Abstract
Purpose of Review
Pharmacogenetics is an important component of precision medicine. Even within the genomic era, several challenges lie ahead in the road towards clinical implementation of pharmacogenetics in the clinic. This review will summarize the current state of knowledge regarding pharmacogenetics of cardiovascular drugs, focusing on those with the most evidence supporting clinical implementation—clopidogrel, warfarin, and simvastatin.
Recent Findings
There is limited translation of pharmacogenetics into clinical practice primarily due to the absence of outcomes data from prospective, randomized, genotype-directed clinical trials. There are several ongoing randomized controlled trials that will provide some answers as to the clinical utility of genotype-directed strategies. Several academic medical centers have pushed towards clinical implementation, where the clinical validity data are strong. Their experiences will inform operational requirements of a clinical pharmacogenetics testing, including the timing of testing, incorporation of test results into the electronic health record, reimbursement, and ethical issues.
Summary
Pharmacogenetics of clopidogrel, warfarin, and simvastatin are three examples, where pharmacogenetics testing may provide added clinical value. Continued accumulation of evidence surrounding clinical utility of pharmacogenetics markers is imperative as this will inform reimbursement policy and drive adoption of pharamcogenetics into routine care.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
An important part of precision medicine includes pharmacogenetics (PGx) or the application of genetic information to individualize medication treatments with the goal of maximizing effectiveness and minimizing adverse drug reactions. The concept of precision medicine is not new, as healthcare providers have routinely been tailoring treatment regimens for patients based on demographic, lifestyle, and clinical factors. However, recent advances in genomics, informatics, and mobile technology can now accelerate the practice of precision medicine to enhance human health [1••]. Capitalizing on the convergence of these technologies, President Obama announced the Precision Medicine Initiative in January 2015 during his State of the Union address [1••]. This concerted effort aims to enhance our understanding of health and disease and improves disease prevention, risk assessment, and treatment taking into account individual variability in genes, environment, and lifestyle. The Precision Medicine Initiative’s million patient cohort is poised to provide a strong foundation to advance PGx. This is by far, the largest effort supported by the National Institute of Health. Previous large-scale projects include the Pharmacogenomics Research Network (PGRN) [2], focused on the discovery of genetic variation that influence drug response, and the Pharmacogenomics Knowledgebase (PharmGKB) [2], an expertly curated database cataloging the impact of genetic variation on drug response. To aid clinicians with the translation of genetic test results into drug treatment decisions, the Clinical Pharmacogenetic Implementation Consortium (CPIC) was established [3]. The group has published guidelines for over 30 drug-gene pairs on how genetic test results should be used to optimize drug therapy, including several cardiovascular drugs [4•, 5, 6•].
This review will summarize the current state of knowledge regarding PGx of cardiovascular drugs, focusing on those with the most evidence supporting clinical implementation—clopidogrel, warfarin, and simvastatin. We will discuss barriers to future PGx marker discovery and issues to consider in the road to clinical implementation [7].
Clopidogrel and CYP2C19
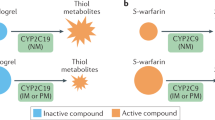
Clopidogrel is a thienopyridine prodrug, which upon biotransformation to its active metabolite irreversibly inhibits the P2Y12 adenosine diphosphate (ADP) receptor on platelets, preventing their aggregation [4•]. Clopidogrel has been shown to prevent thromboembolism-related cardiovascular events in patients experiencing acute coronary syndromes (ACS), especially in the setting of percutaneous coronary interventions (PCI) [8]. Clopidogrel’s effect on inhibition of platelet aggregation displays wide inter-individual variability. Approximately 25 % of the population is non-responsive to the drug, displaying residual on-treatment platelet reactivity [9]. Genetic polymorphisms in the hepatic cytochrome P450 2C19 (CYP2C19) enzyme, one of the principal enzymes involved with the biotransformation of clopidogrel, are associated with lower active clopidogrel metabolite levels and residual platelet aggregation [10–13]. The most common loss-of-function (LOF) allele in CYP2C19 gene is *2 (c.681G > A; rs4244285), with frequencies of ~15 % in Caucasians and Africans and 29–35 % in Asians [4, 14, 15]. Data from a large meta-analysis involving patients undergoing PCI showed that CYP2C19*2 carriers are at increased risk of myocardial infarction, death, and stent thrombosis compared to non-carriers [16••]. Another meta-analysis, including patients, treated with clopidogrel with stable coronary disease or with ACS managed without PCI found no association with CYP2C19*2 genotype and cardiovascular disease (CVD) events (RR 0.97; 95 % CI 0.86–1.09). However, this study confirmed the association with CYP2C19 LOF alleles and stent thrombosis (RR 1.75, 95 % CI 1.5–2.03) [17]. The United States Food and Drug Administration (FDA) has instituted a “black box” warning for clopidogrel, recommending consideration of alternate antiplatelet agents among carriers of two loss-of-function alleles (poor metabolizers). The American College of Cardiology Foundation task force suggests that PGx testing for CYP2C19 may be considered in patients believed to be at moderate or high risk for poor outcomes (e.g., high-risk PCI procedures and complex disease) awaiting further evidence from outcomes trials [18•]. Peer reviewed guidelines published by the CPIC recommend the use of alternative therapies, such as prasugrel or ticagrelor, in CYP2C19 LOF allele carriers, specifically for patients with ACS undergoing PCI [19]. The newer P2Y12 inhibitors, prasugrel and ticagrelor, do provide more potent antiplatelet activity and are not influenced by the presence of CYP2C19 polymorphisms; however, they are associated with increased bleeding risk [20, 21], and since no generic is available, they are substantially more expensive than clopidogrel.
