Abstract
Background
The field of liquid biopsies in oncology is rapidly expanding, with the application of cell-free circulating tumour DNA (ctDNA) showing promise in this era of precision medicine. Compared with traditional clinical and radiographic tumour monitoring methods, the analysis of ctDNA provides a minimally-invasive and technically feasible approach to assess temporal and spatial molecular evolutions of the tumour landscape. The constantly advancing technological platforms available for ctDNA extraction and analysis allow greater analytical sensitivities than ever before. The potential translational impact of ctDNA as a blood-based biomarker for the identification, characterization and monitoring of cancer has been demonstrated in numerous proof-of-concept studies, with ctDNA analysis beginning to be applied clinically across multiple facets of oncology.
Conclusions
In this review we discuss the biology, recent advancements, technical considerations and clinical implications of ctDNA in the context of cancer, and highlight important challenges and future directions for the integration of ctDNA into standardised patient care.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
It is now well-accepted that cancer is fundamentally a genetic disease, where the sequential accumulation of somatic mutations and epigenetic changes disrupts the delicate homeostasis of cellular functioning. Advances in technological platforms such as next-generation sequencing (NGS) have allowed in-depth interrogations of such genetic aberrations and have, subsequently, led to significant advancements in our understanding of the key regulators of oncogenesis. As a consequence, we have entered the era of ‘precision medicine’, whereby therapeutic strategies may be tailored to patients based on their tumour molecular landscapes. With this comes the necessity to collect tumour samples for genomic analysis. Tissue biopsy remains the current gold-standard method for sampling a patient’s tumour for genotypic analysis [1, 2]. There are, however, multiple shortcomings to this method. Such invasive and costly sampling methods are associated with a high degree of complications. For example, 17% of lung cancer patients undergoing intra-thoracic image-guided biopsy suffer from an adverse event [3]. Additionally, owing to spatial and temporal tumour heterogeneity, a single-site biopsy may inadequately represent the total genetic landscape, and therapeutically actionable mutations or potentially resistant clones may thus be missed [1]. There is also a risk that a biopsy fails to obtain a sufficient number of cancerous cells to carry out adequate downstream analyses [4]. Moreover, current tissue preservation methods such as formalin fixation can introduce significant artefacts that may confound molecular testing [5]. In solution to these drawbacks, minimally-invasive biopsy methods have risen in popularity over recent years alongside the rise in molecular tumour profiling. Termed ‘liquid biopsies’, these methods include collecting blood or other bodily fluids from which to isolate various circulating components. In particular, cell-free DNA (cfDNA) holds promise as a biopsy surrogate, and has the potential to become a rapid, cost-effective and non-invasive means to assess the tumour landscape (Table 1).
2 Cell-free DNA biology
Cell-free DNA, double-stranded, highly fragmented extracellular DNA was first described in blood plasma in 1948 [25]. Subsequently, it has been detected in urine, saliva, peritoneal fluid, pleural fluid, uterine lavage fluid and cerebrospinal fluid [26,27,28,29,30], where it is presumably released from cells via both active secretion and apoptosis [31, 32]. Healthy individuals harbour between 1 and 10 nanograms of cfDNA in each millilitre of blood, with the major contributors being hematopoietic cells [30, 33,34,35]. Further studies are, however, required in order to definitively determine the threshold of ‘normal’ cfDNA concentrations. Whilst cfDNA has a short half-life of approximately 16 minutes, associations with various lipids and proteins can increase its longevity up to 10-fold [30, 36, 37]. cfDNA clearance from the body primarily involves nuclease action, followed by renal excretion [37, 38].
2.1 Biology and origin of tumour-derived cell-free DNA
Whilst a number of non-malignant causes of elevated cfDNA exist due to increases in cellular death and turnover (infection, autoimmune disease, trauma), it wasn’t until 1977 that the first connection between cfDNA and cancer was uncovered, when it was noted that circulating levels were increased in various types of cancers [39, 40]. This initial study by Leon et al. also provided a first indication that cfDNA levels may be proportional to tumour staging [41]. The presence of tumour-specific DNA among the pool of extracellular nucleic acids in the circulation was not confirmed until sensitive, mutant-specific PCR technologies were introduced nearly 15 years later [40, 42]. These early observations hinted at the possibility of using cfDNA to analyse tumour genomes in a minimally-invasive manner. Further technological advances such as the advent of next-generation sequencing (NGS) have brought these initial ideas to fruition.
In a malignant state, levels of cfDNA can increase to 50-1000 ng/ml blood, with tumour DNA comprising 3-90% of the total [43]. Increased levels are presumably due to a high turnover rate of tumour cells and an inefficient clearing of dead and dying cells [40]. The principal mechanisms through which tumour-derived cfDNA (herein referred to as ctDNA) is released in both solid and haematological malignancies are still under dispute, but multiple properties suggest cell death as the major contributor. The most common size of double-stranded ctDNA fragments is 160-180bp, as determined by sequencing-based approaches in humans and xenograft mouse models [30, 44, 45]. This size corresponds to the length of DNA occupying a nucleosome, the structural units of the eukaryotic chromosome [40]. Additionally, prominent but smaller fragment populations of ~330 and ~500 bp can be seen, corresponding to a linear progression of di- and tri-nucleosomal fragments [46] (Fig. 1). These observations implicate caspase-activated DNase, an endonuclease active in cellular apoptosis which cleaves chromatin DNA in an inter-nucleosomal pattern [46]. Another suggested method of ctDNA release is active secretion, in which newly-synthesised fragments are released in a lipid-protein complex [47]. This homeostatic mechanism, which was demonstrated early on by Abolhassani et al., can also produce DNA fragments of comparable lengths to those arising from apoptosis [48, 49]. Some studies have suggested that a large proportion of ctDNA is derived from active cellular secretion, such as the one published by Bronkhorst et al. who identified ctDNA fragments of novel length predominantly being released by viable, non-apoptotic osteosarcoma cells from [50]. There is also a case for necrotic release of tumour DNA, especially when considering the hypoxic environment of solid lesions. Necrotic cell death leads to the release of rapidly-cleaved, predominantly large (> 10,000 bp) nucleic acid fragments [43, 51]. A recent study using an in vivo inducible necrotic liver injury model demonstrated that large DNA fragments greater than 10,000 bp can be identified in the circulation which can significantly contribute to the pool of cfDNA fragments [43]. Current extraction methods, however, poorly recover larger ctDNA fragments and, therefore, the contribution of tumour cell necrosis to the ctDNA population is still incompletely understood [34, 52]. It is currently accepted that a combination of all three mechanisms may lead to ctDNA deposition, although further research is required to determine the relative contribution each makes to the total ctDNA levels in the context of different fluid compartments. Such knowledge may inform choices for cfDNA extraction methods in order to enrich for ctDNA.
3 Technical considerations for ctDNA isolation and analysis
There are a number of factors that need to be considered when isolating patient ctDNA for analysis, and a thorough investigation of each may assist in the selection of standardized operating procedures to help control for preanalytical variables common during cfDNA processing. Whilst ctDNA has been identified in multiple fluid compartments, the most common source for analysis is blood (Box 1).
3.1 Plasma versus serum
ctDNA is, by nature, diluted in cfDNA present due to normal cellular turnover. Therefore, further dilution of the ctDNA by inadvertent lysis of leukocytes and other cells after sample collection is undesirable, as it can severely hinder the ability to detect low frequency mutations [58]. Past studies have favoured ctDNA collection from serum, as the total cfDNA yield is consistently greater compared to that of plasma [59, 60]. However, in recent years it has been demonstrated that this difference is due to the higher level of leukocyte lysis during blood coagulation [60]. For this reason, plasma has become the desired medium from which ctDNA may be extracted, and has since been shown to contain higher proportions of tumour-derived nucleic acids [45].
3.2 Collection tube and anti-coagulant
Between venepuncture and sample processing, the concentration of cfDNA will increase over time due to cellular lysis [61, 62]. To minimise genomic DNA contamination of ctDNA, prompt separation of plasma from the cellular components of blood via centrifugation is essential. The maximum time that a sample can be left unprocessed before it becomes compromised is still under debate, and ranges from 2 to 24 hours [63,64,65,66,67]. The type of blood collection tube should be taken into consideration when assessing the available infrastructure, or lack thereof, that may form a barrier to timely plasma isolation after venepuncture. The most common plasma collection tubes are ethylenediaminetetraacetic acid (EDTA) and cell-free DNA™ tubes (Streck), both of which contain the K3EDTA anti-coagulant, with the latter containing an additional preservative to reduce cellular DNA release [68]. Comparative studies in the fields of obstetrics and oncology have demonstrated the equivalence of both collection tube types if plasma isolation occurs within 6 hours of venepuncture [69]. Beyond this, cell-free DNA™ tubes outperform conventional EDTA tubes at preventing significant genomic DNA contamination [69,70,71], and have been shown to preserve the quality of cfDNA for five days at room temperature [72]. Other blood collection tubes include those containing citrate or heparin anti-coagulants, but K3EDTA surmounts both when considering cfDNA preservation and downstream analysis [65, 73].
3.3 Extraction and storage
Circulating DNA levels can be extremely variable and, therefore, all possible measures should be taken to maximise yield and minimise degradation in order to allow for sensitive detection of mutant alleles [74, 75]. Consequently, careful selection of a nucleic acid isolation kit suitable for low cfDNA is critical. There are currently numerous kits principally designed for the purpose of extracting cfDNA (Table 2), each with is own merits and limitations. After extraction, cfDNA can be stored between -20 and -80°C for extended periods of time, but multiple freeze-thaw cycles will invariably cause cfDNA fragmentation and should be avoided unless the DNA is stored in a low-salt elution buffer [61, 66]. Additionally, due to the hydrophilic nature of double-stranded helical DNA, use of plastic-ware containing hydrophobic polymers such as polypropylene can lead to reduced yields [61]. In contrast, polyallomer or specially-treated polypropylene plastic-ware has been shown to be more suitable for DNA storage and for maximising nucleic acid recovery during pre-analytical stages [84, 85].
3.4 Analysis platform
Similar to solid tumour biopsy analysis, the genomic interrogation of ctDNA ranges from single point mutation to whole-genome investigation. At present, there are numerous methods used to interrogate ctDNA, which can broadly be separated into candidate-gene approaches (for assessing 1-10 loci) and deep-sequencing approaches (Table 3). Whilst deep-sequencing methods provide the ability to assess a high number of loci without any prior knowledge of molecular alterations, and also affords a good platform for discovery of novel genes for precision medicine, it is expensive and requires comprehensive informatics analysis. Due to the low complexity and highly-damaged nature of ctDNA, whole genome studies are relatively scarce, although low-depth whole-genome sequencing has been used to identify somatic copy-number alterations in prostate cancer and neuroblastoma [86, 87]. As such, targeted sequencing approaches have become the preferred method when analysing ctDNA via deep sequencing [88, 89]. Candidate gene analysis approaches such as digital PCR have a comparable or increased sensitivity, are more cost-effective, and no bioinformatic analysis is required. However, such approaches only enable monitoring of known mutations, and multiplexing capabilities are low. It is likely that high-throughput, deep-sequencing methodologies will continue to be used for research purposes, and resulting data should inform a candidate-gene analysis approach that can easily be implemented in clinical practice where specific genomic expertise may be lacking. An assessment of the merits and limitations of current technologies used for ctDNA analysis is provided in Table 3.
