Abstract
The Trigonella species possess medicinal, nutraceutical and pharmaceutical properties due to the presence of many bioactive compounds. Its therapeutic effects are mostly valuable in medicine, cosmetics and the functional food industry. Correct genetic characterisation of plant material is needed to increase the potential of Trigonella species by breeding and conservation programs. The aim of this study was to develop a reliable marker system to support the morphological and phytochemical analysis in Trigonella taxonomic research, species identification and characterization as well as determination of the interspecific variation within this genus along with relationships between species. For this purpose, flow cytometry and SCoT molecular markers were combined. Flow cytometric analyses revealed that Trigonella species possess very small and small genomes. The range of genome sizes was from 1.10 to 5.76 pg/2C, with most species possessing very small genomes (< 2.8 pg/2C). In seeds of 14 species endopolyploid nuclei were detected. Flow cytometric analysis of genome size enabled quick identification of four out of 20 species, while combined with endopolyploidy detection in seeds, facilitated distinction of the next seven species. ScoT molecular markers helped to identify closely related species with similar genome size and cell cycle activity. Therefore, flow cytometry was proposed as the first-choice method for quick accession screening, while the more detailed genetic classification was obtained using SCoT molecular markers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Trigonella L. belongs to the Fabaceae family. It consists of many perennial and annual species distributed in the dry regions around the Mediterranean Sea, West Asia, Europe, Africa, North America, and South Australia (Mabberley 1997). The exact number of species that comprise the genus Trigonella is still being analyzed, but the most relevant classification indicates 62 species in the genus (Dangi et al. 2016).
The Trigonella species have medicinal, nutraceutical and pharmaceutical properties due to the presence of many bioactive compounds (e.g. sapogenins, flavonoids, polysaccharide galactomannans, phenolics; Dangi et al. 2004; Ahari et al. 2009; Mehrafarin et al. 2011; Al-Asadi 2014). Within this genus, T. foenum-graecum (fenugreek) is the best-known species with the greatest economic importance since its long history in natural medicine and as a food condiment. High protein content and favourable amino acid composition make fenugreek seeds equal in nutritive value to soybean (Niknam et al. 2004). Trigonella seeds, particularly those of Trigonella foenum-graecum (fenugreek), have a variety of uses as food. Fenugreek seeds are widely used as a spice in culinary preparations. They add a distinct flavour to dishes and are commonly used in Indian, Middle Eastern, and Mediterranean cuisines. The seeds can be used as a food additive and preservative due to their flavour and potential antimicrobial properties. Fenugreek leaves and seeds can be used directly as a vegetable in various dishes (Syed et al. 2020; Ahmad et al. 2023). Moreover, it was proved that this plant has multiple therapeutic properties, mainly against atherosclerosis, hypertension, obesity, and cholestasis. The therapeutic potential in anti-cancerous, anti-inflammatory, antioxidant, antiulcer and antipyretic properties was also proved (Sushma and Davasena 2010; Naidu et al. 2011; Moradikor and Moradi 2013).
Taxonomic classification of species within the Trigonella genus was initially based on morphological characteristics. Sirjaev (1928–1932) divided the genus into three subgenera and 15 sections. Subgenus I: Trigonella sections: Falcatulae, Callicerates, Uncinatae, Cylindricae, Samaroideae, Pectinatae, Erosae, Verae, Spinosae. Subgenus II: Trifoliastrum: section Capitatae. Subgenus III: Foenum-graecum sections: Foenum-graecum, Medicagoids, Medicago, Melilotus, Trifolium. Nevertheless, karyogenetic and molecular data suggested that this taxonomic concept is insufficient because of the high variation of morphological features under environmental factors (Dangi et al. 2016; Al-Maamari et al. 2020). Therefore, environmentally independent karyological and molecular markers are searched to supplement morphological characterization. Among the karyological markers chromosome number and morphology, genome size and endopolyploidy level were used to support the taxonomic classification of other species (e.g. Rewers and Jedrzejczyk 2016; Ducar et al. 2018; Jedrzejczyk and Rewers 2018; Jedrzejczyk 2020). So far, only chromosome number and morphology have been used to support the taxonomic classification of Trigonella genus (Yilmaz et al. 2009; Martin et al. 2011a, 2011b; Najafi et al. 2013; Ranjbar and Hajmoradi 2016). Most species of this genus are diploids with the number of chromosomes 2n = 2x = 14 or 2n = 2x = 16 (Martin et al. 2011a, 2011b) though polyploid species/cytotypes were also observed (Malhotra 2011).
