Abstract
We examine the origin of honey bee (Apis mellifera) populations in Kangaroo Island (Australia), Norfolk Island (Australia) and the Kingdom of Tonga using a highly polymorphic mitochondrial DNA region and a panel of 37 single nucleotide polymorphisms that assigns ancestry to three evolutionary lineages: Eastern Europe, Western Europe and Africa. We also examine inbreeding coefficients and genetic variation using microsatellites and mitochondrial sequencing. The honey bees of Kangaroo Island have a high proportion of Eastern European ancestry (90.2%), consistent with claims that they are of the subspecies A. m. ligustica. The honey bees of Norfolk Island also had a majority of ancestry from Eastern Europe (73.1%) with some contribution from Western Europe (21.2%). The honey bees of Tonga are mainly of Western European (70.3%) origin with some Eastern European ancestry (27.4%). Despite the suspected severe bottlenecks experienced by these island population, inbreeding coefficients were low.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The Western honey bee Apis mellifera is the world’s most widely distributed and well-known pollinator. Honey bees contribute hundreds of billions of dollars to the world’s economy each year by increasing the quantity, quality and value of crops that require pollination (vanEngelsdorp et al. 2008; Gallai et al. 2009; Keogh et al. 2010). Its importance for honey and wax production, and for crop pollination has seen its human-assisted spread from its native range of Africa, the Middle East and Europe to cover most of the world (Crane 1983; Winston 1987).
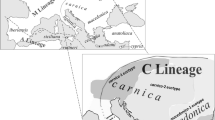
The origin and genetic differences of the world’s honey bee populations is of considerable interest (e.g. Ruttner 1988; Whitfield et al. 2006; Han et al. 2012). There are more than 26 named subspecies (Ruttner 1988; Meixner et al. 2013), which can be assigned to 4 evolutionary lineages: Eastern European (C), Western European (M), African (A) and Middle Eastern (O) (Garnery et al. 1992; Franck et al. 2000b, 2001; Alburaki et al. 2013; Meixner et al. 2013). Different subspecies have different traits that make them more or less suitable for commercial apiculture and pollination (Ruttner 1988; Uzunov et al. 2014), and therefore more or less favoured by beekeepers.
Initial introductions (prior to 1850) into the New World and Australasia were mostly from Western Europe (e.g. Hopkins 1886; Seeley 1985; Cornuet 1986). Later introductions were largely from Eastern Europe, where the genotypes are more suited to commercial beekeeping. In the twentieth century, there was considerable interest in trying new strains and subspecies in various locations, leading to numerous imports into and out of subspecies’ natural ranges (Adam 1983; Seeley 1985; Cornuet 1986). Often, there were secondary introductions via another nation, for example from the UK (native) to Australia (introduced) then to New Zealand (introduced; Hopkins 1886; Barrett 1996). Hence, these often-small introductions underwent two genetic bottlenecks. This is particularly the case for small island nations or communities that tended to get their honey bees from nearby populations.
In this paper, we examine the population genetics and origins of three island populations, Kangaroo Island, Norfolk Island and the Kingdom of Tonga, that were likely derived from Australia and/or New Zealand. Australia has a diverse honey bee population (Oldroyd et al. 1992, 1995; Koulianos and Crozier 1997; Chapman et al. 2008; Oxley and Oldroyd 2009; Chapman et al. 2016), having imported honey bees from all over the world (e.g. Hopkins 1886; Weatherhead 1986; Barrett 1996). Australia has a large feral population (Oldroyd et al. 1994, 1997; Oldroyd 1998; Arundel et al. 2012; Hinson et al. 2015) with a different genetic background from the commercial population, as determined through mitochondrial, microsatellite and SNP (single nucleotide polymorphism) markers (Chapman et al. 2008, 2015, 2016). At least four subspecies of honey bee were introduced to New Zealand (Hopkins 1886; Cook 1967; Donovan 1980; Barrett 1996; Newstrom-Lloyd 2013).
Kangaroo Island is located 13.5 km from the Australian mainland, 112 km southwest of Adelaide. It is a large island of 4405 km2. The economy of the island is mainly based on agriculture and tourism. The first honey bees introduced to the island were likely A. m. mellifera from the Western European lineage (Hopkins 1886; Eckert 1958; Glatz 2015). In the 1880s, a small number of A. m. ligustica colonies were introduced to Kangaroo Island from Italy via Brisbane (Queensland) and Adelaide (South Australia) (Hopkins 1886; Glatz 2015). The source in Italy is reported as Bologna, Naples, or ‘Italy’ (Eckert 1958; Woodward 1993; Koulianos and Crozier 1996; Jolly 2004; Barrett 2010b; Glatz 2015). Regardless of the area they were imported from, all are expected to be A. m. ligustica. There are reports of a concerted effort to remove colonies of A. m. mellifera origin from the island, and to institute a bee breeding program and genetic ‘sanctuary’ based on A. m. ligustica (Woodward 1993; Jolly 2004; Barrett 2010b; Glatz 2015).