3.4.1 Increasing analytical sensitivity
At present, both workflow approaches are lacking in the sensitivity required for ctDNA analysis to become imbedded in clinical practice. False-positives still occur [90], and it is suspected that many false-negatives arise due to insufficient detection of ctDNA in the larger pool of cfDNA, especially in the case of primary, low burden tumours. The addition of unique molecular identifiers (UMIs) in next generation sequencing approaches such as Safe-SeqS are expected to increase the sensitivity of deep-sequencing approaches for ctDNA detection. This molecular barcoding technique assigns unique sequences to each original DNA fragment, allowing the formation of UMI families following sequencing and the generation of corrected consensus reads without PCR or sequencing errors [91]. This allows utilisation of PCR replicates to reduce background errors and, therefore, increase true variant detection at lower allelic frequencies, which is useful when trying to identify ctDNA in a larger pool of normal cfDNA. Likewise, CAPP-seq uses a similar method, but each strand of the DNA template molecules is assigned a unique identifier allowing even pre-PCR errors to be removed, which further increases the sensitivity of the assay [92, 93]. Utilisation of targeted sequencing over whole genome or whole exome sequencing will also improve the sensitivity of NGS assays, as it allows increased coverage of specific regions of interest and, therefore, increases the likelihood of picking up a variant at low allelic frequency within the total pool of cfDNA. However, it should be noted that targeted methods reduce the discovery of novel cancer-associated loci.
Other methods that can be adapted to increase analytical sensitivity include in silico enrichment of shorter fragments from the cfDNA pool, as ctDNA fragments have been shown to be, in general, shorter than other cfDNA fragments [44, 94]. Additionally, when detecting a smaller tumour burden such as in early-stage malignancy, MRD or disease recurrence, a larger-volume sample from patients should be considered. Future studies should be aimed at assessing the best sample source and the utility of taking multiple samples concurrently (i.e., plasma and urine) to improve ctDNA yield.
4 Clinical applications of ctDNA
The clinical applications of circulating DNA in a cancer setting are numerous and varied. These applications include detection and localisation of neoplastic growth, prediction of tumour staging and prognosis, tumour genotyping and identification of potential targeted therapeutics, monitoring treatment efficacy and resistance, as well as identification of early-stage recurrence.
4.1 For early detection of disease
Early identification and intervention of cancer growth has the potential to improve therapy response and overall survival rates. ctDNA can be detected in patient plasma before conventional screening methods identify a cancerous lesion, as shown in one particular prospective study that found that KRAS2 and P53 mutations in otherwise ‘healthy’ individuals were associated with a higher risk of developing clinically detectable bladder cancer within 6 years (OR, 4.25 and 1.81, respectively) [95]. Whether this study represents the identification of a biomarker for cancer development, or a true identification of malignancy in pre-symptomatic individuals is still unclear. Nevertheless, it does represent a proof of concept for the potential incorporation of ctDNA into future screening programs. A deficit of this and similar studies is the interrogation of only 1 or 2 cancer-associated loci, making it likely that some true-positive patients may be overlooked for further investigative procedures in the absence of a ‘hallmark’ mutation [96, 97]. Moreover, a small number of healthy individuals harbour cancer-associated genomic alterations [98, 99], suggesting that analysis of a limited gene set is insufficient to confirm the presence or absence of ctDNA.
Assessment of the mutational and/or copy number status of a panel of pan-cancer-associated genes could potentially provide a more reliable method of identifying early-stage disease by increasing the sensitivity for detecting ctDNA. A multi-gene targeted NGS panel was recently utilised by Nair et al. on ctDNA isolated from uterine lavage fluid of women undergoing hysteroscopy for endometrial cancer screening. Ultra-deep sequencing revealed that all 7 women who received a positive early-stage diagnosis by classical histopathological assessment also had ctDNA harbouring significant cancer-associated gene mutations [26]. On a much larger scale, Cohen et al. reported the advent of CancerSEEK, a blood test with the ability to detect eight common cancer types across 1005 newly-diagnosed patients. CancerSEEK combines data collected from amplicon-based sequencing of cfDNA at 61 pan-cancer loci with levels of 8 circulating protein markers [100]. Of these eight cancer types, five currently have no effective screening test. With an assay specificity of over 99% and a sensitivity ranging from 69-98% (depending on cancer type), the possibility of using such liquid biopsies for early cancer detection is gaining momentum.
Still, whilst these early proof-of-concept studies are promising, a number of technical and biomedical challenges need to be overcome before a ctDNA screening assay becomes clinically attainable. Assay sensitivity requires ongoing improvement to detect early-stage cancer with high confidence and a low frequency of false-negative determinations. Currently, deep sequencing techniques meet this challenge better than low-throughput assays such as digital PCR. Additionally, the type and amount of specimen used (e.g. urine versus plasma) is also an important consideration when trying to identify small quantities of tumour DNA in an asymptomatic individual [26, 101]. Additionally, current patient follow-up tests may be of little use to ctDNA positive patients in which no neoplasm is detectable by current methods, except to pinpoint patients in need of close surveillance.
4.2 For cancers of unknown primary
Malignant cancers in which the disseminated site(s) are identified but not the primary location are associated with an extremely poor prognosis (mean survival < 12 months) due to a lack of tailored therapeutic strategies [102]. Epigenetic patterns of ctDNA are often highly stable and unique to particular cell or tissue types even in a cancer setting, and thus may facilitate the identification of the primary mass in CUPs (cancers of unknown primary). Recently, Lehmann-Werman et al. interrogated methylome databases to generate a list of methylation patterns than can be used to distinguish between different cell and tissue types. Consequently, through bisulfite sequencing, they demonstrated that methylation markers specific for exocrine pancreatic cells corresponded to those found in ctDNA of patients with pancreatic cancer [33]. This notion provides preliminary evidence that the ctDNA epigenome can be used to inform on cell-of-origin. Additionally, patterns of nucleosomal spacing of ctDNA fragments can also be used to infer cell types contributing to a pathological state. Snyder et al. assessed nucleosomal positioning in various solid tumours via deep sequencing and reported that the nucleosomal footprint of the tumour cells corresponds to ‘nucleosome maps’ derived from cancer cell lines [103]. In addition to epigenetic patterns, information on microsatellite instability has also been used to provide a primary source of diagnosis, as was the case in two trophoblastic tumours without a definitive histopathological diagnosis [104]. These proof-of-concept studies have revealed the potential utility of ctDNA for the localisation of a primary neoplastic site, and future studies should include larger cohort sizes and CUP patients to fully elucidate the clinical utility of ctDNA in providing a tissue-specific signal.
4.3 For tumour genotyping
Increasingly, there has been a transition away from anatomical and histological classification of tumours to a system based on molecular classification. This paradigm shift has allowed the introduction of precision medicine – customisation of healthcare and therapies based on genetic and molecular landscapes. The analysis of ctDNA has provided an avenue by which genetic and molecular signatures can be detected in a minimally invasive fashion. However, in order for ctDNA to become clinically relevant and play an integral part in day-to-day practice, there must be demonstration of a high concordance between mutational profiles found in ctDNAs and those found in solid biopsy samples. Studies in NSCLC have so far had the most success in this regard, with a recent trial showing the ability to detect epidermal growth factor receptor (EGFR) mutations in patient cfDNA with a concordance of 94.3% to matched solid biopsies. Additionally, 12 patients with a previously undefined mutational status due to technical difficulties with obtaining a tissue biopsy were identified as mutation-positive, and were subsequently treated with the EGFR inhibitor gefitinib [105]. There have been similar studies performed in a variety of other malignancies, including colon cancer and melanoma, were KRAS and BRAF mutations were interrogated, respectively [106,107,108,109]. Within these studies, variable concordance rates, sensitivities and specificities were reported, likely, in part, due to the assay types and technological platforms used. It should also be noted that rates of concordance between solid and liquid biopsies may not be an ideal method of assay validation, as ctDNA is expected to report on the genetic profile of all malignant populations [110], whereas a single tissue biopsy is unlikely to represent the full molecular landscape of a cancer.
4.4 For tumour staging and prognosis
Clinicopathological characteristics such as tumour stage and burden have been found to be significantly associated with both relative amounts and absolute concentrations of ctDNA [111,112,113]. Specifically, a 0.04% increase in plasma ctDNA levels (measured via TP53 mutant allele fraction) has been reported for every additional 1 cm3 of high grade serous ovarian tumour, as measured by computed tomography scans [114]. Such a statistic cannot be directly applied to other tumour types, due to the inherent differences in rates of cellular turnover, location and vascularity. Nevertheless, a liquid biopsy to determine tumour burden has clear advantages over conventional imaging and biomarker analyses as its short half-life allows real-time reporting [37]. In fact, in metastatic breast cancer, ctDNA has a greater correlation with tumour size when compared to both circulating tumour cells (CTCs) and the conventional protein biomarker CA 15-3 (Cancer Antigen 15-3) [115]. Studies measuring ctDNA levels to investigate tumour size almost always occur in a high burden setting, with the clinical utility of using ctDNA to report on precise tumour volumes in a minimal disease setting still being largely unclear.
The link between ctDNA levels and tumour burden suggests that cfDNA has prognostic ability. Indeed, many studies have observed a relationship between either total cfDNA or ctDNA levels and tumour staging (based on tumour size and extent of metastatic spread) [30, 74, 116, 117]. One particular study in 640 patients with various solid tumours found mutant fragments within plasma ctDNA increasing significantly between tumour stages (I through IV). The study also reported a significant association between mutant allele count and survival probability, with a two-fold rise reducing the 2-year survival probability from 25% to 10% [74]. A recent meta-analysis assessing a total of 1,170 non-small cell lung cancer (NSCLC) patients also concluded that a high cfDNA concentration was associated with a worse outcome in stage III-IV disease [118]. Other studies have reported similar findings [116, 119, 120] although, again, focus has been on tumour types with a high mutational burden and a distinct set of ‘hallmark’ alterations such as colorectal cancer. Larger gene panels are required to increase the sensitivity for detecting ctDNA in genetically heterogeneous tumours, or total cfDNA concentrations considered instead, although this comes with its own challenges due to fluctuations that may occur independently of tumour cell release [39, 40]. Additional challenges for using ctDNA for tumour staging arise from the high amount of inter-patient variability of plasma levels within the same stage and type of disease [74]. This is presumably due to differences in tumour vascularisation and extent of metastatic spread and, therefore, tumour burden. Such inter-individual differences need to be considered when using ctDNA as a staging tool, and perhaps a two-armed approach combining ctDNA with conventional methods may have the greatest utility.
