell Neurosci. 2013;7:182. https://doi.org/10.3389/fncel.2013.00182.
Morelli AE, Larregina AT, Shufesky WJ, Sullivan ML, Stolz DB, Papworth GD, et al. Endocytosis, intracellular sorting, and processing of exosomes by dendritic cells. Blood. 2004;104(10):3257–66. https://doi.org/10.1182/blood-2004-03-0824.
Nazarenko I, Rana S, Baumann A, McAlear J, Hellwig A, Trendelenburg M, et al. Cell surface tetraspanin Tspan8 contributes to molecular pathways of exosome-induced endothelial cell activation. Cancer Res. 2010;70(4):1668–78. https://doi.org/10.1158/0008-5472.CAN-09-2470.
Saunderson SC, Dunn AC, Crocker PR, McLellan AD. CD169 mediates the capture of exosomes in spleen and lymph node. Blood. 2014;123(2):208–16. https://doi.org/10.1182/blood-2013-03-489732.
Christianson HC, Svensson KJ, van Kuppevelt TH, Li JP, Belting M. Cancer cell exosomes depend on cell-surface heparan sulfate proteoglycans for their internalization and functional activity. Proc Natl Acad Sci USA. 2013;110(43):17380–5. https://doi.org/10.1073/pnas.1304266110.
Palma J, Yaddanapudi SC, Pigati L, Havens MA, Jeong S, Weiner GA, et al. MicroRNAs are exported from malignant cells in customized particles. Nucleic Acids Res. 2012;40(18):9125–38. https://doi.org/10.1093/nar/gks656.
Sreekumar PG, Kannan R, Kitamura M, Spee C, Barron E, Ryan SJ, et al. alphaB crystallin is apically secreted within exosomes by polarized human retinal pigment epithelium and provides neuroprotection to adjacent cells. PLoS One. 2010;5(10):e12578. https://doi.org/10.1371/journal.pone.0012578.
Tauro BJ, Greening DW, Mathias RA, Mathivanan S, Ji H, Simpson RJ. Two distinct populations of exosomes are released from LIM1863 colon carcinoma cell-derived organoids. Mol Cell Proteomics. 2013;12(3):587–98. https://doi.org/10.1074/mcp.M112.021303.
Gyorgy B, Szabo TG, Pasztoi M, Pal Z, Misjak P, Aradi B, et al. Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci. 2011;68(16):2667–88. https://doi.org/10.1007/s00018-011-0689-3.
Taylor DD, Shah S. Methods of isolating extracellular vesicles impact down-stream analyses of their cargoes. Methods. 2015;87:3–10. https://doi.org/10.1016/j.ymeth.2015.02.019.
Witwer KW, Buzas EI, Bemis LT, Bora A, Lasser C, Lotvall J, et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J Extracell Vesicles. 2013. https://doi.org/10.3402/jev.v2i0.20360.
Böing AN, van der Pol E, Grootemaat AE, Coumans FAW, Sturk A, Nieuwland R. Single-step isolation of extracellular vesicles by size-exclusion chromatography. J Extracell Vesicles. 2014. https://doi.org/10.3402/jev.v3.23430.
Momen-Heravi F, Balaj L, Alian S, Trachtenberg AJ, Hochberg FH, Skog J, et al. Impact of biofluid viscosity on size and sedimentation efficiency of the isolated microvesicles. Fronti Physiol. 2012;3:162. https://doi.org/10.3389/fphys.2012.00162.
Théry C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol. 2006;3:22. https://doi.org/10.1002/0471143030.cb0322s30.
Zarovni N, Corrado A, Guazzi P, Zocco D, Lari E, Radano G, et al. Integrated isolation and quantitative analysis of exosome shuttled proteins and nucleic acids using immunocapture approaches. Methods. 2015;87:46–58. https://doi.org/10.1016/j.ymeth.2015.05.028.
Gardiner C, Di Vizio D, Sahoo S, Théry C, Witwer KW, Wauben M, et al. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J Extracell Vesicles. 2016;5(1):32945. https://doi.org/10.3402/jev.v5.32945.
Li P, Kaslan M, Lee SH, Yao J, Gao Z. Progress in exosome isolation techniques. Theranostics. 2017;7(3):789–804. https://doi.org/10.7150/thno.18133.
Rood IM, Deegens JK, Merchant ML, Tamboer WP, Wilkey DW, Wetzels JF, et al. Comparison of three methods for isolation of urinary microvesicles to identify biomarkers of nephrotic syndrome. Kidney Int. 2010;78(8):810–6. https://doi.org/10.1038/ki.2010.262.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9(6):654–9. https://doi.org/10.1038/ncb1596.
Johnsen KB, Gudbergsson JM, Skov MN, Pilgaard L, Moos T, Duroux M. A comprehensive overview of exosomes as drug delivery vehicles—endogenous nanocarriers for targeted cancer therapy. Biochim Biophys Acta. 2014;1846(1):75–87. https://doi.org/10.1016/j.bbcan.2014.04.005.
Alvarez ML, Khosroheidari M, Kanchi Ravi R, DiStefano JK. Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int. 2012;82(9):1024–32. https://doi.org/10.1038/ki.2012.256.
Li M, Zeringer E, Barta T, Schageman J, Cheng A, Vlassov AV. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos Trans R Soc Lond B Biol Sci. 2014;369(1652):20130502. https://doi.org/10.1098/rstb.2013.0502.
Neurauter A, Kullmann A, Kierulf B, Oksvold MP, Pedersen KW. Magnetic beads for fast and reproducible isolation/characterization of exosomes based on surface protein expression. Institute for Cancer Research, Oslo University Hospital: Life Technologies; 2013. https://assets.thermofisher.com/TFS-Assets/CMD/posters/Exosome-poster-ISEV-2013-Boston.pdf. Accessed 15 Feb 2019.
Tauro BJ, Greening DW, Mathias RA, Ji H, Mathivanan S, Scott AM, et al. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods. 2012;56(2):293–304. https://doi.org/10.1016/j.ymeth.2012.01.002.
Koliha N, Wiencek Y, Heider U, Jungst C, Kladt N, Krauthauser S, et al. A novel multiplex bead-based platform highlights the diversity of extracellular vesicles. J Extracell Vesicles. 2016;5:29975. https://doi.org/10.3402/jev.v5.29975.
Chen C, Skog J, Hsu CH, Lessard RT, Balaj L, Wurdinger T, et al. Microfluidic isolation and transcriptome analysis of serum microvesicles. Lab Chip. 2010;10(4):505–11. https://doi.org/10.1039/b916199f.
Wu M, Ouyang Y, Wang Z, Zhang R, Huang P-H, Chen C, et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc Natl Acad Sci USA. 2017;114(40):10584–9. https://doi.org/10.1073/pnas.1709210114.
Enderle D, Spiel A, Coticchia CM, Berghoff E, Mueller R, Schlumpberger M, Sprenger-Haussels M, Shaffer JM, Lader E, Skog J, Noerholm M. Characterization of RNA from exosomes and other extracellular vesicles isolated by a novel spin column-based method. PLoS One. 2015;10(8):e0136133. https://doi.org/10.1371/journal.pone.0136133.
Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol. 2013;200(4):373–83. https://doi.org/10.1083/jcb.201211138.
Gámez-Valero A, Monguió-Tortajada M, Carreras-Planella L, Franquesa ML, Beyer K, Borràs FE. Size-exclusion chromatography-based isolation minimally alters extracellular vesicles’ characteristics compared to precipitating agents. Sci Rep. 2016;6:33641. https://doi.org/10.1038/srep33641.
Lobb RJ, Becker M, Wen SW, Wong CS, Wiegmans AP, Leimgruber A, et al. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles. 2015;4:27031. https://doi.org/10.3402/jev.v4.27031.
