Abstract
Mesenchymal stem cell (MSCs)-derived exosomes have been frequently used as useful tools in disease control. This research aimed to study the function of MSC-derived exosomes (Exo) in the stemness of cancer stem cells (CSCs) of hepatocellular carcinoma (HCC) and the molecular mechanism. Exo from the procured human bone marrow-MSCs were extracted and identified. CSCs from HCC cell lines were collected. The CSCs were treated with Exo, and then the proliferation, migration, invasion, angiogenesis-stimulating and self-renewal abilities of the Hep3B-CSCs and HuH7-CSCs were significantly reduced. C5orf66-AS1 was found as the most upregulated long noncoding RNAs (lncRNAs) in CSCs after Exo treatment. The integrated bioinformatic analyses and luciferase assays suggested that C5orf66-AS1 upregulated DUSP1 expression through sequestering microRNA-127-3p (miR-127-3p). Either artificial overexpression of miR-127-3p or silencing of DUSP1 blocked the inhibitory functions of Exo in the CSCs. DUSP1 inhibition increased the phosphorylation of ERK. Similar results were reproduced in vivo where Exo reduced the growth of xenograft formed by CSCs in nude mice, and this reduction was blocked upon miR-127-3p overexpression or DUSP1 silencing. To conclude, this research reported that MSC-derived Exo block malignant behaviors of HCC-sourced CSCs through a C5orf66-AS1/miR-127-3p/DUSP1/ERK axis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver cancer represents the seventh commonest cancer and the second leading cause of cancer-related morbidity with over 840,000 new diagnosis and 780,000 deaths in 2018 worldwide [1]. Hepatocellular carcinoma (HCC) represents the predominant type of liver cancer with infections of hepatitis B virus and hepatitis C virus as the major risk factors [2]. Especially, China alone takes up to over a half of death cases [3]. An early diagnosis is crucial for a better prognosis of patients; however, most patients were not diagnosed until the late stages with an extremely low survival rate due to the rapid development, intrahepatic recurrence and metastasis [4]. Cancer stem cells (CSCs) are deemed as a major cause of tumor recurrence, metastasis, and a dismal prognosis [5]. They reside at the top of tumor heterogeneity and are inherently resistant to chemo or radiotherapy, therefore, resulting in treatment resistance and diffusion [6, 7]. Reducing maintenance of the stem cell properties might help enhance the treatment response and provide higher survival opportunity for HCC patients.
Mesenchymal stem/stromal cells (MSCs) have been frequently used as a source of cellular therapy owing to their potent immunosuppressive and regenerative effects [8]. The most frequently used sources of MSCs are bone marrow (BM-MSCs), umbilical cord, and adipose tissues [9]. Interestingly, the inherent tumor-tropic characteristic of MSCs make them as candidate tools to target cancer cells, and they have been reported as a two-edged sword in cancer treatment since both oncogenic and tumor-suppressing roles of MSCs have been revealed with the involvement of the MSC-derived exosomes (hereafter termed Exo, specifically for the exosomes extracted from our procured BM-MSCs) [10, 11]. Exosomes are a type of extracellular vesicles and key performers in intercellular communication, which represent major vehicles recapitulating the biological activity of MSCs [10]. However, the functions of MSCs and Exo in the development of HCC and, especially, in the behaviors of HCC-sourced CSCs remain unknown. Exosomes are well-known as carriers of genetic molecules including proteins, mRNA, microRNA (miRNAs), and other RNA forms including transfer RNA, nucleolar RNA, ribosomal RNA, and long noncoding RNA (lncRNA) are also contained [11, 12]. LncRNAs are a major type of non-coding RNAs whose dysregulation plays crucial roles in tumorigenesis and stemness of CSCs [13]. Here, we focused on the function of Exo in HCC-CSCs and the candidate responsible lncRNAs involved. Using a lncRNA microarray analysis and integrated bioinformatic analyses, we predicted a lncRNA C5orf66-AS1/miR-127-3p/dual-specificity phosphatase 1 (DUSP1) RNA network that is possibly involved in the malignant behaviors of the HCC-CSCs. Therefore, this study aimed examine the role of Exo in HCC-CSCs, and gain- and loss-of function studies were conducted to validate the involvements of these molecules.
Materials and methods
Clinical samples
HCC tumor tissues and the para-cancerous tissues were collected from 50 patients with HCC admitted into and treated in the First Affiliated Hospital of Xinjiang Medical University from September 2018 to December 2019. All included patients did not have a history of chemo- or radio-therapy. Signed informed consent was acquired from each participant. The research was approved by the Ethical Committee of the First Affiliated Hospital of Xinjiang Medical University (Approval number: K201807-05) and performed according to the Declaration of Helsinki.
Cell culture and transfection
Human BM-MSCs (PCS-500-012), human umbilical vein endothelial cells (HUVECs, PCS-100-010), human hepatic epithelial cells (THLE-3, CRL-11233) and HCC cells Hep3B (HB-8064) were procured from ATCC (Manassas, VA, USA), and another HCC cell line HuH7 (CL-0120) was procured from Procell Life Science & Technology Co., Ltd. (Wuhan, Hubei, China). All cells were cultured in DMEM (HyClone, UT, USA)-with 10% FBS (Gibco Company, Grand Island, NY, USA) in a 37 °C incubator with 5% CO2.
The miR-127-3p overexpressing vector miR-127-3p mimic, small interfering (si)-RNA of DUSP1 and C5orf66-AS1, and the negative control (NC) vectors were procured from GenePharma Co., Ltd. (Shanghai, China). The sequences of si-DUSP1 and si-C5orf66-AS1 are as follows: si-DUSP1: Guide sequence: 5'-AAAUAAAUAAGGUAUAUUCUC-3'; Passenger sequence:5'-GAAUAUACCUUAUUUAUUUUU-3'; si-C5orf66-AS1: Guide sequence: 5'- UCGUUUAGGAAAAAUACCCUC-3'; Passenger sequence:5'- GGGUAUUUUUCCUAAACGAAA-3'. All vectors were transfected into cells using a Lipofectamine 2000 kit (Thermo Fisher Scientific Inc., CA, USA).
Identification of the MSCs and Exo
The surface antigens of MSCs were examined by flow cytometry. In short, 1 × 106 MSCs were incubated with phycoerythrin-conjugated antibodies against CD34 (Cat. No. #343505), CD45 (Cat. No. #304007), CD73 (Cat. No. #344003) and CD90 (Cat. No. #328109) at 4 °C in the dark for 30 min. Then, the cells were resuspended in 3 mL phosphate-buffered saline (PBS), centrifuged, and loaded with 300 μL PBS. The background marker was determined using the homomonoclonal antibody immunoglobulin G (IgG, Cat. No. #400113). The fluorescent cells were measured on a flow cytometer. The positive rate of the surface antigens was analyzed using the FlowJo software (Tree Star, Ashland, OR, USA). All the antibodies were acquired from Biolegend (San Diego, CA, USA). The differentiation potential of the MSCs were identified using adipogenic differentiation (GUXMX-90031) and osteogenic differentiation (GUXMX-90021) induction kits procured from Cyagen Biosciences Inc. (Guangdong, China).
The MSCs at passage 3 were incubated in serum-free medium for 24 h, and the supernatants were centrifuged at 2000g at 4 °C for 20 min to discard the cell debris and dead cells and ultra-centrifuged at 100,000g for 1 h. The sediment was then resuspended in serum-free DMEM containing 25 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (pH = 7.4) and further ultra-centrifuged 100,000g at 4 °C for 2 h. Then, the sediment was stored at − 80 °C until further use. The particles were quantified using a bicinchoninic acid (BCA) kit. The morphology, specific biomarkers and the particle size distribution were analyzed using a transmission electron microscope (TEM, Thermo Fisher Scientific), western blot analysis detection, and nanoparticle tracking analysis (NTA).