Among molecular markers, various DNA fingerprinting techniques have been used to identify species/accessions as well as to study the genetic diversity and relationships between and within different plant species/cytotypes (Rewers and Jedrzejczyk 2016; Ducar et al. 2018; Jedrzejczyk and Rewers 2018; Jedrzejczyk 2020; Jedrzejczyk and Rewers 2020). In the Trigonella genus, most of the molecular research were focused on the application of molecular markers like RAPD, AFLP, ITS-rDNA, SRAP, ISSR, SSR and SCoT to assess the genetic diversity and population structure of T. foenum-graecum (e.g. Dangi et al. 2004, 2016; Kumar et al. 2012; Randhawa et al. 2012; Tomar et al. 2014; Hora et al. 2016; Amiriyan et al. 2019; Maloo et al. 2020). So far, only Dangi et al. (2016) have provided evidence of phylogenetic relationships between 22 Trigonella species using nuclear ribosomal ITS and chloroplast trnL intron sequences. The research provided strong support for the monophyly of the genus and revised the previous classification of the genus.
Among different molecular marker systems, the Start Codon Targeted (SCoT) polymorphism method was proved to be suitable for plant identification and assessing their genetic diversity and relationships between genotypes. SCoT markers are based on the amplification of short, conserved regions in plant genes surrounding the translation start (or initiation) ATG codon (Collard and Mackill 2009). The markers have been successfully used to evaluate genetic diversity and structure in for instance: wheat, sugarcane and coneflower (Que et al. 2014; Etminan et al. 2016; Jedrzejczyk 2020). Daneshmand et al. (2017) investigated the genetic diversity within and between different populations of Trigonella foenum-graecum using ISSR and SCoT markers as well as phytochemical profiles, detecting a high level of genetic variation among them. Moreover, it was proved that the SCoT technique is more informative, polymorphic and repetitive than ISSR or RAPD markers for the evaluation of genetic diversity and relationships among fenugreek populations different in respect of trigonelline content characteristic phytochemical for seeds and leaves of fenugreek (Daneshmand et al. 2017).
The study aimed to develop a reliable marker system to support the morphological and phytochemical analysis in Trigonella taxonomic research, to identify and characterize species as well as determine the interspecific variation within this genus along with relationships between accessions. For this purpose, we applied analysis of genome size, endopolyploidy level in seeds, and SCoT molecular markers. Additionally, to our knowledge, this is the first report on combining the genome size estimation, endopolyploidy analysis and SCoT markers for exploring a great number of Trigonella species.
Materials and methods
Plant material
Seeds of twenty Trigonella accessions were received from GRIN-ARS-USDA gene bank (Table 1). To obtain plant material, all seeds were sown in 12 cm pots, mixed with sand and commercial humus (1:2, w/w), and placed in a growth chamber at 26/18°C (day/night) with 16/8 photoperiod.
Genome size measurements
Nuclear DNA content was estimated in fresh and young leaves of all Trigonella accessions (Table 1). Plant material for the flow cytometric analysis was prepared according to the protocol described by Rewers and Jedrzejczyk (2016). Two internal standards were applied for genome size calculation. For three species (T. anguina, T. kotschyi and T. spruneriana ssp. sibthorpii) leaves of T. corniculata (1.77 pg/2C) were used as the internal standard, while for the 17 species leaves of Vicia villosa ‘Minikowska’ (3.32 pg/2C, Dzialuk et al. 2007) were applied. The nuclei suspension was prepared using 1 ml of nuclei isolation buffer (2.5 mM MgCl2×6H2O, 85 mM NaCl, 0.1 M Tris, 0.1% (v/v) Triton X-100, pH 7.0; Rewicz et al. 2018), with the addition of propidium iodide (PI, 50 μg/ml), ribonuclease A (RNase A, 50 μg/ml) and 1.0% (w/v) polyvinylpyrrolidone (PVP-10). Samples were analysed using a CyFlow SL Green (Partec GmbH, Münster, Germany) flow cytometer, equipped with a high-grade solid-state laser, with green light emission at 532 nm, as well as with side (SSC) and forward (FSC) scatters. The DNA content of 5000-7000 nuclei was measured for each accession using linear amplification. The histograms (Mean CV = 3.01-4.83%; Table 1) were evaluated using FloMax program (Partec GmbH, Münster, Germany). Analyses were performed on six individuals per species. Genome size was determined using the linear relationship between the ratio of the 2C peak positions of Trigonella accessions and the internal standard on the histogram of fluorescence intensities. The 2C DNA contents (pg) were converted to the megabase pairs (Mbp) of nucleotides, using the formula: 1 pg = 978 Mbp (Doležel and Bartoš 2005; Table 1). The results have been estimated using a one-way variance analysis and Duncan’s test (P < 0.05; Statistica v. 13.3, StatSoft, Poland).
Cell cycle analysis
The cell cycle was analyzed in mature, dry seeds of all accessions using the flow cytometric method (Table 2). Samples for the analysis were prepared as previously described by Rewers et al. (2009), using nuclei isolation buffer (2.5 mM MgCl2×6H2O, 85 mM NaCl, 0.1 M Tris, 0.1% (v/v) Triton X-100, pH 7.0) supplemented with 4’,6-diamidino-2-phenylindole (DAPI; 2 μg/mL) for DNA staining. Analyses were performed on five biological replicates using a CyFlow Ploidy Analyser flow cytometer (Sysmex-Partec GmbH, Gorlitz, Germany). For each sample fluorescence of 5000–7000 nuclei was recorded. The proportion of nuclei with different DNA contents, number of endocycles, and mean C-value (Lemontey et al. 2000) were calculated. In this work, only nuclei with DNA content higher than 4C were considered endopolyploid (Rewers and Sliwinska 2012, 2014; Ducar et al. 2018). The results were statistically analyzed using a one-way variance analysis and Duncan’s test (P < 0.05; Statistica v. 13.3, StatSoft, Poland).