The Ligurian bee sanctuary was established on Kangaroo Island with The Ligurian Bee Act (Government of South Australia 1885) to prevent A. m. mellifera from being imported so that the island could be used for the production of Italian queens (Eckert 1958; Barrett 2010b; Glatz 2015). The sanctuary is maintained to this day at the behest of beekeepers and on the basis that the isolated population is pure A. m. ligustica. The keeping of any bee other than ‘Italian’ or ‘Ligurian’ (i.e. A. m. ligustica) strains is prohibited by the Act. The low incidence of honey bee disease on the Island (Roberts et al. 2015) provides further support for the protection of the island’s honey bees and the ban on the importation of honey bees and bee products is current.
Because of its isolation, there has been considerable scientific interest in this population (Eckert 1958; Ruttner 1976; Woyke 1976; Oldroyd et al. 1992; Woodward 1993; Koulianos and Crozier 1996, 1997; Jolly 2004; Glatz 2015), and concerns over inbreeding (Woyke 1976). Oldroyd et al. (1992), using samples from 24 colonies, curiously found that honey bees on Kangaroo Island were morphologically similar to A. m. ligustica or A. m. siciliana (C lineage), had malate dehydrogenase (MDH) alleles typical of A. m. siciliana from Sicily and A. m. ligustica from Southern Italy, but mitochondrial restriction enzyme digestion patterns suggestive of an A. m. mellifera origin. Oldroyd et al. (1992) presented a number of hypotheses to explain these somewhat anomalous findings, including that incomplete surveys of the A. m. ligustica population in Italy meant that M lineage mitochondrial haplotypes had not been reported from Italy at that time. Koulianos and Crozier (1996, 1997) confirmed that the honey bees of Kangaroo Island carry mitochondrial haplotypes consistent with M lineage origin. M lineage mitotypes have since been found in Italy, and it has been determined that there was ancient hybridization between the M and C lineages in Italy (Badino et al. 1982; Badino et al. 1983; Franck et al. 2000a). Franck et al. (2000a) concluded that Kangaroo Island is probably a rare example of the introduction of Italian M mitotypes.
Norfolk Island is a small (34.6 km2) volcanic island located 1412 km off the east coast of Australia. Honey bees were probably introduced to the island between 1822 and 1855 (Malfroy et al. 2016). SNP testing (Chapman et al. 2015) of six samples revealed them to be hybrids of mainly Eastern European origin (average 80.7%) with a significant component of Western European origin (average 17.6%) (Malfroy et al. 2016). The only honey bee pests and diseases recorded on Norfolk Island are the lesser wax moth (Achroia grisella), the microsporidian parasite Nosema ceranae and Lake Sinai virus I (Malfroy et al. 2016), suggesting that the population has been isolated for many decades. Currently, there are approximately 130 managed hives on the island. The number of feral colonies has not been estimated but the feral population was described as ‘large’ (Malfroy et al. 2016). The last known bee imports occurred in 1992 (Malfroy et al. 2016).
The Kingdom of Tonga, in Polynesia, is an archipelago of 171 islands, of which about 35 are inhabited. The total land area is 748 km2. Primary production provides the majority of employment, with most rural Tongans relying on agriculture for both subsistence and income (van der Velde et al. 2007). There are anecdotal reports (White et al. 2016) of honey bees being imported from New Zealand in the 1970s and from Fiji to service pollination requirements during a period of high pumpkin production for export to Japan (1988–1998; Murray 2001). The origin of honey bees introduced to New Zealand (Donovan and Macfarlane 1984; Barrett 1996), Fiji (Anderson 1990; Barrett 2010a) and Tonga has not been studied using genetic tools, and so the genetic background of Tongan honey bees is unknown. The use of honey bees for pollination ceased after an outbreak of American foulbrood, resulting in elimination of the majority of managed hives (Pers. Comm. Bruce White to EJR). Feral honey bees and a small number of domestic hives remain on the major island groups of Tongatapu, ‘Eua, Ha’apai and Vava’u. A secondary introduction of queen honey bees from New Zealand to Vava’u in 2006 resulted in the introduction of Varroa destructor mites (Pers. Comm. Bruce White to EJR). The feral population on Vava’u naturally survive Varroa mite infestation without chemical intervention (Remnant et al. 2017).
This paper uses a panel of 37 single nucleotide polymorphisms (SNPs) (Chapman et al. 2017) and mitochondrial sequencing of a highly polymorphic intergenic region between the tRNAleu and cytochrome oxidase II genes (Crozier and Crozier 1993) to investigate the genetic origins of honey bees on Kangaroo Island, Norfolk Island and Tonga. Genetic variability within these populations is assessed using microsatellite markers.
2 Material and methods
2.1 Kangaroo Island
One bee was sampled from each of 99 colonies, from eight beekeepers (Figure 1), and preserved under ethanol before extracting DNA using phenol-chloroform (Pritchard et al. 2000).
Maps of a the region, showing Kangaroo Island, Norfolk Island and the Kingdom of Tonga, sampling locations on b Kangaroo Island and c Norfolk Island, d the Kingdom of Tonga and e Tongatapu, f ‘Eua and g the Vava’u island group. Sampling locations are denoted with red dots. Numbers next to the dots represent the number of colonies sampled per apiary where this is known.