Western blot analysis
Total protein in cells was isolated using RIPA cell lysis buffer. After concentration determination using a BCA kit, the protein samples were separated by SDS-PAGE and loaded onto PVDF membranes. After being blocked in 5% non-fat milk for 1 h, the membranes were bybridized with primary antibodies at 4 °C overnight and with the horseradish peroxidase-conjugated secondary antibody at 37 °C for 2 h. The blot bands were visualized using the ECL reagent (Beyotime Biotechnology Co., Ltd., Shanghai, China) and analyzed using Image J (Bio-Rad, Hercules, CA, USA). The antibodies are listed in Table 1 in which GAPDH was used as the internal control.
Isolation and identification of the CSCs
CSCs were isolated from HCC cells using the flow cytometer. In short, the HCC cells were harvested and resuspended in PBS. Then, the cells were cultured with fluorescein isothiocyanate (FITC, Biolegend)-conjugated anti-CD44 (Cat. No. #338803) and allophycocyanin-conjugated anti-CD90 (Cat. No. #328113) in the dark for 30 min, and the isotype anti-mouse IgG (all acquired from Biolegend) was used as control. The freshly isolated CSCs were harvested using a flow cytometer and secondly identified using phycoerythrin-conjugated anti-CD133 (Cat. No. #372803).
Examination of the internalization of exosomes
The Exo (250 μg) were labeled using PKH26 (red fluorescence) in compliance with the instructions of a PKH26 labeling kit (Sigma-Aldrich) and washed in PBS and centrifuged at 1000 × g at 4 °C for 2 h. Then, the PKH-labeled Exo (10 μg/mL) were added into the CSCs (3 × 105) for 24 h of warm incubation. The cells were immobilized in 4% paraformaldehyde (PFA) and cleared in 0.1% Triton × 100. The nuclei were stained using 4', 6-diamidino-2-phenylindole (Sigma-Aldrich). The staining was observed under a confocal microscope (LSM 780, Zeiss Inc, AG, Oberkochen, Germany).
MTT assay
Cell viability was examined using an MTT kit (Beyotime). In short, CSCs were loaded in 96-well plates at 5 × 103 cells per well. Each well was filled with 10 μL MTT solution (5 mg/mL) for 4 h of cell incubation. After that, each well was supplemented with 100 μL dissolving solution for 3 h to dissolve the formazan. The optical density (OD) at 570 nm was read using a spectrophotometer (Tecan, Grodig, Austria).
Cell counting kit-8 (CCK-8) method
Proliferation ability of CSCs was first examined using a CCK-8 kit (Beyotime). CSCs were resuspended to 2 × 103 cells/100 μL and loaded into the 96-well plates. Each well was added with 100 μL cell suspension and 10 μL CCK-8 solution, followed by further incubation for 0, 24, 48 and 72 h, respectively. After incubation, the OD at 450 nm was examined using the spectrophotometer.
Colony formation assay
CSCs were incubated for 14 d in 6-well plates at 100 cells per well. The cells were fixed in 4% PFA and stained with 0.1% crystal violet (Sigma-Aldrich) at 25 °C for 15 min. The cells were rinsed using distilled water, and the number of colonies formed by CSCs was counted under an inverted microscope (Leica, Solms, Germany).
Wound healing assay
CSCs were incubated in 6-well plates at 1 × 104 cells per well till reaching a 90% confluence. Then, a pipette head was applied to make scratches at an equal distance on the cell surface, and then the cells were cultured in FBS-free DMEM for 24 h. The width of the scratches at 0 h and 24 h was captured under the microscope. Relative migration of cells was examined using the Image J software.
Transwell assay
Invasion of cells was examined using the Transwell assay. In short, 1 × 104 CSCs were resuspended in 200 μL FBS-free DMEM. The cells were loaded into Transwell apical chambers pre-coated with 200 mg/mL Matrigel (Corning Life Sciences, NY, USA). The chambers were inserted into 24-well plates containing 700 μL complete medium in each well. After 24 h of warm incubation, the cells on the inner membrane were discarded, and those invaded to the outer membrane were fixed in 4% PFA and stained with crystal violet for 15 min. The invading cells were counted under the microscope with five random fields included and analyzed using the Image J.
Limited-dilution assay (LDA)
The self-renewal ability of the CSCs was examined using an LDA as previously described [14]. In short, the cell number was adjusted to 10,000 cells/mL and followed by serial dilutions. Then, the cells were sorted to serial dilutions (Corning). The final cell density was maintained at 100 to 400 cells per well in 0.2-mL aliquots. Next, cells were cultured in serum-free DMEM supplemented B27 (Thermo Fisher Scientific), 20 ng/mL basic fibroblast growth factor, 20 ng/mL epidermal growth factor, 4 μg/mL heparin (Sigma-Aldrich) and 1% methyl cellulose. After 7 d, the number of tumor spheres was counted under the microscope.
Angiogenesis assay
The Matrigel was pre-coated on 24-well plates at 37 °C for 30 min of polymerization. HUVECs were seeded on the plates at 1 × 105 per well and cultured in different CSC-conditioned media (CM) for 12 h. The length of the tubes and the number of tube branches were observed under an inverted microscope and calculated using the Image J.
RT-qPCR
Total RNA was isolated using a TRIzol kit (Thermo Fisher Scientific). RNA samples were reverse-transcribed using a PrimeScript® miRNA cDNA Synthesis kit (TaKaRa, Dalian, China) or a PrimeScript® RT reagent kit (TaKaRa). Real-time qPCR was conducted using an SYBR Green qPCR Master Mix kit (Thermo Fisher Scientific) on a 7500 Real-Time PCR System (Applied Biosystems, CA, USA). The primers are shown in Table 2 where U6 and GAPDH were used as the internal controls for miRNA and other genes, respectively. Relative gene expression was examined using the 2−ΔΔCT method.
Microarray analysis
RNA samples from Hep3B-CSCs were reverse-transcribed into cDNA using a rtStar™ First-Strand cDNA Synthesis Kit (Arraystar Inc., Rockville, MD, USA). LncRNAs were hybridized with a nrStar™ Human Functional LncRNA PCR Array (ArrayStar). Differentially expressed lncRNAs were screened according to fold changes in expression, and the functional and disease-related lncRNAs were categorized using the Arraystar-configured database to screen the cancer-related lncRNAs.
Nuclear-cytoplasmic RNA separation assay
The nuclear and cytoplasmic RNA from cells was isolated using a PARIS kit (Life Technologies, Gaithersburg, MD, USA). The expression of C5orf66-AS1 in nuclear and cytoplasmic RNA was determined using RT-qPCR with U6 and GAPDH serving as the internal controls, respectively.
RNA pull-down assay
Biotinylated RNA probes Bio-NC and Bio-miR-127-3p were synthesized by KeyGen Biotech Company (Shanghai, China). The RNA probes were warm incubated with CSC lysates and extracted using streptavidin-conjugated magnet beads according to the instruction manual of a Pierce™ Magnetic RNA Pull-Down Kit. The RNA–RNA compounds were diluted using saline solution and purified using TRIzol. The enrichment of C5orf66-AS1 in the compounds were examined using RT-qPCR.
Dual-luciferase reporter gene assay
Putative binding sequences (wild type, WT) between C5orf66-AS1 or DUSP1 mRNA with miR-217-3p were obtained from the bioinformatic system Starbase (http://starbase.sysu.edu.cn/) and the mutant type (MT) sequences were designed. The WT and MT sequences were inserted to the pMIR-REPORT™ vector (Ambion, Austin, TX, USA) to construct C5orf66-AS1-WT, C5orf66-AS1-MT, DUSP1-WT and DUSP1-MT luciferase reporter vectors. The vectors were co-transfected with NC mimic or miR-217-3p mimic into 293 T cells (ATCC) using Lipofectamine 2000. After 48 h, the luciferase activity in cells was examined using a Dual-Luciferase Reporter Assay Kit (Promega).