Muller L, Hong CS, Stolz DB, Watkins SC, Whiteside TL. Isolation of biologically-active exosomes from human plasma. J Immunol Methods. 2014;411:55–65. https://doi.org/10.1016/j.jim.2014.06.007.
Nordin JZ, Lee Y, Vader P, Mager I, Johansson HJ, Heusermann W, et al. Ultrafiltration with size-exclusion liquid chromatography for high yield isolation of extracellular vesicles preserving intact biophysical and functional properties. Nanomedicine. 2015;11(4):879–83. https://doi.org/10.1016/j.nano.2015.01.003.
Hagel LÖM, Andersson T. Apparent pore size distributions of chromatography media. J Chrom A. 1996;743:33–42.
Williams AHL. Column handbook for size exclusion chromatography. Philadelphia: Academic Press; 1999.
O’Brien JR. Cell membrane damage, platelet stickiness and some effects of aspirin. Br J Haematol. 1969;17(6):610–1.
Ogawa Y, Kanai-Azuma M, Akimoto Y, Kawakami H, Yanoshita R. Exosome-like vesicles with dipeptidyl peptidase IV in human saliva. Biol Pharm Bull. 2008;31(6):1059–62.
Sokolova V, Ludwig AK, Hornung S, Rotan O, Horn PA, Epple M, et al. Characterisation of exosomes derived from human cells by nanoparticle tracking analysis and scanning electron microscopy. Colloids Surf B Biointerfaces. 2011;87(1):146–50. https://doi.org/10.1016/j.colsurfb.2011.05.013.
Taylor DD, Chou IN, Black PH. Isolation of plasma membrane fragments from cultured murine melanoma cells. Biochem Biophys Res Commun. 1983;113(2):470–6.
Taylor DD, Zacharias W, Gercel-Taylor C. Exosome isolation for proteomic analyses and RNA profiling. Methods Mol Biol. 2011;728:235–46. https://doi.org/10.1007/978-1-61779-068-3_15.
Pietrowska M, Funk S, Gawin M, Marczak Ł, Abramowicz A, Widłak P, et al. Isolation of exosomes for the purpose of protein cargo analysis with the use of mass spectrometry. In: Kaufmann M, Klinger C, Savelsbergh A, editors. Functional genomics: methods and protocols. New York: Springer; 2017. p. 291–307.
Worst TS, von Hardenberg J, Gross JC, Erben P, Schnölzer M, Hausser I, et al. Database-augmented mass spectrometry analysis of exosomes identifies claudin 3 as a putative prostate cancer biomarker. Mol Cell Proteomics. 2017;16(6):998–1008. https://doi.org/10.1074/mcp.M117.068577.
Haraszti RA, Didiot M-C, Sapp E, Leszyk J, Shaffer SA, Rockwell HE, et al. High-resolution proteomic and lipidomic analysis of exosomes and microvesicles from different cell sources. J Extracell Vesicles. 2016;5:32570. https://doi.org/10.3402/jev.v5.32570.
Yang J, Hagen J, Guntur KV, Allette K, Schuyler S, Ranjan J, et al. A next generation sequencing based approach to identify extracellular vesicle mediated mRNA transfers between cells. BMC Genomics. 2017;18(1):987. https://doi.org/10.1186/s12864-017-4359-1.
Ogawa Y, Tsujimoto M, Yanoshita R. Next-generation sequencing of protein-coding and long non-protein-coding RNAs in two types of exosomes derived from human whole saliva. Biol Pharm Bull. 2016;39(9):1496–507. https://doi.org/10.1248/bpb.b16-00297.
Jenjaroenpun P, Kremenska Y, Nair VM, Kremenskoy M, Joseph B, Kurochkin IV. Characterization of RNA in exosomes secreted by human breast cancer cell lines using next-generation sequencing. PeerJ. 2013;1:e201. https://doi.org/10.7717/peerj.201.
Vong S, Kalluri R. The role of stromal myofibroblast and extracellular matrix in tumor angiogenesis. Genes Cancer. 2011;2(12):1139–45. https://doi.org/10.1177/1947601911423940.
Webber JP, Spary LK, Sanders AJ, Chowdhury R, Jiang WG, Steadman R, et al. Differentiation of tumour-promoting stromal myofibroblasts by cancer exosomes. Oncogene. 2015;34(3):290–302. https://doi.org/10.1038/onc.2013.560.
Orimo A, Gupta PB, Sgroi DC, Arenzana-Seisdedos F, Delaunay T, Naeem R, et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell. 2005;121(3):335–48. https://doi.org/10.1016/j.cell.2005.02.034.
Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Curry WT, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10(12):1470–6. https://doi.org/10.1038/ncb1800.
Brentnall TA, Lai LA, Coleman J, Bronner MP, Pan S, Chen R. Arousal of cancer-associated stroma: overexpression of palladin activates fibroblasts to promote tumor invasion. PLoS One. 2012;7(1):e30219. https://doi.org/10.1371/journal.pone.0030219.
Webber J, Steadman R, Mason MD, Tabi Z, Clayton A. Cancer exosomes trigger fibroblast to myofibroblast differentiation. Cancer Res. 2010;70(23):9621–30. https://doi.org/10.1158/0008-5472.CAN-10-1722.
Luga V, Wrana JL. Tumor-stroma interaction: revealing fibroblast-secreted exosomes as potent regulators of Wnt-planar cell polarity signaling in cancer metastasis. Cancer Res. 2013;73(23):6843–7. https://doi.org/10.1158/0008-5472.CAN-13-1791.
Park JE, Tan HS, Datta A, Lai RC, Zhang H, Meng W, et al. Hypoxic tumor cell modulates its microenvironment to enhance angiogenic and metastatic potential by secretion of proteins and exosomes. Mol Cell Proteomics. 2010;9(6):1085–99. https://doi.org/10.1074/mcp.M900381-MCP200.
Min L, Shen J, Tu C, Hornicek F, Duan Z. The roles and implications of exosomes in sarcoma. Cancer Metastasis Rev. 2016;35(3):377–90. https://doi.org/10.1007/s10555-016-9630-4.
You Y, Shan Y, Chen J, Yue H, You B, Shi S, et al. Matrix metalloproteinase 13-containing exosomes promote nasopharyngeal carcinoma metastasis. Cancer Sci. 2015;106(12):1669–77. https://doi.org/10.1111/cas.12818.
Borsig L, Wong R, Feramisco J, Nadeau DR, Varki NM, Varki A. Heparin and cancer revisited: mechanistic connections involving platelets, P-selectin, carcinoma mucins, and tumor metastasis. Proc Natl Acad Sci USA. 2001;98(6):3352–7. https://doi.org/10.1073/pnas.061615598.
Nieswandt B, Hafner M, Echtenacher B, Mannel DN. Lysis of tumor cells by natural killer cells in mice is impeded by platelets. Cancer Res. 1999;59(6):1295–300.
Labelle M, Begum S, Hynes RO. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell. 2011;20(5):576–90. https://doi.org/10.1016/j.ccr.2011.09.009.
Karpatkin S, Pearlstein E, Ambrogio C, Coller BS. Role of adhesive proteins in platelet tumor interaction in vitro and metastasis formation in vivo. J Clin Invest. 1988;81(4):1012–9. https://doi.org/10.1172/JCI113411.
Sierko E, Wojtukiewicz MZ. Inhibition of platelet function: does it offer a chance of better cancer progression control? Semin Thromb Hemost. 2007;33(7):712–21. https://doi.org/10.1055/s-2007-991540.
Dean WL, Lee MJ, Cummins TD, Schultz DJ, Powell DW. Proteomic and functional characterisation of platelet microparticle size classes. Thromb Haemost. 2009;102(4):711–8. https://doi.org/10.1160/TH09-04-243.