2.2 Norfolk Island
A sample was collected from 1 to 10 hives from 20 of the 27 apiaries on the island, with at least 60 individuals sampled per apiary (Malfroy et al. 2016). We extracted DNA using phenol-chloroform (Pritchard et al. 2000) from 2 to 6 individuals from each apiary, resulting in 94 individuals for analysis. As it was possible that more than one individual was sampled per colony, we analysed microsatellite data (see below) using COLONY (Jones and Wang 2010) to identify and remove individuals originating from the same colony, resulting in a verified sample size of 34 individuals. The number of verified colonies sampled at each apiary is given in Figure 1.
2.3 Tonga
We collected samples from 34 colonies on three islands: Eua, Tongatapu and Vava’u. We extracted DNA from one bee per colony using phenol-chloroform (Pritchard et al. 2000). Sampling locations and the number of colonies sampled at each location, where known, are given in Figure 1.
2.4 SNP ancestry assignment
We typed samples from Kangaroo Island (n = 99), Norfolk Island (n = 34) and Tonga (n = 34) with a panel of 37 SNPs (Chapman et al. 2017) using the Agena Sequenom MassARRAY MALDI-TOF system at the Australian Genomic Research Facility, with manual review of allele calls. We assigned the proportion of ancestry of each individual to three ancestral lineages (African, Eastern European and Western European) using data for 37 SNPs (Chapman et al. 2017) from reference samples originating from a previous study (Chapman et al. 2015) in STRUCTURE (v2.3.4; Pritchard et al. 2000). We used a burn-in phase of 50,000 iterations with individuals from the three reference populations and then assigned ancestry to our test samples according to an admixture model with uncorrelated allele frequencies in 100,000 Markov chain Monte Carlo iterations using k = 3 populations. Five replicates were run in order to confirm that ancestry assignment was consistent across runs. Admixture was measured for each individual as 1 minus the maximum proportion of ancestry to a cluster. Thus, an individual with 95% C ancestry is 5% admixed and an individual with 57% M ancestry is 43% admixed.
2.5 Microsatellites
We analysed length polymorphisms at microsatellite loci A107, A113, A14, A29, A79, A8, Ap53 and B124 (Estoup et al. 1993; Solignac et al. 2003) according to Chapman et al. (2008) for 99 individuals from Kangaroo Island, 34 from Norfolk Island and 33 from Tonga. We tested for systematic genotyping errors caused by allelic dropout or stuttering and the presence of null alleles using MICRO-CHECKER (van Oosterhout et al. 2004). We enumerated the number of alleles in each population, calculated observed and expected heterozygosity, tested for departures for Hardy-Weinberg equilibrium, linkage disequilibrium and estimated inbreeding coefficients (FIS) and genetic differentiation between the populations (FST) using GENEPOP (Raymond and Rousset 1995) on the web (v4.2). We determined allelic richness using PopGenReport 3.0.0 (Gruber and Adamack 2017) in the R statistical program (R Development Core Team 2017). We assessed population structure in STRUCTURE with 50,000 iterations in a burn-in phase followed by 100,000 Markov chain Monte Carlo iterations using 1 < k < 8 populations with no admixture assumed. We determined the best k following Evanno et al. (2005).
We used BOTTLENECK (v1.2.02; Piry et al. 1999) to test for a recent genetic bottleneck using the Wilcoxon sign-rank test of excess heterozygosity under the two-phase model assuming 95% single-step mutation and 5% multi-step changes with 1000 simulations. Population were also assessed for a mode-shift from an L-shaped distribution of allele frequencies.
2.6 Mitochondria
We sequenced (Macrogen, South Korea) a highly variable region of the mitochondrial genome, containing part of the tRNAleu gene, an intergenic region and part of the cytochrome oxidase II (COII) gene (Crozier and Crozier 1993) using the E2 and H2 primers (Garnery et al. 1992) following the methods of Chapman et al. (2008) from 76 individuals from Kangaroo Island, 34 individuals from Norfolk Island and 33 individuals from Tonga. Sequences were checked using SEQUENCHER v5.1 (Gene Codes Corporation, USA) and aligned using MUSCLE (EMBL-EBI).
3 Results
3.1 SNP analysis
On average, the honey bees on Kangaroo Island have 90.2% Eastern European ancestry, 7.8% Western European ancestry and 2.0% African ancestry when typed with our 37 SNP panel (Figure 2c). Mean admixture of the population was 9.8%.
a tRNAleu—COII intergenic mitochondrial haplotypes, H1-H4 are from the M lineage while H5-H7 are from the C lineage; b STRUCTURE analysis on microsatellite data with k = 2 clusters; c ancestry proportions of honey bees from Kangaroo Island, Norfolk Island and the Kingdom of Tonga (Tongatapu, ‘Eua and Vava’u) from three ancestral lineages (C, M and A) assigned based on 37 SNPs in reference samples using STRUCTURE. Position in each graph corresponds to the same individual, such that mitochondrial haplotype can be matched with microsatellite structure and SNP ancestry testing.
The honey bees of Norfolk Island have an average of 73.1% Eastern European ancestry, 21.2% Western European ancestry and 5.7% African ancestry (Figure 2). Mean admixture of the population was 26.9%.