Xenograft tumors in nude mice
Thirty-two male BALB/c nude mice (6 weeks old, 20–22 g) were procured from Vital River Co., Ltd. (Beijing, China). All animal procedures were conducted in compliance with the Guide for the Care and Use of Laboratory Animals (NIH, Bethesda, Maryland, USA). Significant efforts were made to reduce the suffering of conscious animals. The mice were assigned into four groups (n = 8 in each): Control group, Exo group, Exo + miR-127 mimic group and Exo + miR-127 + si-DUSP1 group. Mice were injected with Hep3B-CSCs with corresponding treatments. The Hep3B-CSC suspension was mixed with an equal volume of Matrigel and adjusted to 1 × 104 cells/200 μL. Next, 200 μL such mixed solution was injected to the ventral side of the nude mice through a subcutaneous injection. The weekly change in the tumor volume was recorded. The volume (V) was calculated as follows: V = L × W2/2 (mm3), in which ‘L’ refers to length and ‘W’ refers to width. After 4 weeks, all mice were killed via an intraperitoneal injection of an overdose of pentobarbital sodium (200 mg/kg) to collect the tumor tissues for subsequent experiments.
Immunohistochemical (IHC) staining
The tissues of xenograft tumors were cut into 4-μm sections. The sections were dewaxed, rehydrated, and incubated with antigen retrieval solution (p0081, Beyotime) at 22 °C for 10 min, with endogenous peroxidase blocker for 10 min (Biodragon, Beijing, China), and with 5% FBS at 22 °C for 1 h. Then, the sections were hybridized with the primary antibodies against Ki67 (1:200, ab16667, Abcam Inc., Cambridge, MA, USA) and p-Erk (1:200, #4370, Cell Signaling Technology, Beverly, MA, USA) at 4 °C overnight, and then with the secondary antibody to IgG (1:2,000, ab205718, Abcam) at 37 °C for 30 min and then treated with 3,3'-diaminobenzidine (4A Biotech, Beijing, China) for 30 min. The nuclei were stained using hematoxylin (B25380, Yuanye Bio-Technology Co. Ltd, Shanghai, China) for 10 min. The staining was examined under the microscope with five random fields included. Positive gene expression rate was evaluated as the ratio of positive cells to total cells.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 8.02 (GraphPad Software, San Diego, CA, USA). To each experiment, at least three repetitions were performed. Data were presented as mean ± standard deviation (SD). Correlations between variables were determined by Pearson’s correlation analysis. Differences were evaluated using the t test (every two groups) or one-way or two-way analysis of variance (ANOVA, over two groups). Tukey’s multiple comparison was used for the post-hoc test. The clinical information was analyzed using the Fisher’s exact test. p < 0.05 was considered to show significant difference.
Results
Identification of the MSCs and the Exo
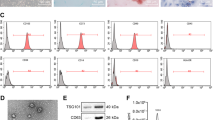
The procured MSCs were identified using flow cytometry concerning the expression of surface antigens. The MSCs showed positive expression of CD73 and CD90 while negative expression of CD34 and CD45 (Fig. 1A), which was aligned with the characteristics of MSCs. In addition, the MSCs showed osteogenic and adipogenic differentiation potentials according to the alizarin red staining and Oil red O staining, respectively (Fig. 1B). Then, the MSC-derived Exo were extracted. Under the TEM, the collected particles were in an oval shape (Fig. 1C). The NTA results showed that the size of the particles was mainly distributed in around 100 nm (Fig. 1D). Positive expression of the exosome marker proteins CD81 and TSG101 whereas negative expression of Calnxin was identified according to western blot analysis (Fig. 1E). Collectively, the Exo were successfully extracted.
Identification of the MSCs and the Exo. A MSC-specific biomarkers CD73 and CD90 in the acquired BM-MSCs determined by flow cytometry (Number of independent experiments; N = 3); B osteogenic and adipogenic differentiation abilities of the MSCs examined by alizarin red staining and Oil red O staining, respectively (N = 3); C shape of the particles under the TEM; D particle size distribution of the collected particles examined by NTA (N = 3); E expression of the exosome surface marker proteins determined by western blot analysis (N = 3); F CSCs from HCC cells screened via flow cytometry (N = 3); G positive rate of CD133 in the screened CSCs examined by flow cytometry (N = 3)
Next, the HCC cell lines HuH7 and Hep3B were subjected to flow cytometry to screen the CD90+ and CD44+ CSCs [15], termed HuH7-CSC and Hep3B-CSC (Fig. 1F), respectively. The flow cytometry results further confirmed high positive rate of CD133 [16] in the screened cells, which confirmed the stemness characteristics of the CSCs (Fig. 1G). Therefore, the HCC-sourced CSCs were successfully collected as well.
Exo treatment reduces proliferation, migration and invasion of CSCs
Before examining the direct impact of Exo on the behaviors of CSCs, we first examined the uptake of Exo by CSCs via immunofluorescence staining (Fig. 2A). The PKH26-labeled Exo were absorbed by the CSCs 24 h after Exo treatment.
Exo treatment reduces proliferation, migration and invasion of CSCs. A absorption of Exo by CSCs examined by immunofluorescence staining (N = 3); B viability of CSCs after different doses of Exo treatment examined by MTT assays (N = 3); C proliferation ability of CSCs determined by CCK-8 methods (N = 3); D colony formation ability of cells examined using colony formation assays; E migration ability of CSCs examined by wound healing assays (N = 3); F invasion ability of CSCs determined by Transwell assays (N = 3). Data were expressed as the mean ± SD. Differences were compared by one-way ANOVA (B) or two-way ANOVA (C–F). *p < 0.05 vs. the control group
Next, the CSCs were treated with an ascending series of Exo with PBS treatment as control. The MTT assay indicated that Exo treatment reduced viability of the CSCs in a dose-dependent manner (Fig. 2B). When the Exo concentration reached 5 μg/mL, the viability of cells reduced by half. Therefore, this concentration was selected in the subsequent experiments, and the role of Exo treatment in the malignant behaviors of CSCs was determined.
After Exo treatment, the proliferation ability of cells, according to the CCK-8 method, was significantly reduced (Fig. 2C). In concert with this, the colony formation ability of CSCs was reduced after Exo treatment (Fig. 2D). The wound healing assay indicated that Exo treatment weakened the migration ability of cells (Fig. 2E). Likewise, the Transwell assay results indicated that the invasion potential of CSCs was reduced as well after Exo treatment (Fig. 2F).
Exo treatment reduced angiogenesis and self-renewal ability of the CSCs
The CM for CSCs were collected for tube formation assays (Fig. 3A). It was found that the tuber formation ability of CSCs was reduced, presented by decreased tube length and reduced branches. Then, the self-renewal ability of cells was examined by LDA (Fig. 3B). At the 100, 150 and 200 cell levels, the number of tumor spheres formed by CSCs was significantly reduced.
Exo treatment reduces angiogenesis and self-renewal ability of the CSCs. A angiogenesis of CSCs after Exo treatment examined by tube formation assays (N = 3); B self-renewal ability of the CSCs after Exo treatment determined by LDA (N = 3). Data were expressed as the mean ± SD. Differences were compared by two-way ANOVA (A–B). *p < 0.05 vs. the control group
Exo treatment enhances expression of C5orf66-AS1 in CSCs
To explore the molecules involved, we first screened the differentially expressed lncRNAs between CSCs with or without Exo treatment. For the sake of reducing experimental costs, we only selected the Hep3B-CSCs which showed higher self-renewal ability and tumorigenesis potential for the lncRNA microarray analysis, and the functional and disease-related lncRNAs were categorized using the Arraystar-configured database. The top 10 differentially expressed lncRNA are shown in Fig. 4A, among which C5orf66-AS1 was upregulated to a highest degree in Hep3B-CSCs after Exo treatment. Thereafter, the expression of C5orf66-AS1 in HuH7-CSCs was examined. It was found that the C5orf66-AS1 expression in HuH7-CSC was significantly elevated after Exo treatment (Fig. 4B). An enrichment of C5orf66-AS1 was found in the Exo (Fig. 4C).