DNA isolation and quantification
Total genomic DNA was extracted from 0.12 g of fresh leaf tissue of three randomly selected plants per accession, using a GeneJET Plant Genomic DNA Purification Mini Kit (Thermo Fischer Scientific, USA). DNA quality and quantity were estimated using a BioPhotometer (Eppendorf, Poland) and agarose gel electrophoresis on 1% agarose gel. Samples with high-quality DNA were used for SCoT-PCR reactions.
DNA amplification
SCoT-PCR analyses were performed using 20 primers (Genomed, Poland), out of which 13 generated stable band patterns and were selected for further studies (Table 3). Amplification was performed in a total reaction mixture of 12 μl, containing 30 ng of genomic DNA template, 5 μl of 2x PCR Master Mix Plus (containing 0.1 U/μl Taq DNA polymerase, 4 mM MgCl2 and 0.5 mM of each dNTPs; A&A Biotechnology, Poland), 1 μl of 10 μM primer and sterile, deionized water. The PCR reactions were run using T100 Thermal Cycler (Bio-Rad, USA) at 94°C for 5 min, followed by 35 cycles of denaturation at 94°C for 1 min, primer annealing at 49.0-63.4°C (depending on the primer) for 1 min., primer elongation at 72°C for 2 min. The final extension step was 7 min at 72°C. The PCR products were detected on 1.5% (w/v) agarose gel stained with ethidium bromide (0.5 μg/ml). A DNA ladder of 3000 bp was used to determine the size of the amplicons (Genoplast Biochemicals, Poland). The bands were visualized and archived using GelDoc XR+ (Bio-Rad, USA).
Data analysis
The banding pattern of the SCoT markers was scored as presence (1) and absence (0) of the band and set in a binary matrix. Only clear and unambiguous fragments for each primer were recorded. The numbers of monomorphic and polymorphic DNA fragments amplified by each primer were determined. The informativeness of the primer was described using the Polymorphism Information Content (PIC), and calculated according to Ghislain et al. (1999) by the formula: PIC = 1 – p2 – q2, where p is the band frequency, and q is no band frequency. According to Nei and Li (1979), genetic distances were calculated for all accessions. The phylogenetic tree was constructed using the unweighted pair group method, with arithmetic average (UPGMA), using the Treecon ver. 3.1 software (Van de Peer and De Wachter 1994). Bootstrapping was performed using 2000 replicates to assess the confidence values of the clusters formed.
Results
2C DNA content
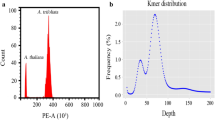
Flow cytometric analysis revealed that the genome size of Trigonella accessions ranged from 1.10 pg/2C in T. arabica (section Pectinatae) to 5.76 pg/2C in T. foenum-graecum (section Foenum-graecum), so the difference between the smallest and the largest nuclear DNA content was 5-fold (Table 1, Fig. 1). This corresponds to 1076 and 5633 Mbp, respectively. Based on the 2C DNA content, four out of 20 studied species could be distinguished.
Cell cycle
Based on the flow cytometric analysis, in mature and dry seeds of the studied species, endopolyploid nuclei were detected in 14 out of the 20 species examined. Only six species (T. calliceras, T. caerulea, T. gracilis, T.kotschyi, T. spicata, T. spruneriana ssp. sibthorpii) possess non-polysomatic seeds where only nuclei with 2C and 4C DNA content were observed. In polysomatic Trigonella seeds, besides nuclei with 2C and 4C, also nuclei with 8C DNA content were detected, indicating the occurrence of one endocycle. However, these species varied in the proportion of 2C, 4C and 8C nuclei. The percentage of 8C nuclei ranged from 1% (T. coerulescens, T. foenum-graecum, T. glabra) up to 11% (T. arabica). The mean C-value varied from 2.3 (T. kotschyi) to over 3.3 (T. creatica, T. macrorrhyncha; Table 2), revealing the lowest and the highest intensity of DNA synthesis in seeds.