The honey bees of the Kingdom of Tonga were of mainly Western European ancestry (70.3%) with some Eastern European (27.4%) and little African ancestry (2.3%). The honey bees of Tongatapu had an average of 41.1% Eastern European ancestry, 57.3% Western European ancestry and 1.6% African ancestry. Mean admixture of the population was 36.1%. The honey bees of ‘Eua had an average 29.6% Eastern European ancestry, 68.1% Western European ancestry and 2.3% African ancestry. Mean admixture of the population was 31.9%. The honey bees of Vava’u had an average 21.7% Eastern European ancestry, 75.6% Western European ancestry and 2.7% African ancestry. Mean admixture of the population was 23.9% (Figure 2).
3.2 Microsatellite analysis
All microsatellite loci were polymorphic, with an average of 12.1 alleles per locus and average allelic richness of populations between 3.07 and 7.46 (Table I). MICRO-CHECKER suggested that locus Ap53 had null alleles in the Kangaroo Island population, in the combined Tonga population and in the ‘Eua population and was therefore excluded from further analysis (Supplementary Information). MICRO-CHECKER suggested that there were no systematic genotyping errors due to allelic dropout or stuttering.
Observed and expected heterozygosity were both high and similar for all three populations (Table I). One locus was not in Hardy-Weinberg equilibrium in the Kangaroo Island population (A113), and one in the combined Tongan population (A107), while all loci were in Hardy-Weinberg equilibrium for Norfolk Island (Table I), or when the three Tonga populations were considered separately (Table I). Four pairs of loci showed significant linkage disequilibrium in the Kangaroo Island population, two in the Norfolk Island population and seven in the Tongan population (Supplementary Information). There were not enough samples to test for linkage disequilibrium on Tongatapu, and as estimates on the other two Tongan island groups are likely to be skewed, we provide data for the combined population only. The inbreeding coefficient (FIS) was low for all populations (Kangaroo Island, 0.012; Norfolk Island, − 0.042; Tonga, − 0.008) and for the three island groups in the Kingdom of Tonga (Tongatapu, − 0.089; ‘Eua, − 0.141; Vava’u, 0.045; Table I).
STRUCTURE found that k = 2 best fit the microsatellite data, with the samples of Kangaroo Island forming one cluster, while samples from Norfolk Island and the Kingdom of Tonga form the other cluster (Figure 2b). FST between Kangaroo Island and Norfolk Island is 0.157, it is 0.315 between Kangaroo Island and the Kingdom of Tonga and 0.141 between Norfolk Island and the Kingdom of Tonga. The FST between Tongatapu and ‘Eua is 0.071, 0.005 between Tongatapu and Vava’u and 0.082 between ‘Eua and Vava’u.
All populations had a normal L-shaped distribution of allele frequencies. No populations show evidence of a recent bottleneck (Kangaroo Island P = 1.000, Norfolk Island P = 0.055, Kingdom of Tonga P = 0.996, Tongatapu P = 0.961, ‘Eua P = 0.945, Vava’u P = 0.996).
3.3 Mitochondrial analysis
We found seven sequence variants (mitochondrial haplotypes) in the P and Q motifs of the intergenic region between the tRNAleu and COII genes (Supplementary Information; GenBank accession numbers MG828857-MG828863; Figure 2a). Three mitochondrial haplotypes were from the C (Eastern Europe) lineage and four from the M lineage (Western Europe). The M7 haplotype is found in Italy and Spain (Franck et al. 2001), the M79a haplotype is found in A. m. iberiensis (Chávez-Galarza et al. 2017), while the two other M haplotypes are previously unreported. The C1 haplotype is found in A. m. ligustica (Franck et al. 2001), the C19 haplotype has been reported in Turkey (Solorzano et al. 2009) and the remaining C haplotype is previously unreported. As expected (Oldroyd et al. 1992; Koulianos and Crozier 1996) the majority (96.1%) of the samples from Kangaroo Island carried M haplotypes (one M7 and 74 unreported M haplotype 2) while one individual carried the C19 haplotype (Supplementary Information). The majority of individuals (22) on Norfolk Island carried the C1 haplotype, while the remainder (12) carried unreported C haplotype 6. The M7, M79a and unrecorded haplotype 2 were found on ‘Eua and Vava’u. Unrecorded haplotype 3 was only found on Tongatapu, and the C1 haplotype was found on Tongatapu and ‘Eua (Supplementary Information).
4 Discussion
Genetic diversity within populations, and indeed within colonies, is important to the maintenance of colony health and productivity (Tarpy and Page 2002; Mattila and Seeley 2007; Harpur et al. 2012; Tarpy et al. 2013). Our study shows that the honey bee populations on Kangaroo Island, Norfolk Island and Tonga have low inbreeding coefficients (FIS), despite the fact that Kangaroo Island and Norfolk Island are closed populations, and imports into Tonga are presumably rare and the number of managed colonies is extremely low. We also note that a single mated queen carries in her spermatheca 75% of the genetic variation present in the population from which she is derived (Ding et al. 2017). Presumably, most island populations are descended from several queens, meaning that genetic bottlenecking at the time of introduction is unlikely to be severe.