There is limited translation of CYP2C19 genotyping in the clinical practice, due primarily to the absence of outcomes data from prospective, randomized, genotype-directed clinical trials [22]. Therefore, PCI guidelines published by the American Heart Association (AHA), American College of Clinical Cardiology (ACC), and Society for Cardiovascular Angiography (SCAI) do not currently recommend PGx testing for CYP2C19 [18•, 23]. There are several ongoing randomized controlled trials (RCTs) that will provide some answers as to the clinical utility of a genotype-directed strategy, including potential economic benefits (Table 1). Results of these studies will inform widespread adoption of personalized antiplatelet therapies.
The largest of the studies, the TAILOR-PCI trial [24], will determine whether prospectively identifying and prescribing ticagrelor for patients with CYP2C19 LOF alleles compared with usual clinical care will prevent major cardiac events at 1 year. Approximately 5200 subjects with stable coronary artery disease (CAD) or ACS undergoing PCI will be enrolled and PGx results provided using a rapid turnaround genotyping platform. Another trial currently enrolling in Europe, POPular Genetics, will compare whether a genotype-directed strategy is superior to prescribing ticagrelor or prasugrel for all subjects without genotyping. In the genotyping group, CYP2C19 LOF carriers will receive ticagrelor and individuals with functional alleles will receive clopidogrel. The trial will include 2700 patients with ST-segment MI who undergo PCI. The primary endpoint is a composite of death, recurrent MI, stent thrombosis, stroke, and major bleeding complications. The investigators will also evaluate the cost-effectiveness of genotype-guided strategy compared with using a non-tailored approach [25]. Both TAILOR-PCI and POPular Genetics will evaluate the efficacy of a genotype-guided approach in preventing major cardiac events. A third RCT will evaluate comparative effectiveness of providing PGx results to physicians to guide decision making. The ADAPT study, which is currently enrolling at the University of Pennsylvania, will randomize patients undergoing PCI to CYP2C19 genotyping or usual care. In the genotype group, physicians are provided PGx test results and antiplatelet therapy recommendations, but ultimately decide the antiplatelet therapy incorporating both genetic and clinical factors. The study will prospectively ascertain major cardiac events, bleeding complications, healthcare costs, and implementation metrics associated with incorporating PGx testing as part of clinical care. The result of this trial will inform healthcare stakeholders of how implementation of a genotype-guided approach will perform in a “real-world” setting.
Two studies evaluating the economics of a genotype-directed strategy, prescribing clopidogrel for patients with functional CYP2C19 alleles who are at low risk for ischemic events and reserving prasugrel and ticagrelor for poor metabolizers, have shown this approach to be cost-effective [26, 27]. However, these studies were conducted by modeling data from previously published clinical trials, and not performed in a prospective setting. Another concern that limits the usefulness of PGx tests in clinical practice is the availability of actionable test results in a time frame conducive to the provision of clinical care. Knowledge of a CYP2C19 genotype result is required emergently in the setting of ACS, ideally within 24 h. Currently, there is one FDA approved device available for clinical use that provides results within 1 h—the Spartan Rx CYP2C19 (Spartan Bioscience, Ottawa, Canada). The test is also reimbursable by Medicare under CPT code 81225. Availability of test results and guidance at point-of-care will enhance the uptake of implementation of these tests in the ACS setting once the clinical utility data show a clinical benefit with prospective genotyping.
There is a need to further understand the mechanisms by which genetic variations impact drug-related outcomes by race. In the TRIUMP genetic substudy, the LOF CYP2C19*2 allele was associated with an increased 1-year mortality in European-ancestry patients treated with clopidogrel following acute MI (HR 1.70, CI 1.01–2.89, p = 0.046), with a trend towards increased rate of recurrent MI. However, among African–American patients, the gain-of-function CYP2C19*17 allele was significantly associated with increased mortality (HR for *17/*17 versus *1/*1: 8.97, CI 3.34–24.10, p < 0.0001), with an increased rate of bleeding events [28••]. This study and the studies on warfarin PGx described below highlight the need to study racially diverse populations, as variants will have different frequencies among ethnic groups and, therefore, will exhibit an effect in one group but not the other. Characterizing PGx differences by race will enable us to develop interventions that will minimize difference in care and eliminate disparities in health outcomes [29].
Warfarin and CYP2C9/VKORC1
Warfarin, the most widely used oral anticoagulant for the treatment of thrombotic disorders, requires intensive monitoring and dosage adjustments based on target international normalized ratio (INR) to avoid drug-related complications [30]. Warfarin-related complications are the most common cause of emergency department visits and hospitalizations for adverse drug events [31]. Warfarin is metabolized by cytochrome P450 2C9 (CYP2C9) in the liver. Two single nucleotide polymorphisms (SNPs) in CYP2C9 denoted as CYP2C9*2 (exon 3; Arg144Cys) and CYP2C9*3 (exon 7; le359Leu) are associated with reduced warfarin metabolism and lower dose requirements [32]. The pharmacological target of warfarin is vitamin K epoxide reductase (VKORC1), the rate-limiting enzyme that controls the formation of vitamin K-dependent clotting factors [33]. A common noncoding variant in VKORC1 (−1639 G > A, rs9923231), which alters a transcription factor binding site and leads to lower protein expression, is associated with warfarin sensitivity and lower dosing requirements [34]. A genome-wide association study (GWAS) performed in 1000 patients taking warfarin confirmed that CYP2C9*2, CYP2C9*3, and VKORC1 are significantly associated with warfarin dose requirements [35•, 36•]. In European-ancestry individuals, CYP2C9 and VKORC1 polymorphisms account for 30–40 % of the variance in warfarin dose [37], but explain less of the variability in other ethnic groups [38]. The FDA updated the warfarin label, including recommendations, for dosing table based on CYP2C9 and VKORC1 genotypes [39]. The current CPIC guidelines also recommend genotype-guided dosing of warfarin when the genotype results are available [5].
