Abstract
Novel laccase-producing thermo-tolerant organisms were identified and its culture conditions optimized for effective delignification. Forty-three isolates were screened for ligninolytic activity, out of which six isolates with higher laccase specific activity (3.27, 5.49, 2.26, 4.54, 4.61, and 4.73 U mg−1 protein) were selected for further analysis. Maximum enzyme production of 450 U/μL was obtained after 120 hours of incubation, at 46°C and pH 6.0. Erianthus bagasse digested with crude enzymes was studied for their structural modifications. The SEM analysis showed cavities (1.28 to 2.56 μm) indicating the disintegration of lignin molecules while the FT-IR showed peaks at 1509 cm−1, 1725 cm−1, and 2894 cm−1 indicating the breakdown of lignin, hemicellulose, and cellulose moieties, respectively. The GC-MS analysis generated derivatives of syringyl and hydroxyphenyl lignin. Sequence analysis of the isolate showed that the isolate S2 is affiliated to A. caespitosus and its purified laccase showed a specific enzyme activity of 7.9 to 19.2 U/mg of protein. Our results suggest that Aspergillus caespitosus S2 has the highest potential for Erianthus delignification. Aspergillus caespitosus and its system biology generated from this study might be beneficial for bio-refinery applications.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
In the bio-energy platform, lignocellulosic biomass plays a pivotal role as feedstock for bio-refineries to replace the fossil fuel demand. Lignocellulosic biomass (LCB) is classified into agricultural, forestry residues, energy crops, and cellulosic wastes. Globally, around 5300 million tons per year of agricultural residues are readily available as feedstock [1]. Erianthus arundinaceus (sweet cane) is a wild cane species recognized as a potential bioenergy crop due to its huge biomass yield with high fiber and lignin content [2, 3]. LCB is composed of cellulose, hemicelluloses, intertwined by complexes of lignin. Lignin are aromatic cross polymers of 4 hydroxyl phenylpropanoid monomers linked by carbon-carbon and ether bonds to form a rigid plant structure [1, 4, 5]. This rigid structure makes lignin recalcitrant to biomass conversion during biofuel production and requires pretreatment to remove considerable lignin.
Pretreatment methods for releasing the polysaccharides for biofuel production can be classified into non-biological and biological processes. Enzymatic pretreatment methods are biological, eco-friendly approaches, compared to physicochemical methods. Microorganisms capable of producing lignin-degrading enzymes are exemplified by the white-rot fungi, as well as the bacteria and the archaeon Haloferax volcanii [1, 6, 7]. The lignin components in the lignocellulosic biomass are decomposed by ligninolytic enzymes such as laccase, lignin peroxidases, polyphenol oxidases, and manganese peroxidases [8,9,10]. Enzymes such as laccase catalyze the oxidation of aromatic thiols, phenols, and anilines present in the biomass. Nevertheless, biological pretreatment are affected by parameters like substrate, types of microorganism, incubation temperature, aeration rate, and moisture content, and these need to be optimized in the process [11]. Among the ligninolytic enzymes, laccase (EC 1.10.3.2) is a multi-copper oxidase enzyme with huge applicability in various biotechnological process, such as bioremediation, biofuels, fiber modified, biosensor, textile, and paper industries [12, 13]. In particular, the textile dyestuffs which were 10–50% highly reactive dyes were discharged in the water bodies by the industrial scale and azo dyes were predominantly found in it and they form toxic aromatic amines and phenolic. The lignin-degrading enzymes such as laccase and peroxidase play pivotal role to cleave this aromatic azo dyes by their oxido-reductive enzymatic mechanism [14]. Due to this wider applicability, laccase enzymes are at huge demand.
Hence, to enhance the laccase production at low cost, optimization of fermentation conditions, methods, and developing new productivity strains are effective methods. Many researchers have used various lignocellulosic wastes, such as coffee shell, cotton seed hull, corncob, cottonseed hull, bamboos, rice straw, and dried leaves, to investigate the production of ligninolytic enzymes at low cost [15, 16]. The different biomass substrates such as birch, alder, aspen from saw dust, and wheat bran were formulated and utilized as lignocellulosic feedstock by Lentinula edodes. The delignification of these combined feedstock corresponds to the degradation of S-lignin unit and reduction of S-G ratio [17].
Sugarcane compost pits/piles are one of the most abundant agricultural wastes, under-exploited, which provide a unique ecological niche, for the identification of ligninolytic microbes that can thrive in relatively high temperatures (45–50°C). Response surface methodology (RSM) is a tool which uses statistical and mathematical technique to optimize the process parameters and their output responses. It aids to design the experiments with minimal trials, to correlate the input and output responses and reduce the time, material, and cost [18]. In this study, we exploited the microbial community in the sugarcane compost pit for the identification and characterization of new thermo-tolerant lignin-degrading isolates. We report the optimization of process variables along with their interaction for effective delignification using RSM. The sequence analysis of the high laccase producing isolate is affiliated to A. caespitosus. Erianthus biomass structural modifications after laccase extracted from A. caespitosus have been documented through scanning electron microscope, Fourier-transform infrared spectroscopy, gas chromatography-mass spectrometry, and degradation of aromatic azo dye. In this study, novel thermo-tolerant Aspergillus caespitosus has been screened and the crude laccase enzyme extracted from the isolate has been used for the digestion of Erianthus bagasse. We have identified that the purified laccase showed excellent specific enzyme activity and, therefore, suggesting that Aspergillus caespitosus has a high potential for Erianthus bagasse delignification. Our results contribute to providing new isolate that holds promise for improved production and activity of laccase enzymes at low cost.
2 Methods
The compost pit samples and Erianthus bagasse from a matured plant (11 months old) were collected from ICAR Sugarcane Breeding Institute, Coimbatore. The chemicals and reagents of molecular grade used for enzyme activity studies were purchased from Hi-media Laboratories (Mumbai, India). PCR reagents, Taq DNA polymerase, and oligonucleotide primers were obtained from Thermo Fisher Scientific, USA. The Trametes versicolor laccase enzyme was procured from Sigma-Aldrich (Merck, Germany).
2.1 Biomass preparation
The pith and rind portion of the Erianthus arundinaceus stems (clone IK-76-81) were removed manually. The juice was extracted, and the bagasse was chopped into smaller pieces. The chopped bagasse material was shade dried for 3 days and ground in a pulverizer machine (model: SF 250, China). The resulting powder was sieved to 200 μm sizes [19].