Exo treatment enhances expression of C5orf66-AS1 in CSCs. A differentially expressed lncRNAs in Hep3B-CSCs with or without Exo treatment screened by a lncRNA microarray analysis (N = 3); B expression of C5orf66-AS1 in HuH7-CSCs after Exo treatment determined by RT-qPCR (N = 3); C enrichment of C5orf66-AS1 in Exo examined by RT-qPCR (N = 3); D expression of C5orf66-AS1 in MSCs after si-C5orf66-AS1 transfection examined by RT-qPCR (N = 3); E expression of C5orf66-AS1 in Exo-NC and Exo-KD examined by RT-qPCR (N = 3); F expression of C5orf66-AS1 in Hep3B-CSCs and HuH7-CSCs after Exo-NC and Exo-KD treatment determined by RT-qPCR (N = 3); G expression of C5orf66-AS1 in MSCs, THLE-3, Hep3B, HuH7, Hep3B-CSCs and HuH7-CSCs examined by RT-qPCR (N = 3); H expression of C5orf66-AS1 in the collected HCC tumor tissues and the adjacent normal tissues determined by RT-qPCR (number of sample size; n = 50). Data were expressed as the mean ± SD. Differences were compared by unpaired t test (B–E), paired t test (H) or one-way ANOVA (G). *p < 0.05 vs. the control group, si-NC group, or the Exo-NC group; ##p < 0.01 vs. supernatant; %p < 0.05 vs. the adjacent tissues; &p < 0.05 vs. the MSC group; @p < 0.05 vs. the THLE-3 group
To determine whether the Exo treatment enhanced the C5orf66-AS1 expression in CSCs by exogenous addition or endogenous induction, the si-C5orf66-AS1) or the si-NC was transfected into MSCs to obtain MSCs with poor expression of C5orf66-AS1 (Fig. 4D). Thereafter, the exosomes from these two sorts of MSCs were collected and named Exo-KD or Exo-NC, respectively. Then, the expression of C5orf66-AS1 in the Exo-KD and Exo-NC was examined and significantly reduced expression of C5orf66-AS1 was identified in Exo-KD (Fig. 4E). After that, the CSCs were treated with the Exo, and the C5orf66-AS1 expression in Exo-KD-treated CSCs was significantly lower than that in Exo-NC treated cells (Fig. 4F). These results indicated that Exo treatment enhanced the C5orf66-AS1 expression in CSCs by exogenous addition rather than endogenous induction.
Following the findings above, we further examined the expression of C5orf66-AS1 in different cell types. It was observed that the C5orf66-AS1 expression was significantly higher than that in other cell lines. Of note, compared to that in the human hepatic epithelial cells THLE-3, the C5orf66-AS1 expression was further reduced in the HCC cell lines (HuH7 and Hep3B) and in particular in the CSCs (HuH7-CSC and Hep3B-CSC) (Fig. 4G). Moreover, the C5orf66-AS1 expression was decreased in the HCC tumor tissues relative to the adjacent normal tissues (Fig. 4H). According to the mean value (0.359) of C5orf66-AS1 expression, the patients were assigned into low C5orf66-AS1 expression group (n = 24) and high C5orf66-AS1 expression group (n = 26). Then, it was found that poor expression of C5orf66-AS1 was linked to increased tumor size, cirrhosis and advanced Barcelona Clinic Liver Cancer (BCLC) stage (Table 3).
C5orf66-AS1 binds to miR-127-3p
To explore the function of C5orf66-AS1 in HCC, we first examined the sub-cellular localization of C5orf66-AS1 in the CSC cells (Fig. 5A). It was found that C5orf66-AS1 was mainly sub-localized in the cytoplasm in Hep3B-CSCs and HuH7-CSCs. Therefore, we predicted the candidate target miRNAs of C5orf66-AS1 using two bioinformatic systems, including Starbase and RNA22 (https://cm.jefferson.edu/rna22/), and three candidate transcripts were predicted: miR-4731-5p, miR-127-3p and miR-149-5p (Fig. 5B). Next, the expression of these three miRNAs in CSCs after Exo treatment was examined. Importantly, it was found that only the expression of miR-127-3p was significantly reduced (Fig. 5C). The expression of miR-125-3p in tissues was then explored. Importantly, high expression of miR-127-3p was identified in the HCC tumor tissues compared to the normal tissues (Fig. 5D), which was negatively correlated with the expression of C5orf66-AS1 (Fig. 5E).
C5orf66-AS1 binds to miR-127-3p. A sub-cellular localization of C5orf66-AS1 in Hep3B-CSCs and HuH7-CSCs examined by a nuclear-cytoplasmic RNA separation assay (N = 3); B candidate miRNAs having binding relationship with C5orf66-AS1 predicted from two bioinformatic systems; C expression of miR-4731-5p, miR-127-3p and miR-149-5p in Hep3B-CSCs and HuH7-CSCs examined by RT-qPCR (N = 3); D expression of miR-127-3p in HCC tumor tissues and the adjacent tissues determined by RT-qPCR (n = 50); E a negative correlation between miR-127-3p and C5orf66-AS1 in HCC tumor tissues; F binding relationship between C5orf66-AS1 and miR-127-3p examined using an RNA pull-down assay (N = 3); G putative (WT) and MT C5orf66-AS1 sequences designed for luciferase assays; H binding relationship between C5orf66-AS1 and miR-127-3p validated using a dual luciferase reporter gene assay (N = 3). Data were expressed as the mean ± SD. Differences were compared by paired t test (D) or two-way ANOVA (C, F and H). *p < 0.05 vs. the control group; #p < 0.05 vs. the adjacent tissues; &&p < 0.01 vs. the Bio-NC group; %p < 0.05 vs. the Bio-NC group; in panel E, correlation between miR-127-3p and C5orf66-AS1 expression was analyzed by Pearson’s correlation analysis, r = − 0.695, p < 0.001
An RNA pull-down assay was then performed to validate the binding relationship between C5orf66-AS1 and miR-127-3p. In CSCs, an enrichment of C5orf66-AS1 sequences was found in the compounds reacted with Bio-miR-127-3p compared to Bio-NC (Fig. 5F). Further, the putative binding sequence between C5orf66-AS1 and miR-127-3p was obtained from Starbase, and the mutant sequence was designed (Fig. 5G) for luciferase assays. Compared to NC mimic, co-transfection of miR-127-3p mimic significantly weakened the luciferase activity of the C5orf66-AS1-WT vector, while the luciferase activity of the C5orf66-AS1-MT vector was not significantly affected (Fig. 5H).
Overexpression of miR-127-3p blocks the inhibitory effects of Exo on HCC-sourced CSCs
To validate if miR-127-3p is the target of the exosomal C5orf66-AS1, the CSCs were transfected with miR-127-3p after Exo treatment. The successful transfection was examined by RT-qPCR, which suggested that the miR-127-3p expression in CSCs was significantly upregulated (Fig. 6A).