The honey bees on Kangaroo Island have a high proportion (90.2%) of ancestry from the C lineages and low admixture (9.8%), as determined with a SNP test (Chapman et al. 2017) for ancestry proportions from three ancestral lineages. This is consistent with this population being of A. m. ligustica origin and is higher than the proportion of C lineage ancestry in commercial (69.4%) and feral (57.4%) populations in the rest of Australia (Chapman et al. 2015, 2016). Previous studies have determined that the honey bees of Kangaroo Island have mitochondrial haplotypes from the M lineage found in Western Europe and Italy (Oldroyd et al. 1992; Koulianos and Crozier 1996; Franck et al. 2000a). The majority (74 of 76) of typed individuals carry a M haplotype that is previously unreported, while one individual carried the M7 haplotype. For the first time, we have identified an individual with a C lineage mitochondrial haplotype on Kangaroo Island. The origin of this haplotype is speculative. However, as this study has more than double the sample size of previous studies (24 samples, Oldroyd et al. 1992; 7 samples, Koulianos and Crozier 1996), its absence from previous studies on Kangaroo Island may merely be an effect of sampling. The haplotype is previously unreported in Australia (Chapman et al. 2008; Oxley and Oldroyd 2009) or Italy (Franck et al. 2000a). A second individual from this colony showed the same haplotype, and therefore this finding is not due to sequencing or other error. The individual in question does not stand out in any way in the SNP data, having 87.1% C lineage, 11.2% M lineage and 1.7% A lineage ancestry. If beekeepers are concerned about propagating this rare mitochondrial haplotype, even though it is likely to be of Italian heritage, then it could potentially be excluded from breeding programs via queen selection based on a simple mitochondrial marker. However, in our view, such a strategy would be pointless.
SNP genotyping revealed that the honey bees of Norfolk Island have ancestry mostly from Eastern Europe (C lineage; 73.1%), with some Western European (M lineage; 21.2%) and little evidence of African ancestry (A lineage; 5.7%). Only two mitochondrial haplotypes were present, both from Eastern Europe. One was the C1 allele associated with A. m. ligustica, and the other is a previously unreported mitotype that differs from the C1 mitotype by one base.
The Tongan populations were in Hardy-Weinberg equilibrium; however, there was linkage disequilibrium between seven microsatellite markers. The last known introduction occurred in 2006 to Vava’u (Pers. Comm. Bruce White to EJR) and it may be that the population has not yet come to equilibrium. More likely is population subdivision across the sampled islands. SNP genotyping revealed that ancestry differs between the three Tonga Island groups sampled; however, all three had a majority Western European origin with negligible African ancestry. Five mitochondrial haplotypes were detected, with likely origins from Spain, Portugal and Italy. Surprisingly, the two closest islands, ‘Eua and Tongatapu were not the most closely related, pointing to ‘Eua having different population origins to Tongatapu and Vava’u.
It is important to note that a low proportion of SNPs of African origin in all of our studied populations does not mean that the populations have had any admixture with African populations now or in the past. It just says that they carry a small number of SNPs that are present in African populations at high frequency.
In conclusion, we found low inbreeding coefficients in A. mellifera populations on Kangaroo Island, Norfolk Island and in Tonga. Future work could consider measuring brood viability or enumerating the number of alleles at the complementary sex determination locus (Lechner et al. 2014) which is central to sex determination and lethal when homozygous (Woyke 1963; Beye et al. 2003). Unsurprisingly, the three populations differed in the proportion of ancestry from the Eastern European, Western European and African lineages, as evidenced by a SNP test that assigns ancestry to these three lineages (but not the O lineage) and from mitochondrial haplotypes. The honey bees of Kangaroo Island have a high proportion of nuclear (SNP) alleles of Eastern European ancestry, consistent with claims that they are of A. m. ligustica origin. The honey bees of Norfolk Island also have a high proportion of Eastern European ancestry, but more Western European ancestry than the Kangaroo Island population. The honey bees of Tonga are of mostly Western European ancestry.
Data availability
The datasets generated during the current study are available at http://hdl.handle.net/2123/18818
References
Adam B. (1983) In Search of the Best Strains of Bees. Northern Bee Books, Hebden Bridge, West Yorkshire, United Kingdom.
Alburaki M., B. Bertrand, H. Legout, S. Moulin, A. Alburaki, et al. (2013) A fifth major genetic group among honeybees revealed in Syria. BMC Genetics 14: 117.
Anderson D.L. (1990) Pests and pathogens of the honeybee (Apis mellifera L.) in Fiji. Journal of Apicultural Research 29: 53–59.
Arundel J., B.P. Oldroyd, S. Winter. (2012) Modelling honey bee queen mating as a measure of feral colony density. Ecological Modelling 247: 48–57.
Badino G., G. Celebrano, A. Manino. (1982) Genetic variability of Apis mellifera ligustica Spin. in a marginal area of its geographic distribution. Experientica 38: 540–541.
Badino G., G. Celebrano, A. Manino. (1983) Population structure and Mdh-1 locus variation in Apis mellifera ligustica. Journal of Heredity 74: 443–446.
Barrett P. (1996) The Immigrant Bees 1788 to 1898: A Cyclopaedia on the Introduction of European Honeybees into Australia and New Zealand. P Barrett, Springwood, NSW, Australia.