2.2 Isolation of lignin-degrading fungi
Thermo-tolerant microbes were isolated from a sugarcane compost pit located at an ICAR Sugarcane Breeding Institute farm (11°0′20″N and 76°54′54″E). The natural compost had been incubated for approximately 14 days at 37–45°C. The thermo-tolerant microbes were isolated from the pit by a serial dilution and plating technique. In this method, compost samples (1 g each) were added to a 250 mL Erlenmeyer flask containing 100 mL of minimal liquid medium with 0.1 Mm CuSO4 for 5 days at 46°C in a rotating shaker of 150 rpm. From the liquid medium, 1 mL of sample was removed and serially diluted from 10−1 to 10−4. After the serial dilution, 100 μL dilution was spread on plates containing minimal agar medium. The plates were incubated at 46°C for five days. The minimal medium, pH 6, was composed per liter of 1 g dextrose, 5 g NaCl, 1 g K2HPO4, 1 g NH4H2PO4, 1 g (NH4)2SO4, 0.2 g MgSO4·7H2O, 3 g KNO3, and 20 g agar, as previously described [20]. The medium was sterilized by autoclaving at 121°C for 1 h prior to use. To obtain pure cultures of the microorganisms, colonies from the mother plate were further isolated by the “zigzag” streaking method, and most of the cultures were isolated from the 10−2 and 10−3 dilutions.
2.3 Screening of laccase-producing fungi
To identify the isolates that produced laccase activity, ABTS (2,2′-azinobis-3-ethyl-benzothiozoline-6 sulfonic acid) was used as a substrate. The isolated strains were grown in a 500 mL baffled Erlenmeyer flask with 50 mL mineral medium along with 0.1 mM CuSO4 for five days at 46°C. The culture was incubated in an orbital shaker at 150 rpm. After five days of growth, the cultures were centrifuged at 6700 × g for 10 min. The resulting culture broth was mixed in a micro titer plate containing 70 μL of 100 mM sodium acetate buffer pH 5.0 and 30 μL of ABTS pipetted into 200 μL of reaction mixture for a final concentration of 5 mM ABTS substrate with final pH of 5. Laccase activity was monitored by a change in absorbance at 420 nm (∆A) over a time of 10 min using a SpectraMax micro plate reader (Molecular Devices, San Jose, CA) and calculated using the following formula:
∆A is the final absorbance value subtracted from the initial absorbance value of the reaction; Ve is the enzyme volume used in the reaction expressed in mL; DF is the dilution factor of the crude enzyme preparation in the culture broth expressed in mL; t is the time of incubation with the unit of minute; ε is the extinction coefficient of 36,000 M−1 cm−1 at 420 nm for the ABTS substrate; l is the path length expressed as cm; and Vr is the total reaction volume with the unit of mL.
2.4 Optimization of laccase production using response surface methodology (RSM)
The response surface methodology using the Box–Behnken design was used to determine the optimum concentration of the selected variables. The variables selected were temperature, pH, inoculum size, and incubation period. A total of 27 experiments were formulated using the statistical software package Design Expert® 13.0 (STAT-EASE, Inc., Minneapolis, MN) [21, 22]. Each selected variable was analyzed at three levels, namely, low, medium, and high coded as −1, 0, and +1 in total of 27 runs (Supplementary Table S1).
2.5 Protein content of crude enzyme
To determine the total protein content of the crude enzymes, the Bradford protein assay was used with bovine serum albumin (BSA) as the standard. The specific activity of the crude enzyme was calculated according to the ratio of volumetric activity to the amount of protein (mg) per mL of culture broth [23].
2.6 Analysis of Erianthus bagasse material
The crude preparations of enzymes from the novel fungal isolates were examined for their influence on the structural composition and degradation of the lignin within the bagasse material. The commercial laccase purified from Trametes versicolor was included for comparison. The biomass slurry was prepared by mixing 1 mL of enzyme with 2 g bagasse in 40 mL of 100 mM sodium acetate buffer at pH 4.5 [18]. The slurry was incubated at 46°C in a 250 mL baffled Erlenmeyer flask for 15 h with orbital shaking at 150 rpm. After incubation, the pretreated bagasse was separated from the supernatant using a muslin cloth (200 μm size). The pretreated material was washed thrice with distilled water (500 mL) and was dried in a hot air oven at 45°C.
2.6.1 Scanning electron microscopy to examine surface morphology
The raw and pretreated biomass materials (treated with the commercial enzyme and the crude enzymes from six isolates identified in this study (S1, S2, S3, S6, S8, and S10)) were analyzed for surface morphology by scanning electron microscopy (SEM; Quanta 250, FEI, Hillsboro, OR, USA).
2.6.2 Fourier-transform infrared spectrometry to assess the functional group
To identify the functional group and structure of compounds within the raw and pretreated biomass samples, the absorption and molecular stretching of the compounds were determined by Fourier-transform infrared spectrometry (FT-IR 6800 JASCO, Japan). In brief, a resolution of 4 cm−1 was used for the analysis with a scanning speed of 2 mm/sec. The absorbance band was documented from the range of 4000–400 cm−1.
2.6.3 Gas chromatography and mass spectroscopy to identify molecular compounds
After digestion of the samples, the biomass was analyzed for lignin degradation by gas chromatography-mass spectrometry (GC-MS) using a DB-5 MS capillary standard nonpolar column (PerkinElmer Clarus, USA) according to [24]. Metabolites identified with high probability and peak area were chosen along with their component name and retention time, for the generation of heat map using ORIGIN 2021b software (Origin Lab, Northampton, MA).
2.7 Residual enzyme activity
The supernatant of the biomass slurry that was pretreated with the enzymes (commercial laccase and crude enzyme preparations from six isolates identified in this study) was collected after passage through a muslin cloth. This residual material was assayed for laccase activity.
2.8 Screening isolates for dye degradation
The fungal strains that produced laccase activity were further screened for azo dye degradation. The azo dyes (malachite green 50 mg L−1 w/v) were incorporated into the PD plates. The cultures were inoculated by spread plating onto PD plates supplemented with the azo dyes and 0.1 mM CuSO4 at 46°C [14, 25].
2.9 Molecular characterization of isolates through ITS primers and DNA sequencing
To determine the taxonomic phyla of the fungi, the genomic DNA was isolated from all the six isolates using a cetyltrimethylammonium bromide (CTAB) method according to (Karakousis et al. [26]) and used as template in a polymerase chain reaction (PCR). The PCR mixture consisted of forward and reverse ITS primers as ITS-1F 5′-CTTGGTCATTTAGAGGAAGTAA-3′ and ITS-4R 5′-TCCTCCGCTTATTGATATGC-3′) 10 μM each, 5 μL of 10x Taq buffer with MgCl2, 2.0 μL of 2 mM dNTPs, 1.0 μL of Taq DNA polymerase (3 U μL−1), and genomic DNA (150 ng) at a final volume of 50 μL. The PCR conditions were 94°C for 3 min (initial denaturation); 34 cycles of 94°C for 1 min (denaturation), 50°C for 1 min (primer annealing to template), and 72°C for 1 min (elongation); and a final extension at 72°C for 10 min. The amplified products were separated by 1% (w/v) agarose gel and documented in G:box EF gel documentation system, Syngene, USA. The PCR product of all the isolates was sequenced using Applied Biosystems Sanger sequencing 3500 Genetic Analyzer (Thermo Fisher Scientific, USA) for further identification of the taxa. Phylogeny analysis was performed with using the molecular evolutionary genetics analysis MEGA11 (https://www.megasoftware.net/).