SCoT markers
A collection of 20 SCoT primers was employed to screen Trigonella species, and 13 of these primers generated consistent polymorphic banding patterns. The chosen primers generated 380 bands. The 12 primers revealed 100% polymorphism. The mean percentage of polymorphism for tested SCoT primers exceeded 99.8% (Table 3). The approximate size of the amplified products ranged from 174 (SCoT-35) to 2924 bp (SCoT-14). The lowest number of bands (22) were generated by SCoT-2 and SCoT-36 primers, while the SCoT-5 primer generated the highest number of bands (35), all polymorphic. The PIC values varied between 0.26 (SCoT-9) to 0.42 (SCoT-34), with an average of 0.33 (Table 3). Four primers: SCoT-7, SCoT-11, SCoT-12, and SCoT-14 were the most effective in species distinction, whereas in genotypes diversification the most accurate were primers: SCoT-7, SCoT-11 and SCoT-14 (the distinction between T. glabra and T. glabra subsp. uncata; Fig. 2), and SCoT-11, SCoT-12 and SCoT-14 (between T. spruneriana and T. spruneriana ssp. sibthorpii).
Selected SCoT band profiles of Trigonella accessions generated by primer SCoT-14. M. 3000 bp ladder; C. negative control; 1 -T. anguina, 2 - T. arabica, 3 - T. balansae, 4 - T. calliceras, 5 - T. caerulea, 6 - T. coerulescens, 7 - T. corniculata, 8 - T. cretica, 9 - T. foenum-graecum, 10 - T. glabra, 11 - T. glabra subsp. uncata, 12 - T. gracilis, 13 - T. kotschyi, 14 - T. macrorrhyncha, 15 - T. schlumbergeri, 16 - T. spicata, 17 - T. spruneriana, 18 - T. spruneriana ssp. sibthorpii, 19 - T. stellata, 20 - T. suavissima
The genetic distance analysis between studied accessions revealed that the highest genetic similarity (0.38) was observed between T. balansae and T. glabra subsp. uncata, whereas the lowest similarity (0.84) was found between T. corniculata and T. spicata, as well as T. arabica and T. caerulea (Table 4). The dendrogram was constructed using binary matrix values as determined from SCoT data and grouped all studied species into three main clusters. The first cluster (I) included 12 species, while the second one (II) had only three species, and the third cluster (III) comprised five species (Fig. 3).
Discussion
The Trigonella genus contains species that are important as medicinal and aromatic plants and are used for multiple purposes. Fresh, dry or powdered leaves, seeds, and extracts induce various pharmacological effects. Therefore, the knowledge about genome size, genetic diversity and the relationship between the species could be very useful in the proper utilization of germplasm resources as well as for its usage in the pharmaceutical industry and breeding programs of Trigonella species.
In this study, the species from 10 sections of Trigonella genus were investigated. The most numerous section Falcatulae, was represented by seven species, while the sections Cylindricae and Foenum-graecum by three species. In the other seven sections (Callicerates, Capitatae, Ellipticae, Erosae, Pectinatae, Samaroideae, Uncinatae), only one species was represented. The genome size measured for seven species of Falcatulae section ranged from 1.34 to 4.02 pg/2C. For three species from Cylindricae section, the 2C DNA content varied from 2.17 to 4.00 pg/2C, while among species of Foenum-graecum section, the values of DNA content ranged from 2.00 to 5.76 pg/2C. In the sections Cellicerates, Ellipticae, Erosae and Uncinatae, the genome size ranged from 2.03 to 2.61 pg/2C, whereas in Pectinatae, Capitatae and Samaroideae was below 2.00 pg/2C. The genome size estimated for 20 Trigonella species could be used to screen genetic diversity within the genus and as an additional parameter for species identification. The nuclear DNA content estimated by FCM analysis allowed to identify four species (T. arabica, T. foenum-graecum, T. spruneriana ssp. sibthorpii and T. stellata). To the best of our knowledge, the 2C DNA content for 19 species of Trigonella genus was estimated for the first time. So far, only for T. stellata the 2C DNA content was measured using the Feulgen method (Bidak and Brandham 1995) and was only 0.06 pg/2C higher than obtained by FCM in the presented study (1.34 pg/2C). According to the categorization proposed by Soltis et al. (2003), 17 species can be considered as having very small genomes (1.10-2.80 pg/2C), while three species as possessing small genome sizes (4.00-5.76 pg/2C). The usefulness of FCM method in plant identification and determination of genetic diversity has also been proved in many herbal plants, e.g. representing Echinacea (Jedrzejczyk 2020), Mentha (Jedrzejczyk and Rewers 2018), Origanum (Jedrzejczyk 2018) and Malva genera (Jedrzejczyk and Rewers 2020). Based on the literature (Martin et al. 2011a, 2011b; Ranjbar and Hajmoradi 2016), most of the studied Trigonella species are diploids with somatic chromosome number 2n=2x=16. Only for T. stellata 18 chromosomes were reported in somatic cells. There is no information about the chromosome number for two species, T. schlumbergeri and T. suavissima. Since the occurrence of extra B chromosomes in some fenugreek lines, the variation in chromosome number can also be observed (Raghuvanshi and Singh 1976; Petropoulos 2002; Martin et al. 2011a, 2011b; Ranjbar and Hajmoradi 2016).