Overexpression of miR-127-3p blocks the inhibitory effects of Exo on HCC-sourced CSCs. A transfection efficiency of miR-127-3p mimic in Exo-treated CSCs examined by RT-qPCR (N = 3); B proliferation ability of CSCs after miR-127-3p mimic transfection determined by CCK-8 methods (N = 3); C colony formation ability of CSCs after miR-127-3p mimic transfection examined by colony formation assays (N = 3); D migration ability of CSCs after miR-127-3p mimic transfection determined by wound healing assays (N = 3); E invasion ability of CSCs after miR-127-3p mimic transfection evaluated by Transwell assays (N = 3); F angiogenesis of CSCs after miR-127-3p mimic transfection examined by tube formation assays (N = 3); G self-renewal ability of the CSCs after miR-127-3p mimic transfection examined by LDA (N = 3). Data were expressed as the mean ± SD. Differences were compared by two-way ANOVA (A–G). *p < 0.05 vs. the Exo + NC mimic group
In the setting of miR-127-3p upregulation, it was found that the proliferation ability of CSCs, which was initially suppressed by Exo, was recovered (Fig. 6B). Likewise, the colony formation ability of the CSCs was restored after miR-127-3p overexpression (Fig. 6C). In addition, the wound healing and Transwell assay results indicated the migration and invasion potentials of CSCs were restored after miR-127-3p overexpression (Fig. 6D-E).
Again, the CM of CSCs were collected for tube formation assay. It was found that the length of tubes newly formed by HUVECs and the number of tube branches were restored after miR-127-3p mimic (Fig. 6F). Also, overexpression of miR-127-3p led to a recovery in the self-renewal ability of CSCs which was first weakened by Exo (Fig. 6G).
miR-127-3p targets DUSP1 mRNA
To explore the downstream molecules mediated by miR-127-3p, we then explored the target genes of miR-127-3p using four bioinformatic systems including miRDIP (http://ophid.utoronto.ca/mirDIP/), Starbase, miRWalk (http://mirwalk.umm.uni-heidelberg.de/) and miRTarBase (http://mirtarbase.cuhk.edu.cn/php/search.php). Consequently, two candidate targets were predicted: integrin subunit alpha 6 (ITGA6) and DUSP1 (Fig. 7A). We then explored the expression profiling of ITGA6 (Fig. 7B) and DUSP1 (Fig. 7C) in HCC patients in the Gene Expression Profiling Interactive Analysis (GEPIA) database. It was predicted that DUSP1 was poorly expressed while ITGA6 was highly expressed in HCC patients. Next, we further explored the correlations between miR-127-3p and ITGA6/DUSP1 in the Pan-cancer system in the Starbase system (Fig. 7D). It was suggested that miR-127-3p showed a negative correlation with DUSP1 expression while a positive correlation with ITGA1 in HCC patients. We therefore, surmised that DUSP1 is the target of miR-127-3p in HCC.
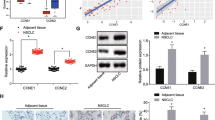
miR-127-3p targets DUSP1 mRNA. A candidate target genes of miR-127-3p predicted in four bioinformatic systems; B–C, expression profiling of ITGA6 (B) and DUSP1 (C) in HCC predicted using the GEPIA database; D correlations between miR-127-3p and ITGA6/DUSP1 predicted according to the data in the Pan-cancer system in the Starbase system; E, F mRNA (E) and protein (F) levels of DUSP1 in THLE-3, Hep3B, HuH7, Hep3B-CSCs and HuH7-CSCs determined by RT-qPCR and western blot analysis, respectively (N = 3); G expression of DUSP1 mRNA in HCC tumor tissues and the adjacent tissues determined by RT-qPCR (N = 3); H a negative correlation between DUSP1 and miR-127-3p in HCC tumor tissues; I putative (WT) and MT DUSP1 mRNA sequences designed for luciferase assays; J binding relationship between miR-127-3p and DUSP1 mRNA validated using a dual-luciferase reporter gene assay (N = 3). Data were expressed as mean ± SD from three independent experiments. Differences were compared by paired t test (G), one-way ANOVA (E and F) or two-way ANOVA (J). *p < 0.05 vs. THLE-3 cells; #p < 0.05 vs. the adjacent tissues; &p < 0.01 vs. the NC mimic group; in panel H, correlation between miR-127-3p and DUSP1 expression was analyzed by Pearson’s correlation analysis, r = − 0.761, p < 0.001
Thereafter, we examined the DUSP1 expression in HCC cells. The RT-qPCR and western blot assay results indicated that the levels of DUSP1 were significantly declined in HCC cells and further reduced in the corresponding CSCs (Fig. 7E, F). Moreover, the expression of DUSP1 was lower in HCC tissues than that in the adjacent tissues (Fig. 7G), which showed a negative correlation with miR-127-3p expression (Fig. 7H).
Likewise, the binding between miR-127-3p and DUSP1 mRNA was validated using a luciferase assay. The putative binding site was obtained from the Starbase system, and the MT sequence was designed for luciferase assays (Fig. 7I). It was found that the luciferase activity of the DUSP1-WT vector was reduced by miR-127-3p, while the activity of the DUSP1-MT luciferase vector was not affected (Fig. 7J).
Silencing of DUSP1 activates ERK phosphorylation to block the inhibitory effect of Exo on CSCs
DUSP1 has been reported to inhibit phosphorylation of ERK to block tumorigenesis of HCC [17]. Then, we examined the protein level of DUSP1 and phosphorylation of ERK in CSCs after Exo treatment or miR-127-3p mimic transfection (Fig. 8). It was found that the level of DUSP1 was elevated, while phosphorylation of ERK was reduced after Exo treatment. However, inverse changes were induced by miR-127-3p mimic (Fig. 8A). Thereafter, the Exo-treated CSCs were transfected with si-DUSP1 to examine if DUSP1 was the functional target of the exosomal C5orf66-AS1. The DUSP1 level, according to western blot analysis, was significantly reduced by si-DUSP1, while the phosphorylation of ERK was enhanced (Fig. 8B).
Silencing of DUSP1 activates ERK phosphorylation to block the inhibitory effect of Exo on CSCs. A protein level of DUSP1 and phosphorylation of ERK in CSCs after Exo treatment or miR-127-3p mimic transfection examined by western blot analysis (N = 3); B protein level of DUSP1 and phosphorylation of ERK in CSCs after si-DUSP1 transfection determined by western blot analysis (N = 3); C proliferation ability of CSCs after si-DUSP1 transfection determined by CCK-8 methods (N = 3); D colony formation ability of CSCs after si-DUSP1 transfection examined by colony formation assays (N = 3); E migration ability of CSCs after si-DUSP1 transfection evaluated by wound healing assays; F invasion ability of CSCs after si-DUSP1 transfection determined by Transwell assays (N = 3); G angiogenesis of CSCs after si-DUSP1 transfection detected by tube formation assays (N = 3); H self-renewal ability of the CSCs after si-DUSP1 transfection evaluated by LDA (N = 3). Data were expressed as the mean ± SD. Differences were compared by two-way ANOVA (A–H). *p < 0.05 vs. the control group; #p < 0.05 vs. the Exo + NC mimic group; &p < 0.05 vs. the Exo + si-NC group
MSC-derived exosomal C5orf66-AS1 enhances DUSP1 expression and suppresses ERK phosphorylation to inhibit tumorigenesis of HCC in vivo. A weekly change in the volume of xenograft tumors (number of mice in each group; n = 8); B weight of the xenograft tumors on the 4th week (n = 8); C mRNA expression of DUSP1 in tumor tissues examined by RT-qPCR; D phosphorylation of ERK and protein level of Ki67 in the tissues examined by IHC staining (n = 8). Data were expressed as mean ± SD from three independent experiments. Differences were compared by one-way ANOVA (B and C) or two-way ANOVA (A and D). *p < 0.05 vs. the control group; #p < 0.05 vs. the Exo group
The relevance of DUSP1 to the malignant behaviors of CSCs was examined. Importantly, the proliferation ability of CSCs suppressed by Exo was restored after DUSP1 inhibition (Fig. 8C). Still, silencing of DUSP1 restored the colony formation ability of the CSCs (Fig. 8D). The migration and invasion abilities of the CSCs, according to the wound healing and Transwell assays, were recovered in the setting of DUSP1 downregulation (Fig. 8E-F). In addition, the tube formation ability of the HUVECs cultured in the CM of CSCs transfected with si-DUSP1 was significantly enhanced (Fig. 8G). The self-renewal ability of CSCs weakened by Exo was recovered upon DUSP1 silencing as well (Fig. 8H).