2.10 Enrichment of laccase enzymes from culture broth and SDS-PAGE analysis
To enrich the laccase enzymes from the culture broth, the fungal isolates were grown in 500 mL baffled Erlenmeyer flask with 50 mL mineral medium for five days at 46°C with orbital shaking at 150 rpm. After the five days of growth, the cultures were centrifuged at 15,000 × g for 20 min at 4°C, filtered with Whatman filter of 3MM size, and precipitated with 1 volume of ice-cold ethanol. The protein was collected by centrifugation at 7000 × g for 10 min at 4°C and suspended in 25 mM Tris-HCl buffer pH 8.4 as a crude protein sample as previously described [27]. The samples were separated based on their molecular weight by 12% reducing SDS-PAGE according to [28]. After electrophoresis, the protein bands were visualized in the gel by Coomassie Blue R-250 staining according to [29] and documented.
3 Results
3.1 Isolation and identification of thermo-tolerant lignin-degrading isolates
A total of forty-three fungal isolates were isolated from sugarcane compost pit and screened for ligninolytic activity using ABTS as chromogenic substrate according to [30] (Fig. S1). The isolates were designated as S1 to S43. Out of the 43 isolates screened, six isolates (S1, S2, S3, S6, S8, and S10) showed an increased level of laccase enzyme activity on the fifth day varying from 0.36 to 0.46 U mL−1 (Fig. 1). The specific activity of these isolates (S1, S2, S3, S6, S8, and S10) was 3.27, 5.49, 2.26, 4.54, 4.61, and 4.73 U mg−1 of protein, respectively. Based on this result, these six isolates were selected for further analysis. By comparison, the commercial laccase enzyme of T. versicolor was determined to have a specific activity of 6 U mg−1.
Laccase activities of the crude enzymes from six isolates (S1, S2, S3, S6, S8, and S10) on days 1, 3, 5, and 7 compared with the commercial laccase enzyme. Error bars represent standard deviation. Student t-test to determine statistical significance of day 5 vs. n = x biological replicates. Data are presented as the mean ± SD (n=3) with degree of freedom (DF) of 2, sum of squares (SS) 149.2, and the mean square (MS) of 74.6 with the statistically significant p value (p < 0.05)
3.2 Optimization of laccase production by the Box–Behnken design
The BBD tool of RSM generated a matrix of 27 experimental runs, each having a combination of the four selected parameters (pH (A), temperature (B), inoculation period (C), and inoculation size (D)) for the optimization of laccase production. The Box–Behnken design matrix for laccase production optimization is presented in Table S1. The response surface plots were generated by keeping the variable at optimal level and other variables at experimental range. The interactions between the model terms (AB, AC, AD, BC, BD, and CD) were expressed as three-dimensional surface models (Figs. 2a–f). The results of the second-order response surface model fitting in the form of analysis of variance (ANOVA) are given in Table S2. Here, the quadratic model of enzyme activity had a predicted and adjusted R2 value of 0.98 and 0.97, respectively. The predicted R2 was in agreement with the adjusted R2, since the difference between the two values was less than 0.2, indicating that the model developed will be able to give a good estimate of the response (laccase enzyme activity). The CV% of 2.11 clearly indicated a very high degree of precision and a good reliability of the experimental values due to closeness of experimental and predicted values. The adequate precision measures the signal to noise ratio and the value greater than 4 which is desirable and indicates adequate model discrimination and implies greater predicted response relative associated error. Hence, in the present study, the proposed model was found to be statistically significant. The optimized results of 3D plot term AD show that the variables at pH of 6, temperature of 46°C, inoculation period of 5 days, and inoculation size of 8 mm provide the highest laccase production of 450 U/μL (Fig. 2c).
3.3 Biomass preparation and analysis
Biomass prepared from Erianthus clone IK-76-81 (Figs. S2A, B) was examined for enzymatic pretreatment. The crude extracellular fractions of these isolates S1, S2, S3, S6, S8, and S10 were used as the enzymatic solutions for the pretreatment of the Erianthus bagasse material. A sample was digested with the purified laccase from T. versicolor as a positive control, and a sample was processed similarly without enzyme as negative control. The biomass was treated overnight with the enzyme preparations (Fig. S2C). The resultant biomass slurry was washed with distilled water to neutralize the pH and dried (Fig. S2D). Further, the biomass samples were analyzed through SEM, FT-IR, and GC-MS.
3.3.1 Biomass surface alterations detected by scanning electron microscope
The physical surface structure of untreated and pretreated Erianthus bagasse was analyzed through SEM. The surface of the untreated Erianthus bagasse (negative control) was smooth with the fiber bundles intact, and we observed slight peel-off due to the overnight digestion (Fig. 3a). The surface of the biomass samples treated with the commercial laccase enzyme (6 U mL−1) had holes and pinholes (Fig. 3b), while the biomass treated with the crude enzyme preparations from the six isolates (S1, S2, S3, S6, S8, and S10) also had numerous pin holes with a range of 1.28 to 2.56 μm and cavities on the surface of the material (Figs. 3c–h) suggesting the action of the enzyme on the biomass disintegration. The integrity of the Erianthus bagasse surface was partially destroyed, forming various pits or holes. These changes indicated that different degrees of degradation occurred on the surface of biomass due to the efficient degradation of lignocellulose caused by the enzymes secreted by these isolates. Likewise, the surface morphology of the Erianthus bagasse treated with the enzymatic extract from S2 isolate showed wider pores (Fig. 3d) indicating the efficiency of the enzyme. These effects are due to the partial removal of lignin, thus exposing cellulose and hemicellulose for accessibility and which in turn leads to increased yield of fermentable sugars. The structural changes and compositional variation were visualized under the SEM after the treatment, and they were further analyzed by FT-IR and GC-MS to assure it was a removal of lignin content from the bagasse which leads to the holes and pinholes [31].