Dovizio M, Alberti S, Sacco A, Guillem-Llobat P, Schiavone S, Maier TJ, et al. Novel insights into the regulation of cyclooxygenase-2 expression by platelet-cancer cell cross-talk. Biochem Soc Trans. 2015;43(4):707–14. https://doi.org/10.1042/BST20140322.
Gay LJ, Felding-Habermann B. Platelets alter tumor cell attributes to propel metastasis: programming in transit. Cancer Cell. 2011;20(5):553–4. https://doi.org/10.1016/j.ccr.2011.11.001.
Matsumoto Y, Kano M, Akutsu Y, Hanari N, Hoshino I, Murakami K, et al. Quantification of plasma exosome is a potential prognostic marker for esophageal squamous cell carcinoma. Oncol Rep. 2016;36(5):2535–43. https://doi.org/10.3892/or.2016.5066.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105(30):10513–8. https://doi.org/10.1073/pnas.0804549105.
Silva J, Garcia V, Rodriguez M, Compte M, Cisneros E, Veguillas P, et al. Analysis of exosome release and its prognostic value in human colorectal cancer. Genes Chromosomes Cancer. 2012;51(4):409–18.
Logozzi M, De Milito A, Lugini L, Borghi M, Calabro L, Spada M, et al. High levels of exosomes expressing CD63 and caveolin-1 in plasma of melanoma patients. PLoS One. 2009;4(4):e5219. https://doi.org/10.1371/journal.pone.0005219.
King HW, Michael MZ, Gleadle JM. Hypoxic enhancement of exosome release by breast cancer cells. BMC Cancer. 2012;12:421. https://doi.org/10.1186/1471-2407-12-421.
Ban JJ, Lee M, Im W, Kim M. Low pH increases the yield of exosome isolation. Biochem Biophys Res Commun. 2015;461(1):76–9. https://doi.org/10.1016/j.bbrc.2015.03.172.
Parolini I, Federici C, Raggi C, Lugini L, Palleschi S, De Milito A, et al. Microenvironmental pH is a key factor for exosome traffic in tumor cells. J Biol Chem. 2009;284(49):34211–22. https://doi.org/10.1074/jbc.M109.041152.
Gutierrez-Vazquez C, Villarroya-Beltri C, Mittelbrunn M, Sanchez-Madrid F. Transfer of extracellular vesicles during immune cell-cell interactions. Immunol Rev. 2013;251(1):125–42. https://doi.org/10.1111/imr.12013.
Qian BZ, Pollard JW. Macrophage diversity enhances tumor progression and metastasis. Cell. 2010;141(1):39–51. https://doi.org/10.1016/j.cell.2010.03.014.
Kang Y. Dissecting tumor-stromal interactions in breast cancer bone metastasis. Endocrinol Metab (Seoul). 2016;31(2):206–12. https://doi.org/10.3803/EnM.2016.31.2.206.
Subramanian A, Gupta V, Sarkar S, Maity G, Banerjee S, Ghosh A, et al. Exosomes in carcinogenesis: molecular palkis carry signals for the regulation of cancer progression and metastasis. J Cell Commun Signal. 2016;10(3):241–9. https://doi.org/10.1007/s12079-016-0338-6.
Ojalvo LS, Whittaker CA, Condeelis JS, Pollard JW. Gene expression analysis of macrophages that facilitate tumor invasion supports a role for Wnt-signaling in mediating their activity in primary mammary tumors. J Immunol. 2010;184(2):702–12. https://doi.org/10.4049/jimmunol.0902360.
Chow A, Zhou W, Liu L, Fong MY, Champer J, Van Haute D, et al. Macrophage immunomodulation by breast cancer-derived exosomes requires Toll-like receptor 2-mediated activation of NF-kappaB. Sci Rep. 2014;4:5750. https://doi.org/10.1038/srep05750.
de Jong OG, Verhaar MC, Chen Y, Vader P, Gremmels H, Posthuma G, et al. Cellular stress conditions are reflected in the protein and RNA content of endothelial cell-derived exosomes. J Extracell Vesicles. 2012. https://doi.org/10.3402/jev.v1i0.18396.
Fais S, Venturi G, Gatenby B. Microenvironmental acidosis in carcinogenesis and metastases: new strategies in prevention and therapy. Cancer Metastasis Rev. 2014;33(4):1095–108. https://doi.org/10.1007/s10555-014-9531-3.
Spugnini E, Fais S. Proton pump inhibition and cancer therapeutics: a specific tumor targeting or it is a phenomenon secondary to a systemic buffering? Semin Cancer Biol. 2017;43:111–8. https://doi.org/10.1016/j.semcancer.2017.01.003.
Logozzi M, Angelini DF, Iessi E, Mizzoni D, Di Raimo R, Federici C, et al. Increased PSA expression on prostate cancer exosomes in in vitro condition and in cancer patients. Cancer Lett. 2017;403:318–29. https://doi.org/10.1016/j.canlet.2017.06.036.
Logozzi M, Mizzoni D, Angelini DF, Di Raimo R, Falchi M, Battistini L, et al. Microenvironmental pH and exosome levels interplay in human cancer cell lines of different histotypes. Cancers (Basel). 2018. https://doi.org/10.3390/cancers10100370.
Hedlund M, Nagaeva O, Kargl D, Baranov V, Mincheva-Nilsson L. Thermal- and oxidative stress causes enhanced release of NKG2D ligand-bearing immunosuppressive exosomes in leukemia/lymphoma T and B cells. PLoS One. 2011;6(2):e16899. https://doi.org/10.1371/journal.pone.0016899.
Salomon C, Ryan J, Sobrevia L, Kobayashi M, Ashman K, Mitchell M, et al. Exosomal signaling during hypoxia mediates microvascular endothelial cell migration and vasculogenesis. PLoS One. 2013;8(7):e68451. https://doi.org/10.1371/journal.pone.0068451.
Kucharzewska P, Christianson HC, Welch JE, Svensson KJ, Fredlund E, Ringner M, et al. Exosomes reflect the hypoxic status of glioma cells and mediate hypoxia-dependent activation of vascular cells during tumor development. Proc Natl Acad Sci U S A. 2013;110(18):7312–7. https://doi.org/10.1073/pnas.1220998110.
Berchem G, Noman MZ, Bosseler M, Paggetti J, Baconnais S, Le Cam E, et al. Hypoxic tumor-derived microvesicles negatively regulate NK cell function by a mechanism involving TGF-beta and miR23a transfer. Oncoimmunology. 2016;5(4):e1062968. https://doi.org/10.1080/2162402X.2015.1062968.
Chen L, Wang Y, Pan Y, Zhang L, Shen C, Qin G, et al. Cardiac progenitor-derived exosomes protect ischemic myocardium from acute ischemia/reperfusion injury. Biochem Biophys Res Commun. 2013;431(3):566–71. https://doi.org/10.1016/j.bbrc.2013.01.015.
Arslan F, Lai RC, Smeets MB, Akeroyd L, Choo A, Aguor EN, et al. Mesenchymal stem cell-derived exosomes increase ATP levels, decrease oxidative stress and activate PI3K/Akt pathway to enhance myocardial viability and prevent adverse remodeling after myocardial ischemia/reperfusion injury. Stem Cell Res. 2013;10(3):301–12. https://doi.org/10.1016/j.scr.2013.01.002.
Sinha A, Alfaro J, Kislinger T. Characterization of protein content present in exosomes isolated from conditioned media and urine. Curr Protoc Protein Sci. 2017;87(1):24–9. https://doi.org/10.1002/cpps.23.