MSC-derived exosomal C5orf66-AS1 enhances DUSP1 expression and suppresses ERK phosphorylation to inhibit tumorigenesis of HCC in vivo
The functions of Exo and the potentially involved molecules in vivo were further explored. The CSCs after Exo treatment, miR-127-3p mimic or si-DUSP1 transfection were injected into nude mice for animal studies. To avoid unnecessary animal kiling, only Hep3B-CSCs were used in animal experiments.
After cell injection, the tumor volume was examined. Importantly, compared to the control cells, Exo treatment in cells reduced growth rate and the weight of xenograft tumors; however, the volume and weight of the xenograft tumors were recovered when CSCs were further transfected with miR-127-3p mimic or si-DUSP1 (Fig. 9A, B).
The tissues of the xenograft tumors were collected for RT-qPCR. Exo treatment in CSCs increased the expression of C5orf66-AS1 and DUSP1 mRNA, whereas it reduced the expression of miR-127-3p in the tumor tissues. Further transfection of miR-127-3p in CSCs, as expected, restored the expression of miR-127-3p, and it reduced the expression of DUSP1 mRNA in tissues, whereas si-DUSP1 reduced DUSP1 mRNA in tissues as well, though it has no impact on the expression of C5orf66-AS1 of miR-127-3p (Fig. 9C). The tissues were collected for IHC staining. The phosphorylation of ERK and protein level of Ki67 in the tissues were reduced by Exo treatment while restored after further miR-127-3p mimic or si-DUSP1 delivery (Fig. 9D).
Discussion
Emerging evidence has suggested that exosomes from MSCs play significant, either oncogenic or tumor-suppressing roles in tumorigenesis, with the involvement of their carried molecular cargoes [11, 18]. CSCs have been deemed as the root of cancers and major regulators for treatment resistance and cancer recurrence. The relevance of the MSC-derived exosomes to stemness property of CSCs, however, has been hardly concerned. This study reports that MSC-derived exosomal C5orf66-AS1 suppresses ERK phosphorylation through a miR-127-3p/DUSP1 axis and inhibits malignant behaviors of CSCs from HCC cells.
MSC therapy has been discussed as an attractive therapeutic option for liver fibrosis and cirrhosis [19]. However, safety concerns on MSC injection, including uncontrolled cell replication and vascular occlusion, have been demonstrated [20]. In addition, the beneficial effects of MSCs can be transient, and the secreted soluble paracrine factors of MSCs including exosomes have been increasingly concerned as alternative tools since they have less disadvantages while recapitulating the biological activity of MSCs [21, 22]. The application of exosomes may have significant advantages over their cellular counterparts owing to a higher safety profile, lower immunogenicity and the incapability of directly forming tumors [23]. Treatment of MSC-derived exosomes has been reported as a novel therapeutic strategy for liver diseases including HCC [24]. Importantly, in this study, the Exo were extracted, identified and administrated into CSCs from HCC cells. The Exo reduced viability of CSCs in a dose-dependent manner and decreased proliferation, colony formation, migration and invasion, angiogenesis-stimulating and self-renewal abilities of the CSCs. Exosomes from BM-MSCs have been found to inhibit proliferation and transition of a human breast cancer cell line BM2 and reduce the abundance of stem cell-like surface markers [25]. The MSC-derived exosomes have also shown anti-angiogenetic role in breast cancer cells via downregulating the expression of vascular endothelial growth factor [26]. Moreover, in a study by Saurabh et al., treatment of encapsulated MSCs reduced viability and migration and conferred an anti-angiogenic role in breast cancer stem cells [27]. More relevantly, exosomes from adipose tissue-derived MSCs have been found to enhance the chemosensitivity of HCC cells [28, 29]. Quite in concert with these findings, our experiments confirmed that the Exo reduced the stemness property and the malignant behaviors of the HCC-sourced CSCs, which validated that MSCs and Exo might serve as potential therapeutic tools for HCC treatment.
As exosomes play a major role in intercellular communication and exert their functions via the carried genetic factors, we then explored the molecules involved in the events above. A lncRNA microarray analysis was performed, and C5orf66-AS1 was screened as the most upregulated lncRNA in CSCs after Exo treatment. C5orf66-AS1 has been reported as a tumor-suppressing gene in gastric cancer [30], oral squamous cell carcinoma [31] and pituitary null cell adenomas [32] by limiting proliferation and aggressiveness of cancer cells. In this paper, we confirmed that C5orf66-AS1 was significantly poorly expressed in HCC cells and tissues compared to the healthy ones. Low expression of C5orf66-AS1 in patients was linked to increased tumor size, aggravated cirrhosis and advanced BCLC stage. These results indicated a similar anti-oncogenic role of C5orf66-AS1 in HCC. In addition, further lower expression of C5orf66-AS1 in CSCs than that in HCCs suggested a possible correlation between poor C5orf66-AS1 expression and the stemness of CSCs.
The subsequent bioinformatic analyses and cellular experiments suggested miR-127-3p as a target miRNA of C5orf66-AS1, and DUSP1 mRNA was determined as a target of miR-127-3p. The binding relationships were validated via luciferase assays. miR-127-3p has been reported as either an oncogene [33] or a tumor suppressor [34] in different cancer types and the different target genes. Although the direct impact of miR-127-3p on HCC has not been concerned, a study by Livingstone MC et al. found that miR-127-3p was significantly upregulated in liver tissues treated with aflatoxin B, a potent human and animal hepatocarcinogen [35]. Here, we confirmed a high expression profiling of miR-127-3p in HCC tissues and cells, especially the CSC cells. On the contrary, DUSP1 mRNA was poorly expressed in the HCC tissues and cells. This was partially in line with a previous study which suggested that DUSP1 was downregulated in HCC and its downregulation was correlated with poor prognosis of patients [36]. Importantly, either upregulation of miR-127-3p or downregulation of DUSP1 in CSCs blocked the anti-tumor role of the Exo, indicating that the exosomal C5orf66-AS1 possibly enhanced DUSP1 expression through sequestering miR-127-3p to reduce the stemness of CSCs. Intriguingly, DUSP1 has been documented to inhibit phosphorylation of ERK to inhibit progression of HCC [17]. The ERK signaling is one of the mitogen-activated kinase-like protein (MAPK) signaling pathways whose aberrant expression is frequently involved in cancer progression via mediating cellular proliferation, differentiation and survival [37, 38]. In addition, increased phosphorylation of ERK has been demonstrated to enhance stemness of CSCs [39, 40]. This is also true for HCC since ERK phosphorylation was helpful for the maintenance of stemness of HCC-CSCs [41, 42]. Here, we confirmed that phosphorylation of ERK was reduced by Exo treatment but it was restored after DUSP1 inhibition, suggesting that ERK inhibition is, at least one of the effectors, of the reduced stem cell property of HCC-CSCs mediated by Exo and the C5orf66-AS1-miR-127-3p-DUSP1 axis.
To conclude, this study evidenced that Exo block malignant behaviors of CSCs from HCC cells through a C5orf66-AS1/miR-127-3p/DUSP1/ERK axis. Several clinical trials of MSCs in cancer therapy have been conducted [10]. This study may offer novel insights into the relevance of MSCs and their derived molecules to cancer progression, and in particular, to the stem cell property of CSCs.