SEM analysis at 5000x magnification of the Erianthus bagasse after enzymatic treatment. (a) Control (without enzyme), (b) digested with commercial laccase, and (c–h) digested with crude enzyme preparations of S1, S2, S3, S6, S8, and S10, respectively. Representative images from n = 3 independent experiments
3.3.2 Structural changes analyzed through Fourier-transform infrared
FT-IR analysis is used to characterize the biomass according to its organic groups, and its assay is mainly associated with lignin modifications [32]. The Erianthus bagasse pretreated with the crude and commercial enzymes was analyzed by FT-IR to determine the stretching of bonds within the lignin components. The spectrum peaks detected in the raw bagasse material served as a control to compare with the enzymatically treated biomass. The reduced and weakened peak intensity was observed at 1509 cm−1. The peaks at the 1450–1300 cm−1 region exhibited molecular coupling super positions around 1458 cm−1 that are associated with the deformation of lignin CH2-CH3 linkages and included spectra linked to the alcohol stretching bands for phenol or tertiary alcohols [33]. The peak observed at 2894 cm−1 can be linked to an asymmetric stretching of C-H and CH2 due to cellulose, while the broad and strong spectrum detected at 3335.28 cm−1 was associated with the stretching of the alcohol O-H hydrogen bond, aliphatic groups, and intermolecular and intramolecular hydrogen bonds within the cellulose structure [31]. The spectrum gradually decreased in intensity in the order of commercially treated on par with the S2, S1, S3, S6, S8, and S10 enzyme-treated biomass when compared to the untreated control. The spectrum observed for the S2 enzyme-treated biomass showed significant peak shifts indicating the breakdown of lignin and cellulosic moieties which are highlighted in Fig. 4. Based on this analysis, the crude enzyme from S2 was found to catalyze the degradation of the lignin components in the biomass on par with the commercial enzyme.
3.3.3 Biomass-derived products detected by gas chromatography-mass spectrometry
The GC-MS analysis was next used to detect the biochemical products generated after pretreatment of the Erianthus bagasse. Pretreatments included a control, crude enzymes from the S1, S2, S3, S6, S8, and S10 isolates, and a commercial laccase. Metabolites were identified based on relative peak area and the probability of the detected component. The GC-MS analysis of the bagasse treated with the enzymes showed 96 metabolites which were totally absent in the control biomass (without enzymatic treatment). Out of the 96 metabolites, 43 metabolites with high probability and peak area, along with their component name and retention time, were identified and selected. These 43 metabolites were compared among the different isolates using a heat map (Fig. 5). Certain compounds, such as tetradecanoic acid, n-hexadecanoic acid, and octadecanoic acid, were identified in all the samples digested with all the six isolates. Derivatives of these components were identified to be of high abundance in the enzyme-treated bagasse when compared to the untreated control that indicate the breakdown of lignin and the formation of derivative compounds [24]. The breakdown products included low molecular weight metabolites, such as propionic acid, likely generated by the breakdown of ester and phenyl-linked components. The GC-MS analysis also detected metabolites with phenolic, acidic, and ester groups due to the breakdown of the β-aryl and benzoyl ether bonds in the lignin [34]. Metabolites associated with the formation of lignin benzene-diols and the breakdown of the cellulose and hemicellulose fractions of biomass were also detected. The metabolites generated from Erianthus bagasse treated with crude enzymes of isolates S1, S3, and S6 predominately showed phenol, phthalates (diisooctyl and diethyl phthalate), and trans-sinapyl alcohol which are highlighted in Fig. 5.
Heat map of lignin-derived metabolites detected by GC-MS after biomass digestion. Green indicates the relative peak maximum of 30, while yellow represents an absence of detecting the compound (peak area of 0) and the gradual coloring represents the intermediate amounts of metabolites detected by GC-MS. The metabolites encircled were used to show the compounds specifically and predominantly reduce due to lignin breakdown in biomass. Origin software was used to generate the heat map as described in methods
3.4 Residual laccase enzyme activity
Recycling of the enzyme biocatalyst during the pretreatment process is desirable as this reduces cost and minimizes waste. To investigate the possibility of reusing the laccase enzyme preparations, the supernatant was collected from the biomass slurry after being treated with the crude enzymes from all the isolates and analyzed for residual laccase activity. Residual laccase activity was detected at 0.006, 0.023, 0.004, 0.021, 0.002, and 0.003 U mg−1 of protein for the isolates (S1, S2, S3, S6, S8, and S10), respectively (Fig. S3). These results reveal that high residual laccase activity was detected in S2 isolate which can be retrieved after the enzymatic digestion process when performed at the lab scale, suggesting these enzymes could be recycled in an active form after pretreatment of bagasse at the industrial scale.
3.5 Degradation of aromatic azo dyes
Aromatic azo dyes used in the textile industry yet often form aromatic amines which are toxic to the environment and human health. Laccases and other ligninolytic enzymes are found to be useful in decolorizing aromatic azo dyes and preventing the formation of toxic aromatic amines [14, 25]. Thus, the thermo-tolerant fungal isolates that were identified in this study to produce laccase activity were grown on plates supplemented with malachite green to screen their activity. These dyes are chromophore compounds that are highly visible even at low concentrations, and certain fungal isolates are distinctly capable of removal and mineralization of azo dyes by their enzymatic machinery [35]. Here, an Escherichia coli (non-laccase producing) isolate was used as a negative control with no inoculation as experimental control which is shown in Figs. 6a, b respectively. Among the six isolates (S1, S2, S3, S6, S8, and S10 shown in Figs. 6c–h, respectively) studied, we were able to observe the zone of clearance for isolates S1 and S3, while isolate S2 showed a more promising zone of clearance. The enzyme-producing fungal S2 isolate was found to positively produce a zone of clearance on malachite green (Fig. 6d) PD plates examined. The azo dyes that were removed by the fungal isolates S2, S3, and S1 were malachite green, which is an industrial textile dye with complex aromatic amine rings and sulfonic groups. These results reveal that isolate S2 is able to oxidize complex aromatic compounds.
Isolate S2 mediates the oxidation of azo dyes as indicated by plate assay. PD plates supplemented with malachite green dye 0.5% w/v. (a) Negative control using E. coli strain, (b) control PD plate with no isolate, (c, d, e) isolates S1, S2, and S3 showing zone of clearance, and (f, g, h) isolates S6, S8, and S10
3.6 Fungal isolates classified to the ascomycetes phyla
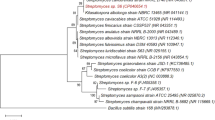
Primers targeting the internal transcribed spacer region (ITS) were designed (ITS1F and ITS4 primers) to distinguish fungi of the ascomycetes, basidiomycetes, and zygomycetes [36]. We were able to observe an amplification of 600 bp for all six isolates (Fig. 7a). All the six isolates were sequenced in which the PCR product derived from isolate S2 revealed 99.16% sequence identity to A. caespitosus. Using this DNA sequence information, a phylogenetic tree was constructed for isolate S2 in which isolate S2 clustered with the fungal species to A. caespitosus. Therefore, with the available data the isolate S2 is affiliated ascomycetes phylum, A. caespitosus (Figs. 7b, c). The morphology of isolate S2 was visualized by growing the culture in czapek yeast extract media for seven days and the microscopic characterization of isolate S2 was done using the lactophenol cotton blue at 40x to visualize the sclerotia, hyphae, and conidiospores of isolate S2 shown in supplementary data (Fig. S4).