Despite that the Trigonella genus belongs to the Fabaceae family, known for high polysomaty, this genus represented a low level of endopolyploidy in seeds. Among the investigated species, two groups could be distinguished based on the cell cycle analysis in seeds. The first group included 14 species with polysomatic seeds where one endocycle occurred and nuclei with DNA content up to 8C were present. While in the second group, six species with non-polysomatic seeds were included. In seeds of this group, no endocycles were observed, and only nuclei with 2C and 4C were detected. Application of this feature in species identification (together with genome size estimation) enables the distinction of the seven next species. Altogether 11 species could be identified based on flow cytometric measurements. Therefore, endopolyploidy level in seeds can be considered a supportive feature for species identification and classification. Additionally, this is the first report on endopolyploidy in Trigonella seeds.
The Trigonella genus variability analysis using different molecular methods (RAPD, ISSR, SCoT, SSR, SCAR, ITS) is mostly concentrated on T. foenum-graecum genotypes (Mirzahosein-Tabrizi et al. 2023). Studies on T. foenum-graecum concerning the application of RAPD markers recorded polymorphism between 43 and 92% (Hora et al. 2016; Mamatha et al. 2017) while for T. caerulea 95% (Dangi et al. 2004). In the case of ISSR markers, the percentage of polymorphism obtained for T. foenum-graecum populations was between 31 and 92% (Marzougui et al. 2009; Randhawa et al. 2012). Only Al-Maamari et al. (2014) reported 99-100% of polymorphism in T. foenum-graecum from Oman using six AFLP markers. This was higher than the values obtained by Kumar et al. (Kumar et al. 2012; 64%). Nevertheless, to our knowledge, this is the first report of using SCoT molecular markers for exploring genetic diversity for such a large group of species from Trigonella genus. In this study, SCoT markers revealed a high level of polymorphic loci (99.8 %) among all investigated species, being higher than results obtained for T. foenum-graecum accessions (82%) using the same markers (Daneshmand et al. 2017). The efficiency of SCoT markers was also estimated based on Polymorphism Information Content, and the obtained value was 0.33, which was similar to PIC estimated by Daneshmand et al. (2017). The significant level of genetic divergence has been calculated by SCoT molecular markers in other herbal plants, e.g. from Echinacea genus (Jedrzejczyk 2020), Crepedium acuminatum (Thakur et al. 2021), Lycoris species (Gao et al. 2014) and Papaver somniferum (Srivastava et al. 2020), what proved the potential of these markers for assessing genetic diversity and relationships in plants.
The results of SCoT-PCR analysis allowed for establishing the relationship between species, considered by genetic distance and dendrogram construction. This phylogenetic analysis discovered the existence of three clusters. The highest genetic distance was observed between T. corniculata and T. spicata, grouped into two separate clusters, I (T. spicata) and III (T. corniculata), as well as between T. arabica and T. caerulea, clustered in II and III group, respectively. The lowest genetic distance was observed between T. balansae and T. glabra subsp. uncata, species (included in the cluster I). This indicated the highest similarity between these species. Based on genetic distance analysis and genome size, we could not reach a consensus on the parameters. In the I cluster, both the highest and the lowest values of genome sizes were found. However, closely related species (e.g. T. corniculata and T. glabra) possess similar genome sizes. Most of the tested primers generated polymorphic bands. However, three proved to be the best for all the species distinction. Moreover, it was also possible to indicate the primers for distinguishing the genotypes of T. glabra and T. glabra subsp. uncata as well as T. spruneriana and T. spruneriana ssp. sibthorpii. This supports previous findings of high-resolution of SCoT markers in analyzing the genetic diversity of species from Echinacea genus (Jedrzejczyk 2020) and the Fabaceae family (Nosair 2016).
Conclusions
The results obtained in the study revealed that the Trigonella genus could be characterized using SCoT molecular markers with the support of genome size estimation and cell cycle analysis performed by FCM. The presented results increased the knowledge about Trigonella genus, particularly its genome size, endopolyploidy and genetic diversity, as well as phylogenetic relationships between the Trigonella species. This could be useful in the conservation of medicinal plants and the application of some species in breeding programs.
Data Availability
The data that support the findings of this study are available from the corresponding author, [MR], upon reasonable request.