Two RCTs were completed in 2013 to determine whether prospective genotyping helped maintain patients in the therapeutic range (INR range of 2–3). The question of whether genotyping will improve clinically relevant outcomes (e.g., hemorrhage) remains to be determined. In the EU-PACT study [40••, 45], 455 patients were randomized to either genotype-guided group, in which warfarin was adjusted based on PGx algorithm for the first 5 days, or a standard dosing group which utilized a 3-day loading dose regimen. After this initial period, subjects were managed according to usual clinical practice and followed for 12 weeks. Subjects in the genotype-guided group achieved significantly greater time within therapeutic range (67 vs. 60 %, p < 0.001), faster time to reach therapeutic range (21 days vs. 29 days, p < 0.001), and fewer instances of supra-therapeutic INRs (INR ≥ 4.0). The study concluded that a genotype-guided approach improved percent time in target range.
In the COAG trial [41••], 1015 subject were randomized to warfarin dosing based on an algorithm that included PGx and clinical factors or one based on clinical factors only. At 4 weeks, the time within the therapeutic range was 45.2 % for the genotype-guided group and 45.4 % in the clinically guided group (p = 0.91). However, African–American subjects assigned to the genotype-guided group did not perform well as well as the clinically guided group (time in range 35 % for genotype group vs. 44 % clinically guided group, p = 0.01). African–American patients comprised 27 % of the study population as opposed to only 1 % in the EU-PACT study. Since then, recent studies have identified additional variants that improve the prediction of warfarin dose requirements in African–Americans (i.e., CYP2C9 rs12777823) [42•, 43]. Conversely, variants that are highly predictive in Europeans (CYP2C9*2) are less important in African–Americans [43, 44]. Therefore, a genetic dosing algorithm based on variants predictive in the European population would not be expected to perform well in other ethnic groups.
In addition to the ethnic differences between the two the studies, there are several important distinctions to highlight. The EU-PACT control group utilized a fixed dose warfarin strategy as compared with a clinical-guided algorithm in COAG which incorporated variables, such as age, sex, ethnicity, and concomitant drug use when providing dose recommendations. One would expect that a clinically guided algorithm would perform better than a fixed dose regimen; however, the time within therapeutic range at the 4-week time point in EU-PACT control subjects was identical (45 %) to that reported in the COAG trial. Perhaps, significant differences may have been observed in the COAG trial if subjects were followed for 12 weeks. In addition, genotype results in EU-PACT were available to investigators within 2 h as contrast to the COAG study, where only 45 % of subjects had genetic results available before the first dose and 94 % before the second dose, resulting in a large proportion of subjects randomized to “genotype” being managed with the clinical dosing algorithm for the first dose. As the intervention period for COAG was the first 3–5 days, this delay may have attenuated the benefits (if any) of PGx. As the clinical utility of warfarin PGx remains unanswered by these studies, implementation efforts remain hindered, as insurance reimbursement is difficult to attain without favorable clinical outcomes data. Perhaps, ongoing studies, such as the Genetics Informatics Trial (GIFT) of Warfarin, to prevent DVT will shed light on the clinical utility of genotype-guided warfarin therapy (Table 1).
Simvastatin and SLCO1B1
3-hydroxy-3-methylglutaryl-coenzyme (HMG-CoA) reductase inhibitors, also known as statins, are prescribed to approximately 25 million people worldwide for primary and secondary prevention of cardiovascular events [45, 46]. The 2014 AHA/ACC guidelines recommended a high-intensity statin in patients with clinical evident CVD, primary LDL-C ≥ 190 mg/dL, and patients with type 1 or type 2 diabetes with an LDL-C ≥ 70 mg/dL [47]. High-intensity statin therapy, which aims to reduce LDL-C by ≥50 %, poses a risk of statin-associated adverse effects. In addition, the guidelines recommend statin therapy for the primary prevention of CVD in patients with an algorithm predicted 10-year risk of atherosclerotic CVD of at least 7.5 %. The new criteria will vastly expand the number of patients eligible for the long-term statin treatment [48]. Although the risk of statin-related myopathy is low [45], the absolute number of people self-reporting statin-associated myalgia is common at 1–5 % [49]. In rare instances, statins can cause muscle pain or weakness (myopathy) which can sometime lead to muscle breakdown (rhabdomyolysis) [50]. In addition, the vast absolute numbers of patients maintained on statin therapy increase the overall number of patients reporting muscle-related complications.
A GWAS in patients with myopathy while taking high-dose simvastatin (80 mg) showed an association with an intronic SNP (rs4363657) in the SLCO1B1 gene, which encodes for the hepatic organic anion-transporting polypeptide OATP1B1, involved with hepatic uptake of statin medications [51]. This SNP was in complete linkage disequilibrium (r 2 = 0.97) with a nonsynonymous variant rs4149056 in exon 6 (c.521T > C, Val174Ala). Patients homozygous for the C variant had an odds ratio for myopathy of 17 (95 % CI 4.7–61.1) as compared to the TT homozygotes. This SNP has been shown to reduce the activity of the transporter in vitro [52] and has a strong impact on simvastatin pharmacokinetics [53]. Polymorphisms in SLCO1B1 are also implicated in milder forms of statin intolerance [54, 55]. However, there was no association of these variants with myopathy in subjects receiving other statin medications [54, 56], likely due to differences in pharmacokinetics properties of the various statins [57]. CPIC guidelines are available to help guide statin initiation in patients with SLCO1B1 variants. Genotyping SLCO1B1 may help titrate statin therapy in patients who do not achieve adequate LDL-C reduction with low-to-moderate doses [6, 49]. Nonetheless, clinical implementation of PGx testing prior to statin initiation has been challenging due to wide therapeutic index of statins, availability of low-cost statin alternatives to simvastatin, and difficulty in defining myalgias in practice. While milder forms of myopathy are self-limiting, they can lead to poor drug compliance and, therefore, result in failure to prevent cardiovascular events [58, 59]. It is unknown if PGx testing may aid in medication adherence for patients taking the long-term statin therapy. An ongoing clinical trial is evaluating whether prospective PGx of SLO1B1 will aid in medication adherence in patients requiring statin therapy to prevent complications of CAD (Table 1).