Molecular identification of fungal isolates using ITS primer and phylogeny of the query isolates clustering. (a) Molecular identification of all 6 isolates using ITS primer with 600 bp. (b) Phylogenic analysis of S2 isolates clustering with the Aspergillus caespitosus. (c) Phylogenic analysis of S3, S6, and S10 isolates clustered with Flammulina velutipes
3.7 Laccase enzymes purified by ethanol precipitation
The laccase enzymes were further purified from the culture broth of the six fungal isolates (S1, S2, S3, S6, S8, and S10) by ethanol precipitation. The spent culture broth containing the laccase enzyme activity was mixed with an equal volume of ethanol and precipitated. The protein composition of this precipitated fraction was then analyzed by 12% reducing SDS-PAGE and Coomassie Blue R250 staining. The major protein bands detected in the ethanol precipitates were found to migrate between 30 and 40 kDa for the six isolates (Fig. 8a). A single protein band of 30 kDa was identified in the S2 isolate that was common to isolates S3, S6, S8, and S10. The S1 sample did not have this 30 kDa protein and was instead composed of a 33 kDa protein suggesting a different homolog or isoform of the laccase. The specific activity of the purified laccase from all the isolates is shown in Fig. 8b, while for the S2 isolate it was 19.22 U mg−1 of protein, a 3.5-fold increase in activity compared to the crude enzyme in the culture broth. Together these results indicate that the laccase enzyme can be readily purified in an active form from the culture broth of the S2 isolate by ethanol precipitation. The specific enzyme activity in the presence of copper inhibitor 0.1% thioglycolic acid for 15 minutes was 10.37, 18.65, 5.89, 11.56, 13.76, and 12.14 U/mg for all six isolates from S1 to S10. The presence of copper inhibitor shows only a slight variation in the specific activity of the enzymes which has been shown in Fig. S5.
Crude enzyme protein molecular weight determination by SDS-PAGE along with the specific activity of laccase enzyme for all 6 isolates. Panel A: lane M shows the pre-stained protein ladder. Crude protein isolated from S1 (lane 1), S2 (lane 2), S3 (lane 3), S6 (lane 4), S8 (lane 5), and S10 (lane 6). Panel B: the specific enzyme activity of all 6 isolates
4 Discussion
The study was aimed at identifying potential thermo-tolerant fungi that could degrade lignin from Erianthus bagasse. In our explorations, we have isolated and purified 43 isolates from sugarcane compost pit. All the isolates were subjected to preliminary screening through laccase enzymatic assay. Based on the high value of lignolytic enzyme activity with ABTS as substrate, we selected six isolates for further analysis. Selection of isolates based on their laccase enzyme activity was in accordance to [37], where they selected two bacteria isolates from 140 isolates with high laccase activity. As the optimization of enzyme production can be done using the response surface methodology which enhance the production of enzyme among which the Box–Behnken design of experiments was used to construct second-order response surfaces with the input parameters such as pH, temperature, and time [21]. The efficiency of these six isolates on the delignification of the Erianthus bagasse was determined through scanning electron microscopy (SEM), FT-IR, and GC-MS. The SEM analysis of the degraded substrate showed an enzyme-dependent alteration. The observed peeling of intermolecular structures is an indicator that the enzymatic pretreatment is breaking down components of the biomass [38]. These enzyme-dependent alterations likely removed lignin from the cell wall resulting in absences of cohesion between adjacent cell walls within the biomass. Cavities generated on the surface of the biomass are found to aid in the access of cellulose and hemicellulose enzymes for saccharification [18]. These results were similar with the earlier reports on various plants [39,40,41]. The alteration in the functional groups for laccase pretreated Erianthus bagasse in relation to the non-treated substrate was qualitatively observed by FT-IR analysis. The reduced and weakened peak was observed at 1509 cm−1 which is in consistent with a stretching of the aromatic rings of lignin [42]. Our results were in accordance with previous findings that reveal lignin can be assigned to a broad region (3300–3600 cm−1) with hydrogen bond in phenolic groups, OH stretching of alcohols, phenols, and acids [43]. The bands detected at ~2900 cm−1 were consistent with the C−H stretching in methyl and methylene groups [44,45,46]. The signal detected at 1725 cm−1 was consistent with carboxylic (C=O stretching vibration) stretching of the hemicelluloses [42]. These findings are indicative of the enzyme-pretreatment step breaking down the biomass, especially the lignin component. Based on this analysis, the crude enzymes from S10, S8, and S2 were found to catalyze the degradation of the lignin components in the biomass on par with the commercial enzyme. The intermediates observed during GC-MS analysis could be a result of the laccase reactions on the phenylpropanoid units. The release of phenolic ketones and heterocyclic aromatic compounds on laccase pretreatment is described natural mediators of non-phenolic lignin biodegradation process [39]. Nevertheless, the metabolite like phthalates has industrial application especially in personal care products and cosmetics and also used in plasticizers [47]. These products are likely derived from the breakdown of S-lignin (syringyl structure lignin) and H-lignin (hydroxyphenyl structure lignin) as this was the composition of lignin component [48]. Metabolites from carbohydrate breakdown, including furaldehyde, furan methanol, furanone, and methyl furfural, were also detected exclusively in the bagasse pretreated with enzymes and not in the control sample. Most of the lignin-derived metabolites generated after pretreatment of the biomass were found in samples treated with the enzymes from S1 and S2 isolates and the commercial source. The laccase enzyme thermal stability was investigated by measuring the residual enzymatic activities after pretreatment of the Erianthus bagasse, at pH 5.0. A comparative study showed that wheat straw delignification of Phlebia brevispora was 30.6% and Trametes trogii was 21.9%, respectively [49], while the laccase enzyme identified from Trichoderma viride had an ability to reduce the lignin content up to 56% in rice straw [50]. Laccase enzyme isolated from Pleurotus djamor was used for the delignification of sugarcane tops which resulted in 79.1% delignification [39].
Large-scale utilization of these crude enzymes would be facilitated in a continuous and batch process where the residual enzymes are reused for modification of the substrate [51]. The laccase derived from the S2 isolate is particularly desirable for this process as it displays the highest residual activity of the isolates isolated in this study and has specific activity values on par with the commercial enzyme. In addition to the increased residual laccase enzyme activity, we observed zone for clearance for the S2 isolate which is in accordance with the earlier numerous reports decolorization of malachite green [52,53,54]. The ITS region was used for phylogenetic classification of fungi, and the results show that isolate S2 is highly similar to A. caespitosus [55] and isolates S3, S6, and S10 show resemblance with Flammulina velutipes [56].
The SDS-PAGE results showed that A. caespitosus has a molecular weight of 30, 33, and 40 kDa close to the laccases from A. nidulans which is reported as 40 kDa [57]. The bands of lesser molecular weight were observed this might be the different isoforms of laccase. A majority of laccases have been reported to be produced as multiple isoforms; three and four laccase isoforms are reported in T. villosa [58] and Pleurotus ostreatus, respectively [59]. The specific activity of purified laccase recorded a 3.5-fold increase as that of the earlier reports [60, 61]. Our results on the specific enzyme activity of laccase in the presence of thioglycolic acid were in accordance with the earlier report [62].