References
Ahari DS, Kashi AK, Hassandokht MR, Amri A, Alizadeh K (2009) Assessment of drought tolerance in Iranian fenugreek landraces. J Food Agric Environ 7(3):414–419. https://doi.org/10.1234/4.2009.2614
Ahmad N, Riaz S, Anwar A, Riaz T (2023) Fenugreek (Trigonella foenum-graecum): An Overview of Food Uses and Health Benefits. In: Neglected Plant Foods Of South Asia, Chapter 18. https://doi.org/10.1007/978-3-031-37077-9_18
Al-Asadi JN (2014) Therapeutic uses of Fenugreek (Trigonella foenum-graecum L.). Am J Soc Issues Hum 2:21–36
Al-Maamari IT, Al-Sadi AM, Al-Saady NA (2014) Assessment of genetic diversity in fenugreek (Trigonella foenum-graecum L.) in Oman. Int J Agric Biol 16(4):813–816
Al-Maamari IT, Khan M, Al-Sadi A, Iqbal Q, Al-Saady N (2020) Morphological characterization and genetic diversity of Fenugreek (Trigonella foenum-graecum L.) accessions in Oman. Bulg J Agric Sci 26(2):375–383
Amiriyan M, Shojaeiyan A, Yadollahi A, Maleki M, Bahari Z (2019) Genetic diversity analysis and population structure of some Iranian fenugreek (Trigonella foenum-graecum L.) landraces using SRAP Markers. Mol Biol Res Commun 8(4):181–190. https://doi.org/10.22099/mbrc.2019.34952.1440
Bidak L, Brandham PE (1995) Intraspecific uniformity of chromosome number and nuclear DNA quantity in two Egyptian weedy species, Malva parviflora (Malvaceae) and Trigonella stellata (Leguminosae). Kew Bull 50:595–599. https://doi.org/10.2307/4110331
Collard BCY, Mackill DJ (2009) Start Codon Targeted (SCoT) Polymorphism: A simple, novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Biol Rep 27:86–93. https://doi.org/10.1007/s11105-008-0060-5
Daneshmand H, Etminan A, Qaderi A (2017) Diversity evaluation of Trigonella foenum-graecum populations using DNA markers and phytochemical characteristics. J Med Plants 16(63):119–132
Dangi R, Tamhankar S, Choudhary RK, Rao S (2016) Molecular phylogenetics and systematics of Trigonella L.(Fabaceae) based on nuclear ribosomal ITS and chloroplast trnL intron sequences. Genet Resour Crop Evol 63:79–96. https://doi.org/10.1007/s10722-015-0236-4
Dangi RS, Lagu MD, Choudhary LB, Ranjekar PK, Gupta VS (2004) Assessment of genetic diversity in Trigonella foenum-graecum and Trigonella caerulea using ISSR nad RAPD markers. BMC Plant Biol 4:13. https://doi.org/10.1186/1471-2229-4-13
Doležel J, Bartoš J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Ann Bot 95(1):99–110. https://doi.org/10.1093/aob/mci005
Ducar E, Rewers M, Jedrzejczyk I, Martonfi P, Sliwinska E (2018) Comparison of the genome size, endoreduplication, and ISSR marker polymorphism in eight Lotus (Fabaceae) species. Turk J Bot 42:1–14. https://doi.org/10.3906/bot-1703-61
Dzialuk A, Chybicki I, Welc M, Sliwinska E, Burczyk J (2007) Presence of triploids among oak species. Ann Bot 99:959–964. https://doi.org/10.1093/aob/mcm043
Etminan A, Pour-Aboughadareh A, Mohammadi R, Ahmadi-Rad A, Noori A (2016) Applicability of start codon targeted (SCoT) and inter-simple sequence repeat (ISSR) markers for genetic diversity analysis in durum wheat genotypes. Biotechnol Biotechnol Equip 30(6):1075–1081. https://doi.org/10.1080/13102818.2016.1228478
Gao YH, Zhu YQ, Tong ZK, Xu ZY, Jiang XF, Huang CHH (2014) Analysis of genetic diversity and relationships among genus Lycoris based on start codon targeted (SCoT) marker. Biochem Syst Ecol 57:221–226. https://doi.org/10.1016/j.bse.2014.08.002
Ghislain M, Zhang D, Fajardo D, Huamán Z, Hijmans RJ (1999) Marker-assisted sampling of the cultivated Andean potato Solanum phureja collection using RAPD markers. Genet Resour Crop Evol 46:547–555. https://doi.org/10.1023/A:1008724007888
Hora A, Malik CP, Kumari B (2016) Assessment of genetic diversity of Trigonella foenum-graecum L. in Northern India using RAPD nad ISSR markers. Int J Pfarm Pfarm Sci 8:179–183
Jedrzejczyk I (2018) Study on genetic diversity between Origanum L. species based on genome size and ISSR markers. Ind Crops Prod 126:201–207. https://doi.org/10.1016/j.indcrop.2018.10.010
Jedrzejczyk I (2020) Genome size and SCoT markers as tools for identification and genetic diversity assessment in Echinacea genus. Ind Crops Prod 144:112055. https://doi.org/10.1016/j.indcrop.2019.112055
Jedrzejczyk I, Rewers M (2018) Genome size and ISSR markers for Mentha L. (Lamiaceae) genetic diversity assessment and species identification. Ind Crops Prod 120:171–179. https://doi.org/10.1016/j.indcrop.2018.04.062