4.5 For guiding treatment selection
The demonstration of reliable concordances between cfDNA and solid biopsies in regards to mutational profiling has given rise to studies investigating whether liquid biopsies can guide the selection of targeted therapies. Although retrospective in nature, multiple studies have showcased the ability of ctDNA to predict outcomes to targeted therapy. One such study in NSCLC reported that patients positive for the T790M EFGR mutation in plasma have outcomes that are equivalent to patients deemed positive by a tissue-based assay when treated with osimertinib, a third-generation EGFR inhibitor (overall response rates of 63% and 62%, respectively) [121]. Likewise, a study assessing BRAF mutations in melanoma ctDNA as a predictive marker of response to dabrafenib across four clinical trials concluded that it may be a useful marker for targeted therapy, although there was a small subset of plasma mutant-negative patients who experienced a therapeutic response [109]. Regardless, the robust data supporting ctDNA profiling in NSCLC has influenced major management guidelines allowing plasma biopsy as a viable alternative for molecular profiling before treatment in cases in which an initial tissue biopsy is non-diagnostic, and repeated biopsy is not feasible [4]. As a result, first-line targeted therapies in EGFR-mutated NSCLC based on liquid biopsy are now a reality due to approval by major regulatory bodies in the United States and Europe [122]. Other cancer types are not far behind, with numerous clinical trials currently underway assessing the predictive utility of ctDNA for approved targeted therapies.
In addition, total ctDNA levels may also be useful for guiding therapy. A recent study assessing plasma ctDNA in 21,807 advanced cancer patients revealed a relationship between ctDNA levels and the total mutational burden of a tumour, which may illuminate patients responsive to immunotherapy [123]. However, this link was only identified in breast cancer, colorectal cancer and NSCLC, and may be less applicable to cancer types with lower ctDNA fractions (e.g. glioma).
4.6 For monitoring treatment efficacy
Quantification and analysis of ctDNA represents a novel and timely method for measuring response (or lack thereof) to therapy. Currently, a combination of patient symptomology, biochemical markers and radiographic evaluation is used to objectively evaluate a patient’s response to treatment. These methods are not without significant drawbacks, however. Definitive symptoms can often lag behind overt disease progression, and tumour markers have been shown to lack specificity. For example, serum prostate-specific antigen (PSA) is widely used for disease monitoring in advanced prostate cancer despite progression regularly occurring in the absence of any detectable changes in PSA [124, 125]. Furthermore, the extended half-lives of tumour markers such as PSA allow them to persist in the circulation at levels that may not reflect treatment outcome, thereby hampering dynamic treatment decisions [126, 127].
In comparison, the unique properties of ctDNA, in particular its short half-life, make it an attractive real-time indicator of treatment efficacy and may lead to the identification of response earlier than clinical detection. Studies monitoring patients throughout therapy have reported that the dynamic changes in ctDNA throughout treatment correlates with response [128,129,130]. One such prospective study in women with metastatic breast cancer concluded that ctDNA is superior to other circulating biomarkers such as CA 15-3 in sensitivity and that, due to its dynamic range, it shows a greater correlation to changes in tumour burden [129]. Similar results have been reported in NSCLC, where reductions in ctDNA concentrations can predate favourable imaging findings by 4-6 weeks [131].
A recent study on metastatic melanoma suggested that clinical responses to anti-PD-1 immunotherapy may be preceded by reductions in ctDNA within 2-4 weeks of treatment initiation, although the cohort of 5 patients needs to be expanded [130]. It is likely that a drop in ctDNA is preceded by an initial peak in circulating ctDNA levels due to an initial increase in tumour cell death. Indeed, an earlier study on metastatic melanoma reported that an early peak in ctDNA level within the first week was indicative of a favourable response to T-cell transfer immunotherapy [132]. If confirmed, these data may be used to shape guidelines as to when ctDNA sampling should take place in order to assess early treatment responses to immunotherapy.
4.7 For identifying treatment resistance
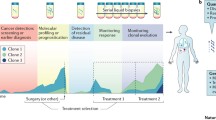
It is well-appreciated that targeted therapies exert selective pressure on sensitive populations of neoplastic cells, ultimately leading to a molecular evolution of a patient’s tumour and the development of treatment resistance [133]. If, during treatment, relative changes in mutations can be observed early, then clinicians may be one step ahead, preparing for a second line of therapy to immediately combat the rise of novel resistant clones. Serial analysis of ctDNA represents an avenue through which clonal evolution can be closely monitored for the manifestation of resistance-conferring mutations that invariably develop during the treatment course [134,135,136] (Fig. 2). The ability of ctDNA to report on the genetic landscape of multiple tumour sites will also ensure that novel metastases arising from sub-clones are not ignored [110, 137]. Most evidence outlining the utility of ctDNA in monitoring treatment resistance currently comes from NSCLC, with the T790M mutation conferring resistance to EGFR tyrosine kinase inhibitors being reliably detected in the ctDNA of patients 16 to 49 weeks before clinical or radiological progression is observed [138,139,140]. There is also emerging evidence that ctDNA can be used to investigate therapeutic resistance in other malignancies, including advanced prostate cancer, where various somatic aberrations have been linked to resistance to the androgen-axis targeting therapeutic compounds enzalutamide and abiraterone acetate [75, 141, 142].
Monitoring of tumour clonal evolution via serial liquid biopsies. Targeted therapeutics puts cancer cells under selective pressure, changing the genetic landscape and, thereby, altering treatment efficacy. Detection of such clonal evolution in circulating tumour DNA can help to guide the selection of ensuing therapies
The preferred method of interrogating the ctDNA mutational landscape for resistance-conferring mutations is via a candidate gene approach such as digital droplet PCR, presumably due to its superior sensitivity (Table 3). However, the application of ctDNA to the identification of treatment resistance warrants deep sequencing studies interrogating multiple targets in order to identify all potential resistance-conferring clones. Whole exome or large-panel sequencing of cfDNA are currently limited by their lower sensitivity and higher costs, with patient-specific mutation panels presenting a more desirable alternative [143]. This, however, also presents the risk of missing de novo events.
4.8 For monitoring MRD
A central question after any anti-tumour therapy (surgery or otherwise) is whether the treatment has left behind any residual cancer cells or microscopic metastatic deposits that are responsible for relapse. Minimal residual disease (MRD) is a prominent concept in haematological malignancies, however the ability to detect small populations of residual tumour cells in solid tumours is still underdeveloped. A highly sensitive and accurate assay capable of detecting the presence of ctDNA can potentially be used to identify patients in need of further adjuvant therapies whilst saving low-risk patients from unnecessary and expensive treatments. It has been estimated that approximately 44 million malignant cells are required for significant plasma ctDNA detection, a number below the limit of resolution of current radiological techniques [144]. The ability to detect ctDNA even in the absence of any other clinical evidence of cancer makes it a prime candidate for measuring early-stage disease recurrence.
A recent prospective correlative biomarker study in 230 early-stage colon cancer patients reported that only 10% of patients with no detectable post-operative ctDNA suffered radiological relapse within three years whilst 100% of ctDNA-positive patients suffered disease recurrence [145]. As a result, post-operative ctDNA detection has been incorporated into a prospective clinical trial of stage II colon cancer patients receiving adjuvant chemotherapy. Such a high predictive value is presumably due to the personalised Safe-SeqS assays developed based on the molecular characteristics of the patient’s tumour resected during surgery, and the large volume of plasma collected at each time point (10 ml). Whilst this study clearly shows the validity of a patient-specific assay to detect MRD, results of similar studies in other cancer types are less clear, such as a separate study in multiple myeloma patients which found plasma ctDNA to be a poor predictor of disease recurrence, although one potential limitation of this study is a lack of sensitivity due to under sampling of patient serum (1 ml) [146]. It should also be noted that the design of a personalised assay for monitoring MRD based on somatic mutations in a resected sample may reduce assay sensitivity due to a potential lack of representation of heterogeneous tumours, and the emergence of novel sub-clones post-surgery.
5 Concluding remarks and future directions
Liquid biopsies are likely to play an increasing role in the implementation of precision medicine and the personalised treatment of various cancers. At present, many proof-of-concept studies have paved the way for the field to move from exploratory studies to implementation in clinical trials where ctDNA profiling guides patient management decisions. Currently, over 200 clinical trials investigating the prognostic and/or predictive biomarker utility of ctDNA across various cancer types are underway or actively recruiting (Fig. 3). However, there remain multiple issues delaying the widespread incorporation of ctDNA into routine patient care. Future focus should be on establishing optimal techniques for sample collection, cfDNA isolation (to improve yield of all relevant fragment sizes) and analysis. In addition, further studies are required to better understand the biological characteristics of ctDNA, such as mechanisms of release and clearance, and a proper characterisation of normal reference ranges. It is also important to understand how non-malignant conditions (e.g. inflammation, illness) contribute to the total cfDNA pool, as this may be informative for the optimal time for liquid biopsy collection.
Current clinical trials utilising cell-free DNA in cancer settings. Trials are either recruiting or underway, as detailed on clinicaltrials.gov. Search performed on 29/06/2018
As with any biomarker, a critical step to incorporation into clinical care is validation of a highly sensitive, universally agreed-upon platform. At present, multi-gene approaches capable of identifying several variant types (e.g. SNVs, CNVs) such as hybrid-capture based deep sequencing appear to be the most promising for biomarker discovery and development, and may soon match candidate-driven approaches in terms of sensitivity. Such assays should be developed with rigorous attention to analytical validity, accompanied by precise reporting on methods in accordance to best practice guidelines [147, 148]. Only by first addressing analytical validity one can begin to broach to the idea of clinical validation in prospective clinical cohorts.
On the challenge of clinical validation, there is an imperative that investigators continue to design biomarker-driven, prospective clinical trials, where patients receive specific treatment(s) on the basis of a predefined marker of interest [149, 150]. This focused approach, where the ctDNA biomarker will define the management plan, enriches study cohorts for patients likely to benefit, potentially streamlining drug development and accelerating drug approval [151]. It is important that ctDNA biomarker testing in such trials is restricted to clinical testing laboratories with a Clinical Laboratory Improvement Amendments (CLIA) certification, preferably with ample prior experience with the assay is question. Finally, robust evidence demonstrating definitively improved clinical outcomes or superior diagnostic capability compared to any current gold-standard is crucial before we can confidently transfer a ctDNA biomarker assay into routine clinical practice.
A future with ctDNA as a validated biomarker is on the horizon. As concordance of detectable driver alterations between ctDNA and solid biopsies improves, it will only be a matter of time before such a minimally-invasive liquid biopsy becomes a vital component of clinical practice and precision medicine.