ExoCarta: exosome protein, RNA and lipid database [database on the Internet]. http://www.exocarta.org/. Accessed 19 Dec 2018.
Wubbolts R, Leckie RS, Veenhuizen PT, Schwarzmann G, Mobius W, Hoernschemeyer J, et al. Proteomic and biochemical analyses of human B cell-derived exosomes. Potential implications for their function and multivesicular body formation. J Biol Chem. 2003;278(13):10963–72. https://doi.org/10.1074/jbc.M207550200.
Øverbye A, Skotland T, Koehler CJ, Thiede B, Seierstad T, Berge V, et al. Identification of prostate cancer biomarkers in urinary exosomes. Oncotarget. 2015;6(30):30357–76.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523(7559):177–82. https://doi.org/10.1038/nature14581.
Hu J, Sheng Y, Kwak KJ, Shi J, Yu B, Lee LJ. A signal-amplifiable biochip quantifies extracellular vesicle-associated RNAs for early cancer detection. Nat Commun. 2017;8(1):1683. https://doi.org/10.1038/s41467-017-01942-1.
Yang KS, Im H, Hong S, Pergolini I, Del Castillo AF, Wang R, et al. Multiparametric plasma EV profiling facilitates diagnosis of pancreatic malignancy. Sci Transl Med. 2017. https://doi.org/10.1126/scitranslmed.aal3226.
Lai X, Wang M, McElyea SD, Sherman S, House M, Korc M. A microRNA signature in circulating exosomes is superior to exosomal glypican-1 levels for diagnosing pancreatic cancer. Cancer Lett. 2017;393:86–93. https://doi.org/10.1016/j.canlet.2017.02.019.
Chaudhary SC, Khalid S, Smethurst V, Monier D, Mobley J, Huet A, et al. Proteomic profiling of extracellular vesicles released from vascular smooth muscle cells during initiation of phosphate-induced mineralization. Connect Tissue Res. 2018;59(sup1):55–61. https://doi.org/10.1080/03008207.2018.1444759.
Poliakov A, Spilman M, Dokland T, Amling CL, Mobley JA. Structural heterogeneity and protein composition of exosome-like vesicles (prostasomes) in human semen. Prostate. 2009;69(2):159–67. https://doi.org/10.1002/pros.20860.
Hong CS, Muller L, Boyiadzis M, Whiteside TL. Isolation and characterization of CD34+ blast-derived exosomes in acute myeloid leukemia. PLoS One. 2014;9(8):e103310. https://doi.org/10.1371/journal.pone.0103310.
Al-Nedawi K, Meehan B, Rak J. Microvesicles: messengers and mediators of tumor progression. Cell Cycle. 2009;8(13):2014–8. https://doi.org/10.4161/cc.8.13.8988.
Liu Q, Yu Z, Yuan S, Xie W, Li C, Hu Z, et al. Circulating exosomal microRNAs as prognostic biomarkers for non-small-cell lung cancer. Oncotarget. 2017;8(8):13048–58. https://doi.org/10.18632/oncotarget.14369.
Wang M, Ji S, Shao G, Zhang J, Zhao K, Wang Z, et al. Effect of exosome biomarkers for diagnosis and prognosis of breast cancer patients. Clin Transl Oncol. 2018;20(7):906–11. https://doi.org/10.1007/s12094-017-1805-0.
Sun Y, Zheng W, Guo Z, Ju Q, Zhu L, Gao J, et al. A novel TP53 pathway influences the HGS-mediated exosome formation in colorectal cancer. Sci Rep. 2016;6:28083. https://doi.org/10.1038/srep28083.
McCready J, Sims JD, Chan D, Jay DG. Secretion of extracellular hsp90alpha via exosomes increases cancer cell motility: a role for plasminogen activation. BMC Cancer. 2010;10:294. https://doi.org/10.1186/1471-2407-10-294.
Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M, et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527(7578):329–35. https://doi.org/10.1038/nature15756.
Lea J, Sharma R, Yang F, Zhu H, Ward ES, Schroit AJ. Detection of phosphatidylserine-positive exosomes as a diagnostic marker for ovarian malignancies: a proof of concept study. Oncotarget. 2017;8(9):14395–407. https://doi.org/10.18632/oncotarget.14795.
Yang J, Liu W, Lu X, Fu Y, Li L, Luo Y. High expression of small GTPase Rab3D promotes cancer progression and metastasis. Oncotarget. 2015;6(13):11125–38. https://doi.org/10.18632/oncotarget.3575.
Hong CS, Muller L, Whiteside TL, Boyiadzis M. Plasma exosomes as markers of therapeutic response in patients with acute myeloid leukemia. Front Immunol. 2014;5:160. https://doi.org/10.3389/fimmu.2014.00160.
Ko H, Jeon H, Lee D, Choi HK, Kang KS, Choi KC. Sanguiin H6 suppresses TGF-beta induction of the epithelial-mesenchymal transition and inhibits migration and invasion in A549 lung cancer. Bioorg Med Chem Lett. 2015;25(23):5508–13. https://doi.org/10.1016/j.bmcl.2015.10.067.
Wang T, Ning K, Lu TX, Sun X, Jin L, Qi X, et al. Increasing circulating exosomes-carrying TRPC5 predicts chemoresistance in metastatic breast cancer patients. Cancer Sci. 2017;108(3):448–54. https://doi.org/10.1111/cas.13150.
Guescini M, Guidolin D, Vallorani L, Casadei L, Gioacchini AM, Tibollo P, et al. C2C12 myoblasts release micro-vesicles containing mtDNA and proteins involved in signal transduction. Exp Cell Res. 2010;316(12):1977–84. https://doi.org/10.1016/j.yexcr.2010.04.006.
Balaj L, Lessard R, Dai L, Cho YJ, Pomeroy SL, Breakefield XO, et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Commun. 2011;2:180. https://doi.org/10.1038/ncomms1180.
Kahlert C, Melo SA, Protopopov A, Tang J, Seth S, Koch M, et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem. 2014;289(7):3869–75. https://doi.org/10.1074/jbc.C113.532267.
Ratajczak J, Miekus K, Kucia M, Zhang J, Reca R, Dvorak P, et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia. 2006;20(5):847–56. https://doi.org/10.1038/sj.leu.2404132.
Deregibus MC, Cantaluppi V, Calogero R, Lo Iacono M, Tetta C, Biancone L, et al. Endothelial progenitor cell derived microvesicles activate an angiogenic program in endothelial cells by a horizontal transfer of mRNA. Blood. 2007;110(7):2440–8. https://doi.org/10.1182/blood-2007-03-078709.
Kislauskis EH, Zhu X, Singer RH. Sequences responsible for intracellular localization of beta-actin messenger RNA also affect cell phenotype. J Cell Biol. 1994;127(2):441–51.
Batagov AO, Kuznetsov VA, Kurochkin IV. Identification of nucleotide patterns enriched in secreted RNAs as putative cis-acting elements targeting them to exosome nano-vesicles. BMC Genomics. 2011;12(Suppl 3):S18. https://doi.org/10.1186/1471-2164-12-S3-S18.
Batagov AO, Kurochkin IV. Exosomes secreted by human cells transport largely mRNA fragments that are enriched in the 3’-untranslated regions. Biol Direct. 2013;8:12. https://doi.org/10.1186/1745-6150-8-12.
Andreassi C, Riccio A. To localize or not to localize: mRNA fate is in 3’UTR ends. Trends Cell Biol. 2009;19(9):465–74. https://doi.org/10.1016/j.tcb.2009.06.001.
Zhou W, Fong MY, Min Y, Somlo G, Liu L, Palomares MR, et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell. 2014;25(4):501–15. https://doi.org/10.1016/j.ccr.2014.03.007.