Jedrzejczyk I, Rewers M (2020) Identification and genetic diversity analysis of edible and medicinal Malva species using flow cytometry and ISSR molecular markers. Agronomy 10(5):650. https://doi.org/10.3390/agronomy10050650
Kumar V, Srivastava N, Singh A, Vyas MK, Gupta S, Katudia K, Vaidya K, Chaudhary S, Ghosh A, Chikara SK (2012) Genetic diversity and identification of variety-specyfic AFLP markers in fenugreek (Trigonella foenum-graecum). Afr J Biotechnol 11(19):4323–4329. https://doi.org/10.5897/AJB11.93095
Lemontey C, Mousset-Déclas C, Munier-Jolain N, Boutin JP (2000) Maternal genotype influences pea seed size by controlling both mitotic activity during early embryogenesis and final endoreduplication level/cotyledon cell size in mature seed. J Exp Bot 51:167–175. https://doi.org/10.1093/jexbot/51.343.167
Mabberley DJ (1997) The plant book: A portable dictionary of the vascular plants, 2nd edn. Cambridge University Press. NY, Cambridge, pp 10011–14211
Malhotra SK (2011) Fenugreek (Trigonella foenum-graecum L.). In: Genetic Resources, Chromosome Engineering, and Crop Improvement. Edition: First, Chapter: 24. CRS Press, pp 801–845
Maloo SR, Sharma R, Jain D, Chaudhary S, Soan H (2020) Assessment of genetic diversity in fenugreek (Trigonella foenum-graecum) genotypes using morphological and molecular markers. Indian J Agric Sci 90(1):25–30. https://doi.org/10.56093/ijas.v90i1.98522
Mamatha NC, Tehlan SK, Srikanth M, Shivaprasad MK, Karthik Reddy P (2017) Molecular characterization of fenugreek (Trigonella foenum-graecum L.) genotypes using RAPD markers. Int J Curr Microbiol App Sci 6(6):2573–2581. https://doi.org/10.20546/ijcmas.2017.606.306
Martin E, Akan H, Ekici M, Aytac Z (2011a) New chromosome numbers in the genus Trigonella L. (Fabaceae) from Turkey. Afr J Biotechnol 10(2):116–125. https://doi.org/10.5897/AJB10.972
Martin E, Akan H, Ekici M, Aytac Z (2011b) Karyotype analyses of ten sections of Trigonella (Fabaceae). Comp Cytogenet 5(2):105–121. https://doi.org/10.3897/compcytogen.v5i2.969
Marzougui N, Boubaya A, Elfalleh W, Guasmi F, Laaraiedh L, Ferchichi A, Triki T, Beji M (2009) Assessment of genetic diversity in Trigonella foenum graecum Tunisian cultivars using ISSR markers. J Food Agric Environ 7(1):101–105
Mehrafarin A, Rezazadeh S, Naghdi Badi H, Noormohammadi G, Zand E, Qaderi A (2011) Review on Biology, Cultivation and Biotechnology of Fenugreek (Trigonella foenum-graecum L.) as a valuable medicinal plant and multipurpose. J Med Plants 10(37):6–24
Mirzahosein-Tabrizi M, Ghanavati F, Azizinezhad R, Etminan A (2023) Genetic diversity revealed by phytochemical and molecular analyses among and within eight Trigonella sp. JCSB 26:345–357. https://doi.org/10.1007/s12892-022-00183-z
Moradikor N, Moradi K (2013) Physiological and pharmaceutical effects of Fenugreek (Trigonella foenum-graecum L.) as a multipurpose and valuable medicinal plant. Glob J Med Plant Res 1(2):199–206
Naidu MM, Shyamala Naik JP, Sulochanamma G, Srinivas P (2011) Chemical composition and antioxidant activity of the husk and endosperm of fenugreek seeds. Food Sci Tech 44(2):451–456. https://doi.org/10.1016/j.lwt.2010.08.013
Najafi S, Anakhatoon EZ, Bırsın MA (2013) Karyotype characterisation of reputed variety of Fenugreek (Trigonella foenum-graecum) in West Azerbaijan-Iran. J Appl Biol Sci 7(1):23–26
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76(10):5269–5273. https://doi.org/10.1073/pnas.76.10.5269
Niknam V, Sharifizadeh B, Ebrahimzadeh H, Zarre S, Izadpanah M (2004) Comparative study of proteins in seeds of some species of Trigonella from Iran. Iranian Int J Sci 5(1):1–11
Nosair H (2016) SCoT polymorphism reveals genetic diversity in some important Fabaceae species. Curr Sci Int 5(4):592–598
Petropoulos GA (2002) Fenugreek: The genus Trigonella, 1st edn. CRC Press, London. https://doi.org/10.4324/9780203217474
Que Y, Pan Y, Lu Y, Yang C, Yang Y, Huang N, Xu L (2014) Genetic analysis of diversity within a Chinese local sugarcane germplasm based on Start Codon Targeted polymorphism. Biomed Res Int:1–10. https://doi.org/10.1155/2014/468375
Raghuvanshi SS, Singh AK (1976) Inheritance of B-chromosomes in Trigonella foenum-graecum L. Caryologia 29(3):277–281. https://doi.org/10.1080/00087114.1976.10796667
Randhawa GJ, Singh M, Gangopadhyay KK, Kumar G, Archak S (2012) Genetic analysis of fenugreek (Trigonella foenum-graecum L.) accession using morphometric and ISSR markers. Indian J Agric Sci 82(5):393–401. https://doi.org/10.56093/ijas.v82i5.17796