References
M. Ilié, P. Hofman, Pros: Can tissue biopsy be replaced by liquid biopsy? Transl Lung Cancer Res 5, 420–423 (2016)
N. Krishnamurthy, E. Spencer, A. Torkamani, L. Nicholson, Liquid biopsies for cancer: Coming to a patient near you. J Clin Med 6, 3 (2017)
M.J. Overman, J. Modak, S. Kopetz, R. Murthy, J.C. Yao, M.E. Hicks, J.L. Abbruzzese, A.L. Tam, Use of research biopsies in clinical trials: Are risks and benefits adequately discussed? J Clin Oncol 31, 17–22 (2013)
P.A. VanderLaan, N. Yamaguchi, E. Folch, D.H. Boucher, M.S. Kent, S.P. Gangadharan, A. Majid, M.A. Goldstein, M.S. Huberman, O.N. Kocher, D.B. Costa, Success and failure rates of tumor genotyping techniques in routine pathological samples with non-small-cell lung cancer. Lung Cancer 84, 39–44 (2014)
S.Q. Wong, J. Li, A.Y.C. Tan, R. Vedururu, J.-M.B. Pang, H. Do, J. Ellul, K. Doig, A. Bell, G.A. McArthur, S.B. Fox, D.M. Thomas, A. Fellowes, J.P. Parisot, A. Dobrovic, Sequence artefacts in a prospective series of formalin-fixed tumours tested for mutations in hotspot regions by massively parallel sequencing. BMC Med Genet 7, 23 (2014)
Z.-D. Hu, Z.-R. Zhou, S. Qian, How to analyze tumor stage data in clinical research. J Thorac Dis 7, 566–575 (2015)
R. Catarino, M.M. Ferreira, H. Rodrigues, A. Coelho, A. Nogal, A. Sousa, R. Medeiros, Quantification of free circulating tumor DNA as a diagnostic marker for breast cancer. DNA Cell Biol 27, 415–421 (2008)
N. Hjortholm, E. Jaddini, K. Hałaburda, E. Snarski, Strategies of pain reduction during the bone marrow biopsy. Ann Hematol Oncol 92, 145–149 (2013)
J.M. Hemmer, J.C. Kelder, H.P.M. van Heesewijk, Stereotactic large-core needle breast biopsy: Analysis of pain and discomfort related to the biopsy procedure. Eur Radiol 18, 351–354 (2008)
S. Perakis, M.R. Speicher, Emerging concepts in liquid biopsies. BMC Med 15, 75 (2017)
F. Janku, Tumor heterogeneity in the clinic: Is it a real problem? Ther Adv Med Oncol 6, 43–51 (2014)
M. Jamal-Hanjani, G.A. Wilson, S. Horswell, R. Mitter, O. Sakarya, T. Constantin, R. Salari, E. Kirkizlar, S. Sigurjonsson, R. Pelham, S. Kareht, B. Zimmermann, C. Swanton, Detection of ubiquitous and heterogeneous mutations in cell-free DNA from patients with early-stage non-small-cell lung cancer. Ann Oncol 27, 862–867 (2016)
M. Murtaza, S.J. Dawson, K. Pogrebniak, O.M. Rueda, E. Provenzano, J. Grant, S.F. Chin, D.W. Tsui, F. Marass, D. Gale, H.R. Ali, P. Shah, T. Contente-Cuomo, H. Farahani, K. Shumansky, Z. Kingsbury, S. Humphray, D. Bentley, S.P. Shah, M. Wallis, N. Rosenfeld, C. Caldas, Multifocal clonal evolution characterized using circulating tumour DNA in a case of metastatic breast cancer. Nat Commun 6, 8760 (2015)
O. Eikrem, C. Beisland, K. Hjelle, A. Flatberg, A. Scherer, L. Landolt, T. Skogstrand, S. Leh, V. Beisvag, H.P. Marti, Transcriptome sequencing (rnaseq) enables utilization of formalin-fixed, paraffin-embedded biopsies with clear cell renal cell carcinoma for exploration of disease biology and biomarker development. PloS One 11, e0149743 (2016)
G. Siravegna, S. Marsoni, S. Siena, A. Bardelli, Integrating liquid biopsies into the management of cancer. Nature Rev Clin Oncol 14, 531–548 (2017)
R. Govindan, L. Ding, M. Griffith, J. Subramanian, N.D. Dees, K.L. Kanchi, C.A. Maher, R. Fulton, L. Fulton, J. Wallis, K. Chen, J. Walker, S. McDonald, R. Bose, D. Ornitz, D. Xiong, M. You, D.J. Dooling, M. Watson, E.R. Mardis, R.K. Wilson, Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150, 1121–1134 (2012)
W. Pan, W. Gu, S. Nagpal, M.H. Gephart, S.R. Quake, Brain tumor mutations detected in cerebral spinal fluid. Clin Chem 61, 514–522 (2015)
F.R. Harris, I.V. Kovtun, J. Smadbeck, F. Multinu, A. Jatoi, F. Kosari, K.R. Kalli, S.J. Murphy, G.C. Halling, S.H. Johnson, M.C. Liu, A. Mariani, G. Vasmatzis, Quantification of somatic chromosomal rearrangements in circulating cell-free DNA from ovarian cancers. Sci Rep 6, 29831 (2016)
J. Li, R.L. Dittmar, S. Xia, H. Zhang, M. Du, C.C. Huang, B.R. Druliner, L. Boardman, L. Wang, Cell-free DNA copy number variations in plasma from colorectal cancer patients. Mol Oncol 11, 1099–1111 (2017)
R. Casadonte, R.M. Caprioli, Proteomic analysis of formalin-fixed paraffin embedded tissue by maldi imaging mass spectrometry. Nat Protoc 6, 1695–1709 (2011)
M. Møller, S.H. Strand, K. Mundbjerg, G. Liang, I. Gill, C. Haldrup, M. Borre, S. Høyer, T.F. Ørntoft, K.D. Sørensen, Heterogeneous patterns of DNA methylation-based field effects in histologically normal prostate tissue from cancer patients. Sci Rep 7, 40636 (2017)
K. Warton, G. Samimi, Methylation of cell-free circulating DNA in the diagnosis of cancer. Front Mol Biosci 2, 13 (2015)
X. Han, J. Wang, Y. Sun, Circulating tumor DNA as biomarkers for cancer detection. Genomics Proteomics Bioinformatics 15, 59–72 (2017)
L. Kubiczkova-Besse, D. Drandi, L. Sedlarikova, S. Oliva, M. Gambella, P. Omedè, Z. Adam, L. Pour, S. Sevcikova, M. Boccadoro, A. Palumbo, R. Hajek, Cell-free DNA for minimal residual disease monitoring in multiple myeloma patients. Blood 124, 3423–3423 (2014)
P. Mandel, P. Metais, Les acides nucléiques du plasma sanguin chez l'homme. Comptes Rendus des Séances de la Société de Biologie et de Ses Filiales 142, 241–243 (1948)
N. Nair, O. Camacho-Vanegas, D. Rykunov, M. Dashkoff, S.C. Camacho, C.A. Schumacher, J.C. Irish, T.T. Harkins, E. Freeman, I. Garcia, E. Pereira, S. Kendall, R. Belfer, T. Kalir, R. Sebra, B. Reva, P. Dottino, J.A. Martignetti, Genomic analysis of uterine lavage fluid detects early endometrial cancers and reveals a prevalent landscape of driver mutations in women without histopathologic evidence of cancer: A prospective cross-sectional study. PLoS Medicine 13, e1002206 (2016)
I. Botezatu, O. Serdyuk, G. Potapova, V. Shelepov, R. Alechina, Y. Molyaka, V. Ananev, I. Bazin, A. Garin, M. Narimanov, V. Knysh, H. Melkonyan, S. Umansky, A. Lichtenstein, Genetic analysis of DNA excreted in urine: A new approach for detecting specific genomic DNA sequences from cells dying in an organism. Clin Chem 46, 1078–1084 (2000)
S.K. Mithani, I.M. Smith, S. Zhou, A. Gray, W.M. Koch, A. Maitra, J.A. Califano, Mitochondrial resequencing arrays detect tumor-specific mutations in salivary rinses of patients with head and neck cancer. Clin Can Res 13, 7335–7340 (2007)
J.D. Krimmel, M.W. Schmitt, M.I. Harrell, K.J. Agnew, S.R. Kennedy, M.J. Emond, L.A. Loeb, E.M. Swisher, R.A. Risques, Ultra-deep sequencing detects ovarian cancer cells in peritoneal fluid and reveals somatic tp53 mutations in noncancerous tissues. Proc Natl Acad Sci U S A 113, 6005–6010 (2016)
J.C.M. Wan, C. Massie, J. Garcia-Corbacho, F. Mouliere, J.D. Brenton, C. Caldas, S. Pacey, R. Baird, N. Rosenfeld, Liquid biopsies come of age: Towards implementation of circulating tumour DNA. Nat Rev Cancer 17, 223–238 (2017)
M. Stroun, J. Lyautey, C. Lederrey, A. Olson-Sand, P. Anker, About the possible origin and mechanism of circulating DNA. Clinica Chimica Acta 313, 139–142 (2001)
D.L. Peters, P.J. Pretorius, Origin, translocation and destination of extracellular occurring DNA — a new paradigm in genetic behaviour. Clinica Chimica Acta 412, 806–811 (2011)
R. Lehmann-Werman, D. Neiman, H. Zemmour, J. Moss, J. Magenheim, A. Vaknin-Dembinsky, S. Rubertsson, B. Nellgård, K. Blennow, H. Zetterberg, K. Spalding, M.J. Haller, C.H. Wasserfall, D.A. Schatz, C.J. Greenbaum, C. Dorrell, M. Grompe, A. Zick, A. Hubert, M. Maoz, V. Fendrich, D.K. Bartsch, T. Golan, S.A. Ben Sasson, G. Zamir, A. Razin, H. Cedar, A.M.J. Shapiro, B. Glaser, R. Shemer, Y. Dor, Identification of tissue-specific cell death using methylation patterns of circulating DNA. Proc Natl Acad Sci U S A 113,E1826-E1834 (2016)
S. Breitbach, S. Tug, S. Helmig, D. Zahn, T. Kubiak, M. Michal, T. Gori, T. Ehlert, T. Beiter, P. Simon, Direct quantification of cell-free, circulating DNA from unpurified plasma. PloS One 9, e87838 (2014)
F. Mouliere, S. El Messaoudi, D. Pang, A. Dritschilo, A.R. Thierry, Multi-marker analysis of circulating cell-free DNA toward personalized medicine for colorectal cancer. Mol Oncol 8, 927–941 (2014)
W. Yao, C. Mei, X. Nan, L. Hui, Evaluation and comparison of in vitro degradation kinetics of DNA in serum, urine and saliva: A qualitative study. Gene 590, 142–148 (2016)
Y.M.D. Lo, J. Zhang, T.N. Leung, T.K. Lau, A.M.Z. Chang, N.M. Hjelm, Rapid clearance of fetal DNA from maternal plasma. Am J Hum Genet 64, 218–224
O.E. Bryzgunova, P.P. Laktionov, Extracellular nucleic acids in urine: Sources, structure, diagnostic potential. Acta Naturae 7, 48–54 (2015)