Barriers to Clinical Implementation
Clinical Utility
There is much debate on what constitutes sufficient evidence for implementing PGx findings into clinical practice [60–62]. In cardiology, RCTs are rightly the gold standard for establishing a new medical intervention as standard of care. Similarly, there is a call for clinical utility data for each drug-gene pair prior to clinical adoption [61]. Conversely, PGx test results have been equated to just one of many clinical factors, such as renal or liver function, all imperfect, but informative when making prescribing decisions [62]. Yet, RCTs are not performed in establishing dose modifications in patients with renal dysfunction. These recommendations are typically based on pharmacokinetic studies alone [63].
The Evaluation of Genomic Applications in Practice and Prevention (EGAPP) is an independent panel, supported by the Centers for Disease Control (CDC), convened to evaluate the evidence base for translation of genetic tests into clinical practice [64]. In their guidance document, they define clinical validity as the capacity of a PGx marker to predict drug response and clinical utility as the ability of the PGx markers to improve clinical outcomes. Considerable progress has been made in strengthening the clinical validity of PGx markers based on efforts by collaborative consortia, such as the PGRN. Based on clinical validity data, the FDA has updated labeling of over 150 medications to include PGx information [65]. A recent study by Wang et al. evaluated the evidence for PGx labeling for 119 drugs based on clinical validity and clinical utility data [66]. Among the eight drugs, they evaluated that are used in cardiovascular disease, they found that only one drug, clopidogrel, that had convincing clinical validity evidence, while none demonstrated convincing clinical utility evidence. The authors concluded that the inclusion of PGx information in the drug label is premature in the absence of clinical utility data.
Several centers have pushed forward toward clinical implementation of drug-gene pairs, where the clinical validity data are robust [67, 68•, 69]. The University of Florida implemented CYP2C19 testing in 2012 for patient undergoing PCI to guide antiplatelet therapy [70]. This was performed initially as a research study, but with the availability of test reimbursement, now constitutes standard of care. Vanderbilt University [71•] performs preemptive genotyping to guide management of antiplatelet agents, anticoagulants, and statins, further described in the next section. These efforts will inform the clinical utility of genotype-guided therapy as implemented in clinical care.
Operational Requirements
Implementation of a health-system wide PGx program entails the cooperation of key administrative and clinical stakeholders, including executive leaders, clinical leaders, laboratory medicine, pharmacy, informatics, and payers [72]. Prioritization of drug-gene pairs for implementation based on clinical validity/utility data requires input from clinical leaders, including physicians, pharmacists, nurses, and physician assistants. Laboratory medicine expertise is critical to conduct PGx testing under Clinical Laboratory Improvements Amendments (CLIA) requirements. The genotyping approach may be reactive, performed at the time the test result is needed, or preemptive, in advance of when the result is needed. The preemptive approach may be the most efficient and cost-effective approach for the future, as 100 s of PGx variants can be ascertained at one time and utilized over an individual’s lifetime [71•, 73]. Using preemptive PGx testing, Vanderbilt University has incorporated PGx results into the electronic medical record (EMR) for five established drug-gene pairs: clopidogrel-CYP2C19, warfarin-CYP2C9/VKORC1, simvastatin- SLCO1B1, thiopurines-TPMT, and tacrolimus-CYP3A5 [71•]. Clinically actionable variants trigger the necessary clinical decision support in the EMR to guide drug selection and dosing. The most important advantage to this approach is that the PGx result is available at the most critical time point for prescription decision making. This requires informatics support to integrate the test result in the EMR with user-friendly clinical decision support tools providing tailored drug and dosing recommendations.
Reimbursement for PGx test results from preemptive genotyping is of concern, as the administration of the testing is not linked to a concrete diagnosis. In this case, reactive genotyping may have the advantage, as it is performed at the time, the test is needed and linked to a specific diagnosis. However, there is still an ongoing need to interface with payers to negotiate PGx test reimbursement as well as additional costs that may arise from switching to more expensive drug therapy, as indicated by PGx test results.
A lack of provider knowledge regarding the interpretation and application of PGx has been documented [74–77]. Therefore, large-scale educational programs and tools would be imperative before PGx would be adopted into clinical practice. Learning resources and teaching tools are available through the NIH Genetics/Genomics Competency Center (http://g-2-c-2.org//) and the Global Genetics and Genomics Community (http://g-3-c.org/en) for clinical and healthcare educators.
Ethical, Legal, Social Implications of PGx testing
Genotyping from high-density genotyping arrays, and in the future whole-genome sequencing, poses additional ethical concerns, as a deluge of genetic data will be generated [78, 79]. Going forward the question will be not be whether patients will be tested, but what information will be provided, how it will be conveyed, and of course who will pay [80, 81]. There is an ongoing debate regarding the return of incidental findings to patients, which results are considered “actionable” [79, 82, 83]. The Genetic Information Nondiscrimination Act (GINA) offers certain protections against discrimination based on genetic information; however, there are still existing loopholes in the law, such as long-term care insurance, that need to be addressed [84]. Issues of privacy and discrimination are of concern to the public and providers alike and will impact implementation of PGx into practice [77, 85, 86].