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26(5):707–21. https://doi.org/10.1016/j.ccell.2014.09.005.
Kawahara Y, Zinshteyn B, Sethupathy P, Iizasa H, Hatzigeorgiou AG, Nishikura K. Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science. 2007;315(5815):1137–40. https://doi.org/10.1126/science.1138050.
Velazquez-Torres G, Shoshan E, Ivan C, Huang L, Fuentes-Mattei E, Paret H, et al. A-to-I miR-378a-3p editing can prevent melanoma progression via regulation of PARVA expression. Nat Commun. 2018;9(1):461. https://doi.org/10.1038/s41467-018-02851-7.
Paul D, Sinha AN, Ray A, Lal M, Nayak S, Sharma A, et al. A-to-I editing in human miRNAs is enriched in seed sequence, influenced by sequence contexts and significantly hypoedited in glioblastoma multiforme. Sci Rep. 2017;7(1):2466. https://doi.org/10.1038/s41598-017-02397-6.
Wang Y, Xu X, Yu S, Jeong KJ, Zhou Z, Han L, et al. Systematic characterization of A-to-I RNA editing hotspots in microRNAs across human cancers. Genome Res. 2017;27(7):1112–25. https://doi.org/10.1101/gr.219741.116.
Pinto Y, Buchumenski I, Levanon EY, Eisenberg E. Human cancer tissues exhibit reduced A-to-I editing of miRNAs coupled with elevated editing of their targets. Nucleic Acids Res. 2018;46(1):71–82. https://doi.org/10.1093/nar/gkx1176.
Nigita G, Distefano R, Veneziano D, Romano G, Rahman M, Wang K, et al. Tissue and exosomal miRNA editing in non-small cell lung cancer. Sci Rep. 2018;8(1):10222. https://doi.org/10.1038/s41598-018-28528-1.
Shen J, Hruby GW, McKiernan JM, Gurvich I, Lipsky MJ, Benson MC, et al. Dysregulation of circulating microRNAs and prediction of aggressive prostate cancer. Prostate. 2012;72(13):1469–77. https://doi.org/10.1002/pros.22499.
Manier S, Liu CJ, Avet-Loiseau H, Park J, Shi J, Campigotto F, et al. Prognostic role of circulating exosomal miRNAs in multiple myeloma. Blood. 2017;129(17):2429–36. https://doi.org/10.1182/blood-2016-09-742296.
Sanchez CA, Andahur EI, Valenzuela R, Castellon EA, Fulla JA, Ramos CG, et al. Exosomes from bulk and stem cells from human prostate cancer have a differential microRNA content that contributes cooperatively over local and pre-metastatic niche. Oncotarget. 2016;7(4):3993–4008. https://doi.org/10.18632/oncotarget.6540.
Muntion S, Ramos TL, Diez-Campelo M, Roson B, Sanchez-Abarca LI, Misiewicz-Krzeminska I, et al. Microvesicles from mesenchymal stromal cells are involved in HPC-microenvironment crosstalk in myelodysplastic patients. PLoS One. 2016;11(2):e0146722. https://doi.org/10.1371/journal.pone.0146722.
Huang Z, Zhu D, Wu L, He M, Zhou X, Zhang L, et al. Six serum-based miRNAs as potential diagnostic biomarkers for gastric cancer. Cancer Epidemiol Biomarkers Prev. 2017;26(2):188–96. https://doi.org/10.1158/1055-9965.EPI-16-0607.
Fong MY, Zhou W, Liu L, Alontaga AY, Chandra M, Ashby J, et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol. 2015;17(2):183–94. https://doi.org/10.1038/ncb3094.
Hannafon BN, Trigoso YD, Calloway CL, Zhao YD, Lum DH, Welm AL, et al. Plasma exosome microRNAs are indicative of breast cancer. Breast Cancer Res. 2016;18(1):90. https://doi.org/10.1186/s13058-016-0753-x.
Madhavan B, Yue S, Galli U, Rana S, Gross W, Muller M, et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer. 2015;136(11):2616–27. https://doi.org/10.1002/ijc.29324.
Liu W, Hu J, Zhou K, Chen F, Wang Z, Liao B, et al. Serum exosomal miR-125b is a novel prognostic marker for hepatocellular carcinoma. Onco Targets Ther. 2017;10:3843–51. https://doi.org/10.2147/OTT.S140062.
Rodriguez M, Silva J, Lopez-Alfonso A, Lopez-Muniz MB, Pena C, Dominguez G, et al. Different exosome cargo from plasma/bronchoalveolar lavage in non-small-cell lung cancer. Genes Chromosomes Cancer. 2014;53(9):713–24. https://doi.org/10.1002/gcc.22181.
Li Z, Ma YY, Wang J, Zeng XF, Li R, Kang W, et al. Exosomal microRNA-141 is upregulated in the serum of prostate cancer patients. Onco Targets Ther. 2016;9:139–48. https://doi.org/10.2147/ott.s95565.
Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110(1):13–21. https://doi.org/10.1016/j.ygyno.2008.04.033.
Fernandez-Mercado M, Manterola L, Lawrie CH. MicroRNAs in lymphoma: regulatory role and biomarker potential. Curr Genomics. 2015;16(5):349–58. https://doi.org/10.2174/1389202916666150707160147.
Que R, Ding G, Chen J, Cao L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J Surg Oncol. 2013;11:219. https://doi.org/10.1186/1477-7819-11-219.
Umezu T, Ohyashiki K, Kuroda M, Ohyashiki JH. Leukemia cell to endothelial cell communication via exosomal miRNAs. Oncogene. 2013;32(22):2747–55. https://doi.org/10.1038/onc.2012.295.
Matsumura T, Sugimachi K, Iinuma H, Takahashi Y, Kurashige J, Sawada G, et al. Exosomal microRNA in serum is a novel biomarker of recurrence in human colorectal cancer. Br J Cancer. 2015;113(2):275–81. https://doi.org/10.1038/bjc.2015.201.
Zhu HT, Liu RB, Liang YY, Hasan AME, Wang HY, Shao Q, et al. Serum microRNA profiles as diagnostic biomarkers for HBV-positive hepatocellular carcinoma. Liver Int. 2017;37(6):888–96. https://doi.org/10.1111/liv.13356.
Zhu M, Huang Z, Zhu D, Zhou X, Shan X, Qi LW, et al. A panel of microRNA signature in serum for colorectal cancer diagnosis. Oncotarget. 2017;8(10):17081–91. https://doi.org/10.18632/oncotarget.15059.
Le MT, Hamar P, Guo C, Basar E, Perdigao-Henriques R, Balaj L, et al. miR-200-containing extracellular vesicles promote breast cancer cell metastasis. J Clin Invest. 2014;124(12):5109–28. https://doi.org/10.1172/JCI75695.
Cappellesso R, Tinazzi A, Giurici T, Simonato F, Guzzardo V, Ventura L, et al. Programmed cell death 4 and microRNA 21 inverse expression is maintained in cells and exosomes from ovarian serous carcinoma effusions. Cancer Cytopathol. 2014;122(9):685–93. https://doi.org/10.1002/cncy.21442.
Liu W, Chen S, Liu B. Diagnostic and prognostic values of serum exosomal microRNA-21 in children with hepatoblastoma: a Chinese population-based study. Pediatr Surg Int. 2016;32(11):1059–65. https://doi.org/10.1007/s00383-016-3960-8.
Zhou X, Wen W, Shan X, Zhu W, Xu J, Guo R, et al. A six-microRNA panel in plasma was identified as a potential biomarker for lung adenocarcinoma diagnosis. Oncotarget. 2017;8(4):6513–25. https://doi.org/10.18632/oncotarget.14311.
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007;133(2):647–58. https://doi.org/10.1053/j.gastro.2007.05.022.