Data availability
The analyzed data sets generated during the study are available from the corresponding author on reasonable request.
Abbreviations
- ANOVA:
-
Analysis of variance
- ATCC:
-
American type culture collection
- BCA:
-
Bicinchoninic acid
- BCLC:
-
Barcelona clinic liver cancer
- BM-MSCs:
-
Bone marrow-derived mesenchymal stem cells
- CCK-8:
-
Cell counting kit-8
- CSCs:
-
Cancer stem cells
- CM:
-
Conditioned media
- DMEM:
-
Dulbecco’s modified Eagle’s medium
- DUSP1:
-
Dual-specificity phosphatase 1
- Exo:
-
Exosomes
- FBS:
-
Fetal bovine serum
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- GEPIA:
-
Gene expression profiling interactive analysis
- HCC:
-
Hepatocellular carcinoma
- HUVECs:
-
Human umbilical vein endothelial cells
- IgG:
-
Immunoglobulin G
- IHC:
-
Immunohistochemical staining
- LDA:
-
Limited-dilution assay
- lncRNA:
-
Long noncoding RNA
- mean ± SD:
-
Mean ± standard deviation
- MT:
-
Mutant type
- MTT:
-
3-(4, 5-Dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide
- NC:
-
Negative control
- NIH:
-
National Institutes of Health
- NTA:
-
Nanoparticle tracking analysis
- OD:
-
Optical density
- PBS:
-
Phosphate-buffered saline
- PFA:
-
Paraformaldehyde
- RIPA:
-
Radio-immunoprecipitation assay
- RT-qPCR:
-
Reverse transcription quantitative polymerase chain reaction
- TEM:
-
Transmission electron microscope
- WT:
-
Wild type
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. https://doi.org/10.3322/caac.21492.
Karadag Soylu N. Update on hepatocellular carcinoma: a brief review from pathologist standpoint. J Gastrointest Cancer. 2020;51(4):1176–86. https://doi.org/10.1007/s12029-020-00499-5.
Fu J, Wang H. Precision diagnosis and treatment of liver cancer in China. Cancer Lett. 2018;412:283–8. https://doi.org/10.1016/j.canlet.2017.10.008.
Zhang X, Xu X, Ge G, Zang X, Shao M, Zou S, Zhang Y, Mao Z, Zhang J, Mao F, Qian H, Xu W. miR498 inhibits the growth and metastasis of liver cancer by targeting ZEB2. Oncol Rep. 2019;41(3):1638–48. https://doi.org/10.3892/or.2018.6948.
Liu G, Luo Q, Li H, Liu Q, Ju Y, Song G. Increased oxidative phosphorylation is required for stemness maintenance in liver cancer stem cells from hepatocellular carcinoma cell line HCCLM3 cells. Int J Mol Sci. 2020. https://doi.org/10.3390/ijms21155276.
Dean M, Fojo T, Bates S. Tumour stem cells and drug resistance. Nat Rev Cancer. 2005;5(4):275–84. https://doi.org/10.1038/nrc1590.
Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, Qian D, Lam JS, Ailles LE, Wong M, Joshua B, Kaplan MJ, Wapnir I, Dirbas FM, Somlo G, Garberoglio C, Paz B, Shen J, Lau SK, Quake SR, Brown JM, Weissman IL, Clarke MF. Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature. 2009;458(7239):780–3. https://doi.org/10.1038/nature07733.
Mendt M, Rezvani K, Shpall E. Mesenchymal stem cell-derived exosomes for clinical use. Bone Marrow Transplant. 2019;54(Suppl 2):789–92. https://doi.org/10.1038/s41409-019-0616-z.
Hass R, Kasper C, Bohm S, Jacobs R. Different populations and sources of human mesenchymal stem cells (MSC): a comparison of adult and neonatal tissue-derived MSC. Cell Commun Signal. 2011;9:12. https://doi.org/10.1186/1478-811X-9-12.
Lin W, Huang L, Li Y, Fang B, Li G, Chen L, Xu L. Mesenchymal stem cells and cancer: clinical challenges and opportunities. Biomed Res Int. 2019;2019:2820853. https://doi.org/10.1155/2019/2820853.
Vakhshiteh F, Atyabi F, Ostad SN. Mesenchymal stem cell exosomes: a two-edged sword in cancer therapy. Int J Nanomedicine. 2019;14:2847–59. https://doi.org/10.2147/IJN.S200036.
Cao X, Xue LD, Di Y, Li T, Tian YJ, Song Y. MSC-derived exosomal lncRNA SNHG7 suppresses endothelial-mesenchymal transition and tube formation in diabetic retinopathy via miR-34a-5p/XBP1 axis. Life Sci. 2021;272: 119232. https://doi.org/10.1016/j.lfs.2021.119232.
Lv H, Lv G, Han Q, Yang W, Wang H. Noncoding RNAs in liver cancer stem cells: the big impact of little things. Cancer Lett. 2018;418:51–63. https://doi.org/10.1016/j.canlet.2018.01.001.
Liu AY, Cai Y, Mao Y, Lin Y, Zheng H, Wu T, Huang Y, Fang X, Lin S, Feng Q, Huang Z, Yang T, Luo Q, Ouyang G. Twist2 promotes self-renewal of liver cancer stem-like cells by regulating CD24. Carcinogenesis. 2014;35(3):537–45. https://doi.org/10.1093/carcin/bgt364.
Yang ZF, Ho DW, Ng MN, Lau CK, Yu WC, Ngai P, Chu PW, Lam CT, Poon RT, Fan ST. Significance of CD90+ cancer stem cells in human liver cancer. Cancer Cell. 2008;13(2):153–66. https://doi.org/10.1016/j.ccr.2008.01.013.
Ran RZ, Chen J, Cui LJ, Lin XL, Fan MM, Cong ZZ, Zhang H, Tan WF, Zhang GQ, Zhang YJ. miR-194 inhibits liver cancer stem cell expansion by regulating RAC1 pathway. Exp Cell Res. 2019;378(1):66–75. https://doi.org/10.1016/j.yexcr.2019.03.007.
Liao B, Zhou H, Liang H, Li C. Regulation of ERK and AKT pathways by hepatitis B virus X protein via the Notch1 pathway in hepatocellular carcinoma. Int J Oncol. 2017;51(5):1449–59. https://doi.org/10.3892/ijo.2017.4126.
de Araujo FV, O’Valle F, Serrano-Saenz S, Anderson P, Andres E, Lopez-Penalver J, Tovar I, Nieto A, Santos A, Martin F, Exposito J, Oliver FJ, de Almodovar JMR. Exosomes derived from mesenchymal stem cells enhance radiotherapy-induced cell death in tumor and metastatic tumor foci. Mol Cancer. 2018;17(1):122. https://doi.org/10.1186/s12943-018-0867-0.
Cao Y, Ji C, Lu L. Mesenchymal stem cell therapy for liver fibrosis/cirrhosis. Ann Transl Med. 2020;8(8):562. https://doi.org/10.21037/atm.2020.02.119.
Du YM, Zhuansun YX, Chen R, Lin L, Lin Y, Li JG. Mesenchymal stem cell exosomes promote immunosuppression of regulatory T cells in asthma. Exp Cell Res. 2018;363(1):114–20. https://doi.org/10.1016/j.yexcr.2017.12.021.
Bagno L, Hatzistergos KE, Balkan W, Hare JM. Mesenchymal stem cell-based therapy for cardiovascular disease: progress and challenges. Mol Ther. 2018;26(7):1610–23. https://doi.org/10.1016/j.ymthe.2018.05.009.