A.N. Butt, R. Swaminathan, Overview of circulating nucleic acids in plasma/serum. Ann N Y Acad Sci 1137, 236–242 (2008)
S. Volik, M. Alcaide, R.D. Morin, C. Collins, Cell-free DNA (cfdna): Clinical significance and utility in cancer shaped by emerging technologies. Mol Cancer Res 14, 898–908 (2016)
S.A. Leon, B. Shapiro, D.M. Sklaroff, M.J. Yaros, Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res 37, 646–650 (1977)
G.D. Sorenson, D.M. Pribish, F.H. Valone, V.A. Memoli, D.J. Bzik, S.L. Yao, Soluble normal and mutated DNA sequences from single-copy genes in human blood. Cancer Epidemiol Biomarkers 3, 67–71 (1994)
S. Jahr, H. Hentze, S. Englisch, D. Hardt, F.O. Fackelmayer, R.-D. Hesch, R. Knippers, DNA fragments in the blood plasma of cancer patients: Quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res 61, 1659–1665 (2001)
H.R. Underhill, J.O. Kitzman, S. Hellwig, N.C. Welker, R. Daza, D.N. Baker, K.M. Gligorich, R.C. Rostomily, M.P. Bronner, J. Shendure, Fragment length of circulating tumor DNA. PLoS Genetics 12, e1006162 (2016)
A.R. Thierry, F. Mouliere, C. Gongora, J. Ollier, B. Robert, M. Ychou, M. Del Rio, F. Molina, Origin and quantification of circulating DNA in mice with human colorectal cancer xenografts. Nucleic Acids Res 38, 6159–6175 (2010)
S. Nagata, Apoptotic DNA fragmentation. Exp Cell Res 256, 12–18 (2000)
C. Tetta, E. Ghigo, L. Silengo, M.C. Deregibus, G. Camussi, Extracellular vesicles as an emerging mechanism of cell-to-cell communication. Endocrine 44, 11–19 (2013)
E.S. Morozkin, P.P. Laktionov, E.Y. Rykova, O.E. Bryzgunova, V.V. Vlassov, Release of nucleic acids by eukaryotic cells in tissue culture. Nucleosides, Nucleotides and Nucleic Acids 23, 927–930 (2004)
M. Abolhassani, J. Tillotson, J. Chiao, Characterization of the release of DNA by a human leukemia-cell line hl-60. Int J Oncol 4, 417–421 (1994)
A.J. Bronkhorst, J.F. Wentzel, J. Aucamp, E. van Dyk, L. du Plessis, P.J. Pretorius, Characterization of the cell-free DNA released by cultured cancer cells. Biochimica et Biophysica Acta 1863, 157–165 (2016)
J.A. Collins, C.A. Schandl, K.K. Young, J. Vesely, M.C. Willingham, Major DNA fragmentation is a late event in apoptosis. J Histochem Cytochem 45, 923–934 (1997)
M. Beranek, I. Sirak, M. Vosmik, J. Petera, M. Drastikova, V. Palicka, Carrier molecules and extraction of circulating tumor DNA for next generation sequencing in colorectal cancer. Acta Medica (Hradec Kralove) 59, 54–58 (2016)
M. Peng, C. Chen, A. Hulbert, M.V. Brock, F. Yu, Non-blood circulating tumor DNA detection in cancer. Oncotarget 8, 69162–69173 (2017)
M.S. Gordon, Managing anemia in the cancer patient: Old problems, future solutions. Oncologist 7, 331–341 (2002)
D.A. Ahlquist, W.R. Taylor, D.W. Mahoney, H. Zou, M. Domanico, S.N. Thibodeau, L.A. Boardman, B.M. Berger, G.P. Lidgard, The stool DNA test is more accurate than the plasma septin 9 test in detecting colorectal neoplasia. Clin Gastroenterol Hepatol 10, 272–277 (2012)
A. Castagnaro, E. Marangio, A. Verduri, A. Chetta, R. D'Ippolito, M. Del Donno, D. Olivieri, G. Di Cola, Microsatellite analysis of induced sputum DNA in patients with lung cancer in heavy smokers and in healthy subjects. Exp Lung Res 33, 289–301 (2007)
L. De Mattos-Arruda, R. Mayor, C.K. Ng, B. Weigelt, F. Martinez-Ricarte, D. Torrejon, M. Oliveira, A. Arias, C. Raventos, J. Tang, E. Guerini-Rocco, E. Martinez-Saez, S. Lois, O. Marin, X. de la Cruz, S. Piscuoglio, R. Towers, A. Vivancos, V. Peg, S. Ramon y Cajal, J. Carles, J. Rodon, M. Gonzalez-Cao, J. Tabernero, E. Felip, J. Sahuquillo, M.F. Berger, J. Cortes, J.S. Reis-Filho, J. Seoane, Cerebrospinal fluid-derived circulating tumour DNA better represents the genomic alterations of brain tumours than plasma. Nat Commun 6, 8839 (2015)
E.C. Hung, R.W. Chiu, Y.M. Lo, Detection of circulating fetal nucleic acids: A review of methods and applications. J Clin Pathol 62, 308–313 (2009)
J. Jen, L. Wu, D. Sidransky, An overview on the isolation and analysis of circulating tumor DNA in plasma and serum. Ann N Y Acad Sci 906, 8–12 (2000)
T.H. Lee, L. Montalvo, V. Chrebtow, M.P. Busch, Quantitation of genomic DNA in plasma and serum samples: Higher concentrations of genomic DNA found in serum than in plasma. Transfusion 41, 276–282 (2001)
A.J. Bronkhorst, J. Aucamp, P.J. Pretorius, Cell-free DNA: Preanalytical variables. Clinica Chimica Acta 450, 243–253 (2015)
H. Markus, T. Contente-Cuomo, M. Farooq, W.S. Liang, M.J. Borad, S. Sivakumar, S. Gollins, N.L. Tran, H.D. Dhruv, M.E. Berens, A. Bryce, A. Sekulic, A. Ribas, J.M. Trent, P.M. LoRusso, M. Murtaza, Evaluation of pre-analytical factors affecting plasma DNA analysis. Scie Rep 8, 7375 (2018)
Y.Y. Lui, K.W. Chik, R.W. Chiu, C.Y. Ho, C.W. Lam, Y.M. Lo, Predominant hematopoietic origin of cell-free DNA in plasma and serum after sex-mismatched bone marrow transplantation. Clin Chem 48, 421–427 (2002)
M. Jung, S. Klotzek, M. Lewandowski, M. Fleischhacker, K. Jung, Changes in concentration of DNA in serum and plasma during storage of blood samples. Clin Chem 49, 1028–1029 (2003)
N.Y. Lam, T.H. Rainer, R.W. Chiu, Y.M. Lo, Edta is a better anticoagulant than heparin or citrate for delayed blood processing for plasma DNA analysis. Clin Chem 50, 256–257 (2004)
K.C. Chan, S.W. Yeung, W.B. Lui, T.H. Rainer, Y.M. Lo, Effects of preanalytical factors on the molecular size of cell-free DNA in blood. Clin Chem 51, 781–784 (2005)
R.E. Board, V.S. Williams, L. Knight, J. Shaw, A. Greystoke, M. Ranson, C. Dive, F.H. Blackhall, A. Hughes, Isolation and extraction of circulating tumor DNA from patients with small cell lung cancer. Ann N Y Acad Sci 1137, 98–107 (2008)
S.E. Norton, K.K. Luna, J.M. Lechner, J. Qin, M.R. Fernando, A new blood collection device minimizes cellular DNA release during sample storage and shipping when compared to a standard device. J Clin Lab Anal 27, 305–311 (2013)
M.R. Fernando, K. Chen, S. Norton, G. Krzyzanowski, D. Bourne, B. Hunsley, W.L. Ryan, C. Bassett, A new methodology to preserve the original proportion and integrity of cell-free fetal DNA in maternal plasma during sample processing and storage. Prenat Diagn 30, 418–424 (2010)
M. Hidestrand, R. Stokowski, K. Song, A. Oliphant, J. Deavers, M. Goetsch, P. Simpson, R. Kuhlman, M. Ames, M. Mitchell, A. Tomita-Mitchell, Influence of temperature during transportation on cell-free DNA analysis. Fetal Diagn Ther 31, 122–128 (2012)
Q. Kang, N.L. Henry, C. Paoletti, H. Jiang, P. Vats, A.M. Chinnaiyan, D.F. Hayes, S.D. Merajver, J.M. Rae, M. Tewari, Comparative analysis of circulating tumor DNA stability in k3edta, streck, and cellsave blood collection tubes. Clin Biochem 49, 1354–1360 (2016)
I. Medina Diaz, A. Nocon, D.H. Mehnert, J. Fredebohm, F. Diehl, F. Holtrup, Performance of streck cfdna blood collection tubes for liquid biopsy testing. PloS One 11, e0166354 (2016)
E. Beutler, T. Gelbart, W. Kuhl, Interference of heparin with the polymerase chain reaction. Biotechniques 9, 166 (1990)
C. Bettegowda, M. Sausen, R.J. Leary, I. Kinde, Y. Wang, N. Agrawal, B.R. Bartlett, H. Wang, B. Luber, R.M. Alani, E.S. Antonarakis, N.S. Azad, A. Bardelli, H. Brem, J.L. Cameron, C.C. Lee, L.A. Fecher, G.L. Gallia, P. Gibbs, D. Le, R.L. Giuntoli, M. Goggins, M.D. Hogarty, M. Holdhoff, S.-M. Hong, Y. Jiao, H.H. Juhl, J.J. Kim, G. Siravegna, D.A. Laheru, C. Lauricella, M. Lim, E.J. Lipson, S.K.N. Marie, G.J. Netto, K.S. Oliner, A. Olivi, L. Olsson, G.J. Riggins, A. Sartore-Bianchi, K. Schmidt, I.-M. Shih, S.M. Oba-Shinjo, S. Siena, D. Theodorescu, J. Tie, T.T. Harkins, S. Veronese, T.-L. Wang, J.D. Weingart, C.L. Wolfgang, L.D. Wood, D. Xing, R.H. Hruban, J. Wu, P.J. Allen, C.M. Schmidt, M.A. Choti, V.E. Velculescu, K.W. Kinzler, B. Vogelstein, N. Papadopoulos, L.A. Diaz, Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 6, 224ra224–224ra224 (2014)
A.A. Azad, S.V. Volik, A.W. Wyatt, A. Haegert, S. Le Bihan, R.H. Bell, S.A. Anderson, B. McConeghy, R. Shukin, J. Bazov, J. Youngren, P. Paris, G. Thomas, E.J. Small, Y. Wang, M.E. Gleave, C.C. Collins, K.N. Chi, Androgen receptor gene aberrations in circulating cell-free DNA: Biomarkers of therapeutic resistance in castration-resistant prostate cancer. Clin Cancer Res 21, 2315–2324 (2015)
D. Charcon-Cortes, L.R. Griffiths, Methods for extracting genomic DNA from whole blood samples: Current perspectives. Br J Appl Sci Technol 2, 1–9 (2014)
C.J. Jorgez, D.D. Dang, J.L. Simpson, D.E. Lewis, F.Z. Bischoff, Quantity versus quality: Optimal methods for cell-free DNA isolation from plasma of pregnant women. Genet Med 8, 615–619 (2006)