Wang H, Hou L, Li A, Duan Y, Gao H, Song X. Expression of serum exosomal microRNA-21 in human hepatocellular carcinoma. Biomed Res Int. 2014;2014:864894. https://doi.org/10.1155/2014/864894.
Li L, Li C, Wang S, Wang Z, Jiang J, Wang W, et al. Exosomes derived from hypoxic oral squamous cell carcinoma cells deliver miR-21 to normoxic cells to elicit a prometastatic phenotype. Cancer Res. 2016;76(7):1770–80. https://doi.org/10.1158/0008-5472.CAN-15-1625.
Masoudi MS, Mehrabian E, Mirzaei H. MiR-21: a key player in glioblastoma pathogenesis. J Cell Biochem. 2018;119(2):1285–90. https://doi.org/10.1002/jcb.26300.
Liu Y, Luo F, Wang B, Li H, Xu Y, Liu X, et al. STAT3-regulated exosomal miR-21 promotes angiogenesis and is involved in neoplastic processes of transformed human bronchial epithelial cells. Cancer Lett. 2016;370(1):125–35. https://doi.org/10.1016/j.canlet.2015.10.011.
Cui H, Seubert B, Stahl E, Dietz H, Reuning U, Moreno-Leon L, et al. Tissue inhibitor of metalloproteinases-1 induces a pro-tumourigenic increase of miR-210 in lung adenocarcinoma cells and their exosomes. Oncogene. 2015;34(28):3640–50. https://doi.org/10.1038/onc.2014.300.
Huang Y, Zhu J, Li W, Zhang Z, Xiong P, Wang H, et al. Serum microRNA panel excavated by machine learning as a potential biomarker for the detection of gastric cancer. Oncol Rep. 2018;39(3):1338–46. https://doi.org/10.3892/or.2017.6163.
Sohn W, Kim J, Kang SH, Yang SR, Cho JY, Cho HC, et al. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Exp Mol Med. 2015;47:e184. https://doi.org/10.1038/emm.2015.68.
Ye SB, Zhang H, Cai TT, Liu YN, Ni JJ, He J, et al. Exosomal miR-24-3p impedes T-cell function by targeting FGF11 and serves as a potential prognostic biomarker for nasopharyngeal carcinoma. J Pathol. 2016;240(3):329–40. https://doi.org/10.1002/path.4781.
Jiao C, Jiao X, Zhu A, Ge J, Xu X. Exosomal miR-34s panel as potential novel diagnostic and prognostic biomarker in patients with hepatoblastoma. J Pediatr Surg. 2017;52(4):618–24. https://doi.org/10.1016/j.jpedsurg.2016.09.070.
Corcoran C, Rani S, O’Driscoll L. miR-34a is an intracellular and exosomal predictive biomarker for response to docetaxel with clinical relevance to prostate cancer progression. Prostate. 2014;74(13):1320–34. https://doi.org/10.1002/pros.22848.
Guduric-Fuchs J, O’Connor A, Camp B, O’Neill CL, Medina RJ, Simpson DA. Selective extracellular vesicle-mediated export of an overlapping set of microRNAs from multiple cell types. BMC Genomics. 2012;13:357. https://doi.org/10.1186/1471-2164-13-357.
Pigati L, Yaddanapudi SC, Iyengar R, Kim DJ, Hearn SA, Danforth D, et al. Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS One. 2010;5(10):e13515. https://doi.org/10.1371/journal.pone.0013515.
Liu C, Eng C, Shen J, Lu Y, Takata Y, Mehdizadeh A, et al. Serum exosomal miR-4772-3p is a predictor of tumor recurrence in stage II and III colon cancer. Oncotarget. 2016;7(46):76250–60. https://doi.org/10.18632/oncotarget.12841.
Sugimachi K, Matsumura T, Hirata H, Uchi R, Ueda M, Ueo H, et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br J Cancer. 2015;112(3):532–8. https://doi.org/10.1038/bjc.2014.621.
Wu H, Zhou J, Mei S, Wu D, Mu Z, Chen B, et al. Circulating exosomal microRNA-96 promotes cell proliferation, migration and drug resistance by targeting LMO7. J Cell Mol Med. 2017;21(6):1228–36. https://doi.org/10.1111/jcmm.13056.
Akers JC, Hua W, Li H, Ramakrishnan V, Yang Z, Quan K, et al. A cerebrospinal fluid microRNA signature as biomarker for glioblastoma. Oncotarget. 2017;8(40):68769–79. https://doi.org/10.18632/oncotarget.18332.
Conigliaro A, Costa V, Lo Dico A, Saieva L, Buccheri S, Dieli F, et al. CD90+ liver cancer cells modulate endothelial cell phenotype through the release of exosomes containing H19 lncRNA. Mol Cancer. 2015;14:155. https://doi.org/10.1186/s12943-015-0426-x.
Isin M, Uysaler E, Ozgur E, Koseoglu H, Sanli O, Yucel OB, et al. Exosomal lncRNA-p21 levels may help to distinguish prostate cancer from benign disease. Front Genet. 2015;6:168. https://doi.org/10.3389/fgene.2015.00168.
Liu T, Zhang X, Gao S, Jing F, Yang Y, Du L, et al. Exosomal long noncoding RNA CRNDE-h as a novel serum-based biomarker for diagnosis and prognosis of colorectal cancer. Oncotarget. 2016;7(51):85551–63. https://doi.org/10.18632/oncotarget.13465.
Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 2009;23(13):1494–504. https://doi.org/10.1101/gad.1800909.
Diepenbruck M, Christofori G. Epithelial-mesenchymal transition (EMT) and metastasis: yes, no, maybe? Curr Opin Cell Biol. 2016;43:7–13. https://doi.org/10.1016/j.ceb.2016.06.002.
Hood JL. Melanoma exosome induction of endothelial cell GM-CSF in pre-metastatic lymph nodes may result in different M1 and M2 macrophage mediated angiogenic processes. Med Hypotheses. 2016;94:118–22. https://doi.org/10.1016/j.mehy.2016.07.009.
Franzen CA, Blackwell RH, Todorovic V, Greco KA, Foreman KE, Flanigan RC, et al. Urothelial cells undergo epithelial-to-mesenchymal transition after exposure to muscle invasive bladder cancer exosomes. Oncogenesis. 2015;4:e163. https://doi.org/10.1038/oncsis.2015.21.
Vella LJ. The emerging role of exosomes in epithelial-mesenchymal-transition in cancer. Front Oncol. 2014;4:361. https://doi.org/10.3389/fonc.2014.00361.
Liu Y, Gu Y, Han Y, Zhang Q, Jiang Z, Zhang X, et al. Tumor exosomal RNAs promote lung pre-metastatic niche formation by activating alveolar epithelial TLR3 to recruit neutrophils. Cancer Cell. 2016;30(2):243–56. https://doi.org/10.1016/j.ccell.2016.06.021.
Rahman MA, Barger JF, Lovat F, Gao M, Otterson GA, Nana-Sinkam P. Lung cancer exosomes as drivers of epithelial mesenchymal transition. Oncotarget. 2016;7(34):54852–66. https://doi.org/10.18632/oncotarget.10243.
Cao M, Seike M, Soeno C, Mizutani H, Kitamura K, Minegishi Y, et al. MiR-23a regulates TGF-beta-induced epithelial-mesenchymal transition by targeting E-cadherin in lung cancer cells. Int J Oncol. 2012;41(3):869–75. https://doi.org/10.3892/ijo.2012.1535.
Kim J, Kim TY, Lee MS, Mun JY, Ihm C, Kim SA. Exosome cargo reflects TGF-beta1-mediated epithelial-to-mesenchymal transition (EMT) status in A549 human lung adenocarcinoma cells. Biochem Biophys Res Commun. 2016;478(2):643–8. https://doi.org/10.1016/j.bbrc.2016.07.124.