Therefore, in this study, six isolates of thermo-tolerant fungi were isolated that secrete laccase activity at high levels in the culture broth. The isolates were classified to the ascomycetes using ITS primers and found to produce laccase specific activity in the culture broth at a level on par with the laccase purified commercially from T. versicolor of the Basidiomycetes. Comparison of the Erianthus biomass pretreated with crude enzymes of the different ascomycetes isolates showed that the breakdown of lignin components was predominantly detected in the enzyme preparations from the S2, S8, and S1 isolates. Further analysis revealed isolate S2 to be superior to the other isolates in its capacity to degrade complex aromatic azo dyes. Thus, isolate S2 was further analyzed by DNA sequencing the ITS PCR product and found to be closely related to A. caespitosus and the morphology of the isolate S2 has been studied using the lacto-phenol cotton blue [63]. The S2 isolate was found to secrete the laccase enzyme at such abundance that it was readily purified to homogeneity and high specific activity (19.2 U mg−1) by ethanol precipitation from the culture broth. Sandrim and his team reported the production of two xylanase (xyl I and II) from thermo-tolerant fungus Aspergillus caespitosus [64]. In an earlier report, A. caespitosus (ASEF14) strain also exhibits high lipid content similar to vegetable oil, when it has been grown in sago processing wastewater which has been employed for biodiesel production [55]. They also showed that these enzymes preferentially hydrolyzed birchwood xylan. Likewise, Alegre and his co-workers have observed the production of thermo-stable invertases by A. caespitosus strains under submerged or solid state fermentation using agro-industrial residues as carbon source [65]. Here, we provide the first report that A. caespitosus strain S2 secretes laccase enzyme at specific activity levels that can modify sugarcane bagasse, a feedstock for biofuels.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Rezania S, Oryani B, Cho J et al (2020) Different pretreatment technologies of lignocellulosic biomass for bioethanol production: an overview. Energy 199:117457. https://doi.org/10.1016/j.energy.2020.117457
Jayabose C, Arumuganathan T, Amalraj VA et al (2017) Compressive force profile of high biomass Erianthus clones. Sugar Tech 19:341–346. https://doi.org/10.1007/s12355-016-0472-8
Kasirajan L, Aruchamy K, Thirugnanasambandam PP, Athiappan S (2017) Molecular cloning, characterization, and expression analysis of lignin genes from sugarcane genotypes varying in lignin content. Appl Biochem Biotechnol 181:1270–1282. https://doi.org/10.1007/s12010-016-2283-5
Ummalyma SB, Supriya RD, Sindhu R et al (2019) Biological pretreatment of lignocellulosic biomass-current trends and future perspectives. Second Third Gener Feed Evol Biofuels 7:197–212. https://doi.org/10.1016/B978-0-12-815162-4.00007-0
Yoo CG, Meng X, Pu Y, Ragauskas AJ (2020) The critical role of lignin in lignocellulosic biomass conversion and recent pretreatment strategies: a comprehensive review. Bioresour Technol 301:122784. https://doi.org/10.1016/j.biortech.2020.122784
Saini A, Aggarwal NK, Sharma A, Yadav A (2015) Actinomycetes: a source of lignocellulolytic enzymes. Enzyme Res 2015:1–15. https://doi.org/10.1155/2015/279381
Uthandi S, Saad B, Humbard MA, Maupin-furlow JA (2010) LccA , an archaeal laccase secreted as a highly stable glycoprotein into the extracellular medium by Haloferax volcanii. 76:733–743. https://doi.org/10.1128/AEM.01757-09
Hasanin MS, Darwesh OM, Matter IA, El-saied H (2019) Isolation and characterization of non-cellulolytic Aspergillus fl avus EGYPTA5 exhibiting selective ligninolytic potential. Biocatal Agric Biotechnol 17:160–167. https://doi.org/10.1016/j.bcab.2018.11.012
Ferdes M, Dinca MN, Moiceanu G et al (2020) Microorganisms and enzymes used in the biological pretreatment of the substrate to enhance biogas production: a review. Sustainability 12:2–25. https://doi.org/10.3390/su12177205
Wagle A, Angove MJ, Mahara A et al (2022) Multi-stage pre-treatment of lignocellulosic biomass for multi-product biorefinery: a review. Sustain Energy Technol Assessments 49:101702. https://doi.org/10.1016/j.seta.2021.101702
Sharma HK, Xu C, Qin W (2019) Biological pretreatment of lignocellulosic biomass for biofuels and bioproducts: an overview. Waste and Biomass Valorization 10:235–251. https://doi.org/10.1007/s12649-017-0059-y
Yang J, Li W, Bun Ng T et al (2017) Laccases: production, expression regulation, and applications in pharmaceutical biodegradation. Front Microbiol 8:832. https://doi.org/10.3389/fmicb.2017.00832
An Q, Qiao J, Bian L-S et al (2020) Comparative study on laccase activity of white rot fungi under submerged fermentation with different lignocellulosic wastes. BioResources 15:9166–9179. https://doi.org/10.15376/biores.15.4.9166-9179
Barapatre A, Aadil KR, Jha H (2017) Biodegradation of malachite green by the ligninolytic fungus Aspergillus flavus. Clean - Soil, Air, Water 45:1–22
Thamvithayakorn P, Phosri C, Pisutpaisal N, et al (2019) Utilization of oil palm decanter cake for valuable laccase and manganese peroxidase enzyme production from a novel white-rot fungus, Pseudolagarobasidium sp. PP17-33. 3 Biotech 9:1–10. https://doi.org/10.1007/s13205-019-1945-8
Xu S, Wang F, Fu Y et al (2020) Effects of mixed agro-residues (corn crop waste) on lignin-degrading enzyme activities, growth, and quality of: Lentinula edodes. RSC Adv 10:9798–9807. https://doi.org/10.1039/c9ra10405d
Chen F, Martín C, Lestander TA et al (2022) Shiitake cultivation as biological preprocessing of lignocellulosic feedstocks – substrate changes in crystallinity, syringyl/guaiacyl lignin and degradation-derived by-products. Bioresour Technol 344. https://doi.org/10.1016/j.biortech.2021.126256
Thangavelu K, Desikan R, Taran OP, Uthandi S (2018) Delignification of corncob via combined hydrodynamic cavitation and enzymatic pretreatment: process optimization by response surface methodology. Biotechnol Biofuels 11:1–13. https://doi.org/10.1186/s13068-018-1204-y
Sidana A, Farooq U (2014) Sugarcane bagasse: a potential medium for fungal cultures. Chinese J Biol 2014:1–5. https://doi.org/10.1155/2014/840505
Wang L, Nie Y, Tang YQ et al (2016) Diverse bacteria with lignin degrading potentials isolated from two ranks of coal. Front Microbiol 7:1–14. https://doi.org/10.3389/fmicb.2016.01428
Bhattacharya SS, Banerjee R (2008) Laccase mediated biodegradation of 2,4-dichlorophenol using response surface methodology. Chemosphere 73:81–85. https://doi.org/10.1016/j.chemosphere.2008.05.005
Arockiasamy S, Krishnan IPG, Anandakrishnan N et al (2008) Enhanced production of laccase from coriolus versicolor NCIM 996 by nutrient optimization using response surface methodology. Appl Biochem Biotechnol 151:371–379. https://doi.org/10.1007/s12010-008-8205-4
Tuck S, Ng R (2005) Protein assay. CRC Press
Du FL, Du QS, Dai J et al (2019) A comparative study for the organic byproducts from hydrothermal carbonizations of sugarcane bagasse and its bio-refined components cellulose and lignin. PLoS One 13:1–16. https://doi.org/10.1371/journal.pone.0197188
Sharma A, Aggarwal NK, Yadav A (2017) Isolation and screening of lignolytic fungi from various ecological niches. Univers J Microbiol Res 5:25–34. https://doi.org/10.13189/ujmr.2017.050202
Karakousis A, Tan L, Ellis D et al (2006) An assessment of the efficiency of fungal DNA extraction methods for maximizing the detection of medically important fungi using PCR. J Microbiol Methods 65:38–48. https://doi.org/10.1016/j.mimet.2005.06.008
Uthandi S, Prunetti L, De Vera IMS et al (2012) Enhanced archaeal laccase production in recombinant Escherichia coli by modification of N-terminal propeptide and twin arginine translocation motifs. J Ind Microbiol Biotechnol 39:1523–1532. https://doi.org/10.1007/s10295-012-1152-7
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the Head of Bacteriophage T4. Nature 227:682–685
Brunelle JL, Green R (2014) Coomassie blue staining, 1st edn. Elsevier Inc.