Barrett P. (2010a) A history of the introduction of honey bees into Fiji. P. Barrett, Caloundra, Queensland, Australia.
Barrett P. (2010b) Kangaroo Island: the isle of (almost) forgotten bees. P. Barrett, Springwood, NSW, Australia.
Beye M., M. Hasselmann, M.K. Fondrk, R.E. Page, S.W. Omholt. (2003) The gene csd is the primary signal for sexual development in the honeybee and encodes an SR-type protein. Cell 114: 419–429.
Chapman N.C., A.L. Bourgeois, L.D. Beaman, J. Lim, B.A. Harpur, et al. (2017) An abbreviated SNP panel for ancestry assignment of honeybees (Apis mellifera). Apidologie 48: 776–783.
Chapman N.C., B.A. Harpur, J. Lim, T.E. Rinderer, M.H. Allsopp, et al. (2015) A SNP test to identify Africanized honeybees via proportion of ‘African’ ancestry. Molecular Ecology Resources 15: 1346–1355.
Chapman N.C., B.A. Harpur, J. Lim, T.E. Rinderer, M.H. Allsopp, et al. (2016) Hybrid origins of Australian honeybees (Apis mellifera). Apidologie 47: 26–34.
Chapman N.C., J. Lim, B.P. Oldroyd. (2008) Population genetics of commerical and feral honey bees in Western Australia. Journal of Economic Entomology 101: 272–277.
Chávez-Galarza J., L. Garnery, D. Henriques, C.J. Neves, W. Loucif-Ayad, et al. (2017) Mitochondrial DNA variation of Apis mellifera iberiensis: further insights from a large-scale study using sequence data of the tRNAleu-cox2 intergenic region. Apidologie 48: 533–544.
Cook V.A. (1967) Facts about beekeeping in New Zealand. Bee World 48: 88–100.
Cornuet J.M. (1986) Population Genetics, in: Rinderer T.E. (Ed.), Bee Genetics and Breeding, Academic Press, Orlando, Florida, USA, pp. 235–254.
Crane E. (1983) The Archaeology of Beekeeping. Cornell University Press, Ithaca, NY, USA.
Crozier R.H., Y.C. Crozier. (1993) The mitochondrial genome of the honeybee Apis mellifera: complete sequence and genome organization. Genetics 133: 97–117.
Ding G., H. Xu, B.P. Oldroyd, R. Gloag. (2017) Extreme polyandry aids the establishment of invasive populations of a social insect. Heredity 119: 318–387.
Donovan B.J. (1980) Interactions between native and introduced bees in New Zealand. New Zealand Journal of Ecology 3: 104–116.
Donovan B.J., R.P. Macfarlane. (1984) Bees and Pollination, in: Scott R.R. (Ed.), New Zealand Pest and Beneficial Insects, Lincoln University of Agriculture, Christchurch, New Zealand, pp. 247–270.
Eckert J.E. (1958) The Kangaroo Island Ligurian bees. Gleanings in Bee Culture 86: 660–663, 722–725.
Estoup A., M. Solignac, M. Harry, J.M. Cornuet. (1993) Characterization of (GT)n and (CT)n microsatellites in two insect species: Apis mellifera and Bombus terrestris. Nucleic Acids Research 21: 1427–1431.
Evanno G., S. Regnaut, J. Goudet. (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: 2611–2620.
Franck P., L. Garnery, G. Celebrano, M. Solignac, J.M. Cornuet. (2000a) Hybrid origins of honeybees from Italy (Apis mellifera ligustica) and Sicily (A. m. sicula). Molecular Ecology 9: 907–921.
Franck P., L. Garnery, A. Loiseau, B.P. Oldroyd, H.R. Hepburn, et al. (2001) Genetic diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86: 420–430.
Franck P., L. Garnery, M. Solignac, J.M. Cornuet. (2000b) Molecular confirmation of a fourth lineage in honeybees from the Near East. Apidologie 31: 167–180.
Gallai N., J.M. Salles, J. Settele, B.E. Vaissière. (2009) Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecological Economics 68: 810–821.
Garnery L., J.M. Cornuet, M. Solignac. (1992) Evolutionary history of the honey bee Apis mellifera inferred from mitochondrial DNA analysis. Molecular Ecology 1: 145–154.
Glatz R.V. (2015) Curious case of the Kangaroo Island honeybee Apis mellifera Linnaeus, 1758 (Hymenoptera: Apidae) sanctuary. Austral Entomology 54: 117–126.
Government of South Australia. (1885) The Ligurian Bee Act - 1885. Government of South Australia, Adelaide, Australia.
Gruber B., A. Adamack. (2017) PopGenReport: a simple framework to analyze population and landscape genetic data, Canberra, ACT, Australia.
Han F., A. Wallberg, M.T. Webster. (2012) From where did the Western honeybee (Apis mellifera) originate?. Ecology and Evolution 2: 1949–1957.
Harpur B.A., S. Minaei, C.F. Kent, A. Zayed. (2012) Management increases genetic diversity of honey bees via admixture. Molecular Ecology 21: 4414–4421.
Hinson E.M., M. Duncan, J. Lim, J. Arundel, B.P. Oldroyd. (2015) The density of feral honey bee (Apis mellifera) colonies in South East Australia is greater in undisturbed than in disturbed habitats. Apidologie 46: 403–413.