A. Psifidi, C.I. Dovas, G. Bramis, T. Lazou, C.L. Russel, G. Arsenos, G. Banos, Comparison of eleven methods for genomic DNA extraction suitable for large-scale whole-genome genotyping and long-term DNA banking using blood samples. PloS One 10, e0115960 (2015)
A.S. Devonshire, A.S. Whale, A. Gutteridge, G. Jones, S. Cowen, C.A. Foy, J.F. Huggett, Towards standardisation of cell-free DNA measurement in plasma: Controls for extraction efficiency, fragment size bias and quantification. Anal Bional Chem 406, 6499–6512 (2014)
J.L. Sherwood, C. Corcoran, H. Brown, A.D. Sharpe, M. Musilova, A. Kohlmann, Optimised pre-analytical methods improve kras mutation detection in circulating tumour DNA (ctdna) from patients with non-small cell lung cancer (nsclc). PloS One 11, e0150197 (2016)
J.-H. Lee, Y. Park, J.R. Choi, E.K. Lee, H.-S. Kim, Comparisons of three automated systems for genomic DNA extraction in a clinical diagnostic laboratory. Yonsei Med J 51, 104–110 (2010)
L. Sorber, K. Zwaenepoel, V. Deschoolmeester, G. Roeyen, F. Lardon, C. Rolfo, P. Pauwels, A comparison of cell-free DNA isolation kits: Isolation and quantification of cell-free DNA in plasma. J Mol Diagn 19, 162–168 (2017)
A. Jena, Pme free-circulating DNA extraction kit (2016)
C. Gaillard, F. Strauss, Avoiding adsorption of DNA to polypropylene tubes and denaturation of short DNA fragments. Technical Tips Online 3, 63–65 (1998)
M. Lecerf, J.L. Goff, Use of eppendorf lobind® tubes to consistently prepare and store standard panels for real-time pcr absolute quantifications (2010)
E. Heitzer, P. Ulz, J. Belic, S. Gutschi, F. Quehenberger, K. Fischereder, T. Benezeder, M. Auer, C. Pischler, S. Mannweiler, Tumor-associated copy number changes in the circulation of patients with prostate cancer identified through whole-genome sequencing. Genome Med 5, 30 (2013)
N. Van Roy, M. Van Der Linden, B. Menten, A. Dheedene, C. Vandeputte, J. Van Dorpe, G. Laureys, M. Renard, T. Sante, T. Lammens, B. De Wilde, F. Speleman, K. De Preter, Shallow whole genome sequencing on circulating cell-free DNA allows reliable noninvasive copy-number profiling in neuroblastoma patients. Clin Cancer Res 23, 6305–6314 (2017)
T. Forshew, M. Murtaza, C. Parkinson, D. Gale, D.W. Tsui, F. Kaper, S.J. Dawson, A.M. Piskorz, M. Jimenez-Linan, D. Bentley, Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med 4, 136 (2012)
X.W. Yongqian Shu, X. Tong, X. Wang, Z. Chang, Y. Mao, X. Chen, J. Sun, Z. Wang, Z. Hong, L. Zhu, C. Zhu, J. Chen, Y. Liang, H. Shao, Y.W. Shao, Circulating tumor DNA mutation profiling by targeted next generation sequencing provides guidance for personalized treatments in multiple cancer types. Sci Rep 7, 583 (2017)
A.A. Chaudhuri, M.S. Binkley, E.C. Osmundson, A.A. Alizadeh, M. Diehn, Predicting radiotherapy responses and treatment outcomes through analysis of circulating tumor DNA. Semin Radiat Oncol 25, 305–312 (2015)
I. Kinde, J. Wu, N. Papadopoulos, K.W. Kinzler, B. Vogelstein, Detection and quantification of rare mutations with massively parallel sequencing. Proc Natl Acad Sci U S A 108, 9530–9535 (2011)
A.M. Newman, S.V. Bratman, J. To, J.F. Wynne, N.C.W. Eclov, L.A. Modlin, C.L. Liu, J.W. Neal, H.A. Wakelee, R.E. Merritt, J.B. Shrager, B.W. Loo Jr., A.A. Alizadeh, M. Diehn, An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med 20, 548–554 (2014)
S.V. Bratman, A.M. Newman, A.A. Alizadeh, M. Diehn, Potential clinical utility of ultrasensitive circulating tumor DNA detection with capp-seq. Expert Rev Mol Diagn 15, 715–719 (2015)
J.S. Bhangu, H. Taghizadeh, T. Braunschmid, T. Bachleitner-Hofmann, C. Mannhalter, Circulating cell-free DNA in plasma of colorectal cancer patients - a potential biomarker for tumor burden. Surg Oncol 26, 395–401 (2017)
E. Gormally, P. Vineis, G. Matullo, F. Veglia, E. Caboux, E. Le Roux, M. Peluso, S. Garte, S. Guarrera, A. Munnia, L. Airoldi, H. Autrup, C. Malaveille, A. Dunning, K. Overvad, A. Tjønneland, E. Lund, F. Clavel-Chapelon, H. Boeing, A. Trichopoulou, D. Palli, V. Krogh, R. Tumino, S. Panico, H.B. Bueno-de-Mesquita, P.H. Peeters, G. Pera, C. Martinez, M. Dorronsoro, A. Barricarte, C. Navarro, J.R. Quirós, G. Hallmans, N.E. Day, T.J. Key, R. Saracci, R. Kaaks, E. Riboli, P. Hainaut, Tp53 and kras2 mutations in plasma DNA of healthy subjects and subsequent cancer occurrence: A prospective study. Cancer Res 66, 6871–6876 (2006)
L. Mao, R.H. Hruban, J.O. Boyle, M. Tockman, D. Sidransky, Detection of oncogene mutations in sputum precedes diagnosis of lung cancer. Cancer Res 54, 1634–1637 (1994)
S. Salvi, G. Gurioli, U. De Giorgi, V. Conteduca, G. Tedaldi, D. Calistri, V. Casadio, Cell-free DNA as a diagnostic marker for cancer: Current insights. Onco targets Ther 9, 6549–6559 (2016)
I. Martincorena, A. Roshan, M. Gerstung, P. Ellis, P. Van Loo, S. McLaren, D.C. Wedge, A. Fullam, L.B. Alexandrov, J.M. Tubio, L. Stebbings, A. Menzies, S. Widaa, M.R. Stratton, P.H. Jones, P.J. Campbell, High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015)
L. Fernandez-Cuesta, S. Perdomo, P.H. Avogbe, N. Leblay, T.M. Delhomme, V. Gaborieau, B. Abedi-Ardekani, E. Chanudet, M. Olivier, D. Zaridze, A. Mukeria, M. Vilensky, I. Holcatova, J. Polesel, L. Simonato, C. Canova, P. Lagiou, C. Brambilla, E. Brambilla, G. Byrnes, G. Scelo, F. Le Calvez-Kelm, M. Foll, J.D. McKay, P. Brennan, Identification of circulating tumor DNA for the early detection of small-cell lung cancer. EBioMedicine 10, 117–123 (2016)
J.D. Cohen, L. Li, Y. Wang, C. Thoburn, B. Afsari, L. Danilova, C. Douville, A.A. Javed, F. Wong, A. Mattox, R.H. Hruban, C.L. Wolfgang, M.G. Goggins, M. Dal Molin, T.-L. Wang, R. Roden, A.P. Klein, J. Ptak, L. Dobbyn, J. Schaefer, N. Silliman, M. Popoli, J.T. Vogelstein, J.D. Browne, R.E. Schoen, R.E. Brand, J. Tie, P. Gibbs, H.-L. Wong, A.S. Mansfield, J. Jen, S.M. Hanash, M. Falconi, P.J. Allen, S. Zhou, C. Bettegowda, L.A. Diaz, C. Tomasetti, K.W. Kinzler, B. Vogelstein, A.M. Lennon, N. Papadopoulos, Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018)
K. Birkenkamp-Demtroder, I. Nordentoft, E. Christensen, S. Hoyer, T. Reinert, S. Vang, M. Borre, M. Agerbaek, J.B. Jensen, T.F. Orntoft, L. Dyrskjot, Genomic alterations in liquid biopsies from patients with bladder cancer. Eur Urol 70, 75–82 (2016)
F.A. Greco, K. Oien, M. Erlander, R. Osborne, G. Varadhachary, J. Bridgewater, D. Cohen, H. Wasan, Cancer of unknown primary: Progress in the search for improved and rapid diagnosis leading toward superior patient outcomes. Ann Oncol 23, 298–304 (2012)
M.W. Snyder, M. Kircher, A.J. Hill, R.M. Daza, J. Shendure, Cell-free DNA comprises an in vivo nucleosome footprint that informs its tissues-of-origin. Cell 164, 57–68 (2016)
M.R. Openshaw, R.A. Harvey, N.J. Sebire, B. Kaur, N. Sarwar, M.J. Seckl, R.A. Fisher, Circulating cell free DNA in the diagnosis of trophoblastic tumors. EBioMedicine 4, 146–152 (2016)
J.-Y. Douillard, G. Ostoros, M. Cobo, T. Ciuleanu, R. Cole, G. McWalter, J. Walker, S. Dearden, A. Webster, T. Milenkova, R. McCormack, Gefitinib treatment in egfr mutated caucasian nsclc: Circulating-free tumor DNA as a surrogate for determination of egfr status. J Thorac Oncol 9, 1345–1353 (2014)
M.F. Sanmamed, S. Fernandez-Landazuri, C. Rodriguez, R. Zarate, M.D. Lozano, L. Zubiri, J.L. Perez-Gracia, S. Martin-Algarra, A. Gonzalez, Quantitative cell-free circulating brafv600e mutation analysis by use of droplet digital pcr in the follow-up of patients with melanoma being treated with braf inhibitors. Clin Chem 61, 297–304 (2015)
K.L. Spindler, N. Pallisgaard, I. Vogelius, A. Jakobsen, Quantitative cell-free DNA, kras, and braf mutations in plasma from patients with metastatic colorectal cancer during treatment with cetuximab and irinotecan. Clin Cancer Res 18, 1177–1185 (2012)
A.R. Thierry, F. Mouliere, S. El Messaoudi, C. Mollevi, E. Lopez-Crapez, F. Rolet, B. Gillet, C. Gongora, P. Dechelotte, B. Robert, M. Del Rio, P.J. Lamy, F. Bibeau, M. Nouaille, V. Loriot, A.S. Jarrousse, F. Molina, M. Mathonnet, D. Pezet, M. Ychou, Clinical validation of the detection of kras and braf mutations from circulating tumor DNA. Nat Med 20, 430–435 (2014)
A. Santiago-Walker, R. Gagnon, J. Mazumdar, M. Casey, G.V. Long, D. Schadendorf, K. Flaherty, R. Kefford, A. Hauschild, P. Hwu, P. Haney, A. O'Hagan, J. Carver, V. Goodman, J. Legos, A.M. Martin, Correlation of braf mutation status in circulating-free DNA and tumor and association with clinical outcome across four brafi and meki clinical trials. Clin Cancer Res 22, 567–574 (2016)