Borger V, Bremer M, Ferrer-Tur R, Gockeln L, Stambouli O, Becic A, Giebel B. Mesenchymal stem/stromal cell-derived extracellular vesicles and their potential as novel immunomodulatory therapeutic agents. Int J Mol Sci. 2017. https://doi.org/10.3390/ijms18071450.
Liew LC, Katsuda T, Gailhouste L, Nakagama H, Ochiya T. Mesenchymal stem cell-derived extracellular vesicles: a glimmer of hope in treating Alzheimer’s disease. Int Immunol. 2017;29(1):11–9. https://doi.org/10.1093/intimm/dxx002.
Lou G, Chen Z, Zheng M, Liu Y. Mesenchymal stem cell-derived exosomes as a new therapeutic strategy for liver diseases. Exp Mol Med. 2017;49(6): e346. https://doi.org/10.1038/emm.2017.63.
Ono M, Kosaka N, Tominaga N, Yoshioka Y, Takeshita F, Takahashi RU, Yoshida M, Tsuda H, Tamura K, Ochiya T. Exosomes from bone marrow mesenchymal stem cells contain a microRNA that promotes dormancy in metastatic breast cancer cells. Sci Signal. 2014;7(332):ra63. https://doi.org/10.1126/scisignal.2005231.
Lee JK, Park SR, Jung BK, Jeon YK, Lee YS, Kim MK, Kim YG, Jang JY, Kim CW. Exosomes derived from mesenchymal stem cells suppress angiogenesis by down-regulating VEGF expression in breast cancer cells. PLoS ONE. 2013;8(12): e84256. https://doi.org/10.1371/journal.pone.0084256.
Mandal S, Arfuso F, Sethi G, Dharmarajan A, Warrier S. Encapsulated human mesenchymal stem cells (eMSCs) as a novel anti-cancer agent targeting breast cancer stem cells: development of 3D primed therapeutic MSCs. Int J Biochem Cell Biol. 2019;110:59–69. https://doi.org/10.1016/j.biocel.2019.02.001.
Lou G, Chen L, Xia C, Wang W, Qi J, Li A, Zhao L, Chen Z, Zheng M, Liu Y. MiR-199a-modified exosomes from adipose tissue-derived mesenchymal stem cells improve hepatocellular carcinoma chemosensitivity through mTOR pathway. J Exp Clin Cancer Res. 2020;39(1):4. https://doi.org/10.1186/s13046-019-1512-5.
Lou G, Song X, Yang F, Wu S, Wang J, Chen Z, Liu Y. Exosomes derived from miR-122-modified adipose tissue-derived MSCs increase chemosensitivity of hepatocellular carcinoma. J Hematol Oncol. 2015;8:122. https://doi.org/10.1186/s13045-015-0220-7.
Zhou Q, Li H, Jing J, Yuan Y, Sun L. Evaluation of C5orf66-AS1 as a potential biomarker for predicting early gastric cancer and its role in gastric carcinogenesis. Onco Targets Ther. 2020;13:2795–805. https://doi.org/10.2147/OTT.S239965.
Lu T, Liu H, You G. Long non-coding RNA C5orf66-AS1 prevents oral squamous cell carcinoma through inhibiting cell growth and metastasis. Int J Mol Med. 2018;42(6):3291–9. https://doi.org/10.3892/ijmm.2018.3913.
Yu G, Li C, Xie W, Wang Z, Gao H, Cao L, Hao L, Zhang Y. Long non-coding RNA C5orf66-AS1 is downregulated in pituitary null cell adenomas and is associated with their invasiveness. Oncol Rep. 2017;38(2):1140–8. https://doi.org/10.3892/or.2017.5739.
Jiang H, Hua D, Zhang J, Lan Q, Huang Q, Yoon JG, Han X, Li L, Foltz G, Zheng S, Lin B. MicroRNA-127-3p promotes glioblastoma cell migration and invasion by targeting the tumor-suppressor gene SEPT7. Oncol Rep. 2014;31(5):2261–9. https://doi.org/10.3892/or.2014.3055.
Ji L, Zhu ZN, He CJ, Shen X. MiR-127-3p targets KIF3B to inhibit the development of oral squamous cell carcinoma. Eur Rev Med Pharmacol Sci. 2019;23(2):630–40. https://doi.org/10.26355/eurrev_201901_16877.
Livingstone MC, Johnson NM, Roebuck BD, Kensler TW, Groopman JD. Profound changes in miRNA expression during cancer initiation by aflatoxin B1 and their abrogation by the chemopreventive triterpenoid CDDO-Im. Mol Carcinog. 2017;56(11):2382–90. https://doi.org/10.1002/mc.22635.
Hao PP, Li H, Lee MJ, Wang YP, Kim JH, Yu GR, Lee SY, Leem SH, Jang KY, Kim DG. Disruption of a regulatory loop between DUSP1 and p53 contributes to hepatocellular carcinoma development and progression. J Hepatol. 2015;62(6):1278–86. https://doi.org/10.1016/j.jhep.2014.12.033.
Dhillon AS, Hagan S, Rath O, Kolch W. MAP kinase signalling pathways in cancer. Oncogene. 2007;26(22):3279–90. https://doi.org/10.1038/sj.onc.1210421.
Samatar AA, Poulikakos PI. Targeting RAS-ERK signalling in cancer: promises and challenges. Nat Rev Drug Discov. 2014;13(12):928–42. https://doi.org/10.1038/nrd4281.
Jang HJ, Bak Y, Pham TH, Kwon SB, Kim BY, Hong J, Yoon DY. STK899704 inhibits stemness of cancer stem cells and migration via the FAK-MEK-ERK pathway in HT29 cells. BMB Rep. 2018;51(11):596–601.
Wei F, Zhang T, Deng SC, Wei JC, Yang P, Wang Q, Chen ZP, Li WL, Chen HC, Hu H, Cao J. PD-L1 promotes colorectal cancer stem cell expansion by activating HMGA1-dependent signaling pathways. Cancer Lett. 2019;450:1–13. https://doi.org/10.1016/j.canlet.2019.02.022.
Guo J, Guo M, Zheng J. Inhibition of bone morphogenetic Protein 2 suppresses the stemness maintenance of cancer stem cells in hepatocellular carcinoma via the MAPK/ERK pathway. Cancer Manag Res. 2021;13:773–85. https://doi.org/10.2147/CMAR.S281969.
Tsai CF, Hsieh TH, Lee JN, Hsu CY, Wang YC, Kuo KK, Wu HL, Chiu CC, Tsai EM, Kuo PL. Curcumin suppresses phthalate-induced metastasis and the proportion of cancer stem cell (CSC)-like cells via the inhibition of AhR/ERK/SK1 signaling in hepatocellular carcinoma. J Agric Food Chem. 2015;63(48):10388–98. https://doi.org/10.1021/acs.jafc.5b04415.
Funding
This work was supported by Natural Science Foundation of Xinjiang Uygur Autonomous Region (No. 2017D01C306).
Author information
Authors and Affiliations
Contributions
HG and CY: conceptualization, methodology, software, and writing—original draft preparation; LW: data curation; JJL: visualization and investigation; ZQZ: software and validation; DZG and HJW: conceptualization, writing—reviewing & editing, and writing—original draft preparation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
Signed informed consent was acquired from each participant. The research was approved by the Ethical Committee of the First Affiliated Hospital of Xinjiang Medical University (Approval number: K201807-05) and performed according to the Declaration of Helsinki.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gu, H., Yan, C., Wan, H. et al. Mesenchymal stem cell-derived exosomes block malignant behaviors of hepatocellular carcinoma stem cells through a lncRNA C5orf66-AS1/microRNA-127-3p/DUSP1/ERK axis. Human Cell 34, 1812–1829 (2021). https://doi.org/10.1007/s13577-021-00599-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-021-00599-9