Future Challenges
Despite the advances in genomics, drug therapy is still initiated and titrated based on a trial and error [87]. Pharmacogenetic markers can enable a better prediction of drug efficacy and toxicity. Early PGx studies of drug response utilized a targeted approach, focusing on known drug metabolism pathways, such as CYP2C9 and warfarin. Genetic variants influencing drug dose or response to warfarin, clopidogrel, and simvastatin were confirmed using GWAS [36, 51, 88]. However, the application of GWAS to the discovery of novel PGx makers has not been straight forward [60, 89]. Some of the issues in applying GWAS to identify variants linked to drug response include heterogeneity in drug response phenotypes, finding suitable replication cohorts that have received similar drug treatment and the large burden of multiple hypothesis testing as most drug trials enroll smaller sample sizes as compared to the hundreds of thousands of subjects analyzed in GWAS studies of disease risk [89, 90]. Identification of PGx markers for beta blockers, thiazide diuretics, and angiotensin-converting enzyme inhibitors for the treatment of hypertension and heart failure has been on the focus of intense research for the past two decades and the reader is referred to recent reviews in this area [91–93].
Conclusions
The prevalence of CVD and the large numbers of patients affected provide a large cohort at risk, where a scalable PGx model can be implemented and effectiveness and cost-effectiveness can be assessed. There are several examples, as discussed herein, where PGx testing may provide added clinical value. Continued accumulation of evidence surrounding clinical utility of PGx markers is imperative as this will inform reimbursement policy and drive adoption of PGx into routine care. Practical considerations, such as integration of PGx into electronic health records, provider education, and financial considerations, will need to be addressed. The Precision Medicine Initiative is poised to enable more genomic discoveries. Parallel efforts that enable the implementation of these discovers in clinical care are needed. Both are needed to optimize therapeutic outcomes.
References
Papers of particular interest, published recently, have been highlight as: • Of importance; •• Of major importance
•• Precision Medicine Initiative Cohort Program. Working Group Report 2015. https://www.nih.gov/precision-medicine-initiative-cohort-program. Accessed 8 Mar 2016. The Precision Medicine Working Group Report outlines the plan for building the one million Americans cohort, enabling precision medicine discoveries at an accerlated pace.
PGRN: Pharmacogenomics Research Network. http://www.nhlbi.nih.gov/research/resources/genetics-genomics/pgrn. Accessed 8 Mar 2016.
CPIC: Clinical pharmacogenetics implementation consortium. https://cpicpgx.org/. Accessed 8 Mar 2016.
• Scott SA, Sangkuhl K, Stein CM, et al. Clinical Pharmacogenetics Implementation Consortium Guidelines for CYP2C19 Genotype and Clopidogrel Therapy: 2013 Update. Clin Pharmacol Ther 2013;94:317–23. The CPIC guidelines for warfarin provide peer-reviewed recommendations for translating pharmacogenetic test results into actionable prescribing descisions for warfarin management.
Johnson JA, Gong L, Whirl-Carrillo M, et al. Clinical pharmacogenetics implementation consortium guidelines for CYP2C9 and VKORC1 genotypes and warfarin dosing. Clin Pharmacol Ther. 2011;90:625–9.
• Wilke RA, Ramsey LB, Johnson SG, et al. The clinical pharmacogenomics implementation consortium: CPIC guideline for SLCO1B1 and simvastatin-induced myopathy. Clin Pharmacol Ther 2012;92:112–7. The CPIC guidelines for statin myopathy provide peer-reviewed recommendations for translating pharmacogenetic test results into actionable prescribing descisions for statin management.
Shuldiner AR, Palmer K, Pakyz RE, et al. Implementation of pharmacogenetics: the University of Maryland Personalized Anti-platelet Pharmacogenetics Program. Am J Med Genet C. 2014;166C:76–84.
Kushner FG, Hand M, Smith SC Jr, et al. 2009 focused updates: ACC/AHA guidelines for the management of patients with ST-elevation myocardial infarction (updating the 2004 guideline and 2007 focused update) and ACC/AHA/SCAI guidelines on percutaneous coronary intervention (updating the 2005 guideline and 2007 focused update) a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. J Am Coll Cardiol. 2009;54:2205–41.
Gurbel PA, Becker RC, Mann KG, Steinhubl SR, Michelson AD. Platelet function monitoring in patients with coronary artery disease. J Am Coll Cardiol. 2007;50:1822–34.
Angiolillo DJ, Fernandez-Ortiz A, Bernardo E, et al. Variability in individual responsiveness to clopidogrel: clinical implications, management, and future perspectives. J Am Coll Cardiol. 2007;49:1505–16.
Kazui M, Nishiya Y, Ishizuka T, et al. Identification of the human cytochrome P450 enzymes involved in the two oxidative steps in the bioactivation of clopidogrel to its pharmacologically active metabolite. Drug Metab Dispos. 2010;38:92–9.
Hulot JS, Bura A, Villard E, et al. Cytochrome P450 2C19 loss-of-function polymorphism is a major determinant of clopidogrel responsiveness in healthy subjects. Blood. 2006;108:2244–7.
Gurbel PA, Bliden KP, Hiatt BL, O’Connor CM. Clopidogrel for coronary stenting: response variability, drug resistance, and the effect of pretreatment platelet reactivity. Circulation. 2003;107:2908–13.
Sibbing D, Koch W, Gebhard D, et al. Cytochrome 2C19*17 allelic variant, platelet aggregation, bleeding events, and stent thrombosis in clopidogrel-treated patients with coronary stent placement. Circulation. 2010;121:512–8.
Momary KM, Dorsch MP, Bates ER. Genetic causes of clopidogrel nonresponsiveness: which ones really count? Pharmacotherapy. 2010;30:265–74.
•• Mega JL, Simon T, Collet JP, et al. Reduced-function CYP2C19 genotype and risk of adverse clinical outcomes among patients treated with clopidogrel predominantly for PCI: a meta-analysis. JAMA 2010;304:1821–30. This was one of the largest meta-analyses providing evidence for increased risk of cardiac events in CYP2C19 loss-of-function allele carriers treated with clopidogrel.