Ranjbar M, Hajmoradi Z (2016) Chromosome numbers and biogeography of the genus Trigonella (Fabaceae). Caryologia 69(3):223–234. https://doi.org/10.1080/00087114.2016.1169090
Rewers M, Jedrzejczyk I (2016) Genetic characterization of Ocimum genus using flow cytometry and inter-simple sequence repeat markers. Ind Crops Prod 91:142–151. https://doi.org/10.1016/j.indcrop.2016.07.006
Rewers M, Sadowski J, Sliwinska E (2009) Endoreduplication in cucumber (Cucumis sativus) seeds during development, after processing and storage, and during germination. Ann Appl Biol 155:431–438. https://doi.org/10.1111/j.1744-7348.2009.00362.x
Rewers M, Sliwinska E (2012) Endoreduplication intensity as a marker of seed developmental stage in the Fabaceae. Cytometry 81A:1067–1075. https://doi.org/10.1002/cyto.a.22202
Rewers M, Sliwinska E (2014) Endoreduplication in germinating embryo and young seedling is related to the type of seedling establishment but is not coupled with superoxide radical accumulation. J Exp Bot 65:4385–4396. https://doi.org/10.1093/jxb/eru210
Rewicz A, Rewers M, Jędrzejczyk I, Rewicz T, Kołodziejek J, Jakubska-Busse A (2018) Morphology and genome size of Epipactis helleborine (L.) Crantz (Orchidaceae) growing in anthropogenic and natural habitats. PeerJ 6:e5992. https://doi.org/10.7717/peerj.5992
Sirjaev GI (1928–1932) Generis Trigonella L. Revisio critica I. Spisy vydavane´ Prirodovedecka´ Fakultou Masarykovy University. Brno 102:1–57
Soltis DE, Soltis PS, Bennett MD, Leitch IJ (2003) Evolution of genome size in the angiosperms. Am J Bot 90(11):1596–1603. https://doi.org/10.3732/ajb.90.11.1596
Srivastava A, Singh Z, Verma V, Choedon T (2020) Potential health benefits of fenugreek with multiple pharmacological properties. In: Mishra N (ed) Ethnopharmacological Investigation of Indian spices, 1st edn. IGI Global, pp 137–153. https://doi.org/10.4018/978-1-7998-2524-1.ch011
Sushma N, Davasena T (2010) Aqueous extract of Trigonella foenum-graecum (fenugreek) prevents cypermethrin-induced hepatotoxicity and nephrotoxicity. Hum Exp Toxicol 29(4):311–319. https://doi.org/10.1177/0960327110361502
Syed QA, Rashid Z, Ahmad MH, Shukat R, Ishaq A, Muhammad N, Rahman HUU (2020) Nutritional and therapeutic properties of fenugreek (Trigonella foenum-graecum): a review. Int J Food Prop 23(1):1777–1791. https://doi.org/10.1080/10942912.2020.1825482
Thakur J, Dwivedi MD, Singh N, Uniyal PL, Goel S, Pandey AK (2021) Applicability of Start Codon Targeted (SCoT) and Inter Simple Sequence Repeat (ISSR) markers in assessing genetic diversity in Crepidium acuminatum (D. Don) Szlach. J Appl Res Med Aromat Plants 23:100310. https://doi.org/10.1016/j.jarmap.2021.100310
Tomar RS, Parakhia MV, Rathod MV, Thakkai JR, Golakiya BA (2014) A comparative of ISSR nad RAPD markers for studying genetic diversity in Trigonella foenum-graecum genotypes. Res J Biotechnol 9(10):89–95
Van de Peer Y, De Wachter Y (1994) TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Appl Biosci 10(5):569–570. https://doi.org/10.1093/bioinformatics/10.5.569
Yilmaz AY, Martin E, Ünal F, Akan H (2009) Karyological study on six Trigonella L. species (Leguminosae) in Turkey. Caryologia 62(2):89–94. https://doi.org/10.1080/00087114.2004.10589673
Acknowledgements
The authors are grateful to the GRIN-ARS-USDA gene bank for providing Trigonella accessions.
Author information
Authors and Affiliations
Contributions
MR: contribution in establishing the experimental design, material preparation, data analysis and writing of the manuscript; AL: performing the experiments and evaluation of FCM histograms; DO: contribution in writing of the manuscript; AN: material preparation and performing the experiments; IJ: contribution in establishing the experimental design, data analysis, supervising the performance of experiments, writing of the manuscript.
Corresponding author
Ethics declarations
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript. The authors have no relevant financial or non-financial interests to disclose.
Competing interests
All authors declare that they have no conflicts of interest.
Additional information
Communicated by: Izabela Pawłowicz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rewers, M., Lojko, A., Olszewska, D. et al. Diversity of genome size, endopolyploidy and SCoT markers in 20 Trigonella (Fabaceae) species. J Appl Genetics (2024). https://doi.org/10.1007/s13353-024-00886-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13353-024-00886-9