Hopkins I. (1886) Illustrated Australasian Bee Manual. I. Hopkins, Auckland, New Zealand.
Jolly B. (2004) South Australia’s early Ligurian beekeeping - and a lingering Kangaroo Island fable. Journal of the Historical Society of South Australia 32: 69–81.
Jones O., J. Wang. (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Molecular Ecology Resources 10: 551–555.
Keogh R.C., A.P.W. Robinson, I.J. Mullins. (2010) Pollination aware: the real value of pollination in Australia, Rural Industries Research and Development Corporation, Canberra, ACT, Australia.
Koulianos S., R. Crozier. (1996) Mitochondrial DNA sequence data provides further evidence that the honeybees of Kangaroo Island, Australia are of hybrid origin. Apidologie 27: 165–174.
Koulianos S., R. Crozier. (1997) Mitochondrial sequence characterisation of Australian commercial and feral honeybee strains, Apis mellifera L. (Hymenoptera: Apidae), in the context of the species worldwide. Australian Journal of Entomology 36: 359–363.
Lechner S., L. Ferretti, C. Schöning, W. Kinuthia, D. Willemsen, et al. (2014) Nucleotide variability at its limit? Insights into the number and evolutionary dynamics of the sex-determining specificities of the honey bee Apis mellifera. Molecular Biology and Evolution 31: 272–287.
Malfroy S.F., J.M.K. Roberts, S. Perrone, G. Maynard, N. Chapman. (2016) A pest and disease survey of the isolated Norfolk Island honey bee (Apis mellifera) population. Journal of Apicultural Research 55: 202–211.
Mattila H.R., T.D. Seeley. (2007) Genetic diversity in honey bee colonies enhances productivity and fitness. Science 317: 362–364.
Meixner M.D., M.A. Pinto, M. Bouga, P. Kryger, E. Ivanova, et al. (2013) Standard methods for characterising subspecies and ecotypes of Apis mellifera, in: Dietemann V., Ellis J.D., and Neumann P. (Eds.), The COLOSS BEEBOOK, Volume I: standard methods for Apis mellifera research. Journal of Apicultural Research 52, pp. IBRA.1.52.54.05.
Murray W.E. (2001) The second wave of globalisation and agrarian change in the Pacific Islands. Journal of Rural Studies 17: 135–148.
Newstrom-Lloyd L.E. (2013) Pollination in New Zealand, Ecosystem services in New Zealand - conditions and trends, Manaaki Whenua Press, Lincoln, New Zealand.
Oldroyd B.P. (1998) Controlling feral honey bee, Apis mellifera L. (Hymenoptera: Apidae), populations in Australia: methodologies and costs. Australian Journal of Entomology 37: 97–100.
Oldroyd B.P., J.M. Cornuet, D. Rowe, T.E. Rinderer, R.H. Crozier. (1995) Racial admixture of Apis mellifera in Tasmania, Australia: similaritites and differences with natural hybrid zones in Europe. Heredity 74: 315–325.
Oldroyd B.P., S.H. Lawler, R.H. Crozier. (1994) Do feral honey-bees (Apis mellifera) and regent parrots (Polytelis anthopeplus) compete for nest sites. Australian Journal of Ecology 19: 444–450.
Oldroyd B.P., W.S. Sheppard, J.A. Stelzer. (1992) Genetic characterization of the bees of Kangaroo Island, South Australia. Journal of Apicultural Research 31: 141–148.
Oldroyd B.P., E.G. Thexton, S.H. Lawler, R.H. Crozier. (1997) Population demography of Australian feral bees (Apis mellifera). Oecologia 111: 381–387.
Oxley P.R., B.P. Oldroyd. (2009) Mitochondrial sequencing reveals five separate origins of ‘Black’ Apis mellifera (Hymenoptera: Apidae) in Eastern Australian commercial colonies. Journal of Economic Entomology 102: 480–484.
Piry S., G. Luikart, J.M. Cornuet. (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. Journal of Heredity 90: 502–503.
Pritchard J.K., M. Stephens, P. Donnelly. (2000) Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
R Development Core Team. (2017) R: a language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria.
Raymond M., F. Rousset. (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity 86: 248–249.
Remnant E.J., M. Shi, G. Buchmann, T. Blaquiere, E.C. Holmes, et al. (2017) A diverse range of novel RNA viruses in geographically distinct honey bee populations. Journal of Virology 91: 16.
Roberts J., D. Anderson, P. Durr. (2015) Upgrading knowledge on pathogens (particularly viruses) of Australian honey bees, Rural Industries Research and Development Corporation, Canberra, Australia.
Ruttner F. (1976) Isolated populations of honeybee in Australia. Journal of Apicultural Research 15: 97–104.
Ruttner F. (1988) Biogeography and Taxonomy of Honeybees. Springer-Verlag, Berlin, Germany.
Seeley T.D. (1985) Honeybee Ecology: A Study of Adaptation in Social Life. Princeton University Press, Princeton, New Jersey, USA.