Yadav S, Gangwar N, Mittal P et al (2014) Isolation, screening and biochemical characterization of laccase producing bacteria for degradation of lignin. Int J Educ Sci Res Rev 1:17–21
Chandel AK, Antunes FAF, Anjos V et al (2014) Multi-scale structural and chemical analysis of sugarcane bagasse in the process of sequential acid-base pretreatment and ethanol production by Scheffersomyces shehatae and Saccharomyces cerevisiae. Biotechnol Biofuels 7:1–17. https://doi.org/10.1186/1754-6834-7-63
Smith BC (1996) Fundamentals of Fourier transform infrared spectroscopy. CRC Press, London, Boca Raton
Guilherme AA, Dantas PVF, Santos ES et al (2015) Evaluation of composition, characterization and enzymatic hydrolysis of pretreated sugar cane bagasse. Braz J Chem Eng 32:23–33. https://doi.org/10.1590/0104-6632.20150321s00003146
Zhang W, Ren X, Lei Q, Wang L (2021) Screening and comparison of lignin degradation microbial consortia from wooden antiques. Molecules 26. https://doi.org/10.3390/molecules26102862
Zubbair N, Ajao A, Adeyemo E, Adeniyi O (2020) Biodegradation of malachite green by white-rot fungus, Pleurotus pulminarious. Egypt Acad J Biol Sci G Microbiol 12:79–90. https://doi.org/10.21608/eajbsg.2020.96943
Manter DK, Vivanco JM (2007) Use of the ITS primers, ITS1F and ITS4, to characterize fungal abundance and diversity in mixed-template samples by qPCR and length heterogeneity analysis. J Microbiol Methods 71:7–14. https://doi.org/10.1016/j.mimet.2007.06.016
Huang XF, Santhanam N, Badri DV et al (2013) Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnol Bioeng 110:1616–1626. https://doi.org/10.1002/bit.24833
Faria SP, de Melo GR, Cintra LC et al (2020) Production of cellulases and xylanases by Humicola grisea var. thermoidea and application in sugarcane bagasse arabinoxylan hydrolysis. Ind Crop Prod 158:112968. https://doi.org/10.1016/j.indcrop.2020.112968
Sherpa KC, Ghangrekar MM, Banerjee R (2018) A green and sustainable approach on statistical optimization of laccase mediated delignification of sugarcane tops for enhanced saccharification. J Environ Manag 217:700–709. https://doi.org/10.1016/j.jenvman.2018.04.008
Banerjee R, Chintagunta AD, Ray S (2019) Laccase mediated delignification of pineapple leaf waste: an ecofriendly sustainable attempt towards valorization. BMC Chem 13:1–11. https://doi.org/10.1186/s13065-019-0576-9
Brar KK, Espirito Santo MC, Pellegrini VOA et al (2020) Enhanced hydrolysis of hydrothermally and autohydrolytically treated sugarcane bagasse and understanding the structural changes leading to improved saccharification. Biomass Bioenergy 139:105639. https://doi.org/10.1016/j.biombioe.2020.105639
Zhang H, Wu S (2015) Generation of lignin and enzymatically digestible cellulose from ethanol-based organosolv pretreatment of sugarcane bagasse. Cellulose 22:2409–2418. https://doi.org/10.1007/s10570-015-0678-z
Kubovský I, Kačíková D, Kačík F (2020) Structural changes of oak wood main components caused by thermal modification. Polymers (Basel) 12:485. https://doi.org/10.3390/polym12020485
Shen DK, Gu S (2009) The mechanism for thermal decomposition of cellulose and its main products. Bioresour Technol 100:6496–6504. https://doi.org/10.1016/j.biortech.2009.06.095
Esteves B, Marques AV, Domingos I, Pereira H (2013) Chemical changes of heat treated pine and eucalypt wood monitored by ftir. Maderas Cienc y Tecnol 15:245–258. https://doi.org/10.4067/S0718-221X2013005000020
Mattos BD, Lourençon TV, Serrano L et al (2015) Chemical modification of fast-growing eucalyptus wood. Wood Sci Technol 49:273–288. https://doi.org/10.1007/s00226-014-0690-8
Saeidnia S (2014) Phthalates. In: Encyclopedia of Toxicology, Third edn. Academic Press, pp 928–933
Kasirajan L, Hoang NV, Furtado A et al (2018) Transcriptome analysis highlights key differentially expressed genes involved in cellulose and lignin biosynthesis of sugarcane genotypes varying in fiber content. Sci Rep 8:1–16. https://doi.org/10.1038/s41598-018-30033-4
Tian SQ, Zhao RY, Chen ZC (2018) Review of the pretreatment and bioconversion of lignocellulosic biomass from wheat straw materials. Renew Sust Energ Rev 91:483–489. https://doi.org/10.1016/j.rser.2018.03.113
Ghorbani F, Karimi M, Biria D et al (2015) Enhancement of fungal delignification of rice straw by Trichoderma viride sp. to improve its saccharification. Biochem Eng J 101:77–84. https://doi.org/10.1016/j.bej.2015.05.005
Henning J and, Manuel P (2016) Enzyme recycling in lignocellulosic biorefineries. Biofuels Bioprod Biorefin 11:150–167. https://doi.org/10.1002/bbb
Sneha U, Poornima R, Sridhar S (2014) Optimization and decolorization of malachite green using Pseudomonas putida. J Chem Pharm Res 6:50–57
Chaturvedi V, Verma P (2015) Biodegradation of malachite green by a novel copper-tolerant Ochrobactrum pseudogrignonense strain GGUPV1 isolated from copper mine waste water. Bioresour Bioprocess 2. https://doi.org/10.1186/s40643-015-0070-8
Wanyonyi WC, Onyari JM, Shiundu PM, Mulaa FJ (2017) Biodegradation and detoxification of malachite green dye using novel enzymes from Bacillus cereus strain KM201428: kinetic and metabolite analysis. Energy Procedia 119:38–51. https://doi.org/10.1016/j.egypro.2017.07.044
Srinivasan N, Thangavelu K, Sekar A et al (2021) Aspergillus caespitosus ASEF14, an oleaginous fungus as a potential candidate for biodiesel production using sago processing wastewater (SWW). Microb Cell Factories 20:1–23. https://doi.org/10.1186/s12934-021-01667-3