Holmes MV, Perel P, Shah T, Hingorani AD, Casas JP. CYP2C19 genotype, clopidogrel metabolism, platelet function, and cardiovascular events: a systematic review and meta-analysis. JAMA. 2011;306:2704–14.
• Holmes DR, Jr., Dehmer GJ, Kaul S, Leifer D, O’Gara PT, Stein CM. ACCF/AHA clopidogrel clinical alert: approaches to the FDA “boxed warning”: a report of the American College of Cardiology Foundation Task Force on clinical expert consensus documents and the American Heart Association endorsed by the Society for Cardiovascular Angiography and Interventions and the Society of Thoracic Surgeons. J Am Coll Cardiol 2010;56:321–41. This consensus report from the major cardiology groups provides a thorough overview of the controversy of pharmacogenetic testing to guide antiplatelet therapy prescribing following percutaneous coronary interventions.
Scott SA, Sangkuhl K, Gardner EE, et al. Clinical Pharmacogenetics Implementation Consortium guidelines for cytochrome P450-2C19 (CYP2C19) genotype and clopidogrel therapy. Clin Pharmacol Ther. 2011;90:328–32.
Wiviott SD, Braunwald E, McCabe CH, et al. Prasugrel versus clopidogrel in patients with acute coronary syndromes. N Engl J Med. 2007;357:2001–15.
Wallentin L, Becker RC, Budaj A, et al. Ticagrelor versus clopidogrel in patients with acute coronary syndromes. N Engl J Med. 2009;361:1045–57.
Manolio TA, Chisholm RL, Ozenberger B, et al. Implementing genomic medicine in the clinic: the future is here. Genet Med. 2013;15:258–67.
Levine GN, Bates ER, Blankenship JC, et al. 2011 ACCF/AHA/SCAI guideline for percutaneous coronary intervention: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines and the Society for Cardiovascular Angiography and Interventions. J Am Coll Cardiol. 2011;58:e44–122.
Pereira NL, Sargent DJ, Farkouh ME, Rihal CS. Genotype-based clinical trials in cardiovascular disease. Nat Rev Cardiol. 2015;12:475–87.
Bergmeijer TO, Janssen PW, Schipper JC, et al. CYP2C19 genotype-guided antiplatelet therapy in ST-segment elevation myocardial infarction patients-Rationale and design of the Patient Outcome after primary PCI (POPular) Genetics study. Am Heart J. 2014;168(16–22):e1.
Kazi DS, Garber AM, Shah RU, et al. Cost-effectiveness of genotype-guided and dual antiplatelet therapies in acute coronary syndrome. Ann Intern Med. 2014;160:221–32.
Lala A, Berger JS, Sharma G, Hochman JS, Scott Braithwaite R, Ladapo JA. Genetic testing in patients with acute coronary syndrome undergoing percutaneous coronary intervention: a cost-effectiveness analysis. J Thromb Haemost JTH. 2013;11:81–91.
•• Cresci S, Depta JP, Lenzini PA, et al. Cytochrome p450 gene variants, race, and mortality among clopidogrel-treated patients after acute myocardial infarction. Circ Cardiovasc Genet 2014;7:277–86. This study highlights the importance of investigating pharmacogenetic effects in several racial groups as they may differ by race.
Nelson A. Unequal treatment: confronting racial and ethnic disparities in health care. J Natl Med Assoc. 2002;94:666–8.
Wittkowsky AK, Devine EB. Frequency and causes of overanticoagulation and underanticoagulation in patients treated with warfarin. Pharmacotherapy. 2004;24:1311–6.
Budnitz DS, Lovegrove MC, Shehab N, Richards CL. Emergency hospitalizations for adverse drug events in older Americans. N Engl J Med. 2011;365:2002–12.
Daly AK, King BP. Pharmacogenetics of oral anticoagulants. Pharmacogenetics. 2003;13:247–52.
Li T, Chang CY, Jin DY, Lin PJ, Khvorova A, Stafford DW. Identification of the gene for vitamin K epoxide reductase. Nature. 2004;427:541–4.
Rieder MJ, Reiner AP, Gage BF, et al. Effect of VKORC1 haplotypes on transcriptional regulation and warfarin dose. N Engl J Med. 2005;352:2285–93.
• Cooper GM, Johnson JA, Langaee TY, et al. A genome-wide scan for common genetic variants with a large influence on warfarin maintenance dose. Blood 2008;112:1022–7. This is the first genomewide association study in European subjects which confirmed that VKORC1 and CYP2C9 are the major genetic contributors to warfarin dose requirements.
• Takeuchi F, McGinnis R, Bourgeois S, et al. A genome-wide association study confirms VKORC1, CYP2C9, and CYP4F2 as principal genetic determinants of warfarin dose. PLoS Genet 2009;5:e1000433. This genome-wide assocation study in European individuals identified CYP4F2 as a third gene that contributes to warfarin dose requirements.
Manolopoulos VG, Ragia G, Tavridou A. Pharmacogenetics of coumarinic oral anticoagulants. Pharmacogenomics. 2010;11:493–6.
Cavallari LH, Langaee TY, Momary KM, et al. Genetic and clinical predictors of warfarin dose requirements in African Americans. Clin Pharmacol Ther. 2010;87:459–64.
Warfarin (Coumadin) [package insert]. Princeton, NJ, USA: Bristol-Myers Squibb; 2010.
•• Pirmohamed M, Burnside G, Eriksson N, et al. A Randomized Trial of Genotype-Guided Dosing of Warfarin. New England Journal of Medicine 2013;369:2294–303. This was a European prospective genotype directed trial aimed at maintaining patients within a target INR range.