Ohshima K, Inoue K, Fujiwara A, Hatakeyama K, Kanto K, Watanabe Y, et al. Let-7 microRNA family is selectively secreted into the extracellular environment via exosomes in a metastatic gastric cancer cell line. PLoS One. 2010;5(10):e13247. https://doi.org/10.1371/journal.pone.0013247.
Tanaka S, Hosokawa M, Yonezawa T, Hayashi W, Ueda K, Iwakawa S. Induction of epithelial-mesenchymal transition and down-regulation of miR-200c and miR-141 in oxaliplatin-resistant colorectal cancer cells. Biol Pharm Bull. 2015;38(3):435–40. https://doi.org/10.1248/bpb.b14-00695.
Xiao D, Barry S, Kmetz D, Egger M, Pan J, Rai SN, et al. Melanoma cell-derived exosomes promote epithelial-mesenchymal transition in primary melanocytes through paracrine/autocrine signaling in the tumor microenvironment. Cancer Lett. 2016;376(2):318–27. https://doi.org/10.1016/j.canlet.2016.03.050.
Liao J, Liu R, Shi YJ, Yin LH, Pu YP. Exosome-shuttling microRNA-21 promotes cell migration and invasion-targeting PDCD4 in esophageal cancer. Int J Oncol. 2016;48(6):2567–79. https://doi.org/10.3892/ijo.2016.3453.
Lu X, Kang Y. Organotropism of breast cancer metastasis. J Mammary Gland Biol Neoplasia. 2007;12(2–3):153–62. https://doi.org/10.1007/s10911-007-9047-3.
Fidler IJ. Selection of successive tumour lines for metastasis. Nat New Biol. 1973;242(118):148–9.
Paget S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 1989;8(2):98–101.
Hood JL, San RS, Wickline SA. Exosomes released by melanoma cells prepare sentinel lymph nodes for tumor metastasis. Cancer Res. 2011;71(11):3792–801. https://doi.org/10.1158/0008-5472.CAN-10-4455.
Javle M, Li Y, Tan D, Dong X, Chang P, Kar S, et al. Biomarkers of TGF-beta signaling pathway and prognosis of pancreatic cancer. PLoS One. 2014;9(1):e85942. https://doi.org/10.1371/journal.pone.0085942.
Ellermeier J, Wei J, Duewell P, Hoves S, Stieg MR, Adunka T, et al. Therapeutic efficacy of bifunctional siRNA combining TGF-beta1 silencing with RIG-I activation in pancreatic cancer. Cancer Res. 2013;73(6):1709–20. https://doi.org/10.1158/0008-5472.CAN-11-3850.
Lugini L, Valtieri M, Federici C, Cecchetti S, Meschini S, Condello M, et al. Exosomes from human colorectal cancer induce a tumor-like behavior in colonic mesenchymal stromal cells. Oncotarget. 2016;7(31):50086–98. https://doi.org/10.18632/oncotarget.10574.
Peinado H, Aleckovic M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G, et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med. 2012;18(6):883–91. https://doi.org/10.1038/nm.2753.
Abusamra AJ, Zhong Z, Zheng X, Li M, Ichim TE, Chin JL, et al. Tumor exosomes expressing Fas ligand mediate CD8+ T-cell apoptosis. Blood Cells Mol Dis. 2005;35(2):169–73. https://doi.org/10.1016/j.bcmd.2005.07.001.
Wieckowski EU, Visus C, Szajnik M, Szczepanski MJ, Storkus WJ, Whiteside TL. Tumor-derived microvesicles promote regulatory T cell expansion and induce apoptosis in tumor-reactive activated CD8+ T lymphocytes. J Immunol. 2009;183(6):3720–30. https://doi.org/10.4049/jimmunol.0900970.
Szajnik M, Czystowska M, Szczepanski MJ, Mandapathil M, Whiteside TL. Tumor-derived microvesicles induce, expand and up-regulate biological activities of human regulatory T cells (Treg). PLoS One. 2010;5(7):e11469. https://doi.org/10.1371/journal.pone.0011469.
Whiteside TL. Immune modulation of T-cell and NK (natural killer) cell activities by TEXs (tumour-derived exosomes). Biochem Soc Trans. 2013;41(1):245–51. https://doi.org/10.1042/BST20120265.
Chalmin F, Ladoire S, Mignot G, Vincent J, Bruchard M, Remy-Martin JP, et al. Membrane-associated Hsp72 from tumor-derived exosomes mediates STAT3-dependent immunosuppressive function of mouse and human myeloid-derived suppressor cells. J Clin Invest. 2010;120(2):457–71. https://doi.org/10.1172/JCI40483.
Nagaraj S, Nelson A, Youn JI, Cheng P, Quiceno D, Gabrilovich DI. Antigen-specific CD4(+) T cells regulate function of myeloid-derived suppressor cells in cancer via retrograde MHC class II signaling. Cancer Res. 2012;72(4):928–38. https://doi.org/10.1158/0008-5472.CAN-11-2863.
Lv LH, Wan YL, Lin Y, Zhang W, Yang M, Li GL, et al. Anticancer drugs cause release of exosomes with heat shock proteins from human hepatocellular carcinoma cells that elicit effective natural killer cell antitumor responses in vitro. J Biol Chem. 2012;287(19):15874–85. https://doi.org/10.1074/jbc.M112.340588.
Goler-Baron V, Assaraf YG. Structure and function of ABCG2-rich extracellular vesicles mediating multidrug resistance. PLoS One. 2011;6(1):e16007. https://doi.org/10.1371/journal.pone.0016007.
Bebawy M, Combes V, Lee E, Jaiswal R, Gong J, Bonhoure A, et al. Membrane microparticles mediate transfer of P-glycoprotein to drug sensitive cancer cells. Leukemia. 2009;23(9):1643–9. https://doi.org/10.1038/leu.2009.76.
Corcoran C, Rani S, O’Brien K, O’Neill A, Prencipe M, Sheikh R, et al. Docetaxel-resistance in prostate cancer: evaluating associated phenotypic changes and potential for resistance transfer via exosomes. PLoS One. 2012;7(12):e50999. https://doi.org/10.1371/journal.pone.0050999.
Safaei R, Larson BJ, Cheng TC, Gibson MA, Otani S, Naerdemann W, et al. Abnormal lysosomal trafficking and enhanced exosomal export of cisplatin in drug-resistant human ovarian carcinoma cells. Mol Cancer Ther. 2005;4(10):1595–604. https://doi.org/10.1158/1535-7163.MCT-05-0102.
Lehmann BD, Paine MS, Brooks AM, McCubrey JA, Renegar RH, Wang R, et al. Senescence-associated exosome release from human prostate cancer cells. Cancer Res. 2008;68(19):7864–71. https://doi.org/10.1158/0008-5472.CAN-07-6538.
Khan S, Jutzy JM, Aspe JR, McGregor DW, Neidigh JW, Wall NR. Survivin is released from cancer cells via exosomes. Apoptosis. 2011;16(1):1–12. https://doi.org/10.1007/s10495-010-0534-4.
Jaiswal R, Luk F, Gong J, Mathys JM, Grau GE, Bebawy M. Microparticle conferred microRNA profiles–implications in the transfer and dominance of cancer traits. Mol Cancer. 2012;11:37. https://doi.org/10.1186/1476-4598-11-37.
Hergenreider E, Heydt S, Treguer K, Boettger T, Horrevoets AJ, Zeiher AM, et al. Atheroprotective communication between endothelial cells and smooth muscle cells through miRNAs. Nat Cell Biol. 2012;14(3):249–56. https://doi.org/10.1038/ncb2441.