Martin KJ, Rygiewicz PT (2005) Fungal-specific PCR primers developed for analysis of the ITS region of environmental DNA extracts. BMC Microbiol 5:28. https://doi.org/10.1186/1471-2180-5-28
Sahay S, Chaurse V, Chauhan D (2020) Laccase from Aspergillus nidulans TTF6 showing Pb activation for smaller substrates and dyes remediation in all climates. Proc Natl Acad Sci India Sect B - Biol Sci 90:143–150. https://doi.org/10.1007/s40011-019-01092-y
Bourbonnais R, Paice MG, Reid ID et al (1995) Lignin oxidation by laccase isozymes from Trametes versicolor and role of the mediator 2,2’-azinobis(3-ethylbenzthiazoline-6-sulfonate) in kraft lignin depolymerization. Appl Environ Microbiol 61:1876–1880. https://doi.org/10.1128/aem.61.5.1876-1880.1995
Palmieri G, Giardina P, Bianco C et al (2000) Copper induction of laccase isoenzymes in the ligninolytic fungus Pleurotus ostreatus. Appl Environ Microbiol 66:920–924. https://doi.org/10.1128/AEM.66.3.920-924.2000
Mukhopadhyay M, Banerjee R (2015) Purification and biochemical characterization of a newly produced yellow laccase from Lentinus squarrosulus MR13. 3 Biotech 5:227–236. https://doi.org/10.1007/s13205-014-0219-8
Wu MH, Lin MC, Lee CC et al (2019) Enhancement of laccase activity by pre-incubation with organic solvents. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-45118-x
Afreen S, Shamsi TN, Baig MA et al (2017) A novel multicopper oxidase (laccase) from cyanobacteria: purification, characterization with potential in the decolorization of anthraquinonic dye. PLoS One 12:1–20. https://doi.org/10.1371/journal.pone.0175144
Shamly V, Kali A, Srirangaraj S, Umadevi S (2014) Comparison of microscopic morphology of fungi using lactophenol cotton blue, iodine glycerol and Congo red formaldehyde staining. J Clin Diagn Res 8:7–8. https://doi.org/10.7860/JCDR/2014/8521.4535
Sandrim VC, Rizzatti ACS, Terenzi HF et al (2005) Purification and biochemical characterization of two xylanases produced by Aspergillus caespitosus and their potential for kraft pulp bleaching. Process Biochem 40:1823–1828. https://doi.org/10.1016/j.procbio.2004.06.061
Alegre ACP, Polizeli M de LT de M, Terenzi HF, et al (2009) Production of thermostable Invertase by Aspergillus caespitosus under submerged or solid state fermentation using agroindustrialresidues as carbon source. Braz J Microbiol 40:612–622. https://doi.org/10.1590/S1517-83822009000300025
Acknowledgements
This work has been supported by the DST SERB project EEQ/2019/000225. We thank Dr. G Hemaprabha, Director ICAR-Sugarcane Breeding Institute, Coimbatore, for providing bagasse material for the study. We also thank Sanjeev, from Metabolomic and Proteomic Analytical Facility of the Department of Agricultural Microbiology, and Velautham, from Nanotechnology of Tamil Nadu Agricultural University, Coimbatore, for their assistance in SEM and GC-MS analysis, respectively.
Funding
This work has been supported by the DST SERB project EEQ/2019/000225 granted to Dr. Lakshmi Kasirajan.
Author information
Authors and Affiliations
Contributions
LK: conceptualization, funding acquisition, investigation, writing—original draft. JAMF: supervision, review, and editing. KK: formal analysis, initial draft. US: formal analysis, review, and editing. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

Figure S1.
Laccase enzyme activity of forty-three isolates identified from sugarcane compost pit. (PNG 341 kb)

Figure S2.
Preparation and pretreatment of Erianthus bagasse as a biomass feedstock. a) The Erianthus clone was collected from the field, chopped, juice extracted, and dried. b) The ground biomass was sieved through and selected for 200 μm size. c) Biomass was digested with commercial enzyme, and the crude enzymes extracted from the isolates S1, S2, S3, S6, S8, S10 and a control (without any addition of enzymes). d). Dried biomass was collected after the enzymatic digestion. (PNG 1876 kb)

Figure S3.
Residual enzyme activity of the biomass slurry after enzymatic treatment. Enzymatic activity from the six thermophilic isolates compared to the purified commercial laccase enzyme. Error bar in the graph represents the standard error of the mean value. Data are presented as the mean ± SD (n=3) with Degree of freedom DF of 2 with the statistically insignificant significant p-value (p< 0.05). (PNG 394 kb)

Figure S4.
Morphological and the microscopic characterization of isolate S2 (a) Morphological image of S2 isolate in the Czapex yeast agar media. (b &c) Microscopic image of isolate S2 using lacto phenol cotton blue staining at 40X (PNG 2185 kb)

Figure S5.
The specific enzyme activity of all six isolates in the presence of copper inhibitor thioglycolic acid (PNG 206 kb)
Table S1.
Box-Behnken design matrix for laccase production optimization. The variables were analyzed at three levels namely low, medium and high coded as -1, 0 and +1. (PDF 416 kb)
Table S2.
Analysis of variance (ANOVA) for the quadratic model for optimization of laccase production. (PDF 404 kb)
Table S3.
GC-MS metabolites with high probability and their retention time (PDF 613 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kasirajan, L., Kamaraj, K., Maupin-Furlow, J.A. et al. Isolation, purification, and identification of novel lignin-degrading Aspergillus caespitosus strain S2. Biomass Conv. Bioref. (2022). https://doi.org/10.1007/s13399-022-03510-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13399-022-03510-0