•• Kimmel SE, French B, Kasner SE, et al. A Pharmacogenetic versus a Clinical Algorithm for Warfarin Dosing. New England Journal of Medicine 2013;369:2283–93. This was a prospective genotype directed trial aimed at maintaining patients within a target INR range performed in the United States and included an ethnically diverse population.
• Perera MA, Cavallari LH, Limdi NA, et al. Genetic variants associated with warfarin dose in African-American individuals: a genome-wide association study. Lancet 2013;382:790–6. This is the first genome-wide assocation study performed in African ancestry individuals which discovered a novel CYP2C variant that predicted warfarin dose requirments in this ethnic group.
Limdi NA, Brown TM, Yan Q, et al. Race influences warfarin dose changes associated with genetic factors. Blood. 2015;126:539–45.
Ramirez AH, Shi Y, Schildcrout JS, et al. Predicting warfarin dosage in European-Americans and African-Americans using DNA samples linked to an electronic health record. Pharmacogenomics. 2012;13:407–18.
Alfirevic A, Neely D, Armitage J, et al. Phenotype standardization for statin-induced myotoxicity. Clin Pharmacol Ther. 2014;96:470–6.
Third Report of the National Cholesterol Education Program (NCEP). Expert panel on detection, evaluation, and treatment of high blood cholesterol in adults (adult treatment panel III) final report. Circulation. 2002;106:3143–421.
Stone NJ, Robinson JG, Lichtenstein AH, et al. 2013 ACC/AHA guideline on the treatment of blood cholesterol to reduce atherosclerotic cardiovascular risk in adults: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J Am Coll Cardiol. 2014;63:2889–934.
Ridker PM, Cook NR. Statins: new American guidelines for prevention of cardiovascular disease. Lancet. 2014;382:1762–5.
Ramsey LB, Johnson SG, Caudle KE, et al. The clinical pharmacogenetics implementation consortium guideline for SLCO1B1 and simvastatin-induced myopathy: 2014 update. Clin Pharmacol Ther. 2014;96:423–8.
Thompson PD, Clarkson P, Karas RH. Statin-associated myopathy. JAMA. 2003;289:1681–90.
SLCO1B1 Variants and Statin-Induced Myopathy—A Genomewide Study. New England Journal of Medicine 2008;359:789–99.
Kameyama Y, Yamashita K, Kobayashi K, Hosokawa M, Chiba K. Functional characterization of SLCO1B1 (OATP-C) variants, SLCO1B1*5, SLCO1B1*15 and SLCO1B1*15 + C1007G, by using transient expression systems of HeLa and HEK293 cells. Pharmacogenet Genom. 2005;15:513–22.
Pasanen MK, Neuvonen M, Neuvonen PJ, Niemi M. SLCO1B1 polymorphism markedly affects the pharmacokinetics of simvastatin acid. Pharmacogenet Genom. 2006;16:873–9.
Voora D, Shah SH, Spasojevic I, et al. The SLCO1B1*5 genetic variant is associated with statin-induced side effects. J Am Coll Cardiol. 2009;54:1609–16.
Donnelly LA, Doney AS, Tavendale R, et al. Common nonsynonymous substitutions in SLCO1B1 predispose to statin intolerance in routinely treated individuals with type 2 diabetes: a go-DARTS study. Clin Pharmacol Ther. 2011;89:210–6.
Danik JS, Chasman DI, MacFadyen JG, Nyberg F, Barratt BJ, Ridker PM. Lack of association between SLCO1B1 polymorphisms and clinical myalgia following rosuvastatin therapy. Am Heart J. 2013;165:1008–14.
Pasanen MK, Fredrikson H, Neuvonen PJ, Niemi M. Different effects of SLCO1B1 polymorphism on the pharmacokinetics of atorvastatin and rosuvastatin. Clin Pharmacol Ther. 2007;82:726–33.
Jung K, McBean AM, Kim JA. Comparison of statin adherence among beneficiaries in MA-PD plans versus PDPs. J Manag Care Pharm JMCP. 2012;18:106–15.
Li JH, Suchindran S, Shah SH, Kraus WE, Ginsburg GS, Voora D. SLCO1B1 genetic variants, long-term low-density lipoprotein cholesterol levels and clinical events in patients following cardiac catheterization. Pharmacogenomics. 2015;16:449–58.
Aslibekyan S, Claas SA, Arnett DK. To replicate or not to replicate: the case of pharmacogenetic studies: Establishing validity of pharmacogenomic findings: from replication to triangulation. Circ Cardiovasc Genet. 2013;6:409–12.
Ioannidis JP. To replicate or not to replicate: the case of pharmacogenetic studies: Have pharmacogenomics failed, or do they just need larger-scale evidence and more replication? Circ Cardiovasc Genet. 2013;6:413–8.
Relling MV, Altman RB, Goetz MP, Evans WE. Clinical implementation of pharmacogenomics: overcoming genetic exceptionalism. Lancet Oncol. 2010;11:507–9.
US Food and Drug Administration Drug Guidance. Pharmacokinetics in patients with impaired renal function- study design, data analysis, and impact on dosing and labeling. http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/UCM204959.pdf. Accessed 9 Mar 2016
Teutsch SM, Bradley LA, Palomaki GE, et al. The evaluation of genomic applications in practice and prevention (EGAPP) initiative: methods of the EGAPP Working Group. Genet Med. 2009;11:3–14.
United States Food and Drug Administration. Table of pharmacogenomics biomarkers in drug lavels. http://www.fda.gov/drugs/scienceresearch/researchareas/pharmacogenetics/ucm083378.htm. Accessed 9 Mar 2016.
Wang B, Canestaro WJ, Choudhry NK. Clinical evidence supporting pharmacogenomic biomarker testing provided in US Food and Drug Administration drug labels. JAMA internal medicine. 2014;174:1938–44.