Solignac M., D. Vautrin, A. Loiseau, F. Mougel, E. Baudry, et al. (2003) Five hundred and fifty microsatellite markers for the study of the honeybee (Apis mellifera L.) genome. Molecular Ecology Notes 3: 307–311.
Solorzano C.D., A.L. Szalanski, M. Kence, J.A. McKern, J.W. Austin, et al. (2009) Phylogeography and population genetics of honey bees (Apis mellifera) from Turkey based on COI-COII sequence data. Sociobiology 53: 237–246.
Tarpy D.R., R.E. Page. (2002) Sex determination and the evolution of polyandry in honey bees (Apis mellifera). Behavioral Ecology and Sociobiology 52: 143–150.
Tarpy D.R., D. vanEngelsdorp, J.S. Pettis. (2013) Genetic diversity affects colony survivorship in commercial honey bee colonies. Naturwissenschaften 100: 723–728.
Uzunov A., C. Costa, B. Panasiuk, M. Meixner, P. Kryger, et al. (2014) Swarming, defensive and hygenic behaviour in honey bee colonies of different genetic origin in a pan-European experiment. Journal of Apicultural Research 53: 248–260.
van der Velde M., S.R. Green, M. Vanclooster, B.E. Clothier. (2007) Sustainable development in small island developing states: agricultural intensification, economic development, and freshwater resources management on the coral atoll of Tongatapu. Ecological Economics 61: 456–468.
van Oosterhout C., W.F. Hutchinson, D.P.M. Wills, P. Shipley. (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes 4: 535–538.
vanEngelsdorp D., J. Hayes, R.M. Underwood, J. Pettis. (2008) A survey of honey bee colony losses in the US, fall 2007 to spring 2008. PLoS One 3: e4071.
Weatherhead T. (1986) Boxes to Bar Hives: Beekeeping History or Queensland. International Colour Productions, Stanthorpe, QLD, Australia.
White B., E. Remnant, L. Osborne. (2016) Beekeeping in the Kingdom of Tonga. Honeybee News 9: 34–38.
Whitfield C.W., S.K. Behura, S.H. Berlocher, A.G. Clark, J.S. Johnston, et al. (2006) Thrice out of Africa: ancient and recent expansions of the honey bee, Apis mellifera. Science 314: 643–645.
Winston M.L. (1987) The Biology of the Honey Bee. Harvard University Press, Cambridge, MA, USA.
Woodward D. (1993) Ligurian bees. American Bee Journal 133: 124–125.
Woyke J. (1963) What happens to diploid drone larvae in a honeybee colony? Journal of Apicultural Research 2: 73–75.
Woyke J. (1976) Population genetic studies on sex alleles in the honeybee using the example of the Kangaroo Island bee sanctuary. Journal of Apicultural Research 15: 105–123.
Acknowledgements
We thank Peter Davis for collecting bees from Kangaroo Island, Bruce White and Lamorna Osborne for collecting bees from Tonga and the beekeepers of Norfolk Island for providing access to their colonies to SFM. We thank two anonymous reviewers for their helpful comments which improved the manuscript.
Funding
This project was supported by Rural Industries Research and Development Corporation (AgriFutures) Australia PRJ-007774 (BPO), Science and Innovation Award for Young People in Agriculture, Fisheries and Forestry: CSIRO Biosecurity Flagship 2015 (NCC), Natural Sciences and Engineering Research Council Canada Discovery grant (AZ), Ontario Ministry of Research and Innovation’s Early Researcher Award (AZ), Natural Sciences and Engineering Research Council Canada Postdoctoral Fellowship (BAH), Australian and Pacific Science Foundation grant APSF15-02 (EJR), the Australian Government and the Norfolk Island Administration (SFM) and Rural Industries Research and Development Corporation (AgrFutures) Australia PRJ-008540 (JMKR).
Author information
Authors and Affiliations
Contributions
NCC, BPO, SFM, JMKR and EJR designed the experiment. NCC and JS extracted DNA and performed genotyping. NCC performed data analysis. AZ and BAH provided SNPs and data. NCC, BPO, SFM, JMKR, EJR, JL, MHA and TER provided samples. NCC and BPO wrote the paper, with suggestions from co-authors. All authors read and approved the article prior to publication.
Corresponding author
Additional information
Manuscript editor: Marina Meixner
Origines génétiques des abeilles mellifères ( Apis mellifera ) sur les îles Kangourou et Norfolk (Australie) et au Royaume de Tonga
Apis mellifera / assignation d’ascendance / génétique des populations / populations isolées / polymorphismes nucléotidiques simples
Genetischer Ursprung der Honigbienen ( Apis mellifera ) auf Kangaroo Island und Norfolk Island (Australien) und im Königreich von Tonga
Apis mellifera / Vorfahrensbestimmung / Populationsgenetik / isolierte Populationen / Single Nucleotide Polymorphismus
Electronic supplementary material
ESM 1.
(DOCX 18 kb)
Rights and permissions
About this article
Cite this article
Chapman, N.C., Sheng, J., Lim, J. et al. Genetic origins of honey bees (Apis mellifera) on Kangaroo Island and Norfolk Island (Australia) and the Kingdom of Tonga. Apidologie 50, 28–39 (2019). https://doi.org/10.1007/s13592-018-0615-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-018-0615-x