L. De Mattos-Arruda, B. Weigelt, J. Cortes, H.H. Won, C.K. Ng, P. Nuciforo, F.C. Bidard, C. Aura, C. Saura, V. Peg, S. Piscuoglio, M. Oliveira, Y. Smolders, P. Patel, L. Norton, J. Tabernero, M.F. Berger, J. Seoane, J.S. Reis-Filho, Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: A proof-of-principle. Ann Oncol 25, 1729–1735 (2014)
J.A. García-Saenz, P. Ayllón, M. Laig, D. Acosta-Eyzaguirre, M. García-Esquinas, M. Montes, J. Sanz, M. Barquín, F. Moreno, V. Garcia-Barberan, E. Díaz-Rubio, T. Caldes, A. Romero, Tumor burden monitoring using cell-free tumor DNA could be limited by tumor heterogeneity in advanced breast cancer and should be evaluated together with radiographic imaging. BMC Cancer 17, 210 (2017)
D.J. McBride, A.K. Orpana, C. Sotiriou, H. Joensuu, P.J. Stephens, L.J. Mudie, E. Hämäläinen, L.A. Stebbings, L.C. Andersson, A.M. Flanagan, V. Durbecq, M. Ignatiadis, O. Kallioniemi, C.A. Heckman, K. Alitalo, H. Edgren, P.A. Futreal, M.R. Stratton, P.J. Campbell, Use of cancer-specific genomic rearrangements to quantify disease burden in plasma from patients with solid tumors. Genes Chromosomes Cancer 49, 1062–1069 (2010)
X. Yi, J. Ma, Y. Guan, R. Chen, L. Yang, X. Xia, The feasibility of using mutation detection in ctdna to assess tumor dynamics. Int J Cancer 140, 2642–2647 (2017)
C.A. Parkinson, D. Gale, A.M. Piskorz, H. Biggs, C. Hodgkin, H. Addley, S. Freeman, P. Moyle, E. Sala, K. Sayal, K. Hosking, I. Gounaris, M. Jimenez-Linan, H.M. Earl, W. Qian, N. Rosenfeld, J.D. Brenton, Exploratory analysis of tp53 mutations in circulating tumour DNA as biomarkers of treatment response for patients with relapsed high-grade serous ovarian carcinoma: A retrospective study. PLoS Medicine 13, e1002198 (2016)
S.-J. Dawson, D.W.Y. Tsui, M. Murtaza, H. Biggs, O.M. Rueda, S.-F. Chin, M.J. Dunning, D. Gale, T. Forshew, B. Mahler-Araujo, S. Rajan, S. Humphray, J. Becq, D. Halsall, M. Wallis, D. Bentley, C. Caldas, N. Rosenfeld, Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med 368, 1199–1209 (2013)
B. Ai, H. Liu, Y. Huang, P. Peng, Circulating cell-free DNA as a prognostic and predictive biomarker in non-small cell lung cancer. Oncotarget 7, 44583–44595 (2016)
S. Basnet, Z.-y. Zhang, W.-q. Liao, S.-h. Li, P.-s. Li, H.-y. Ge, The prognostic value of circulating cell-free DNA in colorectal cancer: A meta-analysis. J Cancer 7, 1105–1113 (2016)
Z. Yi, B. Liu, X. Guan, F. Ma, Plasma cell-free DNA and survival in non-small-cell lung cancer: A meta-analysis. Mol Clin Oncol 7, 167–172 (2017)
C. Bedin, M.V. Enzo, P. Del Bianco, S. Pucciarelli, D. Nitti, M. Agostini, Diagnostic and prognostic role of cell-free DNA testing for colorectal cancer patients. International Journal of Cancer 140, 1888–1898 (2017)
J.H. No, K. Kim, K.H. Park, Y.B. Kim, Cell-free DNA level as a prognostic biomarker for epithelial ovarian cancer. Anticancer Res 32, 3467–3471 (2012)
G.R. Oxnard, K.S. Thress, R.S. Alden, R. Lawrance, C.P. Paweletz, M. Cantarini, J.C. Yang, J.C. Barrett, P.A. Janne, Association between plasma genotyping and outcomes of treatment with osimertinib (azd9291) in advanced non-small-cell lung cancer. J Clin Oncol 34, 3375–3382 (2016)
D. Kwapisz, The first liquid biopsy test approved. Is it a new era of mutation testing for non-small cell lung cancer? Ann Transl Med 5, 46 (2017)
O.A. Zill, K.C. Banks, S.R. Fairclough, S. Mortimer, J.V. Vowles, R. Mokhtari, D.R. Gandara, P.C. Mack, J.I. Odegaard, R.J. Nagy, A.M. Baca, H. Eltoukhy, D.I. Chudova, R.B. Lanman, A. Talasaz, The landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin Cancer Res 24, 3528–3538 (2018)
D.K. Lee, J.H. Park, J.H. Kim, S.J. Lee, M.K. Jo, M.C. Gil, K.H. Song, J.W. Park, Progression of prostate cancer despite an extremely low serum level of prostate-specific antigen. Korean J Radiol 51, 358–361 (2010)
A.H. Bryce, J.J. Alumkal, A. Armstrong, C.S. Higano, P. Iversen, C.N. Sternberg, D. Rathkopf, Y. Loriot, J. de Bono, B. Tombal, S. Abhyankar, P. Lin, A. Krivoshik, D. Phung, T.M. Beer, Radiographic progression with nonrising psa in metastatic castration-resistant prostate cancer: Post hoc analysis of prevail. Prostate Cancer Prostatic Dis 20, 221–227 (2017)
T.A. Stamey, N. Yang, A.R. Hay, J.E. McNeal, F.S. Freiha, E. Redwine, Prostate-specific antigen as a serum marker for adenocarcinoma of the prostate. New Eng J Med 317, 909–916 (1987)
J. Lokich, S. Ellenberg, B. Gerson, W.E. Knox, N. Zamcheck, Plasma clearance of carcinoembryonic antigen following hepatic metastatectomy. J Clin Oncol 2, 462–465 (1984)
E.S. Gray, H. Rizos, A.L. Reid, S.C. Boyd, M.R. Pereira, J. Lo, V. Tembe, J. Freeman, J.H. Lee, R.A. Scolyer, K. Siew, C. Lomma, A. Cooper, M.A. Khattak, T.M. Meniawy, G.V. Long, M.S. Carlino, M. Millward, M. Ziman, Circulating tumor DNA to monitor treatment response and detect acquired resistance in patients with metastatic melanoma. Oncotarget 6, 42008–42018 (2015)
S.-J. Dawson, D.W.Y. Tsui, M. Murtaza, H. Biggs, O.M. Rueda, S.-F. Chin, M.J. Dunning, D. Gale, T. Forshew, B. Mahler-Araujo, S. Rajan, S. Humphray, J. Becq, D. Halsall, M. Wallis, D. Bentley, C. Caldas, N. Rosenfeld, Analysis of circulating tumor DNA to monitor metastatic breast cancer. New Eng J Med 368, 1199–1209 (2013)
A. Ashida, K. Sakaizawa, H. Uhara, R. Okuyama, Circulating tumour DNA for monitoring treatment response to anti-pd-1 immunotherapy in melanoma patients. Acta Derm Venereol 97, 1212–1218 (2017)
J. Tie, I. Kinde, Y. Wang, H.L. Wong, J. Roebert, M. Christie, M. Tacey, R. Wong, M. Singh, C.S. Karapetis, J. Desai, B. Tran, R.L. Strausberg, L.A. Diaz Jr., N. Papadopoulos, K.W. Kinzler, B. Vogelstein, P. Gibbs, Circulating tumor DNA as an early marker of therapeutic response in patients with metastatic colorectal cancer. Ann Oncol 26, 1715–1722 (2015)
L. Xi, T.H. Pham, E.C. Payabyab, R.M. Sherry, S.A. Rosenberg, M. Raffeld, Circulating tumor DNA as an early indicator of response to t-cell transfer immunotherapy in metastatic melanoma. Clin Cancer Res 22, 5480–5486 (2016)
G. Housman, S. Byler, S. Heerboth, K. Lapinska, M. Longacre, N. Snyder, S. Sarkar, Drug resistance in cancer: An overview. Cancers 6, 1769–1792 (2014)
E. Kirkizlar, B. Zimmermann, T. Constantin, R. Swenerton, B. Hoang, N. Wayham, J.E. Babiarz, Z. Demko, R.J. Pelham, S. Kareht, A.L. Simon, K.N. Jinnett, M. Rabinowitz, S. Sigurjonsson, M. Hill, Detection of clonal and subclonal copy-number variants in cell-free DNA from patients with breast cancer using a massively multiplexed pcr methodology. Transl Oncol 8, 407–416 (2015)
J.S. Frenel, S. Carreira, J. Goodall, D. Roda, R. Perez-Lopez, N. Tunariu, R. Riisnaes, S. Miranda, I. Figueiredo, D. Nava-Rodrigues, A. Smith, C. Leux, I. Garcia-Murillas, R. Ferraldeschi, D. Lorente, J. Mateo, M. Ong, T.A. Yap, U. Banerji, D. Gasi Tandefelt, N. Turner, G. Attard, J.S. de Bono, Serial next-generation sequencing of circulating cell-free DNA evaluating tumor clone response to molecularly targeted drug administration. Clin Cancer Res 21, 4586–4596 (2015)
G. Siravegna, B. Mussolin, M. Buscarino, G. Corti, A. Cassingena, G. Crisafulli, A. Ponzetti, C. Cremolini, A. Amatu, C. Lauricella, S. Lamba, S. Hobor, A. Avallone, E. Valtorta, G. Rospo, E. Medico, V. Motta, C. Antoniotti, F. Tatangelo, B. Bellosillo, S. Veronese, A. Budillon, C. Montagut, P. Racca, S. Marsoni, A. Falcone, R.B. Corcoran, F. Di Nicolantonio, F. Loupakis, S. Siena, A. Sartore-Bianchi, A. Bardelli, Clonal evolution and resistance to egfr blockade in the blood of colorectal cancer patients. Nat Med 21, 827 (2015)
R. Lebofsky, C. Decraene, V. Bernard, M. Kamal, A. Blin, Q. Leroy, T. Rio Frio, G. Pierron, C. Callens, I. Bieche, A. Saliou, J. Madic, E. Rouleau, F.-C. Bidard, O. Lantz, M.-H. Stern, C. Le Tourneau, J.-Y. Pierga, Circulating tumor DNA as a non-invasive substitute to metastasis biopsy for tumor genotyping and personalized medicine in a prospective trial across all tumor types. Mol Oncol 9, 783–790 (2015)
K.S. Thress, R. Brant, T.H. Carr, S. Dearden, S. Jenkins, H. Brown, T. Hammett, M. Cantarini, J.C. Barrett, Egfr mutation detection in ctdna from nsclc patient plasma: A cross-platform comparison of leading technologies to support the clinical development of azd9291. Lung Cancer 90, 509–515 (2015)
B.S. Sorensen, L. Wu, W. Wei, J. Tsai, B. Weber, E. Nexo, P. Meldgaard, Monitoring of epidermal growth factor receptor tyrosine kinase inhibitor-sensitizing and resistance mutations in the plasma DNA of patients with advanced non-small cell lung cancer during treatment with erlotinib. Cancer 120, 3896–3901 (2014)