Dunnenberger HM, Crews KR, Hoffman JM, et al. Preemptive clinical pharmacogenetics implementation: current programs in five US medical centers. Annu Rev Pharmacol Toxicol. 2015;55:89–106.
• Shuldiner AR, Relling MV, Peterson JF, et al. The Pharmacogenomics Research Network Translational Pharmacogenetics Program: overcoming challenges of real-world implementation. Clin Pharmacol Ther 2013;94:207–10. This paper discusses logistical barriers to implementation of pharmacogenomics in clinical practice.
Hoffman JM, Haidar CE, Wilkinson MR, et al. PG4KDS: a model for the clinical implementation of pre-emptive pharmacogenetics. Am J Med Genet C. 2014;166C:45–55.
Johnson JA, Elsey AR, Clare-Salzler MJ, Nessl D, Conlon M, Nelson DR. Institutional profile: University of Florida and Shands Hospital Personalized Medicine Program: clinical implementation of pharmacogenetics. Pharmacogenomics. 2013;14:723–6.
• Van Driest SL, Shi Y, Bowton EA, et al. Clinically actionable genotypes among 10,000 patients with preemptive pharmacogenomic testing. Clin Pharmacol Ther 2014;95:423–31. This study shows that preemptive genotyping would yield a high number of pharmacogenetic variants that would be clinically actionable.
Levy KD, Decker BS, Carpenter JS, et al. Prerequisites to implementing a pharmacogenomics program in a large health-care system. Clin Pharmacol Ther. 2014;96:307–9.
Johnson JA, Burkley BM, Langaee TY, Clare-Salzler MJ, Klein TE, Altman RB. Implementing personalized medicine: development of a cost-effective customized pharmacogenetics genotyping array. Clin Pharmacol Ther. 2012;92:437–9.
Gurwitz D, Lunshof JE, Dedoussis G, et al. Pharmacogenomics education: International Society of Pharmacogenomics recommendations for medical, pharmaceutical, and health schools deans of education. Pharmacogenom J. 2005;5:221–5.
Stanek EJ, Sanders CL, Taber KA, et al. Adoption of pharmacogenomic testing by US physicians: results of a nationwide survey. Clin Pharmacol Ther. 2012;91:450–8.
Haga SB, Burke W, Ginsburg GS, Mills R, Agans R. Primary care physicians’ knowledge of and experience with pharmacogenetic testing. Clin Genet. 2012;82:388–94.
Tuteja S, Haynes K, Zayac C, Sprague JE, Bernhardt B, Pyeritz R. Community pharmacists’ attitudes towards clinical utility and ethical implications of pharmacogenetic testing. Personal Med 2013;10
Kullo IJ, Haddad R, Prows CA, et al. Return of results in the genomic medicine projects of the eMERGE network. Front Genet. 2014;5:50.
Fullerton SM, Wolf WA, Brothers KB, et al. Return of individual research results from genome-wide association studies: experience of the Electronic Medical Records and Genomics (eMERGE) Network. Genet Med. 2012;14:424–31.
Peterson JF, Field JR, Shi Y, et al. Attitudes of clinicians following large-scale pharmacogenomics implementation. Pharmacogenomics J 2015.
Pacanowski M, Huang SM. Precision Medicine. Clin Pharmacol Ther. 2016;99:124–9.
Bredenoord AL, Kroes HY, Cuppen E, Parker M, van Delden JJ. Disclosure of individual genetic data to research participants: the debate reconsidered. Trends Genet. 2011;27:41–7.
Green RC, Berg JS, Grody WW, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15:565–74.
Rothstein MA. Currents in contemporary ethics. GINA, the ADA, and genetic discrimination in employment. J Law Med Ethics. 2008;36:841–840.
Patel HN, Ursan ID, Zueger PM, Cavallari LH, Pickard AS. Stakeholder views on pharmacogenomic testing. Pharmacotherapy. 2014;34:151–65.
Laedtke AL, O’Neill SM, Rubinstein WS, Vogel KJ. Family physicians’ awareness and knowledge of the Genetic Information Non-Discrimination Act (GINA). J Genet Couns. 2012;21:345–52.
Roden DM, Johnson JA, Kimmel SE, et al. Cardiovascular Pharmacogenomics. Circ Res. 2011;109:807–20.
Shuldiner AR, O’Connell JR, Bliden KP, et al. Association of cytochrome P450 2C19 genotype with the antiplatelet effect and clinical efficacy of clopidogrel therapy. JAMA. 2009;302:849–57.
Daly AK. Genome-wide association studies in pharmacogenomics. Nat Rev Genet. 2010;11:241–6.
Global Lipids Genetics C, Willer CJ, Schmidt EM, et al. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45:1274–83.
Arwood MJ, Cavallari LH, Duarte JD. Pharmacogenomics of hypertension and heart disease. Curr Hypertens Rep. 2015;17:586.
Oni-Orisan A, Lanfear DE. Pharmacogenomics in heart failure: where are we now and how can we reach clinical application? Cardiol Rev. 2014;22:193–8.
Cooper-DeHoff RM, Johnson JA. Hypertension pharmacogenomics: in search of personalized treatment approaches. Nat Rev Nephrol. 2016;12:110–22.
Funding
Supported in part by grants from the National Heart, Lung, and Blood Institute (RO1HL092173; 1K24HL133373).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
Sony Tuteja reports grants from the University of Pennsylvania Health System for the study of the pharmacogenomics of antiplatelet drugs, during the conduct of study. Nita Limdi reports grants from NIH (NHLBI).
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical collection on Cardiovascular Genetics.
Rights and permissions
About this article
Cite this article
Tuteja, S., Limdi, N. Pharmacogenetics in Cardiovascular Medicine. Curr Genet Med Rep 4, 119–129 (2016). https://doi.org/10.1007/s40142-016-0096-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40142-016-0096-z