Clynes RA, Towers TL, Presta LG, Ravetch JV. Inhibitory Fc receptors modulate in vivo cytotoxicity against tumor targets. Nat Med. 2000;6(4):443–6. https://doi.org/10.1038/74704.
Aung T, Chapuy B, Vogel D, Wenzel D, Oppermann M, Lahmann M, et al. Exosomal evasion of humoral immunotherapy in aggressive B-cell lymphoma modulated by ATP-binding cassette transporter A3. Proc Natl Acad Sci U S A. 2011;108(37):15336–41. https://doi.org/10.1073/pnas.1102855108.
Battke C, Ruiss R, Welsch U, Wimberger P, Lang S, Jochum S, et al. Tumour exosomes inhibit binding of tumour-reactive antibodies to tumour cells and reduce ADCC. Cancer Immunol Immunother. 2011;60(5):639–48. https://doi.org/10.1007/s00262-011-0979-5.
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem. 2010;285(23):17442–52. https://doi.org/10.1074/jbc.M110.107821.
Cho DY, Lin SZ, Yang WK, Hsu DM, Lee HC, Lee WY, et al. Recent advances of dendritic cells (DCs)-based immunotherapy for malignant gliomas. Cell Transplant. 2009;18(9):977–83. https://doi.org/10.3727/096368909X12483162196962.
Cho JA, Yeo DJ, Son HY, Kim HW, Jung DS, Ko JK, et al. Exosomes: a new delivery system for tumor antigens in cancer immunotherapy. Int J Cancer. 2005;114(4):613–22. https://doi.org/10.1002/ijc.20757.
Elsner L, Muppala V, Gehrmann M, Lozano J, Malzahn D, Bickeboller H, et al. The heat shock protein HSP70 promotes mouse NK cell activity against tumors that express inducible NKG2D ligands. J Immunol. 2007;179(8):5523–33.
Escudier B, Dorval T, Chaput N, Andre F, Caby MP, Novault S, et al. Vaccination of metastatic melanoma patients with autologous dendritic cell (DC) derived-exosomes: results of thefirst phase I clinical trial. J Transl Med. 2005;3(1):10. https://doi.org/10.1186/1479-5876-3-10.
Morse MA, Garst J, Osada T, Khan S, Hobeika A, Clay TM, et al. A phase I study of dexosome immunotherapy in patients with advanced non-small cell lung cancer. J Transl Med. 2005;3(1):9. https://doi.org/10.1186/1479-5876-3-9.
de la Fuente A, Alonso-Alconada L, Costa C, Cueva J, Garcia-Caballero T, Lopez-Lopez R, et al. M-Trap: exosome-based capture of tumor cells as a new technology in peritoneal metastasis. J Natl Cancer Inst. 2015;107(9):djv184. https://doi.org/10.1093/jnci/djv184.
Kim MS, Haney MJ, Zhao Y, Mahajan V, Deygen I, Klyachko NL, et al. Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine. 2016;12(3):655–64. https://doi.org/10.1016/j.nano.2015.10.012.
Kooijmans SA, Vader P, van Dommelen SM, van Solinge WW, Schiffelers RM. Exosome mimetics: a novel class of drug delivery systems. Int J Nanomedicine. 2012;7:1525–41. https://doi.org/10.2147/IJN.S29661.
Zhang W, Li C, Shen C, Liu Y, Zhao X, Liu Y, et al. Prodrug-based nano-drug delivery system for co-encapsulate paclitaxel and carboplatin for lung cancer treatment. Drug Deliv. 2016;23(7):2575–80. https://doi.org/10.3109/10717544.2015.1035466.
Airoldi M, Amadori D, Barni S, Cinieri S, De Placido S, Di Leo A, et al. Clinical activity and cardiac tolerability of non-pegylated liposomal doxorubicin in breast cancer: a synthetic review. Tumori. 2011;97(6):690–2. https://doi.org/10.1700/1018.11082.
Swenson CE, Perkins WR, Roberts P, Janoff AS. Liposome technology and the development of Myocet™ (liposomal doxorubicin citrate). The Breast. 2001;10:1–7. https://doi.org/10.1016/S0960-9776(01)80001-1.
Doxorubicin (Myocet) 50 mg powder, dispersion and solvent for concentrate for dispersion for infusion. Teva Pharma B.V., Electronic Medicines Compendium. 2015. https://www.medicines.org.uk/emc/product/5378/smpc#companyDetails. Accessed 4 Jan 2019.
Trivedi M, Talekar M, Shah P, Ouyang Q, Amiji M. Modification of tumor cell exosome content by transfection with wt-p53 and microRNA-125b expressing plasmid DNA and its effect on macrophage polarization. Oncogenesis. 2016;5(8):e250. https://doi.org/10.1038/oncsis.2016.52.
Federici C, Petrucci F, Caimi S, Cesolini A, Logozzi M, Borghi M, D'Ilio S, Lugini L, Violante N, Azzarito T, Majorani C, Brambilla D, Fais S. Exosome release and low pH belong to a framework of resistance of human melanoma cells to cisplatin. PLoS One. 2014;9(2):e88193. https://doi.org/10.1371/journal.pone.0088193.
Iessi E, Logozzi M, Lugini L, Azzarito T, Federici C, Spugnini EP, et al. Acridine Orange/exosomes increase the delivery and the effectiveness of acridine orange in human melanoma cells: a new prototype for theranostics of tumors. J Enzyme Inhib Med Chem. 2017;32(1):648–57. https://doi.org/10.1080/14756366.2017.1292263.
Kusuzaki K, Hosogi S, Ashihara E, Matsubara T, Satonaka H, Nakamura T, et al. Translational research of photodynamic therapy with acridine orange which targets cancer acidity. Curr Pharm Des. 2012;18(10):1414–20.
Blakely CM, Watkins TBK, Wu W, Gini B, Chabon JJ, McCoach CE, McGranahan N, Wilson GA, Birkbak NJ, Olivas VR, Rotow J, Maynard A, Wang V, Gubens MA, Banks KC, Lanman RB, Caulin AF, St John J, Cordero AR, Giannikopoulos P, Simmons AD, Mack PC, Gandara DR, Husain H, Doebele RC, Riess JW, Diehn M, Swanton C, Bivona TG. Evolution and clinical impact of co-occurring genetic alterations in advanced-stage EGFR-mutant lung cancers. Nat. Genet. 2017;49(12):1693–1704.
Zocco D, Ferruzzi P, Cappello F, Kuo WP, Fais S. Extracellular vesicles as shuttles of tumor biomarkers and anti-tumor drugs. Front Oncol. 2014;4:267. https://doi.org/10.3389/fonc.2014.00267.
Lener T, Gimona M, Aigner L, Borger V, Buzas E, Camussi G, et al. Applying extracellular vesicles based therapeutics in clinical trials - an ISEV position paper. J Extracell Vesicles. 2015;4:30087. https://doi.org/10.3402/jev.v4.30087.
Author information
Authors and Affiliations
Contributions
WWW: conception and planning, manuscript writing, and final approval of manuscript. TG: conception and planning, manuscript writing, and final approval of manuscript. HTT: conception and planning, manuscript editing, and final approval of manuscript.
Corresponding author
Ethics declarations
Funding
No funding was received for the preparation of this review.
Conflict of Interest
Wendy W. Weston, Timothy Ganey, and H. Thomas Temple have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Weston, W.W., Ganey, T. & Temple, H.T. The Relationship between Exosomes and Cancer: Implications for Diagnostics and Therapeutics. BioDrugs 33, 137–158 (2019). https://doi.org/10.1007/s40259-019-00338-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40259-019-00338-5