G.R. Oxnard, C.P. Paweletz, Y. Kuang, S.L. Mach, A. O'Connell, M.M. Messineo, J.J. Luke, M. Butaney, P. Kirschmeier, D.M. Jackman, P.A. Janne, Noninvasive detection of response and resistance in egfr-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res 20, 1698–1705 (2014)
A.W. Wyatt, A.A. Azad, S.V. Volik, M. Annala, K. Beja, B. McConeghy, A. Haegert, E.W. Warner, F. Mo, S. Brahmbhatt, R. Shukin, S. Le Bihan, M.E. Gleave, M. Nykter, C.C. Collins, K.N. Chi, Genomic alterations in cell-free DNA and enzalutamide resistance in castration-resistant prostate cancer. JAMA Oncol 2, 1598–1606 (2016)
M. Annala, G. Vandekerkhove, D. Khalaf, S. Taavitsainen, K. Beja, E.W. Warner, K. Sunderland, C. Kollmannsberger, B.J. Eigl, D. Finch, C.D. Oja, J. Vergidis, M. Zulfiqar, A.A. Azad, M. Nykter, M.E. Gleave, A.W. Wyatt, K.N. Chi, Circulating tumor DNA genomics correlate with resistance to abiraterone and enzalutamide in prostate cancer. Cancer Discov 8, 444–457 (2018)
G. Gremel, R.J. Lee, M.R. Girotti, A.K. Mandal, S. Valpione, G. Garner, M. Ayub, S. Wood, D.G. Rothwell, A. Fusi, A. Wallace, G. Brady, C. Dive, N. Dhomen, P. Lorigan, R. Marais, Distinct subclonal tumour responses to therapy revealed by circulating cell-free DNA. Ann Oncol 27, 1959–1965 (2016)
L.A. Diaz, R. Williams, J. Wu, I. Kinde, J.R. Hecht, J. Berlin, B. Allen, I. Bozic, J.G. Reiter, M.A. Nowak, K.W. Kinzler, K.S. Oliner, B. Vogelstein, The molecular evolution of acquired resistance to targeted egfr blockade in colorectal cancers. Nature 486, 537–540 (2012)
J. Tie, Y. Wang, C. Tomasetti, L. Li, S. Springer, I. Kinde, N. Silliman, M. Tacey, H.L. Wong, M. Christie, S. Kosmider, I. Skinner, R. Wong, M. Steel, B. Tran, J. Desai, I. Jones, A. Haydon, T. Hayes, T.J. Price, R.L. Strausberg, L.A. Diaz Jr., N. Papadopoulos, K.W. Kinzler, B. Vogelstein, P. Gibbs, Circulating tumor DNA analysis detects minimal residual disease and predicts recurrence in patients with stage ii colon cancer. Sci Transl Med 8, 346ra392 (2016)
L. Kubiczkova-Besse, D. Drandi, L. Sedlarikova, S. Oliva, M. Gambella, P. Omedè, Z. Adam, L. Pour, S. Sevcikova, M. Boccadoro, A. Palumbo, R. Hajek, Cell-free DNA for minimal residual disease monitoring in multiple myeloma patients. Blood 124, 3423 (2014)
L.M. McShane, D.G. Altman, W. Sauerbrei, S.E. Taube, M. Gion, G.M. Clark, REporting recommendations for tumour MARKer prognostic studies (REMARK). Br J Cancer 22, 387–391 (2005)
D.G. Altman, L.M. McShane, W. Sauerbrei, S.E. Taube, Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK): explanation and elaboration. PLoS Med 9, e1001216 (2012)
D.S. Tan, G.V. Thomas, M.D. Garrett, U. Banerji, J.S. de Bono, S.B. Kaye, P. Workman, Biomarker-driven early clinical trials in oncology: a paradigm shift in drug development. Cancer J 15, 406–420 (2009)
J. Cummings, F. Raynaud, L. Jones, R. Sugar, C. Dive, Fit-for-purpose biomarker method validation for application in clinical trials of anticancer drugs. Br J Cancer 26, 1313–1317 (2010)
D.L. Jardim, M. Schwaederle, D.S. Hong, R. Kurzrock, An appraisal of drug development timelines in the Era of precision oncology. Oncotarget 16, 53037–53046 (2016)
E. Heitzer, P. Ulz, J. Belic, S. Gutschi, F. Quehenberger, K. Fischereder, T. Benezeder, M. Auer, C. Pischler, S. Mannweiler, M. Pichler, F. Eisner, M. Haeusler, S. Riethdorf, K. Pantel, H. Samonigg, G. Hoefler, H. Augustin, J.B. Geigl, M.R. Speicher, Tumor-associated copy number changes in the circulation of patients with prostate cancer identified through whole-genome sequencing. Genome Med 5, 30 (2013)
R.J. Leary, M. Sausen, I. Kinde, N. Papadopoulos, J.D. Carpten, D. Craig, J. O'Shaughnessy, K.W. Kinzler, G. Parmigiani, B. Vogelstein, L.A. Diaz Jr., V.E. Velculescu, Detection of chromosomal alterations in the circulation of cancer patients with whole-genome sequencing. Sci Transl Med 4, 162ra154 (2012)
J.A. Vendrell, F.T. Mau-Them, B. Béganton, S. Godreuil, P. Coopman, J. Solassol, Circulating cell free tumor DNA detection as a routine tool for lung cancer patient management. Int J Mol Sci 18, 264 (2017)
R. Salani, C.-L. Chang, L. Cope, T.-L. Wang, Digital karyotyping: An update of its applications in cancer. Mol Diagn Ther 10, 231 (2006)
M. Murtaza, S.J. Dawson, D.W. Tsui, D. Gale, T. Forshew, A.M. Piskorz, C. Parkinson, S.F. Chin, Z. Kingsbury, A.S. Wong, F. Marass, S. Humphray, J. Hadfield, D. Bentley, T.M. Chin, J.D. Brenton, C. Caldas, N. Rosenfeld, Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature 497, 108–112 (2013)
C. Vnencak-Jones, M. Berger, W. Pao. Types of molecular tumor testing (2016)
A. Narayan, N.J. Carriero, S.N. Gettinger, J. Kluytenaar, K.R. Kozak, T.I. Yock, N.E. Muscato, P. Ugarelli, R.H. Decker, A.A. Patel, Ultrasensitive measurement of hotspot mutations in tumor DNA in blood using error-suppressed multiplexed deep sequencing. Cancer Res 72, 3492–3498 (2012)
D. Gale, V. Plagnol, A. Lawson, M. Pugh, S. Smalley, K. Howarth, M. Madi, B. Durham, V. Kumanduri, K. Lo, J. Clark, E. Green, N. Rosenfeld, T. Forshew, Abstract 3639: Analytical performance and validation of an enhanced tam-seq circulating tumor DNA sequencing assay. Cancer Res 76, 3639–3639 (2016)
A.M. Newman, A.F. Lovejoy, D.M. Klass, D.M. Kurtz, J.J. Chabon, F. Scherer, H. Stehr, C.L. Liu, S.V. Bratman, C. Say, L. Zhou, J.N. Carter, R.B. West, G.W. Sledge, J.B. Shrager, B.W. Loo Jr., J.W. Neal, H.A. Wakelee, M. Diehn, A.A. Alizadeh, Integrated digital error suppression for improved detection of circulating tumor DNA. Nature Biotechnol 34, 547–555 (2016)
R.B. Lanman, S.A. Mortimer, O.A. Zill, D. Sebisanovic, R. Lopez, S. Blau, E.A. Collisson, S.G. Divers, D.S. Hoon, E.S. Kopetz, J. Lee, P.G. Nikolinakos, A.M. Baca, B.G. Kermani, H. Eltoukhy, A. Talasaz, Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PloS One 10, e0140712 (2015)
F. Rothe, J.F. Laes, D. Lambrechts, D. Smeets, D. Vincent, M. Maetens, D. Fumagalli, S. Michiels, S. Drisis, C. Moerman, J.P. Detiffe, D. Larsimont, A. Awada, M. Piccart, C. Sotiriou, M. Ignatiadis, Plasma circulating tumor DNA as an alternative to metastatic biopsies for mutational analysis in breast cancer. Ann Oncol 25, 1959–1965 (2014)
H.R. Kim, S.Y. Lee, D.S. Hyun, M.K. Lee, H.K. Lee, C.M. Choi, S.H. Yang, Y.C. Kim, Y.C. Lee, S.Y. Kim, S.H. Jang, J.C. Lee, K.Y. Lee, Detection of egfr mutations in circulating free DNA by pna-mediated pcr clamping. J Exp Clin Cancer Res 32, 50 (2013)
D. Sefrioui, F. Mauger, L. Leclere, L. Beaussire, F. Di Fiore, J.-F. Deleuze, N. Sarafan-Vasseur, J. Tost, Comparison of the quantification of kras mutations by digital pcr and e-ice-cold-pcr in circulating-cell-free DNA from metastatic colorectal cancer patients. Clinica Chimica Acta 465, 1–4 (2017)
K. Watanabe, T. Fukuhara, Y. Tsukita, M. Morita, A. Suzuki, N. Tanaka, H. Terasaki, T. Nukiwa, M. Maemondo, Egfr mutation analysis of circulating tumor DNA using an improved pna-lna pcr clamp method. Can Respir J 2016, 1–7 (2016)
K. Taniguchi, J. Uchida, K. Nishino, T. Kumagai, T. Okuyama, J. Okami, M. Higashiyama, K. Kodama, F. Imamura, K. Kato, Quantitative detection of egfr mutations in circulating tumor DNA derived from lung adenocarcinomas. Clin Cancer Res 17, 7808–7815 (2011)
T.K. Yung, K.C. Chan, T.S. Mok, J. Tong, K.F. To, Y.M. Lo, Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital pcr in non-small cell lung cancer patients. Clin Cancer Res 15, 2076–2084 (2009)
X. Chai, P. Ren, B. Wei, J. Ma, L. Mai, D.S. Cram, Y. Song, Y. Guo, A comparative study of egfr oncogenic mutations in matching tissue and plasma samples from patients with advanced non-small cell lung carcinoma. Clinica Chimica Acta 457, 106–111 (2016)
J. Luo, L. Shen, D. Zheng, Diagnostic value of circulating free DNA for the detection of egfr mutation status in nsclc: A systematic review and meta-analysis. Sci Rep 4, 6269 (2014)
E. Kidess, K. Heirich, M. Wiggin, V. Vysotskaia, B.C. Visser, A. Marziali, B. Wiedenmann, J.A. Norton, M. Lee, S.S. Jeffrey, G.A. Poultsides, Mutation profiling of tumor DNA from plasma and tumor tissue of colorectal cancer patients with a novel, high-sensitivity multiplexed mutation detection platform. Oncotarget 6, 2549–2561 (2015)
I. Ladas, M. Fitarelli-Kiehl, C. Song, V.A. Adalsteinsson, H.A. Parsons, N.U. Lin, N. Wagle, G.M. Makrigiorgos, Multiplexed elimination of wild-type DNA and high-resolution melting prior to targeted resequencing of liquid biopsies. Clin Chem 63, 1605–1613 (2017)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Fettke, H., Kwan, E.M. & Azad, A.A. Cell-free DNA in cancer: current insights. Cell Oncol. 42, 13–28 (2019). https://doi.org/10.1007/s13402-018-0413-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-018-0413-5